Supplementary figure legends
|
|
|
- Phillip Dickerson
- 5 years ago
- Views:
Transcription
1 Supplementary figure legends SUPPLEMENTRY FIGURE S1. Lentiviral construct. Schematic representation of the PCR fragment encompassing the genomic locus of mir-33a that was introduced in the lentiviral construct. SUPPLEMENTRY FIGURE S2. Reporter gene constructs. Schematic representation of the PCR product inserted in the reporter constructs for. human BC1, B. human CROT, C. human HDHB, D. human CPT1, E. Drosophila melanogaster CPTI, F. human TP8B1, G. NPC1 and H. SLC2525. Numbers denote the nucleotide position in GRCh37/hg19 human and BDGP R5/dm3 drosophila genome assembly. Feathered lines denote introns, and black boxes the PCR product, the mir-33 binding site or the open reading frame (ORF) as indicated. SUPPLEMENTRY FIGURE S3. mir-33b is localized in the SREBP1 gene. Schematic representation of the human SREBP1 locus with embedded mir-33b sequence including evolutionary conservation of the latter. SUPPLEMENTRY FIGURE S4. The binding site for mir-33 in BC1 is highly conserved and consists of tandem repetitions of seed-binding sites.. Evolutionary conservation of the predicted mir- 33 binding site in the 3 UTR of BC1. Light grey indicates possibility of classical Watson-Crick pairing with mir-33 and dark grey indicates GU pairs. B. The predicted mir-33 binding site in the 3 UTR of BC1 contains three tandem repeats of a sequence perfectly complementary to mir-33 (circled). The most favorable annealing is depicted. Since mutagenesis of one, two or three of these repeats might lead to alternative site usage in a potentially suboptimal annealing, which does not allow any conclusions about normal seed binding, we decided to mutagenize all three seed sequences in parallel. SUPPLEMENTRY FIGURE S5. mir-33 targets BC1 and HDHB in mouse adrenocortical cancer cell line Y1. Western Blot analysis of Y1 mouse adrenocortical cancer cell lines engineered to overexpress mir-33a. Note that the HDHB antibody crossreacts with a band around 45kDa in mouse samples. The band that corresponds to the size of human HDHB is marked with an arrow. SUPPLEMENTRY FIGURE S6. Conservation of mir-33 binding site in HDHB 3 UTR. Evolutionary conservation of the predicted mir-33 binding site in the 3 UTR of HDHB. Light grey shading indicates a residue that can undergo a classical Watson-Crick pair with mir-33 while dark grey shading indicates GU pairs. SUPPLEMENTRY FIGURE S7. Conservation of mir-33 binding site in CPT1 3 UTR.. and B. Predicted binding of mir-33a and mir-33b to the () unconserved and (B) conserved mir-33 binding site in the CPT1 3 UTR. C. lignment of the conserved mir-33 binding site in Drosophila and Ciona intestinalis CPT1 gene. D. Predicted seed binding region in the 3 UTR of Xenopus tropicalis CPT1. Shading represents the same as in Figure 5 SUPPLEMENTRY FIGURE S8. Conservation of mir-33 binding site in CROT 3 UTR. Evolutionary conservation of the predicted mir-33 binding site in the 3 UTR of CROT. B. and C. Predicted binding of mir-33b (B) shows two additional Watson-Crick pairs than binding to mir-33a (C). D. Evolutionary conservation of second mir-33 binding site in CROT 3 UTR. Shading represents the same as in Figure 5. SUPPLEMENTRY FIGURE S9. The splice variant carrying the predicted mir-33 binding site is dominant.. Schematic representation of the CPT1 alternative transcripts affecting the 3 UTR B. 1
2 bsolute Ct values of quantitative RT PCR for transcripts with or without the mir-33 binding site in different human organs SUPPLEMENTRY FIGURE S10. mir-33 potentially targets additional genes involved in lipid metabolism.. D. Evolutionary conservation of predicted binding sites in the 3 UTR of NPC1 ( and B), TP8B1 (C) and SLC2525 (D). Formating is identical to Figure 7. E. and F. Reporter assays to assess influence of premir-33 on the indicated 3 UTRs cloned downstream of a constitutively active firefly luciferase cassette in the pgl3-control plasmid. Experimental setup and presentation is identical to Figure 2B. SUPPLEMENTRY FIGURE S11. TLC analysis of cellular lipids. Representative Thin layer chromatography scan used to quantify cellular lipids after staining with Primulin and scanning using a fluorescence scanner. SUPPLEMENTRY FIGURE S12. Influence of culture conditions and mouse diet on mir-33 levels.. mir-33 levels are increased upon culture at high cell density and in medium containing lipid free FCS. B. Northern Blot analysis of mice fed with high fat diet or control chow for 16 weeks starting at age 4 weeks. s a positive control, HepG2 overexpressing mir-33a were used. Hybridization to mir-33a LN probe and for U6 RN are shown. C. Northern Blot analysis of mice fed with high fat diet or control chow for 16 weeks starting at age 8 weeks as in (B) D. Northern Blot analysis of mice on chow diet, after 16h and 39h of starvation or refeeding for 12h. SUPPLEMENTRY FIGURE S13. Tissue distribution of mir-33a. Equal amounts (10µg) of mouse tissue total RN were analyzed by northern blot for expression of mir-33a and U6. Position of mature mir-33a and premir-33a are indicated. Ethidium Bromide staining of polyacrylamide gels is shown to demonstrate equal loading. HepG2 cell transduced with a control lentivirus (HepG2 GIPZ) and a lentivirus driving mir-33a (HepG2 mir-33a) were loaded to show the expression level of mir-33a in overexpression experiments. Please note that the HepG2 mir-33a sample on the second gel is from the same gel and exposure as the remainder of this northern blot. 2
3 Chromosome 22: 1 kb SREBF2 Exon mir-33a PCR product Exon Supplemental Figure S1
4 Chromosome 9: kb PCR product seed 1 seed 2 seed 3 BC1 ORF 3 UTR B Chromosome 7: 500 bases PCR site1 site2 CROT ORF 3 UTR C Chromosome 2: 200 bases PCR site HDHB ORF 3 UTR D 1 kb Chromosome 11: PCR unconserved Supplemental Figure S2 -D conserved C-terminal splice variants CPT1
5 E Drosophila Chromosome 2R bases PCR site CPTI ORF 3 UTR F 1 kb Chromosome 18: PCR site 2 site 1 TP8B1 ORF 3 UTR G Chromosome 18: bases PCR site NPC1 ORF 3 UTR H 1 kb Chromosome 9: SLC PCR site ORF 3 UTR Supplemental Figure S2 E-H
6 chr17: SREBP-1 mir-33b Homo sapiens GUGCUUGCUGUUGCUUG Macaca mulatta GUGCUUGCUGUUGCUUG Pan troglodytes GUGCUUGCUGUUGCUUG Canis familiaris GUGCUUGCUGUUGCUUG Xenopus tropicalis GUGCUUGUUGUUGCUUG Tetraodon nigricans GUGCUUGUGUUGCUUG Supplemental Figure S3
7 B 3 CGUUCGUUGUCGUUCGUG 5 mir-33b 3 CGUUCGUUGUGUUCGUG 5 mir-33a : _ UUCUGCUGCUU-CUGC Homo sapiens (BC1 3 UTR) UUCUGCUGCCUU-CUGC Mus musculus (BC1 3 UTR) UUCUGCUGCCUU-CUGC Rattus norvegicus (BC1 3 UTR) UUCUGCUGCUU-CUGC Canis familiaris (BC1 3 UTR) CUCUGCUGCU-CUGC Gallus gallus (BC1 3 UTR) CCCUGCU-CUGC Danio rerio (BC1 3 UTR) GCUUCUGCG-GCUGC Tetraodon nigroviridis (BC1 3 UTR) CUCUGCUGCU-CUGC Xenopus tropicalis (BC1 3 UTR) U C U U U C U G C U G C U U C G U U C G U U G U C U G C U G C G U U C G U G Supplementary Figure S4
8 control mir-33a BC1 HDHB HDH CTIN Supplemental Figure S5
9 3 CGUUCGUUGUCGUUCGUG 5 mir-33b 3 CGUUCGUUGUGUUCGUG 5 mir-33a _ : _ : UCCCUGGCUGCCUUUCUGCU Homo sapiens (HDHB 3 UTR) UUGCCUGGGCUGCCUUUCUGCC Mus musculus (HDHB 3 UTR) UCGCCUGGGCUGCCUUUCUGCC Rattus norvegicus (HDHB 3 UTR) UCGCCUGGCUGCCUUCCUGCU Canis familiaris (HDHB 3 UTR) CUGUGCCUGGCUGCCUUUCUGCC Gallus gallus (HDHB 3 UTR) Supplemental Figure S6
10 B C 3 CGUUCGUUGUC-GUUCGUG 5 mir-33b 3'CGUUCGUUGU-GUUCGUG 5 mir-33a :: : GUCUCUUCGUUUUUGC Homo sapiens (CPT1 3 UTR) (conserved only in primates) 3 CGUUCGUUGUC-GUUCGUG 5 mir-33b 3'CGUUCGUUGU-GUUCGUG 5 mir-33a : :: : CUCGUGCGUUUCUGC Homo sapiens (CPT1 3 UTR) CUCCUGGCGUUCCUGC Mus musculus (CPT1 3 UTR) CUCCUGGCGUUCCUGC Rattus norvegicus (CPT1 3 UTR) CCGUGCGUUUCUGC Canis familiaris (CPT1 3 UTR) CUUGUGCGUUUCUGC Gallus gallus (CPT1 3 UTR) 3'CGUUCGCUGUGUUCGUG 5 Drosophila melanogaster mir-33 CUCUGC Drosophila melanogaster CUCUGC Drosophila simulans CUCUGC Drosophila sechellia CUUCUGC Drosophila yakuba CUCUGC Drosophila erecta GCCUGC Drosophila ananassae CUGCUGC Drosophila pseudo-obscura CUGCUGC Drosophila persimilis CUGC Drosophila willistoni UGGCUGC Drosophila virilis UGGCUGC Drosophila grimshawi UGCUGC Drosophila mojavensis UCUGC Ciona intestinalis (Ciona intestinalis sequence is derived from chr14q: 522, ,669 (assembly Mar (JGI 2.1/ci2)) unconserved site conserved site D Xenopus tropicalis CPT1 3 UTR (scaffold_1374:1,500-1,520; assembly JGI 4.1/xenTro2) 3 CGUUCGCUGUCGUUCGUG 5 mir-33b (Xenopus) 3 CGUUCGCUGUGUUCGUG 5 mir-33a : 5 GTTCTGTCCCTGCT 3 CPT1 3 UTR Xenopus Supplemental Figure S7
11 B C D 3 CGUUCGUUGUCGUUCGUG 5 mirn33b 3'CGUUCGUUGUGUUCGUG 5 mirn33a -:: : -: UCCUCUGUGCGCGCUGC Homo sapiens (CROT) UCCUGUGUUGCCCCUGC Mus musculus (CROT) UUCUGUGUUGCCGCUGC Rattus norvegicus (CROT) UCCUCUGUGCGUGCGUGCG Canis familiaris (CROT) UGGCCUUUGCCGCUGCG Gallus gallus (CROT) 3 CGUUCGUUGUCGUUCGUG 5 mirn33b :: : UCCUCUGUGCGCGCUGC Homo sapiens (CROT) 3'CGUUCGUUGUGUUCGUG 5 mirn33a -:: : _: UCCUCUGUGCGCGCUGC Homo sapiens (CROT) 3 CGUUCGUUGUCGUUCGUG 5 mirn33b 3'CGUUCGUUGUGUUCGUG 5 mirn33a _ UCUC-CUUUCUGC Homo sapiens (CROT) UGUUUU-GCCUCCUGCC Rattus norvegicus (CROT) UCUCU-UUUUUUGC Canis familiaris (CROT) conserved site unconserved site Supplemental Figure S8
12 2 kb SITE 2 SITE 1 CPT1 CPT1 CPT1 Sus CPT1 I I L Rattus Cpt1a Equus CPT1 Mus Cpt1a Ovis CPT1 Xenopus cpt1a Gallus CPT1 Danio LOC BE D BG variant 1 variant 2 variant 3 pig rat horse mouse sheep frog chicken zebrafish human ESTs human other species Rhesus Mouse Dog Elephant Opossum Chicken X_tropicalis Zebrafish rs SINE LINE LTR DN Simple Low Complexity evolutionary conservation SNPs genomic repeats B variant 2 variant 1 difference in Ct Brain Colon Heart Kidney Liver Lung Spleen Testis Thymus Supplemental Figure S9
13 3 CGUUCGUUGUCGUUCGUG 5 mir-33b 3 CGUUCGUUGUGUUCGUG 5 mir-33a : CUCUGUGGCCU--CUGCC Homo sapiens (NPC1-3 UTR) CUUUUUGGCCU--CUGC Canis familiaris (NPC1-3 UTR) UUUUUUGCGGU--CUGC Gallus gallus (NPC1-3 UTR) UUUUGCUGCCU--CUGCC Xenopus tropicalis(npc1-3 UTR) (not conserved in rodents) B 3 CGUUCGUUGUCGUUCGUG 5 mir-33b 3 CGUUCGUUGUGUUCGUG 5 mir-33a : UGUUUGGCUUUUUUGC Homo sapiens (NPC1-3 UTR) UGUUUGGCUUUUUUGC Mus musculus (NPC1-3 UTR) UGUUUGGCUUUUUUGC Rattus norvegicus (NPC1-3 UTR) UGUUUGGCUUUUUUGC Canis familiaris (NPC1-3 UTR) UGUUCUCUCUUUGC Gallus gallus (NPC1-3 UTR) C D 3 CGUUCGUUGUCGUUCGUG 5 mir-33b 3'CGUUCGUUGU-GUUCGUG 5 mir-33a : : : UGGGGUUGCUGC Homo sapiens (TP8B1 3 UTR) UGGGGGGUUGGCUGC Mus musculus (TP8B1 3 UTR) CGGGGGGUUGCUGC Rattus norvegicus (TP8B1 3 UTR) 3 CGUUCGUUGUC-GUUCGUG 5 mir-33b 3'CGUUCGUUGU-GUUCGUG 5 mir-33a : CUGGUGCGUCUGC Homo sapiens (SLC UTR) CGGUGCGUCUGC Mus musculus (SLC UTR) UGGUGCGUCUGC Rattus norvegicus (SLC UTR) UGGUGCGUCUGC Canis familiaris (SLC UTR) E F Relative luciferase activity control control mir-33 * SLC2525 NPC1 Supplemental Figure S10 * Relative luciferase activity * control PT8B1 control mir-33
14 Triacylglycerol Free fatty acids Cholesterol Phospholipids control mir-33a HepG2 Supplemental Figure S11
15 HepG2 mir-33a Normal serum sparse dense Delipidized serum sparse dense mir-33a U6 B HepG2 mir-33a Control High Fat diet diet HepG2 control mir-33a U6 C HepG2 mir-33a Control High Fat diet diet HepG2 control mir-33a U6 D HepG2 mir-33a chow diet 15h fasting 39h fasting refed HepG2 control mir-33a U6 Supplemental figure S12
16 Supplemental Figure S13 B HepG2 mir-33a brain stem brain cortex heart testis lung spleen stomach HepG2 HepG2 mir-33a muscle kidney brown adipose tissue gonadal white adipose tissue liver large intestine small intestine HepG2 pre-mir-33 mature mir-33 U6 Ethidium Bromide pre-mir-33 mature mir-33 U6 Ethidium Bromide
SUPPLEMENTARY MATERIAL
 SYPPLEMENTARY FIGURE LEGENDS SUPPLEMENTARY MATERIAL Figure S1. Phylogenic studies of the mir-183/96/182 cluster and 3 -UTR of Casp2. (A) Genomic arrangement of the mir-183/96/182 cluster in vertebrates.
SYPPLEMENTARY FIGURE LEGENDS SUPPLEMENTARY MATERIAL Figure S1. Phylogenic studies of the mir-183/96/182 cluster and 3 -UTR of Casp2. (A) Genomic arrangement of the mir-183/96/182 cluster in vertebrates.
Supplementary Figure 1. CFTR protein structure and domain architecture.
 A Plasma Membrane NH ₂ COOH Supplementary Figure. CFT protein structure and domain architecture. (A) Open state CFT homology model, ribbon representation from Serohijos et al. 8 PNAS 5:356. CFT domains
A Plasma Membrane NH ₂ COOH Supplementary Figure. CFT protein structure and domain architecture. (A) Open state CFT homology model, ribbon representation from Serohijos et al. 8 PNAS 5:356. CFT domains
Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein
 Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein content relative to GAPDH in two independent experiments.
Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein content relative to GAPDH in two independent experiments.
Mouse Clec9a ORF sequence
 Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Supplementary Data to: Marco Mariotti and Roderic Guigó
 Supplementary Data to: Selenoprofiles: profile-based scanning of eukaryotic genome sequences for selenoprotein genes Marco Mariotti and Roderic Guigó Section S1: patterns used with SECISearch We report
Supplementary Data to: Selenoprofiles: profile-based scanning of eukaryotic genome sequences for selenoprotein genes Marco Mariotti and Roderic Guigó Section S1: patterns used with SECISearch We report
Uninformative BRCA Tests. Rebecca Sutphen, M.D. Professor, USF College of Medicine Chief Medical Officer, InformedDNA
 Uninformative RCA Tests Rebecca Sutphen, M.D. Professor, USF College of Medicine Chief Medical Officer, InformedDNA Uninformative RCA tests 1. R/OV cancer patient with normal results 2. Cancer-free person
Uninformative RCA Tests Rebecca Sutphen, M.D. Professor, USF College of Medicine Chief Medical Officer, InformedDNA Uninformative RCA tests 1. R/OV cancer patient with normal results 2. Cancer-free person
Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent Protein.
 prfp-vector RFP Exon1 Intron RFP Exon2 prfp-mir-124 mir-93/124 RFP Exon1 Intron RFP Exon2 Untransfected prfp-vector prfp-mir-93 prfp-mir-124 Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent
prfp-vector RFP Exon1 Intron RFP Exon2 prfp-mir-124 mir-93/124 RFP Exon1 Intron RFP Exon2 Untransfected prfp-vector prfp-mir-93 prfp-mir-124 Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12005 a S Ψ ΨΨ ΨΨ ΨΨ Ψ Ψ Ψ Ψ Ψ ΨΨ Ψ Ψ ΨΨΨΨΨΨ S1 (a.a. 1 751) S2 (a.a. 752 1353) S1 Fc S1 (a.a. 1-747) Fc b HCoV-EMC-S1-Fc SARS-CoV-S1-Fc HCoV-EMC-S1-Fc SARS-CoV-S1-Fc kda - 170 - - 130
doi:10.1038/nature12005 a S Ψ ΨΨ ΨΨ ΨΨ Ψ Ψ Ψ Ψ Ψ ΨΨ Ψ Ψ ΨΨΨΨΨΨ S1 (a.a. 1 751) S2 (a.a. 752 1353) S1 Fc S1 (a.a. 1-747) Fc b HCoV-EMC-S1-Fc SARS-CoV-S1-Fc HCoV-EMC-S1-Fc SARS-CoV-S1-Fc kda - 170 - - 130
The Emergence of Alternative 39 and 59 Splice Site Exons from Constitutive Exons
 The Emergence of Alternative 39 and 59 Splice Site Exons from Constitutive Exons Eli Koren, Galit Lev-Maor, Gil Ast * Department of Human Molecular Genetics, Sackler Faculty of Medicine, Tel Aviv University,
The Emergence of Alternative 39 and 59 Splice Site Exons from Constitutive Exons Eli Koren, Galit Lev-Maor, Gil Ast * Department of Human Molecular Genetics, Sackler Faculty of Medicine, Tel Aviv University,
Supporting Information
 Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Kirschner, 2005). Briefly, parallel MEF cultures were isolated from single littermate
 SUPPLEMENTAL MATERIALS AND METHODS Generation of MEFs, osteoblasts and cell culture Prkar1a -/- and control MEFs were generated and cultured as described (Nadella and Kirschner, 2005). Briefly, parallel
SUPPLEMENTAL MATERIALS AND METHODS Generation of MEFs, osteoblasts and cell culture Prkar1a -/- and control MEFs were generated and cultured as described (Nadella and Kirschner, 2005). Briefly, parallel
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3073 LATS2 Binding ability to SIAH2 Degradation by SIAH2 1-160 161-402 403-480 -/ 481-666 1-666 - - 667-1088 -/ 1-0 Supplementary Figure 1 Schematic drawing of LATS2 deletion mutants and
DOI: 10.1038/ncb3073 LATS2 Binding ability to SIAH2 Degradation by SIAH2 1-160 161-402 403-480 -/ 481-666 1-666 - - 667-1088 -/ 1-0 Supplementary Figure 1 Schematic drawing of LATS2 deletion mutants and
Table S1. Quantitative RT-PCR primers
 Table S1. Quantitative RT-PCR primers Gene Forward Primer Reverse Primer Human ApoB gcaagcagaagccagaagta ccatttggagaagcagtttgg Human ApoA1 gaaagctgcggtgctgac agtggccaggtccttcact Human MTP acggccattcccattgtg
Table S1. Quantitative RT-PCR primers Gene Forward Primer Reverse Primer Human ApoB gcaagcagaagccagaagta ccatttggagaagcagtttgg Human ApoA1 gaaagctgcggtgctgac agtggccaggtccttcact Human MTP acggccattcccattgtg
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/2/1/ra81/dc1 Supplementary Materials for Delivery of MicroRNA-126 by Apoptotic Bodies Induces CXCL12- Dependent Vascular Protection Alma Zernecke,* Kiril Bidzhekov,
www.sciencesignaling.org/cgi/content/full/2/1/ra81/dc1 Supplementary Materials for Delivery of MicroRNA-126 by Apoptotic Bodies Induces CXCL12- Dependent Vascular Protection Alma Zernecke,* Kiril Bidzhekov,
WDR62 is associated with the spindle pole and mutated in human microcephaly
 WDR62 is associated with the spindle pole and mutated in human microcephaly Adeline K. Nicholas, Maryam Khurshid, Julie Désir, Ofélia P. Carvalho, James J. Cox, Gemma Thornton, Rizwana Kausar, Muhammad
WDR62 is associated with the spindle pole and mutated in human microcephaly Adeline K. Nicholas, Maryam Khurshid, Julie Désir, Ofélia P. Carvalho, James J. Cox, Gemma Thornton, Rizwana Kausar, Muhammad
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2211 a! mir-143! b! mir-103/107! let-7a! mir-144! mir-122a! mir-126-3p! mir-194! mir-27a! mir-30c! Figure S1 Northern blot analysis of mir-143 expression dependent on feeding conditions.
DOI: 10.1038/ncb2211 a! mir-143! b! mir-103/107! let-7a! mir-144! mir-122a! mir-126-3p! mir-194! mir-27a! mir-30c! Figure S1 Northern blot analysis of mir-143 expression dependent on feeding conditions.
Alternative splicing. Biosciences 741: Genomics Fall, 2013 Week 6
 Alternative splicing Biosciences 741: Genomics Fall, 2013 Week 6 Function(s) of RNA splicing Splicing of introns must be completed before nuclear RNAs can be exported to the cytoplasm. This led to early
Alternative splicing Biosciences 741: Genomics Fall, 2013 Week 6 Function(s) of RNA splicing Splicing of introns must be completed before nuclear RNAs can be exported to the cytoplasm. This led to early
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Supplementary Figure 1. Quantile-quantile (Q-Q) plots. (Panel A) Q-Q plot graphical
 Supplementary Figure 1. Quantile-quantile (Q-Q) plots. (Panel A) Q-Q plot graphical representation using all SNPs (n= 13,515,798) including the region on chromosome 1 including SORT1 which was previously
Supplementary Figure 1. Quantile-quantile (Q-Q) plots. (Panel A) Q-Q plot graphical representation using all SNPs (n= 13,515,798) including the region on chromosome 1 including SORT1 which was previously
Research Article Base Composition Characteristics of Mammalian mirnas
 Journal of Nucleic Acids Volume 2013, Article ID 951570, 6 pages http://dx.doi.org/10.1155/2013/951570 Research Article Base Composition Characteristics of Mammalian mirnas Bin Wang Department of Chemistry,
Journal of Nucleic Acids Volume 2013, Article ID 951570, 6 pages http://dx.doi.org/10.1155/2013/951570 Research Article Base Composition Characteristics of Mammalian mirnas Bin Wang Department of Chemistry,
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1 Schematic depiction of the tandem Fc GDF15. Supplementary Figure 2 Supplementary Figure 2 Gfral mrna levels in the brains of both wild-type and knockout Gfral
Supplementary Figure 1 Supplementary Figure 1 Schematic depiction of the tandem Fc GDF15. Supplementary Figure 2 Supplementary Figure 2 Gfral mrna levels in the brains of both wild-type and knockout Gfral
Figure mouse globin mrna PRECURSOR RNA hybridized to cloned gene (genomic). mouse globin MATURE mrna hybridized to cloned gene (genomic).
 Splicing Figure 14.3 mouse globin mrna PRECURSOR RNA hybridized to cloned gene (genomic). mouse globin MATURE mrna hybridized to cloned gene (genomic). mrna Splicing rrna and trna are also sometimes spliced;
Splicing Figure 14.3 mouse globin mrna PRECURSOR RNA hybridized to cloned gene (genomic). mouse globin MATURE mrna hybridized to cloned gene (genomic). mrna Splicing rrna and trna are also sometimes spliced;
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
Supplementary Information. MicroRNA-33b knock-in mice for an intron of sterol regulatory
 Supplementary Information MicroRNA-33b knock-in mice for an intron of sterol regulatory element-binding factor 1 (Srebf1) exhibit reduced HDL-C in vivo Takahiro Horie, Tomohiro Nishino, Osamu Baba, Yasuhide
Supplementary Information MicroRNA-33b knock-in mice for an intron of sterol regulatory element-binding factor 1 (Srebf1) exhibit reduced HDL-C in vivo Takahiro Horie, Tomohiro Nishino, Osamu Baba, Yasuhide
Figure 1: Final annotation map of Contig 9
 Introduction With rapid advances in sequencing technology, particularly with the development of second and third generation sequencing, genomes for organisms from all kingdoms and many phyla have been
Introduction With rapid advances in sequencing technology, particularly with the development of second and third generation sequencing, genomes for organisms from all kingdoms and many phyla have been
Data and text mining applied to the computational study of protein interaction networks
 Data and text mining applied to the computational study of protein interaction networks Miguel Andrade Faculty of Biology, Johannes Gutenberg University Institute of Molecular Biology Mainz, Germany andrade@uni-mainz.de
Data and text mining applied to the computational study of protein interaction networks Miguel Andrade Faculty of Biology, Johannes Gutenberg University Institute of Molecular Biology Mainz, Germany andrade@uni-mainz.de
Génétique des DCP. Deciphering the molecular bases of ciliopathies. Estelle Escudier, INSERM U 681 Serge Amselem, INSERM U 654
 Génétique des DCP Estelle Escudier, INSERM U 681 Serge Amselem, INSERM U 654 Deciphering the molecular bases of ciliopathies Linkage analyses Candidate gene approaches Chromosome abnormalities Comparative
Génétique des DCP Estelle Escudier, INSERM U 681 Serge Amselem, INSERM U 654 Deciphering the molecular bases of ciliopathies Linkage analyses Candidate gene approaches Chromosome abnormalities Comparative
Males- Western Diet WT KO Age (wks) Females- Western Diet WT KO Age (wks)
 Relative Arv1 mrna Adrenal 33.48 +/- 6.2 Skeletal Muscle 22.4 +/- 4.93 Liver 6.41 +/- 1.48 Heart 5.1 +/- 2.3 Brain 4.98 +/- 2.11 Ovary 4.68 +/- 2.21 Kidney 3.98 +/-.39 Lung 2.15 +/-.6 Inguinal Subcutaneous
Relative Arv1 mrna Adrenal 33.48 +/- 6.2 Skeletal Muscle 22.4 +/- 4.93 Liver 6.41 +/- 1.48 Heart 5.1 +/- 2.3 Brain 4.98 +/- 2.11 Ovary 4.68 +/- 2.21 Kidney 3.98 +/-.39 Lung 2.15 +/-.6 Inguinal Subcutaneous
Supplementary Figure 1
 Supplementary Figure 1 how HFD how HFD Epi WT p p Hypothalamus p p Inguinal WT T Liver Lean mouse adipocytes p p p p p p Obese mouse adipocytes Kidney Muscle Spleen Heart p p p p p p p p Extracellular
Supplementary Figure 1 how HFD how HFD Epi WT p p Hypothalamus p p Inguinal WT T Liver Lean mouse adipocytes p p p p p p Obese mouse adipocytes Kidney Muscle Spleen Heart p p p p p p p p Extracellular
SUPPLEMENTARY INFORMATION. Supp. Fig. 1. Autoimmunity. Tolerance APC APC. T cell. T cell. doi: /nature06253 ICOS ICOS TCR CD28 TCR CD28
 Supp. Fig. 1 a APC b APC ICOS ICOS TCR CD28 mir P TCR CD28 P T cell Tolerance Roquin WT SG Icos mrna T cell Autoimmunity Roquin M199R SG Icos mrna www.nature.com/nature 1 Supp. Fig. 2 CD4 + CD44 low CD4
Supp. Fig. 1 a APC b APC ICOS ICOS TCR CD28 mir P TCR CD28 P T cell Tolerance Roquin WT SG Icos mrna T cell Autoimmunity Roquin M199R SG Icos mrna www.nature.com/nature 1 Supp. Fig. 2 CD4 + CD44 low CD4
Annotation of Chimp Chunk 2-10 Jerome M Molleston 5/4/2009
 Annotation of Chimp Chunk 2-10 Jerome M Molleston 5/4/2009 1 Abstract A stretch of chimpanzee DNA was annotated using tools including BLAST, BLAT, and Genscan. Analysis of Genscan predicted genes revealed
Annotation of Chimp Chunk 2-10 Jerome M Molleston 5/4/2009 1 Abstract A stretch of chimpanzee DNA was annotated using tools including BLAST, BLAT, and Genscan. Analysis of Genscan predicted genes revealed
SUPPLEMENTARY INFORMATION
 Testing the accuracy of ancestral state reconstruction The accuracy of the ancestral state reconstruction with maximum likelihood methods can depend on the underlying model used in the reconstruction.
Testing the accuracy of ancestral state reconstruction The accuracy of the ancestral state reconstruction with maximum likelihood methods can depend on the underlying model used in the reconstruction.
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna
 Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna targeted for hamster PAQR3. At 24 h after the transfection,
Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna targeted for hamster PAQR3. At 24 h after the transfection,
SC-L-H shared(37) Specific (1)
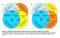 A. Brain (total 2) Tissue-specific (2) Brain-heart shared (8) Specific (2) CNS-heart shared(68) Specific () Heart (total 4) Tissue-specific () B-L-H shared (37) specific () B. Brain (total 2) Tissue-specific
A. Brain (total 2) Tissue-specific (2) Brain-heart shared (8) Specific (2) CNS-heart shared(68) Specific () Heart (total 4) Tissue-specific () B-L-H shared (37) specific () B. Brain (total 2) Tissue-specific
Supplementary information
 Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated
 Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
FINAL ANNOTATION REPORT: Drosophila virilis Fosmid 11 (48P14) Robert Carrasquillo Bio 4342
 FINAL ANNOTATION REPORT: Drosophila virilis Fosmid 11 (48P14) Robert Carrasquillo Bio 4342 2006 TABLE OF CONTENTS I. Overview... 3 II. Genes... 4 III. Clustal Analysis... 15 IV. Repeat Analysis... 17 V.
FINAL ANNOTATION REPORT: Drosophila virilis Fosmid 11 (48P14) Robert Carrasquillo Bio 4342 2006 TABLE OF CONTENTS I. Overview... 3 II. Genes... 4 III. Clustal Analysis... 15 IV. Repeat Analysis... 17 V.
Supplementary Information. Supplementary Figure 1
 Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma.
 Supplementary Figure 1 Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma. (a) Breeding scheme. R26-LSL-SB11 homozygous mice were bred to Trp53 LSL-R270H/+
Supplementary Figure 1 Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma. (a) Breeding scheme. R26-LSL-SB11 homozygous mice were bred to Trp53 LSL-R270H/+
A Central Role of MG53 in Metabolic Syndrome. and Type-2 Diabetes
 A Central Role of MG53 in Metabolic Syndrome and Type-2 Diabetes Yan Zhang, Chunmei Cao, Rui-Ping Xiao Institute of Molecular Medicine (IMM) Peking University, Beijing, China Accelerated Aging in China
A Central Role of MG53 in Metabolic Syndrome and Type-2 Diabetes Yan Zhang, Chunmei Cao, Rui-Ping Xiao Institute of Molecular Medicine (IMM) Peking University, Beijing, China Accelerated Aging in China
Supporting Information Table of Contents
 Supporting Information Table of Contents Supporting Information Figure 1 Page 2 Supporting Information Figure 2 Page 4 Supporting Information Figure 3 Page 5 Supporting Information Figure 4 Page 6 Supporting
Supporting Information Table of Contents Supporting Information Figure 1 Page 2 Supporting Information Figure 2 Page 4 Supporting Information Figure 3 Page 5 Supporting Information Figure 4 Page 6 Supporting
Chapter 2. Investigation into mir-346 Regulation of the nachr α5 Subunit
 15 Chapter 2 Investigation into mir-346 Regulation of the nachr α5 Subunit MicroRNA s (mirnas) are small (< 25 base pairs), single stranded, non-coding RNAs that regulate gene expression at the post transcriptional
15 Chapter 2 Investigation into mir-346 Regulation of the nachr α5 Subunit MicroRNA s (mirnas) are small (< 25 base pairs), single stranded, non-coding RNAs that regulate gene expression at the post transcriptional
Supplementary methods:
 Supplementary methods: Primers sequences used in real-time PCR analyses: β-actin F: GACCTCTATGCCAACACAGT β-actin [11] R: AGTACTTGCGCTCAGGAGGA MMP13 F: TTCTGGTCTTCTGGCACACGCTTT MMP13 R: CCAAGCTCATGGGCAGCAACAATA
Supplementary methods: Primers sequences used in real-time PCR analyses: β-actin F: GACCTCTATGCCAACACAGT β-actin [11] R: AGTACTTGCGCTCAGGAGGA MMP13 F: TTCTGGTCTTCTGGCACACGCTTT MMP13 R: CCAAGCTCATGGGCAGCAACAATA
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1. Differential expression of mirnas from the pri-mir-17-92a locus.
 Supplementary Figure 1 Differential expression of mirnas from the pri-mir-17-92a locus. (a) The mir-17-92a expression unit in the third intron of the host mir-17hg transcript. (b,c) Impact of knockdown
Supplementary Figure 1 Differential expression of mirnas from the pri-mir-17-92a locus. (a) The mir-17-92a expression unit in the third intron of the host mir-17hg transcript. (b,c) Impact of knockdown
Supplementary Table 1. List of primers used in this study
 Supplementary Table 1. List of primers used in this study Gene Forward primer Reverse primer Rat Met 5 -aggtcgcttcatgcaggt-3 5 -tccggagacacaggatgg-3 Rat Runx1 5 -cctccttgaaccactccact-3 5 -ctggatctgcctggcatc-3
Supplementary Table 1. List of primers used in this study Gene Forward primer Reverse primer Rat Met 5 -aggtcgcttcatgcaggt-3 5 -tccggagacacaggatgg-3 Rat Runx1 5 -cctccttgaaccactccact-3 5 -ctggatctgcctggcatc-3
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Lack of cadherins Celsr2 and Celsr3 impairs ependymal ciliogenesis, leading to fatal
 Lack of cadherins Celsr2 and Celsr3 impairs ependymal ciliogenesis, leading to fatal hydrocephalus Fadel TISSIR, Yibo QU, Mireille MONTCOUQUIOL, Libing ZHOU, Kouji KOMATSU, Dongbo SHI, Toshihiko FUJIMORI,
Lack of cadherins Celsr2 and Celsr3 impairs ependymal ciliogenesis, leading to fatal hydrocephalus Fadel TISSIR, Yibo QU, Mireille MONTCOUQUIOL, Libing ZHOU, Kouji KOMATSU, Dongbo SHI, Toshihiko FUJIMORI,
RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice
 SUPPLEMENTARY INFORMATION RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice Paul N Valdmanis, Shuo Gu, Kirk Chu, Lan Jin, Feijie Zhang,
SUPPLEMENTARY INFORMATION RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice Paul N Valdmanis, Shuo Gu, Kirk Chu, Lan Jin, Feijie Zhang,
Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University
 Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
mirna-guided regulation at the molecular level
 molecular level Hervé Seitz IGH du CNRS, Montpellier, France March 3, 2016 microrna target prediction . microrna target prediction mirna: target: 2 7 5 N NNNNNNNNNNNNNN 3 NNNNNN the seed microrna target
molecular level Hervé Seitz IGH du CNRS, Montpellier, France March 3, 2016 microrna target prediction . microrna target prediction mirna: target: 2 7 5 N NNNNNNNNNNNNNN 3 NNNNNN the seed microrna target
Supplementary Figures for
 mirns regulate s Supplementary igures for MicroRNs Reprogram Normal ibroblasts into Cancer ssociated ibroblasts in Ovarian Cancer nirban K. Mitra, Marion Zillhardt, Youjia Hua, Payal iwari, ndrea E. Murmann,
mirns regulate s Supplementary igures for MicroRNs Reprogram Normal ibroblasts into Cancer ssociated ibroblasts in Ovarian Cancer nirban K. Mitra, Marion Zillhardt, Youjia Hua, Payal iwari, ndrea E. Murmann,
microrna-200b and microrna-200c promote colorectal cancer cell proliferation via
 Supplementary Materials microrna-200b and microrna-200c promote colorectal cancer cell proliferation via targeting the reversion-inducing cysteine-rich protein with Kazal motifs Supplementary Table 1.
Supplementary Materials microrna-200b and microrna-200c promote colorectal cancer cell proliferation via targeting the reversion-inducing cysteine-rich protein with Kazal motifs Supplementary Table 1.
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Supplementary Information
 Supplementary Information Notch deficiency decreases hepatic lipid accumulation by induction of fatty acid oxidation No-Joon Song,#, Ui Jeong Yun,#, Sunghee Yang, Chunyan Wu, Cho-Rong Seo, A-Ryeong Gwon,,
Supplementary Information Notch deficiency decreases hepatic lipid accumulation by induction of fatty acid oxidation No-Joon Song,#, Ui Jeong Yun,#, Sunghee Yang, Chunyan Wu, Cho-Rong Seo, A-Ryeong Gwon,,
The clathrin adaptor Numb regulates intestinal cholesterol. absorption through dynamic interaction with NPC1L1
 The clathrin adaptor Numb regulates intestinal cholesterol absorption through dynamic interaction with NPC1L1 Pei-Shan Li 1, Zhen-Yan Fu 1,2, Ying-Yu Zhang 1, Jin-Hui Zhang 1, Chen-Qi Xu 1, Yi-Tong Ma
The clathrin adaptor Numb regulates intestinal cholesterol absorption through dynamic interaction with NPC1L1 Pei-Shan Li 1, Zhen-Yan Fu 1,2, Ying-Yu Zhang 1, Jin-Hui Zhang 1, Chen-Qi Xu 1, Yi-Tong Ma
MicroRNA and Male Infertility: A Potential for Diagnosis
 Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Supplementary Table 1. Metabolic parameters in GFP and OGT-treated mice
 Supplementary Table 1. Metabolic parameters in GFP and OGT-treated mice Fasted Refed GFP OGT GFP OGT Liver G6P (mmol/g) 0.03±0.01 0.04±0.02 0.60±0.04 0.42±0.10 A TGs (mg/g of liver) 20.08±5.17 16.29±0.8
Supplementary Table 1. Metabolic parameters in GFP and OGT-treated mice Fasted Refed GFP OGT GFP OGT Liver G6P (mmol/g) 0.03±0.01 0.04±0.02 0.60±0.04 0.42±0.10 A TGs (mg/g of liver) 20.08±5.17 16.29±0.8
Supplemental Information For: The genetics of splicing in neuroblastoma
 Supplemental Information For: The genetics of splicing in neuroblastoma Justin Chen, Christopher S. Hackett, Shile Zhang, Young K. Song, Robert J.A. Bell, Annette M. Molinaro, David A. Quigley, Allan Balmain,
Supplemental Information For: The genetics of splicing in neuroblastoma Justin Chen, Christopher S. Hackett, Shile Zhang, Young K. Song, Robert J.A. Bell, Annette M. Molinaro, David A. Quigley, Allan Balmain,
Supplementary Information
 Supplementary Information Supplementary Figure 1! a! b! Nfatc1!! Nfatc1"! P1! P2! pa1! pa2! ex1! ex2! exons 3-9! ex1! ex11!!" #" Nfatc1A!!" Nfatc1B! #"!" Nfatc1C! #" DN1! DN2! DN1!!A! #A!!B! #B!!C! #C!!A!
Supplementary Information Supplementary Figure 1! a! b! Nfatc1!! Nfatc1"! P1! P2! pa1! pa2! ex1! ex2! exons 3-9! ex1! ex11!!" #" Nfatc1A!!" Nfatc1B! #"!" Nfatc1C! #" DN1! DN2! DN1!!A! #A!!B! #B!!C! #C!!A!
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/9/430/ra57/dc1 Supplementary Materials for The 4E-BP eif4e axis promotes rapamycinsensitive growth and proliferation in lymphocytes Lomon So, Jongdae Lee, Miguel
www.sciencesignaling.org/cgi/content/full/9/430/ra57/dc1 Supplementary Materials for The 4E-BP eif4e axis promotes rapamycinsensitive growth and proliferation in lymphocytes Lomon So, Jongdae Lee, Miguel
Nature Immunology: doi: /ni Supplementary Figure 1. DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells.
 Supplementary Figure 1 DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells. (a) Scheme for the retroviral shrna screen. (b) Histogram showing CD4 expression (MFI) in WT cytotoxic
Supplementary Figure 1 DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells. (a) Scheme for the retroviral shrna screen. (b) Histogram showing CD4 expression (MFI) in WT cytotoxic
Supplemental Information. LEAP2 Is an Endogenous Antagonist. of the Ghrelin Receptor
 Cell Metabolism, Volume 27 Supplemental Information LEAP2 Is an Endogenous Antagonist of the Ghrelin Receptor Xuecai Ge, Hong Yang, Maria A. Bednarek, Hadas Galon-Tilleman, Peirong Chen, Michael Chen,
Cell Metabolism, Volume 27 Supplemental Information LEAP2 Is an Endogenous Antagonist of the Ghrelin Receptor Xuecai Ge, Hong Yang, Maria A. Bednarek, Hadas Galon-Tilleman, Peirong Chen, Michael Chen,
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004
 Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Bioinformatics Laboratory Exercise
 Bioinformatics Laboratory Exercise Biology is in the midst of the genomics revolution, the application of robotic technology to generate huge amounts of molecular biology data. Genomics has led to an explosion
Bioinformatics Laboratory Exercise Biology is in the midst of the genomics revolution, the application of robotic technology to generate huge amounts of molecular biology data. Genomics has led to an explosion
genomics for systems biology / ISB2020 RNA sequencing (RNA-seq)
 RNA sequencing (RNA-seq) Module Outline MO 13-Mar-2017 RNA sequencing: Introduction 1 WE 15-Mar-2017 RNA sequencing: Introduction 2 MO 20-Mar-2017 Paper: PMID 25954002: Human genomics. The human transcriptome
RNA sequencing (RNA-seq) Module Outline MO 13-Mar-2017 RNA sequencing: Introduction 1 WE 15-Mar-2017 RNA sequencing: Introduction 2 MO 20-Mar-2017 Paper: PMID 25954002: Human genomics. The human transcriptome
Transient cold shock enhances zinc-finger nuclease mediated gene disruption
 nature methods Transient cold shock enhances zincfinger nuclease mediated gene disruption Yannick Doyon, Vivian M Choi, Danny Xia, Thuy D Vo, Philip D Gregory & Michael C Holmes Supplementary figures and
nature methods Transient cold shock enhances zincfinger nuclease mediated gene disruption Yannick Doyon, Vivian M Choi, Danny Xia, Thuy D Vo, Philip D Gregory & Michael C Holmes Supplementary figures and
MODULE 3: TRANSCRIPTION PART II
 MODULE 3: TRANSCRIPTION PART II Lesson Plan: Title S. CATHERINE SILVER KEY, CHIYEDZA SMALL Transcription Part II: What happens to the initial (premrna) transcript made by RNA pol II? Objectives Explain
MODULE 3: TRANSCRIPTION PART II Lesson Plan: Title S. CATHERINE SILVER KEY, CHIYEDZA SMALL Transcription Part II: What happens to the initial (premrna) transcript made by RNA pol II? Objectives Explain
Doctor of Philosophy
 Regulation of Gene Expression of the 25-Hydroxyvitamin D la-hydroxylase (CYP27BI) Promoter: Study of A Transgenic Mouse Model Ivanka Hendrix School of Molecular and Biomedical Science The University of
Regulation of Gene Expression of the 25-Hydroxyvitamin D la-hydroxylase (CYP27BI) Promoter: Study of A Transgenic Mouse Model Ivanka Hendrix School of Molecular and Biomedical Science The University of
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients.
 Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12215 Supplementary Figure 1. The effects of full and dissociated GR agonists in supporting BFU-E self-renewal divisions. BFU-Es were cultured in self-renewal medium with indicated GR
doi:10.1038/nature12215 Supplementary Figure 1. The effects of full and dissociated GR agonists in supporting BFU-E self-renewal divisions. BFU-Es were cultured in self-renewal medium with indicated GR
MicroRNA in Cancer Karen Dybkær 2013
 MicroRNA in Cancer Karen Dybkær RNA Ribonucleic acid Types -Coding: messenger RNA (mrna) coding for proteins -Non-coding regulating protein formation Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
MicroRNA in Cancer Karen Dybkær RNA Ribonucleic acid Types -Coding: messenger RNA (mrna) coding for proteins -Non-coding regulating protein formation Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Annotation of Drosophila mojavensis fosmid 8 Priya Srikanth Bio 434W
 Annotation of Drosophila mojavensis fosmid 8 Priya Srikanth Bio 434W 5.1.2007 Overview High-quality finished sequence is much more useful for research once it is annotated. Annotation is a fundamental
Annotation of Drosophila mojavensis fosmid 8 Priya Srikanth Bio 434W 5.1.2007 Overview High-quality finished sequence is much more useful for research once it is annotated. Annotation is a fundamental
The Alternative Choice of Constitutive Exons throughout Evolution
 The Alternative Choice of Constitutive Exons throughout Evolution Galit Lev-Maor 1[, Amir Goren 1[, Noa Sela 1[, Eddo Kim 1, Hadas Keren 1, Adi Doron-Faigenboim 2, Shelly Leibman-Barak 3, Tal Pupko 2,
The Alternative Choice of Constitutive Exons throughout Evolution Galit Lev-Maor 1[, Amir Goren 1[, Noa Sela 1[, Eddo Kim 1, Hadas Keren 1, Adi Doron-Faigenboim 2, Shelly Leibman-Barak 3, Tal Pupko 2,
Supplementary Figure 1. Generation of knockin mice expressing L-selectinN138G. (a) Schematics of the Sellg allele (top), the targeting vector, the
 Supplementary Figure 1. Generation of knockin mice expressing L-selectinN138G. (a) Schematics of the Sellg allele (top), the targeting vector, the targeted allele in ES cells, and the mutant allele in
Supplementary Figure 1. Generation of knockin mice expressing L-selectinN138G. (a) Schematics of the Sellg allele (top), the targeting vector, the targeted allele in ES cells, and the mutant allele in
SUPPLEMENTARY INFORMATION
 Supplementary Figure 1. Formation of the AA5x. a, Camera lucida drawing of embryo at 48 hours post fertilization (hpf, modified from Kimmel et al. Dev Dyn. 1995 203:253-310). b, Confocal microangiogram
Supplementary Figure 1. Formation of the AA5x. a, Camera lucida drawing of embryo at 48 hours post fertilization (hpf, modified from Kimmel et al. Dev Dyn. 1995 203:253-310). b, Confocal microangiogram
Cross species analysis of genomics data. Computational Prediction of mirnas and their targets
 02-716 Cross species analysis of genomics data Computational Prediction of mirnas and their targets Outline Introduction Brief history mirna Biogenesis Why Computational Methods? Computational Methods
02-716 Cross species analysis of genomics data Computational Prediction of mirnas and their targets Outline Introduction Brief history mirna Biogenesis Why Computational Methods? Computational Methods
Supplemental Materials. for. Conservation of an RNA Regulatory Map between Drosophila and. Mammals
 Supplemental Materials for Conservation of an RNA Regulatory Map between Drosophila and Mammals Angela N. Brooks, Li Yang, Michael O. Duff, Kasper Daniel Hansen, Jung W. Park, Sandrine Dudoit, Steven E.
Supplemental Materials for Conservation of an RNA Regulatory Map between Drosophila and Mammals Angela N. Brooks, Li Yang, Michael O. Duff, Kasper Daniel Hansen, Jung W. Park, Sandrine Dudoit, Steven E.
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
Islets of Langerhans consist mostly of insulin-producing β cells. They will appear to be densely labeled
 Are adult pancreatic beta cells formed by self-duplication or stem cell differentiation? Introduction Researchers have long been interested in how tissues produce and maintain the correct number of cells
Are adult pancreatic beta cells formed by self-duplication or stem cell differentiation? Introduction Researchers have long been interested in how tissues produce and maintain the correct number of cells
Supplementary Figure 1
 S U P P L E M E N TA R Y I N F O R M AT I O N DOI: 10.1038/ncb2896 Supplementary Figure 1 Supplementary Figure 1. Sequence alignment of TERB1 homologs in vertebrates. M. musculus TERB1 was derived from
S U P P L E M E N TA R Y I N F O R M AT I O N DOI: 10.1038/ncb2896 Supplementary Figure 1 Supplementary Figure 1. Sequence alignment of TERB1 homologs in vertebrates. M. musculus TERB1 was derived from
Soluble ADAM33 initiates airway remodeling to promote susceptibility for. Elizabeth R. Davies, Joanne F.C. Kelly, Peter H. Howarth, David I Wilson,
 Revised Suppl. Data: Soluble ADAM33 1 Soluble ADAM33 initiates airway remodeling to promote susceptibility for allergic asthma in early life Elizabeth R. Davies, Joanne F.C. Kelly, Peter H. Howarth, David
Revised Suppl. Data: Soluble ADAM33 1 Soluble ADAM33 initiates airway remodeling to promote susceptibility for allergic asthma in early life Elizabeth R. Davies, Joanne F.C. Kelly, Peter H. Howarth, David
Supplementary Figure 1. IHC and proliferation analysis of pten-deficient mammary tumors
 Wang et al LEGENDS TO SUPPLEMENTARY INFORMATION Supplementary Figure 1. IHC and proliferation analysis of pten-deficient mammary tumors A. Induced expression of estrogen receptor α (ERα) in AME vs PDA
Wang et al LEGENDS TO SUPPLEMENTARY INFORMATION Supplementary Figure 1. IHC and proliferation analysis of pten-deficient mammary tumors A. Induced expression of estrogen receptor α (ERα) in AME vs PDA
VIROLOGY. Engineering Viral Genomes: Retrovirus Vectors
 VIROLOGY Engineering Viral Genomes: Retrovirus Vectors Viral vectors Retrovirus replicative cycle Most mammalian retroviruses use trna PRO, trna Lys3, trna Lys1,2 The partially unfolded trna is annealed
VIROLOGY Engineering Viral Genomes: Retrovirus Vectors Viral vectors Retrovirus replicative cycle Most mammalian retroviruses use trna PRO, trna Lys3, trna Lys1,2 The partially unfolded trna is annealed
HIF-inducible mir-191 promotes migration in breast cancer through complex regulation of TGFβ-signaling in hypoxic microenvironment.
 HIF-inducible mir-9 promotes migration in breast cancer through complex regulation of TGFβ-signaling in hypoxic microenvironment. Neha Nagpal, Hafiz M Ahmad, Shibu Chameettachal3, Durai Sundar, Sourabh
HIF-inducible mir-9 promotes migration in breast cancer through complex regulation of TGFβ-signaling in hypoxic microenvironment. Neha Nagpal, Hafiz M Ahmad, Shibu Chameettachal3, Durai Sundar, Sourabh
Highly Significant Antiviral Activity of HIV-1 LTR-Specific Tre-
 Supporting Information Hauber et al. page 1 Text S1 - Supporting Information Highly Significant Antiviral Activity of HIV-1 LTR-Specific Tre- Recombinase in Humanized Mice Ilona Hauber 1, Helga Hofmann-Sieber
Supporting Information Hauber et al. page 1 Text S1 - Supporting Information Highly Significant Antiviral Activity of HIV-1 LTR-Specific Tre- Recombinase in Humanized Mice Ilona Hauber 1, Helga Hofmann-Sieber
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/393/rs9/dc1 Supplementary Materials for Identification of potential drug targets for tuberous sclerosis complex by synthetic screens combining CRISPR-based knockouts
www.sciencesignaling.org/cgi/content/full/8/393/rs9/dc1 Supplementary Materials for Identification of potential drug targets for tuberous sclerosis complex by synthetic screens combining CRISPR-based knockouts
Identification and characterization of multiple splice variants of Cdc2-like kinase 4 (Clk4)
 Identification and characterization of multiple splice variants of Cdc2-like kinase 4 (Clk4) Vahagn Stepanyan Department of Biological Sciences, Fordham University Abstract: Alternative splicing is an
Identification and characterization of multiple splice variants of Cdc2-like kinase 4 (Clk4) Vahagn Stepanyan Department of Biological Sciences, Fordham University Abstract: Alternative splicing is an
Kerdiles et al - Figure S1
 Kerdiles et al - Figure S1 a b Homo sapiens T B ce ce l ls c l M ls ac r PM oph N ag es Mus musculus Foxo1 PLCγ Supplementary Figure 1 Foxo1 expression pattern is conserved between mouse and human. (a)
Kerdiles et al - Figure S1 a b Homo sapiens T B ce ce l ls c l M ls ac r PM oph N ag es Mus musculus Foxo1 PLCγ Supplementary Figure 1 Foxo1 expression pattern is conserved between mouse and human. (a)
Table S2. Baseline characteristics of the Rotterdam Study for interaction analysis
 Supplementary data Table S1. List of the primers for the cloning of precursor mir-4707 Table S2. Baseline characteristics of the Rotterdam Study for interaction analysis Table S3. List of the primers for
Supplementary data Table S1. List of the primers for the cloning of precursor mir-4707 Table S2. Baseline characteristics of the Rotterdam Study for interaction analysis Table S3. List of the primers for
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project
 Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis. Midbrain Forebrain/ Forebrain/ Hindbrain Spinal cord Hindbrain Hindbrain
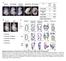 A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis NTD Number of embryos % among NTD Embryos Exencephaly 52 74.3% Craniorachischisis 6 8.6% Spina bifida 5 7.1% Microcephaly 7 1% B Normal
A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis NTD Number of embryos % among NTD Embryos Exencephaly 52 74.3% Craniorachischisis 6 8.6% Spina bifida 5 7.1% Microcephaly 7 1% B Normal
Structural Variation and Medical Genomics
 Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Nature Structural & Molecular Biology: doi: /nsmb.1529
 0.0389 0.0927 0.2641 0.4115 0.3630 0.3618 0.5153 0.3840 0.0357 1.026 0.0866 0.0423 0.1296 0.2052 0.2869 0.0948 0.6285 0.0835 0.0543 0.1135 0.1445 0.1501 0.3272 0.0418 0.4993 0.0624 0.0938 0.1189 0.0768
0.0389 0.0927 0.2641 0.4115 0.3630 0.3618 0.5153 0.3840 0.0357 1.026 0.0866 0.0423 0.1296 0.2052 0.2869 0.0948 0.6285 0.0835 0.0543 0.1135 0.1445 0.1501 0.3272 0.0418 0.4993 0.0624 0.0938 0.1189 0.0768
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
Biology of Cancer Writing Assignment
 1 Biology of Cancer Writing Assignment Based on the information below, you will write a research proposal 3- pages in length (double-spaced, not including references). Draw upon information that you have
1 Biology of Cancer Writing Assignment Based on the information below, you will write a research proposal 3- pages in length (double-spaced, not including references). Draw upon information that you have
CHAPTER 5 RESULTS Previous study: cell culture and organotypical slices
 45 CHAPTER 5 RESULTS 5.1. Previous study: cell culture and organotypical slices Initial experiments have been conducted to ensure that the tet-on system works. A neuronal cell culture from mice expressing
45 CHAPTER 5 RESULTS 5.1. Previous study: cell culture and organotypical slices Initial experiments have been conducted to ensure that the tet-on system works. A neuronal cell culture from mice expressing
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A.
 Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
