On the Reproducibility of TCGA Ovarian Cancer MicroRNA Profiles
|
|
|
- Alban Fitzgerald
- 5 years ago
- Views:
Transcription
1 On the Reproducibility of TCGA Ovarian Cancer MicroRNA Profiles Ying-Wooi Wan 1,2,4, Claire M. Mach 2,3, Genevera I. Allen 1,7,8, Matthew L. Anderson 2,4,5 *, Zhandong Liu 1,5,6,7 * 1 Departments of Pediatrics 2 Obstetrics and Gynecology and 4 Pathology and Immunology, 5 Dan L. Duncan Cancer Center, 6 Computational and Integrative Biomedical Research (CIBR) Center, Baylor College of Medicine. 3 College of Pharmacy, University of Houston 7 Neurological Research Institute, Texas Children s Hospital 8 Department of Statistics and Electrical & Computer Engineering, Rice University *Correspondence: One Baylor Plaza, BCM320 Phone: FAX: Zhandonl@bcm.edu or matthew@bcm.edu 1
2 ABSTRACT Dysregulated microrna (mirna) expression is a well-established feature of human cancer. However, the role of specific mirnas in determining cancer outcomes remains unclear. Using Level 3 expression data from the Cancer Genome Atlas (TCGA), we identified 61 mirnas that are associated with overall survival in 469 ovarian cancers profiled by microarray (p<0.01). We also identified 12 mirnas that are associated with survival when mirnas were profiled in the same specimens using Next Generation Sequencing (mirna-seq) (p<0.01). Surprisingly, only 1 mirna transcript is associated with ovarian cancer survival in both datasets. Our analyses indicate that this discrepancy is due to the fact that mirna levels reported by the two platforms correlate poorly, even after correcting for potential issues inherent to signal detection algorithms. Further investigation is warranted. 2
3 INTRODUCTION MicroRNAs (mirnas) are endogenous RNA transcripts that regulate diverse patterns of gene expression. 1 Most human mirnas are transcribed as long precursors known as primirnas. Starting in the nucleus, pri-mirnas undergo a series of processing events that ultimately result in the cytoplasmic release of mature transcripts ~22 nucleotides in length. Mature mirnas catalyze translational inhibition by directly binding to messenger RNAs (mrnas) and promoting their degradation. 2 Recent data also indicate that mirnas can inhibit translation independent of their ability to induce mrna degradation. Patterns of mirna expression have now been extensively profiled in many different human tissues. It is now clear that dysregulated mirna expression is a feature of many different cancers, including carcinomas of the breast, ovary and lung. 3-5 However, determining the mechanisms by which individual mirnas contribute to cancer outcomes remains a key challenge for biologists hoping to exploit their power. Recently, the Cancer Genome Atlas Consortium (TCGA) reported that ovarian cancers cluster into distinct molecular subtypes based on their patterns of gene and microrna expression. 6 However, we have discovered an alarming lack of consistency between the microrna (mirna) expression profiles initially used by the TCGA and a subsequent profile of mirna expression generated by this group for the same ovarian cancer specimens using mirna-seq. As these observations challenge the validity of the underlying data, they also challenge scientific discoveries based on this data. RESULTS To delineate mirnas associated with ovarian cancer patient survival, we performed a univariate Cox regression analysis using Level 3 TCGA mirna data for 469 ovarian cancers profiled using Agilent microarray technology. We found that 61 mature mirnas are significantly associated with ovarian cancer survival (p<0.01) (Figure 1A). Of these, mir-505, mir-652 and mir-551b* demonstrate the most robust associations. Hazard ratios (HR) calculated for these 3
4 mirnas were -1.73, -1.8, and 9.3, respectively, indicating that each mirna potentially plays an important role in determining ovarian cancer survival. To validate these observations, we next interrogated a second dataset of mirna expression generated for the same ovarian cancer specimens using Next Generation Sequencing (mirna- Seq). The TCGA ovarian cancer project is unique in that mirna expression has been profiled using both mirna array and mirna-seq. Use of these technically distinct platforms creates a unique opportunity to validate discoveries made using one dataset against the other. Ideally, the results obtained should correlate tightly. Using Cox Proportional Hazards analysis, we found that 12 mirna transcripts are associated with survival when mirnas were profiled in ovarian cancers using mirna-seq (Figure 1B). However, the hazard ratios estimated for the 12 mirnas identified from mirna-seq data are all very close to 1.0. Surprisingly, only mir-652 is associated with survival in both the mirna-seq and microarray datasets. To correct for multiple hypothesis testing, we adjusted our Cox model p-values using Benjamini Hochberg procedure. 7 After completing these analyses, no mirnas are correlated with survival in both datasets when the false discovery rate was set at 10%. To elucidate potential causes for this unexpected discrepancy, we examined the reproducibility of mirna expression between the two TCGA files that describe this data. Pearson correlation coefficients (r) were calculated for each of the 359 mature human mirnas for which Level 3 expression data was available in both the mirna-seq and microarray databases. We found that correlation coefficients for levels of individual mirnas reported by each technique varied widely. For example, mir-505 is the mirna most robustly associated with patient outcome in our analyses of the mirna array data (HR = -1.7, p< 9e-5). However, when assessed using sequencing data, the hazard ratio for mir-505 was (p=0.03). Levels of mir-505 measured by mirna-array and mirna-seq data correlated only modestly (r = 0.59) (Figure 2B). Discrepancies were also observed in a number of other mirnas that have been previously implicated in ovarian cancer, such as mir The correlation coefficient for mir- 4
5 143 in our analyses was 0.39 (Figure 2C). Another mirna well-studied in ovarian cancer is mir-141, which has been previously reported to target p38α and modulate the oxidative stress response. 9,10 However, the correlation between levels of mir-141 in TCGA microarray and mirna-seq expression data is only 0.32 (Figure 2D). Overall, we found that correlation coefficients for ~72% of mirnas profiled in both datasets were 0.5 (Figure 3A, 3C), indicating poor reproducibility. In contrast, only 22% of the mrnas measured by Agilent microarray and Illumina HiSeq using the same ovarian cancer specimens correlate poorly (r 0.5; Figure 3B, 3C). Thus, the discrepancy we report here appears to be limited to the TCGA mirna dataset. One potential cause for poor reproducibility may be the signal detection algorithm used to report levels of mirna expression. Level 3 TCGA mirna data are reported in two formats. The first, labeled as a Quantification Data," reports levels for individual human mirnas. However, one of the advantages of mirna-seq is that transcripts retrieved by this technique can be precisely mapped. A second file, labeled as Isoform Data," has also been released by the TCGA. This file reports read counts for transcripts according to their genomic location. As part of this file, transcripts are identified as either mature mirna, mirna* (3p arms of human mirnas), stem-loop transcript or precursor. While working through this data, we learned that mirna levels reported in the TCGA quantification file include read counts for mirna precursors as well as mature mirnas. Because mirna precursors typically lack biologic activity, inclusion of precursors with counts for mature mirnas could confound survival analyses. To address this issue, we retrieved read counts for mature mirnas only from the isoform data file and repeated our analyses. However, the proportion of mirna correlation coefficients 0.5 remained as high as 71% despite the use of this more precisely defined data. A second possible explanation for the discrepancy we observed might be that correlations between measures of mirna expression depend on the frequency with which individual mirna transcripts are expressed. If so, infrequently expressed mirnas might be reported by one or 5
6 both of the platforms used to profile mirna expression randomly or inaccurately. To explore this hypothesis, we re-calculated correlation coefficients for each mirna identified by both platforms after excluding any transcript in the mirna-seq dataset with a read count less than 5. This reduced the number of distinct mirnas available for analysis in the mirna-seq data file from 705 to 380. However, the proportion of mirnas with correlation coefficients 0.5 also decreased from 72% to 56%. Similarly removing poorly expressed transcripts from the pool of mrnas profiled by Illumina HiSeq reduces the proportion of mrnas whose correlation coefficients 0.5 from 22% to 20%. These observations indicate that problems detecting infrequently expressed mirna may impact the ability or one or both platforms to reliably report mirna expression. However, the fact that more than half of mirna transcripts still had correlation coefficients 0.5 even after correcting for this issue indicates that poorly expressed transcripts are not solely responsible for the discordant patterns of mirna expression reported by the two platforms. DISCUSSION Much to our surprise, our analyses indicate that the micrornas associated with survival in ovarian cancer depend highly on whether specimens were profiled by the TCGA using microarray or mirna-seq. Our analyses indicate that this discrepancy exists because mirna- Seq and microarray have generated very different profiles of mirna expression, even though the data is based on the same ovarian cancer specimens. We do not currently have a clear explanation for why mirna expression profiles reported by the TCGA are discordant. However, understanding this discrepancy will ultimately be important for identifying which mirnas if any are important for determining ovarian cancer outcomes. A variety of DNA microarray technologies have been previously validated by investigators examining within platform and cross-platform reproducibility Spearman correlation 6
7 coefficients reported in these studies range from 0.59 to 0.94 with a mean of These results are similar to what we have observed for correlations between patterns of gene expression profiled using microarray and Illumina HiSeq platforms by the TCGA. Both mirna-seq and microarray technologies are associated with multiple technical limitations that might account for the differences we have observed. For example, cross-hybridization is a well-recognized issue that can reduce signal specificity when profiling RNA transcripts by microarray. 14 However, it seems unlikely that cross-hybridization can fully explain the discrepancy we observed, as the number of transcripts correlated with survival by array is greater than the number associated with survival by mirna-seq. One alternate explanation might be that the signal extraction algorithm used to analyze mirna-seq data does not accurately report mirna levels. In general, mirna-seq allows for precise transcript mapping with much greater confidence. The signal extraction algorithm currently used by the TCGA to report mirna levels includes read counts for both a mature mirna and its corresponding precursor. Precursors account for fewer than 1% of the total mirna counts in the TCGA isoform file, likely reflecting the use of sizefractionated total RNA to prepare small RNA libraries for mirna-seq. 5 Our analyses indicate that their inclusion has little bearing on which mirnas are associated with ovarian cancer survival. These observations underscore the urgent need for well-defined algorithms for processing signals generated by mirna-seq and transcriptional profiling platforms. Our understanding is that the same analyses have been performed by TCGA for other cancers, including colon, breast and lung Because mirna expression in these other cancers has not been profiled by microarray, it is not possible to repeat our analyses to determine whether the discrepancy we report is observed in other cancers. Ultimately, consistent and reliable genomic data is critical for constructing testable hypotheses and achieving the full potential of the TCGA. Our observations identify an important hazard of which investigators should be aware as they utilize the TCGA mirna data to study 7
8 ovarian cancer. This hazard underscores the need to validate observations made with one or both of TCGA mirna datasets. Over the long term, resolution of this discrepancy will be important for determining the most effective platform and signal extraction algorithms for profiling mirna expression as part of large scale genomic profiling efforts. MATERIALS AND METHODS Gene and microrna Expression Data. Level 3 data documenting patterns of gene expression for 296 ovarian cancer specimens profiled using Agilent G4502A arrays and Illumina HiSeq were downloaded from the TCGA data portal. Level 3 microrna expression data were also retrieved for 469 ovarian cancer specimens profiled using the Agilent 4X15k array and mirna- Seq. Level 3 mirna data profiled by mirna-seq were retrieved from both the mirna quantification and isoform files available at the TCGA data portal along with metafiles annotating each dataset. Permission to access all data was obtained from the Data Access Committee for the National Center for Biotechnology Information Genotypes and Phenotypes Database (dbgap) at the National Institutes of Health. Survival Analyses. Coded patient survival data was extracted from the TCGA clinical information file. A Cox Proportional Hazards model was used to estimate association between levels of individual mirnas. Patient survival was calculated as time in months elapsed from date of diagnosis until date of last contact. Statistical Analyses. Spearman s rank correlation coefficients, histograms, and the empirical cumulative distribution were computed and plotted for each mirna and gene using r. Sequencing data were log transformed for plotting. Both direct read counts and counts normalized according to millions of mirnas were examined as part of our analyses. All analyses were performed using both raw and normalized read counts reported as part of the TCGA mirna-seq datasets. ACKNOWLEDGEMENTS 8
9 The authors gratefully acknowledge communication from David Wheeler, Rehan Akban, Gordon Robertson and Andy Chu regarding TCGA mirna data analysis algorithms. REFERENCES 1 Wu, L. & Belasco, J. G. Let me count the ways: mechanisms of gene regulation by mirnas and sirnas. Molecular cell 29, 1-7, doi:s (07) [pii] /j.molcel (2008). 2 Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, , doi: /j.cell (2009). 3 Eder, M. & Scherr, M. MicroRNA and lung cancer. The New England journal of medicine 352, , doi: /nejmcibr (2005). 4 Zhang, L. et al. micrornas exhibit high frequency genomic alterations in human cancer. Proc Natl Acad Sci U S A 103, (2006). 5 Creighton, C. J. et al. Molecular profiling uncovers a p53- associated role for microrna- 31 in inhibiting the proliferation of serous ovarian carcinomas and other cancers. Cancer Res 70, , doi: can [pii] / CAN (2010). 6 Integrated genomic analyses of ovarian carcinoma. Nature 474, , doi: /nature10166 (2011). 7 Benjamini, Y. & Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society. Series B (Methodological) 57, , doi: / (1995). 8 Marchini, S. et al. Association between mir- 200c and the survival of patients with stage I epithelial ovarian cancer: a retrospective study of two independent tumour tissue collections. Lancet Oncol 12, , doi: /s (11) (2011). 9 Mateescu, B. et al. mir- 141 and mir- 200a act on ovarian tumorigenesis by controlling oxidative stress response. Nature Medicine 17, , doi: /nm.2512 (2011). 10 Yang, D. et al. Integrated Analyses Identify a Master MicroRNA Regulatory Network for the Mesenchymal Subtype in Serous Ovarian Cancer. Cancer Cell 23, , doi: /j.ccr (2013). 11 Sato, F., Tsuchiya, S., Terasawa, K. & Tsujimoto, G. Intra- platform repeatability and inter- platform comparability of microrna microarray technology. PloS one 4, e5540, doi: /journal.pone (2009). 12 Yauk, C. L., Rowan- Carroll, A., Stead, J. D. & Williams, A. Cross- platform analysis of global microrna expression technologies. BMC genomics 11, 330, doi: / (2010). 13 Shi, L. et al. The MicroArray Quality Control (MAQC) project shows inter- and intraplatform reproducibility of gene expression measurements. Nature Biotechnology 24, , doi: /nbt1239 (2006). 14 Wu, C., Carta, R. & Zhang, L. Sequence dependence of cross- hybridization on short oligo microarrays. Nucleic Acids Research 33, e84, doi: /nar/gni082 (2005). 15 Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, , doi: /nature11252 (2012). 16 Comprehensive genomic characterization of squamous cell lung cancers. Nature 489, , doi: /nature11404 (2012). 9
10 17 Comprehensive molecular portraits of human breast tumours. Nature 490, 61-70, doi: /nature11412 (2012). 10
11 FIGURES FIGURE 1. MicroRNAs associated with ovarian cancer survival. P-value plots of univariate Cox regression for micrornas associated with ovarian cancer survival identified by microarray (A) or mirna-seq (B) data. P-value < 0.01 (Solid line). False discovery rate (FDR) < 0.1 (Dotted line). In both A&B, blue dots indicate mirnas associated with survival by mirna array, while red dots indicate mirnas associated with survival by mir-seq. Green stars are mirnas associated with survival in both datasets. 11
12 FIGURE 2. Scatter-plots of microrna expression measured by microarray and mirna- Seq. (A) mir-98, (B) mir-505 (C) mir-143 and (D) mir
13 FIGURE 3. Distribution of correlations between microarray and sequencing profiles for mirna and gene expression. (A) Histogram of correlation coefficients for individual mirnas measured by mirna-seq and mirna array. (B) Histogram of correlation coefficients for mrnas profiled by Illumina HiSeq and mrna array. (C) The empirical cumulative distribution function (ECDF) of the correlation between array and sequencing for mirna (black) and mrna (gray) measurements. Nearly, 72% of mirnas demonstrate a correlation coefficient 0.5 whereas 22% of RNAs have a correlation coefficient
High AU content: a signature of upregulated mirna in cardiac diseases
 https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
Patnaik SK, et al. MicroRNAs to accurately histotype NSCLC biopsies
 Patnaik SK, et al. MicroRNAs to accurately histotype NSCLC biopsies. 2014. Supplemental Digital Content 1. Appendix 1. External data-sets used for associating microrna expression with lung squamous cell
Patnaik SK, et al. MicroRNAs to accurately histotype NSCLC biopsies. 2014. Supplemental Digital Content 1. Appendix 1. External data-sets used for associating microrna expression with lung squamous cell
HALLA KABAT * Outreach Program, mircore, 2929 Plymouth Rd. Ann Arbor, MI 48105, USA LEO TUNKLE *
 CERNA SEARCH METHOD IDENTIFIED A MET-ACTIVATED SUBGROUP AMONG EGFR DNA AMPLIFIED LUNG ADENOCARCINOMA PATIENTS HALLA KABAT * Outreach Program, mircore, 2929 Plymouth Rd. Ann Arbor, MI 48105, USA Email:
CERNA SEARCH METHOD IDENTIFIED A MET-ACTIVATED SUBGROUP AMONG EGFR DNA AMPLIFIED LUNG ADENOCARCINOMA PATIENTS HALLA KABAT * Outreach Program, mircore, 2929 Plymouth Rd. Ann Arbor, MI 48105, USA Email:
Benchmark study: Exiqon mircury LNA microrna arrays vs. Supplier A s DNA based capture probes. Includes comparison of:
 Benchmark study: Exiqon mircury LNA microrna arrays vs. Supplier A s DNA based capture probes Includes comparison of: I. II. III. Signal levels Relative quantification False differences For complete experimental
Benchmark study: Exiqon mircury LNA microrna arrays vs. Supplier A s DNA based capture probes Includes comparison of: I. II. III. Signal levels Relative quantification False differences For complete experimental
National Surgical Adjuvant Breast and Bowel Project (NSABP) Foundation Annual Progress Report: 2009 Formula Grant
 National Surgical Adjuvant Breast and Bowel Project (NSABP) Foundation Annual Progress Report: 2009 Formula Grant Reporting Period July 1, 2011 June 30, 2012 Formula Grant Overview The National Surgical
National Surgical Adjuvant Breast and Bowel Project (NSABP) Foundation Annual Progress Report: 2009 Formula Grant Reporting Period July 1, 2011 June 30, 2012 Formula Grant Overview The National Surgical
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes
 Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Human breast milk mirna, maternal probiotic supplementation and atopic dermatitis in offsrping
 Human breast milk mirna, maternal probiotic supplementation and atopic dermatitis in offsrping Melanie Rae Simpson PhD candidate Department of Public Health and General Practice Norwegian University of
Human breast milk mirna, maternal probiotic supplementation and atopic dermatitis in offsrping Melanie Rae Simpson PhD candidate Department of Public Health and General Practice Norwegian University of
Digitizing the Proteomes From Big Tissue Biobanks
 Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
MicroRNA and Male Infertility: A Potential for Diagnosis
 Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Inferring condition-specific mirna activity from matched mirna and mrna expression data
 Inferring condition-specific mirna activity from matched mirna and mrna expression data Junpeng Zhang 1, Thuc Duy Le 2, Lin Liu 2, Bing Liu 3, Jianfeng He 4, Gregory J Goodall 5 and Jiuyong Li 2,* 1 Faculty
Inferring condition-specific mirna activity from matched mirna and mrna expression data Junpeng Zhang 1, Thuc Duy Le 2, Lin Liu 2, Bing Liu 3, Jianfeng He 4, Gregory J Goodall 5 and Jiuyong Li 2,* 1 Faculty
MicroRNA expression profiling and functional analysis in prostate cancer. Marco Folini s.c. Ricerca Traslazionale DOSL
 MicroRNA expression profiling and functional analysis in prostate cancer Marco Folini s.c. Ricerca Traslazionale DOSL What are micrornas? For almost three decades, the alteration of protein-coding genes
MicroRNA expression profiling and functional analysis in prostate cancer Marco Folini s.c. Ricerca Traslazionale DOSL What are micrornas? For almost three decades, the alteration of protein-coding genes
Introduction. Introduction
 Introduction We are leveraging genome sequencing data from The Cancer Genome Atlas (TCGA) to more accurately define mutated and stable genes and dysregulated metabolic pathways in solid tumors. These efforts
Introduction We are leveraging genome sequencing data from The Cancer Genome Atlas (TCGA) to more accurately define mutated and stable genes and dysregulated metabolic pathways in solid tumors. These efforts
Genetic alterations of histone lysine methyltransferases and their significance in breast cancer
 Genetic alterations of histone lysine methyltransferases and their significance in breast cancer Supplementary Materials and Methods Phylogenetic tree of the HMT superfamily The phylogeny outlined in the
Genetic alterations of histone lysine methyltransferases and their significance in breast cancer Supplementary Materials and Methods Phylogenetic tree of the HMT superfamily The phylogeny outlined in the
Gene-microRNA network module analysis for ovarian cancer
 Gene-microRNA network module analysis for ovarian cancer Shuqin Zhang School of Mathematical Sciences Fudan University Oct. 4, 2016 Outline Introduction Materials and Methods Results Conclusions Introduction
Gene-microRNA network module analysis for ovarian cancer Shuqin Zhang School of Mathematical Sciences Fudan University Oct. 4, 2016 Outline Introduction Materials and Methods Results Conclusions Introduction
Simple, rapid, and reliable RNA sequencing
 Simple, rapid, and reliable RNA sequencing RNA sequencing applications RNA sequencing provides fundamental insights into how genomes are organized and regulated, giving us valuable information about the
Simple, rapid, and reliable RNA sequencing RNA sequencing applications RNA sequencing provides fundamental insights into how genomes are organized and regulated, giving us valuable information about the
developing new tools for diagnostics Join forces with IMGM Laboratories to make your mirna project a success
 micrornas developing new tools for diagnostics Join forces with IMGM Laboratories to make your mirna project a success Dr. Carola Wagner IMGM Laboratories GmbH Martinsried, Germany qpcr 2009 Symposium
micrornas developing new tools for diagnostics Join forces with IMGM Laboratories to make your mirna project a success Dr. Carola Wagner IMGM Laboratories GmbH Martinsried, Germany qpcr 2009 Symposium
Li-Xuan Qin 1* and Douglas A. Levine 2
 Qin and Levine BMC Medical Genomics (2016) 9:27 DOI 10.1186/s12920-016-0187-4 RESEARCH ARTICLE Open Access Study design and data analysis considerations for the discovery of prognostic molecular biomarkers:
Qin and Levine BMC Medical Genomics (2016) 9:27 DOI 10.1186/s12920-016-0187-4 RESEARCH ARTICLE Open Access Study design and data analysis considerations for the discovery of prognostic molecular biomarkers:
The Cancer Genome Atlas & International Cancer Genome Consortium
 The Cancer Genome Atlas & International Cancer Genome Consortium Session 3 Dr Jason Wong Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 31 st July 2014 1
The Cancer Genome Atlas & International Cancer Genome Consortium Session 3 Dr Jason Wong Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 31 st July 2014 1
The Biology and Genetics of Cells and Organisms The Biology of Cancer
 The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Session 4 Rebecca Poulos
 The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 20
The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 20
Session 4 Rebecca Poulos
 The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 28
The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 28
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
National Surgical Adjuvant Breast and Bowel Project (NSABP) Foundation Annual Progress Report: 2009 Formula Grant
 National Surgical Adjuvant Breast and Bowel Project (NSABP) Foundation Annual Progress Report: 2009 Formula Grant Reporting Period July 1, 2012 June 30, 2013 Formula Grant Overview The National Surgical
National Surgical Adjuvant Breast and Bowel Project (NSABP) Foundation Annual Progress Report: 2009 Formula Grant Reporting Period July 1, 2012 June 30, 2013 Formula Grant Overview The National Surgical
SUPPLEMENTARY FIGURE LEGENDS
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1 Negative correlation between mir-375 and its predicted target genes, as demonstrated by gene set enrichment analysis (GSEA). 1 The correlation between
SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1 Negative correlation between mir-375 and its predicted target genes, as demonstrated by gene set enrichment analysis (GSEA). 1 The correlation between
ncounter Assay Automated Process Immobilize and align reporter for image collecting and barcode counting ncounter Prep Station
 ncounter Assay ncounter Prep Station Automated Process Hybridize Reporter to RNA Remove excess reporters Bind reporter to surface Immobilize and align reporter Image surface Count codes Immobilize and
ncounter Assay ncounter Prep Station Automated Process Hybridize Reporter to RNA Remove excess reporters Bind reporter to surface Immobilize and align reporter Image surface Count codes Immobilize and
Profiles of gene expression & diagnosis/prognosis of cancer. MCs in Advanced Genetics Ainoa Planas Riverola
 Profiles of gene expression & diagnosis/prognosis of cancer MCs in Advanced Genetics Ainoa Planas Riverola Gene expression profiles Gene expression profiling Used in molecular biology, it measures the
Profiles of gene expression & diagnosis/prognosis of cancer MCs in Advanced Genetics Ainoa Planas Riverola Gene expression profiles Gene expression profiling Used in molecular biology, it measures the
he micrornas of Caenorhabditis elegans (Lim et al. Genes & Development 2003)
 MicroRNAs: Genomics, Biogenesis, Mechanism, and Function (D. Bartel Cell 2004) he micrornas of Caenorhabditis elegans (Lim et al. Genes & Development 2003) Vertebrate MicroRNA Genes (Lim et al. Science
MicroRNAs: Genomics, Biogenesis, Mechanism, and Function (D. Bartel Cell 2004) he micrornas of Caenorhabditis elegans (Lim et al. Genes & Development 2003) Vertebrate MicroRNA Genes (Lim et al. Science
STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells
 CAMDA 2009 October 5, 2009 STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells Guohua Wang 1, Yadong Wang 1, Denan Zhang 1, Mingxiang Teng 1,2, Lang Li 2, and Yunlong Liu 2 Harbin
CAMDA 2009 October 5, 2009 STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells Guohua Wang 1, Yadong Wang 1, Denan Zhang 1, Mingxiang Teng 1,2, Lang Li 2, and Yunlong Liu 2 Harbin
The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis
 The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis Tieliu Shi tlshi@bio.ecnu.edu.cn The Center for bioinformatics
The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis Tieliu Shi tlshi@bio.ecnu.edu.cn The Center for bioinformatics
MicroRNA-mediated incoherent feedforward motifs are robust
 The Second International Symposium on Optimization and Systems Biology (OSB 8) Lijiang, China, October 31 November 3, 8 Copyright 8 ORSC & APORC, pp. 62 67 MicroRNA-mediated incoherent feedforward motifs
The Second International Symposium on Optimization and Systems Biology (OSB 8) Lijiang, China, October 31 November 3, 8 Copyright 8 ORSC & APORC, pp. 62 67 MicroRNA-mediated incoherent feedforward motifs
SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer.
 Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
m 6 A mrna methylation regulates AKT activity to promote the proliferation and tumorigenicity of endometrial cancer
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
Bi 8 Lecture 17. interference. Ellen Rothenberg 1 March 2016
 Bi 8 Lecture 17 REGulation by RNA interference Ellen Rothenberg 1 March 2016 Protein is not the only regulatory molecule affecting gene expression: RNA itself can be negative regulator RNA does not need
Bi 8 Lecture 17 REGulation by RNA interference Ellen Rothenberg 1 March 2016 Protein is not the only regulatory molecule affecting gene expression: RNA itself can be negative regulator RNA does not need
microrna PCR System (Exiqon), following the manufacturer s instructions. In brief, 10ng of
 SUPPLEMENTAL MATERIALS AND METHODS Quantitative RT-PCR Quantitative RT-PCR analysis was performed using the Universal mircury LNA TM microrna PCR System (Exiqon), following the manufacturer s instructions.
SUPPLEMENTAL MATERIALS AND METHODS Quantitative RT-PCR Quantitative RT-PCR analysis was performed using the Universal mircury LNA TM microrna PCR System (Exiqon), following the manufacturer s instructions.
microrna Presented for: Presented by: Date:
 microrna Presented for: Presented by: Date: 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions
microrna Presented for: Presented by: Date: 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded
 Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
A Panoptic Approach to Studying MicroRNA Expression in Breast. Tumors. Priyanka Bhatt
 A Panoptic Approach to Studying MicroRNA Expression in Breast Tumors By Priyanka Bhatt A Dissertation Submitted to Rutgers University School of Health Related Professions In partial fulfillment of the
A Panoptic Approach to Studying MicroRNA Expression in Breast Tumors By Priyanka Bhatt A Dissertation Submitted to Rutgers University School of Health Related Professions In partial fulfillment of the
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Multi-omics data integration colon cancer using proteogenomics approach
 Dept. of Medical Oncology Multi-omics data integration colon cancer using proteogenomics approach DTL Focus meeting, 29 August 2016 Thang Pham OncoProteomics Laboratory, Dept. of Medical Oncology VU University
Dept. of Medical Oncology Multi-omics data integration colon cancer using proteogenomics approach DTL Focus meeting, 29 August 2016 Thang Pham OncoProteomics Laboratory, Dept. of Medical Oncology VU University
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
OVARIAN CANCER POSSIBLE TUMOR-SUPPRESSIVE ROLE OF BATF2 IN OVARIAN CANCER
 POSSIBLE TUMOR-SUPPRESSIVE ROLE OF BATF2 IN OVARIAN CANCER Rissa Fedora, OMS-III OVARIAN CANCER 5 th leading cause of death among American women Risk factors: Age Gender female Greater lifetime ovulations
POSSIBLE TUMOR-SUPPRESSIVE ROLE OF BATF2 IN OVARIAN CANCER Rissa Fedora, OMS-III OVARIAN CANCER 5 th leading cause of death among American women Risk factors: Age Gender female Greater lifetime ovulations
Clinical Oncology - Science in focus - Editorial. Understanding oestrogen receptor function in breast cancer, and its interaction with the
 Clinical Oncology - Science in focus - Editorial TITLE: Understanding oestrogen receptor function in breast cancer, and its interaction with the progesterone receptor. New preclinical findings and their
Clinical Oncology - Science in focus - Editorial TITLE: Understanding oestrogen receptor function in breast cancer, and its interaction with the progesterone receptor. New preclinical findings and their
Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities
 Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities Gábor Boross,2, Katalin Orosz,2 and Illés J. Farkas 2, Department of Biological Physics,
Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities Gábor Boross,2, Katalin Orosz,2 and Illés J. Farkas 2, Department of Biological Physics,
Obstacles and challenges in the analysis of microrna sequencing data
 Obstacles and challenges in the analysis of microrna sequencing data (mirna-seq) David Humphreys Genomics core Dr Victor Chang AC 1936-1991, Pioneering Cardiothoracic Surgeon and Humanitarian The ABCs
Obstacles and challenges in the analysis of microrna sequencing data (mirna-seq) David Humphreys Genomics core Dr Victor Chang AC 1936-1991, Pioneering Cardiothoracic Surgeon and Humanitarian The ABCs
Cancer Cell Research 19 (2018)
 Available at http:// www.cancercellresearch.org ISSN 2161-2609 LncRNA RP11-597D13.9 expression and clinical significance in serous Ovarian Cancer based on TCGA database Xiaoyan Gu 1, Pengfei Xu 2, Sujuan
Available at http:// www.cancercellresearch.org ISSN 2161-2609 LncRNA RP11-597D13.9 expression and clinical significance in serous Ovarian Cancer based on TCGA database Xiaoyan Gu 1, Pengfei Xu 2, Sujuan
mirna Whole Transcriptome Assay
 mirna Whole Transcriptome Assay HTG EdgeSeq mirna Whole Transciptome Assay The HTG EdgeSeq mirna Whole Transcriptome Assay (WTA) is a next generation sequencing (NGS) application that measures the expression
mirna Whole Transcriptome Assay HTG EdgeSeq mirna Whole Transciptome Assay The HTG EdgeSeq mirna Whole Transcriptome Assay (WTA) is a next generation sequencing (NGS) application that measures the expression
Statistical Assessment of the Global Regulatory Role of Histone. Acetylation in Saccharomyces cerevisiae. (Support Information)
 Statistical Assessment of the Global Regulatory Role of Histone Acetylation in Saccharomyces cerevisiae (Support Information) Authors: Guo-Cheng Yuan, Ping Ma, Wenxuan Zhong and Jun S. Liu Linear Relationship
Statistical Assessment of the Global Regulatory Role of Histone Acetylation in Saccharomyces cerevisiae (Support Information) Authors: Guo-Cheng Yuan, Ping Ma, Wenxuan Zhong and Jun S. Liu Linear Relationship
micrornas (mirna) and Biomarkers
 micrornas (mirna) and Biomarkers Small RNAs Make Big Splash mirnas & Genome Function Biomarkers in Cancer Future Prospects Javed Khan M.D. National Cancer Institute EORTC-NCI-ASCO November 2007 The Human
micrornas (mirna) and Biomarkers Small RNAs Make Big Splash mirnas & Genome Function Biomarkers in Cancer Future Prospects Javed Khan M.D. National Cancer Institute EORTC-NCI-ASCO November 2007 The Human
starbase Pan-Cancer Tutorial
 starbase Pan-Cancer Tutorial 2010-2013, Jianhua Yang at Sun Yat-sen University For comments, suggestions on the starbase platform, please contact us: lsp03yjh@gmail.com On commercial use, contact: yangjh7@mail.sysu.edu.cn
starbase Pan-Cancer Tutorial 2010-2013, Jianhua Yang at Sun Yat-sen University For comments, suggestions on the starbase platform, please contact us: lsp03yjh@gmail.com On commercial use, contact: yangjh7@mail.sysu.edu.cn
A Statistical Framework for Classification of Tumor Type from microrna Data
 DEGREE PROJECT IN MATHEMATICS, SECOND CYCLE, 30 CREDITS STOCKHOLM, SWEDEN 2016 A Statistical Framework for Classification of Tumor Type from microrna Data JOSEFINE RÖHSS KTH ROYAL INSTITUTE OF TECHNOLOGY
DEGREE PROJECT IN MATHEMATICS, SECOND CYCLE, 30 CREDITS STOCKHOLM, SWEDEN 2016 A Statistical Framework for Classification of Tumor Type from microrna Data JOSEFINE RÖHSS KTH ROYAL INSTITUTE OF TECHNOLOGY
Reduced expression of SOX7 in ovarian cancer: a novel tumor suppressor through the Wnt/β-catenin signaling pathway
 Reduced expression of SOX7 in ovarian cancer: a novel tumor suppressor through the Wnt/β-catenin signaling pathway Huidi Liu, M.D. & Ph.D Genomics Research Centre Harbin Medical University, China 15804606535@163.com
Reduced expression of SOX7 in ovarian cancer: a novel tumor suppressor through the Wnt/β-catenin signaling pathway Huidi Liu, M.D. & Ph.D Genomics Research Centre Harbin Medical University, China 15804606535@163.com
Screening for novel oncology biomarker panels using both DNA and protein microarrays. John Anson, PhD VP Biomarker Discovery
 Screening for novel oncology biomarker panels using both DNA and protein microarrays John Anson, PhD VP Biomarker Discovery Outline of presentation Introduction to OGT and our approach to biomarker studies
Screening for novel oncology biomarker panels using both DNA and protein microarrays John Anson, PhD VP Biomarker Discovery Outline of presentation Introduction to OGT and our approach to biomarker studies
MicroRNA in Cancer Karen Dybkær 2013
 MicroRNA in Cancer Karen Dybkær RNA Ribonucleic acid Types -Coding: messenger RNA (mrna) coding for proteins -Non-coding regulating protein formation Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
MicroRNA in Cancer Karen Dybkær RNA Ribonucleic acid Types -Coding: messenger RNA (mrna) coding for proteins -Non-coding regulating protein formation Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
MicroRNA dysregulation in cancer. Systems Plant Microbiology Hyun-Hee Lee
 MicroRNA dysregulation in cancer Systems Plant Microbiology Hyun-Hee Lee Contents 1 What is MicroRNA? 2 mirna dysregulation in cancer 3 Summary What is MicroRNA? What is MicroRNA? MicroRNAs (mirnas) -
MicroRNA dysregulation in cancer Systems Plant Microbiology Hyun-Hee Lee Contents 1 What is MicroRNA? 2 mirna dysregulation in cancer 3 Summary What is MicroRNA? What is MicroRNA? MicroRNAs (mirnas) -
genomics for systems biology / ISB2020 RNA sequencing (RNA-seq)
 RNA sequencing (RNA-seq) Module Outline MO 13-Mar-2017 RNA sequencing: Introduction 1 WE 15-Mar-2017 RNA sequencing: Introduction 2 MO 20-Mar-2017 Paper: PMID 25954002: Human genomics. The human transcriptome
RNA sequencing (RNA-seq) Module Outline MO 13-Mar-2017 RNA sequencing: Introduction 1 WE 15-Mar-2017 RNA sequencing: Introduction 2 MO 20-Mar-2017 Paper: PMID 25954002: Human genomics. The human transcriptome
Deciphering the Role of micrornas in BRD4-NUT Fusion Gene Induced NUT Midline Carcinoma
 www.bioinformation.net Volume 13(6) Hypothesis Deciphering the Role of micrornas in BRD4-NUT Fusion Gene Induced NUT Midline Carcinoma Ekta Pathak 1, Bhavya 1, Divya Mishra 1, Neelam Atri 1, 2, Rajeev
www.bioinformation.net Volume 13(6) Hypothesis Deciphering the Role of micrornas in BRD4-NUT Fusion Gene Induced NUT Midline Carcinoma Ekta Pathak 1, Bhavya 1, Divya Mishra 1, Neelam Atri 1, 2, Rajeev
Transcriptome Analysis
 Transcriptome Analysis Data Preprocessing Sample Preparation Illumina Sequencing Demultiplexing Raw FastQ Reference Genome (fasta) Reference Annotation (GTF) Reference Genome Analysis Tophat Accepted hits
Transcriptome Analysis Data Preprocessing Sample Preparation Illumina Sequencing Demultiplexing Raw FastQ Reference Genome (fasta) Reference Annotation (GTF) Reference Genome Analysis Tophat Accepted hits
Set the stage: Genomics technology. Jos Kleinjans Dept of Toxicogenomics Maastricht University, the Netherlands
 Set the stage: Genomics technology Jos Kleinjans Dept of Toxicogenomics Maastricht University, the Netherlands Amendment to the latest consolidated version of the REACH legislation REACH Regulation 1907/2006:
Set the stage: Genomics technology Jos Kleinjans Dept of Toxicogenomics Maastricht University, the Netherlands Amendment to the latest consolidated version of the REACH legislation REACH Regulation 1907/2006:
Inferring Biological Meaning from Cap Analysis Gene Expression Data
 Inferring Biological Meaning from Cap Analysis Gene Expression Data HRYSOULA PAPADAKIS 1. Introduction This project is inspired by the recent development of the Cap analysis gene expression (CAGE) method,
Inferring Biological Meaning from Cap Analysis Gene Expression Data HRYSOULA PAPADAKIS 1. Introduction This project is inspired by the recent development of the Cap analysis gene expression (CAGE) method,
Introduction to Systems Biology of Cancer Lecture 2
 Introduction to Systems Biology of Cancer Lecture 2 Gustavo Stolovitzky IBM Research Icahn School of Medicine at Mt Sinai DREAM Challenges High throughput measurements: The age of omics Systems Biology
Introduction to Systems Biology of Cancer Lecture 2 Gustavo Stolovitzky IBM Research Icahn School of Medicine at Mt Sinai DREAM Challenges High throughput measurements: The age of omics Systems Biology
Integrated Analysis of Copy Number and Gene Expression
 Integrated Analysis of Copy Number and Gene Expression Nexus Copy Number provides user-friendly interface and functionalities to integrate copy number analysis with gene expression results for the purpose
Integrated Analysis of Copy Number and Gene Expression Nexus Copy Number provides user-friendly interface and functionalities to integrate copy number analysis with gene expression results for the purpose
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library
 Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Identification of mirnas in Eucalyptus globulus Plant by Computational Methods
 International Journal of Pharmaceutical Science Invention ISSN (Online): 2319 6718, ISSN (Print): 2319 670X Volume 2 Issue 5 May 2013 PP.70-74 Identification of mirnas in Eucalyptus globulus Plant by Computational
International Journal of Pharmaceutical Science Invention ISSN (Online): 2319 6718, ISSN (Print): 2319 670X Volume 2 Issue 5 May 2013 PP.70-74 Identification of mirnas in Eucalyptus globulus Plant by Computational
Computational Approaches to Transcriptome Signatures in the Human Brain
 Computational Approaches to Transcriptome Signatures in the Human Brain Agilent Technologies eseminar February 18, 2016 Mike Hawrylycz, Ph.D. ALLEN Adult Human Atlas Online tools An Anatomic Transcriptional
Computational Approaches to Transcriptome Signatures in the Human Brain Agilent Technologies eseminar February 18, 2016 Mike Hawrylycz, Ph.D. ALLEN Adult Human Atlas Online tools An Anatomic Transcriptional
Single SNP/Gene Analysis. Typical Results of GWAS Analysis (Single SNP Approach) Typical Results of GWAS Analysis (Single SNP Approach)
 High-Throughput Sequencing Course Gene-Set Analysis Biostatistics and Bioinformatics Summer 28 Section Introduction What is Gene Set Analysis? Many names for gene set analysis: Pathway analysis Gene set
High-Throughput Sequencing Course Gene-Set Analysis Biostatistics and Bioinformatics Summer 28 Section Introduction What is Gene Set Analysis? Many names for gene set analysis: Pathway analysis Gene set
Integrated genomic analysis of human osteosarcomas
 Integrated genomic analysis of human osteosarcomas Leonardo A. Meza-Zepeda Project Leader Genomic Section Department of Tumor Biology The Norwegian Radium Hospital Head Microarray Core Facility Norwegian
Integrated genomic analysis of human osteosarcomas Leonardo A. Meza-Zepeda Project Leader Genomic Section Department of Tumor Biology The Norwegian Radium Hospital Head Microarray Core Facility Norwegian
ncounter Assay Automated Process Capture & Reporter Probes Bind reporter to surface Remove excess reporters Hybridize CodeSet to RNA
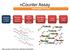 ncounter Assay Automated Process Hybridize CodeSet to RNA Remove excess reporters Bind reporter to surface Immobilize and align reporter Image surface Count codes mrna Capture & Reporter Probes slides
ncounter Assay Automated Process Hybridize CodeSet to RNA Remove excess reporters Bind reporter to surface Immobilize and align reporter Image surface Count codes mrna Capture & Reporter Probes slides
Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well
 Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well plate at cell densities ranging from 25-225 cells in
Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well plate at cell densities ranging from 25-225 cells in
Identification of Tissue Independent Cancer Driver Genes
 Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies
 OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
Identifying microrna signatures of chemo-resistance in colorectal cancer
 Identifying microrna signatures of chemo-resistance in colorectal cancer Presented by: Team Miscreant (MicroRNA Screening and Targeting) Allan Lo, Andy Mungall, Angela Hussainkhel, Linh Phan, Simon Haile
Identifying microrna signatures of chemo-resistance in colorectal cancer Presented by: Team Miscreant (MicroRNA Screening and Targeting) Allan Lo, Andy Mungall, Angela Hussainkhel, Linh Phan, Simon Haile
CONTRACTING ORGANIZATION: Cold Spring Harbor Laboratory Cold Spring Harbor, NY 11724
 AD Award Number: W81XWH-06-1-0249 TITLE: Detection of genes modifying sensitivity to proteasome inhibitors using a shrna Library in Breast Cancer PRINCIPAL INVESTIGATOR: Gregory J. Hannon, Ph.D. CONTRACTING
AD Award Number: W81XWH-06-1-0249 TITLE: Detection of genes modifying sensitivity to proteasome inhibitors using a shrna Library in Breast Cancer PRINCIPAL INVESTIGATOR: Gregory J. Hannon, Ph.D. CONTRACTING
Gene Ontology and Functional Enrichment. Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein
 Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Cancer Problems in Indonesia
 mirna and Cancer : mirna as a Key Regulator in Cancer Sofia Mubarika 2 nd Symposium Biomolecular Update in Cancer PERABOI Padang 18 Mei 2013 Cancer Problems in Indonesia 1. Chemoresistency / recurrency
mirna and Cancer : mirna as a Key Regulator in Cancer Sofia Mubarika 2 nd Symposium Biomolecular Update in Cancer PERABOI Padang 18 Mei 2013 Cancer Problems in Indonesia 1. Chemoresistency / recurrency
Analysis of RNA Expression of Normal and Cancer Tissues Reveals High Correlation of COP9 Gene Expression with Respiratory Chain Complex Components
 University of Kentucky UKnowledge Toxicology and Cancer Biology Faculty Publications Toxicology and Cancer Biology 12-1-2016 Analysis of RNA Expression of Normal and Cancer Tissues Reveals High Correlation
University of Kentucky UKnowledge Toxicology and Cancer Biology Faculty Publications Toxicology and Cancer Biology 12-1-2016 Analysis of RNA Expression of Normal and Cancer Tissues Reveals High Correlation
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression. Chromatin Array of nuc
 Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Breast cancer. Risk factors you cannot change include: Treatment Plan Selection. Inferring Transcriptional Module from Breast Cancer Profile Data
 Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
Mature microrna identification via the use of a Naive Bayes classifier
 Mature microrna identification via the use of a Naive Bayes classifier Master Thesis Gkirtzou Katerina Computer Science Department University of Crete 13/03/2009 Gkirtzou K. (CSD UOC) Mature microrna identification
Mature microrna identification via the use of a Naive Bayes classifier Master Thesis Gkirtzou Katerina Computer Science Department University of Crete 13/03/2009 Gkirtzou K. (CSD UOC) Mature microrna identification
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
TCGA. The Cancer Genome Atlas
 TCGA The Cancer Genome Atlas TCGA: History and Goal History: Started in 2005 by the National Cancer Institute (NCI) and the National Human Genome Research Institute (NHGRI) with $110 Million to catalogue
TCGA The Cancer Genome Atlas TCGA: History and Goal History: Started in 2005 by the National Cancer Institute (NCI) and the National Human Genome Research Institute (NHGRI) with $110 Million to catalogue
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project
 Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
CRS4 Seminar series. Inferring the functional role of micrornas from gene expression data CRS4. Biomedicine. Bioinformatics. Paolo Uva July 11, 2012
 CRS4 Seminar series Inferring the functional role of micrornas from gene expression data CRS4 Biomedicine Bioinformatics Paolo Uva July 11, 2012 Partners Pharmaceutical company Fondazione San Raffaele,
CRS4 Seminar series Inferring the functional role of micrornas from gene expression data CRS4 Biomedicine Bioinformatics Paolo Uva July 11, 2012 Partners Pharmaceutical company Fondazione San Raffaele,
DiffVar: a new method for detecting differential variability with application to methylation in cancer and aging
 Genome Biology This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. DiffVar: a new method for detecting
Genome Biology This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. DiffVar: a new method for detecting
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Expert-guided Visual Exploration (EVE) for patient stratification. Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland
 Expert-guided Visual Exploration (EVE) for patient stratification Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland Oncoscape.sttrcancer.org Paul Lisa Ken Jenny Desert Eric The challenge Given - patient clinical
Expert-guided Visual Exploration (EVE) for patient stratification Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland Oncoscape.sttrcancer.org Paul Lisa Ken Jenny Desert Eric The challenge Given - patient clinical
FISH mcgh Karyotyping ISH RT-PCR. Expression arrays RNA. Tissue microarrays Protein arrays MS. Protein IHC
 Classification of Breast Cancer in the Molecular Era Susan J. Done University Health Network, Toronto Why classify? Prognosis Prediction of response to therapy Pathogenesis Invasive breast cancer can have
Classification of Breast Cancer in the Molecular Era Susan J. Done University Health Network, Toronto Why classify? Prognosis Prediction of response to therapy Pathogenesis Invasive breast cancer can have
Cellecta Overview. Started Operations in 2007 Headquarters: Mountain View, CA
 Cellecta Overview Started Operations in 2007 Headquarters: Mountain View, CA Focus: Development of flexible, scalable, and broadly parallel genetic screening assays to expedite the discovery and characterization
Cellecta Overview Started Operations in 2007 Headquarters: Mountain View, CA Focus: Development of flexible, scalable, and broadly parallel genetic screening assays to expedite the discovery and characterization
Association of mir-21 with esophageal cancer prognosis: a meta-analysis
 Association of mir-21 with esophageal cancer prognosis: a meta-analysis S.-W. Wen 1, Y.-F. Zhang 1, Y. Li 1, Z.-X. Liu 2, H.-L. Lv 1, Z.-H. Li 1, Y.-Z. Xu 1, Y.-G. Zhu 1 and Z.-Q. Tian 1 1 Department of
Association of mir-21 with esophageal cancer prognosis: a meta-analysis S.-W. Wen 1, Y.-F. Zhang 1, Y. Li 1, Z.-X. Liu 2, H.-L. Lv 1, Z.-H. Li 1, Y.-Z. Xu 1, Y.-G. Zhu 1 and Z.-Q. Tian 1 1 Department of
Title: Human breast cancer associated fibroblasts exhibit subtype specific gene expression profiles
 Author's response to reviews Title: Human breast cancer associated fibroblasts exhibit subtype specific gene expression profiles Authors: Julia Tchou (julia.tchou@uphs.upenn.edu) Andrew V Kossenkov (akossenkov@wistar.org)
Author's response to reviews Title: Human breast cancer associated fibroblasts exhibit subtype specific gene expression profiles Authors: Julia Tchou (julia.tchou@uphs.upenn.edu) Andrew V Kossenkov (akossenkov@wistar.org)
omiras: MicroRNA regulation of gene expression
 omiras: MicroRNA regulation of gene expression Sören Müller, Goethe University of Frankfurt am Main Molecular Bioinformatics Group, Institute of Computer Science Plant Molecular Biology Group, Institute
omiras: MicroRNA regulation of gene expression Sören Müller, Goethe University of Frankfurt am Main Molecular Bioinformatics Group, Institute of Computer Science Plant Molecular Biology Group, Institute
A novel and universal method for microrna RT-qPCR data normalization
 A novel and universal method for microrna RT-qPCR data normalization Jo Vandesompele professor, Ghent University co-founder and CEO, Biogazelle 4 th International qpcr Symposium Weihenstephan, March 1,
A novel and universal method for microrna RT-qPCR data normalization Jo Vandesompele professor, Ghent University co-founder and CEO, Biogazelle 4 th International qpcr Symposium Weihenstephan, March 1,
About OMICS Group Conferences
 About OMICS Group OMICS Group International is an amalgamation of Open Access publications and worldwide international science conferences and events. Established in the year 2007 with the sole aim of
About OMICS Group OMICS Group International is an amalgamation of Open Access publications and worldwide international science conferences and events. Established in the year 2007 with the sole aim of
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
The Cancer Genome Atlas
 The Cancer Genome Atlas July 14, 2011 Kenna M. Shaw, Ph.D. Deputy Director The Cancer Genome Atlas Program TCGA: Core Objectives Launched in 2006 as a pilot and expanded in 2009, the goals of TCGA are
The Cancer Genome Atlas July 14, 2011 Kenna M. Shaw, Ph.D. Deputy Director The Cancer Genome Atlas Program TCGA: Core Objectives Launched in 2006 as a pilot and expanded in 2009, the goals of TCGA are
RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
 Xu et al. Molecular Cancer (2019) 18:8 https://doi.org/10.1186/s12943-018-0932-8 LETTER TO THE EDITOR RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
Xu et al. Molecular Cancer (2019) 18:8 https://doi.org/10.1186/s12943-018-0932-8 LETTER TO THE EDITOR RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
