Supplemental File. TRAF6 is an amplified oncogene bridging the Ras and nuclear factor-κb cascade in human lung cancer
|
|
|
- Dorothy Davidson
- 5 years ago
- Views:
Transcription
1 Supplemental File TRAF6 is an amplified oncogene bridging the Ras and nuclear factor-κb cascade in human lung cancer Daniel T. Starczynowski, William W. Lockwood, Sophie Delehouzee, Raj Chari, Joanna Wegrzyn, Megan Fuller, Ming-Sound Tsao, Stephen Lam, Adi F. Gazdar, Wan L. Lam, and Aly Karsan Supplemental Table 1 Supplemental Table 2 Supplemental Table 3 Supplemental Figure 1 Supplemental Figure 2 Supplemental Figure 3 Supplemental Figure 4
2 Supplemental Table 1. Candidate genes within the minimally amplified region on 11p Gene BP Start BP End Corrected p-value n-log 2 (p-value) WT WIT CCDC PRRG QSER DEPDC TCP11L HIPK C11orf CD FBXO LMO NAT ABTB CAT ELF EHF APIP PDHX NA CD SLC1A DKFZP586H FJX TRIM COMMD FLJ TRAF RAG RAG C11orf
3 Supplemental Table 2. Lung cancer cell lines. Name ATCC Number Classifier amp 11p13* A549 CCL-185 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) Yes Calu-1 HTB-54 Epidermoid Carcinoma Calu-3 HTB-55 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) Calu-6 HTB-56 Anaplastic Carcinoma EKVX N/A Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC), NCI-60 H1155 CRL-5818 Non-Small Cell Lung Cancer (NSCLC), Large Cell Carcinoma (LCC) H1184 CRL-5858 Small Cell Lung Cancer (SCLC) H1299 CRL-5803 Non-Small Cell Lung Cancer (NSCLC), Large Cell Carcinoma (LCC) H1355 CRL-5865 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H1395 CRL-5868 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H1437 CRL-5872 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H146 HTB-173 Small Cell Lung Cancer (SCLC) H157 CRL-5802 Non-Small Cell Lung Cancer (NSCLC), Squamous Cell Carcinoma (SqCC) Yes H1607 N/A Small Cell Lung Cancer (SCLC) H1648 CRL-5882 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H1650 CRL-5883 Non-Small Cell Lung Cancer (NSCLC), Bronchioalveolar carcinoma (BAC) Yes H1666 CRL-5885 Non-Small Cell Lung Cancer (NSCLC), Bronchioalveolar carcinoma (BAC) H1672 CRL-5886 Small Cell Lung Cancer (SCLC) H1755 CRL-5892 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H1770 CRL-5893 Neuroendocrine H1781 CRL-5894 Non-Small Cell Lung Cancer (NSCLC), Bronchioalveolar carcinoma (BAC) H1819 CRL-5897 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H187 CRL-5804 Small Cell Lung Cancer (SCLC) H1975 CRL-5908 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H1993 CRL-5909 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H2009 CRL-5911 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H2052 CRL-5915 Mesothelioma (ME) H2073 CRL-5918 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H2085 CRL-5921 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H2087 CRL-5922 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H2107 CRL-5983 Small Cell Lung Cancer (SCLC) H2122 CRL-5985 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H2126 CCL-256 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H2141 CRL-5927 Small Cell Lung Cancer (SCLC) H2170 CRL-5928 Non-Small Cell Lung Cancer (NSCLC), Squamous Cell Carcinoma (SqCC) H2171 CRL-5929 Small Cell Lung Cancer (SCLC) Yes H2228 CRL-5935 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H226 CRL-5826 Non-Small Cell Lung Cancer (NSCLC), Squamous Cell Carcinoma (SqCC) H2347 CRL-5942 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H2882 N/A Non-Small Cell Lung Cancer (NSCLC) H2887 N/A Non-Small Cell Lung Cancer (NSCLC) Yes H289 N/A Small Cell Lung Cancer (SCLC) H3122 N/A Non-Small Cell Lung Cancer (NSCLC) H322 N/A Non-Small Cell Lung Cancer (NSCLC), Bronchioalveolar carcinoma (BAC) H3255 N/A Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) Yes H358 CRL-5807 Non-Small Cell Lung Cancer (NSCLC), Bronchioalveolar carcinoma (BAC) H378 CRL-5808 Small Cell Lung Cancer (SCLC) H441 HTB-174 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) H460 HTB-177 Non-Small Cell Lung Cancer (NSCLC), Large Cell Carcinoma (LCC) H520 HTB-182 Non-Small Cell Lung Cancer (NSCLC), Squamous Cell Carcinoma (SqCC) H524 CRL-5831 Small Cell Lung Cancer (SCLC) H526 CRL-5811 Small Cell Lung Cancer (SCLC) Yes H82 HTB-175 Small Cell Lung Cancer (SCLC) H820 N/A Non-Small Cell Lung Cancer (NSCLC), Bronchioalveolar carcinoma (BAC) H841 CRL-5845 Small Cell Lung Cancer (SCLC) H889 CRL-5817 Small Cell Lung Cancer (SCLC) HCC1171 N/A Non-Small Cell Lung Cancer (NSCLC) HCC1195 N/A Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) HCC1359 N/A Non-Small Cell Lung Cancer (NSCLC), Large Cell Carcinoma (LCC) HCC15 N/A Non-Small Cell Lung Cancer (NSCLC), Squamous Cell Carcinoma (SqCC) HCC1719 N/A Lung Cancer Yes HCC1833 N/A Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) HCC193 N/A Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) Yes HCC2279 N/A Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) HCC2450 N/A Non-Small Cell Lung Cancer (NSCLC), Squamous Cell Carcinoma (SqCC) HCC2935 CRL-2869 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) HCC33 N/A Small Cell Lung Cancer (SCLC) HCC366 N/A Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) Yes HCC4006 CRL-2871 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) Yes HCC4011 N/A Non-Small Cell Lung Cancer (NSCLC) HCC4017 N/A Non-Small Cell Lung Cancer (NSCLC) HCC44 N/A Non-Small Cell Lung Cancer (NSCLC) HCC461 N/A Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) Yes HCC515 N/A Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) HCC78 N/A Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) HCC827 CRL-2868 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC) HCC95 N/A Non-Small Cell Lung Cancer (NSCLC), Squamous Cell Carcinoma (SqCC) Yes HOP 62 N/A Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC), NCI-60 HOP 92 N/A Non-Small Cell Lung Cancer (NSCLC), Large Cell Carcinoma (LCC), NCI-60 NCI-H23 CRL-5800 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC), NCI-60 NCI-H522 CRL-5810 Non-Small Cell Lung Cancer (NSCLC), Adenocarcinoma (AC), NCI-60 Yes PC3 N/A Lung Cancer PC9 N/A Lung Cancer Yes SK-MES-1 HTB-58 Non-Small Cell Lung Cancer (NSCLC), Squamous Cell Carcinoma (SqCC) Yes SW900 HTB-59 Non-Small Cell Lung Cancer (NSCLC), Squamous Cell Carcinoma (SqCC) Yes N/A, not available; *, amplification includes TRAF6
4 Supplemental Table 3. Description of TRAF6 and KRAS alterations in the lung cancer cell lines.
5 Supplemental Figure 1. TRAF6 is overexpressed in lung cancer with chr 11p13 amplifications. A. The copy number status of chromosome 11p is shown for six representative lung cancer tumors (#1-6). The minimally amplified region spanning chr 11p13 (32,126,542) to chr 11p12 (37,251,933) is indicated with the arrow. Normalized log 2 signal intensity ratios were plotted and visualized by SIGMA 2 Vertical lines denote log 2 signal ratios from 1 to +1. Copy number gains are shown to the right of 0 (red). Each black dot represents a single BAC clone. B. Alignment of acgh profiles at chromosome 11p13 from four lung cancer samples. C. Expression of TRAF6 mrna (log 2 ) in lung cancer cell lines with (n = 13) and without amplification of the TRAF6 locus, chr 11p13 (n = 31) (p = ).
6 Supplemental Figure 2. Validation of TRAF6 inhibition by RNAi in lung cancer cells. Quantitative PCR (TRAF6 and GAPDH mrna) (A) and immunoblotting (TRAF6 and actin protein) (B) is shown after inhibition of TRAF6 (sht6) in the indicated cell lines.
7 Supplemental Figure 3. Inhibition of candidate genes on chromosome 11p13 in lung cancer cells. Lung cancer cells with amplification of 11p13 (HCC95) were transduced with virus encoding shrna constructs specific for CD44, COMMD9, TCP11L1, and TRAF6. Transduced cell lines were selected in puromycin and analyzed for colony formation in 0.3% soft agar and confirmation of target gene knockdown by qpcr. A. Colony numbers for the indicated transduced cell lines are shown relative to vector-transduced cells (1.0). The corresponding target gene knockdown is shown below. Colony formation of shcommd9-expressing cells may be in part explained by the less efficient knockdown of COMMD9. B. Representative colonies are shown for the indicated transduced cell lines at 20X magnification. Data is shown from one experiment performed in triplicate.p values are indicated above the histogram. Data are represented as mean ± SEM.
8 Supplemental Figure 4. Inhibition of TRAF6 suppresses breast cancer lines. A. Alignment of acgh profiles at chromosome 11p13 from 4 breast cancer samples. The arrow indicates the location of the TRAF6 locus. Normalized log 2 signal intensity ratios were plotted and visualized by SIGMA 2. Vertical lines denote log 2 signal ratios from 1 to +1. Copy number decreases are shown to the left of 0 (green). Each black dot represents a single BAC clone. B. Expression of TRAF6 mrna in breast cancer samples with (n = 2) amplification at band 11p13, as well as normal controls (n = 6). C. TRAF6 mrna from MCF7 and SKBR3 was measured by qpcr and shown relative to SKBR3 (1.0). As shown, MCF7 cells exhibit higher (two-fold) expression of TRAF6 mrna. D. MCF7 and SKBR3 were transduced with vector- or TRAF6- dominant negative (FLAG-T6DN)-containing virus. Expression of T6DN was determined by anti-flag immunoblotting. E-F. Vector- and T6DN-expressing MCF7 and SKBR3 cells were analyzed for colony formation in 0.3% soft agar. Data are represented as mean ± SEM. *, p = 0.024
TRAF6 is an amplified oncogene bridging the RAS and NF-κB pathways in human lung cancer
 TRAF6 is an amplified oncogene bridging the RAS and NF-κB pathways in human lung cancer Daniel T. Starczynowski, 1,2 William W. Lockwood, 1,2 Sophie Deléhouzée, 2 Raj Chari, 1,2 Joanna Wegrzyn, 1,2 Megan
TRAF6 is an amplified oncogene bridging the RAS and NF-κB pathways in human lung cancer Daniel T. Starczynowski, 1,2 William W. Lockwood, 1,2 Sophie Deléhouzée, 2 Raj Chari, 1,2 Joanna Wegrzyn, 1,2 Megan
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables. File Name: Peer Review File Description:
 File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables File Name: Peer Review File Description: Primer Name Sequence (5'-3') AT ( C) RT-PCR USP21 F 5'-TTCCCATGGCTCCTTCCACATGAT-3'
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables File Name: Peer Review File Description: Primer Name Sequence (5'-3') AT ( C) RT-PCR USP21 F 5'-TTCCCATGGCTCCTTCCACATGAT-3'
Supplementary Information
 Supplementary Information Lin28 Enhances Tumorigenesis and is Associated With Advanced Human Malignancies Srinivas R. Viswanathan 1, John T. Powers 1, William Einhorn 1, Yujin Hoshida 2,5, Tony Ng 15,
Supplementary Information Lin28 Enhances Tumorigenesis and is Associated With Advanced Human Malignancies Srinivas R. Viswanathan 1, John T. Powers 1, William Einhorn 1, Yujin Hoshida 2,5, Tony Ng 15,
Supplemental Information. NRF2 Is a Major Target of ARF. in p53-independent Tumor Suppression
 Molecular Cell, Volume 68 Supplemental Information NRF2 Is a Major Target of ARF in p53-independent Tumor Suppression Delin Chen, Omid Tavana, Bo Chu, Luke Erber, Yue Chen, Richard Baer, and Wei Gu Figure
Molecular Cell, Volume 68 Supplemental Information NRF2 Is a Major Target of ARF in p53-independent Tumor Suppression Delin Chen, Omid Tavana, Bo Chu, Luke Erber, Yue Chen, Richard Baer, and Wei Gu Figure
Functional genomics reveal that the serine synthesis pathway is essential in breast cancer
 Functional genomics reveal that the serine synthesis pathway is essential in breast cancer Results Presented by Stacey Lin Lloyd Lab http://www.amsbio.com/expression-ready-lentiviral-particles.aspx Overview
Functional genomics reveal that the serine synthesis pathway is essential in breast cancer Results Presented by Stacey Lin Lloyd Lab http://www.amsbio.com/expression-ready-lentiviral-particles.aspx Overview
Supplementary Information
 Supplementary Information Supplementary Figure 1. Effect of mir mimics and anti-mirs on DTPs a, Representative fluorescence microscopy images of GFP vector control or mir mimicexpressing parental and DTP
Supplementary Information Supplementary Figure 1. Effect of mir mimics and anti-mirs on DTPs a, Representative fluorescence microscopy images of GFP vector control or mir mimicexpressing parental and DTP
Supplemental Figure S1. A. Venn diagram depicting overlap between anti-correlated genes of
 Supplemental Figure S1. A. Venn diagram depicting overlap between anti-correlated genes of 1,000 most differentially expressed genes with NKX2-1 amplification in lung adenocarcinoma cell lines and anti-correlated
Supplemental Figure S1. A. Venn diagram depicting overlap between anti-correlated genes of 1,000 most differentially expressed genes with NKX2-1 amplification in lung adenocarcinoma cell lines and anti-correlated
Supplementary Information
 Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
(a) Significant biological processes (upper panel) and disease biomarkers (lower panel)
 Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. a. b. Relative cell viability. Nature Genetics: doi: /ng SCR shyap1-1 shyap
 Supplementary Figure 1. a. b. p-value for depletion in vehicle (DMSO) 1e-05 1e-03 1e-01 1 0 1000 2000 3000 4000 5000 Genes log2 normalized shrna counts in T0 0 2 4 6 8 sh1 shluc 0 2 4 6 8 log2 normalized
Supplementary Figure 1. a. b. p-value for depletion in vehicle (DMSO) 1e-05 1e-03 1e-01 1 0 1000 2000 3000 4000 5000 Genes log2 normalized shrna counts in T0 0 2 4 6 8 sh1 shluc 0 2 4 6 8 log2 normalized
supplementary information
 DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs.
 Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs. (a) CNA analysis of expression microarray data obtained from 15 tumors in the SV40Tag
Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs. (a) CNA analysis of expression microarray data obtained from 15 tumors in the SV40Tag
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A.
 Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
Functional Genomics : PTPRF-Mediated Contact Inhibition in HCC development
 Functional Genomics : PTPRF-Mediated Contact Inhibition in HCC development Sen-Yung Hsieh, MD, PhD Liver Research Unit Chang Gung Memorial Hospital Taoyuan, Taiwan Aims To identify genes controlling tumor
Functional Genomics : PTPRF-Mediated Contact Inhibition in HCC development Sen-Yung Hsieh, MD, PhD Liver Research Unit Chang Gung Memorial Hospital Taoyuan, Taiwan Aims To identify genes controlling tumor
Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3
 Supplemental Figure Legends. Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3 ErbB3 gene copy number gain. Supplemental Figure S1. ERBB3 mrna levels are elevated in
Supplemental Figure Legends. Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3 ErbB3 gene copy number gain. Supplemental Figure S1. ERBB3 mrna levels are elevated in
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Correlation between LKB1 and YAP expression in human lung cancer samples. (a) Representative photos showing LKB1 and YAP immunohistochemical staining in human
Supplementary Figures Supplementary Figure 1 Correlation between LKB1 and YAP expression in human lung cancer samples. (a) Representative photos showing LKB1 and YAP immunohistochemical staining in human
Supplementary methods:
 Supplementary methods: Primers sequences used in real-time PCR analyses: β-actin F: GACCTCTATGCCAACACAGT β-actin [11] R: AGTACTTGCGCTCAGGAGGA MMP13 F: TTCTGGTCTTCTGGCACACGCTTT MMP13 R: CCAAGCTCATGGGCAGCAACAATA
Supplementary methods: Primers sequences used in real-time PCR analyses: β-actin F: GACCTCTATGCCAACACAGT β-actin [11] R: AGTACTTGCGCTCAGGAGGA MMP13 F: TTCTGGTCTTCTGGCACACGCTTT MMP13 R: CCAAGCTCATGGGCAGCAACAATA
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
Figure S1 Time-dependent down-modulation of HER3 by EZN No Treatment. EZN-3920, 2 μm. Time, h
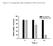 Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 DOT1L regulates the expression of epithelial and mesenchymal markers. (a) The expression levels and cellular localizations of EMT markers were confirmed by
Supplementary Figures Supplementary Figure 1 DOT1L regulates the expression of epithelial and mesenchymal markers. (a) The expression levels and cellular localizations of EMT markers were confirmed by
of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed.
 Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Supplementary Figure 1: STAT3 suppresses Kras-induced lung tumorigenesis
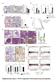 Supplementary Figure 1: STAT3 suppresses Kras-induced lung tumorigenesis (a) Immunohistochemical (IHC) analysis of tyrosine 705 phosphorylation status of STAT3 (P- STAT3) in tumors and stroma (all-time
Supplementary Figure 1: STAT3 suppresses Kras-induced lung tumorigenesis (a) Immunohistochemical (IHC) analysis of tyrosine 705 phosphorylation status of STAT3 (P- STAT3) in tumors and stroma (all-time
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH
 EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
Table S1. Primer sequences used for qrt-pcr. CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT ACTB AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT LCOR
 Table S1. Primer sequences used for qrt-pcr. ACTB LCOR KLF6 CTBP1 CDKN1A CDH1 ATF3 PLAU MMP9 TFPI2 CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT CGGCTGCAGGAAAGTTTACA
Table S1. Primer sequences used for qrt-pcr. ACTB LCOR KLF6 CTBP1 CDKN1A CDH1 ATF3 PLAU MMP9 TFPI2 CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT CGGCTGCAGGAAAGTTTACA
Supplemental Table 1. List of genes and mature shrna sequences identified from the shrna screen in BRCA2-mutant PEO1 cells.
 Guillemette_Supplemental Table 1 Gene Mature Sequence Gene Mature Sequence ATTCATAGGATGTCAGCAG LOC157193 TATGCGTAAGTAATAAATTGCT TTAGTTCTGTCTTGGAGG LOC152225 CGCATATATGTTCTGACTT ALDH2 CACTTCAGTGTATGCCT
Guillemette_Supplemental Table 1 Gene Mature Sequence Gene Mature Sequence ATTCATAGGATGTCAGCAG LOC157193 TATGCGTAAGTAATAAATTGCT TTAGTTCTGTCTTGGAGG LOC152225 CGCATATATGTTCTGACTT ALDH2 CACTTCAGTGTATGCCT
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 MSI2 interactors are associated with the riboproteome and are functionally relevant. (a) Coomassie blue staining of FLAG-MSI2 immunoprecipitated complexes. (b) GO analysis of MSI2-interacting
Supplementary Figure 1 MSI2 interactors are associated with the riboproteome and are functionally relevant. (a) Coomassie blue staining of FLAG-MSI2 immunoprecipitated complexes. (b) GO analysis of MSI2-interacting
fl/+ KRas;Atg5 fl/+ KRas;Atg5 fl/fl KRas;Atg5 fl/fl KRas;Atg5 Supplementary Figure 1. Gene set enrichment analyses. (a) (b)
 KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
Supplemental Figure S1. RANK expression on human lung cancer cells.
 Supplemental Figure S1. RANK expression on human lung cancer cells. (A) Incidence and H-Scores of RANK expression determined from IHC in the indicated primary lung cancer subgroups. The overall expression
Supplemental Figure S1. RANK expression on human lung cancer cells. (A) Incidence and H-Scores of RANK expression determined from IHC in the indicated primary lung cancer subgroups. The overall expression
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
PKCζ Promotes Breast Cancer Invasion by Regulating Expression of E-cadherin and Zonula Occludens-1 (ZO-1) via NFκB-p65
 SUPPLEMENTARY INFORMATION TITLE: PKCζ Promotes Breast Cancer Invasion by Regulating Expression of E-cadherin and Zonula Occludens-1 (ZO-1) via NFκB-p65 RUNNING TITLE: PKCζ-NFκB Signaling in Breast Cancer
SUPPLEMENTARY INFORMATION TITLE: PKCζ Promotes Breast Cancer Invasion by Regulating Expression of E-cadherin and Zonula Occludens-1 (ZO-1) via NFκB-p65 RUNNING TITLE: PKCζ-NFκB Signaling in Breast Cancer
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
SUPPLEMENTARY INFORMATION. Supplementary Figures S1-S9. Supplementary Methods
 SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma.
 Supplementary Figure 1 Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma. (a) Breeding scheme. R26-LSL-SB11 homozygous mice were bred to Trp53 LSL-R270H/+
Supplementary Figure 1 Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma. (a) Breeding scheme. R26-LSL-SB11 homozygous mice were bred to Trp53 LSL-R270H/+
The mir-199a/brm/egr1 axis is a determinant of anchorage-independent growth in epithelial tumor cell lines
 Supplementary information Supplementary Figure -9 Supplementary Table -4 The mir-99a/brm/egr axis is a determinant of anchorage-independent growth in epithelial tumor cell lines Kazuyoshi Kobayashi, Kouhei
Supplementary information Supplementary Figure -9 Supplementary Table -4 The mir-99a/brm/egr axis is a determinant of anchorage-independent growth in epithelial tumor cell lines Kazuyoshi Kobayashi, Kouhei
Developments in small cell lung cancer G. Giaccone, MD PhD Chief, Medical Oncology Branch and Affiliates National Cancer Institute Bethesda MD USA
 Developments in small cell lung cancer G. Giaccone, MD PhD Chief, Medical Oncology Branch and Affiliates National Cancer Institute Bethesda MD USA Geneva April 20, 2012 Neuroendocrine tumors of lung Typical
Developments in small cell lung cancer G. Giaccone, MD PhD Chief, Medical Oncology Branch and Affiliates National Cancer Institute Bethesda MD USA Geneva April 20, 2012 Neuroendocrine tumors of lung Typical
Supplemental Data Macrophage Migration Inhibitory Factor MIF Interferes with the Rb-E2F Pathway
 Supplemental Data Macrophage Migration Inhibitory Factor MIF Interferes with the Rb-E2F Pathway S1 Oleksi Petrenko and Ute M. Moll Figure S1. MIF-Deficient Cells Have Reduced Transforming Ability (A) Soft
Supplemental Data Macrophage Migration Inhibitory Factor MIF Interferes with the Rb-E2F Pathway S1 Oleksi Petrenko and Ute M. Moll Figure S1. MIF-Deficient Cells Have Reduced Transforming Ability (A) Soft
S1a S1b S1c. S1d. S1f S1g S1h SUPPLEMENTARY FIGURE 1. - si sc Il17rd Il17ra bp. rig/s IL-17RD (ng) -100 IL-17RD
 SUPPLEMENTARY FIGURE 1 0 20 50 80 100 IL-17RD (ng) S1a S1b S1c IL-17RD β-actin kda S1d - si sc Il17rd Il17ra rig/s15-574 - 458-361 bp S1f S1g S1h S1i S1j Supplementary Figure 1. Knockdown of IL-17RD enhances
SUPPLEMENTARY FIGURE 1 0 20 50 80 100 IL-17RD (ng) S1a S1b S1c IL-17RD β-actin kda S1d - si sc Il17rd Il17ra rig/s15-574 - 458-361 bp S1f S1g S1h S1i S1j Supplementary Figure 1. Knockdown of IL-17RD enhances
Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients.
 Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
Inhibition of Gastric Cancer Invasion and Metastasis by PLA2G2A, a Novel β-catenin/tcf Target Gene
 Supplementary Information Inhibition of Gastric Cancer Invasion and Metastasis by PLA2G2A, a Novel β-catenin/tcf Target Gene Kumaresan Ganesan, 1 Tatiana Ivanova, 1 Yonghui Wu, 1 Vikneswari Rajasegaran,
Supplementary Information Inhibition of Gastric Cancer Invasion and Metastasis by PLA2G2A, a Novel β-catenin/tcf Target Gene Kumaresan Ganesan, 1 Tatiana Ivanova, 1 Yonghui Wu, 1 Vikneswari Rajasegaran,
Guangdong Medical University, Zhanjiang, China; 5 Guangxi Medical University, Nanning, China; 6 Department of Pathology, University of Michigan
 Overexpression of FAM83H-AS1 indicates poor patient survival and knockdown impairs cell proliferation and invasion via MET/EGFR signaling in lung cancer Jie Zhang 1,2, Shumei Feng 3, Wenmei Su 4, Shengbin
Overexpression of FAM83H-AS1 indicates poor patient survival and knockdown impairs cell proliferation and invasion via MET/EGFR signaling in lung cancer Jie Zhang 1,2, Shumei Feng 3, Wenmei Su 4, Shengbin
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2566 Figure S1 CDKL5 protein expression pattern and localization in mouse brain. (a) Multiple-tissue western blot from a postnatal day (P) 21 mouse probed with an antibody against CDKL5.
DOI: 10.1038/ncb2566 Figure S1 CDKL5 protein expression pattern and localization in mouse brain. (a) Multiple-tissue western blot from a postnatal day (P) 21 mouse probed with an antibody against CDKL5.
SUPPLEMENTARY FIGURE LEGENDS. atypical adenomatous hyperplasias (AAH); Grade II: adenomas; Grade III: adenocarcinomas;
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure S1: Tumor grades in Ras G12D ; p53 / lung tumors. Representative histology (H&E) of K-Ras G12D ; p53 / lung tumors 13 weeks after tumor initiation. Grade
SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure S1: Tumor grades in Ras G12D ; p53 / lung tumors. Representative histology (H&E) of K-Ras G12D ; p53 / lung tumors 13 weeks after tumor initiation. Grade
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
SUPPLEMENTARY INFORMATION
 doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Figure 1. A. Bar graph representing the expression levels of the 19 indicated genes in the microarrays analyses comparing human lung
 Supplementary Figure 1. A. Bar graph representing the expression levels of the 19 indicated genes in the microarrays analyses comparing human lung immortalized broncho-epithelial cells (AALE cells) expressing
Supplementary Figure 1. A. Bar graph representing the expression levels of the 19 indicated genes in the microarrays analyses comparing human lung immortalized broncho-epithelial cells (AALE cells) expressing
Genetic Alteration Panels
 TM Genetic Alteration Panels GENETIC ALTERATION PANELS Table of Contents AKT Genetic Alteration Panel (ATCC No. TCP-1029 )...1 BRAF Genetic Alteration Panel (ATCC No. TCP-1032 )...5 EGFR Genetic Alteration
TM Genetic Alteration Panels GENETIC ALTERATION PANELS Table of Contents AKT Genetic Alteration Panel (ATCC No. TCP-1029 )...1 BRAF Genetic Alteration Panel (ATCC No. TCP-1032 )...5 EGFR Genetic Alteration
Chapter 4 Cellular Oncogenes ~ 4.6 -
 Chapter 4 Cellular Oncogenes - 4.2 ~ 4.6 - Many retroviruses carrying oncogenes have been found in chickens and mice However, attempts undertaken during the 1970s to isolate viruses from most types of
Chapter 4 Cellular Oncogenes - 4.2 ~ 4.6 - Many retroviruses carrying oncogenes have been found in chickens and mice However, attempts undertaken during the 1970s to isolate viruses from most types of
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Supplementary Figure 1
 A B D Relative TAp73 mrna p73 Supplementary Figure 1 25 2 15 1 5 p63 _-tub. MDA-468 HCC1143 HCC38 SUM149 MDA-468 HCC1143 HCC38 SUM149 HCC-1937 MDA-MB-468 ΔNp63_ TAp73_ TAp73β E C Relative ΔNp63 mrna TAp73
A B D Relative TAp73 mrna p73 Supplementary Figure 1 25 2 15 1 5 p63 _-tub. MDA-468 HCC1143 HCC38 SUM149 MDA-468 HCC1143 HCC38 SUM149 HCC-1937 MDA-MB-468 ΔNp63_ TAp73_ TAp73β E C Relative ΔNp63 mrna TAp73
Supplemental Methods LC-MS Assay for NAD Concentration
 Supplemental Methods LC-MS Assay for NAD Concentration The concentration of NAD in cells was determined by a non-validated liquid chromatography tandem mass spectrometry (LC-MS/MS) assay using 13 C 5 -NAD
Supplemental Methods LC-MS Assay for NAD Concentration The concentration of NAD in cells was determined by a non-validated liquid chromatography tandem mass spectrometry (LC-MS/MS) assay using 13 C 5 -NAD
BRIEF REPORT. mir-101 DNA Copy Loss is a Prominent Subtype Specific Event in Lung Cancer
 BRIEF REPORT mir-101 DNA Copy Loss is a Prominent Subtype Specific Event in Lung Cancer Kelsie L. Thu, BSc,* Raj Chari, PhD,* William W. Lockwood, PhD,* Stephen Lam, MD,* and Wan L. Lam, PhD* Introduction:
BRIEF REPORT mir-101 DNA Copy Loss is a Prominent Subtype Specific Event in Lung Cancer Kelsie L. Thu, BSc,* Raj Chari, PhD,* William W. Lockwood, PhD,* Stephen Lam, MD,* and Wan L. Lam, PhD* Introduction:
Discovery Dataset. PD Liver Luminal B/ Her-2+ Letrozole. PD Supraclavicular Lymph node. PD Supraclavicular Lymph node Luminal B.
 Discovery Dataset 11T pt1cpn2am1(liver) 2009 2010 Liver / Her-2+ 2011 Death Letrozole CHT pt1cpn0(sn)m0 Supraclavicular Lymph node Death 12T 2003 2006 2006 2008 Anastrozole Local RT+Examestane Fulvestrant
Discovery Dataset 11T pt1cpn2am1(liver) 2009 2010 Liver / Her-2+ 2011 Death Letrozole CHT pt1cpn0(sn)m0 Supraclavicular Lymph node Death 12T 2003 2006 2006 2008 Anastrozole Local RT+Examestane Fulvestrant
(a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable
 Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Hepatitis B virus X gene in the development of hepatocellular carcinoma. Citation Hong Kong Medical Journal, 2011, v. 17 n. 6, suppl. 6, p.
 Title Hepatitis B virus X gene in the development of hepatocellular carcinoma Author(s) Ma, NF; Lau, SH; Hu, L; Dong, SS; Guan, XY Citation Hong Kong Medical Journal, 2011, v. 17 n. 6, suppl. 6, p. 44-47
Title Hepatitis B virus X gene in the development of hepatocellular carcinoma Author(s) Ma, NF; Lau, SH; Hu, L; Dong, SS; Guan, XY Citation Hong Kong Medical Journal, 2011, v. 17 n. 6, suppl. 6, p. 44-47
America, Hershey, PA Australia, Melbourne, VIC Europe, Munich
 America, Hershey, PA Australia, Melbourne, VIC Europe, Munich info@vivopharm.com www.vivopharm.com Bladder MB-9-luc 2 Mouse urinary bladder carcinoma yes yes yes C57BL/6 2 Bone UMR-106 Rat osteosarcoma
America, Hershey, PA Australia, Melbourne, VIC Europe, Munich info@vivopharm.com www.vivopharm.com Bladder MB-9-luc 2 Mouse urinary bladder carcinoma yes yes yes C57BL/6 2 Bone UMR-106 Rat osteosarcoma
Supplementary Information
 Supplementary Information - chimeric fusion transcript in human gastric cancer promotes tumorigenesis through activation of PI3K/AKT signaling Sun Mi Yun, Kwiyeom Yoon, Sunghoon Lee, Eunjeong Kim, Seong-Ho
Supplementary Information - chimeric fusion transcript in human gastric cancer promotes tumorigenesis through activation of PI3K/AKT signaling Sun Mi Yun, Kwiyeom Yoon, Sunghoon Lee, Eunjeong Kim, Seong-Ho
Species Tumor Type Comment for in vivo work Lead Time for in vivo studies [weeks] MB-49-luc-2 Mouse urinary bladder carcinoma C57BL/6 2
![Species Tumor Type Comment for in vivo work Lead Time for in vivo studies [weeks] MB-49-luc-2 Mouse urinary bladder carcinoma C57BL/6 2 Species Tumor Type Comment for in vivo work Lead Time for in vivo studies [weeks] MB-49-luc-2 Mouse urinary bladder carcinoma C57BL/6 2](/thumbs/88/116309357.jpg) America, Hershey, PA Australia, Melbourne, VIC Europe, Munich info@vivopharm.com www.vivopharm.com Tissue Bladder Species Tumor Type Comment for in vivo work Lead Time for in vivo studies [weeks] MB-49-luc-
America, Hershey, PA Australia, Melbourne, VIC Europe, Munich info@vivopharm.com www.vivopharm.com Tissue Bladder Species Tumor Type Comment for in vivo work Lead Time for in vivo studies [weeks] MB-49-luc-
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Neuregulin-1-Mediated Autocrine Signaling Underlies Sensitivity to HER2 Kinase Inhibitors in a Subset of Human Cancers
 Article Neuregulin-1-Mediated Autocrine Signaling Underlies Sensitivity to HER2 Kinase Inhibitors in a Subset of Human Cancers Timothy R. Wilson, 1,2 Diana Y. Lee, 1 Leanne Berry, 2 David S. Shames, 2
Article Neuregulin-1-Mediated Autocrine Signaling Underlies Sensitivity to HER2 Kinase Inhibitors in a Subset of Human Cancers Timothy R. Wilson, 1,2 Diana Y. Lee, 1 Leanne Berry, 2 David S. Shames, 2
Title: Cytosolic DNA-mediated, STING-dependent pro-inflammatory gene. Fig. S1. STING ligands-mediated signaling response in MEFs. (A) Primary MEFs (1
 1 Supporting Information 2 3 4 Title: Cytosolic DNA-mediated, STING-dependent pro-inflammatory gene induction necessitates canonical NF-κB activation through TBK1 5 6 Authors: Abe et al. 7 8 9 Supporting
1 Supporting Information 2 3 4 Title: Cytosolic DNA-mediated, STING-dependent pro-inflammatory gene induction necessitates canonical NF-κB activation through TBK1 5 6 Authors: Abe et al. 7 8 9 Supporting
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
mtor Inhibition Specifically Sensitizes Colorectal Cancers with KRAS or BRAF Mutations to BCL-2/BCL-
 Supplementary Material for mtor Inhibition Specifically Sensitizes Colorectal Cancers with KRAS or BRAF Mutations to BCL-2/BCL- XL Inhibition by Suppressing MCL-1 Anthony C. Faber 1,2 *, Erin M. Coffee
Supplementary Material for mtor Inhibition Specifically Sensitizes Colorectal Cancers with KRAS or BRAF Mutations to BCL-2/BCL- XL Inhibition by Suppressing MCL-1 Anthony C. Faber 1,2 *, Erin M. Coffee
The Changing Landscape of Lung Cancer and its Treatment. G. Giaccone, MD PhD Medical Oncology Branch National Cancer Institute Bethesda, MD USA
 The Changing Landscape of Lung Cancer and its Treatment G. Giaccone, MD PhD Medical Oncology Branch National Cancer Institute Bethesda, MD USA Lung Cancer 2011 USA 190,000 new cases 165,000 deaths 165,300
The Changing Landscape of Lung Cancer and its Treatment G. Giaccone, MD PhD Medical Oncology Branch National Cancer Institute Bethesda, MD USA Lung Cancer 2011 USA 190,000 new cases 165,000 deaths 165,300
Supplemental figures
 Supplemental figures Supplemental Figure 1. Impact of immunogenic-llc tumors (LLC-OVA) on peripheral OVA-specific CD8 T cells. Tetramer analysis showing percent OVA-specific CD8 T cells in peripheral blood
Supplemental figures Supplemental Figure 1. Impact of immunogenic-llc tumors (LLC-OVA) on peripheral OVA-specific CD8 T cells. Tetramer analysis showing percent OVA-specific CD8 T cells in peripheral blood
(A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC,
 Supplemental Figure Legends Figure S1. Cell line characterization (A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC, green) and co-stained with DAPI to visualize the nuclei.
Supplemental Figure Legends Figure S1. Cell line characterization (A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC, green) and co-stained with DAPI to visualize the nuclei.
Supplementary Information. Induction of p53-independent apoptosis by ectopic expression of HOXA5
 Supplementary Information Induction of p53-independent apoptosis by ectopic expression of in human liposarcomas Dhong Hyun Lee 1, *, Charles Forscher 1, Dolores Di Vizio 2, 3, and H. Phillip Koeffler 1,
Supplementary Information Induction of p53-independent apoptosis by ectopic expression of in human liposarcomas Dhong Hyun Lee 1, *, Charles Forscher 1, Dolores Di Vizio 2, 3, and H. Phillip Koeffler 1,
RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh-
 1 a b Supplementary Figure 1. Effects of GSK3b knockdown on poly I:C-induced cytokine production. RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh- GSK3b) were stimulated
1 a b Supplementary Figure 1. Effects of GSK3b knockdown on poly I:C-induced cytokine production. RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh- GSK3b) were stimulated
p.r623c p.p976l p.d2847fs p.t2671 p.d2847fs p.r2922w p.r2370h p.c1201y p.a868v p.s952* RING_C BP PHD Cbp HAT_KAT11
 ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus
 Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
GENETIC ALTERATIONS AND LINEAGE SPECIFICITY IN LUNG CANCER WILLIAM W. LOCKWOOD. B.Sc., The University of British Columbia, 2004
 GENETIC ALTERATIONS AND LINEAGE SPECIFICITY IN LUNG CANCER by WILLIAM W. LOCKWOOD B.Sc., The University of British Columbia, 2004 A THESIS SUBMITTED IN PARTIAL FULFILMENT OF THE REQUIREMENTS FOR THE DEGREE
GENETIC ALTERATIONS AND LINEAGE SPECIFICITY IN LUNG CANCER by WILLIAM W. LOCKWOOD B.Sc., The University of British Columbia, 2004 A THESIS SUBMITTED IN PARTIAL FULFILMENT OF THE REQUIREMENTS FOR THE DEGREE
Supplementary Table S1: Primers for DNA sequencing (A), quantification of. relative mrna expression (B), and copy number analysis (C)
 Supplementary Table S1: Primers for DNA sequencing (A), quantification of relative mrna expression (B), and copy number analysis (C) A. DNA sequencing (1-4) Genes Primers Sequence EGFR exon 18 EGFR E18F
Supplementary Table S1: Primers for DNA sequencing (A), quantification of relative mrna expression (B), and copy number analysis (C) A. DNA sequencing (1-4) Genes Primers Sequence EGFR exon 18 EGFR E18F
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES 1 Supplementary Figure 1, Adult hippocampal QNPs and TAPs uniformly express REST a-b) Confocal images of adult hippocampal mouse sections showing GFAP (green), Sox2 (red), and REST
SUPPLEMENTARY FIGURES 1 Supplementary Figure 1, Adult hippocampal QNPs and TAPs uniformly express REST a-b) Confocal images of adult hippocampal mouse sections showing GFAP (green), Sox2 (red), and REST
Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well
 Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well plate at cell densities ranging from 25-225 cells in
Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well plate at cell densities ranging from 25-225 cells in
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/322/ra38/dc1 Supplementary Materials for Dynamic Reprogramming of Signaling Upon Met Inhibition Reveals a Mechanism of Drug Resistance in Gastric Cancer Andrea
www.sciencesignaling.org/cgi/content/full/7/322/ra38/dc1 Supplementary Materials for Dynamic Reprogramming of Signaling Upon Met Inhibition Reveals a Mechanism of Drug Resistance in Gastric Cancer Andrea
MOLECULAR PREDICTIVE MARKERS OF LUNG CARCINOMA: KFSH&RC EXPERIENCE
 25 th IAP-Arab Division Conference 07-09 November 2013, Amman, Jordan MOLECULAR PREDICTIVE MARKERS OF LUNG CARCINOMA: KFSH&RC EXPERIENCE Fouad Al Dayel, MD, FRCPA, FRCPath Professor and Chairman Department
25 th IAP-Arab Division Conference 07-09 November 2013, Amman, Jordan MOLECULAR PREDICTIVE MARKERS OF LUNG CARCINOMA: KFSH&RC EXPERIENCE Fouad Al Dayel, MD, FRCPA, FRCPath Professor and Chairman Department
SUPPLEMENTARY INFORMATION
 DOI: 1.138/ncb3355 a S1A8 + cells/ total.1.8.6.4.2 b S1A8/?-Actin c % T-cell proliferation 3 25 2 15 1 5 T cells Supplementary Figure 1 Inter-tumoral heterogeneity of MDSC accumulation in mammary tumor
DOI: 1.138/ncb3355 a S1A8 + cells/ total.1.8.6.4.2 b S1A8/?-Actin c % T-cell proliferation 3 25 2 15 1 5 T cells Supplementary Figure 1 Inter-tumoral heterogeneity of MDSC accumulation in mammary tumor
Nature Immunology: doi: /ni Supplementary Figure 1. DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells.
 Supplementary Figure 1 DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells. (a) Scheme for the retroviral shrna screen. (b) Histogram showing CD4 expression (MFI) in WT cytotoxic
Supplementary Figure 1 DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells. (a) Scheme for the retroviral shrna screen. (b) Histogram showing CD4 expression (MFI) in WT cytotoxic
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was
 Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was painted on the shaved back skin of CBL/J and BALB/c mice for consecutive days. (a, b) Phenotypic presentation of mouse back skin
Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was painted on the shaved back skin of CBL/J and BALB/c mice for consecutive days. (a, b) Phenotypic presentation of mouse back skin
Supplementary Information Supplementary Fig. 1. Elevated Usp9x in melanoma and NRAS mutant melanoma cells are dependent on NRAS for 3D growth.
 Supplementary Information Supplementary Fig. 1. Elevated Usp9x in melanoma and NRAS mutant melanoma cells are dependent on NRAS for 3D growth. a. Immunoblot for Usp9x protein in NRAS mutant melanoma cells
Supplementary Information Supplementary Fig. 1. Elevated Usp9x in melanoma and NRAS mutant melanoma cells are dependent on NRAS for 3D growth. a. Immunoblot for Usp9x protein in NRAS mutant melanoma cells
University of Miami Miller School of Medicine, Miami, FL 33136, USA, 3 State Key Laboratory
 Supplementary File ASXL1 plays an important role in erythropoiesis Hui Shi 1,2,3, Shohei Yamamoto 1,2,4, Mengyao Sheng 3, Jie Bai 3, Peng Zhang 1,2, Runze Chen 1,2, Shi Chen 1,2, Lihong Shi 3, Omar Abdel-Wahab
Supplementary File ASXL1 plays an important role in erythropoiesis Hui Shi 1,2,3, Shohei Yamamoto 1,2,4, Mengyao Sheng 3, Jie Bai 3, Peng Zhang 1,2, Runze Chen 1,2, Shi Chen 1,2, Lihong Shi 3, Omar Abdel-Wahab
Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein
 Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein content relative to GAPDH in two independent experiments.
Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein content relative to GAPDH in two independent experiments.
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland
 AD Award Number: DAMD17-03-1-0392 TITLE: The Role of Notch Signaling Pathway in Breast Cancer Pathogenesis PRINCIPAL INVESTIGATOR: Annapoorni Rangarajan, Ph.D. CONTRACTING ORGANIZATION: Indian Institute
AD Award Number: DAMD17-03-1-0392 TITLE: The Role of Notch Signaling Pathway in Breast Cancer Pathogenesis PRINCIPAL INVESTIGATOR: Annapoorni Rangarajan, Ph.D. CONTRACTING ORGANIZATION: Indian Institute
Award Number: W81XWH
 AD Award Number: W81XWH-11-1-0285 TITLE: Multiple Cooperating Oncogenes Drive Recurrent Breast Cancer-Associated Chromosomal Amplifications: Creation of Isogenic Human Cell Line Models PRINCIPAL INVESTIGATOR:
AD Award Number: W81XWH-11-1-0285 TITLE: Multiple Cooperating Oncogenes Drive Recurrent Breast Cancer-Associated Chromosomal Amplifications: Creation of Isogenic Human Cell Line Models PRINCIPAL INVESTIGATOR:
MOLECULAR PHARMACOLOGY SUPPLEMENT
 MOLECULAR PHARMACOLOGY SUPPLEMENT Therapeutic targeting of a novel 6-substituted pyrrolo[2,3-d]pyrimidine thienoyl antifolate to human solid tumors based on selective uptake by the proton-coupled folate
MOLECULAR PHARMACOLOGY SUPPLEMENT Therapeutic targeting of a novel 6-substituted pyrrolo[2,3-d]pyrimidine thienoyl antifolate to human solid tumors based on selective uptake by the proton-coupled folate
Characterizing the cancer genome in lung adenocarcinoma
 Characterizing the cancer genome in lung adenocarcinoma Supplementary information Table of contents Page Supplemental Figures 1-11 Supplementary Figure 1 (Binomial analysis) 1 Supplementary Figure 2 (Tertile
Characterizing the cancer genome in lung adenocarcinoma Supplementary information Table of contents Page Supplemental Figures 1-11 Supplementary Figure 1 (Binomial analysis) 1 Supplementary Figure 2 (Tertile
Nature Immunology: doi: /ni Supplementary Figure 1. Huwe1 has high expression in HSCs and is necessary for quiescence.
 Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
Supplementary Figure S1. Generation of LSL-EZH2 conditional transgenic mice.
 Downstream Col1A locus S P P P EP Genotyping with P1, P2 frt PGKneopA + frt hygro-pa Targeting vector Genotyping with P3, P4 P1 pcag-flpe P2 P3 P4 frt SApA CAG LSL PGKATG frt hygro-pa C. D. E. ormal KRAS
Downstream Col1A locus S P P P EP Genotyping with P1, P2 frt PGKneopA + frt hygro-pa Targeting vector Genotyping with P3, P4 P1 pcag-flpe P2 P3 P4 frt SApA CAG LSL PGKATG frt hygro-pa C. D. E. ormal KRAS
Nuclear Factor I/B is an Oncogene. in Small Cell Lung Cancer
 Nuclear Factor I/B is an Oncogene in Small Cell Lung Cancer Alison L. Dooley 1, Monte M. Winslow 1, Derek Y. Chiang 2,3,4, Shantanu Banerji 2,3, Nicolas Stransky 2, Talya L. Dayton 1, Eric L. Snyder 1,
Nuclear Factor I/B is an Oncogene in Small Cell Lung Cancer Alison L. Dooley 1, Monte M. Winslow 1, Derek Y. Chiang 2,3,4, Shantanu Banerji 2,3, Nicolas Stransky 2, Talya L. Dayton 1, Eric L. Snyder 1,
Supplementary Figure 1 Cell line TRIB2 status. Supplementary Figure 2 TRIB2 status has no impact on the cell cycle after PI3K inhibition. a. b.
 Supplementary Figure 1 Cell line TRIB2 status. TRIB2 protein expression to determine endogenous expression and to determine the effectiveness of each of our TRIB2 knockdown constructs. Supplementary Figure
Supplementary Figure 1 Cell line TRIB2 status. TRIB2 protein expression to determine endogenous expression and to determine the effectiveness of each of our TRIB2 knockdown constructs. Supplementary Figure
Supplementary Figure 1. Blood glucose and insulin levels in mice during 4-day infusion.
 Supplementary Figure 1. Blood glucose and insulin levels in mice during 4-day infusion. (A-B) WT and HT mice infused with saline or glucose had overlapping achieved blood glucose and insulin levels, necessitating
Supplementary Figure 1. Blood glucose and insulin levels in mice during 4-day infusion. (A-B) WT and HT mice infused with saline or glucose had overlapping achieved blood glucose and insulin levels, necessitating
Title: MYBBP1A suppresses breast cancer tumorigenesis by enhancing the p53 dependent anoikis
 Author's response to reviews Title: MYBBP1A suppresses breast cancer tumorigenesis by enhancing the p53 dependent anoikis Authors: Kensuke Akaogi (kensuke.akaogi@gmail.com) Wakana Ono (wakana315@gmail.com)
Author's response to reviews Title: MYBBP1A suppresses breast cancer tumorigenesis by enhancing the p53 dependent anoikis Authors: Kensuke Akaogi (kensuke.akaogi@gmail.com) Wakana Ono (wakana315@gmail.com)
oocytes were pooled for RT-PCR analysis. The number of PCR cycles was 35. Two
 Supplementary Fig. 1 a Kdm3a Kdm4b β-actin Oocyte Testis Oocyte Testis Oocyte Testis b 1.8 Relative expression.6.4.2 Kdm3a Kdm4b RT-PCR analysis of Kdm3a and Kdm4b expression in oocytes and testes. Twenty
Supplementary Fig. 1 a Kdm3a Kdm4b β-actin Oocyte Testis Oocyte Testis Oocyte Testis b 1.8 Relative expression.6.4.2 Kdm3a Kdm4b RT-PCR analysis of Kdm3a and Kdm4b expression in oocytes and testes. Twenty
Nature Genetics: doi: /ng.3731
 Supplementary Figure 1 Circadian profiles of Adarb1 transcript and ADARB1 protein in mouse tissues. (a) Overlap of rhythmic transcripts identified in the previous transcriptome analyses. The mouse liver
Supplementary Figure 1 Circadian profiles of Adarb1 transcript and ADARB1 protein in mouse tissues. (a) Overlap of rhythmic transcripts identified in the previous transcriptome analyses. The mouse liver
SUPPLEMENTAL TABLE AND FIGURES
 SUPPLEMENTAL TABLE AND FIGURES Zhang et al. (29) - Enzymes in the NAD + Salvage Pathway Regulate SIRT1 Activity at Target Gene Promoters This document contains supplemental data (1 table and 6 figures)
SUPPLEMENTAL TABLE AND FIGURES Zhang et al. (29) - Enzymes in the NAD + Salvage Pathway Regulate SIRT1 Activity at Target Gene Promoters This document contains supplemental data (1 table and 6 figures)
URI Is an Oncogene Amplified in Ovarian Cancer Cells and Is Required for Their Survival
 Article Is an Oncogene Amplified in Ovarian Cancer Cells and Is Required for Their Survival Jean-Philippe Theurillat,,, Stefan Christian Metzler,, Nico Henzi, Nabil Djouder, Marianne Helbling, Anna-Kathrin
Article Is an Oncogene Amplified in Ovarian Cancer Cells and Is Required for Their Survival Jean-Philippe Theurillat,,, Stefan Christian Metzler,, Nico Henzi, Nabil Djouder, Marianne Helbling, Anna-Kathrin
