The CRISPR/Cas9 system inactivates latent HIV-1 proviral DNA
|
|
|
- Arabella Neal
- 5 years ago
- Views:
Transcription
1 Zhu et al. Retrovirology (2015) 12:22 DOI /s z SHORT REPORT Open Access The CRISPR/Cas9 system inactivates latent HIV-1 proviral DNA Weijun Zhu 1*, Rongyue Lei 1, Yann Le Duff 2,3, Jian Li 1, Fei Guo 1, Mark A Wainberg 2,3 and Chen Liang 2,3* Abstract Background: Highly active antiretroviral therapy (HAART) has transformed HIV-1 infection from a deadly disease to a manageable chronic illness, albeit does not provide a cure. The recently developed genome editing system called CRISPR/Cas9 offers a new tool to inactivate the integrated latent HIV-1 DNA and may serve as a new avenue toward cure. Findings: We tested 10 sites in HIV-1 DNA that can be targeted by CRISPR/Cas9. The engineered CRISPR/Cas9 system was introduced into the JLat10.6 cells that are latently infected by HIV-1. The sequencing results showed that each target site in HIV-1 DNA was efficiently mutated by CRISPR/Cas9 with the target site in the second exon of Rev (called T10) exhibiting the highest degree of mutation. As a result, HIV-1 gene expression and virus production were significantly diminished with T10 causing a 20-fold reduction. Conclusions: The CRISPR/Cas9 complex efficiently mutates and deactivates HIV-1 proviral DNA in latently infected Jurkat cells. Our results also revealed a highly efficient Cas9 target site within the second exon of Rev that represents a promising target to be further explored in the CRISPR/Cas9-based cure strategy. Keywords: HIV-1, Provirus, Reservoir, CRISPR/Cas9, Genome editing Findings Current antiretroviral therapy (ART) enables control of HIV infection at both the individual level and on a global scale. As a result, steady declines have been seen in both numbers of new HIV infections as well as in HIV mortality rates [1,2]. Unfortunately, a cure of HIV infection has not yet been attained. One block to this ultimate goal is the persistence of HIV reservoirs that cannot be cleared by current ART [3]. The establishment of viral reservoirs requires the integration of HIV DNA into cellular genome [4]. Theoretically, deleting or deactivating proviral DNA should eliminate the source of HIV persistence, and may thus provide a valuable tool toward cure. Two approaches have been explored in this context. One is based on the zinc-finger nucleases (ZFNs) that were engineered to recognize and cleave specific HIV DNA sequences [5]. The second approach is called TALENS (transcription * Correspondence: zhuwjun@gmail.com; chen.liang@mcgill.ca 1 MOH Key Laboratory of Systems Biology of Pathogens and AIDS Research Center, Institute of Pathogen Biology, Chinese Academy of Medical Sciences & Peking Union Medical College, Beijing , PR China 2 McGill University AIDS Centre, Lady Davis Institute, Jewish General Hospital, Montreal H3T 1E2, Canada Full list of author information is available at the end of the article activator-like effector nucleases) that program transcription factors to recognize specific DNA sequences [6]. Both approaches exploit the principle of protein-dna recognition that often involves a certain degree of ambiguity. A recent development in genome editing is the CRISPR (Type II microbial clustered regularly interspaced short palindromic repeat)/cas9 system that utilizes a guide RNA strand to recognize and mutate target DNA; this therefore now offers a new strategy to inactivate HIV DNA [7-10]. This CRISPR/Cas9 system arms a DNA endonuclease called Cas9 with an RNA component that utilizes a 20- nucleotide guide sequence (grna) to recognize a specific DNA site [7,8]. Cleavage of DNA by Cas9 generates double-stranded DNA breaks that, upon non-homologous end joining (NHEJ) repair, cause insertions or deletions at the DNA target sites. This approach has been demonstrated highly specific and efficient in targeting multiple cellular genes and has been further developed for genome-wide screens to study gene function [11-14]. In this study, we have tested the efficiency of the CRISPR/ Cas9 system in inactivating the integrated HIV-1 DNA in a HIV-GFP Jurkat cell line called JLat10.6 that was developed to study HIV latency [15] Zhu et al.; licensee BioMed Central. This is an Open Access article distributed under the terms of the Creative Commons Attribution License ( which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly credited. The Creative Commons Public Domain Dedication waiver ( applies to the data made available in this article, unless otherwise stated.
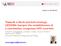
2 Zhu et al. Retrovirology (2015) 12:22 Page 2 of 7 This JLat10.6 HIV-latent cell line harbors the full-length viral DNA that is transcription silent in the absence of external stimulation. Treatment with cytokines such as TNFα activates viral gene expression, which can be measured by monitoring the expression of GFP that has been inserted into HIV-1 DNA as a reporter gene. In order to target HIV-1 DNA, we surveyed the HIV-1 genome and identified more than 80 PAM (protospacer adjacent motif) sites that potentially allow the CRISPR/Cas9 complex to recognize HIV-1 DNA. After considering the conservation of these target DNA sequences across different HIV-1 strains as well as excluding off-target possibilities by blasting the human genome, 10 grnas (named T1 to T10) were selected and inserted into the grna-cloning vector [8]. Of these 10 target sites, 3 are located within the LTR, 5 in the pol gene, and 2 in the second exon of tat/rev (Figure 1). We then nucleotransfected these grna constructs, together with the hcas9 plasmid DNA (expressing the humanized Cas9 enzyme) [8], into JLat10.6 cells. SURVEYOR assay was first performed to measure the frequency of NHEJ as a result of the targeted Cas9 cleavage of HIV-1 DNA. NHEJ was demonstrated for all 10 grnas. The frequency ranged from 10% to 30% (Figure 2A). The targeted viral DNA regions were also amplified by PCR and further examined by sequencing. Both insertions and deletions of various lengths of nucleotides were observed, with the grna T10 causing the most types of deletions and insertions (Figure 2B). These data illustrate the effectiveness of the 10 grnas in targeting Cas9 to cleave and mutate HIV-1 DNA at distinct regions. In order to measure the effects of grna/cas9-induced DNA mutations on the function of integrated HIV-1 DNA, we first transfected the JLat10.6 cells with grna/ Cas9 DNA followed by TNF-α (10 ng/ml) treatment to induce viral gene expression which is monitored by scoring GFP-positive cells by flow cytometry. The results showed that the grna targeting GFP DNA (named T GFP) reduced GFP expression by 5-fold (Figure 3A). No effect was observed for a grna that targets the renilla luciferase (T RL) DNA. The grnas targeting HIV-1 DNA diminished the number of GFP-positive cells to different degrees, ranging from 3-fold (grna T3) to 20-fold (grna T10) (Figure 3A). These grnas alone, without the help of Cas9, exerted no effect on GFP expression (Figure 3B). When levels of HIV-1 in culture supernatants were measured by p24 ELISA, grna T3 led to 3-fold diminution as Figure 1 Illustration of the ten HIV-1 guide RNAs tested in this study. (A) Locations of the 10 guide RNAs (T1 to T10) in HIV-1 genome. (B) Schematic depiction of binding of the T1 grna guide sequence (20 nucleotides, underlined) to the HIV-1 DNA in the context of Cas9 (in yellow). The PAM (protospacer adjacent motif) is highlighted in green letters. The red arrow indicates the cleavage site by Cas9. (C) Sequences of the 10 target sites in the genome HIV-1 strain HXB2 (T1 to T10). Conservation of each target sequence in HIV database is presented. The CRISPR score of each grna was calculated using the program at
3 Zhu et al. Retrovirology (2015) 12:22 Page 3 of 7 Figure 2 Analysis of mutations in HIV-1 DNA caused by CRISPR/Cas9 treatment. (A) Results of the SURVEYOR assays to measure the NHEJ events resulting from HIV-1 guide RNA treatment. Data of two or three independent transfection experiments are shown. The percentage of the mutated DNA (indicated by arrows) is presented at the bottom of each lane. (B) Mutations at each grna target site. The mutated sequences (deletions, insertions and substitutions) are shown in red letters. The PAMs are highlighted in green letters. The wild type HIV-1 DNA is underlined and shown at the top of each aligned sequence panel. compared to 20-fold decrease associated with grnas T4, T8 and T10 (Figure 3C). The grnas alone, in the absence of Cas9, did not affect HIV-1 production (Figure 3D), which further confirms that the grna molecules act on HIV-1 DNA through arming the Cas9 enzyme. Together, these data demonstrate the high efficacy of the CRISPR/Cas9 system in targeting and inactivating HIV-1 proviral DNA. In JLat10.6 cells, without any external stimulation, HIV1 DNA is transcription silent as a result of the inaccessibility of cellular transcriptional machinery to HIV-1 LTR promoter [15,16]. We suspect that this inaccessibility, likely a result of modified chromatin structure, may hinder the ability of the CRISPR/Cas9 complex to target HIV-1 DNA. Were this the case, then pretreatment with HDAC inhibitors such as SAHA may deem necessary to activate HIV-1 gene expression in order to enhance the effectiveness of CRISPR/Cas9 system. We therefore treated the JLat10.6 cells with TNF-α for 16 hours prior to nucleotransfection with the grna and hcas9 plasmid DNA. In contrary to our expectation, activation of HIV-1 transcription by TNF-α did not render HIV-1 DNA more susceptible to inhibition by the CRISPR/Cas9 machinery (Figure 3E and F). This observation suggests that the CRISPR/Cas9 system has the ability to access transcription silent HIV-1 DNA without the need to activate viral gene expression by TNF-α or related agents such as SAHA. This property of the CRISPR/Cas9 system supports its possible utility in eradication of integrated HIV-1 DNA from latently infected immune cells such as CD4+ T cells and macrophages, in which HIV-1 transcription is also inert. Given that these HIV-1 latently-infected cells are often non-cycling, further studies are warranted to assess whether cell cycling may affect the efficacy of CRISPR/Cas9. We next tested whether combinations of different grnas further increased the potency of inhibition. This combination strategy is also expected to diminish the chances of HIV-1 to escape from grna targeting. Indeed, certain combinations, such as T1/T6/T9, T2/T4/T10 and
4 Zhu et al. Retrovirology (2015) 12:22 Page 4 of 7 Figure 3 (See legend on next page.)
5 Zhu et al. Retrovirology (2015) 12:22 Page 5 of 7 (See figure on previous page.) Figure 3 Suppression of HIV-1 gene expression and virus production by grna/cas9. (A, B) Effects of grna/cas9 on GFP expression. JLat10.6 cells were transfected with grna and hcas9 plasmid DNA using Neon (Life Technologies). TNF-α (10 ng/ml) was added 16 hours after to stimulate HIV-1 gene expression from HIV-1 LTR promoter, which was monitored by flow cytometry. A grna targeting the GFP DNA (called T GFP (GTGAACCGCATCGAGCTGAA)) was included as a positive control. The grna targeting renilla luciferase DNA (called T RL (GTAGCGCGGTGTATTATACC)) was utilized as a negative control. Results obtained with the empty grna vector were arbitrarily set as 100%. The results shown are the average of three independent transfection experiments. Experiments were also performed with grna alone (without Cas9) and the results are shown in (B). (C, D) EffectsofgRNA/Cas9onvirusproduction.Levels of viruses in the supernatants were determined by ELISA that measures HIV-1 p24 antigen. Results of three independent transfections are shown. (D) represents the results of experiments performed with grna alone (without Cas9). (E, F) Pretreatment with TNF-α does not increase the inhibition of HIV-1 by grna/cas9. JLat10.6 cells were treated with TNF-α (10 ng/ml) for 16 hours before nucleotransfection with grna and hcas9 plasmid DNA. GFP-positive cells were scored by flow cytometry and the results are shown in (E). Levels of viruses in the supernatants were determined by HIV-1 p24 ELISA and the results shown in (F). Results are the average of three independent transfections. (G, H) Suppression of HIV-1 by multiple grnas that target different HIV-1 DNA sites. Three different grnas were co-transfected with hcas9 plasmid DNA. HIV-1 gene expression (shown in (G)) and virus production (shown in (H)) after TNF-α treatment were measured as described above. Results shown are the average of three independent transfection experiments. Asterisks indicate P <0.05 (Mann Whitney U test, SPSS 16.0). T3/T6/T9, reduced the numbers of GFP-positive cells and the yields of viruses by more than 24-fold (Figure 3G and H). It is noted that all three combinations contain grnas that target tat/rev, which indicates that tat and rev sequences may represent vulnerable sites for CRISPR/ Cas9 targeting. We noted that targeting the pol DNA (T5 to T8) also led to significant reduction in GFP expression that is driven by the LTR and viral Tat protein (Figure 3). One possibility is that Cas9 cleavage of the pol DNA, as guided by T5, T6, T7 or T8, triggers DNA damage response which causes histone modification (such as H3K9me3) and consequent transcription silencing of adjacent genes [17]. To test this, we treated JLat10.6 cells with control guide RNA or the T5 to T8 HIV-1 guide RNA together with Cas9, followed by ChIP (chromatin immunoprecipitation) analysis of histone modification at HIV-1 LTR and GFP regions. Two types of histone modification were studied, H3K9me3 that marks transcription suppression and H3K36me3 that marks transcription activation [18]. The results of Figure 4 show that the LTR DNA or GFP DNA is enriched by more than 2- fold by the anti-h3k9me3 antibodies in the HIV-1 guide RNA treated cells as compared to the cells that were treated with either the empty vector or grna targeting luciferase (T RL). In contrast, neither anti-h3 nor anti- H3K36me3 antibodies enriched HIV-1 LTR DNA or GFP Figure 4 Effect of grna/cas9 treatment on histone modification at HIV-1 LTR. JLat10.6 cells were treated with grnas that target HIV-1 pol gene (T5 to T8). Cells were then harvested and subject to ChIP analysis using control IgG, anti-h3, anti-h3k9me3, or anti-h3k36me3 antibodies. Levels of HIV-1 LTR DNA (shown in (A)) or GFP DNA (shown in (B)) in the immunoprecipitated materials were determined by quantitative PCR. Results shown are the average of three independent experiments. Asterisks indicate P < 0.05 (Mann Whitney U test, SPSS 16.0).
6 Zhu et al. Retrovirology (2015) 12:22 Page 6 of 7 DNAasaresultofHIV-1guideRNAtreatment(Figure4). Interestingly, treatment with the GFP grna (T GFP) led to specific enrichment of GFP DNA associated with anti- H3K9me3 antibody; in contrast, HIV-1 LTR DNA was not significantly enriched (Figure 4). These data suggest that, in addition to generate mutations in the target DNA, the CRISPR/Cas9 machinery also induces H3K9me3 histone modification at the target site, which contributes to transcription suppression. In conclusion, results of this study demonstrate the great potency of the CRISPR/Cas9 system to target and inactivate HIV-1 proviral DNA in the latent JLat10.6 cell line. Since JLat10.6 is a clonal cell line in which HIV-1 is integrated at a specific site in cellular DNA, results in this study cannot conclude whether the CRISRP/Cas9 system may target HIV-1 DNA that is integrated at different cellular DNA loci with different efficiency. Since the CRISPR/Cas9 complex targets both the transcription inert and the transcription active HIV-1 DNA with similar efficiency, CRISPR/Cas9 should be effective in eliminating HIV-1 reservoirs in which HIV-1 exists in a latent state. In support of our data, recent studies reported suppression of the HIV-based vectors using the CRISPR/Cas9 system [19,20]. Both studies tested grnas targeting HIV-1 LTR. We have herein tested 10 target sites across HIV-1 genome and identified one site in the second exon of Rev (called T10) that exhibits the highest level of mutation by Cas9. In a separate study, the CRISPR/Cas9 system was exploited to successfully generate the CCR5Δ32 deletion in pluripotent stem cells with 33% efficiency, as compared to 14% efficiency when the TALENS method was employed [21]. Together, all these findings support the utility of the CRISPR/Cas9 system as a new genome editing approach to eradicate HIV-1. Challenges also exist to target a highly mutable virus like HIV-1 using the CRISPR/Cas9 system whose efficacy is largely dependent on how well the grna matches the target viral DNA sequence. One solution can be a personalized approach in which the grna is designed to match the HIV-1 sequences that are archived in the reservoir of the individual patient. This effort may especially pay off if such an approach can be used to cure infected individuals. A second strategy may involve exploiting multiple grnas to target several relatively conserved sites in the HIV-1 genome in order to maximize efficacy and minimize virus escape. With the CRISPR/Cas9 field quickly evolving, new tools will hopefully emerge to help eradicate HIV in clinical settings. Competing interests The authors declare that they have no competing interests. Authors contributions WZ and CL conceived the study, WZ, RL, JL and YD performed the experiments, WZ, FG and CL analyzed the data, CL, WZ and MW wrote the manuscript. All authors read and approved the final manuscript. Acknowledgments We thank Li li and Yan Xiao for their technical assistance, Dr. George M Church for providing the grna and hcas9 plasmids. This study was supported by the National 12th 5-year project 2012ZX , basic scientific research grant 2012IPB301 from the Institute of Pathogen Biology, 2012C01 and a grant for a distinguished visiting professor from the Chinese Academy of Medical Sciences & Peking Union Medical College, the Program for Changjiang Scholars and Innovative Research Team in University (IRT13007), the Ministry of Science and Technology of China (2012CB911103, 2012ZX and 2013ZX ), as well as the Canadian Institutes of Health Research. Author details 1 MOH Key Laboratory of Systems Biology of Pathogens and AIDS Research Center, Institute of Pathogen Biology, Chinese Academy of Medical Sciences & Peking Union Medical College, Beijing , PR China. 2 McGill University AIDS Centre, Lady Davis Institute, Jewish General Hospital, Montreal H3T 1E2, Canada. 3 Departments of Medicine, Microbiology & Immunology, McGill University, Montreal H3A 2B4, Canada. Received: 25 November 2014 Accepted: 9 February 2015 References 1. Wainberg MA, Zaharatos GJ, Brenner BG. Development of antiretroviral drug resistance. N Engl J Med. 2011;365: Maartens G, Celum C, Lewin SR. HIV infection: epidemiology, pathogenesis, treatment, and prevention. Lancet. 2014;384: Siliciano RF, Greene WC. HIV latency. Cold Spring Harb Perspect Med. 2012;1:a Donahue DA, Wainberg MA. Cellular and molecular mechanisms involved in the establishment of HIV-1 latency. Retrovirology. 2013;10: Qu X, Wang P, Ding D, Li L, Wang H, Ma L, et al. Zinc-finger-nucleases mediate specific and efficient excision of HIV-1 proviral DNA from infected and latently infected human T cells. Nucleic Acids Res. 2013;41: Miller JC, Tan S, Qiao G, Barlow KA, Wang J, Xia DF, et al. A TALE nuclease architecture for efficient genome editing. Nat Biotechnol. 2011;29: Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339: Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, et al. RNA-guided human genome engineering via Cas9. Science. 2013;339: Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science. 2007;315: Grissa I, Vergnaud G, Pourcel C. The CRISPRdb database and tools to display CRISPRs and to generate dictionaries of spacers and repeats. BMC Bioinformatics. 2007;8: Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelsen TS, et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science. 2014;343: Wang T, Wei JJ, Sabatini DM, Lander ES. Genetic screens in human cells using the CRISPR-Cas9 system. Science. 2014;343: Zhou Y, Zhu S, Cai C, Yuan P, Li C, Huang Y, et al. High-throughput screening of a CRISPR/Cas9 library for functional genomics in human cells. Nature. 2014;509: Koike-Yusa H, Li Y, Tan EP, Velasco-Herrera Mdel C, Yusa K. Genome-wide recessive genetic screening in mammalian cells with a lentiviral CRISPR-guide RNA library. Nat Biotechnol. 2014;32: Jordan A, Bisgrove D, Verdin E. HIV reproducibly establishes a latent infection after acute infection of T cells in vitro. EMBO J. 2003;22: Spina CA, Anderson J, Archin NM, Bosque A, Chan J, Famiglietti M, et al. An in-depth comparison of latent HIV-1 reactivation in multiple cell model systems and resting CD4+ T cells from aviremic patients. PLoS Pathog. 2013;9:e Ayrapetov MK, Gursoy-Yuzugullu O, Xu C, Xu Y, Price BD. DNA double-strand breaks promote methylation of histone H3 on lysine 9 and transient formation of repressive chromatin. Proc Natl Acad Sci U S A. 2014;111: Edmunds JW, Mahadevan LC, Clayton AL. Dynamic histone H3 methylation during gene induction: HYPB/Setd2 mediates all H3K36 trimethylation. EMBO J. 2008;27: Ebina H, Misawa N, Kanemura Y, Koyanagi Y. Harnessing the CRISPR/Cas9 system to disrupt latent HIV-1 provirus. Sci Rep. 2013;3:2510.
7 Zhu et al. Retrovirology (2015) 12:22 Page 7 of Hu W, Kaminski R, Yang F, Zhang Y, Cosentino L, Li F, et al. RNA-directed gene editing specifically eradicates latent and prevents new HIV-1 infection. Proc Natl Acad Sci U S A. 2014;111: Ye L, Wang J, Beyer AI, Teque F, Cradick TJ, Qi Z, et al. Seamless modification of wild-type induced pluripotent stem cells to the natural CCR5Delta32 mutation confers resistance to HIV infection. Proc Natl Acad Sci U S A. 2014;4:4. Submit your next manuscript to BioMed Central and take full advantage of: Convenient online submission Thorough peer review No space constraints or color figure charges Immediate publication on acceptance Inclusion in PubMed, CAS, Scopus and Google Scholar Research which is freely available for redistribution Submit your manuscript at
Right view a copy of this license, visit
 Title Harnessing the CRISPR/Cas9 system t provirus. Author(s) Ebina, Hirotaka; Misawa, Naoko; Kan Yoshio Citation Scientific reports (2013), 3 Issue Date 2013-08-26 URL http://hdl.handle.net/2433/178737
Title Harnessing the CRISPR/Cas9 system t provirus. Author(s) Ebina, Hirotaka; Misawa, Naoko; Kan Yoshio Citation Scientific reports (2013), 3 Issue Date 2013-08-26 URL http://hdl.handle.net/2433/178737
Supplementary Information. Supplementary Figure 1
 Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Promises and challenges of gene editing in the age of CRISPR. Neville Sanjana New York Genome Center & NYU Biology
 Promises and challenges of gene editing in the age of CRISPR Neville Sanjana New York Genome Center & NYU Biology We have developed advanced tools to manipulate information stored in books or on computers
Promises and challenges of gene editing in the age of CRISPR Neville Sanjana New York Genome Center & NYU Biology We have developed advanced tools to manipulate information stored in books or on computers
Development of 5 LTR DNA methylation of latent HIV-1 provirus in cell line models and in long-term-infected individuals
 Trejbalová et al. Clinical Epigenetics (2016) 8:19 DOI 10.1186/s13148-016-0185-6 RESEARCH Development of 5 LTR DNA methylation of latent HIV-1 provirus in cell line models and in long-term-infected individuals
Trejbalová et al. Clinical Epigenetics (2016) 8:19 DOI 10.1186/s13148-016-0185-6 RESEARCH Development of 5 LTR DNA methylation of latent HIV-1 provirus in cell line models and in long-term-infected individuals
UNDERSTANDING THE ROLES OF NUCLEAR RECEPTORS IN THE MAINTENANCE OF HIV PROVIRAL LATENCY USING NOVEL GENE EDITING TECHONOLOGY STEPHANIE C.
 UNDERSTANDING THE ROLES OF NUCLEAR RECEPTORS IN THE MAINTENANCE OF HIV PROVIRAL LATENCY USING NOVEL GENE EDITING TECHONOLOGY By STEPHANIE C. MILNE Submitted in partial fulfillment of the requirements for
UNDERSTANDING THE ROLES OF NUCLEAR RECEPTORS IN THE MAINTENANCE OF HIV PROVIRAL LATENCY USING NOVEL GENE EDITING TECHONOLOGY By STEPHANIE C. MILNE Submitted in partial fulfillment of the requirements for
Tel: ; Fax: ;
 Tel.: +98 216 696 9291; Fax: +98 216 696 9291; E-mail: mrasadeghi@pasteur.ac.ir Tel: +98 916 113 7679; Fax: +98 613 333 6380; E-mail: abakhshi_e@ajums.ac.ir A Soluble Chromatin-bound MOI 0 1 5 0 1 5 HDAC2
Tel.: +98 216 696 9291; Fax: +98 216 696 9291; E-mail: mrasadeghi@pasteur.ac.ir Tel: +98 916 113 7679; Fax: +98 613 333 6380; E-mail: abakhshi_e@ajums.ac.ir A Soluble Chromatin-bound MOI 0 1 5 0 1 5 HDAC2
Towards an HIV Cure. Steven G. Deeks Professor of Medicine University of California, San Francisco
 Towards an HIV Cure Steven G. Deeks Professor of Medicine University of California, San Francisco Why are we now talking about a cure? Emerging recognition that HAART does not fully restore health and/or
Towards an HIV Cure Steven G. Deeks Professor of Medicine University of California, San Francisco Why are we now talking about a cure? Emerging recognition that HAART does not fully restore health and/or
Host Double Strand Break Repair Generates HIV-1 Strains Resistant to CRISPR/Cas9
 Host Double Strand Break Repair Generates HIV-1 Strains Resistant to CRISPR/Cas9 Kristine E. Yoder, a * and Ralf Bundschuh b a Department of Molecular Virology, Immunology and Medical Genetics, Center
Host Double Strand Break Repair Generates HIV-1 Strains Resistant to CRISPR/Cas9 Kristine E. Yoder, a * and Ralf Bundschuh b a Department of Molecular Virology, Immunology and Medical Genetics, Center
cure research HIV & AIDS
 Glossary of terms HIV & AIDS cure research Antiretroviral Therapy (ART) ART involves the use of several (usually a cocktail of three or more) antiretroviral drugs to halt HIV replication. ART drugs may
Glossary of terms HIV & AIDS cure research Antiretroviral Therapy (ART) ART involves the use of several (usually a cocktail of three or more) antiretroviral drugs to halt HIV replication. ART drugs may
The updated incidences and mortalities of major cancers in China, 2011
 DOI 10.1186/s40880-015-0042-6 REVIEW Open Access The updated incidences and mortalities of major cancers in China, 2011 Wanqing Chen *, Rongshou Zheng, Hongmei Zeng and Siwei Zhang Abstract Introduction:
DOI 10.1186/s40880-015-0042-6 REVIEW Open Access The updated incidences and mortalities of major cancers in China, 2011 Wanqing Chen *, Rongshou Zheng, Hongmei Zeng and Siwei Zhang Abstract Introduction:
DNA context and promoter activity affect gene expression in lentiviral vectors
 ACTA BIOMED 2008; 79: 192-196 Mattioli 1885 O R I G I N A L A R T I C L E DNA context and promoter activity affect gene expression in lentiviral vectors Gensheng Mao 1, Francesco Marotta 2, Jia Yu 3, Liang
ACTA BIOMED 2008; 79: 192-196 Mattioli 1885 O R I G I N A L A R T I C L E DNA context and promoter activity affect gene expression in lentiviral vectors Gensheng Mao 1, Francesco Marotta 2, Jia Yu 3, Liang
Oncolytic Viruses as a Potential Approach to Eliminate the HIV Reservoir
 Oncolytic Viruses as a Potential Approach to Eliminate the HIV Reservoir Costiniuk CT, Côté SC, Al-Ghazawi FM, Carrasco-Medina L,Young CD, & Angel JB University of Ottawa, November 12 th, 2012 HIV Reservoirs
Oncolytic Viruses as a Potential Approach to Eliminate the HIV Reservoir Costiniuk CT, Côté SC, Al-Ghazawi FM, Carrasco-Medina L,Young CD, & Angel JB University of Ottawa, November 12 th, 2012 HIV Reservoirs
CRISPR/CAS9 based high-throughput screening. Journal club Caihong Zhu
 CRISPR/CAS9 based high-throughput screening Journal club Caihong Zhu 29.04.2014 High Throughput Screening (HTS) HTS is a method for scientific experimentation especially used in drug discovery and relevant
CRISPR/CAS9 based high-throughput screening Journal club Caihong Zhu 29.04.2014 High Throughput Screening (HTS) HTS is a method for scientific experimentation especially used in drug discovery and relevant
Professor Jonathan Weber
 HIV/AIDS at 30: Back to the Future BHIVA / Wellcome Trust Multidisciplinary Event to mark World AIDS Day 2011 British HIV Association (BHIVA) 2011 HIV/AIDS at 30: Back to the Future BHIVA / Wellcome Trust
HIV/AIDS at 30: Back to the Future BHIVA / Wellcome Trust Multidisciplinary Event to mark World AIDS Day 2011 British HIV Association (BHIVA) 2011 HIV/AIDS at 30: Back to the Future BHIVA / Wellcome Trust
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
CRISPR/Cas9 cleavage of viral DNA efficiently suppresses hepatitis B virus
 0 0 CRISPR/Cas cleavage of viral DNA efficiently suppresses hepatitis B virus Authors: Vyas Ramanan,, Amir Shlomai,, David B.T. Cox,,,, Robert E. Schwartz,,, Eleftherios Michailidis, Ankit Bhatta, David
0 0 CRISPR/Cas cleavage of viral DNA efficiently suppresses hepatitis B virus Authors: Vyas Ramanan,, Amir Shlomai,, David B.T. Cox,,,, Robert E. Schwartz,,, Eleftherios Michailidis, Ankit Bhatta, David
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Choosing Between Lentivirus and Adeno-associated Virus For DNA Delivery
 Choosing Between Lentivirus and Adeno-associated Virus For DNA Delivery Presenter: April 12, 2017 Ed Davis, Ph.D. Senior Application Scientist GeneCopoeia, Inc. Outline Introduction to GeneCopoeia Lentiviral
Choosing Between Lentivirus and Adeno-associated Virus For DNA Delivery Presenter: April 12, 2017 Ed Davis, Ph.D. Senior Application Scientist GeneCopoeia, Inc. Outline Introduction to GeneCopoeia Lentiviral
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Specific Reactivation of Latent HIV-1 by dcas9-suntag-vp64-mediated Guide RNA Targeting the HIV-1 Promoter
 original article The American Society of Gene & Cell Therapy Specific Reactivation of Latent HIV- by dcas9-suntag-vp64-mediated Guide RNA Targeting the HIV- Promoter Haiyan Ji, Zhengtao Jiang, Panpan Lu,
original article The American Society of Gene & Cell Therapy Specific Reactivation of Latent HIV- by dcas9-suntag-vp64-mediated Guide RNA Targeting the HIV- Promoter Haiyan Ji, Zhengtao Jiang, Panpan Lu,
The HIV life cycle. integration. virus production. entry. transcription. reverse transcription. nuclear import
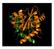 The HIV life cycle entry reverse transcription transcription integration virus production nuclear import Hazuda 2012 Integration Insertion of the viral DNA into host chromosomal DNA, essential step in
The HIV life cycle entry reverse transcription transcription integration virus production nuclear import Hazuda 2012 Integration Insertion of the viral DNA into host chromosomal DNA, essential step in
CRISPRaTest Functional dcas9-activator Assay Kit v1 Last update: 2018/07/04 Cellecta, Inc.
 CRISPRaTest Functional dcas9-activator Assay Kit v1 Last update: 2018/07/04 Cellecta, Inc. Copyright (c) 2018 Cellecta, Inc. All Rights Reserved. Table of Contents 1. CRISPRaTest Functional dcas9-activator
CRISPRaTest Functional dcas9-activator Assay Kit v1 Last update: 2018/07/04 Cellecta, Inc. Copyright (c) 2018 Cellecta, Inc. All Rights Reserved. Table of Contents 1. CRISPRaTest Functional dcas9-activator
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
HIV INFECTION: An Overview
 HIV INFECTION: An Overview UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL MBBS II SEMINAR VJ
HIV INFECTION: An Overview UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL MBBS II SEMINAR VJ
7.012 Quiz 3 Answers
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
Approaching a Cure Daniel R. Kuritzkes, MD
 Approaching a Cure Daniel R. Kuritzkes, MD Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Disclosures The speaker is a consultant and/or has received speaking honoraria
Approaching a Cure Daniel R. Kuritzkes, MD Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Disclosures The speaker is a consultant and/or has received speaking honoraria
IAS 2015 Towards an HIV Cure symposium Vancouver Immune recognition following latency reversal
 IAS 2015 Towards an HIV Cure symposium Vancouver Immune recognition following latency reversal Marcus Altfeld Professor of Medicine Outline Immune recognition of HIV-1-infected cells Kinetics of antigen
IAS 2015 Towards an HIV Cure symposium Vancouver Immune recognition following latency reversal Marcus Altfeld Professor of Medicine Outline Immune recognition of HIV-1-infected cells Kinetics of antigen
Citation for published version (APA): Von Eije, K. J. (2009). RNAi based gene therapy for HIV-1, from bench to bedside
 UvA-DARE (Digital Academic Repository) RNAi based gene therapy for HIV-1, from bench to bedside Von Eije, K.J. Link to publication Citation for published version (APA): Von Eije, K. J. (2009). RNAi based
UvA-DARE (Digital Academic Repository) RNAi based gene therapy for HIV-1, from bench to bedside Von Eije, K.J. Link to publication Citation for published version (APA): Von Eije, K. J. (2009). RNAi based
About OMICS Group Conferences
 About OMICS Group OMICS Group International is an amalgamation ofopen Access publicationsand worldwide international science conferences and events. Established in the year 2007 with the sole aim of making
About OMICS Group OMICS Group International is an amalgamation ofopen Access publicationsand worldwide international science conferences and events. Established in the year 2007 with the sole aim of making
Relationship between incident types and impact on patients in drug name errors: a correlational study
 Tsuji et al. Journal of Pharmaceutical Health Care and Sciences (2015) 1:11 DOI 10.1186/s40780-015-0011-x RESEARCH ARTICLE Open Access Relationship between incident types and impact on patients in drug
Tsuji et al. Journal of Pharmaceutical Health Care and Sciences (2015) 1:11 DOI 10.1186/s40780-015-0011-x RESEARCH ARTICLE Open Access Relationship between incident types and impact on patients in drug
Regulation of cell signaling cascades by influenza A virus
 The Second International Symposium on Optimization and Systems Biology (OSB 08) Lijiang, China, October 31 November 3, 2008 Copyright 2008 ORSC & APORC, pp. 389 394 Regulation of cell signaling cascades
The Second International Symposium on Optimization and Systems Biology (OSB 08) Lijiang, China, October 31 November 3, 2008 Copyright 2008 ORSC & APORC, pp. 389 394 Regulation of cell signaling cascades
HIV & AIDS: Overview
 HIV & AIDS: Overview UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL SEMINAR VJ TEMPLE 1 What
HIV & AIDS: Overview UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL SEMINAR VJ TEMPLE 1 What
Repressive Transcription
 Repressive Transcription The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Guenther, M. G., and R. A.
Repressive Transcription The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Guenther, M. G., and R. A.
I declare that I have no financial conflicts of interest
 I declare that I have no financial conflicts of interest Cytotoxic T-Lymphocytes Eliminate Defective HIV Proviruses Without Impacting Infectious Reservoirs R. Brad Jones Assistant Professor The George
I declare that I have no financial conflicts of interest Cytotoxic T-Lymphocytes Eliminate Defective HIV Proviruses Without Impacting Infectious Reservoirs R. Brad Jones Assistant Professor The George
Compounds producing an effective combinatorial regimen for disruption of HIV-1 latency
 Research Article Compounds producing an effective combinatorial regimen for disruption of HIV-1 latency Pargol Hashemi 1, Kris Barreto 1, Wendy Bernhard 1, Adam Lomness 1, Nicolette Honson 2, Tom A Pfeifer
Research Article Compounds producing an effective combinatorial regimen for disruption of HIV-1 latency Pargol Hashemi 1, Kris Barreto 1, Wendy Bernhard 1, Adam Lomness 1, Nicolette Honson 2, Tom A Pfeifer
Chronic viral infections cause enormous suffering among infected
 MINIREVIEW Targeted DNA Mutagenesis for the Cure of Chronic Viral Infections Joshua T. Schiffer, a,b Martine Aubert, a Nicholas D. Weber, a,c Esther Mintzer, a,d Daniel Stone, a and Keith R. Jerome a,c
MINIREVIEW Targeted DNA Mutagenesis for the Cure of Chronic Viral Infections Joshua T. Schiffer, a,b Martine Aubert, a Nicholas D. Weber, a,c Esther Mintzer, a,d Daniel Stone, a and Keith R. Jerome a,c
RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
 Xu et al. Molecular Cancer (2019) 18:8 https://doi.org/10.1186/s12943-018-0932-8 LETTER TO THE EDITOR RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
Xu et al. Molecular Cancer (2019) 18:8 https://doi.org/10.1186/s12943-018-0932-8 LETTER TO THE EDITOR RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
Painting a specific chromosome with CRISPR/Cas9 for live-cell imaging
 Painting a specific chromosome with CRISPR/Cas9 for live-cell imaging The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation
Painting a specific chromosome with CRISPR/Cas9 for live-cell imaging The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation
Recent Insights into HIV Pathogenesis and Treatment: Towards a Cure
 Recent Insights into HIV Pathogenesis and Treatment: Towards a Cure Deborah Persaud, M.D. Associate Professor Department of Pediatrics Division of Infectious Diseases Johns Hopkins University School of
Recent Insights into HIV Pathogenesis and Treatment: Towards a Cure Deborah Persaud, M.D. Associate Professor Department of Pediatrics Division of Infectious Diseases Johns Hopkins University School of
Potential Role of Human Endogenous Retrovirus K102 (HERV-K102) Particles in Resistance to HIV-1 Transmission
 Potential Role of Human Endogenous Retrovirus K102 (HERV-K102) Particles in Resistance to HIV-1 Transmission M. Laderoute, L. Larocque, A. Giulivi, K.R. Fowke, F.A. Plummer & F. Diaz-Mitoma As presented
Potential Role of Human Endogenous Retrovirus K102 (HERV-K102) Particles in Resistance to HIV-1 Transmission M. Laderoute, L. Larocque, A. Giulivi, K.R. Fowke, F.A. Plummer & F. Diaz-Mitoma As presented
Towards a block and lock strategy: LEDGINs hamper the establishment of a reactivation competent HIV reservoir.
 Abstract no. MOLBPEA13 Towards a block and lock strategy: LEDGINs hamper the establishment of a reactivation competent HIV reservoir. G. Vansant,,A. Bruggemans, L. Vranckx, S. Saleh, I. Zurnic, F. Christ,
Abstract no. MOLBPEA13 Towards a block and lock strategy: LEDGINs hamper the establishment of a reactivation competent HIV reservoir. G. Vansant,,A. Bruggemans, L. Vranckx, S. Saleh, I. Zurnic, F. Christ,
Supplementary information. MARCH8 inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins
 Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
HIV 101: Fundamentals of HIV Infection
 HIV 101: Fundamentals of HIV Infection David H. Spach, MD Professor of Medicine University of Washington Seattle, Washington Learning Objectives After attending this presentation, learners will be able
HIV 101: Fundamentals of HIV Infection David H. Spach, MD Professor of Medicine University of Washington Seattle, Washington Learning Objectives After attending this presentation, learners will be able
Impact of Vorinostat Treatment of Non- Hodgkin s Lymphoma on HIV-1 Latent Reservoir
 Impact of Vorinostat Treatment of Non- Hodgkin s Lymphoma on HIV-1 Latent Reservoir Adam Capoferri, Juan Carlos Ramos, Daniel Xu, Daniel I.S. Rosenbloom, Janet D. Siliciano, Robert F. Siliciano, Ariela
Impact of Vorinostat Treatment of Non- Hodgkin s Lymphoma on HIV-1 Latent Reservoir Adam Capoferri, Juan Carlos Ramos, Daniel Xu, Daniel I.S. Rosenbloom, Janet D. Siliciano, Robert F. Siliciano, Ariela
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP
 Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP 1 Learning Objectives Recognize hazards associated with viral vectors in research and animal
Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP 1 Learning Objectives Recognize hazards associated with viral vectors in research and animal
Control of Stochastic Gene Expression by Host Factors at the HIV Promoter
 Control of Stochastic Gene Expression by Host Factors at the HIV Promoter John C. Burnett 1, Kathryn Miller-Jensen 1, Priya S. Shah 1, Adam P. Arkin 2,3 *, David V. Schaffer 1,2,3 * 1 Department of Chemical
Control of Stochastic Gene Expression by Host Factors at the HIV Promoter John C. Burnett 1, Kathryn Miller-Jensen 1, Priya S. Shah 1, Adam P. Arkin 2,3 *, David V. Schaffer 1,2,3 * 1 Department of Chemical
1. Engineering Foot-and-Mouth Disease Viruses with Improved
 Engineering Foot-and-Mouth Disease Virus with Improved Properties for the Development of Effective Vaccine Haixue Zheng, Fan Yang, Ye Jin, Jianhong Guo, Jijun He, Lvlv, Xuepeng Cai, Xiangtao Liu, Hong
Engineering Foot-and-Mouth Disease Virus with Improved Properties for the Development of Effective Vaccine Haixue Zheng, Fan Yang, Ye Jin, Jianhong Guo, Jijun He, Lvlv, Xuepeng Cai, Xiangtao Liu, Hong
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
The Canadian contribution to the otolaryngology literature: a five year bibliometric analysis
 Gurberg et al. Journal of Otolaryngology - Head and Neck Surgery 2014, 43:47 ORIGINAL RESEARCH ARTICLE Open Access The Canadian contribution to the otolaryngology literature: a five year bibliometric analysis
Gurberg et al. Journal of Otolaryngology - Head and Neck Surgery 2014, 43:47 ORIGINAL RESEARCH ARTICLE Open Access The Canadian contribution to the otolaryngology literature: a five year bibliometric analysis
Immunodeficiency. (2 of 2)
 Immunodeficiency (2 of 2) Acquired (secondary) immunodeficiencies More common Many causes such as therapy, cancer, sarcoidosis, malnutrition, infection & renal disease The most common of which is therapy-related
Immunodeficiency (2 of 2) Acquired (secondary) immunodeficiencies More common Many causes such as therapy, cancer, sarcoidosis, malnutrition, infection & renal disease The most common of which is therapy-related
Li et al. Journal of Experimental & Clinical Cancer Research (2018) 37:108
 Li et al. Journal of Experimental & Clinical Cancer Research (2018) 37:108 https://doi.org/10.1186/s13046-018-0774-7 CORRECTION Correction to: Novel smac mimetic APG- 1387 elicits ovarian cancer cell killing
Li et al. Journal of Experimental & Clinical Cancer Research (2018) 37:108 https://doi.org/10.1186/s13046-018-0774-7 CORRECTION Correction to: Novel smac mimetic APG- 1387 elicits ovarian cancer cell killing
Supplemental Information. Augmentation of Antitumor Immunity by Human. and Mouse CAR T Cells Secreting IL-18
 Cell Reports, Volume 20 Supplemental Information Augmentation of Antitumor Immunity by Human and Mouse CAR T Cells Secreting IL-18 Biliang Hu, Jiangtao Ren, Yanping Luo, Brian Keith, Regina M. Young, John
Cell Reports, Volume 20 Supplemental Information Augmentation of Antitumor Immunity by Human and Mouse CAR T Cells Secreting IL-18 Biliang Hu, Jiangtao Ren, Yanping Luo, Brian Keith, Regina M. Young, John
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Histones modifications and variants
 Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
HIV Cure Update. Christine Durand, MD 14 de abril de 2016, XIII Conferência Brasil Johns Hopkins University em HIV/AIDS
 HIV Cure Update Christine Durand, MD 14 de abril de 2016, XIII Conferência Brasil Johns Hopkins University em HIV/AIDS Financial Disclosures Research grants paid to my institution: Gilead Sciences, Bristol
HIV Cure Update Christine Durand, MD 14 de abril de 2016, XIII Conferência Brasil Johns Hopkins University em HIV/AIDS Financial Disclosures Research grants paid to my institution: Gilead Sciences, Bristol
With over 20 drugs and several viable regimens, the mo6vated pa6ent with life- long access to therapy can control HIV indefinitely, elimina6ng the
 Towards an HIV Cure Steven G. Deeks, MD Professor of Medicine in Residence HIV/AIDS Division University of California, San Francisco (UCSF) WorldMedSchool; July 22, 2013 1 With over 20 drugs and several
Towards an HIV Cure Steven G. Deeks, MD Professor of Medicine in Residence HIV/AIDS Division University of California, San Francisco (UCSF) WorldMedSchool; July 22, 2013 1 With over 20 drugs and several
IAS 2013 Towards an HIV Cure Symposium
 Entinostat is a histone deacetylase (HDAC) inhibitor selective for class 1 HDACs and activates HIV production from latently infected primary T-cells Fiona Wightman Monash University, Melbourne, Australia
Entinostat is a histone deacetylase (HDAC) inhibitor selective for class 1 HDACs and activates HIV production from latently infected primary T-cells Fiona Wightman Monash University, Melbourne, Australia
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Human Immunodeficiency Virus
 Human Immunodeficiency Virus Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Viruses and hosts Lentivirus from Latin lentis (slow), for slow progression of disease
Human Immunodeficiency Virus Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Viruses and hosts Lentivirus from Latin lentis (slow), for slow progression of disease
HIV Remission: Compound screening and optimization. Michael D. Miller
 HIV Remission: Compound screening and optimization Michael D. Miller Can we drive HIV into remission in large numbers of patients? "George Bernard Shaw, speaking as an Irishman, summed up an approach to
HIV Remission: Compound screening and optimization Michael D. Miller Can we drive HIV into remission in large numbers of patients? "George Bernard Shaw, speaking as an Irishman, summed up an approach to
Engineering Foot-and-Mouth Disease Virus with Improved Properties for the Development of Effective Vaccine Introduction: Materials and methods:
 Engineering Foot-and-Mouth Disease Virus with Improved Properties for the Development of Effective Vaccine Haixue Zheng, Fan Yang, Ye Jin, Jianhong Guo, Jijun He, Kaiqi Lian, Zixiang Zhu, Weijun Cao, Lvlv,
Engineering Foot-and-Mouth Disease Virus with Improved Properties for the Development of Effective Vaccine Haixue Zheng, Fan Yang, Ye Jin, Jianhong Guo, Jijun He, Kaiqi Lian, Zixiang Zhu, Weijun Cao, Lvlv,
Egr-1 regulates RTA transcription through a cooperative involvement of transcriptional regulators
 /, 2017, Vol. 8, (No. 53), pp: 91425-91444 Egr-1 regulates RTA transcription through a cooperative involvement of transcriptional regulators Roni Sarkar 1 and Subhash C. Verma 1 1 Department of Microbiology
/, 2017, Vol. 8, (No. 53), pp: 91425-91444 Egr-1 regulates RTA transcription through a cooperative involvement of transcriptional regulators Roni Sarkar 1 and Subhash C. Verma 1 1 Department of Microbiology
IDENTIFICATION OF BIOMARKERS FOR EARLY DIAGNOSIS OF BREAST CANCER"
 IDENTIFICATION OF BIOMARKERS FOR EARLY DIAGNOSIS OF BREAST CANCER" Edmond Marzbani, MD December 2, 2008 Early diagnosis of cancer Many solid tumors are potentially curable if diagnosed at an early stage
IDENTIFICATION OF BIOMARKERS FOR EARLY DIAGNOSIS OF BREAST CANCER" Edmond Marzbani, MD December 2, 2008 Early diagnosis of cancer Many solid tumors are potentially curable if diagnosed at an early stage
Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases
 Brief Communication Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases Qinghai Zeng 1 *, Cuihong Jin 2 *, Wenhang Chen 2, Fang Xia 3, Qi Wang 3, Fan Fan 4,
Brief Communication Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases Qinghai Zeng 1 *, Cuihong Jin 2 *, Wenhang Chen 2, Fang Xia 3, Qi Wang 3, Fan Fan 4,
The relation between HIV- 1 integration and latency
 The relation between HIV- 1 integration and latency Linos Vandekerckhove HIV translational research, Ghent Department of General Internal Medicine, Infectious Diseases and Psychosomatic Medicine University
The relation between HIV- 1 integration and latency Linos Vandekerckhove HIV translational research, Ghent Department of General Internal Medicine, Infectious Diseases and Psychosomatic Medicine University
Combined use of AFP, CEA, CA125 and CAl9-9 improves the sensitivity for the diagnosis of gastric cancer
 He et al. BMC Gastroenterology 2013, 13:87 RESEARCH ARTICLE Open Access Combined use of AFP, CEA, CA125 and CAl9-9 improves the sensitivity for the diagnosis of gastric cancer Chao-Zhu He 1,2, Kun-He Zhang
He et al. BMC Gastroenterology 2013, 13:87 RESEARCH ARTICLE Open Access Combined use of AFP, CEA, CA125 and CAl9-9 improves the sensitivity for the diagnosis of gastric cancer Chao-Zhu He 1,2, Kun-He Zhang
Chapter 7 Conclusions
 VII-1 Chapter 7 Conclusions VII-2 The development of cell-based therapies ranging from well-established practices such as bone marrow transplant to next-generation strategies such as adoptive T-cell therapy
VII-1 Chapter 7 Conclusions VII-2 The development of cell-based therapies ranging from well-established practices such as bone marrow transplant to next-generation strategies such as adoptive T-cell therapy
TITLE: Influenza A (H7N9) virus evolution: Which genetic mutations are antigenically important?
 TITLE: Influenza A (H7N9) virus evolution: Which genetic mutations are antigenically important? AUTHORS: Joshua G. Petrie 1, Adam S. Lauring 2,3 AFFILIATIONS: 1 Department of Epidemiology, University of
TITLE: Influenza A (H7N9) virus evolution: Which genetic mutations are antigenically important? AUTHORS: Joshua G. Petrie 1, Adam S. Lauring 2,3 AFFILIATIONS: 1 Department of Epidemiology, University of
Lecture 2: Virology. I. Background
 Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
HIV transcription, Tat transactivation mrna processing and latency
 HIV transcription, Tat transactivation mrna processing and latency Tat transactivation Roles of CTD kinases in HIV replication Latency and reservoir Strategies to eliminate the reservoir of HIV in the
HIV transcription, Tat transactivation mrna processing and latency Tat transactivation Roles of CTD kinases in HIV replication Latency and reservoir Strategies to eliminate the reservoir of HIV in the
Lecture 11. Immunology and disease: parasite antigenic diversity
 Lecture 11 Immunology and disease: parasite antigenic diversity RNAi interference video and tutorial (you are responsible for this material, so check it out.) http://www.pbs.org/wgbh/nova/sciencenow/3210/02.html
Lecture 11 Immunology and disease: parasite antigenic diversity RNAi interference video and tutorial (you are responsible for this material, so check it out.) http://www.pbs.org/wgbh/nova/sciencenow/3210/02.html
Clinical Significance of Human Immunodeficiency Virus Type 1 Replication Fitness
 CLINICAL MICROBIOLOGY REVIEWS, Oct. 2007, p. 550 578 Vol. 20, No. 4 0893-8512/07/$08.00 0 doi:10.1128/cmr.00017-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Clinical Significance
CLINICAL MICROBIOLOGY REVIEWS, Oct. 2007, p. 550 578 Vol. 20, No. 4 0893-8512/07/$08.00 0 doi:10.1128/cmr.00017-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Clinical Significance
Fig. 1: Schematic diagram of basic structure of HIV
 UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL SEMINAR HIV & AIDS: An Overview What is HIV?
UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL SEMINAR HIV & AIDS: An Overview What is HIV?
Recombinant Protein Expression Retroviral system
 Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
CRISPR-mediated Editing of Hematopoietic Stem Cells for the Treatment of β-hemoglobinopathies
 CRISPR-mediated Editing of Hematopoietic Stem Cells for the Treatment of β-hemoglobinopathies Jennifer Gori American Society of Gene & Cell Therapy May 11, 2017 editasmedicine.com 1 Highlights Developed
CRISPR-mediated Editing of Hematopoietic Stem Cells for the Treatment of β-hemoglobinopathies Jennifer Gori American Society of Gene & Cell Therapy May 11, 2017 editasmedicine.com 1 Highlights Developed
Supplemental Figure 1
 Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data
 Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier ChIP-seq experiment To
Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier ChIP-seq experiment To
Eukaryotic Gene Regulation
 Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Comparison of latent HIV-1 reactivation in multiple cell model systems
 Comparison of latent HIV-1 reactivation in multiple cell model systems Celsa A. Spina, Jenny Anderson, Nancie M. Archin, Alberto Bosque, Jonathan Chan, Marylinda Famiglietti, Warner C. Greene, Angela Kashuba,
Comparison of latent HIV-1 reactivation in multiple cell model systems Celsa A. Spina, Jenny Anderson, Nancie M. Archin, Alberto Bosque, Jonathan Chan, Marylinda Famiglietti, Warner C. Greene, Angela Kashuba,
Effectiveness of the surgical torque limiter: a model comparing drill- and hand-based screw insertion into locking plates
 Ioannou et al. Journal of Orthopaedic Surgery and Research (2016) 11:118 DOI 10.1186/s13018-016-0458-y RESEARCH ARTICLE Effectiveness of the surgical torque limiter: a model comparing drill- and hand-based
Ioannou et al. Journal of Orthopaedic Surgery and Research (2016) 11:118 DOI 10.1186/s13018-016-0458-y RESEARCH ARTICLE Effectiveness of the surgical torque limiter: a model comparing drill- and hand-based
Therapeutic Applications and Specificity of Action of Designer Nucleases for Precision Genome Engineering
 University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 1-1-2015 Therapeutic Applications and Specificity of Action of Designer Nucleases for Precision Genome Engineering Chukwuka
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 1-1-2015 Therapeutic Applications and Specificity of Action of Designer Nucleases for Precision Genome Engineering Chukwuka
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Therapeutic Genome Editing with CRISPR-Cas and other programmable gene scissors
 Brussels, April 12 th 2018 Therapeutic Genome Editing with CRISPR-Cas and other programmable gene scissors Toni Cathomen Gene Therapy returns to Center Stage Engineering the Genome Jameson Golliday (source:
Brussels, April 12 th 2018 Therapeutic Genome Editing with CRISPR-Cas and other programmable gene scissors Toni Cathomen Gene Therapy returns to Center Stage Engineering the Genome Jameson Golliday (source:
CRISPR/Cas9-mediated genome editing induces exon skipping by complete or stochastic altering splicing in the migratory locust
 Chen et al. BMC Biotechnology (2018) 18:60 https://doi.org/10.1186/s12896-018-0465-7 METHODOLOGY ARTICLE Open Access CRISPR/Cas9-mediated genome editing induces exon skipping by complete or stochastic
Chen et al. BMC Biotechnology (2018) 18:60 https://doi.org/10.1186/s12896-018-0465-7 METHODOLOGY ARTICLE Open Access CRISPR/Cas9-mediated genome editing induces exon skipping by complete or stochastic
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Part-4. Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death
 Part-4 Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death 95 1. Introduction The process of replicating DNA and dividing cells can be described as a series of coordinated
Part-4 Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death 95 1. Introduction The process of replicating DNA and dividing cells can be described as a series of coordinated
CRISPR way to cut genes speeds advances in autism
 NEWS CRISPR way to cut genes speeds advances in autism BY JESSICA WRIGHT 14 DECEMBER 2015 Less than three years ago, two landmark publications in Science gave researchers a quick and easy recipe for tinkering
NEWS CRISPR way to cut genes speeds advances in autism BY JESSICA WRIGHT 14 DECEMBER 2015 Less than three years ago, two landmark publications in Science gave researchers a quick and easy recipe for tinkering
Mutants and HBV vaccination. Dr. Ulus Salih Akarca Ege University, Izmir, Turkey
 Mutants and HBV vaccination Dr. Ulus Salih Akarca Ege University, Izmir, Turkey Geographic Distribution of Chronic HBV Infection 400 million people are carrier of HBV Leading cause of cirrhosis and HCC
Mutants and HBV vaccination Dr. Ulus Salih Akarca Ege University, Izmir, Turkey Geographic Distribution of Chronic HBV Infection 400 million people are carrier of HBV Leading cause of cirrhosis and HCC
Palindromes drive the re-assortment in Influenza A
 www.bioinformation.net Hypothesis Volume 7(3) Palindromes drive the re-assortment in Influenza A Abdullah Zubaer 1, 2 * & Simrika Thapa 1, 2 1Swapnojaatra Bioresearch Laboratory, DataSoft Systems, Dhaka-15,
www.bioinformation.net Hypothesis Volume 7(3) Palindromes drive the re-assortment in Influenza A Abdullah Zubaer 1, 2 * & Simrika Thapa 1, 2 1Swapnojaatra Bioresearch Laboratory, DataSoft Systems, Dhaka-15,
Optimizing sgrna structure to improve CRISPR-Cas9 knockout efficiency
 Dang et al. Genome Biology (215) 16:28 DOI 1.1186/s1359-15-846-3 RESEARCH Open Access Optimizing sgrna structure to improve CRISPR-Cas9 knockout efficiency Ying Dang, Gengxiang Jia, Jennie Choi, Hongming
Dang et al. Genome Biology (215) 16:28 DOI 1.1186/s1359-15-846-3 RESEARCH Open Access Optimizing sgrna structure to improve CRISPR-Cas9 knockout efficiency Ying Dang, Gengxiang Jia, Jennie Choi, Hongming
For all of the following, you will have to use this website to determine the answers:
 For all of the following, you will have to use this website to determine the answers: http://blast.ncbi.nlm.nih.gov/blast.cgi We are going to be using the programs under this heading: Answer the following
For all of the following, you will have to use this website to determine the answers: http://blast.ncbi.nlm.nih.gov/blast.cgi We are going to be using the programs under this heading: Answer the following
AIDS free generation. Bob Colebunders Institute of Tropical Medicine
 AIDS free generation Bob Colebunders Institute of Tropical Medicine Why this optimism? Rapid scale-up of antiretroviral therapy Treatment can prevent new infections Expanded coverage of programmes to prevent
AIDS free generation Bob Colebunders Institute of Tropical Medicine Why this optimism? Rapid scale-up of antiretroviral therapy Treatment can prevent new infections Expanded coverage of programmes to prevent
CRISPR/Cas9 targeting of the androgen receptor suppresses the growth of LNCaP human prostate cancer cells
 MOLECULAR MEDICINE REPORTS 17: 2901-2906, 2018 CRISPR/Cas9 targeting of the androgen receptor suppresses the growth of LNCaP human prostate cancer cells CHAOGANG WEI 1,2*, FENGJIAO WANG 3*, WEI LIU 1*,
MOLECULAR MEDICINE REPORTS 17: 2901-2906, 2018 CRISPR/Cas9 targeting of the androgen receptor suppresses the growth of LNCaP human prostate cancer cells CHAOGANG WEI 1,2*, FENGJIAO WANG 3*, WEI LIU 1*,
SECTION 25-1 REVIEW STRUCTURE. 1. The diameter of viruses ranges from about a. 1 to 2 nm. b. 20 to 250 nm. c. 1 to 2 µm. d. 20 to 250 µm.
 SECTION 25-1 REVIEW STRUCTURE VOCABULARY REVIEW Define the following terms. 1. virus 2. capsid 3. retrovirus 4. viroid 5. prion MULTIPLE CHOICE Write the correct letter in the blank. 1. The diameter of
SECTION 25-1 REVIEW STRUCTURE VOCABULARY REVIEW Define the following terms. 1. virus 2. capsid 3. retrovirus 4. viroid 5. prion MULTIPLE CHOICE Write the correct letter in the blank. 1. The diameter of
Supporting Information
 Supporting Information Palmisano et al. 10.1073/pnas.1202174109 Fig. S1. Expression of different transgenes, driven by either viral or human promoters, is up-regulated by amino acid starvation. (A) Quantification
Supporting Information Palmisano et al. 10.1073/pnas.1202174109 Fig. S1. Expression of different transgenes, driven by either viral or human promoters, is up-regulated by amino acid starvation. (A) Quantification
Variation in the HindlII Restriction Fragments of DNA from the Chinese Tian Tan Strain of Vaccinia Virus
 J. gen. irol. (1985), 66, 1819-1823. Printed in Great Britain 1819 Key words: vaccinia virus~vaccine~restriction Jragrnent variation ariation in the Hindl Restriction Fragments of DNA from the Chinese
J. gen. irol. (1985), 66, 1819-1823. Printed in Great Britain 1819 Key words: vaccinia virus~vaccine~restriction Jragrnent variation ariation in the Hindl Restriction Fragments of DNA from the Chinese
