Minimalistic encapsulated proteomic sample processing applied to copy number
|
|
|
- Camilla Rogers
- 5 years ago
- Views:
Transcription
1 Nature Methods Minimalistic encapsulated proteomic sample processing applied to copy number estimation in eukaryotic cells Nils A. Kulak, Garwin Pichler, Igor Paron, Nagarjuna Nagaraj and Matthias Mann Supplementary Figure 1 Comparison of SILAC-based ratios using different sample processing conditions, Supplementary Figure 2 Comparison of SILAC-based ratios of HeLa proteins using various StageTip materials. Supplementary Figure 3 Comparison of SILAC-based ratios of HeLa proteins using non-activated SDB-RPS vs. activated SDB-RPS. Supplementary Figure 4 Loading capacity test of StageTip materials. Supplementary Figure 5 Comparison of StageTip based SAX, SCX, and SDB-RPS fractionation techniques. Supplementary Figure 6 Distribution of estimated copy numbers for S. pombe. Supplementary Figure 7 S. pombe correlation of estimated copy numbers using 6-fraction ist-scx analysis to copy numbers reported in Marguerat et al. using 34 synthetic peptide standards (R 2 = 0.89). Supplementary Figure 8 Distribution of median estimated copy numbers per cell for HeLa (21,057), S. pombe (5,137) and S. cerevisiae (822). Supplementary Figure 9 2D annotation enrichment score for GO terms based on estimated copy numbers between orthologs. Supplementary Figure 10 Multi-scatter plot of label-free quantification intensities depicting the reproducibility across four technical replicates of 6-fraction ist-scx analysis of HeLa proteome. Supplementary Figure 11 Protein copy number distribution with respect to individual copy number bins. Supplementary Table 1 Protein identifications and protein copy number estimations of S.cerevisiae, S.pombe and HeLa cells. Supplementary Table 2 Orthologs of S.cerevisiae, S.pombe and HeLa cells and their protein copy number estimations. Supplementary Table 3 Buffer compositions and materials for the ist method. Supplementary Note Supplementary Video 1 ist sample preparation video tutorial. The video tutorial shows all steps of the ist sample preparation workflow. It further shows how to troubleshoot the method in case of need.
2 Supplementary Figure 1 Comparison of SILAC-based ratios using different sample processing conditions. (a) SILAC-based protein ratios of clarified vs. non-clarified HeLa cell lysates prior to proteolytic digestion. (b) SILAC-based protein ratios of FASP vs. ist HeLa cell lysates. (c) SILAC-based protein ratios of StageTip vs. tube-processed HeLa cell lysates. (d) Comparison of the ist-sdc method to a published SDC based in-solution protocol. Shown are median unique peptide identifications of technical triplicates ± s.e.m.
3 Supplementary Figure 2 Comparison of SILAC-based ratios of HeLa proteins using various StageTip materials. (a) SILAC-based ratios of SDB-XC vs. C 18. (b) SILAC-based ratios of SDB-RPS vs. C 18. (c) SILACbased ratios of SCX vs. C 18. Single-shot measurements were performed. Supplementary Figure 3 Comparison of SILAC-based ratios of HeLa proteins using non-activated SDB- RPS vs. activated SDB-RPS. Single-shot measurements were performed.
4 Supplementary Figure 4 Loading capacity test of StageTip materials. Respective quantities of peptides were loaded onto 16-gauge double-plug StageTips. The eluates were dried and re-suspended in corresponding volumes (10 µl, 15 µl, 20 µl, 25 µl Buffer A*: 2% (v/v) ACN, 0.1% TFA (v/v)). 1 µl of the peptides were measured on a 2h gradient. Shown are technical duplicates ± s.d. The data indicate that loadings of up to 20 µg (up to 10 µg per plug) have little effect on peptide identifications using SDB-RPS material.
5 Supplementary Figure 5 Comparison of StageTip based SAX, SCX, and SDB-RPS fractionation techniques. (a) Rounded percent of unique peptides in each fraction. (b) Rounded percent of exclusive peptides in each fraction compared to other fractions. (c) Rounded percent of fractionation efficiencies.
6 Supplementary Figure 6 Distribution of estimated copy numbers for S. pombe. Blue bins represent proteins that are identified uniquely in the 6-fraction ist-scx analysis in comparison to Gunaratne et al 32. Supplementary Figure 7 S. pombe correlation of estimated copy numbers using 6-fraction ist-scx analysis to copy numbers reported in Marguerat et al 27. using 34 synthetic peptide standards (R 2 = 0.89).
7 Supplementary Figure 8 Distribution of median estimated copy numbers per cell for HeLa (21,057), S. pombe (5,137) and S. cerevisiae (822). Box plots show the median (stripe), the 25 th to 75 th percentile (interquartile range, box), 1.5x the interquartile range (1.5 IQR, whiskers), and outliers (open circles). Supplementary Figure 9 2D annotation enrichment score for GO terms based on estimated copy numbers between orthologs. (a) For S. pombe vs. S. cerevisiae, (b) for HeLa vs. S. cerevisiae and (c) HeLa vs. S. pombe. Representative GO terms showing anti-correlating behavior are depicted in red.
8 Supplementary Figure 10 Multi-scatter plot of label-free quantification intensities depicting the reproducibility across four technical replicates of 6-fraction ist-scx analysis of HeLa proteome.
9 Supplementary Figure 11 Protein copy number distribution with respect to individual copy number bins. The bar plot represents the percentage of proteins present in the copy number bins (blue), neighbours with copy numbers within a factor 10 (grey, weighted average = 40%) or 100 (brown, weighted average = 64%).
10 Supplementary Note Interpretation of SILAC comparison plots The different conditions that were tested for the optimization of the sample preparation were carried out by using SILAC label swapping experiments (Supplementary Fig. 1-3). These plots show a cloud around the origin if there is no significant effect of the variable. However when the variable exerts a significant effect on the proteome detected, there will be a cloud along the diagonal line with slope = -1 as the ratios are inverted in a forward and reverse label swap experiment. Excluded fraction in density plots The color schemes of the density plots enable us to slice the data points in a discrete manner. We used excluded fraction based coloring. For example in figure 3c, the data points in black exclude 5% of the total data points and the point in black and grey exclude 25% of the total data points. Calculation of neighboring proteins For any protein, to calculate the percentage of total proteins that are within a factor 10 in copy number of each other, the proteins were divided into bins with each bin spanning an order of magnitude. The relative contribution in terms of number of proteins per bin was then calculated. The percentage of total proteins that are within one order of magnitude from a selected bin was determined by adding the relative contributions from the bins one order of magnitude above and below. The weighted average of this value from all the bins yields the percentage of proteins within one order of magnitude for any intensity bin. For the calculation of proteins within two orders of magnitude, two continuous bins above and below the bin of interest were added up.
Universal sample preparation method for proteome analysis
 nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
Nature Structural & Molecular Biology: doi: /nsmb.3218
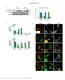 Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Supplementary Figure 1. Nature Neuroscience: doi: /nn.4547
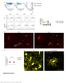 Supplementary Figure 1 Characterization of the Microfetti mouse model. (a) Gating strategy for 8-color flow analysis of peripheral Ly-6C + monocytes from Microfetti mice 5-7 days after TAM treatment. Living
Supplementary Figure 1 Characterization of the Microfetti mouse model. (a) Gating strategy for 8-color flow analysis of peripheral Ly-6C + monocytes from Microfetti mice 5-7 days after TAM treatment. Living
Molecular Cell, Volume 46. Supplemental Information
 Molecular Cell, Volume 46 Supplemental Information Mapping N-Glycosylation Sites across Seven Evolutionary Distant Species Reveals a Divergent Substrate Proteome Despite a Common Core Machinery Dorota
Molecular Cell, Volume 46 Supplemental Information Mapping N-Glycosylation Sites across Seven Evolutionary Distant Species Reveals a Divergent Substrate Proteome Despite a Common Core Machinery Dorota
Nature Methods: doi: /nmeth.3177
 Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Understandable Statistics
 Understandable Statistics correlated to the Advanced Placement Program Course Description for Statistics Prepared for Alabama CC2 6/2003 2003 Understandable Statistics 2003 correlated to the Advanced Placement
Understandable Statistics correlated to the Advanced Placement Program Course Description for Statistics Prepared for Alabama CC2 6/2003 2003 Understandable Statistics 2003 correlated to the Advanced Placement
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Experimental design and workflow utilized to generate the WMG Protein Atlas.
 Supplementary Figure 1 Experimental design and workflow utilized to generate the WMG Protein Atlas. (a) Illustration of the plant organs and nodule infection time points analyzed. (b) Proteomic workflow
Supplementary Figure 1 Experimental design and workflow utilized to generate the WMG Protein Atlas. (a) Illustration of the plant organs and nodule infection time points analyzed. (b) Proteomic workflow
Supporting Information MALDI versus ESI the impact of the ion source on peptide identification
 Supporting Information MALDI versus ESI the impact of the ion source on peptide identification Wiebke Maria Nadler, 1 Dietmar Waidelich, 2 Alexander Kerner, 1 Sabrina Hanke, 1 Regina Berg, 3 Andreas Trumpp
Supporting Information MALDI versus ESI the impact of the ion source on peptide identification Wiebke Maria Nadler, 1 Dietmar Waidelich, 2 Alexander Kerner, 1 Sabrina Hanke, 1 Regina Berg, 3 Andreas Trumpp
APPLICATION NOTE. Highly reproducible and Comprehensive Proteome Profiling of Formalin-Fixed Paraffin-Embedded (FFPE) Tissues Slices
 APPLICATION NOTE Highly reproducible and Comprehensive Proteome Profiling of Formalin-Fixed Paraffin-Embedded (FFPE) Tissues Slices INTRODUCTION Preservation of tissue biopsies is a critical step to This
APPLICATION NOTE Highly reproducible and Comprehensive Proteome Profiling of Formalin-Fixed Paraffin-Embedded (FFPE) Tissues Slices INTRODUCTION Preservation of tissue biopsies is a critical step to This
Supplementary figure 1: LII/III GIN-cells show morphological characteristics of MC
 1 2 1 3 Supplementary figure 1: LII/III GIN-cells show morphological characteristics of MC 4 5 6 7 (a) Reconstructions of LII/III GIN-cells with somato-dendritic compartments in orange and axonal arborizations
1 2 1 3 Supplementary figure 1: LII/III GIN-cells show morphological characteristics of MC 4 5 6 7 (a) Reconstructions of LII/III GIN-cells with somato-dendritic compartments in orange and axonal arborizations
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for
 Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for Proteomics Studies Zhen Wu, Jichang Huang, Jingnan Huang, Qingqing Li, Xumin Zhang *, State Key Laboratory of Genetic Engineering, Department
Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for Proteomics Studies Zhen Wu, Jichang Huang, Jingnan Huang, Qingqing Li, Xumin Zhang *, State Key Laboratory of Genetic Engineering, Department
 a RT:. - 1.1 1 95 9 85 8 27.64 27.87 28.1 27.42 28.24 38.98 Low ph RPLC NL: 2.29E9 TIC MS TrypticBSA_Dig est_replicate1_ 2Mar18_Preci ous_18-1-6 75 7 65 48.75 65.8 Relative Abundance 6 55 5 45 28.54 48.98
a RT:. - 1.1 1 95 9 85 8 27.64 27.87 28.1 27.42 28.24 38.98 Low ph RPLC NL: 2.29E9 TIC MS TrypticBSA_Dig est_replicate1_ 2Mar18_Preci ous_18-1-6 75 7 65 48.75 65.8 Relative Abundance 6 55 5 45 28.54 48.98
Supplementary Figure 1. Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
 Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary material: Materials and suppliers
 Supplementary material: Materials and suppliers Electrophoresis consumables including tris-glycine, acrylamide, SDS buffer and Coomassie Brilliant Blue G-2 dye (CBB) were purchased from Ameresco (Solon,
Supplementary material: Materials and suppliers Electrophoresis consumables including tris-glycine, acrylamide, SDS buffer and Coomassie Brilliant Blue G-2 dye (CBB) were purchased from Ameresco (Solon,
Supporting Information. Lysine Propionylation to Boost Proteome Sequence. Coverage and Enable a Silent SILAC Strategy for
 Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Supplementary Figure 1
 Supplementary Figure 1 6 HE-50 HE-116 E-1 HE-108 Supplementary Figure 1. Targeted drug response curves of endometrial cancer cells. Endometrial cancer cell lines were incubated with serial dilutions of
Supplementary Figure 1 6 HE-50 HE-116 E-1 HE-108 Supplementary Figure 1. Targeted drug response curves of endometrial cancer cells. Endometrial cancer cell lines were incubated with serial dilutions of
Protein and peptide separations,
 A Quarterly Technical Newsletter Spring Grace Vydac: Leading the Way in Separations for Proteomics Protein and peptide separations, always important to protein chemists and enzymologists, have recently
A Quarterly Technical Newsletter Spring Grace Vydac: Leading the Way in Separations for Proteomics Protein and peptide separations, always important to protein chemists and enzymologists, have recently
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded
 Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Edge detection. Gradient-based edge operators
 Edge detection Gradient-based edge operators Prewitt Sobel Roberts Laplacian zero-crossings Canny edge detector Hough transform for detection of straight lines Circle Hough Transform Digital Image Processing:
Edge detection Gradient-based edge operators Prewitt Sobel Roberts Laplacian zero-crossings Canny edge detector Hough transform for detection of straight lines Circle Hough Transform Digital Image Processing:
Nature Medicine: doi: /nm.4078
 Supplementary Figure 1. Cetuximab induces ER stress response in DiFi cells. (a) Scheme of SILAC proteome. (b) MS-base read out of SILAC experiment. The histogram of log 2 -transformed normalized H/L ratios
Supplementary Figure 1. Cetuximab induces ER stress response in DiFi cells. (a) Scheme of SILAC proteome. (b) MS-base read out of SILAC experiment. The histogram of log 2 -transformed normalized H/L ratios
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2
 Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
7. Bivariate Graphing
 1 7. Bivariate Graphing Video Link: https://www.youtube.com/watch?v=shzvkwwyguk&index=7&list=pl2fqhgedk7yyl1w9tgio8w pyftdumgc_j Section 7.1: Converting a Quantitative Explanatory Variable to Categorical
1 7. Bivariate Graphing Video Link: https://www.youtube.com/watch?v=shzvkwwyguk&index=7&list=pl2fqhgedk7yyl1w9tgio8w pyftdumgc_j Section 7.1: Converting a Quantitative Explanatory Variable to Categorical
Supplementary. properties of. network types. randomly sampled. subsets (75%
 Supplementary Information Gene co-expression network analysis reveals common system-level prognostic genes across cancer types properties of Supplementary Figure 1 The robustness and overlap of prognostic
Supplementary Information Gene co-expression network analysis reveals common system-level prognostic genes across cancer types properties of Supplementary Figure 1 The robustness and overlap of prognostic
Nature Genetics: doi: /ng Supplementary Figure 1. Phenotypic characterization of MES- and ADRN-type cells.
 Supplementary Figure 1 Phenotypic characterization of MES- and ADRN-type cells. (a) Bright-field images showing cellular morphology of MES-type (691-MES, 700-MES, 717-MES) and ADRN-type (691-ADRN, 700-
Supplementary Figure 1 Phenotypic characterization of MES- and ADRN-type cells. (a) Bright-field images showing cellular morphology of MES-type (691-MES, 700-MES, 717-MES) and ADRN-type (691-ADRN, 700-
COPD lungs show an attached stratified mucus layer that separate. bacteria from the epithelial cells resembling the protective colonic
 COPD lungs show an attached stratified mucus layer that separate bacteria from the epithelial cells resembling the protective colonic mucus SUPPLEMENTARY TABLES AND FIGURES Tables S1 S8, page 1 and separate
COPD lungs show an attached stratified mucus layer that separate bacteria from the epithelial cells resembling the protective colonic mucus SUPPLEMENTARY TABLES AND FIGURES Tables S1 S8, page 1 and separate
A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis
 APPLICATION NOTE Cell-Free DNA Isolation Kit A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis Abstract Circulating cell-free DNA (cfdna) has been shown
APPLICATION NOTE Cell-Free DNA Isolation Kit A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis Abstract Circulating cell-free DNA (cfdna) has been shown
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood
 Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
SCATTER PLOTS AND TREND LINES
 1 SCATTER PLOTS AND TREND LINES LEARNING MAP INFORMATION STANDARDS 8.SP.1 Construct and interpret scatter s for measurement to investigate patterns of between two quantities. Describe patterns such as
1 SCATTER PLOTS AND TREND LINES LEARNING MAP INFORMATION STANDARDS 8.SP.1 Construct and interpret scatter s for measurement to investigate patterns of between two quantities. Describe patterns such as
Shotgun Proteomics MS/MS. Protein Mixture. proteolysis. Peptide Mixture. Time. Abundance. Abundance. m/z. Abundance. m/z 2. Abundance.
 Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
Digitizing the Proteomes From Big Tissue Biobanks
 Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Total Histone H3 Acetylation Detection Fast Kit (Colorimetric)
 Total Histone H3 Acetylation Detection Fast Kit (Colorimetric) Catalog Number KA1538 48 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use...
Total Histone H3 Acetylation Detection Fast Kit (Colorimetric) Catalog Number KA1538 48 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use...
Q: How do I get the protein concentration in mg/ml from the standard curve if the X-axis is in units of µg.
 Photometry Frequently Asked Questions Q: How do I get the protein concentration in mg/ml from the standard curve if the X-axis is in units of µg. Protein standard curves are traditionally presented as
Photometry Frequently Asked Questions Q: How do I get the protein concentration in mg/ml from the standard curve if the X-axis is in units of µg. Protein standard curves are traditionally presented as
Nature Neuroscience: doi: /nn Supplementary Figure 1. Behavioral training.
 Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplemental Information. 3D-CLEM Reveals that a Major Portion. of Mitotic Chromosomes Is Not Chromatin
 Molecular Cell, Volume 64 Supplemental Information 3D-CLEM Reveals that a Major Portion of Mitotic Chromosomes Is Not Chromatin Daniel G. Booth, Alison J. Beckett, Oscar Molina, Itaru Samejima, Hiroshi
Molecular Cell, Volume 64 Supplemental Information 3D-CLEM Reveals that a Major Portion of Mitotic Chromosomes Is Not Chromatin Daniel G. Booth, Alison J. Beckett, Oscar Molina, Itaru Samejima, Hiroshi
EpiQuik Total Histone H3 Acetylation Detection Fast Kit (Colorimetric)
 EpiQuik Total Histone H3 Acetylation Detection Fast Kit (Colorimetric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Total Histone H3 Acetylation Detection Fast Kit (Colorimetric)
EpiQuik Total Histone H3 Acetylation Detection Fast Kit (Colorimetric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Total Histone H3 Acetylation Detection Fast Kit (Colorimetric)
Quantification with Proteome Discoverer. Bernard Delanghe
 Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
Up to now, gel electrophoresis remains the most powerful
 pubs.acs.org/ac Polyacrylamide Gel with Switchable Trypsin Activity for Analysis of Proteins Fangjie Liu,, Mingliang Ye,*, Chunli Wang,, Zhengyan Hu,, Yi Zhang,, Hongqiang Qin,, Kai Cheng,, and Hanfa Zou*,
pubs.acs.org/ac Polyacrylamide Gel with Switchable Trypsin Activity for Analysis of Proteins Fangjie Liu,, Mingliang Ye,*, Chunli Wang,, Zhengyan Hu,, Yi Zhang,, Hongqiang Qin,, Kai Cheng,, and Hanfa Zou*,
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Effects of UBL5 knockdown on cell cycle distribution and sister chromatid cohesion
 Supplementary Figure S1. Effects of UBL5 knockdown on cell cycle distribution and sister chromatid cohesion A. Representative examples of flow cytometry profiles of HeLa cells transfected with indicated
Supplementary Figure S1. Effects of UBL5 knockdown on cell cycle distribution and sister chromatid cohesion A. Representative examples of flow cytometry profiles of HeLa cells transfected with indicated
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 The timeline of the NGAG method for extraction of N-linked glycans and glycosite-containing peptides. The timeline can be changed based on the number of samples. Supplementary Figure
Supplementary Figure 1 The timeline of the NGAG method for extraction of N-linked glycans and glycosite-containing peptides. The timeline can be changed based on the number of samples. Supplementary Figure
SUPPLEMENTARY INFORMATION
 Supplemental Figure 1. Furin is efficiently deleted in CD4 + and CD8 + T cells. a, Western blot for furin and actin proteins in CD4cre-fur f/f and fur f/f Th1 cells. Wild-type and furin-deficient CD4 +
Supplemental Figure 1. Furin is efficiently deleted in CD4 + and CD8 + T cells. a, Western blot for furin and actin proteins in CD4cre-fur f/f and fur f/f Th1 cells. Wild-type and furin-deficient CD4 +
Expanded View Figures
 EMO Molecular Medicine Proteomic map of squamous cell carcinomas Hanibal ohnenberger et al Expanded View Figures Figure EV1. Technical reproducibility. Pearson s correlation analysis of normalised SILC
EMO Molecular Medicine Proteomic map of squamous cell carcinomas Hanibal ohnenberger et al Expanded View Figures Figure EV1. Technical reproducibility. Pearson s correlation analysis of normalised SILC
Fructose-6-Phosphate Colorimetric Assay Kit
 Fructose-6-Phosphate Colorimetric Assay Kit Catalog No. KM0060 Detection and Quantification of Fructose-6-Phosphate Concentrations in Biological Samples. Research Purposes Only. Not Intended for Diagnostic
Fructose-6-Phosphate Colorimetric Assay Kit Catalog No. KM0060 Detection and Quantification of Fructose-6-Phosphate Concentrations in Biological Samples. Research Purposes Only. Not Intended for Diagnostic
Figure S1. PMVs from THP-1 cells expose phosphatidylserine and carry actin. A) Flow
 SUPPLEMENTARY DATA Supplementary Figure Legends Figure S1. PMVs from THP-1 cells expose phosphatidylserine and carry actin. A) Flow cytometry analysis of PMVs labelled with annexin-v-pe (Guava technologies)
SUPPLEMENTARY DATA Supplementary Figure Legends Figure S1. PMVs from THP-1 cells expose phosphatidylserine and carry actin. A) Flow cytometry analysis of PMVs labelled with annexin-v-pe (Guava technologies)
Insulin SEC analysis. Insulin Samples:
 EE1 Insulin SEC analysis Insulin Samples: Lispro, identical in structure to Insulin Human, except that it has lysine and proline at positions 8 and 9, respectively, of chain B, whereas this sequence is
EE1 Insulin SEC analysis Insulin Samples: Lispro, identical in structure to Insulin Human, except that it has lysine and proline at positions 8 and 9, respectively, of chain B, whereas this sequence is
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
Introduction to Statistical Data Analysis I
 Introduction to Statistical Data Analysis I JULY 2011 Afsaneh Yazdani Preface What is Statistics? Preface What is Statistics? Science of: designing studies or experiments, collecting data Summarizing/modeling/analyzing
Introduction to Statistical Data Analysis I JULY 2011 Afsaneh Yazdani Preface What is Statistics? Preface What is Statistics? Science of: designing studies or experiments, collecting data Summarizing/modeling/analyzing
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose.
 Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose. (a) Relative levels of both new acetylation and all acetylation
Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose. (a) Relative levels of both new acetylation and all acetylation
Kidney. Heart. Lung. Sirt1. Gapdh. Mouse IgG DAPI. Rabbit IgG DAPI
 a e Na V 1.5 Ad-LacZ Ad- 110KD b Scn5a/ (relative to Ad-LacZ) f 150 100 50 0 p = 0.65 Ad-LacZ Ad- c Heart Lung Kidney Spleen 110KD d fl/fl c -/- DAPI 20 µm Na v 1.5 250KD fl/fl Rabbit IgG DAPI fl/fl Mouse
a e Na V 1.5 Ad-LacZ Ad- 110KD b Scn5a/ (relative to Ad-LacZ) f 150 100 50 0 p = 0.65 Ad-LacZ Ad- c Heart Lung Kidney Spleen 110KD d fl/fl c -/- DAPI 20 µm Na v 1.5 250KD fl/fl Rabbit IgG DAPI fl/fl Mouse
Phosphotyrosine biased enrichment of tryptic peptides from cancer cells by combining py-mip and TiO 2 affinity resins
 Phosphotyrosine biased enrichment of tryptic peptides from cancer cells by combining py-mip and TiO 2 affinity resins Loreta Bllaci, Silje B. Torsetnes,, Celina Wierzbicka, Sudhirkumar Shinde, Börje Sellergren,
Phosphotyrosine biased enrichment of tryptic peptides from cancer cells by combining py-mip and TiO 2 affinity resins Loreta Bllaci, Silje B. Torsetnes,, Celina Wierzbicka, Sudhirkumar Shinde, Börje Sellergren,
Frequency distributions
 Applied Biostatistics distributions Martin Bland Professor of Health Statistics University of York http://www-users.york.ac.uk/~mb55/ Types of data Qualitative data arise when individuals may fall into
Applied Biostatistics distributions Martin Bland Professor of Health Statistics University of York http://www-users.york.ac.uk/~mb55/ Types of data Qualitative data arise when individuals may fall into
Human Neurology 3-Plex A
 Human Neurology 3-Plex A SUMMARY AND EXPLANATION OF THE TEST The Human N3PA assay is a digital immunoassay for the quantitative determination of total Tau, Aβ42, and Aβ40 in human plasma and CSF. Determination
Human Neurology 3-Plex A SUMMARY AND EXPLANATION OF THE TEST The Human N3PA assay is a digital immunoassay for the quantitative determination of total Tau, Aβ42, and Aβ40 in human plasma and CSF. Determination
Incomplete immune response to Coxsackie B viruses. associates with early autoimmunity against insulin
 Incomplete immune response to Coxsackie B viruses associates with early autoimmunity against insulin Michelle P. Ashton 1, Anne Eugster 1, Denise Walther 1, Natalie Daehling 1, Stephanie Riethausen 2,
Incomplete immune response to Coxsackie B viruses associates with early autoimmunity against insulin Michelle P. Ashton 1, Anne Eugster 1, Denise Walther 1, Natalie Daehling 1, Stephanie Riethausen 2,
Nature Methods: doi: /nmeth.4257
 Supplementary Figure 1 Screen for polypeptides that affect cellular actin filaments. (a) Table summarizing results from all polypeptides tested. Source shows organism, gene, and amino acid numbers used.
Supplementary Figure 1 Screen for polypeptides that affect cellular actin filaments. (a) Table summarizing results from all polypeptides tested. Source shows organism, gene, and amino acid numbers used.
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Pepsin Microplate Assay Kit User Manual
 Pepsin Microplate Assay Kit User Manual Catalog # CAK1081 Detection and Quantification of Pepsin Activity in Urine, Serum, Plasma, Tissue extracts, Cell lysate, Cell culture media and Other biological
Pepsin Microplate Assay Kit User Manual Catalog # CAK1081 Detection and Quantification of Pepsin Activity in Urine, Serum, Plasma, Tissue extracts, Cell lysate, Cell culture media and Other biological
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow
 Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
2.75: 84% 2.5: 80% 2.25: 78% 2: 74% 1.75: 70% 1.5: 66% 1.25: 64% 1.0: 60% 0.5: 50% 0.25: 25% 0: 0%
 Capstone Test (will consist of FOUR quizzes and the FINAL test grade will be an average of the four quizzes). Capstone #1: Review of Chapters 1-3 Capstone #2: Review of Chapter 4 Capstone #3: Review of
Capstone Test (will consist of FOUR quizzes and the FINAL test grade will be an average of the four quizzes). Capstone #1: Review of Chapters 1-3 Capstone #2: Review of Chapter 4 Capstone #3: Review of
Folin Ciocalteau Phenolic Content Quantification Assay Kit KB tests (96 well plate)
 Folin Ciocalteau Phenolic Content Quantification Assay Kit KB-03-006 400 tests (96 well plate) Index Introduction Pag. 1 Materials Pag. 2 Assay Principle Pag. 3 Assay protocol Pag. 4 Data analysis Pag.
Folin Ciocalteau Phenolic Content Quantification Assay Kit KB-03-006 400 tests (96 well plate) Index Introduction Pag. 1 Materials Pag. 2 Assay Principle Pag. 3 Assay protocol Pag. 4 Data analysis Pag.
Mouse C-peptide EIA. Cat. No. YII-YK013-EX FOR LABORATORY USE ONLY
 Mouse C-peptide EIA Cat. No. YII-YK013-EX FOR LABORATORY USE ONLY TOYO 2CHOME, KOTO-KU, TOKYO, 135-0016, JAPAN http://www.cosmobio.co.jp e-mail : export@cosmobio.co.jp Phone : +81-3-5632-9617 FAX : +81-3-5632-9618
Mouse C-peptide EIA Cat. No. YII-YK013-EX FOR LABORATORY USE ONLY TOYO 2CHOME, KOTO-KU, TOKYO, 135-0016, JAPAN http://www.cosmobio.co.jp e-mail : export@cosmobio.co.jp Phone : +81-3-5632-9617 FAX : +81-3-5632-9618
Summary of behavioral performances for mice in imaging experiments.
 Supplementary Figure 1 Summary of behavioral performances for mice in imaging experiments. (a) Task performance for mice during M2 imaging experiments. Open triangles, individual experiments. Filled triangles,
Supplementary Figure 1 Summary of behavioral performances for mice in imaging experiments. (a) Task performance for mice during M2 imaging experiments. Open triangles, individual experiments. Filled triangles,
Supplementary Figures and Notes for Digestion and depletion of abundant
 Supplementary Figures and Notes for Digestion and depletion of abundant proteins improves proteomic coverage Bryan R. Fonslow, Benjamin D. Stein, Kristofor J. Webb, Tao Xu, Jeong Choi, Sung Kyu Park, and
Supplementary Figures and Notes for Digestion and depletion of abundant proteins improves proteomic coverage Bryan R. Fonslow, Benjamin D. Stein, Kristofor J. Webb, Tao Xu, Jeong Choi, Sung Kyu Park, and
Supplementary Materials
 Supplementary Materials Supplementary Figure S1 Regulation of Ubl4A stability by its assembly partner A, The translation rate of Ubl4A is not affected in the absence of Bag6. Control, Bag6 and Ubl4A CRISPR
Supplementary Materials Supplementary Figure S1 Regulation of Ubl4A stability by its assembly partner A, The translation rate of Ubl4A is not affected in the absence of Bag6. Control, Bag6 and Ubl4A CRISPR
EPIGENTEK. EpiQuik Global Acetyl Histone H3K27 Quantification Kit (Colorimetric) Base Catalog # P-4059 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Global Acetyl Histone H3K27 Quantification Kit (Colorimetric) Base Catalog # P-4059 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Global Acetyl Histone H3K27 Quantification Kit (Colorimetric)
EpiQuik Global Acetyl Histone H3K27 Quantification Kit (Colorimetric) Base Catalog # P-4059 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Global Acetyl Histone H3K27 Quantification Kit (Colorimetric)
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
ab Exosome Isolation and Analysis Kit - Flow Cytometry, Cell Culture (CD63 / CD81)
 Version 1 Last updated 26 September 2018 ab239682 Exosome Isolation and Analysis Kit - Flow Cytometry, Cell Culture (CD63 / For the isolation and analysis of exosome from cell culture. This product is
Version 1 Last updated 26 September 2018 ab239682 Exosome Isolation and Analysis Kit - Flow Cytometry, Cell Culture (CD63 / For the isolation and analysis of exosome from cell culture. This product is
Online Appendix Material and Methods: Pancreatic RNA isolation and quantitative real-time (q)rt-pcr. Mice were fasted overnight and killed 1 hour (h)
 Online Appendix Material and Methods: Pancreatic RNA isolation and quantitative real-time (q)rt-pcr. Mice were fasted overnight and killed 1 hour (h) after feeding. A small slice (~5-1 mm 3 ) was taken
Online Appendix Material and Methods: Pancreatic RNA isolation and quantitative real-time (q)rt-pcr. Mice were fasted overnight and killed 1 hour (h) after feeding. A small slice (~5-1 mm 3 ) was taken
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Aggregated neutrophil extracellular traps limit inflammation by degrading cytokines and chemokines
 CORRECTION NOTICE Nat. Med. doi:10.1038/nm.3547; corrected online 25 August 2014 Aggregated neutrophil extracellular traps limit inflammation by degrading cytokines and chemokines Christine Schauer, Christina
CORRECTION NOTICE Nat. Med. doi:10.1038/nm.3547; corrected online 25 August 2014 Aggregated neutrophil extracellular traps limit inflammation by degrading cytokines and chemokines Christine Schauer, Christina
Supplementary Material for
 Supplementary Material for Selective neuronal lapses precede human cognitive lapses following sleep deprivation Supplementary Table 1. Data acquisition details Session Patient Brain regions monitored Time
Supplementary Material for Selective neuronal lapses precede human cognitive lapses following sleep deprivation Supplementary Table 1. Data acquisition details Session Patient Brain regions monitored Time
Type of file: PDF Title of file for HTML: Supplementary Information Description: Supplementary Figures
 Type of file: PDF Title of file for HTML: Supplementary Information Description: Supplementary Figures Type of file: MOV Title of file for HTML: Supplementary Movie 1 Description: NLRP3 is moving along
Type of file: PDF Title of file for HTML: Supplementary Information Description: Supplementary Figures Type of file: MOV Title of file for HTML: Supplementary Movie 1 Description: NLRP3 is moving along
File name: Supplementary Information Description: Supplementary Figures, Supplementary Table and Supplementary References
 File name: Supplementary Information Description: Supplementary Figures, Supplementary Table and Supplementary References File name: Supplementary Data 1 Description: Summary datasheets showing the spatial
File name: Supplementary Information Description: Supplementary Figures, Supplementary Table and Supplementary References File name: Supplementary Data 1 Description: Summary datasheets showing the spatial
Th17 and Th17/Treg ratio at early HIV infection associate with protective HIV-specific CD8 + T-cell responses and disease progression
 Th17 and Th17/Treg ratio at early HIV infection associate with protective HIV-specific CD8 T-cell responses and disease progression Juliana Falivene 1, Yanina Ghiglione 1, Natalia Laufer 1,3, María Eugenia
Th17 and Th17/Treg ratio at early HIV infection associate with protective HIV-specific CD8 T-cell responses and disease progression Juliana Falivene 1, Yanina Ghiglione 1, Natalia Laufer 1,3, María Eugenia
Cohort 2. Age, years 41.0 (10.2) Diabetes duration, years 26.5 (15.8)
 Supplementary Table. Participant characteristics in cohort Cohort Number Sex, Male Female Age, years. (.) Diabetes duration, years.5 (5.) BMI, kg.m. (3.) HbA c, % mmol.mol. (.7) () C-peptide, nmol.l.3
Supplementary Table. Participant characteristics in cohort Cohort Number Sex, Male Female Age, years. (.) Diabetes duration, years.5 (5.) BMI, kg.m. (3.) HbA c, % mmol.mol. (.7) () C-peptide, nmol.l.3
GLP-2 (Rat) ELISA. For the quantitative determination of glucagon-like peptide 2 (GLP-2) in rat serum and plasma
 GLP-2 (Rat) ELISA For the quantitative determination of glucagon-like peptide 2 (GLP-2) in rat serum and plasma For Research Use Only. Not For Use In Diagnostic Procedures. Catalog Number: 48-GP2RT-E01
GLP-2 (Rat) ELISA For the quantitative determination of glucagon-like peptide 2 (GLP-2) in rat serum and plasma For Research Use Only. Not For Use In Diagnostic Procedures. Catalog Number: 48-GP2RT-E01
Supplementary Figure 1. Localization of face patches (a) Sagittal slice showing the location of fmri-identified face patches in one monkey targeted
 Supplementary Figure 1. Localization of face patches (a) Sagittal slice showing the location of fmri-identified face patches in one monkey targeted for recording; dark black line indicates electrode. Stereotactic
Supplementary Figure 1. Localization of face patches (a) Sagittal slice showing the location of fmri-identified face patches in one monkey targeted for recording; dark black line indicates electrode. Stereotactic
In vitro human regulatory T cell suppression assay
 Human CD4 + CD25 + regulatory T cell isolation, in vitro suppression assay and analysis In vitro human regulatory T cell suppression assay Introduction Regulatory T (Treg) cells are a subpopulation of
Human CD4 + CD25 + regulatory T cell isolation, in vitro suppression assay and analysis In vitro human regulatory T cell suppression assay Introduction Regulatory T (Treg) cells are a subpopulation of
Supporting Information for
 Supporting Information for Rupture of Lipid Membranes Induced by Amphiphilic Janus Nanoparticles Kwahun Lee, Liuyang Zhang, Yi Yi, Xianqiao Wang, Yan Yu* Department of Chemistry, Indiana University, Bloomington,
Supporting Information for Rupture of Lipid Membranes Induced by Amphiphilic Janus Nanoparticles Kwahun Lee, Liuyang Zhang, Yi Yi, Xianqiao Wang, Yan Yu* Department of Chemistry, Indiana University, Bloomington,
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE
 Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
4.3 Measures of Variation
 4.3 Measures of Variation! How much variation is there in the data?! Look for the spread of the distribution.! What do we mean by spread? 1 Example Data set:! Weight of contents of regular cola (grams).
4.3 Measures of Variation! How much variation is there in the data?! Look for the spread of the distribution.! What do we mean by spread? 1 Example Data set:! Weight of contents of regular cola (grams).
The distribution of log 2 ratio (H/L) for quantified peptides. cleavage sites in each bin of log 2 ratio of quantified. peptides
 Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
HD1 (FLU) HD2 (EBV) HD2 (FLU)
 ramer staining + anti-pe beads ramer staining a HD1 (FLU) HD2 (EBV) HD2 (FLU).73.11.56.46.24 1.12 b CD127 + c CD127 + d CD127 - e CD127 - PD1 - PD1 + PD1 + PD1-1 1 1 1 %CD127 + PD1-8 6 4 2 + anti-pe %CD127
ramer staining + anti-pe beads ramer staining a HD1 (FLU) HD2 (EBV) HD2 (FLU).73.11.56.46.24 1.12 b CD127 + c CD127 + d CD127 - e CD127 - PD1 - PD1 + PD1 + PD1-1 1 1 1 %CD127 + PD1-8 6 4 2 + anti-pe %CD127
GLP-2 ELISA. For the quantitative determination of GLP-2 in human serum and plasma samples.
 GLP-2 ELISA For the quantitative determination of GLP-2 in human serum and plasma samples. For Research Use Only. Not For Use In Diagnostic Procedures. Catalog Number: 48-GP2HU-E01.1 Size: 96 wells Version:
GLP-2 ELISA For the quantitative determination of GLP-2 in human serum and plasma samples. For Research Use Only. Not For Use In Diagnostic Procedures. Catalog Number: 48-GP2HU-E01.1 Size: 96 wells Version:
UF#Stats#Club#STA#2023#Exam#1#Review#Packet# #Fall#2013#
 UF#Stats#Club#STA##Exam##Review#Packet# #Fall## The following data consists of the scores the Gators basketball team scored during the 8 games played in the - season. 84 74 66 58 79 8 7 64 8 6 78 79 77
UF#Stats#Club#STA##Exam##Review#Packet# #Fall## The following data consists of the scores the Gators basketball team scored during the 8 games played in the - season. 84 74 66 58 79 8 7 64 8 6 78 79 77
Supplementary Figure 1. BMS enhances human T cell activation in vitro in a
 Supplementary Figure 1. BMS98662 enhances human T cell activation in vitro in a concentration-dependent manner. Jurkat T cells were activated with anti-cd3 and anti-cd28 antibody in the presence of titrated
Supplementary Figure 1. BMS98662 enhances human T cell activation in vitro in a concentration-dependent manner. Jurkat T cells were activated with anti-cd3 and anti-cd28 antibody in the presence of titrated
Astrocyte signaling controls spike timing-dependent depression at neocortical synapses
 Supplementary Information Astrocyte signaling controls spike timing-dependent depression at neocortical synapses Rogier Min and Thomas Nevian Department of Physiology, University of Berne, Bern, Switzerland
Supplementary Information Astrocyte signaling controls spike timing-dependent depression at neocortical synapses Rogier Min and Thomas Nevian Department of Physiology, University of Berne, Bern, Switzerland
Triglyceride Assay Kit-Bulk 5 Plate Kit Cat# TG-5-RB
 Triglyceride Assay Kit-Bulk 5 Plate Kit Cat# TG-5-RB INSTRUCTION MANUAL (ZBM-14) STORAGE CONDITIONS Reagents A & B, Buffers: Store at 2-8 C. Glycerol Standard -20 C For Research Use Only Not For Use In
Triglyceride Assay Kit-Bulk 5 Plate Kit Cat# TG-5-RB INSTRUCTION MANUAL (ZBM-14) STORAGE CONDITIONS Reagents A & B, Buffers: Store at 2-8 C. Glycerol Standard -20 C For Research Use Only Not For Use In
Supplemental Figure S1
 Supplemental Figure S1 Children from Nandi (low pathogen burden area) and Kisumu (high pathogen burden area) have significantly different pathogen-specific serological profiles. Serum antibody titers for
Supplemental Figure S1 Children from Nandi (low pathogen burden area) and Kisumu (high pathogen burden area) have significantly different pathogen-specific serological profiles. Serum antibody titers for
Supplementary Online Content
 Supplementary Online Content Schmidt B, Whyte RK, Asztalos EV, et al; for the Canadian Oxygen Trial (COT) Group. Effects of targeting higher vs lower arterial oxygen saturations on death or disability
Supplementary Online Content Schmidt B, Whyte RK, Asztalos EV, et al; for the Canadian Oxygen Trial (COT) Group. Effects of targeting higher vs lower arterial oxygen saturations on death or disability
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2638 Figure S1 Morphological characteristics of fetal testes and ovaries from 6.5-20 developmental weeks. Representative images of Hematoxylin and Eosin staining of testes and ovaries over
DOI: 10.1038/ncb2638 Figure S1 Morphological characteristics of fetal testes and ovaries from 6.5-20 developmental weeks. Representative images of Hematoxylin and Eosin staining of testes and ovaries over
The Impact of Column Peak Capacity on the Multi- dimensional Chromatography of Complex Peptide Mixtures
 ISPPP 2003 November 12, 2003 The Impact of Column Peak Capacity on the Multi- dimensional Chromatography of Complex Peptide Mixtures Martin Gilar Life Sciences Chemistry, Waters Corporation Outline Highly
ISPPP 2003 November 12, 2003 The Impact of Column Peak Capacity on the Multi- dimensional Chromatography of Complex Peptide Mixtures Martin Gilar Life Sciences Chemistry, Waters Corporation Outline Highly
Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes.
 Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
nature methods Organelle-specific, rapid induction of molecular activities and membrane tethering
 nature methods Organelle-specific, rapid induction of molecular activities and membrane tethering Toru Komatsu, Igor Kukelyansky, J Michael McCaffery, Tasuku Ueno, Lidenys C Varela & Takanari Inoue Supplementary
nature methods Organelle-specific, rapid induction of molecular activities and membrane tethering Toru Komatsu, Igor Kukelyansky, J Michael McCaffery, Tasuku Ueno, Lidenys C Varela & Takanari Inoue Supplementary
