In brief, cells were harvested and proteins were cross-linked to DNA by adding
|
|
|
- Shawn Gilbert
- 5 years ago
- Views:
Transcription
1 Immunity, Volume 31 Supplemental Data The Transcriptional Repressor Bcl-6 Directs T Follicular Helper Cell Lineage Commitment Di Yu, Sudha Rao, Louis M. Tsai, Sau K. Lee, Yiqing He, Elissa L. Sutcliffe, Monika Srivastava, Michelle Linterman, Lei Zheng, Nicholas Simpson, Julia I. Ellyard, Ian A. Parish, Cindy Ma, Qi- Jing Li, Christopher R. Parish, Charles R. Mackay, and Carola G. Vinuesa SUPPLEMENTAL EXPERIMENTAL PROECEDURES Detailed ChIP Assay In brief, cells were harvested and proteins were cross-linked to DNA by adding formaldehyde to a final concentration of 1% for 1 min at room temperature followed by the addition of.25m glycine. Cells were washed three times with phosphatebuffered saline and resuspended in sodium dodecyl sulfate lysis buffer with the addition of complete EDTA-free protease inhibitor cocktail tablets (Roche Molecular Biochemicals). Formaldehyde-fixed cells were incubated on ice for 1 min and sonicated to shear chromatin by using an Utrasonic processor (Cole/Parmer) under optimized conditions that gave a range of DNA fragments from 1 to 5 bp, as determined by 2% agarose gel electrophoresis. Before the addition of antibody, samples were precleared with salmon sperm DNA-protein A-agarose for 3 min at 4 C with rotation. The soluble chromatin fraction was incubated overnight with rotation at 4 C with 5 µg of anti-bcl-6 antibody (SC-858, Santa Cruz) or without antibodies as a control. Immune complexes were collected with salmon sperm DNAprotein A-agarose, washed with 1 ml of low-salt buffer, high-salt buffer, LiCl wash buffer, and Tris-EDTA buffer (2x) and eluted with 4 µl of elution buffer with rotation at room temperature. Protein-DNA cross-links were reversed at 65 C overnight. Samples were recovered by phenol-chloroform extraction and ethanol precipitated. DNA pellets were washed in 7% ethanol, resuspended in 5 µl of Tris, ph 8., and 1
2 subsequently used for SYBR real-time PCR amplification (Applied Biosystems) (Figure S4). Antibody Used in Experiments For cell stimulation and differentiation: anti-cd3 (Clone 5A2, BD Pharmingen), anti-cd28 (Clone 37.51, BD Pharminggen), anti-ifn-γ (Clone XMG1.2, Biolegend), anti-il-4 (Clone 11B11, Biolegend), anti-il-12/il-23 p4 (Clone C17.8, Biolegend). For flow cytometry, antibodies were purchased from BD Pharmingen, ebiosciences and Biolegend except where otherwise indicated: Anti-mouse: CCR7 (Clone 4B7), CD25 (Clone Jo2), CD44 (Clone IM7), (Clone A2), (Clone 14), CD62L (Clone MEL-14), CXCR4 (Clone 2B11), CXCR5 (Clone 2G8), Fas (Clone PC61.5), GATA-3 (Clone L5-823), GL-7 (Clone GL7), ICOS (Clone 7E.17G9), IFN-γ (Clone XMG1.2), IL-17 (Clone TC11-18H1), IL-2 (Clone JES6-5H4), PD-1 (Clone J43), T-bet (Clone ebio4b1), Thy1.1 (Clone HIS51); Anti-human: CCR7 (Clone 1553, R&D Systems), CD45RA (Clone HI1), CD45RO (Clone UCHL1), CXCR4 (Clone 12G5), CXCR5 (Clone RF8B2), ICOS (Clone ISA-3), IFN-γ (Clone 4S.B3), IL- 17 (Clone ebio64dec17), IL-2 (Clone MQ1-17H12), IL-4 (Clone 8D4-8), PD-1 (Clone MIH4). For immunofluorescence: anti-mouse biotin (Clone 14, ebioscience) and biotinylated Peanut Agglutinin (PNA) (Vector Laboratories) conjugated to Alexa Fluor 555-Streptavidin (Invitrogen), CD4 FITC (Clone RM4-5, BD Bioscience) and IgD Alexa Fluor 647 (Clone 11-26c.2a, Biolegend). 2
3 SUPPLEMENTARY FIGURE LEGENDS Figure S1. GC and Tfh Profiles in Control Unimmunized Chimeric Mice. (A-B) Representative profiles showing GC cells (A) and Tfh cells (B) in unimmunized : (Group A) or : Bcl6 -/- (Group B) fetal liver chimeric mice. Figure S2. Development of Effector CD4 + T Cells from Bcl6 -/- T cells. (A) Flow cytometric profiles showing CD25 vs CD44 expression on CD4+ cells. in 5%:5% : (Group A) or : Bcl6 -/- (right) fetal liver chimeric mice one week after SRBC immunization. Gates identify effector (CD44 hi CD25 - ) CD4 + T cells and the percentages of cells in this gate are indicated. (B) Bar graphs showing the percentage of effector cells gated in (A) amongst and compartments from the same mice. (C) Ratio of CD4 + T effector cells between and compartments in each mouse. Each symbol represents one mouse. The lines are drawn through the median values in each group. P value was calculated by Student's t-test. Figure S3. Effects of Bcl-6 Deficiency on IL-17 and IFN-γ ssecretion by Mouse CD4 + cells (A, B) IFNγ (A) and IL-17 (B) production by total CD4 + T cells from : (Group A) or : Bcl6 -/- (Group B) fetal liver chimeras 8 days after SRBC immunization. Left panel shows representative flow cytometric profiles of IFN-γ vs CD44 expression and gates used to enumerate cytokine-producing cells quantified in the bar graphs (middle panel). The right panel shows ratios calculated by dividing the percentage of cytokine-secreting cells among CD4 + T cells ( or Bcl6 -/- ) by the percentage of cytokine-secreting cells amongst CD4 + T cells ( ) in each individual mouse. 3
4 (C, D) IFN-γ (C) and IL-17 (D) production by CD44 hi CD25 - CD62 lo effector cells sorted from the same mice as in (A-B). The ratios shown are calculated as in (A-B). (E, F) IFN-γ (E) and IL-17 (F) production by naïve CD44 lo CD25 - CD62L hi CD4 + cells sorted from the same mice as the panels above and differentiated for 3 days under Th1 (E) or Th17 (F) conditions. The ratios shown are calculated as in (A-B). In A-F, each symbol represents one mouse and the lines are drawn through the median values in each group. P values were calculated by Student's t-test. Figure S4. FACS Sorting Strategies (A) Strategy used to sort naïve CD4 + T cells (CD4 + B22-7AAD - CD44 lo CD62L hi CD25 - ) and activated CD4 + T cells (CD4 + B22-7AAD - CD44 hi CD62L low CD25 - ) from mouse spleens and lymph nodes. Percentages indicate the purity of populations. (B) Strategy used to sort human tonsil naïve CD4+ T cells (CD4 + 7AAD - CD45RO - CXCR5 lo ); Tfh cells (CD4 + 7AAD - CD45RO + CXCR5 hi PD-1 hi ); non-tfh effector cells (CD4 + 7AAD - CD45RO + CXCR5 int/lo PD-1 int/lo ); and a B cell-enriched population (CD4-7AAD - CXCR5 hi ). (C) Strategy to indentify Tfh cells and non-tfh effectors in mice. Figure S5. Bcl-6 Deficiency Does Not Influence Treg Development. (A) Representative flow cytometric profiles showing CD25 vs FoxP3 gated on CD4 + cells that are either Bcl-6 +/+ or Bcl-6 -/- from the fetal liver chimeras in the previous figure. (B) Numbers indicate the percentage of T regulatory cells (Tregs, FoxP3 + CD4 + ) amongst total CD4 + cells of the indicated genotypes. Each symbol represents one mouse and the lines are drawn through the mean values in each group. Figure S6. Increased Bcl-6 Expression Suppresses Th1 and Th17 cytokine secretion. 4
5 Intracellular IFN-γ and IL-17 production measured by flow cytometric staining of naïve CD4 + T cells sorted from C57BL/6 mice, stimulated with plate-bound anti-cd3ε and CD28 under Th1 or Th17 differentiation conditions in the presence of IL-2 for 2 days, followed by retroviral transduction with a Bcl-6 or empty vector control. After transduction, cells were cultured under Th1 or Th17 differentiation conditions for the indicated times before cytokine production was measured. For analysis, transduced cells were divided into three groups -, Intermediate (int), High (hi) - according to levels of GFP (Bcl-6) transduction as shown in Figure 3C and 3D. Figure S7 Bcl-6 Expression Does Not Suppress IL-2 Secretion. IL-2 production by naïve CD4 + T cells transduced with either a Bcl-6-expressing retrovirus or a vector control and stimulated under Th17 differentiation conditions for the times indicated. Transduced cells were divided into three groups: -, Intermediate (int), High (hi), according to the levels of GFP (Bcl-6). Day 5 representative FACS plots for each of the three groups are shown. Data shown represent mean values ± S.D., with n = 3. Figure S8. Location and Sequences of the Primer Sets Used for ChIP Analysis Figure S9 Bcl-6 Overexpression Suppresses GATA-3 Expression (A) Intracellular GATA-3 expression in sorted mouse naïve CD4 + T cells transduced with either a Bcl-6-expressing retrovirus or a vector control and cultured in Th2 differentiation conditions for 2 days (left panels). Bar-graphs show Geometric Mean Fluorescent Intensity (Geom MFI) and positive percentages in transduced (GFP + ) and non-transduced (GFP - ) cells (right panels). Data shown represent mean values ± S.D., with n = 3. P values were calculated by Student's t-test. (B) Representative flow cytometric profiles showing CD4 vs CD44 expression and CXCR5 vs PD-1 from Bcl6+/+ : Bcl-6-/- mixed chimeric mice 5
6 (described in Fig. 1), used to gate T naive (T N ), Tfh and non-tfh effectors (left panel). The histograms (middle panel) show GATA-3 expression in Bcl-6 +/+ (top) and Bcl-6 -/- (bottom panel) T cells from the chimeric mice; numbers indicate mean fluorescence intensity in each population. The graphs (right panel) show GATA-3 MFI in Bcl-6 +/+ Tnaive, Tfh and non-tfh effector cells from 4 individual mice. Figure S1. Bcl-6 Promotes the Tfh Gene Expression Signature. Flow cytometric analysis of PD-1, CXCR5, CXCR4 and CCR7 expression on naïve CD4 + T cells sorted from C57BL/6 mice, stimulated with plate-bound anti-cd3ε and anti-cd28 in the presence of IL-2, transduced by either a Bcl-6-expressing retrovirus or a vector control and cultured under ThN, Th1 or Th2 differentiation conditions for 2 days. Red, green and blue cells represent cells expressing no, intermediate or high levels of GFP (Bcl-6) respectively, with Geom MFI values of PD-1 or positive percentages of chemokine receptors shown. Data are representative of two experiments. Figure S11. mirna Expression by Non-transduced Cells (A) Naïve CD4 + T cells were sorted from C57BL/6 mice, stimulated with plate-bound anti-cd3ε and CD28 in the presence of IL-2 and transduced with either a Bcl-6- expressing retrovirus or a vector control. After transduction, cells were cultured under Th conditions for 2 days. Cells expressing intermediate to high levels of GFP were sorted to >98% purity to compare mirna gene expression changes in cells transduced with Bcl-6 vs empty vector. These results are shown in Fig. 6. Untransduced cells expressing no GFP (GFP - ) were sorted as an additional control. (B) MiRNA expression in GFP - cells from (A). No mirnas expressed at levels >1 were up- or down-regulated by more than 2-fold. 6
7 Figure S12. Predicted Binding Sites within 3 UTRs of Tfh Signature Molecules for mirnas Downregulated upon forced Bcl-6 Expression Schematic representation of predicted mirna binding sites within the 3 untranslated region (3 UTR) of mouse CXCR5, CXCR4, PD-1 and IL-21 as predicted by miranda and Targetscan. Figure S13 mir-11 and mir-13 Are Downregulated in Cells Expressing Bcl-6 and in Tfh Cells. Relative mrna levels of mir-11 and mir13 in (A) naïve CD4 + T cells transduced with either a Bcl-6-expressing retrovirus or a vector control, cultured under Th conditions for 2 days and sorted for high GFP expression; (B) mouse naïve and Tfh cells. Figure S14 Overexpression of mir Does Not Repress CXCR4 Expression (A) Histograms showing CXCR4 fluorescence on mouse B22 + cells transduced with retrovirus expressing empty vector (filled histogram) or mir (empty histogram). (B) The bar graphs show CXCR4 geometric mean fluorescence intensity in the same cells as in (A). Data shown represent mean values ± S.D., with n = 3. 7
8 Figure S1 A GL-7 B CXCR5 B Bcl6 -/-.3.2 Fas CD Bcl6 -/ Group A Group B Group A Group B PD-1
9 Figure S2 A Group A Group B Bcl6 -/- CD % 14.17% 25.16% 9.35% CD25 CD44 B Teffectors (CD44 hi CD25 - ) in CD4 + (%) Bcl6 CD45. Group A Group B +/+ +/+ +/+ -/ C Ratio of Teffector cells P<.1 Bcl6 -/-
10 Figure S3 A B C IFN-γ IL-17 IFN-γ Bcl6 -/- CD CD Bcl6 -/ Ratio of IFN-γ + cells 1.1 Bcl6 -/- CD CD Group A Group B Group A Group B P <.1 IFN-γ + in CD4 + cells (%) IL-17 + in CD4 + cells (%) Bcl6 -/ Group A Group A D IL-17 Bcl6 -/- Bcl6 -/ Bcl6 -/ Group B Group B Ratio of IL-17 + cells Ratio of IFN-γ + cells Ratio of IL-17 + cells P =.3 Group A 1.. P =.5 Bcl6 -/ Bcl6 -/- Group B Group A Group B.5 P =.2 Bcl6 -/- E IFN-γ Bcl6 -/ Ratio of IFN-γ + cells P <.1 Bcl6 -/- F IL Bcl6 -/ Ratio of IL-17 + cells P <.1. Bcl6 -/-
11 Figure S4 A 7AAD - CD4 + B22-7AAD - CD4 + B22 - CD25 - Effector CD25 CD62L 97.1% 97.6% Naïve CD44 CD44 B 7AAD - 7AAD - CD4 + 7AAD - CD4 + CD45RO + Tfh 92.% CXCR5 B cells CXCR5 CXCR5 PD % Naïve non-tfh 93.4% 92.8% CD4 CD45RO PD-1 C Tnaive CD4 CXCR5 Tfh non-tfh CD44 PD1
12 Figure S5 A Bcl6+/+ Bcl6-/- + CD Foxp3 CD25 B Treg (Foxp3 + ) in CD4 + (%) Bcl6 -/-
13 Figure S6 INF-γ + cells (%) GFP Bcl-6 - GFP Bcl-6 int GFP Bcl-6 hi Vector Bcl Time (days) IL-17 + cells (%) GFP Bcl-6 nil Vector Bcl-6 GFP Bcl-6 int GFP Bcl-6 hi Time (days)
14 Figure S7 8 GFP Bcl-6 nil GFP Bcl-6 int GFP Bcl-6 hi Vector Bcl-6 IL-2 + cells (%) Time (days) IL-2 Vector Bcl-6 GFP Bcl-6
15 Figure S8 A Predicted Bcl-6 binding sites: Human Primer: TSS TSS Tbx21 Rorc B Gene Rorc Tbx21 Primer Sets F: GTA GAA GCA TCA GGG GGT CA R: GAG ATG GAG AGG GCA GGT TT F: CCT CTG AGT GCC AGG AGA AT R: CTT GCT TTC TGG GAG TGT CA
16 Figure S9 A Cell number GFP + cells positive GFP - cells positive GATA-3 GATA-3 (Geom MFI) GFP + cells GFP - cells ** Bcl-6 Vector Bcl-6 Vector GATA-3 + cells (%) 1 75 GFP + cells GFP - cells ** 5 25 Bcl-6 Vector Bcl-6 Vector B CXCR5 CD4 Tnaive CD44 Tfh non-tfh PD1 Bcl6 -/- Gata-3 Tnaive = 35 non-tfh= 74 Tfh= 94 Tnaive = 39 Non-Tfh=9 Gata-3 MFI Tnaive Non-Tfh Tfh
17 % 1.59% 2.27% 5.36% 1.99% 1.39% 1.81% 4.17% 2.2% 1.68% 3.19% 6.4% Vector Bcl-6 GFP 22.7% 24.6% 25.2% 24.7% 3.7% 59.1% 21.1% 24.9% 21.9% 21.2% 24.6% 23.2% 15.8% 16.8% 17.9% 27.6% 35.4% 68.9% CCR7 22.4% 25.5% 22.3% 19.1% 21.4% 24.% 32.5% 32.2% 53.1% 69.5% 91.7% 5.2% 51.7% 57.5% 14.3% 22.% 35.4% Vector 29.8% CXCR4 2.35% 1.69% 1.97% 2.36% 1.78% Bcl-6 2.% CXCR Vector PD-1 Figure S1 Bcl-6 Vector Bcl-6 Thn Th1 Th2
18 Figure S11 A GFP 98.7% Figure 6 FSC 99.9% B Bcl-6 (probe signal) Vector (probe signal)
19 Figure S12 Mouse CXCR5 3 UTR (length: 1446) mir-18a, 18b mir-93, 17/2a, 2b, 16a, 16b Mouse CXCR4 3 UTR (length: 642) mir-132, mir-212 mir-23a Mouse PD-1 3 UTR (length: 956) mir-17-3p let-7d, let-7f, let-7a, let-7i, let-7g Mouse IL-21 3 UTR (length: 2578) mir-27a, mir-27b mir-21 mir-25 mir-21 mir-3e mir-142-5p mir-142-5p Predicted mirna target site
20 Figure S13 A 2.5 mir-11 5 mir-13 Relative to U6 (x1-4 ) Relative to U6 (x1-2 ) Vector Bcl-6 Vector Bcl-6 B 12 mir-11 5 mir-13 Relative to U6 (x1-4 ) Relative to U6 (x1-2 ) Tnaïve Tfh Tnaïve Tfh
21 Figure S14 A GFP + cells Cell number GFP - cells CXCR4 B 25 GFP - cells GFP + cells GMFI (x1 3 ) Vector mir Vector mir-17-92
22 Table S1 Details of the putative Bcl-6 binding sites generated by MatInspector Gene (Human) Location of Bcl-6 binding sites From To Optimised Core Similarity Matrix Similarity Strand Sequence (capitals: core sequence) Stat (+) atatacctagaaaagca Stat (+) ggactcctagagtcttc (+) gactttctggagaaaag (+) ggagtcctggcaagagc RORγt (+) tgcatcctagagaagag (+) tcttaggtagaaatcac (-) gccttccgagatggccc T-bet (+) aatttccccgagggtcc (+) acactcccagaaagcaa (Mouse) Gene Location of Bcl-6 binding sites From To Optimised Core Similarity Matrix Similarity Strand Sequence (capitals: core sequence) Stat (-) gtattccttgatatgga Stat (+) cttttcctccaagggac RORgt (+) ctcttccttgaggttgc T-bet (+) ttcctcctagacagggc
23 Table S2 MicroRNAs downregulated by Bcl-6 (by 2-fold) Clustered mirnas Bcl-6_1 Bcl-6_2 Control_1 Control_2 Fold change (mirna) Fold change (cluster) mmu-mir mmu-mir-23b mmu-mir-27b mmu-mir mmu-mir mmu-mir mmu-mir-2b mmu-mir-16a mmu-mir-19b mmu-mir-18a mmu-mir mmu-mir-19a mmu-mir-2a mmu-mir-92a mmu-mir-17* mmu-mir-31a mmu-mir-13b mmu-mir-142-5p mmu-mir-142-3p mmu-mir-23a mmu-mir-27a mmu-mir mmu-let-7g mmu-let-7i mmu-mir mmu-mir mmu-mir-16b mmu-let-7f mmu-let-7d mmu-let-7a mmu-mir mmu-mir-15b mmu-mir-3c mmu-mir-3e mmu-mir-15a mmu-mir
24 Table S3 MicroRNAs downregulated by Bcl-6 (by less than 2-fold) and upregulated by Bcl-6 Fold change (mirna) Fold change (cluster) Clustered mirnas Bcl-6_1 Bcl-6_2 Control_1 Control_2 mmu-mir-487b mmu-mir-342-3p mmu-mir-3b mmu-mir-3d mmu-mir-338-5p mmu-mir mmu-mir mmu-mir mmu-mir-22-3p mmu-mir-26a mmu-mir mmu-mir mmu-mir mmu-mir mmu-mir mmu-mir-146a mmu-mir mmu-mir-29c mmu-mir-29b mmu-mir-466c-5p mmu-mir-669c mmu-mir-669a mmu-mir mmu-mir mmu-mir-29a mmu-mir-29b mmu-mir-574-5p mmu-mir mmu-mir-466f-3p
25 Table S4 Genomic location of mirnas downregulated by BCL-6 Coordinates Gene / Intergenic Protein encoded by overlapping transcript OTTMUSG33261 No protein product 14: [+] (non-coding exon). mir-18a 14: [+] mir-17-5p/17-3p mir-19b 14: [+] mir-19a 14: [+] mir-2a 14: [+] mir-92a 14: [+] mir-93 5: mir-25 5: mir- 5: [-] 16b mir : [+] mir : [+] mir-23a 8: [+] mir-27a 8: [+] let-7d 13: [-] let-7f_1 13: [-] let-7a_1 13: [-] OTTMUSG22672 (intronic) OTTMUSG6193 (intronic) Intergenic Intergenic let-7i 1: [-] Intergenic let-7g 9: [+] ENSMUSG2257 (intronic) mir-23b 13: [+] mir-27b 13: [+] mir-24 13: [+] OTTMUSG3849 (intronic) ENSMUSG56199 (intronic) ENSMUSG21458 (intronic) mir-21 11: [-] OTTMUSG125 3'UTR (exonic) mir-3e 4: [-] OTTMUSG8852 mir-3c 4: [-] (intronic) mir-3d 15: [-] mir-3b 15: [-] mir p Intergenic 11: [+] OTTMUSG1327 (intronic) mir-15b 3: [+] OTTMUSG24475 (intronic) mir-16 14: [-] Intergenic mir-15a 14: [-] mir-2b X: [-] Intergenic mir- 16a mir- 31a mir- 13b X: [-] OTTMUSG17339 (exonic) 11: [+] OTTMUSG1218 (intronic) 16: [-] ENSMUSG49916 (intronic) Mcm7 No protein product Wdr82 WU:AC AC I1Rik-21 Tmem4 Nfyc (nuclear transcription factor-y gamma) No protein product Smc4 Kis2 locus (nonprotein coding) RP23-352L N2Rik-22 Function of protein DNA replication Gene methylation Unknown Unknown Transcription activation (cmyc, p53 ) Organisation of mitotic chromosomes unknown unknown
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Supplementary Figure 1: Chemokine receptor expression profiles of CCR6 + and CCR6 - CD4 + IL-17A +/ex and Treg cells. Quantitative PCR analysis of chemokine receptor transcript abundance
SUPPLEMENTARY FIGURES Supplementary Figure 1: Chemokine receptor expression profiles of CCR6 + and CCR6 - CD4 + IL-17A +/ex and Treg cells. Quantitative PCR analysis of chemokine receptor transcript abundance
SUPPLEMENTARY INFORMATION. Supp. Fig. 1. Autoimmunity. Tolerance APC APC. T cell. T cell. doi: /nature06253 ICOS ICOS TCR CD28 TCR CD28
 Supp. Fig. 1 a APC b APC ICOS ICOS TCR CD28 mir P TCR CD28 P T cell Tolerance Roquin WT SG Icos mrna T cell Autoimmunity Roquin M199R SG Icos mrna www.nature.com/nature 1 Supp. Fig. 2 CD4 + CD44 low CD4
Supp. Fig. 1 a APC b APC ICOS ICOS TCR CD28 mir P TCR CD28 P T cell Tolerance Roquin WT SG Icos mrna T cell Autoimmunity Roquin M199R SG Icos mrna www.nature.com/nature 1 Supp. Fig. 2 CD4 + CD44 low CD4
Nature Immunology: doi: /ni Supplementary Figure 1. Huwe1 has high expression in HSCs and is necessary for quiescence.
 Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
Blocking antibodies and peptides. Rat anti-mouse PD-1 (29F.1A12, rat IgG2a, k), PD-
 Supplementary Methods Blocking antibodies and peptides. Rat anti-mouse PD-1 (29F.1A12, rat IgG2a, k), PD- L1 (10F.9G2, rat IgG2b, k), and PD-L2 (3.2, mouse IgG1) have been described (24). Anti-CTLA-4 (clone
Supplementary Methods Blocking antibodies and peptides. Rat anti-mouse PD-1 (29F.1A12, rat IgG2a, k), PD- L1 (10F.9G2, rat IgG2b, k), and PD-L2 (3.2, mouse IgG1) have been described (24). Anti-CTLA-4 (clone
Supplementary Figure 1. Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Nature Immunology: doi: /ni.
 Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supplementary Information:
 Supplementary Information: Follicular regulatory T cells with Bcl6 expression suppress germinal center reactions by Yeonseok Chung, Shinya Tanaka, Fuliang Chu, Roza Nurieva, Gustavo J. Martinez, Seema
Supplementary Information: Follicular regulatory T cells with Bcl6 expression suppress germinal center reactions by Yeonseok Chung, Shinya Tanaka, Fuliang Chu, Roza Nurieva, Gustavo J. Martinez, Seema
CD31 5'-AGA GAC GGT CTT GTC GCA GT-3' 5 ' -TAC TGG GCT TCG AGA GCA GT-3'
 Table S1. The primer sets used for real-time RT-PCR analysis. Gene Forward Reverse VEGF PDGFB TGF-β MCP-1 5'-GTT GCA GCA TGA ATC TGA GG-3' 5'-GGA GAC TCT TCG AGG AGC ACT T-3' 5'-GAA TCA GGC ATC GAG AGA
Table S1. The primer sets used for real-time RT-PCR analysis. Gene Forward Reverse VEGF PDGFB TGF-β MCP-1 5'-GTT GCA GCA TGA ATC TGA GG-3' 5'-GGA GAC TCT TCG AGG AGC ACT T-3' 5'-GAA TCA GGC ATC GAG AGA
Plasmids Western blot analysis and immunostaining Flow Cytometry Cell surface biotinylation RNA isolation and cdna synthesis
 Plasmids psuper-retro-s100a10 shrna1 was constructed by cloning the dsdna oligo 5 -GAT CCC CGT GGG CTT CCA GAG CTT CTT TCA AGA GAA GAA GCT CTG GAA GCC CAC TTT TTA-3 and 5 -AGC TTA AAA AGT GGG CTT CCA GAG
Plasmids psuper-retro-s100a10 shrna1 was constructed by cloning the dsdna oligo 5 -GAT CCC CGT GGG CTT CCA GAG CTT CTT TCA AGA GAA GAA GCT CTG GAA GCC CAC TTT TTA-3 and 5 -AGC TTA AAA AGT GGG CTT CCA GAG
Supplementary. presence of the. (c) mrna expression. Error. in naive or
 Figure 1. (a) Naive CD4 + T cells were activated in the presence of the indicated cytokines for 3 days. Enpp2 mrna expression was measured by qrt-pcrhr, infected with (b, c) Naive CD4 + T cells were activated
Figure 1. (a) Naive CD4 + T cells were activated in the presence of the indicated cytokines for 3 days. Enpp2 mrna expression was measured by qrt-pcrhr, infected with (b, c) Naive CD4 + T cells were activated
MATERIALS AND METHODS. Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All
 MATERIALS AND METHODS Antibodies (Abs), flow cytometry analysis and cell lines Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All other antibodies used
MATERIALS AND METHODS Antibodies (Abs), flow cytometry analysis and cell lines Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All other antibodies used
Supplemental Information. T Cells Enhance Autoimmunity by Restraining Regulatory T Cell Responses via an Interleukin-23-Dependent Mechanism
 Immunity, Volume 33 Supplemental Information T Cells Enhance Autoimmunity by Restraining Regulatory T Cell Responses via an Interleukin-23-Dependent Mechanism Franziska Petermann, Veit Rothhammer, Malte
Immunity, Volume 33 Supplemental Information T Cells Enhance Autoimmunity by Restraining Regulatory T Cell Responses via an Interleukin-23-Dependent Mechanism Franziska Petermann, Veit Rothhammer, Malte
CD44
 MR1-5-OP-RU CD24 CD24 CD44 MAIT cells 2.78 11.2 WT RORγt- GFP reporter 1 5 1 4 1 3 2.28 1 5 1 4 1 3 4.8 1.6 8.1 1 5 1 4 1 3 1 5 1 4 1 3 3.7 3.21 8.5 61.7 1 2 1 3 1 4 1 5 TCRβ 2 1 1 3 1 4 1 5 CD44 1 2 GFP
MR1-5-OP-RU CD24 CD24 CD44 MAIT cells 2.78 11.2 WT RORγt- GFP reporter 1 5 1 4 1 3 2.28 1 5 1 4 1 3 4.8 1.6 8.1 1 5 1 4 1 3 1 5 1 4 1 3 3.7 3.21 8.5 61.7 1 2 1 3 1 4 1 5 TCRβ 2 1 1 3 1 4 1 5 CD44 1 2 GFP
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Supplementary Table 3. 3 UTR primer sequences. Primer sequences used to amplify and clone the 3 UTR of each indicated gene are listed.
 Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice
 Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
and follicular helper T cells is Egr2-dependent. (a) Diagrammatic representation of the
 Supplementary Figure 1. LAG3 + Treg-mediated regulation of germinal center B cells and follicular helper T cells is Egr2-dependent. (a) Diagrammatic representation of the experimental protocol for the
Supplementary Figure 1. LAG3 + Treg-mediated regulation of germinal center B cells and follicular helper T cells is Egr2-dependent. (a) Diagrammatic representation of the experimental protocol for the
Nature Immunology: doi: /ni Supplementary Figure 1. DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells.
 Supplementary Figure 1 DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells. (a) Scheme for the retroviral shrna screen. (b) Histogram showing CD4 expression (MFI) in WT cytotoxic
Supplementary Figure 1 DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells. (a) Scheme for the retroviral shrna screen. (b) Histogram showing CD4 expression (MFI) in WT cytotoxic
c Tuj1(-) apoptotic live 1 DIV 2 DIV 1 DIV 2 DIV Tuj1(+) Tuj1/GFP/DAPI Tuj1 DAPI GFP
 Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Figure 1. Immune profiles of untreated and PD-1 blockade resistant EGFR and Kras mouse lung tumors (a) Total lung weight of untreated
 1 Supplementary Figure 1. Immune profiles of untreated and PD-1 blockade resistant EGFR and Kras mouse lung tumors (a) Total lung weight of untreated (U) EGFR TL mice (n=7), Kras mice (n=7), PD-1 blockade
1 Supplementary Figure 1. Immune profiles of untreated and PD-1 blockade resistant EGFR and Kras mouse lung tumors (a) Total lung weight of untreated (U) EGFR TL mice (n=7), Kras mice (n=7), PD-1 blockade
ECM1 controls T H 2 cell egress from lymph nodes through re-expression of S1P 1
 ZH, Li et al, page 1 ECM1 controls T H 2 cell egress from lymph nodes through re-expression of S1P 1 Zhenhu Li 1,4,Yuan Zhang 1,4, Zhiduo Liu 1, Xiaodong Wu 1, Yuhan Zheng 1, Zhiyun Tao 1, Kairui Mao 1,
ZH, Li et al, page 1 ECM1 controls T H 2 cell egress from lymph nodes through re-expression of S1P 1 Zhenhu Li 1,4,Yuan Zhang 1,4, Zhiduo Liu 1, Xiaodong Wu 1, Yuhan Zheng 1, Zhiyun Tao 1, Kairui Mao 1,
Supplemental Figure 1
 Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
Nature Immunology: doi: /ni Supplementary Figure 1. Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice.
 Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
Abbreviations: P- paraffin-embedded section; C, cryosection; Bio-SA, biotin-streptavidin-conjugated fluorescein amplification.
 Supplementary Table 1. Sequence of primers for real time PCR. Gene Forward primer Reverse primer S25 5 -GTG GTC CAC ACT ACT CTC TGA GTT TC-3 5 - GAC TTT CCG GCA TCC TTC TTC-3 Mafa cds 5 -CTT CAG CAA GGA
Supplementary Table 1. Sequence of primers for real time PCR. Gene Forward primer Reverse primer S25 5 -GTG GTC CAC ACT ACT CTC TGA GTT TC-3 5 - GAC TTT CCG GCA TCC TTC TTC-3 Mafa cds 5 -CTT CAG CAA GGA
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1: Cryopreservation alters CD62L expression by CD4 T cells. Freshly isolated (left) or cryopreserved PBMCs (right) were stained with the mix of antibodies described
Supplementary Figure 1 Supplementary Figure 1: Cryopreservation alters CD62L expression by CD4 T cells. Freshly isolated (left) or cryopreserved PBMCs (right) were stained with the mix of antibodies described
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
for six pairs of mice. (b) Representative FACS analysis of absolute number of T cells (CD4 + and
 SUPPLEMENTARY DATA Supplementary Figure 1: Peripheral lymphoid organs of SMAR1 -/- mice have an effector memory phenotype. (a) Lymphocytes collected from MLNs and Peyer s patches (PPs) of WT and SMAR1
SUPPLEMENTARY DATA Supplementary Figure 1: Peripheral lymphoid organs of SMAR1 -/- mice have an effector memory phenotype. (a) Lymphocytes collected from MLNs and Peyer s patches (PPs) of WT and SMAR1
The autoimmune disease-associated PTPN22 variant promotes calpain-mediated Lyp/Pep
 SUPPLEMENTARY INFORMATION The autoimmune disease-associated PTPN22 variant promotes calpain-mediated Lyp/Pep degradation associated with lymphocyte and dendritic cell hyperresponsiveness Jinyi Zhang, Naima
SUPPLEMENTARY INFORMATION The autoimmune disease-associated PTPN22 variant promotes calpain-mediated Lyp/Pep degradation associated with lymphocyte and dendritic cell hyperresponsiveness Jinyi Zhang, Naima
Effector T Cells and
 1 Effector T Cells and Cytokines Andrew Lichtman, MD PhD Brigham and Women's Hospital Harvard Medical School 2 Lecture outline Cytokines Subsets of CD4+ T cells: definitions, functions, development New
1 Effector T Cells and Cytokines Andrew Lichtman, MD PhD Brigham and Women's Hospital Harvard Medical School 2 Lecture outline Cytokines Subsets of CD4+ T cells: definitions, functions, development New
Supplementary Document
 Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Figure 1
 Supplementary Figure 1 3 3 3 1 1 Bregma -1.6mm 3 : Bregma Ref) Http://www.mbl.org/atlas165/atlas165_start.html Bregma -.18mm Supplementary Figure 1 Schematic representation of the utilized brain slice
Supplementary Figure 1 3 3 3 1 1 Bregma -1.6mm 3 : Bregma Ref) Http://www.mbl.org/atlas165/atlas165_start.html Bregma -.18mm Supplementary Figure 1 Schematic representation of the utilized brain slice
Supplementary Information
 Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression
 Supplementary Figure 1 Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression. Quantitative real-time PCR of indicated mrnas in DCs stimulated with TLR2-Dectin-1 agonist zymosan
Supplementary Figure 1 Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression. Quantitative real-time PCR of indicated mrnas in DCs stimulated with TLR2-Dectin-1 agonist zymosan
Eosinophils are required. for the maintenance of plasma cells in the bone marrow
 Eosinophils are required for the maintenance of plasma cells in the bone marrow Van Trung Chu, Anja Fröhlich, Gudrun Steinhauser, Tobias Scheel, Toralf Roch, Simon Fillatreau, James J. Lee, Max Löhning
Eosinophils are required for the maintenance of plasma cells in the bone marrow Van Trung Chu, Anja Fröhlich, Gudrun Steinhauser, Tobias Scheel, Toralf Roch, Simon Fillatreau, James J. Lee, Max Löhning
Supplementary Figure 1. mrna expression of chitinase and chitinase-like protein in splenic immune cells. Each splenic immune cell population was
 Supplementary Figure 1. mrna expression of chitinase and chitinase-like protein in splenic immune cells. Each splenic immune cell population was sorted by FACS. Surface markers for sorting were CD11c +
Supplementary Figure 1. mrna expression of chitinase and chitinase-like protein in splenic immune cells. Each splenic immune cell population was sorted by FACS. Surface markers for sorting were CD11c +
Supplementary Figure 1 Lymphocytes can be tracked for at least 4 weeks after
 Supplementary Figure 1 Lymphocytes can be tracked for at least 4 weeks after photoconversion by using H2B-Dendra2. 4-5 PPs of H2B-Dendra2 BM chimeras were photoconverted and analyzed 7 days (upper panel)
Supplementary Figure 1 Lymphocytes can be tracked for at least 4 weeks after photoconversion by using H2B-Dendra2. 4-5 PPs of H2B-Dendra2 BM chimeras were photoconverted and analyzed 7 days (upper panel)
Supplementary Figure 1. Ex vivo IFNγ production by Tregs. Nature Medicine doi: /nm % CD127. Empty SSC 98.79% CD25 CD45RA.
 SSC CD25 1.8% CD127 Empty 98.79% FSC CD45RA CD45RA Foxp3 %IFNγ + cells 4 3 2 1 + IL-12 P =.3 IFNγ p=.9 %IL-4+ cells 3 2 1 IL-4 P =.4 c %IL-1 + cells IFNγ 4 3 2 1 Control Foxp3 IL-1 P =.41.64 4.76 MS 2.96
SSC CD25 1.8% CD127 Empty 98.79% FSC CD45RA CD45RA Foxp3 %IFNγ + cells 4 3 2 1 + IL-12 P =.3 IFNγ p=.9 %IL-4+ cells 3 2 1 IL-4 P =.4 c %IL-1 + cells IFNγ 4 3 2 1 Control Foxp3 IL-1 P =.41.64 4.76 MS 2.96
a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53
 1 2 3 4 5 6 7 8 9 10 Supplementary Figure 1. Induction of p53 LOH by MADM. a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53 mouse revealed increased p53 KO/KO (green,
1 2 3 4 5 6 7 8 9 10 Supplementary Figure 1. Induction of p53 LOH by MADM. a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53 mouse revealed increased p53 KO/KO (green,
Supplementary Table; Supplementary Figures and legends S1-S21; Supplementary Materials and Methods
 Silva et al. PTEN posttranslational inactivation and hyperactivation of the PI3K/Akt pathway sustain primary T cell leukemia viability Supplementary Table; Supplementary Figures and legends S1-S21; Supplementary
Silva et al. PTEN posttranslational inactivation and hyperactivation of the PI3K/Akt pathway sustain primary T cell leukemia viability Supplementary Table; Supplementary Figures and legends S1-S21; Supplementary
Nature Immunology: doi: /ni Supplementary Figure 1. Id2 and Id3 define polyclonal T H 1 and T FH cell subsets.
 Supplementary Figure 1 Id2 and Id3 define polyclonal T H 1 and T FH cell subsets. Id2 YFP/+ (a) or Id3 GFP/+ (b) mice were analyzed 7 days after LCMV infection. T H 1 (SLAM + CXCR5 or CXCR5 PD-1 ), T FH
Supplementary Figure 1 Id2 and Id3 define polyclonal T H 1 and T FH cell subsets. Id2 YFP/+ (a) or Id3 GFP/+ (b) mice were analyzed 7 days after LCMV infection. T H 1 (SLAM + CXCR5 or CXCR5 PD-1 ), T FH
well for 2 h at rt. Each dot represents an individual mouse and bar is the mean ±
 Supplementary data: Control DC Blimp-1 ko DC 8 6 4 2-2 IL-1β p=.5 medium 8 6 4 2 IL-2 Medium p=.16 8 6 4 2 IL-6 medium p=.3 5 4 3 2 1-1 medium IL-1 n.s. 25 2 15 1 5 IL-12(p7) p=.15 5 IFNγ p=.65 4 3 2 1
Supplementary data: Control DC Blimp-1 ko DC 8 6 4 2-2 IL-1β p=.5 medium 8 6 4 2 IL-2 Medium p=.16 8 6 4 2 IL-6 medium p=.3 5 4 3 2 1-1 medium IL-1 n.s. 25 2 15 1 5 IL-12(p7) p=.15 5 IFNγ p=.65 4 3 2 1
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Sherman SI, Wirth LJ, Droz J-P, et al. Motesanib diphosphate
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Sherman SI, Wirth LJ, Droz J-P, et al. Motesanib diphosphate
Nature Immunology: doi: /ni.3836
 Supplementary Figure 1 Recombinant LIGHT-VTP induces pericyte contractility and endothelial cell activation. (a) Western blot showing purification steps for full length murine LIGHT-VTP (CGKRK) protein:
Supplementary Figure 1 Recombinant LIGHT-VTP induces pericyte contractility and endothelial cell activation. (a) Western blot showing purification steps for full length murine LIGHT-VTP (CGKRK) protein:
Supplementary Figure S1. Flow cytometric analysis of the expression of Thy1 in NH cells. Flow cytometric analysis of the expression of T1/ST2 and
 Supplementary Figure S1. Flow cytometric analysis of the expression of Thy1 in NH cells. Flow cytometric analysis of the expression of T1/ST2 and Thy1 in NH cells derived from the lungs of naïve mice.
Supplementary Figure S1. Flow cytometric analysis of the expression of Thy1 in NH cells. Flow cytometric analysis of the expression of T1/ST2 and Thy1 in NH cells derived from the lungs of naïve mice.
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1175194/dc1 Supporting Online Material for A Vital Role for Interleukin-21 in the Control of a Chronic Viral Infection John S. Yi, Ming Du, Allan J. Zajac* *To whom
www.sciencemag.org/cgi/content/full/1175194/dc1 Supporting Online Material for A Vital Role for Interleukin-21 in the Control of a Chronic Viral Infection John S. Yi, Ming Du, Allan J. Zajac* *To whom
SUPPLEMENTARY INFORMATION
 Supplemental Figure 1. Furin is efficiently deleted in CD4 + and CD8 + T cells. a, Western blot for furin and actin proteins in CD4cre-fur f/f and fur f/f Th1 cells. Wild-type and furin-deficient CD4 +
Supplemental Figure 1. Furin is efficiently deleted in CD4 + and CD8 + T cells. a, Western blot for furin and actin proteins in CD4cre-fur f/f and fur f/f Th1 cells. Wild-type and furin-deficient CD4 +
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk
 Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Figure S1. Analysis of genomic and cdna sequences of the targeted regions in WT-KI and
 Figure S1. Analysis of genomic and sequences of the targeted regions in and indicated mutant KI cells, with WT and corresponding mutant sequences underlined. (A) cells; (B) K21E-KI cells; (C) D33A-KI cells;
Figure S1. Analysis of genomic and sequences of the targeted regions in and indicated mutant KI cells, with WT and corresponding mutant sequences underlined. (A) cells; (B) K21E-KI cells; (C) D33A-KI cells;
T H 1, T H 2 and T H 17 polarization of naïve CD4 + mouse T cells
 A complete workflow for cell preparation, isolation, polarization and analysis T H 1, T H 2 and T H 17 polarization of naïve CD4 + mouse T cells Introduction Workflow CD4 + T helper (T H) cells play a
A complete workflow for cell preparation, isolation, polarization and analysis T H 1, T H 2 and T H 17 polarization of naïve CD4 + mouse T cells Introduction Workflow CD4 + T helper (T H) cells play a
Supplemental Figures: Supplemental Figure 1
 Supplemental Figures: Supplemental Figure 1 Suppl. Figure 1. BM-DC infection with H. pylori does not induce cytotoxicity and treatment of BM-DCs with H. pylori sonicate, but not heat-inactivated bacteria,
Supplemental Figures: Supplemental Figure 1 Suppl. Figure 1. BM-DC infection with H. pylori does not induce cytotoxicity and treatment of BM-DCs with H. pylori sonicate, but not heat-inactivated bacteria,
Supplementary Table e-1. Flow cytometry reagents and staining combinations
 Supplementary data Supplementary Table e-1. Flow cytometry reagents and staining combinations Reagents Antibody Fluorochrome Clone Source conjugation CD3 FITC UCHT1 BD Biosciences CD3 PerCP-Cy5.5 SK7 Biolegend
Supplementary data Supplementary Table e-1. Flow cytometry reagents and staining combinations Reagents Antibody Fluorochrome Clone Source conjugation CD3 FITC UCHT1 BD Biosciences CD3 PerCP-Cy5.5 SK7 Biolegend
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Supplementary Data Table of Contents:
 Supplementary Data Table of Contents: - Supplementary Methods - Supplementary Figures S1(A-B) - Supplementary Figures S2 (A-B) - Supplementary Figures S3 - Supplementary Figures S4(A-B) - Supplementary
Supplementary Data Table of Contents: - Supplementary Methods - Supplementary Figures S1(A-B) - Supplementary Figures S2 (A-B) - Supplementary Figures S3 - Supplementary Figures S4(A-B) - Supplementary
Supplementary Figure 1 Cytokine receptors on developing thymocytes that can potentially signal Runx3d expression.
 Supplementary Figure 1 Cytokine receptors on developing thymocytes that can potentially signal Runx3d expression. (a) Characterization of c-independent SP8 cells. Stainings for maturation markers (top)
Supplementary Figure 1 Cytokine receptors on developing thymocytes that can potentially signal Runx3d expression. (a) Characterization of c-independent SP8 cells. Stainings for maturation markers (top)
Nature Immunology: doi: /ni Supplementary Figure 1. Transcriptional program of the TE and MP CD8 + T cell subsets.
 Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
<10. IL-1β IL-6 TNF + _ TGF-β + IL-23
 3 ns 25 ns 2 IL-17 (pg/ml) 15 1 ns ns 5 IL-1β IL-6 TNF
3 ns 25 ns 2 IL-17 (pg/ml) 15 1 ns ns 5 IL-1β IL-6 TNF
ILC1 and ILC3 isolation and culture Following cell sorting, we confirmed that the recovered cells belonged to the ILC1, ILC2 and
 Supplementary Methods and isolation and culture Following cell sorting, we confirmed that the recovered cells belonged to the, ILC2 and subsets. For this purpose we performed intracellular flow cytometry
Supplementary Methods and isolation and culture Following cell sorting, we confirmed that the recovered cells belonged to the, ILC2 and subsets. For this purpose we performed intracellular flow cytometry
Kerdiles et al - Figure S1
 Kerdiles et al - Figure S1 a b Homo sapiens T B ce ce l ls c l M ls ac r PM oph N ag es Mus musculus Foxo1 PLCγ Supplementary Figure 1 Foxo1 expression pattern is conserved between mouse and human. (a)
Kerdiles et al - Figure S1 a b Homo sapiens T B ce ce l ls c l M ls ac r PM oph N ag es Mus musculus Foxo1 PLCγ Supplementary Figure 1 Foxo1 expression pattern is conserved between mouse and human. (a)
Supplementary Figure S1. PTPN2 levels are not altered in proliferating CD8+ T cells. Lymph node (LN) CD8+ T cells from C57BL/6 mice were stained with
 Supplementary Figure S1. PTPN2 levels are not altered in proliferating CD8+ T cells. Lymph node (LN) CD8+ T cells from C57BL/6 mice were stained with CFSE and stimulated with plate-bound α-cd3ε (10µg/ml)
Supplementary Figure S1. PTPN2 levels are not altered in proliferating CD8+ T cells. Lymph node (LN) CD8+ T cells from C57BL/6 mice were stained with CFSE and stimulated with plate-bound α-cd3ε (10µg/ml)
Supplementary Figures
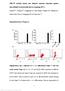 MiR-29 controls innate and adaptive immune responses against intracellular bacterial infection by targeting IFN-γ Feng Ma 1,2,5, Sheng Xu 1,5, Xingguang Liu 1, Qian Zhang 1, Xiongfei Xu 1, Mofang Liu 3,
MiR-29 controls innate and adaptive immune responses against intracellular bacterial infection by targeting IFN-γ Feng Ma 1,2,5, Sheng Xu 1,5, Xingguang Liu 1, Qian Zhang 1, Xiongfei Xu 1, Mofang Liu 3,
Supplementary Table 1. Oligonucleotide primers used in quantitative and qualitative PCRs of this study.
 Supplementary Table 1. Oligonucleotide primers used in quantitative and qualitative PCRs of this study. Primer Sequence Qualitative PCR primers Universal 16S rdna forward 5 - GACGCTGAGGCATGAGAGCA-3 Universal
Supplementary Table 1. Oligonucleotide primers used in quantitative and qualitative PCRs of this study. Primer Sequence Qualitative PCR primers Universal 16S rdna forward 5 - GACGCTGAGGCATGAGAGCA-3 Universal
Transcription factor Foxp3 and its protein partners form a complex regulatory network
 Supplementary figures Resource Paper Transcription factor Foxp3 and its protein partners form a complex regulatory network Dipayan Rudra 1, Paul deroos 1, Ashutosh Chaudhry 1, Rachel Niec 1, Aaron Arvey
Supplementary figures Resource Paper Transcription factor Foxp3 and its protein partners form a complex regulatory network Dipayan Rudra 1, Paul deroos 1, Ashutosh Chaudhry 1, Rachel Niec 1, Aaron Arvey
Suppl Video: Tumor cells (green) and monocytes (white) are seeded on a confluent endothelial
 Supplementary Information Häuselmann et al. Monocyte induction of E-selectin-mediated endothelial activation releases VE-cadherin junctions to promote tumor cell extravasation in the metastasis cascade
Supplementary Information Häuselmann et al. Monocyte induction of E-selectin-mediated endothelial activation releases VE-cadherin junctions to promote tumor cell extravasation in the metastasis cascade
Generation of ST2-GFP reporter mice and characterization of ILC1 cells following infection
 Supplementary Figure 1 Generation of ST2-GFP reporter mice and characterization of ILC1 cells following infection with influenza virus. (a) ST2-GFP reporter mice were generated as described in Methods.
Supplementary Figure 1 Generation of ST2-GFP reporter mice and characterization of ILC1 cells following infection with influenza virus. (a) ST2-GFP reporter mice were generated as described in Methods.
Toluidin-Staining of mast cells Ear tissue was fixed with Carnoy (60% ethanol, 30% chloroform, 10% acetic acid) overnight at 4 C, afterwards
 Toluidin-Staining of mast cells Ear tissue was fixed with Carnoy (60% ethanol, 30% chloroform, 10% acetic acid) overnight at 4 C, afterwards incubated in 100 % ethanol overnight at 4 C and embedded in
Toluidin-Staining of mast cells Ear tissue was fixed with Carnoy (60% ethanol, 30% chloroform, 10% acetic acid) overnight at 4 C, afterwards incubated in 100 % ethanol overnight at 4 C and embedded in
Supplementary Figures. T Cell Factor-1 initiates T helper 2 fate by inducing GATA-3 and repressing Interferon-γ
 Supplementary Figures T Cell Factor-1 initiates T helper 2 fate by inducing GATA-3 and repressing Interferon-γ Qing Yu, Archna Sharma, Sun Young Oh, Hyung-Geun Moon, M. Zulfiquer Hossain, Theresa M. Salay,
Supplementary Figures T Cell Factor-1 initiates T helper 2 fate by inducing GATA-3 and repressing Interferon-γ Qing Yu, Archna Sharma, Sun Young Oh, Hyung-Geun Moon, M. Zulfiquer Hossain, Theresa M. Salay,
Supplementary Information
 Supplementary Information Supplementary Figure 1! a! b! Nfatc1!! Nfatc1"! P1! P2! pa1! pa2! ex1! ex2! exons 3-9! ex1! ex11!!" #" Nfatc1A!!" Nfatc1B! #"!" Nfatc1C! #" DN1! DN2! DN1!!A! #A!!B! #B!!C! #C!!A!
Supplementary Information Supplementary Figure 1! a! b! Nfatc1!! Nfatc1"! P1! P2! pa1! pa2! ex1! ex2! exons 3-9! ex1! ex11!!" #" Nfatc1A!!" Nfatc1B! #"!" Nfatc1C! #" DN1! DN2! DN1!!A! #A!!B! #B!!C! #C!!A!
Supplemental Table I.
 Supplemental Table I Male / Mean ± SEM n Mean ± SEM n Body weight, g 29.2±0.4 17 29.7±0.5 17 Total cholesterol, mg/dl 534.0±30.8 17 561.6±26.1 17 HDL-cholesterol, mg/dl 9.6±0.8 17 10.1±0.7 17 Triglycerides,
Supplemental Table I Male / Mean ± SEM n Mean ± SEM n Body weight, g 29.2±0.4 17 29.7±0.5 17 Total cholesterol, mg/dl 534.0±30.8 17 561.6±26.1 17 HDL-cholesterol, mg/dl 9.6±0.8 17 10.1±0.7 17 Triglycerides,
Optimizing Intracellular Flow Cytometry:
 Optimizing Intracellular Flow Cytometry: Simultaneous Detection of Cytokines and Transcription Factors An encore presentation by Jurg Rohrer, PhD, BD Biosciences 10.26.10 Outline Introduction Cytokines
Optimizing Intracellular Flow Cytometry: Simultaneous Detection of Cytokines and Transcription Factors An encore presentation by Jurg Rohrer, PhD, BD Biosciences 10.26.10 Outline Introduction Cytokines
B220 CD4 CD8. Figure 1. Confocal Image of Sensitized HLN. Representative image of a sensitized HLN
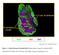 B220 CD4 CD8 Natarajan et al., unpublished data Figure 1. Confocal Image of Sensitized HLN. Representative image of a sensitized HLN showing B cell follicles and T cell areas. 20 µm thick. Image of magnification
B220 CD4 CD8 Natarajan et al., unpublished data Figure 1. Confocal Image of Sensitized HLN. Representative image of a sensitized HLN showing B cell follicles and T cell areas. 20 µm thick. Image of magnification
Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to
 Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most
 Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most differentially expressed between human synovial fibroblasts
Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most differentially expressed between human synovial fibroblasts
B6.SJL (Ly5.2) mice were obtained from Taconic Farms. Rag1-deficient mice were
 Supplementary Methods Mice. B6.SJL (Ly5.2) mice were obtained from Taconic Farms. Rag1-deficient mice were purchased from The Jackson Laboratory. Real-time PCR Total cellular RNA was extracted from the
Supplementary Methods Mice. B6.SJL (Ly5.2) mice were obtained from Taconic Farms. Rag1-deficient mice were purchased from The Jackson Laboratory. Real-time PCR Total cellular RNA was extracted from the
PBMC from each patient were suspended in AIM V medium (Invitrogen) with 5% human
 Anti-CD19-CAR transduced T-cell preparation PBMC from each patient were suspended in AIM V medium (Invitrogen) with 5% human AB serum (Gemini) and 300 international units/ml IL-2 (Novartis). T cell proliferation
Anti-CD19-CAR transduced T-cell preparation PBMC from each patient were suspended in AIM V medium (Invitrogen) with 5% human AB serum (Gemini) and 300 international units/ml IL-2 (Novartis). T cell proliferation
SUPPLEMENT Supplementary Figure 1: (A) (B)
 SUPPLEMENT Supplementary Figure 1: CD4 + naïve effector T cells (CD4 effector) were labeled with CFSE, stimulated with α-cd2/cd3/cd28 coated beads (at 2 beads/cell) and cultured alone or cocultured with
SUPPLEMENT Supplementary Figure 1: CD4 + naïve effector T cells (CD4 effector) were labeled with CFSE, stimulated with α-cd2/cd3/cd28 coated beads (at 2 beads/cell) and cultured alone or cocultured with
L-selectin Is Essential for Delivery of Activated CD8 + T Cells to Virus-Infected Organs for Protective Immunity
 Cell Reports Supplemental Information L-selectin Is Essential for Delivery of Activated CD8 + T Cells to Virus-Infected Organs for Protective Immunity Rebar N. Mohammed, H. Angharad Watson, Miriam Vigar,
Cell Reports Supplemental Information L-selectin Is Essential for Delivery of Activated CD8 + T Cells to Virus-Infected Organs for Protective Immunity Rebar N. Mohammed, H. Angharad Watson, Miriam Vigar,
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
Akt and mtor pathways differentially regulate the development of natural and inducible. T H 17 cells
 Akt and mtor pathways differentially regulate the development of natural and inducible T H 17 cells Jiyeon S Kim, Tammarah Sklarz, Lauren Banks, Mercy Gohil, Adam T Waickman, Nicolas Skuli, Bryan L Krock,
Akt and mtor pathways differentially regulate the development of natural and inducible T H 17 cells Jiyeon S Kim, Tammarah Sklarz, Lauren Banks, Mercy Gohil, Adam T Waickman, Nicolas Skuli, Bryan L Krock,
Supplementary Figure 1
 Supplementary Figure 1 Identification of IFN-γ-producing CD8 + and CD4 + T cells with naive phenotype by alternative gating and sample-processing strategies. a. Contour 5% probability plots show definition
Supplementary Figure 1 Identification of IFN-γ-producing CD8 + and CD4 + T cells with naive phenotype by alternative gating and sample-processing strategies. a. Contour 5% probability plots show definition
Supplementary Materials for
 immunology.sciencemag.org/cgi/content/full/2/16/eaan6049/dc1 Supplementary Materials for Enzymatic synthesis of core 2 O-glycans governs the tissue-trafficking potential of memory CD8 + T cells Jossef
immunology.sciencemag.org/cgi/content/full/2/16/eaan6049/dc1 Supplementary Materials for Enzymatic synthesis of core 2 O-glycans governs the tissue-trafficking potential of memory CD8 + T cells Jossef
Supplemental Materials
 Supplemental Materials Programmed death one homolog maintains the pool size of regulatory T cells by promoting their differentiation and stability Qi Wang 1, Jianwei He 1, Dallas B. Flies 2, Liqun Luo
Supplemental Materials Programmed death one homolog maintains the pool size of regulatory T cells by promoting their differentiation and stability Qi Wang 1, Jianwei He 1, Dallas B. Flies 2, Liqun Luo
Human and mouse T cell regulation mediated by soluble CD52 interaction with Siglec-10. Esther Bandala-Sanchez, Yuxia Zhang, Simone Reinwald,
 Human and mouse T cell regulation mediated by soluble CD52 interaction with Siglec-1 Esther Bandala-Sanchez, Yuxia Zhang, Simone Reinwald, James A. Dromey, Bo Han Lee, Junyan Qian, Ralph M Böhmer and Leonard
Human and mouse T cell regulation mediated by soluble CD52 interaction with Siglec-1 Esther Bandala-Sanchez, Yuxia Zhang, Simone Reinwald, James A. Dromey, Bo Han Lee, Junyan Qian, Ralph M Böhmer and Leonard
Supplementary Figure 1 a
 Supplementary Figure a Normalized expression/tbp (A.U.).6... Trip-br transcripts Trans Trans Trans b..5. Trip-br Ctrl LPS Normalized expression/tbp (A.U.) c Trip-br transcripts. adipocytes.... Trans Trans
Supplementary Figure a Normalized expression/tbp (A.U.).6... Trip-br transcripts Trans Trans Trans b..5. Trip-br Ctrl LPS Normalized expression/tbp (A.U.) c Trip-br transcripts. adipocytes.... Trans Trans
Supplementary Fig. 1 No relative growth advantage of Foxp3 negative cells.
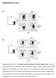 Supplementary Fig. 1 Supplementary Figure S1: No relative growth advantage of Foxp3 negative cells. itreg were induced from WT (A) or FIR (B) CD4 + T cells. FIR itregs were then removed from the TCR signal
Supplementary Fig. 1 Supplementary Figure S1: No relative growth advantage of Foxp3 negative cells. itreg were induced from WT (A) or FIR (B) CD4 + T cells. FIR itregs were then removed from the TCR signal
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Interleukin-2-Dependent Allergen-Specific Tissue-Resident Memory Cells Drive Asthma
 Immunity Supplemental Information Interleukin-2-Dependent Allergen-Specific Tissue-Resident Memory Cells Drive Asthma Brian D. Hondowicz, Dowon An, Jason M. Schenkel, Karen S. Kim, Holly R. Steach, Akshay
Immunity Supplemental Information Interleukin-2-Dependent Allergen-Specific Tissue-Resident Memory Cells Drive Asthma Brian D. Hondowicz, Dowon An, Jason M. Schenkel, Karen S. Kim, Holly R. Steach, Akshay
SUPPLEMENTARY INFORMATION
 Complete but curtailed T-cell response to very-low-affinity antigen Dietmar Zehn, Sarah Y. Lee & Michael J. Bevan Supp. Fig. 1: TCR chain usage among endogenous K b /Ova reactive T cells. C57BL/6 mice
Complete but curtailed T-cell response to very-low-affinity antigen Dietmar Zehn, Sarah Y. Lee & Michael J. Bevan Supp. Fig. 1: TCR chain usage among endogenous K b /Ova reactive T cells. C57BL/6 mice
Supplementary Information. Supplementary Figure 1
 Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Figure 1. IL-12 serum levels and frequency of subsets in FL patients. (A) IL-12
 1 Supplementary Data Figure legends Supplementary Figure 1. IL-12 serum levels and frequency of subsets in FL patients. (A) IL-12 serum levels measured by multiplex ELISA (Luminex) in FL patients before
1 Supplementary Data Figure legends Supplementary Figure 1. IL-12 serum levels and frequency of subsets in FL patients. (A) IL-12 serum levels measured by multiplex ELISA (Luminex) in FL patients before
Online Appendix Material and Methods: Pancreatic RNA isolation and quantitative real-time (q)rt-pcr. Mice were fasted overnight and killed 1 hour (h)
 Online Appendix Material and Methods: Pancreatic RNA isolation and quantitative real-time (q)rt-pcr. Mice were fasted overnight and killed 1 hour (h) after feeding. A small slice (~5-1 mm 3 ) was taken
Online Appendix Material and Methods: Pancreatic RNA isolation and quantitative real-time (q)rt-pcr. Mice were fasted overnight and killed 1 hour (h) after feeding. A small slice (~5-1 mm 3 ) was taken
Optimizing Intracellular Flow Cytometry:
 Optimizing Intracellular Flow Cytometry: Simultaneous Detection of Cytokines and Transcription Factors Presented by Jurg Rohrer, PhD, BD Biosciences 23-10780-00 Outline Introduction Cytokines Transcription
Optimizing Intracellular Flow Cytometry: Simultaneous Detection of Cytokines and Transcription Factors Presented by Jurg Rohrer, PhD, BD Biosciences 23-10780-00 Outline Introduction Cytokines Transcription
ice-cold 70% ethanol with gentle vortexing, incubated at -20 C for 4 hours, and washed with PBS.
 Cell cycle analysis For cell cycle analysis, single cell suspensions of E12.5 fetal liver cells were suspended in 4 ml ice-cold 7% ethanol with gentle vortexing, incubated at -2 C for 4 hours, and washed
Cell cycle analysis For cell cycle analysis, single cell suspensions of E12.5 fetal liver cells were suspended in 4 ml ice-cold 7% ethanol with gentle vortexing, incubated at -2 C for 4 hours, and washed
Supplementary Figure 1. Efficient DC depletion in CD11c.DOG transgenic mice
 Supplementary Figure 1. Efficient DC depletion in CD11c.DOG transgenic mice (a) CD11c.DOG transgenic mice (tg) were treated with 8 ng/g body weight (b.w.) diphtheria toxin (DT) i.p. on day -1 and every
Supplementary Figure 1. Efficient DC depletion in CD11c.DOG transgenic mice (a) CD11c.DOG transgenic mice (tg) were treated with 8 ng/g body weight (b.w.) diphtheria toxin (DT) i.p. on day -1 and every
IMMUNOLOGICAL MEMORY. CD4 T Follicular Helper Cells. Memory CD8 T Cell Differentiation
 IMMUNOLOGICAL MEMORY CD4 T Follicular Helper Cells Memory CD8 T Cell Differentiation CD4 T Cell Differentiation Bcl-6 T-bet GATA-3 ROR t Foxp3 CD4 T follicular helper (Tfh) cells FUNCTION Provide essential
IMMUNOLOGICAL MEMORY CD4 T Follicular Helper Cells Memory CD8 T Cell Differentiation CD4 T Cell Differentiation Bcl-6 T-bet GATA-3 ROR t Foxp3 CD4 T follicular helper (Tfh) cells FUNCTION Provide essential
Nature Medicine doi: /nm.3957
 Supplementary Fig. 1. p38 alternative activation, IL-21 expression, and T helper cell transcription factors in PDAC tissue. (a) Tissue microarrays of pancreatic tissue from 192 patients with pancreatic
Supplementary Fig. 1. p38 alternative activation, IL-21 expression, and T helper cell transcription factors in PDAC tissue. (a) Tissue microarrays of pancreatic tissue from 192 patients with pancreatic
Transduction of lentivirus to human primary CD4+ T cells
 Transduction of lentivirus to human primary CD4 + T cells Human primary CD4 T cells were stimulated with anti-cd3/cd28 antibodies (10 µl/2 5 10^6 cells of Dynabeads CD3/CD28 T cell expander, Invitrogen)
Transduction of lentivirus to human primary CD4 + T cells Human primary CD4 T cells were stimulated with anti-cd3/cd28 antibodies (10 µl/2 5 10^6 cells of Dynabeads CD3/CD28 T cell expander, Invitrogen)
Categorical analysis of human T cell heterogeneity with One-SENSE
 1 2 3 4 Supplementary Information Categorical analysis of human T cell heterogeneity with One-SENSE 5 6 Running title: T cell Analysis by One-SENSE 7 8 9 1 11 12 13 14 15 16 17 18 19 2 21 22 23 24 25 26
1 2 3 4 Supplementary Information Categorical analysis of human T cell heterogeneity with One-SENSE 5 6 Running title: T cell Analysis by One-SENSE 7 8 9 1 11 12 13 14 15 16 17 18 19 2 21 22 23 24 25 26
Supplemental Figure 1. Signature gene expression in in vitro differentiated Th0, Th1, Th2, Th17 and Treg cells. (A) Naïve CD4 + T cells were cultured
 Supplemental Figure 1. Signature gene expression in in vitro differentiated Th0, Th1, Th2, Th17 and Treg cells. (A) Naïve CD4 + T cells were cultured under Th0, Th1, Th2, Th17, and Treg conditions. mrna
Supplemental Figure 1. Signature gene expression in in vitro differentiated Th0, Th1, Th2, Th17 and Treg cells. (A) Naïve CD4 + T cells were cultured under Th0, Th1, Th2, Th17, and Treg conditions. mrna
SUPPORTING INFORMATIONS
 SUPPORTING INFORMATIONS Mice MT/ret RetCD3ε KO α-cd25 treated MT/ret Age 1 month 3 mnths 6 months 1 month 3 months 6 months 1 month 3 months 6 months 2/87 Survival 87/87 incidence of 17/87 1 ary tumor
SUPPORTING INFORMATIONS Mice MT/ret RetCD3ε KO α-cd25 treated MT/ret Age 1 month 3 mnths 6 months 1 month 3 months 6 months 1 month 3 months 6 months 2/87 Survival 87/87 incidence of 17/87 1 ary tumor
Supplementary Data. Treg phenotype
 Supplementary Data Additional Experiment An additional experiment was performed using cryopreserved peripheral blood mononuclear cells (PBMC) derived from five renal cell carcinoma (RCC) patients [see
Supplementary Data Additional Experiment An additional experiment was performed using cryopreserved peripheral blood mononuclear cells (PBMC) derived from five renal cell carcinoma (RCC) patients [see
