Supplementary figures legends
|
|
|
- Mildred Cannon
- 5 years ago
- Views:
Transcription
1 Supplementary Information to Nijmegen Breakage Syndrome fibroblasts and ipscs: cellular models for uncovering diseaseassociated signaling pathways and establishing a screening platform for anti-oxidants by Barbara Mlody, Wasco Wruck, Soraia Martins, Karl Sperling and James Adjaye Supplementary figures legends Supplementary Figure S1: Characterization of the NBS mutation. (Modified from our previous publication 1 ) a) PCR-based analysis of genomic DNA with primers flanking the deletion, results in either a 60 bp (wt) or 55 bp (657del5) amplicon. b) Sanger sequencing results comparing the genomic DNA sequence of fibroblast sample NBS8 against HFF1 fibroblasts (normal), on the top the blasting result ( from HFF1 Exon 6 (fw) against NBN mrna (NM_ ), then NBS 1 Exon 6 (fw) against HFF1 Exon 6 (fw) and NBS 8 Exon 6 (fw) against HFF1 Exon 6 (fw). Below the chromatogram of the sequence is shown. c) Immunoflurescent detection of full-length (p95) NBN in patient fibroblasts. Even loading of the SDS-gel is shown by house-keeping protein GAPDH. d) Western Blot detection of p95-nbn in reprogrammed cells. ß-Actin serves as a loading control. For the sake of better readability western blots were cropped. Supplementary Figure S2: Sanger sequencing confirms the NBS causing deletion 657del5 in the NBS-iPSCs. Results of Sanger sequencing of the NBS8-iPSCs showing the chromatogram of the region around the heterozygous 657del5 deletion in the NBS gene. Supplementary Figure S3: NBS-iPSCs: Differentially regulated genes involved in Glycolysis. KEGG pathway (hsa00010) 2 of Glycolysis/Gluconeogenesis. In red: up-regulated; in green: down-regulated genes in NBS-iPSCs vs. control, classified by differential p-value < 0.05 and fold-change > 1.5. Supplementary Figure S4: NBS-iPSCs vs. ESCs: Differentially expressed genes from Pathways in cancer and Cell cycle. Genes differentially expressed in NBS-iPSCs vs. control, classified by differential p-value < 0.05 and fold-change > 1.5 and functionally annotated as Pathways in cancer were further refined by another DAVID analysis. Resulting genes annotated with the KEGG pathway Cell cycle 2 were mapped to the chart of this pathway. Supplementary Figure S5: NBS-iPSCs vs. ESCs: Differentially expressed genes from Pathways in cancer and p53-signaling. Genes differentially expressed in NBS-iPSCs vs. control, classified by differential p-value < 0.05 and fold-change > 1.5 and functionally annotated as Pathways in cancer were further refined by another DAVID analysis. Resulting genes annotated with the KEGG pathway p53-signaling 2 were mapped to the chart of this pathway. Supplementary Figure S6: Reprogramming of NBS fibroblasts into ipscs shifts energy supply to Glycolysis. Hierarchical cluster analysis and heatmap of regulated genes from the KEGG pathway Glycolysis 2 in a) NBS fibroblasts and b) NBS-iPSCs shows a change in the Glycolysis pathway from predominantly down-regulated in NBS fibroblasts to predominantly up-regulated in NBS-iPSCs. The
2 sample clustering displays a clear separation of NBS and healthy control clusters. (Color bars: blue NBS, red control; high expression: red, low expression; green). Supplementary Figure S7: Cluster analysis of Oxidative phosphorylation pathway in NBS fibroblasts and ipscs. Hierarchical cluster analysis and heatmap of genes from the KEGG pathway Oxidative phosphorylation 2 in a) NBS fibroblasts and b) NBS-iPSCs shows clear separation of NBS and healthy control clusters. If at all, there is only a slight tendency of more down-regulation of the NBS resulting from the reprogramming because the predominantly down-regulated cluster is bigger in the NBSiPSCs than in the NBS fibroblasts. (Color bars: blue NBS, red control; high expression: red, low expression; green). Supplementary Figure S8: Cluster analysis of p53 signaling pathway in NBS fibroblasts and ipscs. Hierarchical cluster analysis and heatmap of genes from the KEGG pathway p53-signaling 2 in NBS fibroblasts and NBS-iPSCs show a cluster of pluripotent stem cells and another cluster of fibroblasts. Within these clusters NBS and healthy control samples were separated. Passage 8 and 15 of the only NBS line which could be reprogrammed showed differences to the other NBS fibroblast lines (Color bars: blue NBS, red control; high expression: red, low expression; green). References 1. Mlody, B. & Adjaye, J. Generation of ipsc lines from a Nijmegen Breakage Syndrome patient. Stem Cell Res. 15, (2015). 2. Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353 D361 (2017).
3 Supplementary figures
4 Supplementary Figure S1: Characterization of the NBS mutation
5 Supplementary Figure S2: Sanger sequencing confirms the NBS causing deletion 657del5 in the NBS-iPSCs
6 Supplementary Figure S3: NBS-iPSCs: Differentially regulated genes involved in Glycolysis
7 Supplementary Figure S4: NBS-iPSCs vs. ESCs: Differentially expressedgenes from Pathways in cancer and Cell cycle Supplementary Figure S5: NBS-iPSCs vs. ESCs: Differentially expressed genes from Pathways in cancer and p53-signaling
8 Supplementary Figure S6: Reprogramming of NBS fibroblasts into ipscs shifts energy supply to Glycolysis.
9 Supplementary Figure S7: Cluster analysis of Oxidative phosphorylation pathway in NBS fibroblasts and ipscs.
10 Supplementary Figure S8: Cluster analysis of p53 signaling pathway in NBS fibroblasts and ipscs.
11 Supplementary table S1 to Supplementary Figure S3: Gene information PROBE_ID SYMBOL Ratio EC Definition Name ILMN_ AKR1A aldo-keto reductase family 1, member A1 ILMN_ ALDOA aldolase A, fructosebisphosphate ILMN_ ALDOA aldolase A, fructosebisphosphate ILMN_ ALDOC aldolase C, fructosebisphosphate ILMN_ ENO enolase 2 (gamma, neuronal) ILMN_ GAPDH glyceraldehyde-3- phosphate dehydrogenase ILMN_ GAPDH glyceraldehyde-3- phosphate dehydrogenase ILMN_ GPI glucose-6-phosphate isomerase ILMN_ HK hexokinase 2 ILMN_ LDHA ILMN_ PFKL ILMN_ PFKM ILMN_ PGK1 ILMN_ PGK1 ILMN_ TPI1 ILMN_ TPI1 ILMN_ FBP lactate dehydrogenase A phosphofructokinase, liver phosphofructokinase, muscle phosphoglycerate kinase phosphoglycerate kinase triosephosphate isomerase triosephosphate isomerase fructose-1,6- bisphosphatase 1
12 Supplementary table S2: Refinement of genes differentially expressed in NBS-iPSCs vs. HESCs and annotated with pathways in cancer (DAVID funtional annotations) Term PValue Genes Benjamini hsa05222:small cell lung cancer CCNE1, E2F2, SKP2, NFKB1, CDK6, PIAS2, CDK4, LAMB1, MYC, ITGB1, 1.58E-12 PIK3R1, CHUK 1.02E-10 hsa05220:chronic myeloid leukemia 8.93E-12 E2F2, CBLC, HRAS, CBLB, CTBP2, NFKB1, CDK6, CDK4, MYC, PIK3R1, CHUK 3.87E-10 hsa04151:pi3k-akt signaling pathway FGF19, HRAS, FGF8, FGF11, KITLG, NFKB1, CDK6, FGF13, KIT, CDK4, 5.38E-11 ITGB1, CCNE1, VEGFA, LAMB1, MYC, PIK3R1, CHUK 1.75E-09 hsa05166:htlv-i infection E2F2, HRAS, WNT10B, WNT3, SLC2A1, NFKB1, NFKB2, FZD5, CDK4, FZD4, 1.79E-09 MYC, CHUK, FZD7, PIK3R1 4.65E-08 hsa05212:pancreatic cancer 3.61E-09 E2F2, RALBP1, VEGFA, TGFA, NFKB1, CDK6, CDK4, PIK3R1, CHUK 7.82E-08 hsa04014:ras signaling pathway FGF19, HRAS, FGF8, RALBP1, RASSF1, VEGFA, FGF11, KITLG, NFKB1, 5.20E-09 FGF13, KIT, CHUK, PIK3R1 9.66E-08 hsa05218:melanoma 7.41E-09 FGF19, E2F2, HRAS, FGF8, FGF11, FGF13, CDK6, CDK4, PIK3R1 1.20E-07 hsa05205:proteoglycans in cancer CBLC, HRAS, CBLB, WNT10B, WNT3, VEGFA, FZD5, FZD4, MYC, ITGB1, 1.77E-08 FZD7, PIK3R1 2.56E-07 hsa05211:renal cell carcinoma 9.75E-08 HRAS, VHL, VEGFA, SLC2A1, TCEB2, TGFA, EGLN1, PIK3R1 1.27E-06 hsa04550:signaling pathways regulating pluripotency of stem cells 1.12E-07 BMP4, BMP2, HRAS, WNT10B, WNT3, FZD5, FZD4, MYC, FZD7, PIK3R1 1.33E-06 hsa05161:hepatitis B 1.52E-07 CCNE1, E2F2, HRAS, BIRC5, NFKB1, CDK6, CDK4, MYC, PIK3R1, CHUK 1.65E-06 hsa04390:hippo signaling pathway 2.16E-07 BMP4, BMP2, WNT10B, WNT3, RASSF1, BIRC5, FZD5, FZD4, MYC, FZD7 2.16E-06 hsa05217:basal cell carcinoma 8.09E-07 BMP4, BMP2, WNT10B, WNT3, FZD5, FZD4, FZD7 7.51E-06 hsa05223:non-small cell lung cancer 9.02E-07 E2F2, HRAS, RASSF1, TGFA, CDK6, CDK4, PIK3R1 7.82E-06 hsa04916:melanogenesis 1.92E-06 HRAS, WNT10B, WNT3, KITLG, KIT, FZD5, FZD4, FZD7 1.56E-05 hsa04015:rap1 signaling pathway 3.47E-06 FGF19, HRAS, FGF8, VEGFA, FGF11, KITLG, FGF13, KIT, ITGB1, PIK3R1 2.66E-05 hsa05219:bladder cancer 4.14E-06 E2F2, HRAS, RASSF1, VEGFA, CDK4, MYC 2.99E-05 hsa05215:prostate cancer 1.30E-05 CCNE1, E2F2, HRAS, TGFA, NFKB1, PIK3R1, CHUK 8.93E-05 hsa05221:acute myeloid leukemia 1.97E-05 HRAS, NFKB1, KIT, MYC, PIK3R1, CHUK 1.28E-04 hsa04066:hif-1 signaling pathway 2.42E-05 VHL, VEGFA, SLC2A1, TCEB2, NFKB1, EGLN1, PIK3R1 1.50E-04 hsa04660:t cell receptor signaling pathway 3.22E-05 CBLC, HRAS, CBLB, NFKB1, CDK4, PIK3R1, CHUK 1.90E-04 hsa05214:glioma 4.10E-05 E2F2, HRAS, TGFA, CDK6, CDK4, PIK3R1 2.32E-04 hsa04010:mapk signaling pathway 1.22E-04 FGF19, HRAS, FGF8, FGF11, FGF13, NFKB1, NFKB2, MYC, CHUK 6.62E-04 hsa04310:wnt signaling pathway 1.65E-04 WNT10B, WNT3, CTBP2, FZD5, FZD4, MYC, FZD7 8.59E-04 hsa04012:erbb signaling pathway 1.66E-04 CBLC, HRAS, CBLB, TGFA, MYC, PIK3R1 8.30E-04 hsa05203:viral carcinogenesis 2.04E-04 CCNE1, HRAS, SKP2, NFKB1, CDK6, NFKB2, CDK4, PIK3R1 9.80E-04 hsa05206:micrornas in cancer 2.63E-04 CCNE1, E2F2, HRAS, WNT3, RASSF1, VEGFA, NFKB1, CDK6, MYC hsa05230:central carbon metabolism in cancer 5.75E-04 HRAS, SLC2A1, KIT, MYC, PIK3R hsa05216:thyroid cancer 6.84E-04 HRAS, TPR, MYC, TPM hsa04110:cell cycle 8.58E-04 CCNE1, E2F2, SKP2, CDK6, CDK4, MYC hsa05162:measles CCNE1, NFKB1, CDK6, CDK4, PIK3R1, CHUK hsa04120:ubiquitin mediated proteolysis CBLC, CBLB, VHL, TCEB2, SKP2, PIAS hsa04810:regulation of actin cytoskeleton FGF19, HRAS, FGF8, FGF11, FGF13, ITGB1, PIK3R hsa05213:endometrial cancer HRAS, MLH1, MYC, PIK3R hsa05145:toxoplasmosis NFKB1, LAMB1, ITGB1, PIK3R1, CHUK hsa05169:epstein-barr virus infection SKP2, NFKB1, NFKB2, MYC, PIK3R1, CHUK hsa04210:apoptosis BID, NFKB1, PIK3R1, CHUK hsa05210:colorectal cancer MLH1, BIRC5, MYC, PIK3R hsa04115:p53 signaling pathway BID, CCNE1, CDK6, CDK hsa04662:b cell receptor signaling pathway HRAS, NFKB1, PIK3R1, CHUK hsa05100:bacterial invasion of epithelial cells CBLC, CBLB, ITGB1, PIK3R
Chapter 15 Glycolysis and The Pentose Phosphate Pathway
 Principles of Biochemistry Fourth Edition Donald Voet Judith G. Voet harlotte W. Pratt hapter 15 Glycolysis and The Pentose Phosphate Pathway Page No. 47-490 Introduction Glucose: is major source of metabolic
Principles of Biochemistry Fourth Edition Donald Voet Judith G. Voet harlotte W. Pratt hapter 15 Glycolysis and The Pentose Phosphate Pathway Page No. 47-490 Introduction Glucose: is major source of metabolic
SUPPLEMENTARY DATA. Supplementary Table 1. Primers used in qpcr
 Supplementary Table 1. Primers used in qpcr Gene forward primer (5'-3') reverse primer (5'-3') β-actin AGAGGGAAATCGTGCGTGAC CAATAGTGATGACCTGGCCGT Hif-p4h-2 CTGGGCAACTACAGGATAAAC GCGTCCCAGTCTTTATTTAGATA
Supplementary Table 1. Primers used in qpcr Gene forward primer (5'-3') reverse primer (5'-3') β-actin AGAGGGAAATCGTGCGTGAC CAATAGTGATGACCTGGCCGT Hif-p4h-2 CTGGGCAACTACAGGATAAAC GCGTCCCAGTCTTTATTTAGATA
MBioS 303 Recitation Introductory Biochemistry, Summer 2008 Practice Problem Set #7: General Metabolism Concepts, Glycolysis and the TCA Cycle
 MBioS 303 Recitation Introductory Biochemistry, Summer 2008 Practice Problem Set #7: General Metabolism Concepts, Glycolysis and the TCA Cycle (1) Glucose 1-pohsphate is converted to fructose 6-phosphate
MBioS 303 Recitation Introductory Biochemistry, Summer 2008 Practice Problem Set #7: General Metabolism Concepts, Glycolysis and the TCA Cycle (1) Glucose 1-pohsphate is converted to fructose 6-phosphate
ATP. And how do we do that? Cellular Respiration Stage 1: Glycolysis. The point is to make ATP! What s the point? ATP synthase.
 ellular Respiration Stage 1: Glycolysis A 007-008 Biology What s the point? The point is to make! A 007-008 Biology And how do we do that? synthase set up a + gradient allow + to flow through synthase
ellular Respiration Stage 1: Glycolysis A 007-008 Biology What s the point? The point is to make! A 007-008 Biology And how do we do that? synthase set up a + gradient allow + to flow through synthase
Cellular Respiration Stage 1: Glycolysis
 Cellular Respiration Stage 1: Glycolysis 2007-2008 What s the point? The point is to make! 2007-2008 Glycolysis Breaking down glucose glyco lysis (splitting sugar) glucose pyruvate 6C 2x 3C In the cytosol?
Cellular Respiration Stage 1: Glycolysis 2007-2008 What s the point? The point is to make! 2007-2008 Glycolysis Breaking down glucose glyco lysis (splitting sugar) glucose pyruvate 6C 2x 3C In the cytosol?
Cellular Respiration Stage 1: (Glycolysis) AP Biology
 Cellular Respiration Stage 1: (Glycolysis) What s the point? The point is to make! Glycolysis: Breaking down glucose glyco lysis (splitting sugar) glucose pyruvate 6C 2x 3C In the cytosol? Why does that
Cellular Respiration Stage 1: (Glycolysis) What s the point? The point is to make! Glycolysis: Breaking down glucose glyco lysis (splitting sugar) glucose pyruvate 6C 2x 3C In the cytosol? Why does that
Aerobic Respiration. The four stages in the breakdown of glucose
 Aerobic Respiration The four stages in the breakdown of glucose 1 I. Aerobic Respiration Why can t we break down Glucose in one step? (Flaming Gummy Bear) Enzymes gently lower the potential energy until
Aerobic Respiration The four stages in the breakdown of glucose 1 I. Aerobic Respiration Why can t we break down Glucose in one step? (Flaming Gummy Bear) Enzymes gently lower the potential energy until
Glycolysis. Degradation of Glucose to yield pyruvate
 Glycolysis Degradation of Glucose to yield pyruvate After this Lecture you will be able to answer: For each step of glycolysis: How does it occur? Why does it occur? Is it Regulated? How? What are the
Glycolysis Degradation of Glucose to yield pyruvate After this Lecture you will be able to answer: For each step of glycolysis: How does it occur? Why does it occur? Is it Regulated? How? What are the
0.40. Biochemistry of Carbohydrates
 0.40 Biochemistry of Carbohydrates Biochemistry of Carbohydrates ATP ADP Glycolysis The Breakdown of Glucose Primary Energy Source of Cells Central Metabolic Pathway All Reactions Occur in Cytoplasm Two
0.40 Biochemistry of Carbohydrates Biochemistry of Carbohydrates ATP ADP Glycolysis The Breakdown of Glucose Primary Energy Source of Cells Central Metabolic Pathway All Reactions Occur in Cytoplasm Two
How To Use SubpathwayGMir
 How To Use SubpathwayGMir Li Feng, Chunquan Li and Xia Li May 20, 2015 Contents 1 Overview 1 2 The experimentally verified mirna-target interactions 2 3 Reconstruct KEGG metabolic pathways 2 3.1 Embed
How To Use SubpathwayGMir Li Feng, Chunquan Li and Xia Li May 20, 2015 Contents 1 Overview 1 2 The experimentally verified mirna-target interactions 2 3 Reconstruct KEGG metabolic pathways 2 3.1 Embed
Interpreting Diverse Genomic Data Using Gene Sets
 Interpreting Diverse Genomic Data Using Gene Sets giovanni parmigiani@dfci.harvard.edu FHCRC February 2012 Gene Sets Leary 2008 PNAS Alterations in the combined FGF, EGFR, ERBB2 and PIK3 pathways. Red:
Interpreting Diverse Genomic Data Using Gene Sets giovanni parmigiani@dfci.harvard.edu FHCRC February 2012 Gene Sets Leary 2008 PNAS Alterations in the combined FGF, EGFR, ERBB2 and PIK3 pathways. Red:
Interpreting Diverse Genomic Data Using Gene Sets
 Interpreting Diverse Genomic Data Using Gene Sets giovanni parmigiani@dfci.harvard.edu FHCRC February 2012 Gene Sets Leary 2008 PNAS Alterations in the combined FGF, EGFR, ERBB2 and PIK3 pathways. Red:
Interpreting Diverse Genomic Data Using Gene Sets giovanni parmigiani@dfci.harvard.edu FHCRC February 2012 Gene Sets Leary 2008 PNAS Alterations in the combined FGF, EGFR, ERBB2 and PIK3 pathways. Red:
Glycolysis. Glycolysis Expectations. Glycolysis 10/20/2015. Chapter 16, Stryer Short Course. Memorize/learn Figure 16.1
 Glycolysis Chapter 16, Stryer Short Course Glycolysis Expectations Memorize/learn Figure 16.1 Know overall reaction and stages Explain chemical/physiological purpose of each step Learn structures Reversible/Irreversible
Glycolysis Chapter 16, Stryer Short Course Glycolysis Expectations Memorize/learn Figure 16.1 Know overall reaction and stages Explain chemical/physiological purpose of each step Learn structures Reversible/Irreversible
Chem 109 C. Fall Armen Zakarian Office: Chemistry Bldn 2217
 Chem 109 C Fall 2014 Armen Zakarian ffice: Chemistry Bldn 2217 o Catabolism of carbohydrates: 10 reactions of glycolysis Chapter 25 C C 2 C 2 D-glucose α-d-glucopyranose aworth projection α-d-glucopyranose
Chem 109 C Fall 2014 Armen Zakarian ffice: Chemistry Bldn 2217 o Catabolism of carbohydrates: 10 reactions of glycolysis Chapter 25 C C 2 C 2 D-glucose α-d-glucopyranose aworth projection α-d-glucopyranose
Review of Carbohydrate Digestion
 Review of Carbohydrate Digestion Glycolysis Glycolysis is a nine step biochemical pathway that oxidizes glucose into two molecules of pyruvic acid. During this process, energy is released and some of it
Review of Carbohydrate Digestion Glycolysis Glycolysis is a nine step biochemical pathway that oxidizes glucose into two molecules of pyruvic acid. During this process, energy is released and some of it
Supplementary Table 1: List of the 242 hypoxia/reoxygenation marker genes collected from literature and databases
 Supplementary Tables: Supplementary Table 1: List of the 242 hypoxia/reoxygenation marker genes collected from literature and databases Supplementary Table 2: Contingency tables used in the fisher exact
Supplementary Tables: Supplementary Table 1: List of the 242 hypoxia/reoxygenation marker genes collected from literature and databases Supplementary Table 2: Contingency tables used in the fisher exact
Chapter 13 Carbohydrate Metabolism
 Chapter 13 Carbohydrate Metabolism Metabolism of Foods Food is broken down into carbohydrates, lipids, and proteins and sent through catabolic pathways to produce energy. Glycolysis glucose 2 P i 2 ADP
Chapter 13 Carbohydrate Metabolism Metabolism of Foods Food is broken down into carbohydrates, lipids, and proteins and sent through catabolic pathways to produce energy. Glycolysis glucose 2 P i 2 ADP
Photosynthesis in chloroplasts. Cellular respiration in mitochondria ATP. ATP powers most cellular work
 Light energy ECOSYSTEM CO + H O Photosynthesis in chloroplasts Cellular respiration in mitochondria Organic molecules + O powers most cellular work Heat energy 1 becomes oxidized (loses electron) becomes
Light energy ECOSYSTEM CO + H O Photosynthesis in chloroplasts Cellular respiration in mitochondria Organic molecules + O powers most cellular work Heat energy 1 becomes oxidized (loses electron) becomes
Supplementary Figure 1: Detailed schematic of the synthetic biochemistry system for the conversion of glucose to monoterpenes.
 Supplementary Figure 1: Detailed schematic of the synthetic biochemistry system for the conversion of glucose to monoterpenes. Glycolytic and mevalonate enzymes are highlighted in blue and orange respectively.
Supplementary Figure 1: Detailed schematic of the synthetic biochemistry system for the conversion of glucose to monoterpenes. Glycolytic and mevalonate enzymes are highlighted in blue and orange respectively.
SOPten flox/flox (KO) Pten flox/flox (WT) flox allele 6.0 kb. Pten. Actin. ! allele 2.3 kb. Supplementary Figure S1. Yanagi, et al.
 s1 A Pten flox/flox () SOPten flox/flox () flox allele 6. kb B Pten flox/flox () SOPten flox/flox () Pten Actin! allele 2.3 kb Supplementary Figure S1. Yanagi, et al. A B BrdU BrdU positive cells ( ) 3
s1 A Pten flox/flox () SOPten flox/flox () flox allele 6. kb B Pten flox/flox () SOPten flox/flox () Pten Actin! allele 2.3 kb Supplementary Figure S1. Yanagi, et al. A B BrdU BrdU positive cells ( ) 3
CHAPTER 16. Glycolysis
 CHAPTER 16 Glycolysis Net reaction of Glycolysis Converts: 1 Glucose Hexose stage 2 pyruvate - Two molecules of ATP are produced - Two molecules of NAD + are reduced to NADH Triose stage Glucose + 2 ADP
CHAPTER 16 Glycolysis Net reaction of Glycolysis Converts: 1 Glucose Hexose stage 2 pyruvate - Two molecules of ATP are produced - Two molecules of NAD + are reduced to NADH Triose stage Glucose + 2 ADP
Biochemistry of carbohydrates
 Biochemistry of carbohydrates الفريق الطبي األكاديمي Done By: - Hanan Jamal لكية الطب البرشي البلقاء التطبيقية / املركز 6166 6102/ In the last lecture we talked about Pyruvate, pyruvate is a central intermediate;
Biochemistry of carbohydrates الفريق الطبي األكاديمي Done By: - Hanan Jamal لكية الطب البرشي البلقاء التطبيقية / املركز 6166 6102/ In the last lecture we talked about Pyruvate, pyruvate is a central intermediate;
High expression of cellular retinol binding protein-1 in lung adenocarcinoma is associated with poor prognosis
 High expression of cellular retinol binding protein-1 in lung adenocarcinoma is associated with poor prognosis Supplementary Material Supplementary Figure S1. Representative CRBP-1 immunostaining of non-neoplastic
High expression of cellular retinol binding protein-1 in lung adenocarcinoma is associated with poor prognosis Supplementary Material Supplementary Figure S1. Representative CRBP-1 immunostaining of non-neoplastic
Human recombinat MIF protein (hrmif), MW: Da. m/z. hrmif ( Da) + 4-IPP (282 Da) MWtot ~ Da. m/z.
 Intensity % Intensity % A Human recombinat MIF protein (hrmif), MW: 12428.31 Da m/z hrmif (12428.31 Da) + 4-IPP (282 Da) MWtot ~ 12715.21 Da m/z B HTC/C3 DAPI phistone-h3 Merge HTC/C3 DAPI phistone-h3
Intensity % Intensity % A Human recombinat MIF protein (hrmif), MW: 12428.31 Da m/z hrmif (12428.31 Da) + 4-IPP (282 Da) MWtot ~ 12715.21 Da m/z B HTC/C3 DAPI phistone-h3 Merge HTC/C3 DAPI phistone-h3
Glycolysis. BCH 340 lecture 3 Chapter 8 in Lippincott 5 th edition
 Glycolysis B 40 lecture hapter 8 in Lippincott 5 th edition All carbohydrates to be catabolized must enter the glycolytic pathway Glycolysis is degradation of glucose to generate energy (ATP) and to provide
Glycolysis B 40 lecture hapter 8 in Lippincott 5 th edition All carbohydrates to be catabolized must enter the glycolytic pathway Glycolysis is degradation of glucose to generate energy (ATP) and to provide
Chem Lecture 8 Carbohydrate Metabolism Part I: Glycolysis
 Chem 352 - Lecture 8 Carbohydrate Metabolism Part I: Glycolysis Introduction Carbohydrate metabolism involves a collection of pathways. Glycolysis Hexoses 3-Carbon molecules Gluconeogenesis 3-Carbon molecules
Chem 352 - Lecture 8 Carbohydrate Metabolism Part I: Glycolysis Introduction Carbohydrate metabolism involves a collection of pathways. Glycolysis Hexoses 3-Carbon molecules Gluconeogenesis 3-Carbon molecules
GLYCOLYSIS Generation of ATP from Metabolic Fuels
 GLYCOLYSIS Generation of ATP from Metabolic Fuels - Catabolic process degradative pathway - Energy stored in sugars (carbohydrates) released to perform biological work - Transforms GLUCOSE to PYRUVATE
GLYCOLYSIS Generation of ATP from Metabolic Fuels - Catabolic process degradative pathway - Energy stored in sugars (carbohydrates) released to perform biological work - Transforms GLUCOSE to PYRUVATE
Hexose Metabolism. An overview of sugar metabolism and how these sugars enter glycolysis.
 Hexose Metabolism An overview of sugar metabolism and how these sugars enter glycolysis. See chapter 15 of Fundamentals of Biochemisty: Life at the Molecular Level, 4 th Ed by Voet, Voet, and Pratt. Overview
Hexose Metabolism An overview of sugar metabolism and how these sugars enter glycolysis. See chapter 15 of Fundamentals of Biochemisty: Life at the Molecular Level, 4 th Ed by Voet, Voet, and Pratt. Overview
Photosynthesis in chloroplasts CO2 + H2O. Cellular respiration in mitochondria ATP. powers most cellular work. Heat energy
 Figure 9-01 LE 9-2 Light energy ECOSYSTEM Photosynthesis in chloroplasts CO2 + H2O Cellular respiration in mitochondria Organic + O molecules 2 powers most cellular work Heat energy LE 9-UN161a becomes
Figure 9-01 LE 9-2 Light energy ECOSYSTEM Photosynthesis in chloroplasts CO2 + H2O Cellular respiration in mitochondria Organic + O molecules 2 powers most cellular work Heat energy LE 9-UN161a becomes
Cellular Respiration: Harvesting Chemical Energy
 Chapter 9 Cellular Respiration: Harvesting Chemical Energy PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with
Chapter 9 Cellular Respiration: Harvesting Chemical Energy PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with
Biology 638 Biochemistry II Exam-1
 Biology 638 Biochemistry II Exam-1 Using the following values, answer questions 1-3. ATP + H 2 O ADP + P i ΔG = -30 kj/mol Creatine-phosphate + H 2 O Creatine + P i ΔG = -12 kj/mol ½O 2 + 2H + + 2e - H
Biology 638 Biochemistry II Exam-1 Using the following values, answer questions 1-3. ATP + H 2 O ADP + P i ΔG = -30 kj/mol Creatine-phosphate + H 2 O Creatine + P i ΔG = -12 kj/mol ½O 2 + 2H + + 2e - H
Biochemistry: A Short Course
 Tymoczko Berg Stryer Biochemistry: A Short Course Second Edition CHAPTER 16 Glycolysis 2013 W. H. Freeman and Company Chapter 16 Outline Why is glucose such a prominent fuel in all life forms? 1. Glucose
Tymoczko Berg Stryer Biochemistry: A Short Course Second Edition CHAPTER 16 Glycolysis 2013 W. H. Freeman and Company Chapter 16 Outline Why is glucose such a prominent fuel in all life forms? 1. Glucose
number Done by Corrected by Doctor Nayef Karadsheh
 number 11 Done by حسام أبو عوض Corrected by Moayyad Al-Shafei Doctor Nayef Karadsheh 1 P a g e General Regulatory Aspects in Metabolism: We can divide all pathways in metabolism to catabolicand anabolic.
number 11 Done by حسام أبو عوض Corrected by Moayyad Al-Shafei Doctor Nayef Karadsheh 1 P a g e General Regulatory Aspects in Metabolism: We can divide all pathways in metabolism to catabolicand anabolic.
Cellular Respiration Stage 1: Glycolysis (Ch. 6)
 Cellular Respiration Stage 1: Glycolysis (Ch. 6) What s the point? The point is to make! 2007-2008 Harvesting stored energy Energy is stored in organic molecules carbohydrates, fats, proteins Heterotrophs
Cellular Respiration Stage 1: Glycolysis (Ch. 6) What s the point? The point is to make! 2007-2008 Harvesting stored energy Energy is stored in organic molecules carbohydrates, fats, proteins Heterotrophs
Products List We have more enzymes other than listed here. Contact us freely.
 Glycolysis and Ethanol Fermentation Volume EC No. Thermostability Example of use product Glucokinase YK1 glucose 7.5~11.0 1 ml 2.7.1.2 phosphorylation Stable after incubation at 80 o C for 60 min GLK-97-01
Glycolysis and Ethanol Fermentation Volume EC No. Thermostability Example of use product Glucokinase YK1 glucose 7.5~11.0 1 ml 2.7.1.2 phosphorylation Stable after incubation at 80 o C for 60 min GLK-97-01
Carbohydrate Metabolism I
 Carbohydrate Metabolism I Outline Glycolysis Stages of glycolysis Regulation of Glycolysis Carbohydrate Metabolism Overview Enzyme Classification Dehydrogenase - oxidizes substrate using cofactors as
Carbohydrate Metabolism I Outline Glycolysis Stages of glycolysis Regulation of Glycolysis Carbohydrate Metabolism Overview Enzyme Classification Dehydrogenase - oxidizes substrate using cofactors as
CHEM121 Unit 2: Carbohydrate Metabolism
 CHEM121 Unit 2: Carbohydrate Metabolism Lecture 3 At the end of the lecture, students should be able to: Define metabolism Discuss the structure and function of ATP in metabolism Discuss glycolysis in
CHEM121 Unit 2: Carbohydrate Metabolism Lecture 3 At the end of the lecture, students should be able to: Define metabolism Discuss the structure and function of ATP in metabolism Discuss glycolysis in
ANSC 689 PHYSIOLOGICAL CHEMISTRY OF LIVESTOCK SPECIDS. Enzyme Kinetics and Control Reactions
 Handout Enzyme Kinetics and Control Reactions ANSC 689 PHYSIOLOGICAL CHEMISTRY OF LIVESTOCK SPECIDS Enzyme Kinetics and Control Reactions I. Kinetics A. Reaction rates 1. First order (reaction rate is
Handout Enzyme Kinetics and Control Reactions ANSC 689 PHYSIOLOGICAL CHEMISTRY OF LIVESTOCK SPECIDS Enzyme Kinetics and Control Reactions I. Kinetics A. Reaction rates 1. First order (reaction rate is
Fate of glucose in living systems. Glycolysis: Derived from Greek words; Glucose + 6O 2 = 6CO 2 + 6H 2 O δg o = kj/mol
 Glycolysis: Derived from Greek words; Glykys = Sweet, Lysis = splitting During this process one molecule of glucose (6 carbon molecule) is degraded into two molecules of pyruvate (three carbon molecule).
Glycolysis: Derived from Greek words; Glykys = Sweet, Lysis = splitting During this process one molecule of glucose (6 carbon molecule) is degraded into two molecules of pyruvate (three carbon molecule).
Carbohydrate. Metabolism
 Carbohydrate Metabolism Dietary carbohydrates (starch, glycogen, sucrose, lactose Mouth salivary amylase Summary of Carbohydrate Utilization Utilization for energy (glycolysis) ligosaccharides and disaccharides
Carbohydrate Metabolism Dietary carbohydrates (starch, glycogen, sucrose, lactose Mouth salivary amylase Summary of Carbohydrate Utilization Utilization for energy (glycolysis) ligosaccharides and disaccharides
Part III => METABOLISM and ENERGY. 3.2 Glucose Catabolism 3.2a Glycolysis Pathway 3.2b Glycolysis Regulation 3.2c Fermentation
 Part III => METABOLISM and ENERGY 3.2 Glucose Catabolism 3.2a Glycolysis Pathway 3.2b Glycolysis Regulation 3.2c Fermentation Section 3.2a: Glycolysis Synopsis 3.2a - Dietary starch (eg bread, rice and
Part III => METABOLISM and ENERGY 3.2 Glucose Catabolism 3.2a Glycolysis Pathway 3.2b Glycolysis Regulation 3.2c Fermentation Section 3.2a: Glycolysis Synopsis 3.2a - Dietary starch (eg bread, rice and
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
Genes of glycolysis are ubiquitously overexpressed in 24 cancer classes
 Genomics 84 (2004) 1014 1020 www.elsevier.com/locate/ygeno Genes of glycolysis are ubiquitously overexpressed in 24 cancer classes B. Altenberg a, *, K.. Greulich b a Bioinformatics Group, European Molecular
Genomics 84 (2004) 1014 1020 www.elsevier.com/locate/ygeno Genes of glycolysis are ubiquitously overexpressed in 24 cancer classes B. Altenberg a, *, K.. Greulich b a Bioinformatics Group, European Molecular
Meta-analysis of prostate cancer gene expression data identifies a novel discriminatory signature enriched for glycosylating enzymes
 Meta-analysis of prostate cancer gene expression data identifies a novel discriminatory signature enriched for glycosylating enzymes Barfeld, S. J., East, P., Zuber, V., & Mills, I. G. (2014). Meta-analysis
Meta-analysis of prostate cancer gene expression data identifies a novel discriminatory signature enriched for glycosylating enzymes Barfeld, S. J., East, P., Zuber, V., & Mills, I. G. (2014). Meta-analysis
Integration of Metabolism
 Integration of Metabolism Metabolism is a continuous process. Thousands of reactions occur simultaneously in order to maintain homeostasis. It ensures a supply of fuel, to tissues at all times, in fed
Integration of Metabolism Metabolism is a continuous process. Thousands of reactions occur simultaneously in order to maintain homeostasis. It ensures a supply of fuel, to tissues at all times, in fed
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
TEB. Id4 p63 DAPI Merge. Id4 CK8 DAPI Merge
 a Duct TEB b Id4 p63 DAPI Merge Id4 CK8 DAPI Merge c d e Supplementary Figure 1. Identification of Id4-positive MECs and characterization of the Comma-D model. (a) IHC analysis of ID4 expression in the
a Duct TEB b Id4 p63 DAPI Merge Id4 CK8 DAPI Merge c d e Supplementary Figure 1. Identification of Id4-positive MECs and characterization of the Comma-D model. (a) IHC analysis of ID4 expression in the
Tema 10A: Fermentations. Chapter 14 and Chapter 8 Pages
 Tema 10A: Fermentations hapter 14 and hapter 8 Pages 383-402 Lactic Acid Bacteria haracteristics: Gram positive, carbohydrate users, proteolysis rare, nonmotile, non-spore forming Strict fermentors, unable
Tema 10A: Fermentations hapter 14 and hapter 8 Pages 383-402 Lactic Acid Bacteria haracteristics: Gram positive, carbohydrate users, proteolysis rare, nonmotile, non-spore forming Strict fermentors, unable
Metabolism. Metabolic pathways. BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Metabolism Metabolism is the chemical change of
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Metabolism Metabolism is the chemical change of
Glucose is the only source of energy in red blood cells. Under starvation conditions ketone bodies become a source of energy for the brain
 Glycolysis 4 / The Text :- Some Points About Glucose Glucose is very soluble source of quick and ready energy. It is a relatively stable and easily transported. In mammals, the brain uses only glucose
Glycolysis 4 / The Text :- Some Points About Glucose Glucose is very soluble source of quick and ready energy. It is a relatively stable and easily transported. In mammals, the brain uses only glucose
Supplementary Material
 Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Dr. DerVartanian is ill and will likely not be able to give lectures this week.
 Dr. DerVartanian is ill and will likely not be able to give lectures this week. Today s slides will be put on-line today, and are designed to introduce you to glycolysis. You should use these slides, along
Dr. DerVartanian is ill and will likely not be able to give lectures this week. Today s slides will be put on-line today, and are designed to introduce you to glycolysis. You should use these slides, along
BCH 4054 Chapter 19 Lecture Notes
 BCH 4054 Chapter 19 Lecture Notes 1 Chapter 19 Glycolysis 2 aka = also known as verview of Glycolysis aka The Embden-Meyerhoff Pathway First pathway discovered Common to almost all living cells ccurs in
BCH 4054 Chapter 19 Lecture Notes 1 Chapter 19 Glycolysis 2 aka = also known as verview of Glycolysis aka The Embden-Meyerhoff Pathway First pathway discovered Common to almost all living cells ccurs in
AP VP DLP H&E. p-akt DLP
 A B AP VP DLP H&E AP AP VP DLP p-akt wild-type prostate PTEN-null prostate Supplementary Fig. 1. Targeted deletion of PTEN in prostate epithelium resulted in HG-PIN in all three lobes. (A) The anatomy
A B AP VP DLP H&E AP AP VP DLP p-akt wild-type prostate PTEN-null prostate Supplementary Fig. 1. Targeted deletion of PTEN in prostate epithelium resulted in HG-PIN in all three lobes. (A) The anatomy
Enzymatic Assay of FRUCTOSE-6-PHOSPHATE KINASE, PYROPHOSPHATE DEPENDENT (EC ) from Mung Bean
 PRINCIPLE: PP i + F-6-P PP i -PFK > F-1,6-DP + P i F-2,6-DP 1 F-1,6-DP Aldolase > GAP + DHAP GAP TPI > DHAP 2DHAP + 2 ß-NADH GDH > 2 Glycerol-3-Phosphate + 2 ß-NAD Abbreviations used: PP i = Pyrophosphate
PRINCIPLE: PP i + F-6-P PP i -PFK > F-1,6-DP + P i F-2,6-DP 1 F-1,6-DP Aldolase > GAP + DHAP GAP TPI > DHAP 2DHAP + 2 ß-NADH GDH > 2 Glycerol-3-Phosphate + 2 ß-NAD Abbreviations used: PP i = Pyrophosphate
Probe. Hind III Q,!?R'!! /0!!!!D1"?R'! vector. Homologous recombination
 Supple-Zhang Page 1 Wild-type locus Targeting construct Targeted allele Exon Exon3 Exon Probe P1 P P3 FRT FRT loxp loxp neo vector amh I Homologous recombination neo P1 P P3 FLPe recombination Q,!?R'!!
Supple-Zhang Page 1 Wild-type locus Targeting construct Targeted allele Exon Exon3 Exon Probe P1 P P3 FRT FRT loxp loxp neo vector amh I Homologous recombination neo P1 P P3 FLPe recombination Q,!?R'!!
Protein SD Units (P-value) Cluster order
 SUPPLEMENTAL TABLE AND FIGURES Table S1. Signature Phosphoproteome of CD22 E12 Transgenic Mouse BPL Cells. T-test vs. Other Protein SD Units (P-value) Cluster order ATPase (Ab-16) 1.41 0.000880 1 mtor
SUPPLEMENTAL TABLE AND FIGURES Table S1. Signature Phosphoproteome of CD22 E12 Transgenic Mouse BPL Cells. T-test vs. Other Protein SD Units (P-value) Cluster order ATPase (Ab-16) 1.41 0.000880 1 mtor
Portal module: m Glycolysis. First Last. 1 First Half of Glycolysis (Energy-Requiring Steps)
 Portal module: m10399 1 Glycolysis First Last This work is produced by Portal and licensed under the Creative Commons Attribution License 4.0 Abstract By the end of this section, you will be able to do
Portal module: m10399 1 Glycolysis First Last This work is produced by Portal and licensed under the Creative Commons Attribution License 4.0 Abstract By the end of this section, you will be able to do
Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition
 Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition Andrew J Massey Supplementary Information Supplementary Figure S1. Related to Fig. 1. (a) HT29 or U2OS cells
Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition Andrew J Massey Supplementary Information Supplementary Figure S1. Related to Fig. 1. (a) HT29 or U2OS cells
Yield of energy from glucose
 Paper : Module : 05 Yield of Energy from Glucose Principal Investigator, Paper Coordinator and Content Writer Prof. Ramesh Kothari, Professor Dept. of Biosciences, Saurashtra University, Rajkot - 360005
Paper : Module : 05 Yield of Energy from Glucose Principal Investigator, Paper Coordinator and Content Writer Prof. Ramesh Kothari, Professor Dept. of Biosciences, Saurashtra University, Rajkot - 360005
Journal of Cell Science Supplementary information. Arl8b +/- Arl8b -/- Inset B. electron density. genotype
 J. Cell Sci. : doi:.4/jcs.59: Supplementary information E9. A Arl8b /- Arl8b -/- Arl8b Arl8b non-specific band Gapdh Tbp E7.5 HE Inset B D Control al am hf C E Arl8b -/- al am hf E8.5 F low middle high
J. Cell Sci. : doi:.4/jcs.59: Supplementary information E9. A Arl8b /- Arl8b -/- Arl8b Arl8b non-specific band Gapdh Tbp E7.5 HE Inset B D Control al am hf C E Arl8b -/- al am hf E8.5 F low middle high
Chemistry Chapter 28
 hemistry 2100 hapter 28 arbohydrate atabolism glycolysis: glucose pyruvate acetyl TA ycle: acetyl 2 + NAD / FAD 2 oxidative phosphorylation: NAD / FAD 2 ATP Glycolysis ATP ADP isomerase hexokin ase Mg
hemistry 2100 hapter 28 arbohydrate atabolism glycolysis: glucose pyruvate acetyl TA ycle: acetyl 2 + NAD / FAD 2 oxidative phosphorylation: NAD / FAD 2 ATP Glycolysis ATP ADP isomerase hexokin ase Mg
respiration mitochondria mitochondria metabolic pathways reproduction can fuse or split DRP1 interacts with ER tubules chapter DRP1 ER tubule
 mitochondria respiration chapter 3-4 shape highly variable can fuse or split structure outer membrane inner membrane cristae intermembrane space mitochondrial matrix free ribosomes respiratory enzymes
mitochondria respiration chapter 3-4 shape highly variable can fuse or split structure outer membrane inner membrane cristae intermembrane space mitochondrial matrix free ribosomes respiratory enzymes
Ch. 9 Cellular Respira,on BIOL 222
 Ch. 9 Cellular Respira,on BIOL Energy Arrives as sunlight Photosynthesis Energy ECOSYSTEM Light energy Plants capture sunlight organic molecules and generates O Carbs used in cellular respira@on CO + H
Ch. 9 Cellular Respira,on BIOL Energy Arrives as sunlight Photosynthesis Energy ECOSYSTEM Light energy Plants capture sunlight organic molecules and generates O Carbs used in cellular respira@on CO + H
This is an example outline of 3 lectures in BSC (Thanks to Dr. Ellington for sharing this information.)
 This is an example outline of 3 lectures in BSC 2010. (Thanks to Dr. Ellington for sharing this information.) Topic 10: CELLULAR RESPIRATION (lectures 14-16) OBJECTIVES: 1. Know the basic reactions that
This is an example outline of 3 lectures in BSC 2010. (Thanks to Dr. Ellington for sharing this information.) Topic 10: CELLULAR RESPIRATION (lectures 14-16) OBJECTIVES: 1. Know the basic reactions that
Respiration. Organisms can be classified based on how they obtain energy: Autotrophs
 Respiration rganisms can be classified based on how they obtain energy: Autotrophs Able to produce their own organic molecules through photosynthesis Heterotrophs Live on organic compounds produced by
Respiration rganisms can be classified based on how they obtain energy: Autotrophs Able to produce their own organic molecules through photosynthesis Heterotrophs Live on organic compounds produced by
I tried to put as many questions as possible, but unfortunately only answers were found without the questions.
 I tried to put as many questions as possible, but unfortunately only answers were found without the questions. These are some questions from doctor2015 med exam : 1. One of them isn t acute phase protein
I tried to put as many questions as possible, but unfortunately only answers were found without the questions. These are some questions from doctor2015 med exam : 1. One of them isn t acute phase protein
I tried to put as many questions as possible, but unfortunately only answers were found without the questions.
 I tried to put as many questions as possible, but unfortunately only answers were found without the questions. These are some questions from doctor2015 med exam : 1. One of them isn t acute phase protein
I tried to put as many questions as possible, but unfortunately only answers were found without the questions. These are some questions from doctor2015 med exam : 1. One of them isn t acute phase protein
III. Metabolism - Gluconeogenesis
 Department of Chemistry and Biochemistry University of Lethbridge III. Metabolism - Gluconeogenesis Carl & Gertrude Cori Slide 1 Carbohydrate Synthesis Lactate, pyruvate and glycerol are the important
Department of Chemistry and Biochemistry University of Lethbridge III. Metabolism - Gluconeogenesis Carl & Gertrude Cori Slide 1 Carbohydrate Synthesis Lactate, pyruvate and glycerol are the important
Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction. Marker Sequence (5 3 ) Accession No.
 Supplementary Tables Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction Marker Sequence (5 3 ) Accession No. Angiopoietin 1, ANGPT1 A CCCTCCGGTGAATATTGGCTGG NM_001146.3
Supplementary Tables Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction Marker Sequence (5 3 ) Accession No. Angiopoietin 1, ANGPT1 A CCCTCCGGTGAATATTGGCTGG NM_001146.3
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
Glycolysis. Color index: Doctors slides Notes and explanations Extra information Highlights. Biochemistry Team 437
 Glycolysis Color index: Doctors slides Notes and explanations Extra information Highlights Biochemistry Team 437 ﺑ ﺳ م ﷲ اﻟرﺣﻣن اﻟرﺣﯾم Objectives: Recognize glycolysis as the major oxidative pathway of
Glycolysis Color index: Doctors slides Notes and explanations Extra information Highlights Biochemistry Team 437 ﺑ ﺳ م ﷲ اﻟرﺣﻣن اﻟرﺣﯾم Objectives: Recognize glycolysis as the major oxidative pathway of
Supplementary information
 Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
TIGAR's promiscuity Bolaños, Juan P.
 TIGAR's promiscuity Bolaños, Juan P. TIGAR [TP53 (tumour protein 53)-induced glycolysis and apoptosis regulator] is an important survival factor for cancer cells. The enzymatic activity supported by sequence
TIGAR's promiscuity Bolaños, Juan P. TIGAR [TP53 (tumour protein 53)-induced glycolysis and apoptosis regulator] is an important survival factor for cancer cells. The enzymatic activity supported by sequence
Nature Immunology: doi: /ni Supplementary Figure 1. Huwe1 has high expression in HSCs and is necessary for quiescence.
 Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
incorporation into fructose-6-phosphate and glucose-6-phosphate (F6P/G6P); 2-
 Relative abundance (%) Relative abundance (%) Relative abundance (%) Relative abundance (%) 100% 80% 60% 40% F6P / G6P M+0 M+1 M+2 M+3 M+4 M+5 M+6 100% 80% 60% 40% 2PG / 3PG M+0 M+1 M+2 M+3 20% 20% 0%
Relative abundance (%) Relative abundance (%) Relative abundance (%) Relative abundance (%) 100% 80% 60% 40% F6P / G6P M+0 M+1 M+2 M+3 M+4 M+5 M+6 100% 80% 60% 40% 2PG / 3PG M+0 M+1 M+2 M+3 20% 20% 0%
Supplemental Figure S1A Notch1
 Supplemental Figure S1A Notch1 erage) epth of Cove ormalized De Log1(No Notch exons Figure S1: A) Relative coverage of Notch1 and Notch 2 exons in HCC2218, HCC1187, MB157, MDA-MB157 cell lines. Blue color
Supplemental Figure S1A Notch1 erage) epth of Cove ormalized De Log1(No Notch exons Figure S1: A) Relative coverage of Notch1 and Notch 2 exons in HCC2218, HCC1187, MB157, MDA-MB157 cell lines. Blue color
of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed.
 Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Fig. S1. Dose-response effects of acute administration of the β3 adrenoceptor agonists CL316243, BRL37344, ICI215,001, ZD7114, ZD2079 and CGP12177 at
 Fig. S1. Dose-response effects of acute administration of the β3 adrenoceptor agonists CL316243, BRL37344, ICI215,001, ZD7114, ZD2079 and CGP12177 at doses of 0.1, 0.5 and 1 mg/kg on cumulative food intake
Fig. S1. Dose-response effects of acute administration of the β3 adrenoceptor agonists CL316243, BRL37344, ICI215,001, ZD7114, ZD2079 and CGP12177 at doses of 0.1, 0.5 and 1 mg/kg on cumulative food intake
Deregulation of signal transduction and cell cycle in Cancer
 Deregulation of signal transduction and cell cycle in Cancer Tuangporn Suthiphongchai, Ph.D. Department of Biochemistry Faculty of Science, Mahidol University Email: tuangporn.sut@mahidol.ac.th Room Pr324
Deregulation of signal transduction and cell cycle in Cancer Tuangporn Suthiphongchai, Ph.D. Department of Biochemistry Faculty of Science, Mahidol University Email: tuangporn.sut@mahidol.ac.th Room Pr324
Supplementary Figure 1: Expression of NFAT proteins in Nfat2-deleted B cells (a+b) Protein expression of NFAT2 (a) and NFAT1 (b) in isolated splenic
 Supplementary Figure 1: Expression of NFAT proteins in Nfat2-deleted B cells (a+b) Protein expression of NFAT2 (a) and NFAT1 (b) in isolated splenic B cells from WT Nfat2 +/+, TCL1 Nfat2 +/+ and TCL1 Nfat2
Supplementary Figure 1: Expression of NFAT proteins in Nfat2-deleted B cells (a+b) Protein expression of NFAT2 (a) and NFAT1 (b) in isolated splenic B cells from WT Nfat2 +/+, TCL1 Nfat2 +/+ and TCL1 Nfat2
Supplementary Information
 Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
CHE 242 Exam 3 Practice Questions
 CHE 242 Exam 3 Practice Questions Glucose metabolism 1. Below is depicted glucose catabolism. Indicate on the pathways the following: A) which reaction(s) of glycolysis are irreversible B) where energy
CHE 242 Exam 3 Practice Questions Glucose metabolism 1. Below is depicted glucose catabolism. Indicate on the pathways the following: A) which reaction(s) of glycolysis are irreversible B) where energy
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2566 Figure S1 CDKL5 protein expression pattern and localization in mouse brain. (a) Multiple-tissue western blot from a postnatal day (P) 21 mouse probed with an antibody against CDKL5.
DOI: 10.1038/ncb2566 Figure S1 CDKL5 protein expression pattern and localization in mouse brain. (a) Multiple-tissue western blot from a postnatal day (P) 21 mouse probed with an antibody against CDKL5.
Deletion of tyrosine phosphatase Shp2 in Sertoli cells causes infertility. in mice
 Deletion of tyrosine phosphatase Shp2 in Sertoli cells causes infertility in mice Xiaopeng Hu 1 *, Zhenzhou Tang 1,2 *, Yang Li 2 *, Wensheng Liu 2, Shuang Zhang 3, Bingyan Wang 3, Yingpu Tian 2, Yinan
Deletion of tyrosine phosphatase Shp2 in Sertoli cells causes infertility in mice Xiaopeng Hu 1 *, Zhenzhou Tang 1,2 *, Yang Li 2 *, Wensheng Liu 2, Shuang Zhang 3, Bingyan Wang 3, Yingpu Tian 2, Yinan
Simulation of Metabolic Pathway Model Using Petri Net Approach
 2012 International Conference on Bioscience, Biochemistry and Bioinformatics IPCBEE vol.3 1(2012) (2012)IACSIT Press, Singapoore Simulation of Metabolic Pathway Model Using Petri Net Approach Rose Mary
2012 International Conference on Bioscience, Biochemistry and Bioinformatics IPCBEE vol.3 1(2012) (2012)IACSIT Press, Singapoore Simulation of Metabolic Pathway Model Using Petri Net Approach Rose Mary
atty Acid Oxidation (β-oxidation)( in Mitochondria Net Result of Fatty Acid Oxidation Pathway Fatty acid shortened by 2 carbon unit
 ecture 14 10/3/05 ellular Respiration: arvesting hemical Energy hapter 9 II. atabolic Pathways Lecture utline 1. Review Fatty Acid xidation Pathway. xidation of Monosaccharides ( Pathway) 3. Mitochondrial
ecture 14 10/3/05 ellular Respiration: arvesting hemical Energy hapter 9 II. atabolic Pathways Lecture utline 1. Review Fatty Acid xidation Pathway. xidation of Monosaccharides ( Pathway) 3. Mitochondrial
Supplementary Figure 1. DJ-1 modulates ROS concentration in mouse skeletal muscle.
 Supplementary Figure 1. DJ-1 modulates ROS concentration in mouse skeletal muscle. (a) mrna levels of Dj1 measured by quantitative RT-PCR in soleus, gastrocnemius (Gastroc.) and extensor digitorum longus
Supplementary Figure 1. DJ-1 modulates ROS concentration in mouse skeletal muscle. (a) mrna levels of Dj1 measured by quantitative RT-PCR in soleus, gastrocnemius (Gastroc.) and extensor digitorum longus
Synthetic non-oxidative glycolysis (NOG) enables complete carbon conservation
 Synthetic non-oxidative glycolysis (NOG) enables complete carbon conservation Igor W. Bogorad, Tzu-Shyang Lin, James C. Liao Nature, vol 502, 2013 Presented by Elise Jyränoja, Kellen Leskinen and Tia Korhonen
Synthetic non-oxidative glycolysis (NOG) enables complete carbon conservation Igor W. Bogorad, Tzu-Shyang Lin, James C. Liao Nature, vol 502, 2013 Presented by Elise Jyränoja, Kellen Leskinen and Tia Korhonen
(A) RT-PCR for components of the Shh/Gli pathway in normal fetus cell (MRC-5) and a
 Supplementary figure legends Supplementary Figure 1. Expression of Shh signaling components in a panel of gastric cancer. (A) RT-PCR for components of the Shh/Gli pathway in normal fetus cell (MRC-5) and
Supplementary figure legends Supplementary Figure 1. Expression of Shh signaling components in a panel of gastric cancer. (A) RT-PCR for components of the Shh/Gli pathway in normal fetus cell (MRC-5) and
Supplementary methods:
 Supplementary methods: Primers sequences used in real-time PCR analyses: β-actin F: GACCTCTATGCCAACACAGT β-actin [11] R: AGTACTTGCGCTCAGGAGGA MMP13 F: TTCTGGTCTTCTGGCACACGCTTT MMP13 R: CCAAGCTCATGGGCAGCAACAATA
Supplementary methods: Primers sequences used in real-time PCR analyses: β-actin F: GACCTCTATGCCAACACAGT β-actin [11] R: AGTACTTGCGCTCAGGAGGA MMP13 F: TTCTGGTCTTCTGGCACACGCTTT MMP13 R: CCAAGCTCATGGGCAGCAACAATA
OVERVIEW OF THE GLYCOLYTIC PATHWAY Glycolysis is considered one of the core metabolic pathways in nature for three primary reasons:
 Glycolysis 1 Supplemental Reading Key Concepts - Overview of the Glycolytic Pathway Glycolysis generates a small amount of ATP Preview of the ten enzyme-catalyzed reactions of glycolysis - Stage 1: ATP
Glycolysis 1 Supplemental Reading Key Concepts - Overview of the Glycolytic Pathway Glycolysis generates a small amount of ATP Preview of the ten enzyme-catalyzed reactions of glycolysis - Stage 1: ATP
ATP ATP. Cellular Respiration Harvesting Chemical Energy. The point is to make ATP!
 ellular Respiration Harvesting hemical Energy 1 The point is to make! 2 Harvesting stored energy Energy is stored in organic molecules carbohydrates, fats, proteins Heterotrophs eat these organic molecules
ellular Respiration Harvesting hemical Energy 1 The point is to make! 2 Harvesting stored energy Energy is stored in organic molecules carbohydrates, fats, proteins Heterotrophs eat these organic molecules
CHAPTER 24: Carbohydrate, Lipid, & Protein Metabolism. General, Organic, & Biological Chemistry Janice Gorzynski Smith
 CHAPTER 24: Carbohydrate, Lipid, & Protein Metabolism General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 24: Carbohydrate, Lipid, & Protein Metabolism Learning Objectives: q Role in
CHAPTER 24: Carbohydrate, Lipid, & Protein Metabolism General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 24: Carbohydrate, Lipid, & Protein Metabolism Learning Objectives: q Role in
Glycolysis 10/26/2009. Glycolysis I 11/03/09. Historical perspective. Pathway overview
 Glycolysis Glycolysis I 11/03/09 The conversion of glucose to pyruvate to yield 2ATP molecules 10 enzymatic steps hemical interconversion steps Mechanisms of enzyme conversion and intermediates Energetics
Glycolysis Glycolysis I 11/03/09 The conversion of glucose to pyruvate to yield 2ATP molecules 10 enzymatic steps hemical interconversion steps Mechanisms of enzyme conversion and intermediates Energetics
PRINT your Name Student (FAMILY, first name) Midterm 7:00 P.M.
 PRINT your Name Student No. (FAMILY, first name) BIOCHEMISTRY 311A VERSION 1 (ONE) Midterm 7:00 P.M. Examiners: Dr. R. E. MacKenzie (69%) Dr. A. Storer (18%) Dr. W. Mushynski (13%) READ THE QUESTIONS CAREFULLY!!
PRINT your Name Student No. (FAMILY, first name) BIOCHEMISTRY 311A VERSION 1 (ONE) Midterm 7:00 P.M. Examiners: Dr. R. E. MacKenzie (69%) Dr. A. Storer (18%) Dr. W. Mushynski (13%) READ THE QUESTIONS CAREFULLY!!
HepaRG LX2. HepaRG HepaRG LX2 LX2
 C Supporting Figure 1. Experimental design of s between and cells. (A) -hepatocytes were isolated from a 30 days of -progenitors. Differentiation into mature hepatocytes was achieved following a 2-weeks
C Supporting Figure 1. Experimental design of s between and cells. (A) -hepatocytes were isolated from a 30 days of -progenitors. Differentiation into mature hepatocytes was achieved following a 2-weeks
Physiological and Metabolic Background of Speed Training. Loren Seagrave, Director of Track & Field and Speed and Movement, IMG Academy
 Physiological and Metabolic Background of Speed Training Loren Seagrave, Director of Track & Field and Speed and Movement, IMG Academy 1 DISCLAIMER» I am not a scientist. - but I have spent considerable
Physiological and Metabolic Background of Speed Training Loren Seagrave, Director of Track & Field and Speed and Movement, IMG Academy 1 DISCLAIMER» I am not a scientist. - but I have spent considerable
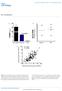 DOI: 10.1038/ncb2210 b. ICAM1 ng ml -1 P = 0.0001 Small RNA (15-30nts) ng ml -1 Cell Lysate Exosome HDL Plasma HDL Normal Human HDL mirnas R = 0.45 P < 0.0001 Normal Human Exosome mirnas Figure S1. Characterization
DOI: 10.1038/ncb2210 b. ICAM1 ng ml -1 P = 0.0001 Small RNA (15-30nts) ng ml -1 Cell Lysate Exosome HDL Plasma HDL Normal Human HDL mirnas R = 0.45 P < 0.0001 Normal Human Exosome mirnas Figure S1. Characterization
Notes CELLULAR RESPIRATION SUMMARY EQUATION C 6 H 12 O 6 + O 2. 6CO 2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION
 AP BIOLOGY CELLULAR ENERGETICS ACTIVITY #2 Notes NAME DATE HOUR SUMMARY EQUATION CELLULAR RESPIRATION C 6 H 12 O 6 + O 2 6CO 2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION Oxidation: partial or complete
AP BIOLOGY CELLULAR ENERGETICS ACTIVITY #2 Notes NAME DATE HOUR SUMMARY EQUATION CELLULAR RESPIRATION C 6 H 12 O 6 + O 2 6CO 2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION Oxidation: partial or complete
