Using a proteomic approach to identify proteasome interacting proteins in mammalian
|
|
|
- Estella Haynes
- 5 years ago
- Views:
Transcription
1 Supplementary Discussion Using a proteomic approach to identify proteasome interacting proteins in mammalian cells, we describe in the present study a novel chaperone complex that plays a key role in the 5 assembly of mammalian 20S proteasomes, named PAC1 and PAC2. The assembly of 20S proteasomes is a multistep-ordered process, and presumptive α-ring formation represents the initial step. At present, the process of α-ring formation is considered spontaneous. This is based on the observations that the α subunit of archaebacteria 1, Trypanosoma brucei 2 and human 3 can form homomeric seven-membered 10 rings when expressed in E. coli. However, no homoheptameric rings occur in vivo in eukaryotic proteasomes, whose α-ring is invariably composed of seven different α subunits. In addition, in yeast all α subunits except α3 are essential for life 4-6. These observations imply that there is a mechanism that correctly arranges the seven α subunits into a heteroheptamer, but such mechanism had remained entirely unknown. 15 In the present study, we identified the complex containing all seven α subunits but no β subunits with the size of approximately 230 kda, which is most likely to represent α-rings associated with PAC1/PAC2 heterodimer (Fig. 1). The PAC complex was most abundantly associated with α-rings, indicating its role in α-ring assembly. Indeed, IVTT experiment suggested that the PAC complex promoted the gathering of α subunits (Supplementary Fig. 20 4). In cells, we demonstrated that it was also bound to early α subunit assembly intermediates that contained a restricted subset of α subunits. These results indicate that the formation of α-rings of mammalian 20S proteasomes is chaperoned by the PAC1/PAC2 complex.
2 Knockdown of the PAC complex revealed its additional but important role in proteasome assembly. Glycerol gradient analysis demonstrated accumulation of α subunits in 25 fractions corresponding to half-proteasomes without any accompanying increase in pro-β subunits or hump1 (Fig. 3b), which are hallmarks of half-proteasomes. Indeed, this fraction in PAC-knockdown cells contained much smaller amounts of pro-β subunits and hump1 relative to α subunits (Fig. 3c). Considering its size and content, it is most likely that this fraction results from dimerization of α-rings. It is uncertain whether the ring is normally 30 organized or not in the absence of the PAC complex, but considering the milder defect in proteasome function in PAC-knockdown cells than hump1-knockdown cells, at least part of the α-rings was normal, which could be incorporated into functional half-proteasomes and 20S proteasomes. Since human α subunits expressed in E. coli tend to form a double ring-like structure 3, prevention of formation of double α-ring can be important for proper assembly of 35 20S proteasomes (see our model, Fig. 4d). Our data indicate that the PAC complex is responsible for suppressing the formation of off-pathway, non-productive α-ring dimers. As for chaperones for the 20S proteasome assembly, it has been shown that Ump1 is included in half-proteasomes and is required for the coordination between correct maturation of β subunits and dimerization of half-proteasomes in yeast 7. Several studies have suggested 40 that the mammalian homolog of Ump1 operates in a similar fashion. We investigated the relationship between the PAC complex and hump1. The PAC complex was directly associated with at least some of the α subunits (Supplementary Fig. 4a), whereas hump1 is reported to bind to certain β subunits but not to any of α subunits in yeast two hybrid analysis 8. Glycerol gradient analysis of cell extracts revealed that hump1 specifically 45 associates with half-proteasomes and is not associated with α-rings, unlike the PAC complex
3 (Fig. 1). Though both the PAC complex and hump1 were included in half-proteasomes, they did not directly associate with each other in vitro (Supplementary Fig. 2c). RNAi experiments further distinguished their functional differences. Knockdown of PACs resulted in loss of normal α-rings, whereas knockdown of hump1 was associated with defective dimerization of 50 half-proteasomes though the α-rings and half-proteasomes were maintained. These results demonstrate that the PAC complex is a chaperone devoted to α-rings, whereas hump1 is dedicated to half-proteasome formation, taking over the duty of the PAC complex (Fig. 4d). MG132 treatment elicited another difference. The PAC complex accumulated in 20S proteasome fractions, whereas hump1 did not. hump1 is likely to be located at the interface 55 between two β-rings, as predicted in yeast analysis 7, and its degradation is tightly coupled with 20S proteasome formation. On the other hand, the PAC complex is likely to be located at the side of α-ring opposite to hump1 and β-ring, since it directly binds to α subunits and is still a component of half-proteasomes without inhibiting the assembly of hump1 and β subunits on α-rings. This indicates that the PAC complex constitutively binds α subunits 60 whether in precursor or mature proteasomes and that it is continually degraded by 20S proteasomes. Therefore, when proteasome activity is inhibited, PACs still associate with the ends of 20S proteasomes (Figs. 1 and 4d). The precise mechanism of PAC1/PAC2 degradation by 20S proteasomes is not clear at present, but it is plausible that 20S proteasomes directly degrade PAC1/PAC2 through a mechanism similar to that employed for 65 the degradation of naturally unfolded polypeptides 9 such as α-synuclein 10, tau 11 and p21 Cip1 (refs 12, 13), or localization of the PAC complex to the proteasome may be sufficient for degradation 14. In this context, it is intriguing whether triplication of PAC1 gene in Down
4 syndrome overloads proteasomes and contributes to the pathogenesis of various disease entities. 70 Another question is whether this system is conserved throughout eukaryotes. PAC2 seems to have a yeast homolog, YKL206C, with 19% identity to the mammalian gene. YKL206C is known to interact with proteasomes, but its null mutant is viable and showed no obvious phenotypes (data not shown). Thus, YKL206C might be the ancestral gene of PAC2 but does not appear to be a functional homolog. As for PAC1, we could find its orthologous 75 genes only in vertebrates. Whereas the individual α subunits and β subunits are approximately 50-60% identical in humans and yeast, Ump1 and PAC2 are only 27% and 19% identical, respectively. Therefore, the apparent lack of yeast or invertebrate orthologues of human PAC1 may simply reflect low amino acid sequence conservation. Recent proteomic approaches in yeast have identified many proteasome interacting proteins with yet unknown 80 functions, among which there may be a functional orthologue of PAC1. Alternatively, it is possible that PAC1 is an invention of vertebrates because there are significant differences between human and yeast in the assembly of 20S proteasomes. As we reported here, hump1 did not accumulate in 20S proteasomes upon inhibition of proteasome activity (Fig. 1b), whereas yeast Ump1 was 7. Instead, hump1 appeared in lighter fractions (fractions 2, 4, and 6 85 of Supplementary Fig. 1b), presumably in a free form. In addition, in human cell lines, disturbance of the assembly pathway did not cause accumulation of unprocessed β subunits as free forms (Figs. 1a and 3), which was clearly observed in yeast 7. In human cells, unassembled subunits may be subjected to rapid degradation by protein quality control system. Structural differences have also been identified in each α subunit between mammals 90 and yeast 15, though the subunit composition and arrangement of mammalian 20S proteasomes
5 are identical to those of yeast 20S proteasomes. Moreover, some proteasome regulators that bind to α-rings, such as PA28 (refs 16,17) and PI31 (refs 18, 19), are found in mammals but not in yeast. Since it is indicated that both PA28 and PI31 do not only regulate 20S proteasome activities but are also involved in immunoproteasome assembly 20, 21, the PAC 95 complex may counteract or work cooperatively with these regulators for proper α-ring formation and subsequent half-proteasome formation Zwickl, P., Kleinz, J. & Baumeister, W. Critical elements in proteasome assembly. Nat Struct Biol 1, (1994). 2. Yao, Y. et al. alpha5 subunit in Trypanosoma brucei proteasome can self-assemble to form a cylinder of four stacked heptamer rings. Biochem J 344 Pt 2, (1999). 3. Gerards, W. L. et al. The human α-type proteasomal subunit HsC8 forms a double ringlike structure, but does not assemble into proteasome-like particles with the β-type subunits HsDelta or HsBPROS26. J Biol Chem 272, (1997). 4. Emori, Y. et al. Molecular cloning and functional analysis of three subunits of yeast proteasome. Mol Cell Biol 11, (1991). 5. Heinemeyer, W., Trondle, N., Albrecht, G. & Wolf, D. H. PRE5 and PRE6, the last missing genes encoding 20S proteasome subunits from yeast? Indication for a set of 14 different subunits in the eukaryotic proteasome core. Biochemistry 33, (1994). 6. Velichutina, I., Connerly, P. L., Arendt, C. S., Li, X. & Hochstrasser, M. Plasticity in eucaryotic 20S proteasome ring assembly revealed by a subunit deletion in yeast. EMBO J 23, (2004). 7. Ramos, P. C., Hockendorff, J., Johnson, E. S., Varshavsky, A. & Dohmen, R. J. Ump1p is required for proper maturation of the 20S proteasome and becomes its substrate upon completion of the assembly. Cell 92, (1998). 8. Jayarapu, K. & Griffin, T. A. Protein-protein interactions among human 20S proteasome subunits and proteassemblin. Biochem Biophys Res Commun 314, (2004). 9. Liu, C. W., Corboy, M. J., DeMartino, G. N. & Thomas, P. J. Endoproteolytic activity of the proteasome. Science 299, (2003).
6 Tofaris, G. K., Layfield, R. & Spillantini, M. G. α-synuclein metabolism and aggregation is linked to ubiquitin-independent degradation by the proteasome. FEBS Lett 509, (2001). 11. David, D. C. et al. Proteasomal degradation of tau protein. J Neurochem 83, (2002). 12. Sheaff, R. J. et al. Proteasomal turnover of p21 Cip1 does not require p21 Cip1 ubiquitination. Mol Cell 5, (2000). 13. Touitou, R. et al. A degradation signal located in the C-terminus of p21 WAF1/CIP1 is a binding site for the C8 α-subunit of the 20S proteasome. EMBO J 20, (2001). 14. Janse, D. M., Crosas, B., Finley, D. & Church, G. M. Localization to the proteasome is sufficient for degradation. J Biol Chem 279, (2004). 15. Unno, M. et al. The structure of the mammalian 20S proteasome at 2.75 Å resolution. Structure (Camb) 10, (2002). 16. Gray, C. W., Slaughter, C. A. & DeMartino, G. N. PA28 activator protein forms regulatory caps on proteasome stacked rings. J Mol Biol 236, 7-15 (1994). 17. Rechsteiner, M., Realini, C. & Ustrell, V. The proteasome activator 11 S REG (PA28) and class I antigen presentation. Biochem J 345 Pt 1, 1-15 (2000). 18. Zaiss, D. M., Standera, S., Holzhutter, H., Kloetzel, P. & Sijts, A. J. The proteasome inhibitor PI31 competes with PA28 for binding to 20S proteasomes. FEBS Lett 457, (1999). 19. McCutchen-Maloney, S. L. et al. cdna cloning, expression, and functional characterization of PI31, a proline-rich inhibitor of the proteasome. J Biol Chem 275, (2000). 20. Preckel, T. et al. Impaired immunoproteasome assembly and immune responses in PA28 -/- mice. Science 286, (1999). 21. Zaiss, D. M., Standera, S., Kloetzel, P. M. & Sijts, A. J. PI31 is a modulator of proteasome formation and antigen processing. Proc Natl Acad Sci U S A 99, (2002).
There are approximately 30,000 proteasomes in a typical human cell Each proteasome is approximately 700 kda in size The proteasome is made up of 3
 Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
Proteasomes. When Death Comes a Knock n. Warren Gallagher Chem412, Spring 2001
 Proteasomes When Death Comes a Knock n Warren Gallagher Chem412, Spring 2001 I. Introduction Introduction The central dogma Genetic information is used to make proteins. DNA RNA Proteins Proteins are the
Proteasomes When Death Comes a Knock n Warren Gallagher Chem412, Spring 2001 I. Introduction Introduction The central dogma Genetic information is used to make proteins. DNA RNA Proteins Proteins are the
The Nobel Prize in Chemistry 2004
 The Nobel Prize in Chemistry 2004 Ubiquitous Quality Control of Life C S Karigar and K R Siddalinga Murthy The Nobel Prize in Chemistry for 2004 is shared by Aaron Ciechanover, Avram Hershko and Irwin
The Nobel Prize in Chemistry 2004 Ubiquitous Quality Control of Life C S Karigar and K R Siddalinga Murthy The Nobel Prize in Chemistry for 2004 is shared by Aaron Ciechanover, Avram Hershko and Irwin
Structure of the Proteasome
 Structure of the Proteasome Tobias Jung and Tilman Grune Institute of Nutrition, Friedrich Schiller University, Jena, Germany I. Introduction... 1 II. The 20S Proteasome... 2 A. The Proteasomal a-subunits...
Structure of the Proteasome Tobias Jung and Tilman Grune Institute of Nutrition, Friedrich Schiller University, Jena, Germany I. Introduction... 1 II. The 20S Proteasome... 2 A. The Proteasomal a-subunits...
Chapter 31. Completing the Protein Life Cycle: Folding, Processing and Degradation. Biochemistry by Reginald Garrett and Charles Grisham
 Chapter 31 Completing the Protein Life Cycle: Folding, Processing and Degradation Biochemistry by Reginald Garrett and Charles Grisham Essential Question How are newly synthesized polypeptide chains transformed
Chapter 31 Completing the Protein Life Cycle: Folding, Processing and Degradation Biochemistry by Reginald Garrett and Charles Grisham Essential Question How are newly synthesized polypeptide chains transformed
2013 John Wiley & Sons, Inc. All rights reserved. PROTEIN SORTING. Lecture 10 BIOL 266/ Biology Department Concordia University. Dr. S.
 PROTEIN SORTING Lecture 10 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University Introduction Membranes divide the cytoplasm of eukaryotic cells into distinct compartments. The endomembrane
PROTEIN SORTING Lecture 10 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University Introduction Membranes divide the cytoplasm of eukaryotic cells into distinct compartments. The endomembrane
Peptide hydrolysis uncatalyzed half-life = ~450 years HIV protease-catalyzed half-life = ~3 seconds
 Uncatalyzed half-life Peptide hydrolysis uncatalyzed half-life = ~450 years IV protease-catalyzed half-life = ~3 seconds Life Sciences 1a Lecture Slides Set 9 Fall 2006-2007 Prof. David R. Liu In the absence
Uncatalyzed half-life Peptide hydrolysis uncatalyzed half-life = ~450 years IV protease-catalyzed half-life = ~3 seconds Life Sciences 1a Lecture Slides Set 9 Fall 2006-2007 Prof. David R. Liu In the absence
UNDERSTANDING THE ROLE OF ATP HYDROLYSIS IN THE SPLICEOSOME
 UNDERSTANDING THE ROLE OF ATP HYDROLYSIS IN THE SPLICEOSOME RNA Splicing Lecture 2, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Eight essential ATPases
UNDERSTANDING THE ROLE OF ATP HYDROLYSIS IN THE SPLICEOSOME RNA Splicing Lecture 2, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Eight essential ATPases
MOLECULAR CELL BIOLOGY
 1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
Key Concept B F. How do peptides get loaded onto the proper kind of MHC molecule?
 Location of MHC class I pockets termed B and F that bind P and P9 amino acid side chains of the peptide Different MHC alleles confer different functional properties on the adaptive immune system by specifying
Location of MHC class I pockets termed B and F that bind P and P9 amino acid side chains of the peptide Different MHC alleles confer different functional properties on the adaptive immune system by specifying
B F. Location of MHC class I pockets termed B and F that bind P2 and P9 amino acid side chains of the peptide
 Different MHC alleles confer different functional properties on the adaptive immune system by specifying molecules that have different peptide binding abilities Location of MHC class I pockets termed B
Different MHC alleles confer different functional properties on the adaptive immune system by specifying molecules that have different peptide binding abilities Location of MHC class I pockets termed B
Martin Rechsteiner Department of Biochemistry University of Utah, Salt Lake, UT, USA
 1 Proteasomes Martin Rechsteiner Department of Biochemistry University of Utah, Salt Lake, UT, USA 1 Proteasomes A Quick Summary 5 2 The 20S Proteasome 5 2.1 Structure 5 2.2 Enzyme Mechanism and Proteasome
1 Proteasomes Martin Rechsteiner Department of Biochemistry University of Utah, Salt Lake, UT, USA 1 Proteasomes A Quick Summary 5 2 The 20S Proteasome 5 2.1 Structure 5 2.2 Enzyme Mechanism and Proteasome
The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein
 THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
General information. Cell mediated immunity. 455 LSA, Tuesday 11 to noon. Anytime after class.
 General information Cell mediated immunity 455 LSA, Tuesday 11 to noon Anytime after class T-cell precursors Thymus Naive T-cells (CD8 or CD4) email: lcoscoy@berkeley.edu edu Use MCB150 as subject line
General information Cell mediated immunity 455 LSA, Tuesday 11 to noon Anytime after class T-cell precursors Thymus Naive T-cells (CD8 or CD4) email: lcoscoy@berkeley.edu edu Use MCB150 as subject line
Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University
 Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
Cellular functions of protein degradation
 Protein Degradation Cellular functions of protein degradation 1. Elimination of misfolded and damaged proteins: Environmental toxins, translation errors and genetic mutations can damage proteins. Misfolded
Protein Degradation Cellular functions of protein degradation 1. Elimination of misfolded and damaged proteins: Environmental toxins, translation errors and genetic mutations can damage proteins. Misfolded
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10643 Supplementary Table 1. Identification of hecw-1 coding polymorphisms at amino acid positions 322 and 325 in 162 strains of C. elegans. WWW.NATURE.COM/NATURE 1 Supplementary Figure
doi:10.1038/nature10643 Supplementary Table 1. Identification of hecw-1 coding polymorphisms at amino acid positions 322 and 325 in 162 strains of C. elegans. WWW.NATURE.COM/NATURE 1 Supplementary Figure
Moving Proteins into Membranes and Organelles
 13 Moving Proteins into Membranes and Organelles Review the Concepts 1. In eukaryotes, protein translocation across the endoplasmic reticulum (ER) membrane is most commonly cotranslational; it can also
13 Moving Proteins into Membranes and Organelles Review the Concepts 1. In eukaryotes, protein translocation across the endoplasmic reticulum (ER) membrane is most commonly cotranslational; it can also
CMB621: Cytoskeleton. Also known as How the cell plays with LEGOs to ensure order, not chaos, is temporally and spatially achieved
 CMB621: Cytoskeleton Also known as How the cell plays with LEGOs to ensure order, not chaos, is temporally and spatially achieved Lecture(s) Overview Lecture 1: What is the cytoskeleton? Membrane interaction
CMB621: Cytoskeleton Also known as How the cell plays with LEGOs to ensure order, not chaos, is temporally and spatially achieved Lecture(s) Overview Lecture 1: What is the cytoskeleton? Membrane interaction
Glycoprotein Maturation and Quality Control in the Endoplasmic Reticulum Dr. Daniel Hebert
 Glycoprotein Maturation and Quality Control in the Endoplasmic Reticulum Department of Biochemistry and Molecular Biology University of Massachusetts, USA 1 Intracellular protein trafficking Plasma membrane
Glycoprotein Maturation and Quality Control in the Endoplasmic Reticulum Department of Biochemistry and Molecular Biology University of Massachusetts, USA 1 Intracellular protein trafficking Plasma membrane
Molecular Graphics Perspective of Protein Structure and Function
 Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Electron Microscopy Imaging Reveals Unique Higher Order Structures of Adalimumab-TNFα and Infliximab-TNFα Complexes
 Electron Microscopy Imaging Reveals Unique Higher Order Structures of Adalimumab-TNFα and Infliximab-TNFα Complexes Siew Leong Chan Protein Analytics, AbbVie Bioresearch Center Worcester MA Acknowledgement
Electron Microscopy Imaging Reveals Unique Higher Order Structures of Adalimumab-TNFα and Infliximab-TNFα Complexes Siew Leong Chan Protein Analytics, AbbVie Bioresearch Center Worcester MA Acknowledgement
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics
 1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics to gain mechanistic insight! 4. Return to step 2, as
1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics to gain mechanistic insight! 4. Return to step 2, as
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Antigen Presentation to T lymphocytes
 Antigen Presentation to T lymphocytes Immunology 441 Lectures 6 & 7 Chapter 6 October 10 & 12, 2016 Jessica Hamerman jhamerman@benaroyaresearch.org Office hours by arrangement Antibodies and T cell receptors
Antigen Presentation to T lymphocytes Immunology 441 Lectures 6 & 7 Chapter 6 October 10 & 12, 2016 Jessica Hamerman jhamerman@benaroyaresearch.org Office hours by arrangement Antibodies and T cell receptors
The ubiquitin-proteasome system. in bone biology. University of Nottingham. ECTS PhD Training July 5 th Dr Rob Layfield
 The ubiquitin-proteasome system in bone biology Dr Rob Layfield University of Nottingham ECTS PhD Training July 5 th 2009 Images reproduced from: http://www.bostonbiochem.com/overview.php?prod=ubchains
The ubiquitin-proteasome system in bone biology Dr Rob Layfield University of Nottingham ECTS PhD Training July 5 th 2009 Images reproduced from: http://www.bostonbiochem.com/overview.php?prod=ubchains
Chapter 10. Regulatory Strategy
 Chapter 10 Regulatory Strategy Regulation of enzymatic activity: 1. Allosteric Control. Allosteric proteins have a regulatory site(s) and multiple functional sites Activity of proteins is regulated by
Chapter 10 Regulatory Strategy Regulation of enzymatic activity: 1. Allosteric Control. Allosteric proteins have a regulatory site(s) and multiple functional sites Activity of proteins is regulated by
Intrinsically Disordered Proteins. Alex Cioffi June 22 nd 2013
 Intrinsically Disordered Proteins Alex Cioffi June 22 nd 2013 Implications in Human Disease Adenovirus Early Region 1A (E1A) and Cancer α-synuclein and Parkinson s Disease Amyloid β and Alzheimer s Disease
Intrinsically Disordered Proteins Alex Cioffi June 22 nd 2013 Implications in Human Disease Adenovirus Early Region 1A (E1A) and Cancer α-synuclein and Parkinson s Disease Amyloid β and Alzheimer s Disease
Chapter 6. Antigen Presentation to T lymphocytes
 Chapter 6 Antigen Presentation to T lymphocytes Generation of T-cell Receptor Ligands T cells only recognize Ags displayed on cell surfaces These Ags may be derived from pathogens that replicate within
Chapter 6 Antigen Presentation to T lymphocytes Generation of T-cell Receptor Ligands T cells only recognize Ags displayed on cell surfaces These Ags may be derived from pathogens that replicate within
Cell Quality Control. Peter Takizawa Department of Cell Biology
 Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Andrea s SI Session PCB Practice Test Test 3
 Practice Test Test 3 READ BEFORE STARTING PRACTICE TEST: Remember to please use this practice test as a tool to measure your knowledge, and DO NOT use it as your only tool to study for the test, since
Practice Test Test 3 READ BEFORE STARTING PRACTICE TEST: Remember to please use this practice test as a tool to measure your knowledge, and DO NOT use it as your only tool to study for the test, since
BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001
 BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001 SS# Name This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. Good luck! 1. (2) The
BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001 SS# Name This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. Good luck! 1. (2) The
Study of different types of ubiquitination
 Study of different types of ubiquitination Rudi Beyaert (rudi.beyaert@irc.vib-ugent.be) VIB UGent Center for Inflammation Research Ghent, Belgium VIB Training Novel Proteomics Tools: Identifying PTMs October
Study of different types of ubiquitination Rudi Beyaert (rudi.beyaert@irc.vib-ugent.be) VIB UGent Center for Inflammation Research Ghent, Belgium VIB Training Novel Proteomics Tools: Identifying PTMs October
Cell Biology. A discipline of biology: 1. Cell structure 2. Cellular processes 3. Cell division
 The Cell Cell Biology 1 A discipline of biology: 1. Cell structure 2. Cellular processes 3. Cell division Tight connection with 1. Molecular biology 2. Biochemistry Cell theory 2 1838, 1839 Theodor Schwann
The Cell Cell Biology 1 A discipline of biology: 1. Cell structure 2. Cellular processes 3. Cell division Tight connection with 1. Molecular biology 2. Biochemistry Cell theory 2 1838, 1839 Theodor Schwann
Cell Biology from an Immune Perspective
 Harvard-MIT Division of Health Sciences and Technology HST.176: Cellular and Molecular Immunology Course Director: Dr. Shiv Pillai Cell Biology from an Immune Perspective In this lecture we will very briefly
Harvard-MIT Division of Health Sciences and Technology HST.176: Cellular and Molecular Immunology Course Director: Dr. Shiv Pillai Cell Biology from an Immune Perspective In this lecture we will very briefly
RNA Processing in Eukaryotes *
 OpenStax-CNX module: m44532 1 RNA Processing in Eukaryotes * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you
OpenStax-CNX module: m44532 1 RNA Processing in Eukaryotes * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you
20S Proteasome Activity Assay Kit
 20S Proteasome Activity Assay Kit For 100 Assays Cat. No. APT280 FOR RESEARCH USE ONLY NOT FOR USE IN DIAGNOSTIC PROCEDURES USA & Canada Phone: +1(800) 437-7500 Fax: +1 (951) 676-9209 Europe +44 (0) 23
20S Proteasome Activity Assay Kit For 100 Assays Cat. No. APT280 FOR RESEARCH USE ONLY NOT FOR USE IN DIAGNOSTIC PROCEDURES USA & Canada Phone: +1(800) 437-7500 Fax: +1 (951) 676-9209 Europe +44 (0) 23
Materials and Methods , The two-hybrid principle.
 The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
Cbl ubiquitin ligase: Lord of the RINGs
 Cbl ubiquitin ligase: Lord of the RINGs Not just quite interesting - really interesting! A cell must be able to degrade proteins when their activity is no longer required. Many eukaryotic proteins are
Cbl ubiquitin ligase: Lord of the RINGs Not just quite interesting - really interesting! A cell must be able to degrade proteins when their activity is no longer required. Many eukaryotic proteins are
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Why chaperone vectors?
 Why chaperone vectors? A protein folding initiative An open discussion with structural biologists Protein Structure Initiative: Pilot Phase Whether the pilot phase achieved its goal depends on how we measure
Why chaperone vectors? A protein folding initiative An open discussion with structural biologists Protein Structure Initiative: Pilot Phase Whether the pilot phase achieved its goal depends on how we measure
Regulators of Cell Cycle Progression
 Regulators of Cell Cycle Progression Studies of Cdk s and cyclins in genetically modified mice reveal a high level of plasticity, allowing different cyclins and Cdk s to compensate for the loss of one
Regulators of Cell Cycle Progression Studies of Cdk s and cyclins in genetically modified mice reveal a high level of plasticity, allowing different cyclins and Cdk s to compensate for the loss of one
brought to you by and REFERENCES
 brought to you by www.thebacteriophages.org and www.phage.org REFERENCES 1. Butcher, S. J., T. Dokland, P. M. Ojala, D. H. Bamford, and S. D. Fuller. 1997. Intermediates in the assembly pathway of the
brought to you by www.thebacteriophages.org and www.phage.org REFERENCES 1. Butcher, S. J., T. Dokland, P. M. Ojala, D. H. Bamford, and S. D. Fuller. 1997. Intermediates in the assembly pathway of the
Antigen presenting cells
 Antigen recognition by T and B cells - T and B cells exhibit fundamental differences in antigen recognition - B cells recognize antigen free in solution (native antigen). - T cells recognize antigen after
Antigen recognition by T and B cells - T and B cells exhibit fundamental differences in antigen recognition - B cells recognize antigen free in solution (native antigen). - T cells recognize antigen after
Cell Cycle, Mitosis, and Microtubules. LS1A Final Exam Review Friday 1/12/07. Processes occurring during cell cycle
 Cell Cycle, Mitosis, and Microtubules LS1A Final Exam Review Friday 1/12/07 Processes occurring during cell cycle Replicate chromosomes Segregate chromosomes Cell divides Cell grows Cell Growth 1 The standard
Cell Cycle, Mitosis, and Microtubules LS1A Final Exam Review Friday 1/12/07 Processes occurring during cell cycle Replicate chromosomes Segregate chromosomes Cell divides Cell grows Cell Growth 1 The standard
The Adaptive Immune Response. T-cells
 The Adaptive Immune Response T-cells T Lymphocytes T lymphocytes develop from precursors in the thymus. Mature T cells are found in the blood, where they constitute 60% to 70% of lymphocytes, and in T-cell
The Adaptive Immune Response T-cells T Lymphocytes T lymphocytes develop from precursors in the thymus. Mature T cells are found in the blood, where they constitute 60% to 70% of lymphocytes, and in T-cell
Significance of the MHC
 CHAPTER 8 Major Histocompatibility Complex (MHC) What is is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) Significance of the MHC role in immune response role in organ
CHAPTER 8 Major Histocompatibility Complex (MHC) What is is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) Significance of the MHC role in immune response role in organ
DOWNLOAD OR READ : EFFECTS OF PHOSPHORYLATION ON THE RECYCLING OF EUKARYOTIC INITIATION FACTOR 2 EIF 2 PDF EBOOK EPUB MOBI
 DOWNLOAD OR READ : EFFECTS OF PHOSPHORYLATION ON THE RECYCLING OF EUKARYOTIC INITIATION FACTOR 2 EIF 2 PDF EBOOK EPUB MOBI Page 1 Page 2 effects of phosphorylation on the recycling of eukaryotic initiation
DOWNLOAD OR READ : EFFECTS OF PHOSPHORYLATION ON THE RECYCLING OF EUKARYOTIC INITIATION FACTOR 2 EIF 2 PDF EBOOK EPUB MOBI Page 1 Page 2 effects of phosphorylation on the recycling of eukaryotic initiation
Supplementary data Multiple hit hypotheses for dopamine neuron loss in Parkinson s disease
 Supplementary data Multiple hit hypotheses for dopamine neuron loss in Parkinson s disease David Sulzer Departments of Neurology, Psychiatry and Pharmacology, Black 309, 650 West, 168th Street, Columbia
Supplementary data Multiple hit hypotheses for dopamine neuron loss in Parkinson s disease David Sulzer Departments of Neurology, Psychiatry and Pharmacology, Black 309, 650 West, 168th Street, Columbia
Biochemistry 2 Recita0on Amino Acid Metabolism
 Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10495 WWW.NATURE.COM/NATURE 1 2 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 3 4 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 5 6 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 7 8 WWW.NATURE.COM/NATURE
doi:10.1038/nature10495 WWW.NATURE.COM/NATURE 1 2 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 3 4 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 5 6 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 7 8 WWW.NATURE.COM/NATURE
Intracellular vesicular traffic. B. Balen
 Intracellular vesicular traffic B. Balen Three types of transport in eukaryotic cells Figure 12-6 Molecular Biology of the Cell ( Garland Science 2008) Endoplasmic reticulum in all eucaryotic cells Endoplasmic
Intracellular vesicular traffic B. Balen Three types of transport in eukaryotic cells Figure 12-6 Molecular Biology of the Cell ( Garland Science 2008) Endoplasmic reticulum in all eucaryotic cells Endoplasmic
Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras
 Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras Tarik Issad, Ralf Jockers and Stefano Marullo 1 Because they play a pivotal role
Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras Tarik Issad, Ralf Jockers and Stefano Marullo 1 Because they play a pivotal role
a) The statement is true for X = 400, but false for X = 300; b) The statement is true for X = 300, but false for X = 200;
 1. Consider the following statement. To produce one molecule of each possible kind of polypeptide chain, X amino acids in length, would require more atoms than exist in the universe. Given the size of
1. Consider the following statement. To produce one molecule of each possible kind of polypeptide chain, X amino acids in length, would require more atoms than exist in the universe. Given the size of
RECAP (1)! In eukaryotes, large primary transcripts are processed to smaller, mature mrnas.! What was first evidence for this precursorproduct
 RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
Novel Targets of disease modifying therapy for Parkinson disease. David G. Standaert, MD, PhD John N. Whitaker Professor and Chair of Neurology
 Novel Targets of disease modifying therapy for Parkinson disease David G. Standaert, MD, PhD John N. Whitaker Professor and Chair of Neurology Disclosures Dr. Standaert has served as a paid consultant
Novel Targets of disease modifying therapy for Parkinson disease David G. Standaert, MD, PhD John N. Whitaker Professor and Chair of Neurology Disclosures Dr. Standaert has served as a paid consultant
III. Metabolism The Citric Acid Cycle
 Department of Chemistry and Biochemistry University of Lethbridge III. Metabolism The Citric Acid Cycle Slide 1 The Eight Steps of the Citric Acid Cycle Enzymes: 4 dehydrogenases (2 decarboxylation) 3
Department of Chemistry and Biochemistry University of Lethbridge III. Metabolism The Citric Acid Cycle Slide 1 The Eight Steps of the Citric Acid Cycle Enzymes: 4 dehydrogenases (2 decarboxylation) 3
PRC2 crystal clear. Matthieu Schapira
 PRC2 crystal clear Matthieu Schapira Epigenetic mechanisms control the combination of genes that are switched on and off in any given cell. In turn, this combination, called the transcriptional program,
PRC2 crystal clear Matthieu Schapira Epigenetic mechanisms control the combination of genes that are switched on and off in any given cell. In turn, this combination, called the transcriptional program,
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus
 Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
supplementary information
 DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
Overview of virus life cycle
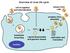 Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Human Genome: Mapping, Sequencing Techniques, Diseases
 Human Genome: Mapping, Sequencing Techniques, Diseases Lecture 4 BINF 7580 Fall 2005 1 Let us review what we talked about at the previous lecture. Please,... 2 The central dogma states that the transfer
Human Genome: Mapping, Sequencing Techniques, Diseases Lecture 4 BINF 7580 Fall 2005 1 Let us review what we talked about at the previous lecture. Please,... 2 The central dogma states that the transfer
Structure of proteins
 Structure of proteins Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Structure of proteins The 20 a.a commonly found
Structure of proteins Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Structure of proteins The 20 a.a commonly found
Effects of Second Messengers
 Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Cahn - Ingold - Prelog system. Proteins: Evolution, and Analysis Lecture 7 9/15/2009. The Fischer Convention (1) G (2) (3)
 Chapter 4 (1) G Proteins: Evolution, and Analysis Lecture 7 9/15/2009 A V L I M P F W Chapter 4 (2) S (3) T N Q Y C K R H D E The Fischer Convention Absolute configuration about an asymmetric carbon related
Chapter 4 (1) G Proteins: Evolution, and Analysis Lecture 7 9/15/2009 A V L I M P F W Chapter 4 (2) S (3) T N Q Y C K R H D E The Fischer Convention Absolute configuration about an asymmetric carbon related
Protein Trafficking in the Secretory and Endocytic Pathways
 Protein Trafficking in the Secretory and Endocytic Pathways The compartmentalization of eukaryotic cells has considerable functional advantages for the cell, but requires elaborate mechanisms to ensure
Protein Trafficking in the Secretory and Endocytic Pathways The compartmentalization of eukaryotic cells has considerable functional advantages for the cell, but requires elaborate mechanisms to ensure
Practice Exam 2 MCBII
 1. Which feature is true for signal sequences and for stop transfer transmembrane domains (4 pts)? A. They are both 20 hydrophobic amino acids long. B. They are both found at the N-terminus of the protein.
1. Which feature is true for signal sequences and for stop transfer transmembrane domains (4 pts)? A. They are both 20 hydrophobic amino acids long. B. They are both found at the N-terminus of the protein.
Homework Hanson section MCB Course, Fall 2014
 Homework Hanson section MCB Course, Fall 2014 (1) Antitrypsin, which inhibits certain proteases, is normally secreted into the bloodstream by liver cells. Antitrypsin is absent from the bloodstream of
Homework Hanson section MCB Course, Fall 2014 (1) Antitrypsin, which inhibits certain proteases, is normally secreted into the bloodstream by liver cells. Antitrypsin is absent from the bloodstream of
RECAP FROM MONDAY AND TUESDAY LECTURES
 RECAP FROM MONDAY AND TUESDAY LECTURES Nucleo-cytoplasmic transport factors: interact with the target macromolecule (signal recognition) interact with the NPC (nucleoporin Phe - Gly repeats) NPC cytoplasmic
RECAP FROM MONDAY AND TUESDAY LECTURES Nucleo-cytoplasmic transport factors: interact with the target macromolecule (signal recognition) interact with the NPC (nucleoporin Phe - Gly repeats) NPC cytoplasmic
Coenzyme Q 10. R. JÜRGEN DOHMEN Institute for Genetics, University of Cologne, Germany
 Coenzyme Q 10 Ubiquitin is a small protein of 76 residues that is conjugated posttranslationally to other proteins. It is ubiquitously present in all eukaryotes, hence its name, and is one of the most
Coenzyme Q 10 Ubiquitin is a small protein of 76 residues that is conjugated posttranslationally to other proteins. It is ubiquitously present in all eukaryotes, hence its name, and is one of the most
ATPases; P-type, V-type & F-type, how does H + power V & F-type 2015
 ATPases; P-type, V-type & F-type, how does H + power V & F-type 2015 1 /2 O 2 +NADH 2 H 2 O + NAD + ΔE O =0.82-(-0.32)= 1.14V ΔG O = -218KJ/mol The oxidation of 1mol of NADH is associated with the release
ATPases; P-type, V-type & F-type, how does H + power V & F-type 2015 1 /2 O 2 +NADH 2 H 2 O + NAD + ΔE O =0.82-(-0.32)= 1.14V ΔG O = -218KJ/mol The oxidation of 1mol of NADH is associated with the release
H C. C α. Proteins perform a vast array of biological function including: Side chain
 Topics The topics: basic concepts of molecular biology elements on Python overview of the field biological databases and database searching sequence alignments phylogenetic trees microarray data analysis
Topics The topics: basic concepts of molecular biology elements on Python overview of the field biological databases and database searching sequence alignments phylogenetic trees microarray data analysis
HLA and antigen presentation. Department of Immunology Charles University, 2nd Medical School University Hospital Motol
 HLA and antigen presentation Department of Immunology Charles University, 2nd Medical School University Hospital Motol MHC in adaptive immunity Characteristics Specificity Innate For structures shared
HLA and antigen presentation Department of Immunology Charles University, 2nd Medical School University Hospital Motol MHC in adaptive immunity Characteristics Specificity Innate For structures shared
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Amino Acids and Proteins Hamad Ali Yaseen, PhD MLS Department, FAHS, HSC, KU Biochemistry 210 Chapter 22
 Amino Acids and Proteins Hamad Ali Yaseen, PhD MLS Department, FAHS, HSC, KU Hamad.ali@hsc.edu.kw Biochemistry 210 Chapter 22 Importance of Proteins Main catalysts in biochemistry: enzymes (involved in
Amino Acids and Proteins Hamad Ali Yaseen, PhD MLS Department, FAHS, HSC, KU Hamad.ali@hsc.edu.kw Biochemistry 210 Chapter 22 Importance of Proteins Main catalysts in biochemistry: enzymes (involved in
Function of aquaporin in the brain and brain edema
 Nagoya Med. J., 199 Function of aquaporin in the brain and brain edema KAZUYA SOBUE Department of Anesthesiology and Medical crisis Management, Nagoya City University Graduate School of Medical Sciences
Nagoya Med. J., 199 Function of aquaporin in the brain and brain edema KAZUYA SOBUE Department of Anesthesiology and Medical crisis Management, Nagoya City University Graduate School of Medical Sciences
This student paper was written as an assignment in the graduate course
 77:222 Spring 2003 Free Radicals in Biology and Medicine Page 0 This student paper was written as an assignment in the graduate course Free Radicals in Biology and Medicine (77:222, Spring 2003) offered
77:222 Spring 2003 Free Radicals in Biology and Medicine Page 0 This student paper was written as an assignment in the graduate course Free Radicals in Biology and Medicine (77:222, Spring 2003) offered
REGULATION OF ENZYME ACTIVITY. Medical Biochemistry, Lecture 25
 REGULATION OF ENZYME ACTIVITY Medical Biochemistry, Lecture 25 Lecture 25, Outline General properties of enzyme regulation Regulation of enzyme concentrations Allosteric enzymes and feedback inhibition
REGULATION OF ENZYME ACTIVITY Medical Biochemistry, Lecture 25 Lecture 25, Outline General properties of enzyme regulation Regulation of enzyme concentrations Allosteric enzymes and feedback inhibition
Signal Transduction: Information Metabolism. Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire
 Signal Transduction: Information Metabolism Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire Introduction Information Metabolism How cells receive, process and respond
Signal Transduction: Information Metabolism Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire Introduction Information Metabolism How cells receive, process and respond
Transcript-indexed ATAC-seq for immune profiling
 Transcript-indexed ATAC-seq for immune profiling Technical Journal Club 22 nd of May 2018 Christina Müller Nature Methods, Vol.10 No.12, 2013 Nature Biotechnology, Vol.32 No.7, 2014 Nature Medicine, Vol.24,
Transcript-indexed ATAC-seq for immune profiling Technical Journal Club 22 nd of May 2018 Christina Müller Nature Methods, Vol.10 No.12, 2013 Nature Biotechnology, Vol.32 No.7, 2014 Nature Medicine, Vol.24,
8 Suppression Analysis
 Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 8 Suppression Analysis OVERVIEW Suppression
Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 8 Suppression Analysis OVERVIEW Suppression
T cell maturation. T-cell Maturation. What allows T cell maturation?
 T-cell Maturation What allows T cell maturation? Direct contact with thymic epithelial cells Influence of thymic hormones Growth factors (cytokines, CSF) T cell maturation T cell progenitor DN DP SP 2ry
T-cell Maturation What allows T cell maturation? Direct contact with thymic epithelial cells Influence of thymic hormones Growth factors (cytokines, CSF) T cell maturation T cell progenitor DN DP SP 2ry
Zn(II) as a minimal chaperone mimic to retard Aβ peptide fibril formation
 Zn(II) as a minimal chaperone mimic to retard Aβ peptide fibril formation Astrid Gräslund Department of Biochemistry and Biophysics Stockholm University Lorentz workshop, Leiden, April 17, 215 Potential
Zn(II) as a minimal chaperone mimic to retard Aβ peptide fibril formation Astrid Gräslund Department of Biochemistry and Biophysics Stockholm University Lorentz workshop, Leiden, April 17, 215 Potential
Introduction to the Proteasome and its Inhibitors
 Chapter 2 / Biochemistry and Cell Biology of Proteasomes 17 2 Introduction to the Proteasome and its Inhibitors Biochemistry and Cell Biology Alfred L. Goldberg CONTENTS INTRODUCTION THE UBIQUITIN PROTEASOME
Chapter 2 / Biochemistry and Cell Biology of Proteasomes 17 2 Introduction to the Proteasome and its Inhibitors Biochemistry and Cell Biology Alfred L. Goldberg CONTENTS INTRODUCTION THE UBIQUITIN PROTEASOME
Protein sorting (endoplasmic reticulum) Dr. Diala Abu-Hsasan School of Medicine
 Protein sorting (endoplasmic reticulum) Dr. Diala Abu-Hsasan School of Medicine dr.abuhassand@gmail.com An overview of cellular components Endoplasmic reticulum (ER) It is a network of membrane-enclosed
Protein sorting (endoplasmic reticulum) Dr. Diala Abu-Hsasan School of Medicine dr.abuhassand@gmail.com An overview of cellular components Endoplasmic reticulum (ER) It is a network of membrane-enclosed
Zool 3200: Cell Biology Exam 4 Part II 2/3/15
 Name:Key Trask Zool 3200: Cell Biology Exam 4 Part II 2/3/15 Answer each of the following questions in the space provided, explaining your answers when asked to do so; circle the correct answer or answers
Name:Key Trask Zool 3200: Cell Biology Exam 4 Part II 2/3/15 Answer each of the following questions in the space provided, explaining your answers when asked to do so; circle the correct answer or answers
High AU content: a signature of upregulated mirna in cardiac diseases
 https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
Muscular Dystrophy. Biol 405 Molecular Medicine
 Muscular Dystrophy Biol 405 Molecular Medicine Duchenne muscular dystrophy Duchenne muscular dystrophy is a neuromuscular disease that occurs in ~ 1/3,500 male births. The disease causes developmental
Muscular Dystrophy Biol 405 Molecular Medicine Duchenne muscular dystrophy Duchenne muscular dystrophy is a neuromuscular disease that occurs in ~ 1/3,500 male births. The disease causes developmental
Antigen processing and presentation. Monika Raulf
 Antigen processing and presentation Monika Raulf Lecture 25.04.2018 What is Antigen presentation? AP is the display of peptide antigens (created via antigen processing) on the cell surface together with
Antigen processing and presentation Monika Raulf Lecture 25.04.2018 What is Antigen presentation? AP is the display of peptide antigens (created via antigen processing) on the cell surface together with
Proteasome activity and its relationship with protein phosphorylation during capacitation and acrosome reaction in human spermatozoa
 Proteasome activity and its relationship with protein phosphorylation Proteasome activity and its relationship with protein phosphorylation during capacitation and acrosome reaction in human spermatozoa
Proteasome activity and its relationship with protein phosphorylation Proteasome activity and its relationship with protein phosphorylation during capacitation and acrosome reaction in human spermatozoa
The major histocompatibility complex (MHC) is a group of genes that governs tumor and tissue transplantation between individuals of a species.
 Immunology Dr. John J. Haddad Chapter 7 Major Histocompatibility Complex The major histocompatibility complex (MHC) is a group of genes that governs tumor and tissue transplantation between individuals
Immunology Dr. John J. Haddad Chapter 7 Major Histocompatibility Complex The major histocompatibility complex (MHC) is a group of genes that governs tumor and tissue transplantation between individuals
Loss of protein association causes cardiolipin degradation in Barth syndrome
 SUPPLEMENTARY INFORMATION Loss of protein association causes cardiolipin degradation in Barth syndrome Yang Xu 1, Colin K.L. Phoon 2, Bob Berno 5, Kenneth D Souza 6, Esthelle Hoedt 4, Guoan Zhang 4, Thomas
SUPPLEMENTARY INFORMATION Loss of protein association causes cardiolipin degradation in Barth syndrome Yang Xu 1, Colin K.L. Phoon 2, Bob Berno 5, Kenneth D Souza 6, Esthelle Hoedt 4, Guoan Zhang 4, Thomas
Sheet #5 Dr. Mamoun Ahram 8/7/2014
 P a g e 1 Protein Structure Quick revision - Levels of protein structure: primary, secondary, tertiary & quaternary. - Primary structure is the sequence of amino acids residues. It determines the other
P a g e 1 Protein Structure Quick revision - Levels of protein structure: primary, secondary, tertiary & quaternary. - Primary structure is the sequence of amino acids residues. It determines the other
Insulin mrna to Protein Kit
 Insulin mrna to Protein Kit A 3DMD Paper BioInformatics and Mini-Toober Folding Activity Student Handout www.3dmoleculardesigns.com Insulin mrna to Protein Kit Contents Becoming Familiar with the Data...
Insulin mrna to Protein Kit A 3DMD Paper BioInformatics and Mini-Toober Folding Activity Student Handout www.3dmoleculardesigns.com Insulin mrna to Protein Kit Contents Becoming Familiar with the Data...
Membrane Biochemistry. Lectures by. John F. Allen. School of Biological and Chemical Sciences, Queen Mary, University of London. jfallen.
 Membrane Biochemistry Lectures by John F. Allen School of Biological and Chemical Sciences, Queen Mary, University of London jfallen.org/lectures 1 Membrane Biochemistry Bioenergetics jfallen.org/lectures
Membrane Biochemistry Lectures by John F. Allen School of Biological and Chemical Sciences, Queen Mary, University of London jfallen.org/lectures 1 Membrane Biochemistry Bioenergetics jfallen.org/lectures
Structure-Function Relationship
 1 P a g e Structure-Function Relationship You have studied the amino acids and their characteristics, but in this part we will study the relation between the structure and the function of protein. Proteins
1 P a g e Structure-Function Relationship You have studied the amino acids and their characteristics, but in this part we will study the relation between the structure and the function of protein. Proteins
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature19102 Supplementary Discussion Benzothiazepine Binding in Ca V Ab Diltiazem and other benzothiazepines inhibit Ca V 1.2 channels in a frequency-dependent manner consistent with pore block
doi:10.1038/nature19102 Supplementary Discussion Benzothiazepine Binding in Ca V Ab Diltiazem and other benzothiazepines inhibit Ca V 1.2 channels in a frequency-dependent manner consistent with pore block
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature22314 Supplementary Discussion In mammals, BCAAs are essential amino acids that must be supplied from food. These dietary BCAAs are subsequently used for protein synthesis or are catabolized
doi:10.1038/nature22314 Supplementary Discussion In mammals, BCAAs are essential amino acids that must be supplied from food. These dietary BCAAs are subsequently used for protein synthesis or are catabolized
Lecture 10 - Protein Turnover and Amino Acid Catabolism
 Lecture 10 - Protein Turnover and Amino Acid Catabolism Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire 1 Introduction 2 Proteins are degraded into amino acids. Protein
Lecture 10 - Protein Turnover and Amino Acid Catabolism Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire 1 Introduction 2 Proteins are degraded into amino acids. Protein
