1 Supplementary Figures
|
|
|
- Loraine Delphia Dalton
- 5 years ago
- Views:
Transcription
1 Supplementary Figures D. simulans (dsim) D. sechellia (dsec) D. melanogaster (dmel) S. cerevisiae (scer) S. paradoxus (spar) S. mikatae (smik) S. bayanus (sbay) S. castellii (scas) C. glabrata (cgla) K. waltii (kwal) A. gossypii (agos) K. lactis (klac) D. yakuba (dyak) D. erecta (dere) D. ananassae (dana) D. pseudoobscura (dpse) D. persimils (dper) D. willistoni (dwil) D. mojavensis (dmoj) D. virilis (dvir) D. grimshawi (dgri) ~ million years ~ million years. Figure S: Species names, abbreviations, and phylogeny
2 Name T T T T T T T T T9 T T T T T T Topology ((((((dere,dyak),dmel),dana),dpse),dwil),((dmoj,dvir),dgri)) ((((((dere,dmel),dyak),dana),dpse),dwil),((dmoj,dvir),dgri)) ((((((dere,dyak),dmel),dana),dpse),dwil),((dgri,dvir),dmoj)) ((((((dmel,dyak),dere),dana),dpse),dwil),((dmoj,dvir),dgri)) ((((((dere,dmel),dyak),dana),dpse),dwil),((dgri,dvir),dmoj)) ((((((dere,dyak),dmel),dana),dpse),dwil),((dgri,dmoj),dvir)) ((((((dmel,dyak),dere),dana),dpse),dwil),((dgri,dvir),dmoj)) ((((((dere,dyak),dmel),dana),dwil),dpse),((dmoj,dvir),dgri)) (((((dere,dyak),dmel),dana),(dpse,dwil)),((dmoj,dvir),dgri)) ((((((dere,dmel),dyak),dana),dpse),dwil),((dgri,dmoj),dvir)) (((((dere,dyak),dmel),dana),(dpse,dwil)),((dgri,dvir),dmoj)) ((((((dere,dmel),dyak),dana),dwil),dpse),((dmoj,dvir),dgri)) ((((((dmel,dyak),dere),dana),dpse),dwil),((dgri,dmoj),dvir)) ((((((dere,dyak),dmel),dana),dwil),dpse),((dgri,dvir),dmoj)) (((((dere,dmel),dyak),dana),(dpse,dwil)),((dmoj,dvir),dgri)) Figure Sa: Naming scheme for ortholog gene-trees for 9 fly species. Ordered by ML frequency. SPIDIR SPIDIR (D=.) SPIDIR (D=.) PHYML BIONJ MrBayes DNAPARS T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.9%) T (.%) T (9.%) T (.%) T (.%) T (.9%) T (.%) T (.%) T (.%) T (.9%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.9%) T (.%) T (.%) T (.9%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.9%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.9%) T (.%) T9 (.%) T (.%) T (.%) T (.%) T (.%) T (.9%) T (.%) T (.%) T9 (.9%) T9 (.%) T (.%) T (.%) T9 (.%) T (.%) T (.%) T (.%) T (.%) T9 (.%) T9 (.%) T (.%) T9 (.%) T (.9%) T (.%) T (.%) T (.%) T (.%) T (.9%) T (.%) Figure Sb: Most frequently constructed ortholog gene-trees for 9 fly species. Different methods reconstruct the same topologies at similar frequencies.
3 Name T T T T T T T T T9 T T T T T T Topology (((((((dsec,dsim),dmel),(dere,dyak)),dana),(dper,dpse)),dwil),((dmoj,dvir),dgri)) ((((((((dsec,dsim),dmel),dere),dyak),dana),(dper,dpse)),dwil),((dmoj,dvir),dgri)) (((((((dsec,dsim),dmel),(dere,dyak)),dana),(dper,dpse)),dwil),((dgri,dvir),dmoj)) ((((((((dsec,dsim),dmel),dyak),dere),dana),(dper,dpse)),dwil),((dmoj,dvir),dgri)) ((((((((dsec,dsim),dmel),dere),dyak),dana),(dper,dpse)),dwil),((dgri,dvir),dmoj)) (((((((dsec,dsim),dmel),(dere,dyak)),dana),(dper,dpse)),dwil),((dgri,dmoj),dvir)) ((((((((dsec,dsim),dmel),dyak),dere),dana),(dper,dpse)),dwil),((dgri,dvir),dmoj)) (((((((dsec,dsim),dmel),(dere,dyak)),dana),dwil),(dper,dpse)),((dmoj,dvir),dgri)) ((((((dsec,dsim),dmel),(dere,dyak)),dana),((dper,dpse),dwil)),((dmoj,dvir),dgri)) ((((((((dsec,dsim),dmel),dere),dyak),dana),(dper,dpse)),dwil),((dgri,dmoj),dvir)) (((((((dmel,dsim),dsec),(dere,dyak)),dana),(dper,dpse)),dwil),((dmoj,dvir),dgri)) (((((((dere,dyak),(dsec,dsim)),dmel),dana),(dper,dpse)),dwil),((dmoj,dvir),dgri)) (((((((dmel,dsec),dsim),(dere,dyak)),dana),(dper,dpse)),dwil),((dmoj,dvir),dgri)) (((((((dsec,dsim),dmel),dere),dyak),dana),((dper,dpse),dwil)),((dmoj,dvir),dgri)) ((((((((dsec,dsim),dmel),dere),dyak),dana),dwil),(dper,dpse)),((dmoj,dvir),dgri)) Figure Sa: Naming scheme for ortholog gene-trees for fly species. Ordered by ML frequency. SPIDIR PHYML BIONJ MrBayes DNAPARS T (.%) T (.9%) T (.9%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (9.%) T (.%) T (.%) T (.%) T (.9%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.9%) T (.9%) T (.%) T (.%) T (.%) T (.9%) T (.%) T (.%) T (.%) T (.9%) T (.%) T (.%) T (.%) T (.%) T (.%) T9 (.%) T (.%) T9 (.%) T9 (.%) T9 (.%) T (.%) T9 (.%) T (.%) T (.%) T (.%) T (.%) T (.%) Figure Sb: Most frequently constructed ortholog gene-trees for fly species. Different methods reconstruct the same topologies at similar frequencies. SPIDIR PHYML BIONJ MrBayes DNAPARS T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T9 (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T9 (.%) T9 (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T9 (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T9 (.%) Figure Sc: Most frequent gene-trees for simulated fly orthologs ( species). Simulated trees have similar errors at nearly the same relative frequencies as those found in real fly gene-trees.
4 PHYML MrBayes BIONJ Occurrence T T T (.%) T T T 9 (.%) T T T 9 (.%) T T T (.%) T T T (.%) T T T 9 (.%) T T T (.%) T T T 9 (.%) T T T (.%) T T T (.%) T T T (.9%) T T T (.%) T T T (.%) T T T (.%) T T T (.%) T T T (.%) T9 T9 T9 (.%) T T T (.%) T T T (.%) T T T (.%) Figure Sd: Most frequently constructed ortholog gene-trees for fly species. When methods are aquired to agree, reconstruction frequency for each topology decreases.
5 Name T T T T T T T T T T T T T T T T T9 T T T Topology ((((((spar,yeast),smik),sbay),scas),cgla),((agos,kwal),klac)) ((((((spar,yeast),smik),sbay),scas),cgla),((agos,klac),kwal)) (((((spar,yeast),smik),sbay),(cgla,scas)),((agos,kwal),klac)) ((((((spar,yeast),smik),sbay),cgla),scas),((agos,kwal),klac)) (((((spar,yeast),smik),sbay),(cgla,scas)),((agos,klac),kwal)) ((((((spar,yeast),smik),sbay),cgla),scas),((agos,klac),kwal)) ((((((spar,yeast),smik),sbay),scas),klac),((agos,kwal),cgla)) ((((((spar,yeast),smik),sbay),scas),cgla),((klac,kwal),agos)) (((((((spar,yeast),smik),sbay),scas),klac),kwal),(agos,cgla)) ((((((spar,yeast),smik),sbay),scas),(agos,kwal)),(cgla,klac)) (((((spar,yeast),smik),sbay),(cgla,scas)),((klac,kwal),agos)) ((((((spar,yeast),sbay),smik),scas),cgla),((agos,klac),kwal)) ((((((smik,yeast),spar),sbay),scas),cgla),((agos,kwal),klac)) ((((((spar,yeast),sbay),smik),cgla),scas),((agos,kwal),klac)) ((((((spar,yeast),smik),sbay),scas),(agos,klac)),(cgla,kwal)) (((((spar,yeast),smik),sbay),(klac,scas)),((agos,kwal),cgla)) ((((((spar,yeast),smik),sbay),scas),(klac,kwal)),(agos,cgla)) (((((spar,yeast),smik),sbay),(agos,kwal)),((klac,scas),cgla)) (((((spar,yeast),smik),sbay),klac),((agos,kwal),(cgla,scas))) (((((spar,yeast),smik),sbay),(agos,kwal)),((cgla,klac),scas)) Figure Sa: Naming scheme for ortholog gene-trees for 9 fungi species. Ordered by ML frequency. SPIDIR SPIDIR (D=.) SPIDIR (D=.) PHYML BIONJ MrBayes DNAPARS T (.%) T (.%) T (.%) T (.%) T (9.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.9%) T (9.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (9.%) T (.%) T (.9%) T (.%) T9 (.9%) T (.%) T (.%) T (.%) T (.9%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T9 (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T9 (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.9%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.9%) T (.%) T9 (.%) T (.%) T (.%) T (.9%) T (.%) T (.%) T (.%) T (.%) T (.%) T9 (.%) T (.9%) T9 (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.9%) T (.9%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.9%) T (.9%) T (.%) T (.%) T9 (.%) T (.%) T9 (.%) T (.%) T9 (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T9 (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T (.%) T9 (.%) T (.%) T (.%) T9 (.%) T (.%) T (.%) T (.%) T (.%) T9 (.%) Figure Sb: Most frequently constructed gene-tree topologies on 9 syntenic fungi orthologs. Topologies are numbered by their frequency across all methods.
6 Effect of gene length on reconstruction avg RF error.... spidir spidir-d. spidir-d. phyml bionj mrbayes parsimony bionj (ds) phyml (pep) bionj (pep) mrbayes (pep) parsimony (pep)..... gene length percentile Figure S: Robinson Foulds (RF) error correlates with sequence length for the 9 fly species. Acuracy improvements made by SPIDIR are also apparent for partial tree correctness (RF error). RF error is calculated as the fraction of internal branches that incorrectly bi-partition the leaves.
7 9 Fly orthologs Effect of percent identity on reconstruction 9 Fungal orthologs Effect of percent identity on reconstruction reconstruction accuracy.... SPIDIR (D=.) SPIDIR (D=.) SPIDIR SPIDIR SPIDIR (D=.) SPIDIR (D=.) PHYML BIONJ MrBayes Parsimony BIONJ(dS) PHYML(pep) BIONJ(pep) MrBayes (pep) Parsimony (pep) reconstruction accuracy.... SPIDIR (D=.) SPIDIR (D=..) SPIDIR SPIDIR SPIDIR (D=.) SPIDIR (D=.) PHYML BIONJ MrBayes Parsimony BIONJ (ds) PHYML (pep) BIONJ (pep) MrBayes (pep) Parsimony (pep) percent identity.... percent identity Figure S: Effect of mutation rate (nucleotide percent identity) on phylogenetic reconstruction accuracy. Reconstruction accuracy drops for both fast and slow evolving genes. Peptide alignments perform worse overall and saturate near. nucleotide percent identity, where most mutations are likely to beonly in the wobble codon position. Neighbor Joining on silent mutations (ds) increases slightly for slower evolving genes.
8 9 fly species (exclude dsec, dsim, dper) branch mean st dev st dev/mean dmel... dyak.9.. dere...9 dana..9. dpse.9.. dwil...9 dvir... dmoj.9..9 dgri fly species branch mean st dev st dev/mean dsim...9 dsec... dmel.9.. dyak... dere... dana... dpse... dper... dwil... dmoj...9 dvir.9.. dgri Figure S: Species-specific rates for the 9 and fly tree. Branches lengths are drawn with the mean species-specific rate. Thick line segments represent one standard deviation of the species-specific rate. Substitutions/site are relative, such that the branch lengths of each tree sums to one. The st dev/mean for each branch measures the tightness of its rate distribution.
9 dmel absolute branch lengths (a=.99, b=., Pval=.e-) dsim absolute branch lengths (a=.9, b=.9, Pval=9.9e-) dsec absolute branch lengths (a=., b=., Pval=.e-) dere absolute branch lengths (a=., b=., Pval=.e-) dyak absolute branch lengths (a=., b=9.9, Pval=.e-) dana absolute branch lengths (a=., b=., Pval=.e-) dpse absolute branch lengths (a=.9, b=., Pval=.e+) dper absolute branch lengths (a=., b=., Pval=.e+) dwil absolute branch lengths (a=., b=., Pval=.9e-) dmoj absolute branch lengths (a=., b=., Pval=.9e-) dvir absolute branch lengths (a=.9, b=., Pval=.e-) dgri absolute branch lengths (a=., b=., Pval=.e-) absolute branch lengths (a=.9, b=.9, Pval=.e-) absolute branch lengths (a=.99, b=., Pval=.e-) absolute branch lengths (a=., b=., Pval=.e-) absolute branch lengths (a=.9, b=.9, Pval=.e-) absolute branch lengths (a=., b=.9, Pval=.e-) absolute branch lengths (a=.9, b=., Pval=.e-) absolute branch lengths (a=., b=., Pval=.9e-) 9 absolute branch lengths (a=.9, b=., Pval=.e-) absolute branch lengths (a=.9, b=.9, Pval=.e-) absolute branch lengths (a=., b=9.9, Pval=.e-) Figure S: Branch length distributions fitted by gamma for every branch in the fly tree. out of absolute branch lengths significantly (p >.) fits the gamma distribution by Kolmogorov- Smirnov test (using middle 9% of lengths). Remaining branch lengths are closely approximated by gamma distribution. 9
10 dmel relative branch lengths (mean=.9, sdev=., pval=.9e-) dsim relative branch lengths (mean=., sdev=., pval=.e-) dsec relative branch lengths (mean=., sdev=., pval=.9e-) dere relative branch lengths (mean=., sdev=., pval=.9e-) rel rel rel rel dyak relative branch lengths (mean=., sdev=., pval=.e-) dana relative branch lengths (mean=., sdev=., pval=.9e-) dpse relative branch lengths (mean=.9, sdev=., pval=.e+) dper relative branch lengths (mean=., sdev=., pval=.e+) rel rel rel rel dwil relative branch lengths (mean=., sdev=., pval=.e-) dmoj relative branch lengths (mean=., sdev=., pval=.e-) dvir relative branch lengths (mean=., sdev=., pval=.9e-) dgri relative branch lengths (mean=., sdev=., pval=.9e-) rel rel rel rel relative branch lengths (mean=., sdev=.9, pval=.e-) relative branch lengths (mean=.9, sdev=.9, pval=.e-) relative branch lengths (mean=., sdev=.9, pval=.e-) relative branch lengths (mean=., sdev=., pval=9.9e-) rel rel rel rel relative branch lengths (mean=., sdev=., pval=.e-) relative branch lengths (mean=., sdev=.9, pval=.e-) relative branch lengths (mean=., sdev=., pval=.e-9) 9 relative branch lengths (mean=.9, sdev=., pval=.e-) rel rel rel rel relative branch lengths (mean=., sdev=.9, pval=.e-) relative branch lengths (mean=., sdev=., pval=.e-) rel rel Figure S9: Normalized length distributions fitted by normal for every branch in the fly tree. 9 out of relative branch lengths significantly (p >.) fits the normal distribution by Kolmogorov- Smirnov test (using middle 9% of lengths). Remaining branch lengths are closely approximated by normal distribution.
11 branch mean st dev st dev / mean scer..9. spar.9.. smik... sbay..9.9 scas.9.9. cgla... agos.99.. kwal.9.. klac Figure S: Species-specific rates for the 9 fungi tree. Branches lengths are drawn with the mean species-specific rate. Thick line segments represent one standard deviation of the species-specific rate. Substitutions/site are relative, such that the branch lengths of each tree sums to one. The st dev/mean for each branch measures the tightness of its rate distribution.
12 yeast absolute branch lengths (a=.9, b=9.99, Pval=.e-) spar absolute branch lengths (a=., b=9., Pval=.999e-) smik absolute branch lengths (a=.9, b=.9, Pval=.9e-) sbay absolute branch lengths (a=., b=., Pval=.e-) scas absolute branch lengths (a=., b=9., Pval=9.e-) cgla absolute branch lengths (a=., b=., Pval=.e-) klac absolute branch lengths (a=., b=., Pval=.e-) kwal absolute branch lengths (a=.9, b=9., Pval=.e-) agos absolute branch lengths (a=., b=., Pval=.9e-) absolute branch lengths (a=., b=., Pval=.e-) absolute branch lengths (a=.9, b=., Pval=.e-) absolute branch lengths (a=.9, b=., Pval=.9e-) absolute branch lengths (a=., b=., Pval=.e-) absolute branch lengths (a=9., b=., Pval=.e-) absolute branch lengths (a=., b=., Pval=.e-) absolute branch lengths (a=.9, b=9., Pval=.e-) Figure S: Branch length distributions fitted by gamma for every branch in the fungal tree. out of absolute branch lengths significantly (p >.) fits the gamma distribution by Kolmogorov- Smirnov test (using middle 9% of lengths).
13 yeast relative branch lengths (mean=., sdev=., pval=.9e-) spar relative branch lengths (mean=.9, sdev=., pval=.e-) smik relative branch lengths (mean=., sdev=., pval=.e-) sbay relative branch lengths (mean=., sdev=.9, pval=.9e-) rel rel rel rel scas relative branch lengths (mean=.9, sdev=., pval=.e-) cgla relative branch lengths (mean=., sdev=., pval=.e-) klac relative branch lengths (mean=., sdev=.9, pval=.e-) kwal relative branch lengths (mean=.9, sdev=., pval=.e-) rel rel rel rel agos relative branch lengths (mean=.99, sdev=., pval=.9e-) relative branch lengths (mean=., sdev=., pval=.9e-) relative branch lengths (mean=., sdev=., pval=.e-) relative branch lengths (mean=., sdev=.9, pval=.e-) rel rel rel rel relative branch lengths (mean=., sdev=., pval=.e-) relative branch lengths (mean=., sdev=., pval=9.e-) relative branch lengths (mean=., sdev=., pval=.e-) relative branch lengths (mean=., sdev=., pval=.e-) rel rel rel rel Figure S: Normalized length distributions fitted by normal for every branch in the fungal tree. out of relative branch lengths significantly (p >.) fits the normal distribution by Kolmogorov-Smirnov test (using middle 9% of lengths).
14 yeast spar smik sbay scas cgla klac kwal agos yeast spar smik sbay scas cgla klac kwal agos yeast spar smik sbay scas cgla klac kwal agos Absolution branch length correlations yeast spar smik sbay scas cgla klac kwal agos Relative branch length correlations.. -. Figure S: Absolute (left) and relative (right) branch length correlations within the fungal tree. Ancerstral species are numbered as shown.
15 gene tree counts 9% of gene trees Correlation with average gene tree Figure S: Histogram of gene tree branch length correlations between species tree branch lengths and fly ortholog gene trees. 9% of gene trees have a correlation greater than. with the species tree, indicating gene trees are highly correlated with each other.
16 fly branch variances fly branch means.... variance... mean dvir dpse dere dmoj dyak dwil dsec dgri dmel dper dsim dana. > gene length > gene length Figure S: Variances of species-specific rates decrease with increasing sequence length, indicating that the variance of a species-specific distribution is partly due to distance estimation error (left). Speciesspecific means are not affected by sequence length (right). Mean and variance were calculated for relative branch lengths of fly gene trees binned by gene length into four bins (<,,, > ).
17 TAB] DATASET : Beta Hemoglobin Orthologs Method Topology chosen Ratio Result Topology Topology SPIDIR likelihood Topology. Correct.9.9 Max likelihood Topology 9.9 Wrong Neighbor Joining Topology. Wrong Parsimony Topology. Wrong Species Expected Topology Topology Likelihood branch Length Stdev Branch Abs length Rel length Likelihood Branch Abs length Rel length Likelihood comparison mouse.9. d... y.... rat.. c... x human.. e.... hmr+human.9.99 w... dog.. v..9.. dog+hmr.. a+f..99. mr.. b.9... hmr+mr.. z.9.9. Family rate alpha=. beta=.99 rate=..9 rate=... Total.9.9. DATASET : Alpha and Beta Hemoglobin Paralogs Method Topology chosen Ratio Result Topology Topology SPIDIR likelihood Topology. Correct. 9. Max likelihood Topology.E+ Correct Neighbor Joining Topology Correct. 99. Parsimony Topology. Correct 9.. Species Expected Topology Topology Likelihood branch Length Stdev Branch Abs length Rel length Likelihood Branch Abs length Rel length Likelihood comparison mouse.9. d... y....9 rat.. c...9 x.... human.. e.... hmr+human.9.99 w...9 dog.. v.... dog+hmr.. a+f... mr.. b.... hmr+mr.. z... Family rate alpha=. beta=.99 rate=.9. rate= Total. 9.. Figure S: (Dataset ) Evaluation of two topologies (Figure b, e) for four hemoglobinbeta orthologs by three phylogenetic methods and SPIDIR. Due to long branch attraction, each method finds the incorrect Topology as more likeliy. In constrast, SPIDIR uses learned branch length distributions (Expected) to compute the likelihood of each branch and concludes that Topology is -fold more likely. (Dataset ) Evaluation of two topologies (Figure b, e) for two hemoglobin-beta orthologs (mouse and rat) and two hemoglobin-alpha orthologs (human and dog). Since these genes are related by an ancestral duplication that is well before the mammalian speciation, the branch lengths support the topology T with a -fold likelihood ratio.
18 branch bootstrap corresponds to branch accuracy on fly orthologous alignments. accuracy (fraction conguent to species tree)... bootstrap Figure S: Branch bootstrap values from reconstructing fly ortholog trees plotted against probability of branch being correct. Probability of correctness for a branch with bootstrap X was estimated by percentage of correct branches with bootstraps within X / to X + /.
19 a comparison of accuracy for each go term. spidir accuracy phyml accuracy b GO Term p-value SPIDIR PHYML total oocyte microtubule cytoskeleton polarization.9 (.%) (.%) mrna cleavage and polyadenylation specificity factor complex. (.%) (.%) establishment and/or maintenance of cytoskeleton polarity. (.%) (.%) establishment and/or maintenance of microtubule cytoskeleton polarity. (.%) (.%) oocyte microtubule cytoskeleton organization. (.%) (.%) mannosyl-oligosaccharide mannosidase activity. (.%) (.%) mannosidase activity.9 (.%) (.%) antifungal humoral response (sensu Protostomia).9 (.%) (.%) endoribonuclease activity. (.%) (.%) structural constituent of peritrophic membrane (sensu Insecta). (.%) (.%) RAB small monomeric GTPase activity. (.%) (.%) SH/SH adaptor protein activity. (.%) (.%) carboxylesterase activity. (.%) (.%) transmembrane receptor protein tyrosine kinase docking protein activity. (.%) (.%) transmembrane receptor protein tyrosine kinase signaling protein activity. (.%) (.%) Figure S: (a) Accuracy comparison of SPIDIR vs PHYML for each GO term. Each tree was associated with each GO term of the D. melanogaster gene. (b) GO terms for which SPIDIR most significantly under performs PHYML. P-values are computed using the hypergeometric test. P-values above have not been corrected for multiple-testing. Doing so would decrease significance even further. No GO term performs significantly worse (p >.) for SPIDIR than for PHYML. 9
20 klac to scer (mu=.9, sigma=.) (mu=., sigma=.) klac to spar (mu=.9, sigma=.) (mu=., sigma=.) klac to smik (mu=.9, sigma=.) (mu=.9, sigma=.) klac to sbay (mu=.9, sigma=.) (mu=., sigma=.) klac to scas (mu=.9, sigma=.) (mu=.9, sigma=.) klac to cgla (mu=., sigma=.) (mu=., sigma=.) Figure S9: Branch distributions of trees with duplication approximated by normal. For each of the gene families in our WGD dataset, we identified the distance between one pre-duplication species, K. lactis, and two ohnologs in a post-duplication species. These distances were normalized by the estimated family rate for each gene tree. The histogram of branch lengths is plotted against the normal expected by the model (sum of several independent normals).
Principles of phylogenetic analysis
 Principles of phylogenetic analysis Arne Holst-Jensen, NVI, Norway. Fusarium course, Ås, Norway, June 22 nd 2008 Distance based methods Compare C OTUs and characters X A + D = Pairwise: A and B; X characters
Principles of phylogenetic analysis Arne Holst-Jensen, NVI, Norway. Fusarium course, Ås, Norway, June 22 nd 2008 Distance based methods Compare C OTUs and characters X A + D = Pairwise: A and B; X characters
Data and text mining applied to the computational study of protein interaction networks
 Data and text mining applied to the computational study of protein interaction networks Miguel Andrade Faculty of Biology, Johannes Gutenberg University Institute of Molecular Biology Mainz, Germany andrade@uni-mainz.de
Data and text mining applied to the computational study of protein interaction networks Miguel Andrade Faculty of Biology, Johannes Gutenberg University Institute of Molecular Biology Mainz, Germany andrade@uni-mainz.de
DATA. Miriah Meyer University of Utah. cs6964 January
 cs6964 January 19 212 DATA Miriah Meyer University of Utah slide acknowledgements: Tamara Munzner, University of British Columbia Hanspeter Pfister, Harvard University 2 LAST TIME 3 target translate design
cs6964 January 19 212 DATA Miriah Meyer University of Utah slide acknowledgements: Tamara Munzner, University of British Columbia Hanspeter Pfister, Harvard University 2 LAST TIME 3 target translate design
Nature Neuroscience: doi: /nn Supplementary Figure 1. Behavioral training.
 Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
The BLAST search on NCBI ( and GISAID
 Supplemental materials and methods The BLAST search on NCBI (http:// www.ncbi.nlm.nih.gov) and GISAID (http://www.platform.gisaid.org) showed that hemagglutinin (HA) gene of North American H5N1, H5N2 and
Supplemental materials and methods The BLAST search on NCBI (http:// www.ncbi.nlm.nih.gov) and GISAID (http://www.platform.gisaid.org) showed that hemagglutinin (HA) gene of North American H5N1, H5N2 and
FINAL ANNOTATION REPORT: Drosophila virilis Fosmid 11 (48P14) Robert Carrasquillo Bio 4342
 FINAL ANNOTATION REPORT: Drosophila virilis Fosmid 11 (48P14) Robert Carrasquillo Bio 4342 2006 TABLE OF CONTENTS I. Overview... 3 II. Genes... 4 III. Clustal Analysis... 15 IV. Repeat Analysis... 17 V.
FINAL ANNOTATION REPORT: Drosophila virilis Fosmid 11 (48P14) Robert Carrasquillo Bio 4342 2006 TABLE OF CONTENTS I. Overview... 3 II. Genes... 4 III. Clustal Analysis... 15 IV. Repeat Analysis... 17 V.
Phylogenetic Methods
 Phylogenetic Methods Multiple Sequence lignment Pairwise distance matrix lustering algorithms: NJ, UPM - guide trees Phylogenetic trees Nucleotide vs. amino acid sequences for phylogenies ) Nucleotides:
Phylogenetic Methods Multiple Sequence lignment Pairwise distance matrix lustering algorithms: NJ, UPM - guide trees Phylogenetic trees Nucleotide vs. amino acid sequences for phylogenies ) Nucleotides:
NJMerge: A generic technique for scaling phylogeny estimation methods and its application to species trees Supplementary Materials
 NJMerge: A generic technique for scaling phylogeny estimation methods and its application to species trees Supplementary Materials Erin K. Molloy 1[ 1 5553 3312] and Tandy Warnow 1[ 1 7717 3514] Department
NJMerge: A generic technique for scaling phylogeny estimation methods and its application to species trees Supplementary Materials Erin K. Molloy 1[ 1 5553 3312] and Tandy Warnow 1[ 1 7717 3514] Department
Cahn - Ingold - Prelog system. Proteins: Evolution, and Analysis Lecture 7 9/15/2009. The Fischer Convention (1) G (2) (3)
 Chapter 4 (1) G Proteins: Evolution, and Analysis Lecture 7 9/15/2009 A V L I M P F W Chapter 4 (2) S (3) T N Q Y C K R H D E The Fischer Convention Absolute configuration about an asymmetric carbon related
Chapter 4 (1) G Proteins: Evolution, and Analysis Lecture 7 9/15/2009 A V L I M P F W Chapter 4 (2) S (3) T N Q Y C K R H D E The Fischer Convention Absolute configuration about an asymmetric carbon related
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project
 Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Evolutionary interactions between haemagglutinin and neuraminidase in avian influenza
 Ward et al. BMC Evolutionary Biology 2013, 13:222 RESEARCH ARTICLE Open Access Evolutionary interactions between haemagglutinin and neuraminidase in avian influenza Melissa J Ward 1*, Samantha J Lycett
Ward et al. BMC Evolutionary Biology 2013, 13:222 RESEARCH ARTICLE Open Access Evolutionary interactions between haemagglutinin and neuraminidase in avian influenza Melissa J Ward 1*, Samantha J Lycett
Name: Due on Wensday, December 7th Bioinformatics Take Home Exam #9 Pick one most correct answer, unless stated otherwise!
 Name: Due on Wensday, December 7th Bioinformatics Take Home Exam #9 Pick one most correct answer, unless stated otherwise! 1. What process brought 2 divergent chlorophylls into the ancestor of the cyanobacteria,
Name: Due on Wensday, December 7th Bioinformatics Take Home Exam #9 Pick one most correct answer, unless stated otherwise! 1. What process brought 2 divergent chlorophylls into the ancestor of the cyanobacteria,
Going Nowhere Fast: Lentivirus genetic sequence evolution does not correlate with phenotypic evolution.
 Going Nowhere Fast: Lentivirus genetic sequence evolution does not correlate with phenotypic evolution. Brian T. Foley, PhD btf@lanl.gov HIV Genetic Sequences, Immunology, Drug Resistance and Vaccine Trials
Going Nowhere Fast: Lentivirus genetic sequence evolution does not correlate with phenotypic evolution. Brian T. Foley, PhD btf@lanl.gov HIV Genetic Sequences, Immunology, Drug Resistance and Vaccine Trials
Adaptive phylogeography: functional divergence between haemoglobins derived from different glacial refugia in the bank vole
 Electronic supplementary material for: Adaptive phylogeography: functional divergence between haemoglobins derived from different glacial refugia in the bank vole Petr Kotlík, Silvia Marková, Libor Vojtek,
Electronic supplementary material for: Adaptive phylogeography: functional divergence between haemoglobins derived from different glacial refugia in the bank vole Petr Kotlík, Silvia Marková, Libor Vojtek,
Nature Neuroscience: doi: /nn Supplementary Figure 1. Neuron class-specific arrangements of Khc::nod::lacZ label in dendrites.
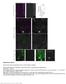 Supplementary Figure 1 Neuron class-specific arrangements of Khc::nod::lacZ label in dendrites. Staining with fluorescence antibodies to detect GFP (Green), β-galactosidase (magenta/white). (a, b) Class
Supplementary Figure 1 Neuron class-specific arrangements of Khc::nod::lacZ label in dendrites. Staining with fluorescence antibodies to detect GFP (Green), β-galactosidase (magenta/white). (a, b) Class
Estimating Phylogenies (Evolutionary Trees) I
 stimating Phylogenies (volutionary Trees) I iol4230 Tues, Feb 27, 2018 ill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Goals of today s lecture: Why estimate phylogenies? Origin of man (woman) Origin of
stimating Phylogenies (volutionary Trees) I iol4230 Tues, Feb 27, 2018 ill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Goals of today s lecture: Why estimate phylogenies? Origin of man (woman) Origin of
Unit 1 Exploring and Understanding Data
 Unit 1 Exploring and Understanding Data Area Principle Bar Chart Boxplot Conditional Distribution Dotplot Empirical Rule Five Number Summary Frequency Distribution Frequency Polygon Histogram Interquartile
Unit 1 Exploring and Understanding Data Area Principle Bar Chart Boxplot Conditional Distribution Dotplot Empirical Rule Five Number Summary Frequency Distribution Frequency Polygon Histogram Interquartile
Annotation of Drosophila mojavensis fosmid 8 Priya Srikanth Bio 434W
 Annotation of Drosophila mojavensis fosmid 8 Priya Srikanth Bio 434W 5.1.2007 Overview High-quality finished sequence is much more useful for research once it is annotated. Annotation is a fundamental
Annotation of Drosophila mojavensis fosmid 8 Priya Srikanth Bio 434W 5.1.2007 Overview High-quality finished sequence is much more useful for research once it is annotated. Annotation is a fundamental
Integrative Biology 200A PRINCIPLES OF PHYLOGENETICS Spring 2012
 Integrative Biology 200A PRINCIPLES OF PHYLOGENETICS Spring 2012 University of California, Berkeley Kipling Will- 1 March Data/Hypothesis Exploration and Support Measures I. Overview. -- Many would agree
Integrative Biology 200A PRINCIPLES OF PHYLOGENETICS Spring 2012 University of California, Berkeley Kipling Will- 1 March Data/Hypothesis Exploration and Support Measures I. Overview. -- Many would agree
Maria D Vibranovski 1,6*, Yong E Zhang 1,2, Claus Kemkemer 1, Nicholas W VanKuren 1,3, Hedibert F Lopes 4, Timothy L Karr 5 and Manyuan Long 1*
 Vibranovski et al. BMC Evolutionary Biology 2012, 12:169 CORRESPONDENCE Open Access Segmental dataset and whole body expression data do not support the hypothesis that non-random movement is an intrinsic
Vibranovski et al. BMC Evolutionary Biology 2012, 12:169 CORRESPONDENCE Open Access Segmental dataset and whole body expression data do not support the hypothesis that non-random movement is an intrinsic
Gene Ontology and Functional Enrichment. Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein
 Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
PSYCH-GA.2211/NEURL-GA.2201 Fall 2016 Mathematical Tools for Cognitive and Neural Science. Homework 5
 PSYCH-GA.2211/NEURL-GA.2201 Fall 2016 Mathematical Tools for Cognitive and Neural Science Homework 5 Due: 21 Dec 2016 (late homeworks penalized 10% per day) See the course web site for submission details.
PSYCH-GA.2211/NEURL-GA.2201 Fall 2016 Mathematical Tools for Cognitive and Neural Science Homework 5 Due: 21 Dec 2016 (late homeworks penalized 10% per day) See the course web site for submission details.
BMC Evolutionary Biology
 BMC Evolutionary Biology This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. Segmental dataset
BMC Evolutionary Biology This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. Segmental dataset
Evolution of influenza
 Evolution of influenza Today: 1. Global health impact of flu - why should we care? 2. - what are the components of the virus and how do they change? 3. Where does influenza come from? - are there animal
Evolution of influenza Today: 1. Global health impact of flu - why should we care? 2. - what are the components of the virus and how do they change? 3. Where does influenza come from? - are there animal
Effects of Branch Length Uncertainty on Bayesian Posterior Probabilities for Phylogenetic Hypotheses
 Effects of Branch Length Uncertainty on Bayesian Posterior Probabilities for Phylogenetic Hypotheses Bryan Kolaczkowski and Joseph W. Thornton Center for Ecology and Evolutionary Biology, University of
Effects of Branch Length Uncertainty on Bayesian Posterior Probabilities for Phylogenetic Hypotheses Bryan Kolaczkowski and Joseph W. Thornton Center for Ecology and Evolutionary Biology, University of
The Alternative Choice of Constitutive Exons throughout Evolution
 The Alternative Choice of Constitutive Exons throughout Evolution Galit Lev-Maor 1[, Amir Goren 1[, Noa Sela 1[, Eddo Kim 1, Hadas Keren 1, Adi Doron-Faigenboim 2, Shelly Leibman-Barak 3, Tal Pupko 2,
The Alternative Choice of Constitutive Exons throughout Evolution Galit Lev-Maor 1[, Amir Goren 1[, Noa Sela 1[, Eddo Kim 1, Hadas Keren 1, Adi Doron-Faigenboim 2, Shelly Leibman-Barak 3, Tal Pupko 2,
WHEN DO MUTATIONS OCCUR?
 WHEN DO MUTATIONS OCCUR? While most DNA replicates with fairly high accuracy, mistakes do happen. DNA polymerase sometimes inserts the wrong nucleotide or too many or too few nucleotides into a sequence.
WHEN DO MUTATIONS OCCUR? While most DNA replicates with fairly high accuracy, mistakes do happen. DNA polymerase sometimes inserts the wrong nucleotide or too many or too few nucleotides into a sequence.
Remarkable expansions of an X-linked reproductive homeobox gene cluster in rodent evolution
 Genomics 88 (2006) 34 43 www.elsevier.com/locate/ygeno Remarkable expansions of an X-linked reproductive homeobox gene cluster in rodent evolution Xiaoxia Wang, Jianzhi Zhang Department of Ecology and
Genomics 88 (2006) 34 43 www.elsevier.com/locate/ygeno Remarkable expansions of an X-linked reproductive homeobox gene cluster in rodent evolution Xiaoxia Wang, Jianzhi Zhang Department of Ecology and
Supplementary figure legends
 Supplementary figure legends SUPPLEMENTRY FIGURE S1. Lentiviral construct. Schematic representation of the PCR fragment encompassing the genomic locus of mir-33a that was introduced in the lentiviral construct.
Supplementary figure legends SUPPLEMENTRY FIGURE S1. Lentiviral construct. Schematic representation of the PCR fragment encompassing the genomic locus of mir-33a that was introduced in the lentiviral construct.
Distinguishing epidemiological dependent from treatment (resistance) dependent HIV mutations: Problem Statement
 Distinguishing epidemiological dependent from treatment (resistance) dependent HIV mutations: Problem Statement Leander Schietgat 1, Kristof Theys 2, Jan Ramon 1, Hendrik Blockeel 1, and Anne-Mieke Vandamme
Distinguishing epidemiological dependent from treatment (resistance) dependent HIV mutations: Problem Statement Leander Schietgat 1, Kristof Theys 2, Jan Ramon 1, Hendrik Blockeel 1, and Anne-Mieke Vandamme
Skala Stress. Putaran 1 Reliability. Case Processing Summary. N % Excluded a 0.0 Total
 Skala Stress Putaran 1 Reliability Case Processing Summary N % Cases Valid Excluded a 0.0 Total a. Listwise deletion based on all variables in the procedure. Reliability Statistics Cronbach's Alpha N of
Skala Stress Putaran 1 Reliability Case Processing Summary N % Cases Valid Excluded a 0.0 Total a. Listwise deletion based on all variables in the procedure. Reliability Statistics Cronbach's Alpha N of
NEUROBLASTOMA DATA -- TWO GROUPS -- QUANTITATIVE MEASURES 38 15:37 Saturday, January 25, 2003
 NEUROBLASTOMA DATA -- TWO GROUPS -- QUANTITATIVE MEASURES 38 15:37 Saturday, January 25, 2003 Obs GROUP I DOPA LNDOPA 1 neurblst 1 48.000 1.68124 2 neurblst 1 133.000 2.12385 3 neurblst 1 34.000 1.53148
NEUROBLASTOMA DATA -- TWO GROUPS -- QUANTITATIVE MEASURES 38 15:37 Saturday, January 25, 2003 Obs GROUP I DOPA LNDOPA 1 neurblst 1 48.000 1.68124 2 neurblst 1 133.000 2.12385 3 neurblst 1 34.000 1.53148
SMPD 287 Spring 2015 Bioinformatics in Medical Product Development. Final Examination
 Final Examination You have a choice between A, B, or C. Please email your solutions, as a pdf attachment, by May 13, 2015. In the subject of the email, please use the following format: firstname_lastname_x
Final Examination You have a choice between A, B, or C. Please email your solutions, as a pdf attachment, by May 13, 2015. In the subject of the email, please use the following format: firstname_lastname_x
SUPPLEMENTARY INFORMATION
 Testing the accuracy of ancestral state reconstruction The accuracy of the ancestral state reconstruction with maximum likelihood methods can depend on the underlying model used in the reconstruction.
Testing the accuracy of ancestral state reconstruction The accuracy of the ancestral state reconstruction with maximum likelihood methods can depend on the underlying model used in the reconstruction.
Phylogenetic Tree Practical Problems
 Phylogenetic Tree Practical Problems Software Tools: MEGA A software package for constructing phylogenetic trees using neighbor-joining, UPGMA, and maximum parsimony. ClustalW A tool for constructing multiple
Phylogenetic Tree Practical Problems Software Tools: MEGA A software package for constructing phylogenetic trees using neighbor-joining, UPGMA, and maximum parsimony. ClustalW A tool for constructing multiple
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Interactive analysis and quality assessment of single-cell copy-number variations
 Interactive analysis and quality assessment of single-cell copy-number variations Tyler Garvin, Robert Aboukhalil, Jude Kendall, Timour Baslan, Gurinder S. Atwal, James Hicks, Michael Wigler, Michael C.
Interactive analysis and quality assessment of single-cell copy-number variations Tyler Garvin, Robert Aboukhalil, Jude Kendall, Timour Baslan, Gurinder S. Atwal, James Hicks, Michael Wigler, Michael C.
Relaxation of Selective Constraints Causes Independent Selenoprotein Extinction in Insect Genomes
 Relaxation of Selective Constraints Causes Independent Selenoprotein Extinction in Insect Genomes Charles E. Chapple 1, Roderic Guigó 2 * 1 Center for Genomic Regulation, Universitat Pompeu Fabra and Institut
Relaxation of Selective Constraints Causes Independent Selenoprotein Extinction in Insect Genomes Charles E. Chapple 1, Roderic Guigó 2 * 1 Center for Genomic Regulation, Universitat Pompeu Fabra and Institut
Reliable reconstruction of HIV-1 whole genome haplotypes reveals clonal interference and genetic hitchhiking among immune escape variants
 Pandit and de Boer Retrovirology 2014, 11:56 RESEARCH Open Access Reliable reconstruction of HIV-1 whole genome haplotypes reveals clonal interference and genetic hitchhiking among immune escape variants
Pandit and de Boer Retrovirology 2014, 11:56 RESEARCH Open Access Reliable reconstruction of HIV-1 whole genome haplotypes reveals clonal interference and genetic hitchhiking among immune escape variants
Lecture 15. Signal Transduction Pathways - Introduction
 Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood
 Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Colon cancer subtypes from gene expression data
 Colon cancer subtypes from gene expression data Nathan Cunningham Giuseppe Di Benedetto Sherman Ip Leon Law Module 6: Applied Statistics 26th February 2016 Aim Replicate findings of Felipe De Sousa et
Colon cancer subtypes from gene expression data Nathan Cunningham Giuseppe Di Benedetto Sherman Ip Leon Law Module 6: Applied Statistics 26th February 2016 Aim Replicate findings of Felipe De Sousa et
Methods for Determining Random Sample Size
 Methods for Determining Random Sample Size This document discusses how to determine your random sample size based on the overall purpose of your research project. Methods for determining the random sample
Methods for Determining Random Sample Size This document discusses how to determine your random sample size based on the overall purpose of your research project. Methods for determining the random sample
Evaluating Classifiers for Disease Gene Discovery
 Evaluating Classifiers for Disease Gene Discovery Kino Coursey Lon Turnbull khc0021@unt.edu lt0013@unt.edu Abstract Identification of genes involved in human hereditary disease is an important bioinfomatics
Evaluating Classifiers for Disease Gene Discovery Kino Coursey Lon Turnbull khc0021@unt.edu lt0013@unt.edu Abstract Identification of genes involved in human hereditary disease is an important bioinfomatics
Sawtooth Software. MaxDiff Analysis: Simple Counting, Individual-Level Logit, and HB RESEARCH PAPER SERIES. Bryan Orme, Sawtooth Software, Inc.
 Sawtooth Software RESEARCH PAPER SERIES MaxDiff Analysis: Simple Counting, Individual-Level Logit, and HB Bryan Orme, Sawtooth Software, Inc. Copyright 009, Sawtooth Software, Inc. 530 W. Fir St. Sequim,
Sawtooth Software RESEARCH PAPER SERIES MaxDiff Analysis: Simple Counting, Individual-Level Logit, and HB Bryan Orme, Sawtooth Software, Inc. Copyright 009, Sawtooth Software, Inc. 530 W. Fir St. Sequim,
List of Figures. List of Tables. Preface to the Second Edition. Preface to the First Edition
 List of Figures List of Tables Preface to the Second Edition Preface to the First Edition xv xxv xxix xxxi 1 What Is R? 1 1.1 Introduction to R................................ 1 1.2 Downloading and Installing
List of Figures List of Tables Preface to the Second Edition Preface to the First Edition xv xxv xxix xxxi 1 What Is R? 1 1.1 Introduction to R................................ 1 1.2 Downloading and Installing
Journal: Nature Methods
 Journal: Nature Methods Article Title: Network-based stratification of tumor mutations Corresponding Author: Trey Ideker Supplementary Item Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure
Journal: Nature Methods Article Title: Network-based stratification of tumor mutations Corresponding Author: Trey Ideker Supplementary Item Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure
RNA-seq: filtering, quality control and visualisation. COMBINE RNA-seq Workshop
 RNA-seq: filtering, quality control and visualisation COMBINE RNA-seq Workshop QC and visualisation (part 1) Slide taken from COMBINE RNAseq workshop on 23/09/2016 RNA-seq of Mouse mammary gland Basal
RNA-seq: filtering, quality control and visualisation COMBINE RNA-seq Workshop QC and visualisation (part 1) Slide taken from COMBINE RNAseq workshop on 23/09/2016 RNA-seq of Mouse mammary gland Basal
Mapping Evolutionary Pathways of HIV-1 Drug Resistance. Christopher Lee, UCLA Dept. of Chemistry & Biochemistry
 Mapping Evolutionary Pathways of HIV-1 Drug Resistance Christopher Lee, UCLA Dept. of Chemistry & Biochemistry Stalemate: We React to them, They React to Us E.g. a virus attacks us, so we develop a drug,
Mapping Evolutionary Pathways of HIV-1 Drug Resistance Christopher Lee, UCLA Dept. of Chemistry & Biochemistry Stalemate: We React to them, They React to Us E.g. a virus attacks us, so we develop a drug,
Supplemental Materials. for. Conservation of an RNA Regulatory Map between Drosophila and. Mammals
 Supplemental Materials for Conservation of an RNA Regulatory Map between Drosophila and Mammals Angela N. Brooks, Li Yang, Michael O. Duff, Kasper Daniel Hansen, Jung W. Park, Sandrine Dudoit, Steven E.
Supplemental Materials for Conservation of an RNA Regulatory Map between Drosophila and Mammals Angela N. Brooks, Li Yang, Michael O. Duff, Kasper Daniel Hansen, Jung W. Park, Sandrine Dudoit, Steven E.
Les phéromones de la Drosophil hile : évoluti tion et rôle dans la spéciation e i Wicker Th omas
 Les phéromones de la Drosophile : évolution et rôle dans la spéciation Claude Wicker-Thomas legs, UPR 9034, 91198 Gif sur Yvette Drosophila courtship pheromones pheromones pheromones pheromones Pheromones
Les phéromones de la Drosophile : évolution et rôle dans la spéciation Claude Wicker-Thomas legs, UPR 9034, 91198 Gif sur Yvette Drosophila courtship pheromones pheromones pheromones pheromones Pheromones
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10776 Supplementary Information 1: Influence of inhibition among blns on STDP of KC-bLN synapses (simulations and schematics). Unconstrained STDP drives network activity to saturation
doi:10.1038/nature10776 Supplementary Information 1: Influence of inhibition among blns on STDP of KC-bLN synapses (simulations and schematics). Unconstrained STDP drives network activity to saturation
Modeling Sentiment with Ridge Regression
 Modeling Sentiment with Ridge Regression Luke Segars 2/20/2012 The goal of this project was to generate a linear sentiment model for classifying Amazon book reviews according to their star rank. More generally,
Modeling Sentiment with Ridge Regression Luke Segars 2/20/2012 The goal of this project was to generate a linear sentiment model for classifying Amazon book reviews according to their star rank. More generally,
SiFit: inferring tumor trees from single-cell sequencing data under finite-sites models
 Zafar et al. Genome Biology (2017) 18:178 DOI 10.1186/s13059-017-1311-2 METHOD Open Access SiFit: inferring tumor trees from single-cell sequencing data under finite-sites models Hamim Zafar 1,2, Anthony
Zafar et al. Genome Biology (2017) 18:178 DOI 10.1186/s13059-017-1311-2 METHOD Open Access SiFit: inferring tumor trees from single-cell sequencing data under finite-sites models Hamim Zafar 1,2, Anthony
It is well known that some pathogenic microbes undergo
 Colloquium Effects of passage history and sampling bias on phylogenetic reconstruction of human influenza A evolution Robin M. Bush, Catherine B. Smith, Nancy J. Cox, and Walter M. Fitch Department of
Colloquium Effects of passage history and sampling bias on phylogenetic reconstruction of human influenza A evolution Robin M. Bush, Catherine B. Smith, Nancy J. Cox, and Walter M. Fitch Department of
Minimalistic encapsulated proteomic sample processing applied to copy number
 Nature Methods Minimalistic encapsulated proteomic sample processing applied to copy number estimation in eukaryotic cells Nils A. Kulak, Garwin Pichler, Igor Paron, Nagarjuna Nagaraj and Matthias Mann
Nature Methods Minimalistic encapsulated proteomic sample processing applied to copy number estimation in eukaryotic cells Nils A. Kulak, Garwin Pichler, Igor Paron, Nagarjuna Nagaraj and Matthias Mann
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
Evolutionary distances between proteins of the Influenza A Virus
 Evolutionary distances between proteins of the Influenza A Virus Hatem Nassrat B00393388 Bioinformatics: Winter 06/07 Dr. Christian Blouin Table of Contents Evolutionary distances between proteins of the
Evolutionary distances between proteins of the Influenza A Virus Hatem Nassrat B00393388 Bioinformatics: Winter 06/07 Dr. Christian Blouin Table of Contents Evolutionary distances between proteins of the
BEAST Bayesian Evolutionary Analysis Sampling Trees
 BEAST Bayesian Evolutionary Analysis Sampling Trees Introduction Revealing the evolutionary dynamics of influenza This tutorial provides a step-by-step explanation on how to reconstruct the evolutionary
BEAST Bayesian Evolutionary Analysis Sampling Trees Introduction Revealing the evolutionary dynamics of influenza This tutorial provides a step-by-step explanation on how to reconstruct the evolutionary
Stats 95. Statistical analysis without compelling presentation is annoying at best and catastrophic at worst. From raw numbers to meaningful pictures
 Stats 95 Statistical analysis without compelling presentation is annoying at best and catastrophic at worst. From raw numbers to meaningful pictures Stats 95 Why Stats? 200 countries over 200 years http://www.youtube.com/watch?v=jbksrlysojo
Stats 95 Statistical analysis without compelling presentation is annoying at best and catastrophic at worst. From raw numbers to meaningful pictures Stats 95 Why Stats? 200 countries over 200 years http://www.youtube.com/watch?v=jbksrlysojo
Yeast Cells Classification Machine Learning Approach to Discriminate Saccharomyces cerevisiae Yeast Cells Using Sophisticated Image Features.
 Yeast Cells Classification Machine Learning Approach to Discriminate Saccharomyces cerevisiae Yeast Cells Using Sophisticated Image Features. Mohamed Tleis Supervisor: Fons J. Verbeek Leiden University
Yeast Cells Classification Machine Learning Approach to Discriminate Saccharomyces cerevisiae Yeast Cells Using Sophisticated Image Features. Mohamed Tleis Supervisor: Fons J. Verbeek Leiden University
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data
 Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Mouse Clec9a ORF sequence
 Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Rajesh Kannangai Phone: ; Fax: ; *Corresponding author
 Amino acid sequence divergence of Tat protein (exon1) of subtype B and C HIV-1 strains: Does it have implications for vaccine development? Abraham Joseph Kandathil 1, Rajesh Kannangai 1, *, Oriapadickal
Amino acid sequence divergence of Tat protein (exon1) of subtype B and C HIV-1 strains: Does it have implications for vaccine development? Abraham Joseph Kandathil 1, Rajesh Kannangai 1, *, Oriapadickal
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE
 Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
Inferring Biological Meaning from Cap Analysis Gene Expression Data
 Inferring Biological Meaning from Cap Analysis Gene Expression Data HRYSOULA PAPADAKIS 1. Introduction This project is inspired by the recent development of the Cap analysis gene expression (CAGE) method,
Inferring Biological Meaning from Cap Analysis Gene Expression Data HRYSOULA PAPADAKIS 1. Introduction This project is inspired by the recent development of the Cap analysis gene expression (CAGE) method,
Chapter 1: Exploring Data
 Chapter 1: Exploring Data Key Vocabulary:! individual! variable! frequency table! relative frequency table! distribution! pie chart! bar graph! two-way table! marginal distributions! conditional distributions!
Chapter 1: Exploring Data Key Vocabulary:! individual! variable! frequency table! relative frequency table! distribution! pie chart! bar graph! two-way table! marginal distributions! conditional distributions!
Early Learning vs Early Variability 1.5 r = p = Early Learning r = p = e 005. Early Learning 0.
 The temporal structure of motor variability is dynamically regulated and predicts individual differences in motor learning ability Howard Wu *, Yohsuke Miyamoto *, Luis Nicolas Gonzales-Castro, Bence P.
The temporal structure of motor variability is dynamically regulated and predicts individual differences in motor learning ability Howard Wu *, Yohsuke Miyamoto *, Luis Nicolas Gonzales-Castro, Bence P.
Research Strategy: 1. Background and Significance
 Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
LOGO. Statistical Modeling of Breast and Lung Cancers. Cancer Research Team. Department of Mathematics and Statistics University of South Florida
 LOGO Statistical Modeling of Breast and Lung Cancers Cancer Research Team Department of Mathematics and Statistics University of South Florida 1 LOGO 2 Outline Nonparametric and parametric analysis of
LOGO Statistical Modeling of Breast and Lung Cancers Cancer Research Team Department of Mathematics and Statistics University of South Florida 1 LOGO 2 Outline Nonparametric and parametric analysis of
Figure 1: Final annotation map of Contig 9
 Introduction With rapid advances in sequencing technology, particularly with the development of second and third generation sequencing, genomes for organisms from all kingdoms and many phyla have been
Introduction With rapid advances in sequencing technology, particularly with the development of second and third generation sequencing, genomes for organisms from all kingdoms and many phyla have been
EVOLUTIONARY TRAJECTORY ANALYSIS: RECENT ENHANCEMENTS. R. Burke Squires
 EVOLUTIONARY TRAJECTORY ANALYSIS: RECENT ENHANCEMENTS R. Burke Squires Pandemic H1N1 2009 Origin? April / May 2009 Cases of an Influenza-like Illness (ILI) occurred in California, Texas and Mexico New
EVOLUTIONARY TRAJECTORY ANALYSIS: RECENT ENHANCEMENTS R. Burke Squires Pandemic H1N1 2009 Origin? April / May 2009 Cases of an Influenza-like Illness (ILI) occurred in California, Texas and Mexico New
Nature Genetics: doi: /ng Supplementary Figure 1. Clinical timeline for the discovery WES cases.
 Supplementary Figure 1 Clinical timeline for the discovery WES cases. This illustrates the timeline of the disease events during the clinical course of each patient s disease, further indicating the available
Supplementary Figure 1 Clinical timeline for the discovery WES cases. This illustrates the timeline of the disease events during the clinical course of each patient s disease, further indicating the available
Annotation of Chimp Chunk 2-10 Jerome M Molleston 5/4/2009
 Annotation of Chimp Chunk 2-10 Jerome M Molleston 5/4/2009 1 Abstract A stretch of chimpanzee DNA was annotated using tools including BLAST, BLAT, and Genscan. Analysis of Genscan predicted genes revealed
Annotation of Chimp Chunk 2-10 Jerome M Molleston 5/4/2009 1 Abstract A stretch of chimpanzee DNA was annotated using tools including BLAST, BLAT, and Genscan. Analysis of Genscan predicted genes revealed
Six Sigma Glossary Lean 6 Society
 Six Sigma Glossary Lean 6 Society ABSCISSA ACCEPTANCE REGION ALPHA RISK ALTERNATIVE HYPOTHESIS ASSIGNABLE CAUSE ASSIGNABLE VARIATIONS The horizontal axis of a graph The region of values for which the null
Six Sigma Glossary Lean 6 Society ABSCISSA ACCEPTANCE REGION ALPHA RISK ALTERNATIVE HYPOTHESIS ASSIGNABLE CAUSE ASSIGNABLE VARIATIONS The horizontal axis of a graph The region of values for which the null
Inter-country mixing in HIV transmission clusters: A pan-european phylodynamic study
 Inter-country mixing in HIV transmission clusters: A pan-european phylodynamic study Prabhav Kalaghatgi Max Planck Institute for Informatics March 20th 2013 HIV epidemic (2009) Prabhav Kalaghatgi 2/18
Inter-country mixing in HIV transmission clusters: A pan-european phylodynamic study Prabhav Kalaghatgi Max Planck Institute for Informatics March 20th 2013 HIV epidemic (2009) Prabhav Kalaghatgi 2/18
A new human immunodeficiency virus derived from gorillas
 A new human immunodeficiency virus derived from gorillas Jean-Christophe Plantier, Marie Leoz, Jonathan E Dickerson, Fabienne De Oliveira, François Cordonnier, Véronique Lemée, Florence Damond, David L
A new human immunodeficiency virus derived from gorillas Jean-Christophe Plantier, Marie Leoz, Jonathan E Dickerson, Fabienne De Oliveira, François Cordonnier, Véronique Lemée, Florence Damond, David L
AP BIOLOGY: READING ASSIGNMENT FOR CHAPTER 5
 1) Complete the following table: Class Monomer Functions Carbohydrates 1. 3. Lipids 1. 3. Proteins 1. 3. 4. 5. 6. Nucleic Acids 1. 2) Circle the atoms of these two glucose molecules that will be removed
1) Complete the following table: Class Monomer Functions Carbohydrates 1. 3. Lipids 1. 3. Proteins 1. 3. 4. 5. 6. Nucleic Acids 1. 2) Circle the atoms of these two glucose molecules that will be removed
Teaching Phylogeny and Direction of Viral Transmission using a Real HIV Criminal Case
 Tested Studies for Laboratory Teaching Proceedings of the Association for Biology Laboratory Education Volume 39, Article 24, 2018 Teaching Phylogeny and Direction of Viral Transmission using a Real HIV
Tested Studies for Laboratory Teaching Proceedings of the Association for Biology Laboratory Education Volume 39, Article 24, 2018 Teaching Phylogeny and Direction of Viral Transmission using a Real HIV
PROTOCOL FOR INFLUENZA A VIRUS GLOBAL SWINE H1 CLADE CLASSIFICATION
 PROTOCOL FOR INFLUENZA A VIRUS GLOBAL SWINE H1 CLADE CLASSIFICATION January 23, 2017 1. Background Swine H1 viruses have diversified into three major genetic lineages over time. Recently, Anderson et al.
PROTOCOL FOR INFLUENZA A VIRUS GLOBAL SWINE H1 CLADE CLASSIFICATION January 23, 2017 1. Background Swine H1 viruses have diversified into three major genetic lineages over time. Recently, Anderson et al.
6/10/2015. Background. Background. Background. Background. Methods
 /1/1 The challenges of diversity: HIV-1 subtype distribution and transmission s within the Australian Molecular Epidemiology Network-HIV -1 Castley A, Sawleshwarkar S, Varma R, Herring B, Thapa K, Chibo
/1/1 The challenges of diversity: HIV-1 subtype distribution and transmission s within the Australian Molecular Epidemiology Network-HIV -1 Castley A, Sawleshwarkar S, Varma R, Herring B, Thapa K, Chibo
PREDICTING THE EXPRESSION AND SOLUBILITY OF MEMBRANE PROTEINS
 PREDICTING THE EXPRESSION AND SOLUBILITY OF MEMBRANE PROTEINS Mark E. Dumont, Michael A. White, Kathy Clark, Elizabeth J. Grayhack, and Eric. M. Phizicky Unknowns in Membrane Protein Expression, and Solubilization
PREDICTING THE EXPRESSION AND SOLUBILITY OF MEMBRANE PROTEINS Mark E. Dumont, Michael A. White, Kathy Clark, Elizabeth J. Grayhack, and Eric. M. Phizicky Unknowns in Membrane Protein Expression, and Solubilization
Explore. sexcntry Sex according to country. [DataSet1] D:\NORA\NORA Main File.sav
![Explore. sexcntry Sex according to country. [DataSet1] D:\NORA\NORA Main File.sav Explore. sexcntry Sex according to country. [DataSet1] D:\NORA\NORA Main File.sav](/thumbs/82/84761267.jpg) EXAMINE VARIABLES=nc228 BY sexcntry /PLOT BOXPLOT HISTOGRAM NPPLOT /COMPARE GROUPS /STATISTICS DESCRIPTIVES /CINTERVAL 95 /MISSING LISTWISE /NOTOTAL. Explore Notes Output Created Comments Input Missing
EXAMINE VARIABLES=nc228 BY sexcntry /PLOT BOXPLOT HISTOGRAM NPPLOT /COMPARE GROUPS /STATISTICS DESCRIPTIVES /CINTERVAL 95 /MISSING LISTWISE /NOTOTAL. Explore Notes Output Created Comments Input Missing
STRUCTURAL CHROMOSOMAL ABERRATIONS
 STRUCTURAL CHROMOSOMAL ABERRATIONS Structural chromosomal aberrations cause structural abnormalities in chromosome structure. They alter the sequence or the kind of genes present in chromosome. These are
STRUCTURAL CHROMOSOMAL ABERRATIONS Structural chromosomal aberrations cause structural abnormalities in chromosome structure. They alter the sequence or the kind of genes present in chromosome. These are
Nature Genetics: doi: /ng Supplementary Figure 1. Rates of different mutation types in CRC.
 Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
Sex Comb Variation in Four Drosophila Species from North India
 International Journal of Biotechnology and Bioengineering Research. ISSN 2231-1238, Volume 4, Number 4 (2013), pp. 329-334 Research India Publications http://www.ripublication.com/ ijbbr.htm Sex Comb Variation
International Journal of Biotechnology and Bioengineering Research. ISSN 2231-1238, Volume 4, Number 4 (2013), pp. 329-334 Research India Publications http://www.ripublication.com/ ijbbr.htm Sex Comb Variation
Host Dependent Evolutionary Patterns and the Origin of 2009 H1N1 Pandemic Influenza
 Host Dependent Evolutionary Patterns and the Origin of 2009 H1N1 Pandemic Influenza The origin of H1N1pdm constitutes an unresolved mystery, as its most recently observed ancestors were isolated in pigs
Host Dependent Evolutionary Patterns and the Origin of 2009 H1N1 Pandemic Influenza The origin of H1N1pdm constitutes an unresolved mystery, as its most recently observed ancestors were isolated in pigs
Measurement Error 2: Scale Construction (Very Brief Overview) Page 1
 Measurement Error 2: Scale Construction (Very Brief Overview) Richard Williams, University of Notre Dame, https://www3.nd.edu/~rwilliam/ Last revised January 22, 2015 This handout draws heavily from Marija
Measurement Error 2: Scale Construction (Very Brief Overview) Richard Williams, University of Notre Dame, https://www3.nd.edu/~rwilliam/ Last revised January 22, 2015 This handout draws heavily from Marija
Molecular Evolution and the Neutral Theory
 Molecular Evolution and the Neutral Theory 1. Observation: DNA and amino-acid sequences evolve at roughly constant rates. 2. Model: The neutral theory explains why this might be expected. 3. Application:
Molecular Evolution and the Neutral Theory 1. Observation: DNA and amino-acid sequences evolve at roughly constant rates. 2. Model: The neutral theory explains why this might be expected. 3. Application:
Use of a camp BRET Sensor to Characterize a Novel Regulation of camp by the Sphingosine-1-phosphate/G 13 Pathway
 Use of a camp BRET Sensor to Characterize a Novel Regulation of camp by the Sphingosine-1-phosphate/G 13 Pathway SUPPLEMENTAL DATA Characterization of the CAMYEL sensor and calculation of intracellular
Use of a camp BRET Sensor to Characterize a Novel Regulation of camp by the Sphingosine-1-phosphate/G 13 Pathway SUPPLEMENTAL DATA Characterization of the CAMYEL sensor and calculation of intracellular
Sum of Neurally Distinct Stimulus- and Task-Related Components.
 SUPPLEMENTARY MATERIAL for Cardoso et al. 22 The Neuroimaging Signal is a Linear Sum of Neurally Distinct Stimulus- and Task-Related Components. : Appendix: Homogeneous Linear ( Null ) and Modified Linear
SUPPLEMENTARY MATERIAL for Cardoso et al. 22 The Neuroimaging Signal is a Linear Sum of Neurally Distinct Stimulus- and Task-Related Components. : Appendix: Homogeneous Linear ( Null ) and Modified Linear
UvA-DARE (Digital Academic Repository)
 UvA-DARE (Digital Academic Repository) Superinfection with drug-resistant HIV is rare and does not contribute substantially to therapy failure in a large European cohort Bartha, I.; Assel, M.; Sloot, P.M.A.;
UvA-DARE (Digital Academic Repository) Superinfection with drug-resistant HIV is rare and does not contribute substantially to therapy failure in a large European cohort Bartha, I.; Assel, M.; Sloot, P.M.A.;
Copyright 2007 IEEE. Reprinted from 4th IEEE International Symposium on Biomedical Imaging: From Nano to Macro, April 2007.
 Copyright 27 IEEE. Reprinted from 4th IEEE International Symposium on Biomedical Imaging: From Nano to Macro, April 27. This material is posted here with permission of the IEEE. Such permission of the
Copyright 27 IEEE. Reprinted from 4th IEEE International Symposium on Biomedical Imaging: From Nano to Macro, April 27. This material is posted here with permission of the IEEE. Such permission of the
Ras and Cell Signaling Exercise
 Ras and Cell Signaling Exercise Learning Objectives In this exercise, you will use, a protein 3D- viewer, to explore: the structure of the Ras protein the active and inactive state of Ras and the amino
Ras and Cell Signaling Exercise Learning Objectives In this exercise, you will use, a protein 3D- viewer, to explore: the structure of the Ras protein the active and inactive state of Ras and the amino
White Paper Estimating Complex Phenotype Prevalence Using Predictive Models
 White Paper 23-12 Estimating Complex Phenotype Prevalence Using Predictive Models Authors: Nicholas A. Furlotte Aaron Kleinman Robin Smith David Hinds Created: September 25 th, 2015 September 25th, 2015
White Paper 23-12 Estimating Complex Phenotype Prevalence Using Predictive Models Authors: Nicholas A. Furlotte Aaron Kleinman Robin Smith David Hinds Created: September 25 th, 2015 September 25th, 2015
aV. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
Comparison of discrimination methods for the classification of tumors using gene expression data
 Comparison of discrimination methods for the classification of tumors using gene expression data Sandrine Dudoit, Jane Fridlyand 2 and Terry Speed 2,. Mathematical Sciences Research Institute, Berkeley
Comparison of discrimination methods for the classification of tumors using gene expression data Sandrine Dudoit, Jane Fridlyand 2 and Terry Speed 2,. Mathematical Sciences Research Institute, Berkeley
