X-Inactivation Choice in Mice
|
|
|
- Hollie Perry
- 6 years ago
- Views:
Transcription
1 REFERENCES CONTENT ALERTS Differential Methylation of Xite and CTCF Sites in Tsix Mirrors the Pattern of X-Inactivation Choice in Mice Rebecca Maxfield Boumil, Yuya Ogawa, Bryan K. Sun, Khanh D. Huynh and Jeannie T. Lee Mol. Cell. Biol. 2006, 26(6):2109. DOI: /MCB Updated information and services can be found at: These include: This article cites 38 articles, 13 of which can be accessed free at: Receive: RSS Feeds, etocs, free alerts (when new articles cite this article), more» Downloaded from on February 27, 2014 by PENN STATE UNIV Information about commercial reprint orders: To subscribe to to another ASM Journal go to:
2 MOLECULAR AND CELLULAR BIOLOGY, Mar. 2006, p Vol. 26, No /06/$ doi: /mcb Copyright 2006, American Society for Microbiology. All Rights Reserved. Differential Methylation of Xite and CTCF Sites in Tsix Mirrors the Pattern of X-Inactivation Choice in Mice Rebecca Maxfield Boumil, Yuya Ogawa, Bryan K. Sun, Khanh D. Huynh, and Jeannie T. Lee* Howard Hughes Medical Institute, Department of Molecular Biology, Massachusetts General Hospital, and Department of Genetics, Harvard Medical School, Boston, Massachusetts Received 21 September 2005/Returned for modification 25 October 2005/Accepted 21 December 2005 During mammalian dosage compensation, one of two X-chromosomes in female cells is inactivated. The choice of which X is silenced can be imprinted or stochastic. Although genetic loci influencing the choice decision have been identified, the primary marks for imprinting and random selection remain undefined. Here, we examined the role of DNA methylation, a mechanism known to regulate imprinting in autosomal loci, and sought to determine whether differential methylation on the two Xs might predict their fates. To identify differentially methylated domains (DMDs) at the X-inactivation center, we used bisulfite sequencing and methylation-sensitive restriction enzyme analyses. We found DMDs in Tsix and Xite, two genes previously shown to influence choice. Interestingly, the DMDs in Tsix lie within CTCF binding sites. Allelic methylation differences occur in gametes and are erased in embryonic stem cells carrying two active Xs. Because the pattern of DNA methylation mirrors events of X-inactivation, we propose that differential methylation of DMDs in Tsix and Xite constitute a primary mark for epigenetic regulation. The discovery of DMDs in CTCF sites draws further parallels between X-inactivation and autosomal imprinting. In mammals, one of two X-chromosomes is inactivated in the developing female embryo to ensure that XX and XY individuals have equal sex chromosome dosage (22). The questions of how a chromosome-counting mechanism differentiates the XX and the XY cell and how one X-chromosome is selected for silencing remain two key epigenetic problems in the X-chromosome inactivation (XCI) field (reviewed in reference 2). With regard to X-chromosome selection, silencing can take place randomly or in an imprinted fashion. In mice, imprinted XCI silences the paternal X (X P ) through preimplantation development and persists in the extraembryonic lineages (11, 30). In the epiblast lineage that gives rise to the embryo proper, the X P is reactivated to enable random selection of one X-chromosome for silencing just after uterine implantation (24). Three loci associated with noncoding RNA have been implicated in random X-chromosome choice. Xist, the gene that initiates silencing along the X (5), has been associated with skewed XCI ratios when specific mutations are incurred (25, 27, 33). Xist s antisense counterpart, Tsix, blocks Xist RNA accumulation (19) and is required for random XCI choice. Tsix deletions lead to biased silencing of the mutated X (19, 36). Required sequences lie within its CpG-rich 5 terminus, a region that includes multiple binding sites for CTCF (6), a chromatin insulator known to regulate autosomal imprinting at the H19/Igf2 locus (40). XCI choice is also affected by Xite, a modulator of Tsix and a candidate locus for the Xce modifier (29). Xite deletions reduce Tsix expression in cis, thereby lowering the probability that the linked X will become the active X (Xa). Although Xist, Tsix, and Xite each play a part in random choice, the molecular underpinnings remain unknown. * Corresponding author. Mailing address: Department of Molecular Biology, Massachusetts General Hospital, Simches 6.624, 185 Cambridge St., Boston, MA Phone: (617) Fax: (617) lee@molbio.mgh.harvard.edu. Superficially, imprinted XCI appears different with regard to mechanism, since it leads to stereotyped inactivation of the X P. One recent work has proposed that imprinted XCI consists of two phases: an Xist-independent phase that initiates silencing in the paternal germ line and is then propagated into the zygote, followed by a presumptively Xist-dependent phase that takes over the task of maintaining X P silence in the early embryo (11, 12). While the Xist-independent phase is posited to find its basis in meiotic sex chromosome inactivation, the presumptively Xist-dependent mechanism is thought to require the more conventional mechanism of gametic imprinting, occurring specifically at the X-inactivation center (Xic). Such an Xic imprint would be acquired some time during gametogenesis. A priori, the gametic Xic mark could be of either parental origin. If maternal, the imprint would promote the activity of the X M (maternal X); if paternal, the imprint would promote silencing of the X P. Strong arguments have been made for a maternal imprint (8, 31). Nuclear transplantation experiments indicate that embryos carrying two X M resist silencing despite having two Xs, while those carrying two X P partially overcome imprinting to silence one X P. This suggests that the X M is rigidly imprinted, whereas the X P is loosely imprinted. While the X M imprint is acquired during the oocyte growth phase (38), the exact nature of the maternal and possible paternal imprints is uncertain. Recent evidence suggests an imprinting center at the 5 end of Tsix, since its deletion on the X M causes ectopic Xist expression in the placenta (15, 36). Mounting evidence reveals that random choice and Xic imprinting may actually have a common mechanism. Most notably, Tsix and Xist mutations that affect random XCI also affect imprinted XCI (15, 19, 25, 36). It has been postulated that random XCI descended directly from imprinted XCI by passing the reigns of Xic control from parent to zygote (9, 16, 23). Genomic imprinting may have in fact originated on the X- chromosome to deal with dosage compensation (16). This idea 2109
3 2110 BOUMIL ET AL. MOL. CELL. BIOL. suggests potential mechanistic parallels between XCI and genomic imprinting in general. At autosomally imprinted loci, it is known that DNA methyltransferases (14, 21) and differential methylation at CpG dinucleotides (reviewed in references 3 and 37) play a critical role in regulation. Furthermore, at the H19/Igf2 locus, differentially methylated domains (DMDs) occur within binding sites for CTCF (40), a chromatin insulator that also binds Tsix (6). Given these striking coincidences, we now sought to determine whether DNA methylation differences also underlie Xic imprinting and choice. This idea makes very clear predictions. If true, there would be obvious methylation differences between sperm and oocyte. In anticipation of dynamic changes in XCI status during early zygotic development, these differences should remain through preimplantation development (X P inactive), possibly persist in the placenta as well (X P remains inactive), and then be erased in epiblast-derived cells (two active Xs) and zygotically reestablished in somatic lineages (one randomly chosen inactive X). To examine epiblast-derived cells, we use a mouse embryonic stem (ES) cell model that faithfully carries out random XCI during cell differentiation in culture. Several CpG islands occur in the murine Xist-Tsix-Xite region. Analyses of the CpG cluster in the Xist promoter have produced variable results, with some studies demonstrating differential methylation (1, 28, 41) and others finding none (26; H. Sasaki, personal communication). Similarly, the DXPas34 cluster downstream of the Tsix start site was originally thought to display differential methylation (7) but was later refuted (34). Thus, the role of differential methylation in XCI has remained ambiguous. Here, we focus on Tsix and Xite, two loci that bias X-chromosome selection. We find strong DMDs whose patterns of methylation correlate with the ongoing events of XCI. Interestingly, the DMDs partially overlap with CTCF binding sites and further highlight close molecular parallels between XCI and autosomal imprinting. MATERIALS AND METHODS Collection and preparation of DNA from cells, embryos, trophoblast-derived stem cells (TS cells), and cell lines. Sperm DNA was collected from C57BL/6J male mice (10). Oocytes were collected from 3- to 4-week-old superovulated B6SJLF1/J female mice, washed three times in Tris-EDTA (TE), and pooled in 20 l of TE, and DNA was extracted (13). Fertilized eggs were recovered at 0.5 days postcoitus from the corresponding matings and cultured in vitro in KSOM (Specialty Media) to the appropriate stages. For allele-specific methylation analysis, embryos were derived from crosses between female Mus musculus castaneus (CAST/Ei) and male M. m. musculus (C57BL/6). A total of 25 to 30 morula-stage embryos were pooled for each methylation-sensitive restriction analysis (MSRA) experiment. Derivation and culture of TS cells were described previously (11). The ES lines 16.7, J1, male (CG7) and female (3F1) CpG and Xite ( L-C2) have been described (18, 19, 29). Southern blot analysis. A total of 20 g of genomic DNA was digested overnight with enzymes outside the region of interest and simultaneously digested with a methylation-sensitive enzyme in the region of interest. The DNA was then Southern blotted, and the relevant fragments detected by probes doubly labeled with [ - 32 P]dCTP and [ - 32 P]dATP. MSRA. A total of 500 ng of DNA (or DNA from the average of 200 to 500 oocytes) was doubly digested overnight with a methylation-insensitive enzyme that cuts outside of region to be amplified (PvuII or EcoRI) and with a methylation-sensitive enzyme of interest. DNA was amplified during 35 cycles of amplification at 95 C for 15 s, 56 C for 45 s, and 72 C for 60 s and analyzed on agarose gels. Real-time PCR was performed on a Bio-Rad icycler machine in triplicate in SYBR green buffer (Stratagene) or with a TaqMan sequence-specific probe. The template was amplified for 50 cycles of 95 C for 15 s, 56 C for 45 s, TABLE 1. Sequences of primers and TaqMan probes used in MSRA and bisulfite analysis Primer or probe Sequence (5 3 ) MRSA primers CC3-1A...GTACCTGCAAGCGCTACACA CC3-1B...CGGGAACGTGGCATGTATGT CC3-1C...AATGCCTGCGTAGTCCCGAA CC3-3A...GGCGGGCAAGGCAAAACAAT CC3-3B...GGAGCCTAAACCTGTCTGTC CC3-3C...GCTACCTGTGTCTCTGTATC CC3-3R...CGAATTTGACTGCATATGTAC CC3-3S (DMR1-for)...CCTCAAACTCGCAAAGCTCTG DMR1-rev...GTCTGCCTACTAACACAGGT CC3-3T...GAGAATTCATTTCCCTGCCCT NG-1...CGCACGCACATATCTGTACGCA NG-2...GCGTTTTACCTTTTTAGGACAGAG Apa-1...GCATTCCATGAGTAAATCCCAC Apa AGCCCACATCCCTGATGATC Apa-2...GAGACAACTGGAAATGGGA Apa GGTGTAGCTTAGTTTCATCAGCTTC Apa-3...CTCCTCAGCCTTCCAGATTT Apa CAATCCTTGAAGCCAGTGAAC Apa-4...AACGGTAGACTGGTTTTGACC Apa AGTTGCCATGGGGAGAGGGGA Apa-5...CGCTGGATATAGCCTGGATTA Apa CACCACACCAGGCCAAAGCC Apa-6...GTGCTCAGTTCTAAATGGAGC Apa TATCAGAGGGCCACACCATTAAC TaqMan probes 1B1C...(6FAM)-TGGCTCCAGGACCCAGCAGAC 3B3C...(VIC)-TAGCTCAAGAGGCTAGACCTCTC-(MGB) 3S3T...(VIC)-CTAGTAATTGCCTTCCTCTATGTCT-(TAMRA) Allele-specific PCR probes KHp4...TAACCACCTGTAAGGGACAG KHp388...GTCTTCTGATTGTAACATGCTG Bisulfite sequencing primers bcc3-1f...tatataaactatctttccaaatccctaac bcc3-1g...gtagaaagatttttaaaatgtttgttagt bcc3-3f...aaataaaacctaaacctatctatctctt bcc3-3g...tttattaagtttttattatttgggataag bcc3-3h...taactacatatatacataaaacctttaaataa bcc3-3i...agttttggaagttgagatttaaaattttggta bcc3-3l...agaaaaggaattatgagataggttaaggta bcc3-3n...cttctataaatctaaaaaattcatttccct bcc3-3o...gtttagaaaggagggaaattaataattagt bcc3-3u...gttttatttttgagataaggttttagtaaataat bcc3-3v...taaaaaatcaatatttatctacctactaacacaaa bng-3...tatattttagattagattggatttaaattt bng ctatactcaaactaaaaacttacaaactaa bng-4...atatataaacataaataataaaatcaaata bng ttagtttgtaagtttttagtttgagtatagat bapa-1...atttttagaagagtagttggtgtttttt bapa ttaaaaatttataacccacatccctaataatc bapa-2...caaataatacctaaaacctaataaaata bapa ttttggtattatgaattgagttagatagt bapa-3...taaatttttatgaatgtttattttttaatgt bapa-4...catctaattataattaacataataaaaata bapa gtttattggttttaaggattgaggagttgt bapa-5...ttattaaatttatttaaataaattttttaag bapa-6...atatcacacaattacatacaaacctcaaaa bapa gattttgtaatttggggaatgttatttttttg and 67 C for 60 s. The fluorescence data were collected during the extension phase. The fluorescence threshold for determination of C is the average standard deviation of the background The cycle number at the threshold is the C. Samples digested with methylation-sensitive enzymes will have C values which correspond to the undigested samples if methylated and C values that diverge from undigested samples if unmethylated. Application of the formula, C undigested C digested C, normalizes the C value of each sample to that of the undigested control (the cycle number at the threshold is the C.) In all MSRA assays, we interpreted differences of less than one cycle as essentially equal amplification (because this is within the standard error of the
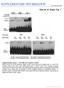
4 VOL. 26, 2006 DIFFERENTIAL METHYLATION OF Xite AND Tsix 2111 FIG. 1. Southern blot analysis of the Tsix region reveals potential DMDs. (A) Map of the Tsix and Xite regions. Oval region, CTCF motifs. DHS, DNase I-hypersensitive sites. Inset: H, HpaII; A, AciI. (B) Genomic Southern blot analysis of the 5 end of Tsix using the combination of enzymes indicated. Probe, 1.1-kb AgeI-HincII fragment at the 5 end of Tsix; Sp, sperm; M, male liver; F, female liver; filled circles, methylated bands; open circles, unmethylated bands. (C) Genomic Southern analysis of undifferentiated (day 0) ES cells and differentiated (day 12) EB cells using the 1.1-kb AgeI-HincII probe. Filled circles, methylated bands; open circles, unmethylated bands. Asterisks, doublet lower bands associated with the female ES and EB DNAs resulted from DXpas34 repeat length polymorphisms between the 129-derived and Mus castaneus-derived Xs in 16.7 ES cells. (D) Genomic Southern analysis of DXPas34 using the same probe. ES cells are female. The restriction sites (HpaII and AciI) are those indicated in the panel A inset. Asterisks indicate sites in ES cells that have become hypermethylated and have shifted upward and lost intensity or disappeared in EB cells. assay), suggesting that the template is methylated. Differences of between one and two cycles are interpreted as partially methylated or equivalent to 25 to 50% of template being methylated. Differences of greater than two cycles are unmethylated, indicating that 25% of template are methylated. Site A sequence was amplified with primer pair CC3-3B/CC3-3C; the real-time PCR analysis was done by using either the TaqMan probe 3B3C or SYBR green buffer. Site C sequence was amplified with the primer pair CC3-1C/CC3-1B; the real-time PCR analysis was done by using either the TaqMan probe 1B1C or SYBR green buffer. DHS6 was amplified with the primer pair NG-1/NG-2, DHS1 2 was amplified with the primer pairs Apa-5/Apa-5.1 and Apa-6/Apa-6.1, DHS3 was amplified with the primer pairs Apa-3/3.1 and Apa-4/4.1, and DHS4 was amplified with the primer pairs Apa-1/Apa-1.1 and Apa-2/Apa-2.1. Primers sequences are indicated in the Table 1. Allele-specific PCR analysis. Allele-specific PCR was done by digesting the MSRA conventional PCR amplified products with specified restriction enzymes that are sensitive to single nucleotide polymorphisms to distinguish between the parental alleles followed by Southern blot analysis. M. m. castaneus (CAST/Ei) DNA was used as the control maternal sample and M. m. musculus (C57BL/6) DNA was the control paternal sample for allele specific polymorphic digestion to compare with assayed DNAs. DHS6 was amplified with the primer pair NG-1/ NG-2 for allele-specific PCR and probed with KHp388. Site C was amplified with the primer pair CC3-1B/CC3-1C and probed with KHp4. All primer and probe sequences are shown in Table 1. Bisulfite sequencing. A total of 500 ng of genomic DNA, or DNA from an average of 500 oocytes, was treated with the CpGenome DNA modification kit (Intergen) as directed by the supplier, with the modification of additional denaturation steps at 95 C during the bisulfite incubation. PCR amplification was done on 2 l of treated sample DNA using primers devoid of CpG dinucleotides which would only amplify converted sequence, as is standard practice. Site A was amplified by two rounds of PCR using either primer pair bcc3-3f/bcc3-3l or primer pair bcc3-3h/bcc3-3l (with similar results). Site C was amplified by two PCR rounds with bcc3-1f/bcc3-1g. DHS6 was amplified by using bng-3/ bng-3.1 and bng-4/bng-4.1, DHS4 was amplified by using bapa-1/bapa-1.1 and bapa-2/bapa-2.1, and DHS3 was amplified by using bapa-4/4.1. Primer sequences are shown in the Table 1. Amplified PCR products were cloned into the pgem-t vector (Promega Corp.), sequenced by using an Applied Biosystems Taq DyeDeoxy Terminator cycle sequencing kit, and resolved on an ABI 3700 Prism automated sequencer. RESULTS Gamete-specific methylation differences in the CTCF binding sites of Tsix. We first focused on the 5 end of Tsix wherein lies a CpG island implicated in imprinting and random choice. This CpG island includes the Tsix promoter and CTCF binding sites in and around DXPas34 (34) (Fig. 1A). To search for DMDs, we performed MSRA, whereby the methylation status is assessed by whether genomic DNA is digestible (unmethylated) or not (methylated) with methylation sensitive enzymes. As a preliminary screen, we performed Southern analysis by digestion of gametic, ES cell, and somatic genomic DNAs with up to four different methylation-sensitive enzymes. Oocyte DNA is limiting so female gametes were not included in this
5 2112 BOUMIL ET AL. MOL. CELL. BIOL. FIG. 2. CTCF sites A and C of Tsix are differentially methylated in gametes. (A) MSRA at sites A and C using conventional PCR. (B) Representative MSRA of at site A using quantitative real-time PCR analysis. Each curve represents the amplification of a single sample. Horizontal orange lines, fluorescence threshold. HgaI-digested (red arrows) and undigested (black arrows) samples were run in triplicates. (C) Bisulfite analysis of site A. Filled circle, methylated; open circle, unmethylated. n, number of strands exhibiting each methylation pattern. initial analysis. DNA was digested with enzymes whose sites fall within the region of interest and with a flanking methylation-insensitive enzyme, PvuII (to reduce fragment sizes for electrophoresis). The results revealed a variegated pattern of DNA methylation across the region (Fig. 1B). No tissue-specific differences were observed at NarI and AgeI: NarI appeared uniformly methylated, and AgeI was undermethylated in the three tissues examined. In contrast, HgaI and SalI revealed potential differences: at HgaI, the sperm and male liver were hypermethylated, whereas female liver appeared to contain methylated and unmethylated epialleles. At SalI, sperm DNA was unmethylated, while the male and female liver DNA looked similar in being partially methylated. HgaI and SalI also exhibited differences in an ES model (Fig. 1C), with undifferentiated male and female ES cells being undermethylated and differentiated male and female EBs being hypermethylated. (Note that the doublet lower bands associated with the female ES and EB DNAs resulted from DXpas34 repeat length polymorphisms between the 129-derived and M. m. castaneus-derived Xs in 16.7 ES cells.) Intriguingly, SalI and HgaI coincide with elements previously shown to bind CTCF in vitro in a methylation-sensitive manner (sites A and B) (6). Furthermore, Southern analysis of DXPas34 suggested that AciI sites might be differentially methylated, since sites present in ES cells carrying two Xa become hypermethylated upon cell differentiation and initiation of XCI (asterisks, Fig. 1D; bands present in ES cells shifted upward and lost intensity or disappeared in EB). Because several CTCF motifs in Tsix have AciI sites and HgaI and SalI are within CTCF sites as well, these results led us to ask whether CTCF sites might constitute regulatory DMDs. To test this possibility, we compared the methylation status of CTCF sites of oocytes to that of sperm (Fig. 2). Because oocyte DNA was limiting, we modified the MSRA to include a PCR step after digestion with a methylation-sensitive enzyme. If the site is methylated, the DNA resists digestion and amplifies. If unmethylated, digestion would render the DNA unamplifiable. We designed primers to flank site A and another CTCF site previously designated site C (6) (Fig. 1A). Other sites, such as site B, could not be tested because they lie within the DXPas34 repeat, precluding design of locus-specific primers. At both sites A and C, PCR analysis showed that oocytes amplified poorly compared to the undigested control, whereas somatic cells and sperm amplified robustly (Fig. 2A). The results of conventional PCR were confirmed by quantitative realtime PCR analysis (Fig. 2B). Sperm DNA remained resistant to cutting at both sites. These findings demonstrated a gametespecific methylation difference in two assayable CTCF sites, with the maternal allele being undermethylated and the paternal allele being hypermethylated. To determine how many Cs are differentially methylated, we carried out bisulfite sequencing analysis, whereby genomic DNA is treated with bisulfite which converts C s to U s (read as T s) when the C is unmethylated. After PCR and sequencing with locus-specific primers, the methylation pattern on the original DNA strand can be inferred. Bisulfite analysis of site A revealed that the single CpG dinucleotide coinciding with the HgaI site is differentially methylated (Fig. 2C). Whereas
6 VOL. 26, 2006 DIFFERENTIAL METHYLATION OF Xite AND Tsix 2113 Downloaded from FIG. 3. Developmentally regulated methylation profiles at CTCF sites A and C. (A) Bisulfite analysis at site A. The methylation state of the single CpG is shown. (B) Bisulfite analysis at site C. CpG residues examined are numbered across; the AciI recognition site is noted. Filled circle, methylated; open circle, unmethylated; n, number of strands exhibiting each methylation pattern. (C) MSRA using real-time PCR of various cell lines and tissues. Filled circle, methylated; half-filled circle, partially methylated; open circle, unmethylated (see the text). ES cells are undifferentiated day 0 (d0) cultures; EB cells are differentiated cultures at days 7 to 12, a time when a vast majority of cells have undergone XCI (19). (D) Representative real-time PCR runs of the MSRA performed in triplicates. Black arrows, undigested samples; red arrows, digested samples. (E) Allele-specific MSRA at site C in ES and EB cells. Samples were digested ( ) or not digested ( ) with AciI before PCR amplification with primers CC3-1B and CC3-1C and probing with end-labeled primer, KHp4. on February 27, 2014 by PENN STATE UNIV sperm was consistently methylated, oocytes were undermethylated. Interestingly, several asymmetric C s within site A showed differential methylation (data not shown). Although CTCF binding can be modulated by non-cpg methylation in vitro (6), the significance of non-cpg methylation remains unclear, since it was predominantly observed in oocytes and not in other cell types (data not shown). Dynamic methylation changes during cell differentiation. We next examined methylation in undifferentiated female ES cells, which would have reactivated the X P and therefore would carry two active Xs, and differentiated EB derivatives, which randomly reinactivate one of two Xs. Bisulfite sequencing showed that site A was undermethylated in ES cells (Fig. 3A). The minority of undifferentiated clones showing methylation may be due to partial differentiation, as ES cells are prone to differentiate in culture. A similar result was observed at site C (Fig. 3B). MSRA PCR confirmed that ES cells were undermethylated at sites A and C (Fig. 3C and D). Differentiation of ES cells for 7 to 12 days into EB cells resulted in de novo DNA methylation at sites A and C. Bisulfite analysis showed various degrees of new methylation at seven CpGs at site C in males and females (Fig. 3A and B).
7 2114 BOUMIL ET AL. MOL. CELL. BIOL. Downloaded from FIG. 4. Differential methylation at the Xite locus correlates with events of XCI. (A) Map of Xite. Methylation-sensitive restriction sites are indicated above the axis; amplicons are indicated below the axis. Black rectangles, novel DMDs. (B) MSRA using real-time PCR at Xite. The deduced methylation status is shown; open circle, unmethylated; filled circles, methylated; half-filled circles, partial methylation. ES cells are day 0 (d0) cultures, EB cells are from days 7 to 12. ND, no data. (C) Bisulfite analysis at DHS6. CpG residues examined are numbered across; the enzyme recognition sites are noted. Filled circle, methylated; open circle, unmethylated; n, number of strands exhibiting each methylation pattern. (D) Allele-specific MSRA of ES and EB cells at DHS6. Enzyme indicates samples that were digested ( ) with either HpaII, AciI, or HgaI or undigested ( ) before PCR amplification with primers NG-1 and NG-2 and then probing with end-labeled primer KHp388. (E) Allele-specific MSRA of morula and female TS cells at DHS6 as described in panel D. For the TS cells, the maternal allele is of 129 origin, the paternal allele of castaneus origin. For morulae, the maternal allele is of castaneus origin, and the paternal allele is of 129 origin. on February 27, 2014 by PENN STATE UNIV These results were supported by MSRA (Fig. 3C and D). To determine which allele was methylated, we carried out quantitative allele-specific PCR after digestion with methylation-sensitive restriction enzymes. The allele-specific PCR was based on a single-nucleotide polymorphism that creates an AluI restriction site on the castaneus chromosome (Fig. 3E). In the 16.7 female ES line, the 129 X is maternally derived and the castaneus Xis paternally derived. The analysis showed that female cells were
8 VOL. 26, 2006 DIFFERENTIAL METHYLATION OF Xite AND Tsix 2115 FIG. 5. Mutant analyses suggest that DNA methylation works genetically upstream to Tsix and Xite. (A) The Tsix CpG and Xite L positions are shown. Proposed DMDs are indicated as filled black rectangles. (B) Effects of Xite L on CTCF sites A and C methylation at Tsix. MSRA analysis using real-time PCR of wild-type and mutant female cells lines at sites A and C. (C) Effects of Tsix CpG on Xite DMDs. MSRA analysis using real-time PCR of wild-type and mutant cell lines at the Xite DMDs. The deduced methylation status is shown. Female mutants are heterozygous. Open circles, unmethylated; filled circles, methylated. Downloaded from undermethylated on both maternal and paternal alleles before the onset of XCI (undifferentiated ES cells). Cell differentiation and the onset of XCI then resulted in hypermethylation of either or both alleles. Thus, in epiblast-derived ES cells, methylation is lost from the previously methylated paternal allele (spermatic methylation), and this timing of demethylation correlates with the developmental stage when the XCI imprint is known to be erased. In males, the single X in undifferentiated cells was undermethylated, a finding consistent with its previously unmethylated state in the oocyte (Fig. 3E). We conclude that DNA methylation is therefore a reasonable candidate for the gametic XCI imprint. We then tested specific embryonic and somatic tissues. Liver DNA exhibited hypermethylation by MSRA (Fig. 3C) and full methylation of all strands tested by bisulfite sequencing (data not shown). Furthermore, MSRA and bisulfite analysis of female extraembryonic yolk sac endoderm at 7.5 days postcoitus showed that site A was hypermethylated and site C was partially methylated (Fig. 3C) and that, in female trophoblastderived stem cell line (i.e., TS cells), both parental alleles were also methylated (data not shown). In morulae, both alleles were slightly methylated (data not shown). These findings demonstrated that, in both tissues subject to imprinted XCI and those subject to random XCI, both Tsix alleles are at least partially methylated in differentiated cells. However, in extraembryonic tissues, DNA methylation does not correlate completely with the XCI status, a finding consistent with studies showing that epigenetic regulation in extra-embryonic tissues may depend less on DNA methylation and rely more on other chromatin modifications (20, 35, 39). Taken together, these data showed that DNA methylation at Tsix is differentially established in cells that carry the primary imprints for choice (oocyte, sperm) and lost in cells that have erased the imprint (ES cells). Remethylation occurs upon cell differentiation and becomes biallelic in somatic cells and also in TS cells. The latter result counters the prediction that the Xi would be uniquely methylated. However, it is known that Tsix expression is eventually turned off in cells that have established XCI. Therefore, it is possible that biallelic methylation in differentiated and TS cells is the result of two sequential events: a primary event on the future Xi and a secondary event on the established Xa after the loss of Tsix expression. The Xite locus contains three DMDs. We next examined the methylation status of Xite. Xite contains two clusters of CpGrich sites, coinciding with two intergenic transcription start sites and associated DNase I-hypersensitive sites, DHS1 to DHS6 (Fig. 4A). For MSRA, we designed primer pairs to span the DHS and compared the PCR efficiency of cut versus uncut (control) samples. Several DMDs were uncovered (Fig. 4B). Between DHS1 and DHS2, analysis of AgeI, HpaII, and AciI showed that oocytes were undermethylated, correlating with a single Xa in oocytes. In contrast, sperm was hypermethylated, a finding consistent with a paternal X marked as Xi. The AvaI site at DHS4 and the HpaII, AciI, and HgaI sites at DHS6 all demonstrated a similar pattern of differential methylation. Bisulfite sequencing analysis confirmed that the oocyte DNA is considerably undermethylated relative to sperm DNA (Fig. 4C). We then examined Xite methylation in ES and EB cells. Undifferentiated pre-xci ES cells were relatively undermethylated, a finding consistent with there being one Xa in male ES on February 27, 2014 by PENN STATE UNIV
9 2116 BOUMIL ET AL. MOL. CELL. BIOL. cells and two Xa in female ES cells (Fig. 4B). Differentiated post-xci EB cells became considerably hypermethylated, with methylation being most pronounced and biallelic in somatic cells (liver) (Fig. 4B and data not shown). Thus, like at the Tsix locus, a secondary methylation event of Xite may occur on both alleles following Tsix repression after XCI (29). To determine which ES and EB alleles were methylated, we carried out quantitative allele-specific PCR after digestion with various methylation-sensitive restriction enzymes (HpaII, AciI, and HgaI; Fig. 4D). This allele-specific PCR was based on a singlenucleotide polymorphism which creates a BstUI restriction site on the 129 X-chromosome. We found that female ES cells were vastly undermethylated on both the maternal and the paternal alleles before the onset of XCI. Cell differentiation and the onset of XCI then resulted in hypermethylation of either or both alleles. Thus, as was the case for Tsix, DNA methylation is lost from the previously methylated paternal Xite allele, and this demethylation occurred around the developmental stage when the XCI imprint is known to be erased. DNA methylation at Xite is therefore also a good candidate for the gametic XCI imprint. We also examined allelic methylation patterns in TS cells and morulae (Fig. 4E). In TS cells, the HpaII and AciI sites was biallelically methylated, whereas the HgaI site was undermethylated on both alleles. These results showed that DNA methylation does not correlate completely with the XCI status in this extraembryonic tissue, a finding consistent with studies showing that epigenetic regulation in extraembryonic tissues may depend less on DNA methylation and rely more on histone modifications (20, 35, 39). In pooled morulae, while HpaII and AciI appeared to be biallelically methylated, the HgaI site showed residual paternal-specific methylation, indicating that the methylation pattern in sperm is partially retained in the preimplantation embryo. The DMDs are not affected by mutations in Tsix and Xite. If DNA methylation were the primary mark for the imprinting and choice decision, it must act genetically upstream of Xite and Tsix. To test this possibility, we examined whether mutations in Tsix and Xite expression affected DNA methylation patterns by MSRA. When the Tsix CpG island, including the major antisense promoter, is deleted in ES cells (Tsix CpG ; Fig. 5A), the pattern of DNA methylation at the Xite locus did not deviate significantly from the wild type (Fig. 5C). Conversely, when a 12-kb region of Xite including both intergenic promoters was deleted in ES cells (Xite L ), Tsix methylation patterns were not altered (Fig. 5B). Bisulfite sequencing analysis also revealed no obvious methylation differences in either mutant cell line (data not shown). Thus, these results are consistent with the model that differential methylation acts genetically upstream of Tsix and Xite expression. DISCUSSION In this study, we have tested whether DNA methylation could serve as a primary mark for Xic imprinting and choice. The results of MRSA and bisulfite analysis revealed DMDs within both Tsix and Xite. Our observations are consistent with the hypothesis in several ways. First, changes in DNA methylation patterns tend to precede XCI and correlate with ongoing events: the marks are differentially imprinted in oocyte and sperm, and the marks are lost in epiblast-derived ES cells, where it is known that the gametic imprint is erased in full. De novo methylation occurs in differentiated cells, in day 7.5 yolk sac endoderm, and in TS cells, with methylation becoming biallelic in these tissues. Biallelic methylation is consistent with biallelic repression of Xite and Tsix in differentiated cells. Second, the DNA methylation marks are not affected by mutations in Tsix and Xite expression. Current understanding of XCI places Xite genetically upstream of Tsix (29), which in turn lies upstream of Xist (19). Therefore, if DNA methylation lay further upstream than all three loci, mutations in the downstream genes would not be expected to affect the DMDs. Finally, the DMDs are located within two genes Xite and Tsix known to be involved in the imprinting and/or choice decisions (15, 19, 29, 36). Whether Xite plays a role in imprinting XCI has not yet been examined. For random XCI, genetic analysis in ES cells suggests that Xite increases the likelihood of the Xa state by prolonging expression of the linked Tsix allele. In turn, Tsix designates the future Xa by blocking Xist RNA accumulation in cis. The observation that the Tsix DMD coincides with CTCF binding sites raises interesting parallels to autosomal imprinting. At H19/Igf2, the differential sensitivity of CTCF to DNA methylation sets up the pattern of parentspecific expression (40). At the Xic, CTCF has also been proposed as a candidate trans-acting factor for regulating allelic choice (6). By analogy to regulation at H19/Igf2, CTCF binding to Tsix could act as either a transcriptional activator for Tsix or a chromatin insulator that blocks Xist from upstream enhancers. In the case of transcriptional activation, DMD methylation could preclude binding of the CTCF activator to Tsix. If CTCF sets up a chromatin insulator, DMD methylation could nullify the insulator and allow upstream enhancers to engage Xist. Thus, gamete-specific methylation patterns in the CTCF sites of Tsix could provide the means for control of imprinted XCI. In the epiblast, demethylation could enable a zygotic mechanism to reset allelic choice in a stochastic fashion. Interestingly, these domains in Xite and Tsix have recently also been implicated in X-chromosome counting (17), so the potential role of DNA methylation in this aspect of XCI would be of future interest. The XCI phenotype seen in ES cells and mice carrying mutations in Dnmt1, the maintenance methyltransferase, is consistent with a role for DNA methylation in random XCI. Male Dnmt1 / ES cells and mice show inappropriate expression of Xist and ectopic Xi formation in some cells (4, 32). In extraembryonic cells, Dnmt1 mutations do not cause a major disruption to imprinted XCI (35). Therefore, if DNA methylation plays a primary role in these cells, other Dnmt genes may be responsible. Ultimately, necessary confirmation of DNA methylation as the primary mark will come with specific mutagenesis of the DMDs in Tsix and Xite. ACKNOWLEDGMENTS We thank M. E. Donohoe, D. E. Cohen, and R. K. Rowntree for critical reading of the manuscript and all members of the laboratory for discussion. R.M.B. was supported by an NRSA award (5F32-HD08541), K.D.H. was supported by an MGH Fund for Discovery, and B.K.S. was supported by the National Institutes of Health Medical Scientist Training Program. This study was funded by grants from the National Institutes
10 VOL. 26, 2006 DIFFERENTIAL METHYLATION OF Xite AND Tsix 2117 of Health (RO1-GM58839), the Howard Hughes Medical Institute, and the Pew Scholars Program to J.T.L. REFERENCES 1. Ariel, M., E. Robinson, J. R. McCarrey, and H. Cedar Gamete-specific methylation correlates with imprinting of the murine Xist gene. Nat. Genet. 9: Avner, P., and E. Heard X-chromosome inactivation: counting, choice, and initiation. Nat. Rev. Genet. 2: Bartolomei, M. S., and S. M. Tilghman Genomic imprinting in mammals. Annu. Rev. Genet. 81: Beard, C., E. Li, and R. Jaenisch Loss of methylation activates Xist in somatic but not in embryonic cells. Genes Dev. 9: Brown, C. J., B. D. Hendrich, J. L. Rupert, R. G. Lafreniere, Y. Xing, J. Lawrence, and H. F. Willard The human XIST gene: analysis of a 17-kb inactive X-specific RNA that contains conserved repeats and is highly localized within the nucleus. Cell 71: Chao, W., K. D. Huynh, R. J. Spencer, L. S. Davidow, and J. T. Lee CTCF, a candidate trans-acting factor for X-inactivation choice. Science 295: Courtier, B., E. Heard, and P. Avner Xce haplotypes show modified methylation in a region of the active X chromosome lying 3 to Xist. Proc. Natl. Acad. Sci. USA 92: Goto, Y., and N. Takagi Maternally inherited X chromosome is not inactivated in mouse blastocysts due to parental imprinting. Chromosome Res. 8: Graves, J. A. M Mammals that break the rules: genetics of marsupials and monotremes. Annu. Rev. Genet. 30: Hanel, M. L., and R. Wevrick Establishment and maintenance of DNA methylation patterns in mouse Ndn: implications for maintenance of imprinting in target genes of the imprinting center. Mol. Cell. Biol. 21: Huynh, K. D., and J. T. Lee Inheritance of a pre-inactivated paternal X-chromosome in early mouse embryos. Nature 426: Huynh, K. D., and J. T. Lee X-chromosome inactivation: linking ontogeny and phylogeny. Nat. Rev. Genet. 6: Kafri, T., M. Ariel, M. Brandeis, R. Shemer, L. Urven, J. McCarrey, H. Cedar, and A. Razin Developmental pattern of gene-specific DNA methylation in the mouse embryo and germ line. Genes Dev. 6: Kaneda, M., M. Okano, K. Hata, T. Sado, N. Tsujimoto, E. Li, and H. Sasaki Essential role for de novo DNA methyltransferase Dnmt3a in paternal and maternal imprinting. Nature 429: Lee, J. T Disruption of imprinted X inactivation by parent-of-origin effects at Tsix. Cell 103: Lee, J. T Molecular links between X-inactivation and autosomal imprinting: X-inactivation as a driving force for the evolution of imprinting. Curr. Biol. 13:R242 R Lee, J. T Regulation of X-chromosome counting by Tsix and Xite sequences. Science 309: Lee, J. T., L. S. Davidow, and D. Warshawsky Tsix, a gene antisense to Xist at the X-inactivation center. Nat. Genet. 21: Lee, J. T., and N. Lu Targeted mutagenesis of Tsix leads to nonrandom X-inactivation. Cell 99: Lewis, A., K. Mitsuya, D. Umlauf, P. Smith, W. Dean, J. Walter, M. J. Higgins, R. Feil, and W. Reik Imprinting on distal chromosome 7 in the placenta involves repressive histone methylation independent of DNA methylation. Nat. Genet. 36: Li, E., C. Beard, and R. Jaenisch Role for DNA methylation in genomic imprinting. Nature 366: Lyon, M. F Gene action in the X-chromosome of the mouse (Mus musculus L.). Nature 190: Lyon, M. F Imprinting and X chromosome inactivation, p In R. Ohlsson (ed.), Results and problems in cell differentiation. Springer-Verlag, Heidelberg, Germany. 24. Mak, W., T. B. Nesterova, M. de Napoles, R. Appanah, S. Yamanaka, A. P. Otte, and N. Brockdorff Reactivation of the paternal X chromosome in early mouse embryos. Science 303: Marahrens, Y., J. Loring, and R. Jaenisch Role of the Xist gene in X chromosome choosing. Cell 92: McDonald, L. E., C. A. Paterson, and G. F. Kay Bisulfite genomic sequencing-derived methylation profile of the Xist gene throughout early mouse development. Genomics 43: Nesterova, T. B., C. M. Johnston, R. Appanah, A. E. T. Newall, J. Godwin, M. Alexiou, and N. Brockdorff Skewing X chromosome choice by modulating sense transcription across the Xist locus. Genes Dev. 17: Norris, D. P., D. Patel, G. F. Kay, G. D. Penny, N. Brockdorff, S. A. Sheardown, and S. Rastan Evidence that random and imprinted Xist expression is controlled by preemptive methylation. Cell 77: Ogawa, Y., and J. T. Lee Xite, X-inactivation intergenic transcription elements that regulate the probability of choice. Mol. Cell 11: Okamoto, I., A. P. Otte, C. D. Allis, D. Reinberg, and E. Heard Epigenetic dynamics of imprinted X inactivation during early mouse development. Science 303: Okamoto, I., S. S. Tan, and N. Takagi X-chromosome inactivation in XX androgenetic mouse embryos surviving implantation. Development 127: Panning, B., and R. Jaenisch DNA hypomethylation can activate Xist expression and silence X-linked genes. Genes Dev. 10: Plenge, R. M., B. D. Hendrich, C. Schwartz, J. F. Arena, A. Naumova, C. Sapienza, R. M. Winter, and H. F. Willard A promoter mutation in the XIST gene in two unrelated families with skewed X-chromosome inactivation. Nat. Genet. 17: Prissette, M., O. El-Maarri, D. Arnaud, J. Walter, and P. Avner Methylation profiles of DXPas34 during the onset of X-inactivation. Hum. Mol. Genet. 10: Sado, T., M. H. Fenner, S. S. Tan, P. Tam, T. Shioda, and E. Li X inactivation in the mouse embryo deficient for Dnmt1: distinct effect of hypomethylation on imprinted and random X inactivation. Dev. Biol. 225: Sado, T., Z. Wang, H. Sasaki, and E. Li Regulation of imprinted X-chromosome inactivation in mice by Tsix. Development 128: Sleutels, F., and D. P. Barlow The origins of genomic imprinting in mammals, p In J. C. Dunlap and C.-T. Wu (ed.), Homology effects. Academic Press, Inc., San Diego, Calif. 38. Tada, T., Y. Obata, M. Tada, Y. Goto, N. Nakatsuji, S. S. Tan, T. Kono, and N. Takagi Imprint switching for non-random X-chromosome inactivation during mouse oocyte growth. Development 127: Umlauf, D., Y. Goto, R. Cao, F. Cerqueira, A. Wagschal, Y. Zhang, and R. Feil Imprinting along the Kcnq1 domain on mouse chromosome 7 involves repressive histone methylation and recruitment of Polycomb group complexes. Nat. Genet. 36: Wolffe, A Transcriptional control: imprinting insulation. Curr. Biol. 10:R463 R Zuccotti, M., and M. Monk Methylation of the mouse Xist gene in sperm and eggs correlates with imprinted Xist expression and paternal X- inactivation. Nat. Genet. 9:
Today. Genomic Imprinting & X-Inactivation
 Today 1. Quiz (~12 min) 2. Genomic imprinting in mammals 3. X-chromosome inactivation in mammals Note that readings on Dosage Compensation and Genomic Imprinting in Mammals are on our web site. Genomic
Today 1. Quiz (~12 min) 2. Genomic imprinting in mammals 3. X-chromosome inactivation in mammals Note that readings on Dosage Compensation and Genomic Imprinting in Mammals are on our web site. Genomic
Lecture 27. Epigenetic regulation of gene expression during development
 Lecture 27 Epigenetic regulation of gene expression during development Development of a multicellular organism is not only determined by the DNA sequence but also epigenetically through DNA methylation
Lecture 27 Epigenetic regulation of gene expression during development Development of a multicellular organism is not only determined by the DNA sequence but also epigenetically through DNA methylation
Transcriptional repression of Xi
 Transcriptional repression of Xi Xist Transcription of Xist Xist RNA Spreading of Xist Recruitment of repression factors. Stable repression Translocated Xic cannot efficiently silence autosome regions.
Transcriptional repression of Xi Xist Transcription of Xist Xist RNA Spreading of Xist Recruitment of repression factors. Stable repression Translocated Xic cannot efficiently silence autosome regions.
Genetics and Genomics in Medicine Chapter 6 Questions
 Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Joanna Hillman Michael Higgins Lab Oncology for Scientists I 10/29/2015
 Joanna Hillman Michael Higgins Lab Oncology for Scientists I 10/29/2015 ! Define Epigenetics & Genomic Imprinting! Discovery! What is the imprint! Lifecycle of an Imprint DMRs and ICEs! 2 main mechanisms
Joanna Hillman Michael Higgins Lab Oncology for Scientists I 10/29/2015 ! Define Epigenetics & Genomic Imprinting! Discovery! What is the imprint! Lifecycle of an Imprint DMRs and ICEs! 2 main mechanisms
Phenotypic Variation in a Genetically Identical Population of Mice
 MOLECULAR AND CELLULAR BIOLOGY, Sept. 1997, p. 5269 5274 Vol. 17, No. 9 0270-7306/97/$04.00 0 Copyright 1997, American Society for Microbiology Phenotypic Variation in a Genetically Identical Population
MOLECULAR AND CELLULAR BIOLOGY, Sept. 1997, p. 5269 5274 Vol. 17, No. 9 0270-7306/97/$04.00 0 Copyright 1997, American Society for Microbiology Phenotypic Variation in a Genetically Identical Population
Loss of DNMT1o Disrupts Imprinted X Chromosome Inactivation and Accentuates Placental Defects in Females
 Loss of DNMT1o Disrupts Imprinted X Chromosome Inactivation and Accentuates Placental Defects in Females Serge McGraw 1., Christopher C. Oakes 2., Josée Martel 1, M. Cecilia Cirio 3, Pauline de Zeeuw 1,
Loss of DNMT1o Disrupts Imprinted X Chromosome Inactivation and Accentuates Placental Defects in Females Serge McGraw 1., Christopher C. Oakes 2., Josée Martel 1, M. Cecilia Cirio 3, Pauline de Zeeuw 1,
X-Chromosome Kiss and Tell: How the Xs Go Their Separate Ways
 X-Chromosome Kiss and Tell: How the Xs Go Their Separate Ways M.C. ANGUERA, B.K. SUN, N. XU, AND J.T. LEE Howard Hughes Medical Institute, Department of Molecular Biology, Massachusetts General Hospital,
X-Chromosome Kiss and Tell: How the Xs Go Their Separate Ways M.C. ANGUERA, B.K. SUN, N. XU, AND J.T. LEE Howard Hughes Medical Institute, Department of Molecular Biology, Massachusetts General Hospital,
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3
 Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
DNA methylation & demethylation
 DNA methylation & demethylation Lars Schomacher (Group Christof Niehrs) What is Epigenetics? Epigenetics is the study of heritable changes in gene expression (active versus inactive genes) that do not
DNA methylation & demethylation Lars Schomacher (Group Christof Niehrs) What is Epigenetics? Epigenetics is the study of heritable changes in gene expression (active versus inactive genes) that do not
Methylation reprogramming dynamics and defects in gametogenesis and embryogenesis: implications for reproductive medicine
 Univ.-Prof. Dr. Thomas Haaf Methylation reprogramming dynamics and defects in gametogenesis and embryogenesis: implications for reproductive medicine Epigenetics and DNA methylation Heritable change of
Univ.-Prof. Dr. Thomas Haaf Methylation reprogramming dynamics and defects in gametogenesis and embryogenesis: implications for reproductive medicine Epigenetics and DNA methylation Heritable change of
Mammalian X-Chromosome Inactivation: An Epigenetics Paradigm
 Mammalian X-Chromosome Inactivation: An Epigenetics Paradigm E. HEARD, * J. CHAUMEIL, O. MASUI, AND I. OKAMOTO CNRS UMR218, Curie Institute, 75248 Paris Cedex 05, France In mammals, dosage compensation
Mammalian X-Chromosome Inactivation: An Epigenetics Paradigm E. HEARD, * J. CHAUMEIL, O. MASUI, AND I. OKAMOTO CNRS UMR218, Curie Institute, 75248 Paris Cedex 05, France In mammals, dosage compensation
Recent advances in X-chromosome inactivation Edith Heard
 Recent advances in X-chromosome inactivation Edith Heard X inactivation is the silencing one of the two X chromosomes in XX female mammals. Initiation of this process during early development is controlled
Recent advances in X-chromosome inactivation Edith Heard X inactivation is the silencing one of the two X chromosomes in XX female mammals. Initiation of this process during early development is controlled
Antagonism between DNA and H3K27 Methylation at the Imprinted Rasgrf1 Locus
 Antagonism between DNA and H3K27 Methylation at the Imprinted Rasgrf1 Locus Anders M. Lindroth 1., Yoon Jung Park 1., Chelsea M. McLean 1, Gregoriy A. Dokshin 1, Jenna M. Persson 1, Herry Herman 1,2, Diego
Antagonism between DNA and H3K27 Methylation at the Imprinted Rasgrf1 Locus Anders M. Lindroth 1., Yoon Jung Park 1., Chelsea M. McLean 1, Gregoriy A. Dokshin 1, Jenna M. Persson 1, Herry Herman 1,2, Diego
Epigenetics: A historical overview Dr. Robin Holliday
 Epigenetics 1 Rival hypotheses Epigenisis - The embryo is initially undifferentiated. As development proceeds, increasing levels of complexity emerge giving rise to the larval stage or to the adult organism.
Epigenetics 1 Rival hypotheses Epigenisis - The embryo is initially undifferentiated. As development proceeds, increasing levels of complexity emerge giving rise to the larval stage or to the adult organism.
OVERVIEW OF EPIGENETICS
 OVERVIEW OF EIENETICS Date: * Time: 9:00 am - 9:50 am * Room: Berryhill 103 Lecturer: Terry Magnuson 4312 MBRB trm4@med.unc.edu 843-6475 *lease consult the online schedule for this course for the definitive
OVERVIEW OF EIENETICS Date: * Time: 9:00 am - 9:50 am * Room: Berryhill 103 Lecturer: Terry Magnuson 4312 MBRB trm4@med.unc.edu 843-6475 *lease consult the online schedule for this course for the definitive
Analysis of sex differences in EGC imprinting
 Developmental Biology 268 (2004) 105 110 www.elsevier.com/locate/ydbio Analysis of sex differences in EGC imprinting Gabriela Durcova-Hills, a Paul Burgoyne, b and Anne McLaren a, * a The Wellcome Trust/Cancer
Developmental Biology 268 (2004) 105 110 www.elsevier.com/locate/ydbio Analysis of sex differences in EGC imprinting Gabriela Durcova-Hills, a Paul Burgoyne, b and Anne McLaren a, * a The Wellcome Trust/Cancer
Epigenetic Regulation of Health and Disease Nutritional and environmental effects on epigenetic regulation
 Epigenetic Regulation of Health and Disease Nutritional and environmental effects on epigenetic regulation Robert FEIL Director of Research CNRS & University of Montpellier, Montpellier, France. E-mail:
Epigenetic Regulation of Health and Disease Nutritional and environmental effects on epigenetic regulation Robert FEIL Director of Research CNRS & University of Montpellier, Montpellier, France. E-mail:
Epigenetics and Chromatin Remodeling
 Epigenetics and Chromatin Remodeling Bradford Coffee, PhD, FACMG Emory University Atlanta, GA Speaker Disclosure Information Grant/Research Support: none Salary/Consultant Fees: none Board/Committee/Advisory
Epigenetics and Chromatin Remodeling Bradford Coffee, PhD, FACMG Emory University Atlanta, GA Speaker Disclosure Information Grant/Research Support: none Salary/Consultant Fees: none Board/Committee/Advisory
Stem Cell Epigenetics
 Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development
 Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
Biology Developmental Biology Spring Quarter Midterm 1 Version A
 Biology 411 - Developmental Biology Spring Quarter 2013 Midterm 1 Version A 75 Total Points Open Book Choose 15 out the 20 questions to answer (5 pts each). Only the first 15 questions that are answered
Biology 411 - Developmental Biology Spring Quarter 2013 Midterm 1 Version A 75 Total Points Open Book Choose 15 out the 20 questions to answer (5 pts each). Only the first 15 questions that are answered
Repressive Transcription
 Repressive Transcription The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Guenther, M. G., and R. A.
Repressive Transcription The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Guenther, M. G., and R. A.
Fragile X Syndrome. Genetics, Epigenetics & the Role of Unprogrammed Events in the expression of a Phenotype
 Fragile X Syndrome Genetics, Epigenetics & the Role of Unprogrammed Events in the expression of a Phenotype A loss of function of the FMR-1 gene results in severe learning problems, intellectual disability
Fragile X Syndrome Genetics, Epigenetics & the Role of Unprogrammed Events in the expression of a Phenotype A loss of function of the FMR-1 gene results in severe learning problems, intellectual disability
Imprinting. Joyce Ohm Cancer Genetics and Genomics CGP-L2-319 x8821
 Imprinting Joyce Ohm Cancer Genetics and Genomics CGP-L2-319 x8821 Learning Objectives 1. To understand the basic concepts of genomic imprinting Genomic imprinting is an epigenetic phenomenon that causes
Imprinting Joyce Ohm Cancer Genetics and Genomics CGP-L2-319 x8821 Learning Objectives 1. To understand the basic concepts of genomic imprinting Genomic imprinting is an epigenetic phenomenon that causes
Xist and the order of silencing Karen Ng*, Dieter Pullirsch*, Martin Leeb* & Anton Wutz +
 review Xist and the order of silencing Karen Ng*, Dieter Pullirsch*, Martin Leeb* & Anton Wutz + Research Institute of Molecular Pathology, Vienna, Austria X inactivation is the mechanism by which mammals
review Xist and the order of silencing Karen Ng*, Dieter Pullirsch*, Martin Leeb* & Anton Wutz + Research Institute of Molecular Pathology, Vienna, Austria X inactivation is the mechanism by which mammals
Review Article Epigenetic Mechanisms of Genomic Imprinting: Common Themes in the Regulation of Imprinted Regions in Mammals, Plants, and Insects
 Genetics Research International Volume 2012, Article ID 585024, 17 pages doi:10.1155/2012/585024 Review Article Epigenetic echanisms of Genomic Imprinting: Common Themes in the Regulation of Imprinted
Genetics Research International Volume 2012, Article ID 585024, 17 pages doi:10.1155/2012/585024 Review Article Epigenetic echanisms of Genomic Imprinting: Common Themes in the Regulation of Imprinted
Recent Advances in X-Chromosome Inactivation
 MINI-REVIEW 1714 Journal of Recent Advances in X-Chromosome Inactivation SUNDEEP KALANTRY* Department of Human Genetics, University of Michigan Medical School, Ann Arbor, Michigan Cellular Physiology X-chromosome
MINI-REVIEW 1714 Journal of Recent Advances in X-Chromosome Inactivation SUNDEEP KALANTRY* Department of Human Genetics, University of Michigan Medical School, Ann Arbor, Michigan Cellular Physiology X-chromosome
BIOL2005 WORKSHEET 2008
 BIOL2005 WORKSHEET 2008 Answer all 6 questions in the space provided using additional sheets where necessary. Hand your completed answers in to the Biology office by 3 p.m. Friday 8th February. 1. Your
BIOL2005 WORKSHEET 2008 Answer all 6 questions in the space provided using additional sheets where necessary. Hand your completed answers in to the Biology office by 3 p.m. Friday 8th February. 1. Your
Epigenetics DNA methylation. Biosciences 741: Genomics Fall, 2013 Week 13. DNA Methylation
 Epigenetics DNA methylation Biosciences 741: Genomics Fall, 2013 Week 13 DNA Methylation Most methylated cytosines are found in the dinucleotide sequence CG, denoted mcpg. The restriction enzyme HpaII
Epigenetics DNA methylation Biosciences 741: Genomics Fall, 2013 Week 13 DNA Methylation Most methylated cytosines are found in the dinucleotide sequence CG, denoted mcpg. The restriction enzyme HpaII
Epigenetic modifications in an imprinting cluster are controlled by a hierarchy of DMRs suggesting long-range chromatin interactions
 Human Molecular Genetics, 2003, Vol. 12, No. 3 295 305 DOI: 10.1093/hmg/ddg022 Epigenetic modifications in an imprinting cluster are controlled by a hierarchy of DMRs suggesting long-range chromatin interactions
Human Molecular Genetics, 2003, Vol. 12, No. 3 295 305 DOI: 10.1093/hmg/ddg022 Epigenetic modifications in an imprinting cluster are controlled by a hierarchy of DMRs suggesting long-range chromatin interactions
Xist function: bridging chromatin and stem cells
 Review TRENDS in Genetics Vol.23 No.9 Xist function: bridging chromatin and stem cells Anton Wutz Research Institute of Molecular Pathology, Dr. Bohr-Gasse 7, 1030 Vienna, Austria In mammals, dosage compensation
Review TRENDS in Genetics Vol.23 No.9 Xist function: bridging chromatin and stem cells Anton Wutz Research Institute of Molecular Pathology, Dr. Bohr-Gasse 7, 1030 Vienna, Austria In mammals, dosage compensation
Nature Immunology: doi: /ni Supplementary Figure 1. DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells.
 Supplementary Figure 1 DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells. (a) Scheme for the retroviral shrna screen. (b) Histogram showing CD4 expression (MFI) in WT cytotoxic
Supplementary Figure 1 DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells. (a) Scheme for the retroviral shrna screen. (b) Histogram showing CD4 expression (MFI) in WT cytotoxic
A Continuity of X-Chromosome Silence from Gamete to Zygote
 A Continuity of X-Chromosome Silence from Gamete to Zygote K.D. HUYNH AND J.T. LEE Howard Hughes Medical Institute, Department of Molecular Biology, Massachusetts General Hospital, and Department of Genetics,
A Continuity of X-Chromosome Silence from Gamete to Zygote K.D. HUYNH AND J.T. LEE Howard Hughes Medical Institute, Department of Molecular Biology, Massachusetts General Hospital, and Department of Genetics,
GENDER James Bier
 GENDER 2005-2008 James Bier Objectives 1. State the method of determining gender in several genetic systems. 2. List the three regions of the Y chromosome. 3. Describe the events that promote sexual development
GENDER 2005-2008 James Bier Objectives 1. State the method of determining gender in several genetic systems. 2. List the three regions of the Y chromosome. 3. Describe the events that promote sexual development
Genetics and Genomics in Medicine Chapter 6. Questions & Answers
 Genetics and Genomics in Medicine Chapter 6 Multiple Choice Questions Questions & Answers Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the
Genetics and Genomics in Medicine Chapter 6 Multiple Choice Questions Questions & Answers Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the
Long Noncoding RNAs in Imprinting and X Chromosome Inactivation
 Biomolecules 2014, 4, 76-100; doi:10.3390/biom4010076 Review OPEN ACCESS biomolecules ISSN 2218-273X www.mdpi.com/journal/biomolecules/ Long Noncoding RNAs in Imprinting and X Chromosome Inactivation Joseph
Biomolecules 2014, 4, 76-100; doi:10.3390/biom4010076 Review OPEN ACCESS biomolecules ISSN 2218-273X www.mdpi.com/journal/biomolecules/ Long Noncoding RNAs in Imprinting and X Chromosome Inactivation Joseph
Epigenetics: Basic Principals and role in health and disease
 Epigenetics: Basic Principals and role in health and disease Cambridge Masterclass Workshop on Epigenetics in GI Health and Disease 3 rd September 2013 Matt Zilbauer Overview Basic principals of Epigenetics
Epigenetics: Basic Principals and role in health and disease Cambridge Masterclass Workshop on Epigenetics in GI Health and Disease 3 rd September 2013 Matt Zilbauer Overview Basic principals of Epigenetics
Biology 2C03 Term Test #3
 Biology 2C03 Term Test #3 Instructors: Dr. Kimberley Dej, Ray Procwat Date: Monday March 22, 2010 Time: 10:30 am to 11:20 am Instructions: 1) This midterm test consists of 9 pages. Please ensure that all
Biology 2C03 Term Test #3 Instructors: Dr. Kimberley Dej, Ray Procwat Date: Monday March 22, 2010 Time: 10:30 am to 11:20 am Instructions: 1) This midterm test consists of 9 pages. Please ensure that all
Bisphenol A Exposure Disrupts Genomic Imprinting in the Mouse
 Bisphenol A Exposure Disrupts Genomic Imprinting in the Mouse Martha Susiarjo 1,2, Isaac Sasson 3, Clementina Mesaros 4, Marisa S. Bartolomei 1,2 * 1 Department of Cell and Developmental Biology, University
Bisphenol A Exposure Disrupts Genomic Imprinting in the Mouse Martha Susiarjo 1,2, Isaac Sasson 3, Clementina Mesaros 4, Marisa S. Bartolomei 1,2 * 1 Department of Cell and Developmental Biology, University
The Chromosomal Basis of Inheritance
 The Chromosomal Basis of Inheritance Factors and Genes Mendel s model of inheritance was based on the idea of factors that were independently assorted and segregated into gametes We now know that these
The Chromosomal Basis of Inheritance Factors and Genes Mendel s model of inheritance was based on the idea of factors that were independently assorted and segregated into gametes We now know that these
Lessons from comparative analysis of X-chromosome inactivation in mammals
 Chromosome Research (2009) 17:659 669 DOI 10.1007/s10577-009-9057-7 Lessons from comparative analysis of X-chromosome inactivation in mammals Ikuhiro Okamoto & Edith Heard # Springer Science + Business
Chromosome Research (2009) 17:659 669 DOI 10.1007/s10577-009-9057-7 Lessons from comparative analysis of X-chromosome inactivation in mammals Ikuhiro Okamoto & Edith Heard # Springer Science + Business
REVIEWS. X chromosome regulation: diverse patterns in development, tissues and disease
 X chromosome regulation: diverse patterns in development, tissues and disease Xinxian Deng 1, Joel B. Berletch 1, Di K. Nguyen 1 and Christine M. Disteche 1,2 Abstract Genes on the mammalian X chromosome
X chromosome regulation: diverse patterns in development, tissues and disease Xinxian Deng 1, Joel B. Berletch 1, Di K. Nguyen 1 and Christine M. Disteche 1,2 Abstract Genes on the mammalian X chromosome
Chromosome-Wide Analysis of Parental Allele-Specific Chromatin and DNA Methylation
 MOLECULAR AND CELLULAR BIOLOGY, Apr. 2011, p. 1757 1770 Vol. 31, No. 8 0270-7306/11/$12.00 doi:10.1128/mcb.00961-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Chromosome-Wide
MOLECULAR AND CELLULAR BIOLOGY, Apr. 2011, p. 1757 1770 Vol. 31, No. 8 0270-7306/11/$12.00 doi:10.1128/mcb.00961-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Chromosome-Wide
Strategic delivery: Setting standards Increasing and. Details: Output: Demonstrating efficiency. informing choice.
 Strategic delivery: Setting standards Increasing and informing choice Demonstrating efficiency economy and value Details: Meeting Scientific and Clinical Advances Advisory Committee Agenda item 6 Paper
Strategic delivery: Setting standards Increasing and informing choice Demonstrating efficiency economy and value Details: Meeting Scientific and Clinical Advances Advisory Committee Agenda item 6 Paper
Chapter 15 Notes 15.1: Mendelian inheritance chromosome theory of inheritance wild type 15.2: Sex-linked genes
 Chapter 15 Notes The Chromosomal Basis of Inheritance Mendel s hereditary factors were genes, though this wasn t known at the time Now we know that genes are located on The location of a particular gene
Chapter 15 Notes The Chromosomal Basis of Inheritance Mendel s hereditary factors were genes, though this wasn t known at the time Now we know that genes are located on The location of a particular gene
Epigenetics. Lyle Armstrong. UJ Taylor & Francis Group. f'ci Garland Science NEW YORK AND LONDON
 ... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
An Unexpected Function of the Prader-Willi Syndrome Imprinting Center in Maternal Imprinting in Mice
 An Unexpected Function of the Prader-Willi Syndrome Imprinting Center in Maternal Imprinting in Mice Mei-Yi Wu 1 *, Ming Jiang 1, Xiaodong Zhai 2, Arthur L. Beaudet 2, Ray-Chang Wu 1 * 1 Department of
An Unexpected Function of the Prader-Willi Syndrome Imprinting Center in Maternal Imprinting in Mice Mei-Yi Wu 1 *, Ming Jiang 1, Xiaodong Zhai 2, Arthur L. Beaudet 2, Ray-Chang Wu 1 * 1 Department of
Long nonoding RNAs in the X-inactivation center
 Chromosome Res DOI 10.1007/s10577-013-9396-2 REVIEW Long nonoding RNAs in the X-inactivation center Emily Maclary & Michael Hinten & Clair Harris & Sundeep Kalantry # Springer Science+Business Media Dordrecht
Chromosome Res DOI 10.1007/s10577-013-9396-2 REVIEW Long nonoding RNAs in the X-inactivation center Emily Maclary & Michael Hinten & Clair Harris & Sundeep Kalantry # Springer Science+Business Media Dordrecht
Functional Characterization of a Testis-Specific DNA Binding Activity at the H19/Igf2 Imprinting Control Region
 MOLECULAR AND CELLULAR BIOLOGY, Nov. 2003, p. 8345 8351 Vol. 23, No. 22 0270-7306/03/$08.00 0 DOI: 10.1128/MCB.23.22.8345 8351.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
MOLECULAR AND CELLULAR BIOLOGY, Nov. 2003, p. 8345 8351 Vol. 23, No. 22 0270-7306/03/$08.00 0 DOI: 10.1128/MCB.23.22.8345 8351.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
Session 2: Biomarkers of epigenetic changes and their applicability to genetic toxicology
 Session 2: Biomarkers of epigenetic changes and their applicability to genetic toxicology Bhaskar Gollapudi, Ph.D The Dow Chemical Company Workshop: Genetic Toxicology: Opportunities to Integrate New Approaches
Session 2: Biomarkers of epigenetic changes and their applicability to genetic toxicology Bhaskar Gollapudi, Ph.D The Dow Chemical Company Workshop: Genetic Toxicology: Opportunities to Integrate New Approaches
Histones modifications and variants
 Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Although DNA methylation has been considered the primary
 Colloquium The insulation of genes from external enhancers and silencing chromatin Bonnie Burgess-Beusse, Catherine Farrell, Miklos Gaszner, Michael Litt, Vesco Mutskov, Felix Recillas-Targa, Melanie Simpson,
Colloquium The insulation of genes from external enhancers and silencing chromatin Bonnie Burgess-Beusse, Catherine Farrell, Miklos Gaszner, Michael Litt, Vesco Mutskov, Felix Recillas-Targa, Melanie Simpson,
Genome-editing via Oviductal Nucleic Acids Delivery (GONAD) system: a novel microinjection-independent genome engineering method in mice
 Supplementary Information Genome-editing via Oviductal Nucleic Acids Delivery (GONAD) system: a novel microinjection-independent genome engineering method in mice Gou Takahashi, Channabasavaiah B Gurumurthy,
Supplementary Information Genome-editing via Oviductal Nucleic Acids Delivery (GONAD) system: a novel microinjection-independent genome engineering method in mice Gou Takahashi, Channabasavaiah B Gurumurthy,
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Results. Abstract. Introduc4on. Conclusions. Methods. Funding
 . expression that plays a role in many cellular processes affecting a variety of traits. In this study DNA methylation was assessed in neuronal tissue from three pigs (frontal lobe) and one great tit (whole
. expression that plays a role in many cellular processes affecting a variety of traits. In this study DNA methylation was assessed in neuronal tissue from three pigs (frontal lobe) and one great tit (whole
The Chromosomal Basis Of Inheritance
 The Chromosomal Basis Of Inheritance Chapter 15 Objectives Explain the chromosomal theory of inheritance and its discovery. Explain why sex-linked diseases are more common in human males than females.
The Chromosomal Basis Of Inheritance Chapter 15 Objectives Explain the chromosomal theory of inheritance and its discovery. Explain why sex-linked diseases are more common in human males than females.
DNA Methylation and Cancer
 DNA Methylation and Cancer October 25, 2016 Dominic Smiraglia, Ph.D. Department of Cancer Genetics From Alan Wolffe, Science and Medicine, 1999 Vital Statistics Human genome contains 3 billion bp ~ 50,000
DNA Methylation and Cancer October 25, 2016 Dominic Smiraglia, Ph.D. Department of Cancer Genetics From Alan Wolffe, Science and Medicine, 1999 Vital Statistics Human genome contains 3 billion bp ~ 50,000
Chromosomes, Mapping, and the Meiosis-Inheritance Connection. Chapter 13
 Chromosomes, Mapping, and the Meiosis-Inheritance Connection Chapter 13 Chromosome Theory Chromosomal theory of inheritance - developed in 1902 by Walter Sutton - proposed that genes are present on chromosomes
Chromosomes, Mapping, and the Meiosis-Inheritance Connection Chapter 13 Chromosome Theory Chromosomal theory of inheritance - developed in 1902 by Walter Sutton - proposed that genes are present on chromosomes
Long noncoding RNA-mediated maintenance of DNA methylation and transcriptional gene silencing
 2792 Development 139, 2792-2803 (2012) doi:10.1242/dev.079566 2012. Published by The Company of Biologists Ltd Long noncoding RNA-mediated maintenance of DNA methylation and transcriptional gene silencing
2792 Development 139, 2792-2803 (2012) doi:10.1242/dev.079566 2012. Published by The Company of Biologists Ltd Long noncoding RNA-mediated maintenance of DNA methylation and transcriptional gene silencing
Altered imprinted gene methylation and expression in completely ES cellderived mouse fetuses: association with aberrant phenotypes
 Development 125, 2273-2282 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV4011 2273 Altered imprinted gene methylation and expression in completely ES cellderived mouse fetuses:
Development 125, 2273-2282 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV4011 2273 Altered imprinted gene methylation and expression in completely ES cellderived mouse fetuses:
Supplementary Materials and Methods
 Supplementary Materials and Methods Whole Mount X-Gal Staining Whole tissues were collected, rinsed with PBS and fixed with 4% PFA. Tissues were then rinsed in rinse buffer (100 mm Sodium Phosphate ph
Supplementary Materials and Methods Whole Mount X-Gal Staining Whole tissues were collected, rinsed with PBS and fixed with 4% PFA. Tissues were then rinsed in rinse buffer (100 mm Sodium Phosphate ph
Alpha thalassemia mental retardation X-linked. Acquired alpha-thalassemia myelodysplastic syndrome
 Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
High resolution melting for methylation analysis
 High resolution melting for methylation analysis Helen White, PhD Senior Scientist National Genetics Reference Lab (Wessex) Why analyse methylation? Genomic imprinting In diploid organisms somatic cells
High resolution melting for methylation analysis Helen White, PhD Senior Scientist National Genetics Reference Lab (Wessex) Why analyse methylation? Genomic imprinting In diploid organisms somatic cells
Parental Allele-Specific Chromatin Configuration in a Boundary Imprinting-Control Element Upstream of the Mouse H19 Gene
 MOLECULAR AND CELLULAR BIOLOGY, Apr. 1999, p. 2556 2566 Vol. 19, No. 4 0270-7306/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Parental Allele-Specific Chromatin Configuration
MOLECULAR AND CELLULAR BIOLOGY, Apr. 1999, p. 2556 2566 Vol. 19, No. 4 0270-7306/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Parental Allele-Specific Chromatin Configuration
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10495 WWW.NATURE.COM/NATURE 1 2 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 3 4 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 5 6 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 7 8 WWW.NATURE.COM/NATURE
doi:10.1038/nature10495 WWW.NATURE.COM/NATURE 1 2 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 3 4 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 5 6 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 7 8 WWW.NATURE.COM/NATURE
Imprinting disorders and assisted reproductive technology
 MODERN TRENDS Imprinting disorders and assisted reproductive technology Somjate Manipalviratn, M.D., Alan DeCherney, M.D., and James Segars, M.D. Reproductive Biology and Medicine Branch, Eunice Kennedy
MODERN TRENDS Imprinting disorders and assisted reproductive technology Somjate Manipalviratn, M.D., Alan DeCherney, M.D., and James Segars, M.D. Reproductive Biology and Medicine Branch, Eunice Kennedy
The Chromosomal Basis of Inheritance
 Chapter 15 The Chromosomal Basis of Inheritance PowerPoint Lectures for Biology, Seventh Edition Neil Campbell and Jane Reece Lectures by Chris Romero Overview: Locating Genes on Chromosomes A century
Chapter 15 The Chromosomal Basis of Inheritance PowerPoint Lectures for Biology, Seventh Edition Neil Campbell and Jane Reece Lectures by Chris Romero Overview: Locating Genes on Chromosomes A century
Methylation Dynamics in the Early Mammalian Embryo: Implications of Genome Reprogramming Defects for Development
 CTMI (2006) 310:13 22 c Springer-Verlag Berlin Heidelberg 2006 Methylation Dynamics in the Early Mammalian Embryo: Implications of Genome Reprogramming Defects for Development T. Haaf ( ) Johannes Gutenberg-Universität
CTMI (2006) 310:13 22 c Springer-Verlag Berlin Heidelberg 2006 Methylation Dynamics in the Early Mammalian Embryo: Implications of Genome Reprogramming Defects for Development T. Haaf ( ) Johannes Gutenberg-Universität
An epigenetic approach to understanding (and predicting?) environmental effects on gene expression
 www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
Problem Set 5 KEY
 2006 7.012 Problem Set 5 KEY ** Due before 5 PM on THURSDAY, November 9, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You are studying the development
2006 7.012 Problem Set 5 KEY ** Due before 5 PM on THURSDAY, November 9, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You are studying the development
BIOLOGY - CLUTCH CH.15 - CHROMOSOMAL THEORY OF INHERITANCE
 !! www.clutchprep.com Chromosomal theory of inheritance: chromosomes are the carriers of genetic material. Independent Assortment alleles for different characters sort independently of each other during
!! www.clutchprep.com Chromosomal theory of inheritance: chromosomes are the carriers of genetic material. Independent Assortment alleles for different characters sort independently of each other during
Analysis of DNA methylation acquisition at the imprinted Dlk1 locus reveals asymmetry at CpG dyads
 Gagne et al. Epigenetics & Chromatin 214, 7:9 RESEARCH Open Access Analysis of DNA methylation acquisition at the imprinted Dlk1 locus reveals asymmetry at CpG dyads Alyssa Gagne, Abigail Hochman, Mahvish
Gagne et al. Epigenetics & Chromatin 214, 7:9 RESEARCH Open Access Analysis of DNA methylation acquisition at the imprinted Dlk1 locus reveals asymmetry at CpG dyads Alyssa Gagne, Abigail Hochman, Mahvish
Human Genetics (Learning Objectives)
 Human Genetics (Learning Objectives) Recognize Mendel s contribution to the field of genetics. Review what you know about a karyotype: autosomes and sex chromosomes. Understand and define the terms: characteristic,
Human Genetics (Learning Objectives) Recognize Mendel s contribution to the field of genetics. Review what you know about a karyotype: autosomes and sex chromosomes. Understand and define the terms: characteristic,
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
PRADER WILLI/ANGELMAN
 SALSA MS-MLPA probemix ME028-B2 PRADER WILLI/ANGELMAN Lot B2-0811: As compared to version B1 (lot B1-0609, B1-1108), the 88 and 96 nt control fragments have been replaced (QDX2). PRADER-WILLI SYNDROME
SALSA MS-MLPA probemix ME028-B2 PRADER WILLI/ANGELMAN Lot B2-0811: As compared to version B1 (lot B1-0609, B1-1108), the 88 and 96 nt control fragments have been replaced (QDX2). PRADER-WILLI SYNDROME
Epigenetics Armstrong_Prelims.indd 1 04/11/2013 3:28 pm
 Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
I) Development: tissue differentiation and timing II) Whole Chromosome Regulation
 Epigenesis: Gene Regulation Epigenesis : Gene Regulation I) Development: tissue differentiation and timing II) Whole Chromosome Regulation (X chromosome inactivation or Lyonization) III) Regulation during
Epigenesis: Gene Regulation Epigenesis : Gene Regulation I) Development: tissue differentiation and timing II) Whole Chromosome Regulation (X chromosome inactivation or Lyonization) III) Regulation during
Parental Chromosome-specific Chromatin Conformation in the Imprinted U2af1-rs1 Gene in the Mouse*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 272, No. 33, Issue of August 15, pp. 20893 20900, 1997 1997 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Parental Chromosome-specific
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 272, No. 33, Issue of August 15, pp. 20893 20900, 1997 1997 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Parental Chromosome-specific
Chapter 11 How Genes Are Controlled
 Chapter 11 How Genes Are Controlled PowerPoint Lectures for Biology: Concepts & Connections, Sixth Edition Campbell, Reece, Taylor, Simon, and Dickey Copyright 2009 Pearson Education, Inc. Lecture by Mary
Chapter 11 How Genes Are Controlled PowerPoint Lectures for Biology: Concepts & Connections, Sixth Edition Campbell, Reece, Taylor, Simon, and Dickey Copyright 2009 Pearson Education, Inc. Lecture by Mary
Characterization of Novel Parent-Specific Epigenetic Modifications Upstream of the Imprinted Mouse H19 Gene
 MOLECULAR AND CELLULAR BIOLOGY, Nov. 1998, p. 6767 6776 Vol. 18, No. 11 0270-7306/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. Characterization of Novel Parent-Specific
MOLECULAR AND CELLULAR BIOLOGY, Nov. 1998, p. 6767 6776 Vol. 18, No. 11 0270-7306/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. Characterization of Novel Parent-Specific
The genetics of heterochromatin. in metazoa. mutations by means of X-ray irradiation" "for the discovery of the production of
 The genetics of heterochromatin in metazoa 1 Hermann Joseph Muller 1946 Nobel Prize in Medicine: "for the discovery of the production of mutations by means of X-ray irradiation" 3 4 The true meaning of
The genetics of heterochromatin in metazoa 1 Hermann Joseph Muller 1946 Nobel Prize in Medicine: "for the discovery of the production of mutations by means of X-ray irradiation" 3 4 The true meaning of
Histone macroh2a1 is concentrated in the inactive X chromosome of female preimplantation mouse embryos
 Development 127, 2283-2289 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV4325 2283 Histone macroh2a1 is concentrated in the inactive X chromosome of female preimplantation mouse
Development 127, 2283-2289 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV4325 2283 Histone macroh2a1 is concentrated in the inactive X chromosome of female preimplantation mouse
Genome - Wide Linkage Mapping
 Biological Sciences Initiative HHMI Genome - Wide Linkage Mapping Introduction This activity is based on the work of Dr. Christine Seidman et al that was published in Circulation, 1998, vol 97, pgs 2043-2048.
Biological Sciences Initiative HHMI Genome - Wide Linkage Mapping Introduction This activity is based on the work of Dr. Christine Seidman et al that was published in Circulation, 1998, vol 97, pgs 2043-2048.
Dominic J Smiraglia, PhD Department of Cancer Genetics. DNA methylation in prostate cancer
 Dominic J Smiraglia, PhD Department of Cancer Genetics DNA methylation in prostate cancer Overarching theme Epigenetic regulation allows the genome to be responsive to the environment Sets the tone for
Dominic J Smiraglia, PhD Department of Cancer Genetics DNA methylation in prostate cancer Overarching theme Epigenetic regulation allows the genome to be responsive to the environment Sets the tone for
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
DNA methylation: a potential clinical biomarker for the detection of human cancers
 DNA methylation: a potential clinical biomarker for the detection of human cancers Name: Tong Samuel Supervisor: Zigui CHEN Date: 1 st December 2016 Department: Microbiology Source: cited from Jakubowski,
DNA methylation: a potential clinical biomarker for the detection of human cancers Name: Tong Samuel Supervisor: Zigui CHEN Date: 1 st December 2016 Department: Microbiology Source: cited from Jakubowski,
Evaluation of the Epigenetic Regulation of Two X- linked, Autism Candidate Genes: X-linked Lymphocyte Regulated 3b and Transketolase-Like 1
 University of Connecticut OpenCommons@UConn Doctoral Dissertations University of Connecticut Graduate School 11-8-2018 Evaluation of the Epigenetic Regulation of Two X- linked, Autism Candidate Genes:
University of Connecticut OpenCommons@UConn Doctoral Dissertations University of Connecticut Graduate School 11-8-2018 Evaluation of the Epigenetic Regulation of Two X- linked, Autism Candidate Genes:
THE CHROMOSOMAL BASIS OF INHERITANCE CHAPTER 15
 THE CHROMOSOMAL BASIS OF INHERITANCE CHAPTER 15 What you must know: Inheritance in sex-linked genes. Inheritance of linked genes and chromosomal mapping. How alteration of chromosome number or structurally
THE CHROMOSOMAL BASIS OF INHERITANCE CHAPTER 15 What you must know: Inheritance in sex-linked genes. Inheritance of linked genes and chromosomal mapping. How alteration of chromosome number or structurally
SALSA MLPA probemix P185-C2 Intersex Lot C2-1015: As compared to the previous version C1 (lot C1-0611), the lengths of four probes have been adjusted.
 mix P185-C2 Intersex Lot C2-1015: As compared to the previous version C1 (lot C1-0611), the lengths of four s have been adjusted. The sex-determining region on chromosome Y (SRY) is the most important
mix P185-C2 Intersex Lot C2-1015: As compared to the previous version C1 (lot C1-0611), the lengths of four s have been adjusted. The sex-determining region on chromosome Y (SRY) is the most important
GCATCCATCTTGGGGCGTCCCAATTGCTGAGTAACAAATGAGACGC TGTGGCCAAACTCAGTCATAACTAATGACATTTCTAGACAAAGTGAC TTCAGATTTTCAAAGCGTACCCTGTTTACATCATTTTGCCAATTTCG
 Lecture 6 GCATCCATCTTGGGGCGTCCCAATTGCTGAGTAACAAATGAGACGC TGTGGCCAAACTCAGTCATAACTAATGACATTTCTAGACAAAGTGAC TTCAGATTTTCAAAGCGTACCCTGTTTACATCATTTTGCCAATTTCG CGTACTGCAACCGGCGGGCCACGCCCCCGTGAAAAGAAGGTTGTT TTCTCCACATTTCGGGGTTCTGGACGTTTCCCGGCTGCGGGGCGG
Lecture 6 GCATCCATCTTGGGGCGTCCCAATTGCTGAGTAACAAATGAGACGC TGTGGCCAAACTCAGTCATAACTAATGACATTTCTAGACAAAGTGAC TTCAGATTTTCAAAGCGTACCCTGTTTACATCATTTTGCCAATTTCG CGTACTGCAACCGGCGGGCCACGCCCCCGTGAAAAGAAGGTTGTT TTCTCCACATTTCGGGGTTCTGGACGTTTCCCGGCTGCGGGGCGG
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Epigenetic Inheritance
 (2) The role of Epigenetic Inheritance Lamarck Revisited Lamarck was incorrect in thinking that the inheritance of acquired characters is the main mechanism of evolution (Natural Selection more common)
(2) The role of Epigenetic Inheritance Lamarck Revisited Lamarck was incorrect in thinking that the inheritance of acquired characters is the main mechanism of evolution (Natural Selection more common)
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
MRC-Holland MLPA. Description version 29;
 SALSA MLPA KIT P003-B1 MLH1/MSH2 Lot 1209, 0109. As compared to the previous lots 0307 and 1006, one MLH1 probe (exon 19) and four MSH2 probes have been replaced. In addition, one extra MSH2 exon 1 probe,
SALSA MLPA KIT P003-B1 MLH1/MSH2 Lot 1209, 0109. As compared to the previous lots 0307 and 1006, one MLH1 probe (exon 19) and four MSH2 probes have been replaced. In addition, one extra MSH2 exon 1 probe,
SALSA MS-MLPA KIT ME011-A1 Mismatch Repair genes (MMR) Lot 0609, 0408, 0807, 0407
 SALSA MS-MLPA KIT ME011-A1 Mismatch Repair genes (MMR) Lot 0609, 0408, 0807, 0407 The Mismatch Repair (MMR) system is critical for the maintenance of genomic stability. MMR increases the fidelity of DNA
SALSA MS-MLPA KIT ME011-A1 Mismatch Repair genes (MMR) Lot 0609, 0408, 0807, 0407 The Mismatch Repair (MMR) system is critical for the maintenance of genomic stability. MMR increases the fidelity of DNA
Maintenance of Paternal Methylation and Repression of the Imprinted H19 Gene Requires MBD3
 Maintenance of Paternal Methylation and Repression of the Imprinted H19 Gene Requires MBD3 Kimberly J. Reese 1, Shu Lin 1, Raluca I. Verona 1, Richard M. Schultz 2, Marisa S. Bartolomei 1* 1 Department
Maintenance of Paternal Methylation and Repression of the Imprinted H19 Gene Requires MBD3 Kimberly J. Reese 1, Shu Lin 1, Raluca I. Verona 1, Richard M. Schultz 2, Marisa S. Bartolomei 1* 1 Department
The Biology and Genetics of Cells and Organisms The Biology of Cancer
 The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
Dissecting gene regulation network in human early embryos. at single-cell and single-base resolution
 Dissecting gene regulation network in human early embryos at single-cell and single-base resolution Fuchou Tang BIOPIC, College of Life Sciences Peking University 07/10/2015 Cockburn and Rossant, 2010
Dissecting gene regulation network in human early embryos at single-cell and single-base resolution Fuchou Tang BIOPIC, College of Life Sciences Peking University 07/10/2015 Cockburn and Rossant, 2010
