Ethnically Diverse Causes of Walker-Warburg Syndrome (WWS): FCMD Mutations Are a More Common Cause of WWS Outside of the Middle East
|
|
|
- May Osborne
- 6 years ago
- Views:
Transcription
1 HUMAN MUTATION Mutation in Brief #1025, 29:E231-E241, (2008) Online MUTATION IN BRIEF Ethnically Diverse Causes of Walker-Warburg Syndrome (WWS): FCMD Mutations Are a More Common Cause of WWS Outside of the Middle East M. Chiara Manzini 1,2, Danielle Gleason 1,2, Bernard S. Chang 2, R. Sean Hill 1,2, Brenda J. Barry 1,2, Jennifer N. Partlow 1,2, Annapurna Poduri 3, Sophie Currier 1, Patricia Galvin-Parton 4, Lawrence R. Shapiro 5, Karen Schmidt 6, Jessica G. Davis 7, Lina Basel-Vanagaite 8,9, Mohamed Z. Seidahmed 10, Mustafa A. M. Salih 11, William B. Dobyns 12, and Christopher A. Walsh 1,2, * 1 Division of Genetics, Children s Hospital Boston, Harvard Medical School, Boston, MA; 2 Department of Neurology, Beth Israel Deaconess Medical Center, and Howard Hughes Medical Institute, Harvard Medical School, Boston, MA; 3 Department of Neurology, Children s Hospital Boston, Harvard Medical School, Boston, MA; 4 Department of Pediatrics, Stony Brook University Medical Center, Stony Brook, NY; 5 Department of Pediatrics, Westchester Medical Center, Valhalla, NY; 6 Department of Genetics, Children s Hospital of Pittsburgh, Pittsburgh, PA; 7 Department of Pediatrics, Weill Cornell Medical College, New York, NY; 8 Schneider Children's Medical Center of Israel and The Raphael Recanati Genetic Institute, Rabin Medical Center, Petach- Tikva, Israel; 9 Sackler School of Medicine, Tel Aviv University, Tel Aviv; 10 Department of Pediatrics, Security Forces Hospital, Riyadh, Saudi Arabia; 11 Division of Pediatric Neurology, College of Medicine, King Saud University, Riyadh, Saudi Arabia; 12 Department of Human Genetics, Neurology and Pediatrics, University of Chicago, Chicago, IL *Correspondence to: Christopher A. Walsh, MD, PhD, Division of Genetics, Children s Hospital Boston, Harvard Medical School, 77 Avenue Louis Pasteur, NRB-266, Boston, MA Tel: +1 (617) ; cwalsh@bidmc.harvard.edu Communicated by Christine Van Broeckhoven Walker-Warburg syndrome (WWS) is a genetically heterogeneous autosomal recessive disease characterized by congenital muscular dystrophy, cobblestone lissencephaly, and ocular malformations. Mutations in six genes involved in the glycosylation of α-dystroglycan (POMT1, POMT2, POMGNT1, FCMD, FKRP and LARGE) have been identified in WWS patients, but account for only a portion of WWS cases. To better understand the genetics of WWS and establish the frequency and distribution of mutations across WWS genes, we genotyped all known loci in a cohort of 43 WWS patients of varying geographical and ethnic origin. Surprisingly, we reached a molecular diagnosis for 40% of our patients and found mutations in POMT1, POMT2, FCMD and FKRP, many of which were novel alleles, but no mutations in POMGNT1 or LARGE. Notably, the FCMD gene was a more common cause of WWS than previously expected in the European/American subset of our cohort, including all Ashkenazi Jewish cases, who carried the same founder mutation Wiley-Liss, Inc. KEY WORDS: Walker-Warburg syndrome; congenital muscular dystrophy; alpha-dystroglycan; POMT1, POMT2, FCMD, FKRP INTRODUCTION Walker-Warburg syndrome (WWS; MIM# ) is an autosomal recessive disorder characterized by cobblestone (type II) lissencephaly, hydrocephalus, severe cerebellar and ocular malformations, and congenital Received 01 November 2007; accepted revised manuscript 18 May WILEY-LISS, INC. DOI: /humu.20844
2 E232 Manzini et al. muscular dystrophy (CMD) (Dobyns, et al., 1989; Cormand, et al., 2001; Vajsar and Schachter, 2006). It is the most severe form of CMD, leading to significant developmental delay and a life expectancy of less than 3 years. While WWS presents with a relatively homogeneous phenotype, it is genetically heterogeneous (Beltran-Valero de Bernabe, et al., 2002; Currier, et al., 2005; Vajsar and Schachter, 2006). The most common known causes of WWS are mutations in two glycosyltransferases, protein O-mannosyltransferase 1 and 2 (POMT1; MIM# , NM_007171; POMT2; MIM# , NM_013382) (Beltran-Valero de Bernabe, et al., 2002; Currier, et al., 2005; van Reeuwijk, et al., 2005b). POMT1 and POMT2 attach the first sugar in the O-mannose-linked glycan moiety of the α-dystroglycan protein, a transmembrane glycoprotein expressed on the surface of muscle cells and neurons (Barresi and Campbell, 2006). α-dystroglycan interacts with several extracellular matrix components in the basal membrane, and disruption of its function is thought to underlie the severe defects in muscle, eye and brain development in WWS patients (Barresi and Campbell, 2006). However, POMT1 mutations have been reported in only 10-20% of WWS cases (Currier, et al., 2005; Vajsar and Schachter, 2006), and only three WWS patients and three fetuses affected by fetal cobblestone lissencephaly have been found to carry POMT2 mutations (van Reeuwijk, et al., 2005b; Bouchet, et al., 2007). Disruption of α-dystroglycan glycosylation can have variable effects on muscle, eye and brain development, leading to other CMD syndromes that affect the nervous system less severely than WWS (van Reeuwijk, et al., 2005a). Additional genes in the α-dystroglycan glycosylation pathway are mutated in autosomal recessive disorders similar to WWS and grouped under the term α-dystroglycanopathies: POMGNT1 (MIM# , NM_017739) in Muscle Eye Brain disease (MEB; MIM# ) (Yoshida, et al., 2001), FCMD (MIM# , NM_ ) in Fukuyama congenital muscular dystrophy (FCMD; MIM# ) (Kobayashi, et al., 1998), fukutin-related protein (FKRP; MIM# , NM_024301) in Muscular Dystrophy Congenital 1C (MDC1C; MIM# ) (Brockington, et al., 2001), and LARGE (MIM# , NM_004737) in MDC1D (MIM# ) (Longman, et al., 2003). Further genetic analyses have revealed that these α-dystroglycanopathy genes are associated with extremely variable phenotypes ranging from mild CMD to WWS (van Reeuwijk, et al., 2005a; Barresi and Campbell, 2006; Godfrey, et al., 2007). FCMD mutations have been identified in four classic cases of WWS each (de Bernabe, et al., 2003; Silan, et al., 2003; Cotarelo, et al., 2008), LARGE mutations in two WWS cases (Godfrey, et al., 2007; van Reeuwijk, et al., 2007) and FKRP and POMGNT1 mutations in one WWS case each (Beltran-Valero de Bernabe, et al., 2004, Godfrey, et al., 2007). Such genetic heterogeneity of WWS and the variability in the genotype/phenotype correlation for each identified gene have greatly complicated the genetic diagnosis of WWS patients. Because of the rarity of the disease, a comprehensive overview of the frequency and distribution of mutations in all known genes in a carefully phenotyped and homogeneous cohort of patients had not previously been undertaken. To explore the genetic heterogeneity of WWS and provide some guidance in the choice of genetic tests in these patients, we collected a cohort of classic WWS cases of different ethnic origins and analyzed both common causes of WWS (POMT1, POMT2) and genes that are more rarely mutated in this disease, such as FCMD, FKRP, POMGNT1 and LARGE. SUBJECTS AND METHODS Subjects All subjects were enrolled after informed consent was obtained, and research was conducted according to protocols approved by the Institutional Review Boards of Beth Israel Deaconess Medical Center and Children s Hospital, Boston. All patients were diagnosed with classic WWS based on the following criteria: (1) brain malformation on neuroimaging characterized by cobblestone lissencephaly, severe ventriculomegaly, and cerebellar hypoplasia; (2) the presence of ocular malformations; and (3) a clinical diagnosis of congenital muscular dystrophy, usually based on severe hypotonia (Dobyns, et al., 1989; Cormand, et al., 2001). Because of the clinical severity and reduced lifespan associated with this condition, serum creatine kinase (CK) and histopathological confirmation of muscular dystrophy were not available in most cases, although CK was highly elevated in all patients tested. Whole genome scans and sequencing analysis Genomic DNA was prepared from peripheral blood samples from patients and available family members according to standard protocols. 600 ng of genomic DNA were used to hybridize Affymetrix 250K StyI SNP arrays at the Broad Institute of MIT and Harvard, Cambridge, MA. Genotyping data was analyzed using the loss
3 Distribution of Mutations in Known WWS Genes E233 of heterozygosity algorithm on dchip software (Lin, et al., 2004) to identify regions of identity by descent (IBD) at known WWS loci. Sequencing of POMT1 (NM_ ), POMT2 (NM_ ), POMGNT1 (NM_ ), LARGE (NM_ ), FKRP (NM_ ) and FCMD (NM_ ) coding regions was performed on PCR products after amplification of genomic DNA by SeqWright (Houston, TX). PCR primers were designed for each exon including at least 50 base pairs of flanking intronic sequences and are available upon request. DNA changes are described according to the nucleotide numbering reflects in the cdna sequence, with +1 corresponding to the A of the ATG translation initiation codon in the reference sequence, according to journal guidelines ( For protein changes the initiation codon is codon 1. Whenever a putative mutation was identified, its inheritance pattern was confirmed in all available family members, and at least 96 control DNAs (192 alleles) were tested to confirm it was not a benign polymorphic change. Unfortunately, no parents were available in the four compound heterozygous cases identified in this study. Multiplex Ligation-dependent Probe Amplification (MLPA) MLPA analysis was performed using the SALSA MLPA EK-1 kit (MRC-Holland, Amsterdam, The Netherlands) according to the manufacturer specifications. Custom probes for each exon of the POMT2 gene were designed following instructions from MRC-Holland and sequences are available upon request. MLPA-PCR products were run on an ABI-3130XL genetic analyzer (Applied Biosystems, Foster City, CA), and MLPA peaks were quantified on GeneMapper software (Applied Biosystems) and analyzed following protocols from MRC- Holland. RESULTS We collected a cohort of 43 affected individuals (from 40 families) of varying provenance. Sixteen patients from 14 families were Middle Eastern children born to parents who were first cousins (9 from Saudi Arabia, 3 from Israel, 1 from Jordan, 1 from Syria). Twenty-seven additional cases from the United States and Europe were mostly Caucasian (20, including 3 of Ashkenazi Jewish origin), with 6 Hispanic patients and 1 African-American patient. All individuals (n=104) received genome-wide screens on Affymetrix 250K StyI SNP arrays at the Broad Institute of MIT and Harvard, Cambridge, MA. We analyzed both consanguineous and non-consanguineous families for the presence of stretches of homozygous SNPs spanning the known WWS loci (POMT1; POMT2; POMGNT1, FCMD; FKRP and LARGE), using the loss of heterozygosity algorithm in dchip software (Lin, et al., 2004). Stretches of homozygosity/identity by descent (IBD) of the order of several centimorgans (cm) were expected in the consanguineous families in our cohort, with the mutation-bearing fragment usually larger than 5 cm for children whose parents are first cousins (Woods, et al., 2006). Smaller regions of IBD may also be present in affected children from non-consanguineous marriages subject to founder effects. Sequencing of the coding sequence of the corresponding gene was performed wherever IBD was observed. In the non-consanguineous cases, all six genes were sequenced in all affected. In addition, the Affymetrix arrays allow for copy-number change analysis and we assessed all known WWS genes for changes, such as heterozygous or homozygous deletion and duplications, using the copy-number function in dchip (Lin, et al., 2004). No copy-number changes were identified in the six WWS loci analyzed in our cohort, however the average distance between SNPs in these arrays is approx. 10 kb and it is possible that smaller deletions/duplications may be present. POMT1 Five Middle Eastern families and one Hispanic family showed IBD at the POMT1 locus and upon sequencing of the POMT1 gene all but one affected carried a homozygous mutation (Table 1, Families 1,2,4-6). All POMT1 mutations identified were predicted to severely disrupt POMT1 translation and function by either splicing disruption (c.280+1g>t, c g>t), frameshift (c.1260_1261delct, p.l421fs) or early termination (c.2163c>a, Y721X). The same mutation (c.280+1g>t) was found in two unrelated Saudi families (Table 1, Family 1, 2) carrying different haplotypes (not shown), suggesting that this mutation either originated twice in the population, or was an extremely remote founder mutation in the Saudi population. Additional sequencing identified only one other patient carrying a POMT1 mutation (Table 1, Patient 3). An African-American male carried two likely POMT1 mutations: a novel missense change at cysteine 366 (c.1097g>a, p.c366y) and a previously reported frameshift mutation (c.2167_2168insg, p.g722fs, (Beltran-
4 E234 Manzini et al. Valero de Bernabe, et al., 2002)). This patient was presumed to be a compound heterozygote, but parental samples were not available to confirm this. Table 1. Mutations in POMT1, POMT2, FCMD and FKRP Family Origin Consanguinity Gene Mutation 1, 2 Saudi Arabia 1st cousin POMT1 homozygous c.280+1g>t (p.e94fs) 3 US (African) No POMT1 c.1097g>a (p.c366y), c.2167_2168insg (p.g722fs)* 4 Israeli Arab 1st cousin POMT1 homozygous c.1260_1261delct (p.l421fs)* 5 Bahrain 1st cousin POMT1 homozygous c g>t (p.q630fs) 6 US (Hispanic) 1st cousin POMT1 homozygous c.2163c>a (p.y721x)* 7 Palestinian 1st cousin POMT2 homozygous c.924-2a>c (p.k307fs) 8 US (Caucasian) No POMT2 c g>a**, c.1890a>g (splicing?) 9 Saudi Arabia 1st cousin POMT2 homozygous c.1159dela (p.i387sfs) 10 US (Caucasian) No FKRP c.341c>g (p.a114g)*, c.437a>c (p.e146a) 11 Saudi Arabia 1st cousin FKRP homozygous c.962_970dup9 (p.a321_r323dup) 12 US (Caucasian) No FCMD c.385dela (p.i129x), c.1176c>a (p.y392x) 13 US (Hispanic) Probable FCMD homozygous c.557a>g (p.h186r) Ashkenazi No FCMD homozygous c.1167_1168insa (p.f390fs)* * previously identified mutations, ** novel mutations at same site of previously identified mutations RefSeq accession number of sequences: POMT1 NM_ ; POMT2 NM_ ; FKRP NM_ ; FCMD NM_ POMT2 POMT2 forms a heterocomplex with POMT1 to initiate glycosylation (Akasaka-Manya, et al., 2006). POMT2 mutations have been reported in only a few WWS cases (van Reeuwijk, et al., 2005b) and more recently in at least six independent MEB cases (Godfrey, et al., 2007). Five Middle Eastern families showed IBD at the POMT2 locus, and two mutations were found in the coding sequence of the POMT2 gene, one in a Saudi family (c.1159dela) causing a frameshift and early termination (p.i387sfs; Table1, Family 9) and one in a Palestinian family disrupting the splice acceptor in intron 7 (c.924-2a>c, p.k307fs, Table 1, Family 7). IBD in four families without a mutation upon sequencing was not unexpected, as the consanguineous families in our cohort often have only one affected child with multiple regions of IBD across the genome due to the high degree of consanguinity. The SNP haplotypes in the four families were all different, ruling out a common undentified mutation. We asked whether these cases carried copy-number changes, such as deletions or duplication in the POMT2 gene. We had ruled out larger changes by copy-number analysis of the SNP data, but small copy-number changes at the single exon level would not be detected. We excluded homozygous deletion, since all exon could be amplified in all patients during PCR-based sequencing, but could not rule out homozygous duplications. Therefore, we further analyzed these patients via Multiplex Ligation-dependent Probe Amplification (MLPA) at the POMT2 locus (Schouten, et al., 2002). No single exon deletions or duplications were identified (not shown). It is possible that non-coding mutations may cause the disorder in these four families or they may carry mutations in yet unidentified genes. Sequencing in our non-consanguineous cohort identified one additional case carrying two heterozygous putative splicing changes in POMT2 (Table 1, Patient 8). The first is a transition at a highly conserved guanine 5 basepairs (bp) downstream of exon 8 (c g>a). While less universally conserved than the splice donor site, this G in intronic position +5 is an important part of the snrnp binding site (Cartegni, et al., 2002) and identical mutations at the same position have been reported to affect mrna splicing in other CMD cases (Thi Tran, et al., 2005; Bouchet, et al., 2007). The second putative mutation is a synonymous change (c.1890a>g) located at position 2 at the end of exon 18, a position that is also highly conserved and often involved in splicing defects (Cartegni, et al., 2002). The patient is deceased and no cell lines are available for this case, therefore direct analysis of possible
5 Distribution of Mutations in Known WWS Genes E235 splicing disruptions could not be perfomed. To confirm that these were not rare benign SNP s, we doubled the number of control DNAs tested and did not find these changes in 188 control individuals (376 chromosomes). LARGE and POMGNT1 The LARGE gene is usually found mutated in a form of CMD (MDC1D) and the POMGNT1 gene is responsible for Muscle-Eye-Brain disease (MEB). Recently both LARGE and POMGNT1 mutations have been identified in rare WWS patients (van Reeuwijk, et al., 2007; Godfrey, et al., 2007). From our consanguineous cohort one Saudi family showed IBD at the LARGE locus and two Hispanic families showed IBD at the POMGNT1 locus, but no mutation was identified upon sequence analysis. Sequencing of the LARGE and POMGNT1 genes in the nonconsanguineous group did not identify any potential mutations. FKRP FKRP mutations are associated with a wide spectrum of congenital muscular dystrophies with or without cognitive defects and brain malformations (Mercuri, et al., 2003; Louhichi, et al., 2004; Mercuri, et al., 2006). Only one case of WWS and two cases of MEB are known to carry FKRP mutations (Beltran-Valero de Bernabe, et al., 2004; Mercuri, et al., 2006). By homozygosity mapping, one Saudi family on our cohort showed IBD at the FKRP locus and carried a 9 bp duplication (c.962_970dup9) predicted to duplicate three amino acids, alanine 321 to arginine 323 (Table 1, Family 11; Figure 1A). Interestingly, the only previously reported FKRP mutation causing WWS is a transition affecting cysteine 318, which is close to the change in Family 11 (Figure 1C; Beltran- Valero de Bernabe, et al., 2004). Mutations at this location in the protein sequence may be particularly deleterious for brain development. One additional WWS case carrying heterozygous FKRP changes was identified in the European/American nonconsanguineous pool (Table 1, Patient 10). A girl from the United States presented a known mutation (c.341c>g, p.a114g, Figure 1B-C (Mercuri, et al., 2006)), accompanied by a novel missense changes, c.437a>c (p.e146a). Both changes were tested in 188 control individuals (376 chromosomes). The A114G mutation has already been described in homozygosity in one MEB case (Mercuri, et al., 2006). A114 is another residue in the FKRP protein which, when mutated, may severely perturb brain development. FCMD Mutations in the putative glycotransferase FCMD were initially described in FCMD patients in Japan, where an ancestral retrotransposon insertion in the 3 UTR of the gene is present in all affected individuals, usually in homozygosity or less commonly in heterozygosity in combination with mutations in the coding region (Figure 2C) (Kobayashi, et al., 1998; Kondo-Iida, et al., 1999). No homozygous mutations in the coding sequence were identified in FCMD patients, suggesting that such mutations may be embryonic lethal or cause a more severe disorder, such as WWS (Kondo-Iida, et al., 1999). The identification of homozygous coding mutations in the FCMD gene outside of Japan, in two Turkish families affected by WWS (de Bernabe, et al., 2003; Silan, et al., 2003) indicated that WWS may be allelic to FCMD. However, the relative contribution of FCMD mutations to the etiology of WWS has not been studied. Recently, two additional WWS patients were reported to carry FCMD mutations (Cotarelo, et al., 2008). No IBD was observed at the FCMD locus in our consanguineous families. Due to the high resolution of our SNP screen, we wondered whether small regions of IBD with identical haplotype could be used to identify a founder mutation in non-consanguineous families with the same ethnic origin. All three Ashkenazi Jewish families in our group carried the same homozygous haplotype at the FCMD locus (Figure 2A). Upon sequencing of the FCMD gene, we found the same homozygous insertion in all three affected children (c.1167_1168insa, Table 1, Family 14-16). This insertion had been described as a de novo heterozygous change coupled with the 3 UTR insertion in a Japanese FCMD patient (Kondo-Iida, et al., 1999) and was concurrently identified by another group in Family 14, which was obtained from Coriell Cell Repositories, Camden, NJ (Cotarelo, et al., 2008). In our patient cohort, this mutation appears to represent an inherited founder mutation common to all Ashkenazi families (Figure 2A). We therefore analyzed 299 samples from the control Ashkenazi population in Israel to determine carrier frequency and found that 2/299 (0.7%) of the Israeli Ashkenazim carry this FCMD mutation in heterozygosity. Sequencing in the remaining patients identified two additional cases carrying FCMD mutations. A Caucasian boy from the United States carried a deletion (c.381dela, p.i129x) and an early termination (c.1176c>a,
6 E236 Manzini et al. p.y392x) (Table 1, Patient 11; Figure 2B) and was presumed to be a compound heterozygote, but parental samples were not available to confirm this. A Mexican boy from the United States originally reported as being born from a non-consanguineous union showed large stretches of homozygosity in his genome scan, suggesting that the parents share a common ancestor. He carried a homozygous mutation in exon 4 (c.557a>g, p.h186r) (Table 1, Patient 12; Figure 2B). Figure 1. FKRP mutations in classic WWS patients. A. Patient 11 was born to first-cousin parents and carried a homozygous 9 basepair duplication starting at bp.962 in the FKRP gene. This mutation leads to a 3 amino acid duplicagtion. B. Two heterozygous missense mutations were identified in Patient 10. C. Schematic representation of the FKRP protein and mutation summary. The initiation codon is codon 1. Mutations listed above the protein cause WWS: previously reported mutations are in blue, mutations identified in this study are in red. Below the protein mutations causing MEB and CMD associated with variable brain malformations are shown. (h) means heterozygous mutations. Compound heterozygous mutations are connected by an arrow. FKRP mutations causing only CMD are not listed. TM: transmembrane domain.
7 Distribution of Mutations in Known WWS Genes E237 Figure 2. FCMD mutations in classic WWS patients. A. Patient 15 is of Ashkenazi Jewish origin. She carried a homozygous insertion in exon 8 of the FCMD gene, which was inherited from both parents. SNP genotyping results are indicated below each individual. The same mutation and the same SNP haplotype were found in two additional Ashkenazim affected in our cohort suggesting a founder effect in this population. B. Patient 12 carried two heterozygous nonsense mutations, while Patient 13 carried a homozygous missense mutation. C. Schematic representation of the fukutin protein and mutation summary. Mutations listed above the protein cause WWS: previously reported mutations are in blue (G125S was identified in heterozygosity with a small 3 UTR deletion, which is different from the classic FCMD mutations (Cotarelo, et al., 2008)), mutations identified in this study are in red. (h) means heterozygous mutations. Compound heterozygous mutations are connected by an arrow. Below the protein coding mutations causing FCMD are shown; (+3 ) next to each mutation indicates that the mutation is in heterozygosity and associated with the 3 retransposon insertion characteristic of Japanese FCMD patients. TM: transmembrane domain. DISCUSSION To date, there has been no comprehensive analysis of the frequency and distribution of mutations in all known WWS genes in a large group of carefully phenotyped WWS patients. Previous reports indicated that only 10-20% of WWS cases could be explained by mutations in POMT1 and POMT2 (Currier, et al., 2005; Vajsar and Schachter, 2006). Mutations in FCMD, FKRP and LARGE were reported in a few WWS patients each (Silan, et al., 2003; Beltran-Valero de Bernabe, et al., 2003 and 2004; Godfrey, et al., 2007; van Reeuwijk, et al., 2007; Cotarelo, et al., 2008), and were presumed to be extremely rare. We assembled a large cohort of patients with typical WWS, drawn from Middle Eastern consanguineous families and from consanguineous and nonconsanguineous families from Europe and the Americas. We found that 40% (16/40) of patients in our cohort carried mutations in the coding sequence of POMT1, POMT2, FKRP or FCMD with no POMGNT1 or LARGE mutations detected. FCMD and FKRP mutations in particular were much more common than previously
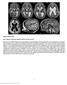
8 E238 Manzini et al. suggested and were mostly identified in non-consanguineous patients of European descent (Table 2). All Ashkenazi Jewish patients in our group shared an identical haplotype at the FCMD locus and the same mutation suggesting a founder effect in this population (Table 2). We identified the carrier frequency of this mutation to be 0.7% in the Ashkenazi population in Israel, which will be extremely informative for genetic testing. A striking difference was observed in the geographic distribution of mutations, as Middle Eastern families were mostly carriers of POMT1 mutations, while the most common cause of European/American cases was FCMD mutations (Table 2). Table 2. Frequency of coding mutations according to origin POMT1 POMT2 FCMD FKRP Middle East (14) 35.7% (5/14) 14.3% (2/14) 0.0% 7.1% (1/14) Europe/Americas (27) 7.4% (2/27) 3.7% (1/27) 18.5% (5/27) 3.7% (1/27) Ashkenazi (3) 0.0% 0.0% 100.0% (3/3) 0.0% Hispanics (6) 16.7% (1/6) 0.0% 16.7% (1/6) 0.0% Our data support the hypothesis that WWS is an extremely genetically heterogeneous disease. Analysis of the medical records and neuroimaging data available for our cohort revealed that patients carrying POMT1, POMT2, FKRP and FCMD mutations are clinically and radiologically very similar (Table 3 and Figure 3). Moreover, they are indistinguishable from WWS patients not carrying mutations in any of these genes.. Figure 3. Brain imaging of children with WWS of varying genetic etiologies. Magnetic resonance imaging (MRI) using T2-weighted axial (top row) and T1-weighted sagittal (bottom row) sequences demonstrates that affected individuals with mutations in the POMT1 (left), FCMD (middle), and FKRP (right) genes have indistinguishable radiological phenotypes. All demonstrate classic WWS findings including cobblestone lissencephaly with an absence of sulcation and an irregular graywhite junction, marked ventriculomegaly, and severe dysplasia of posterior fossa structures, including cerebellar hypoplasia and a kinked appearance of the brainstem.
9 Distribution of Mutations in Known WWS Genes E239 Table 3. Summary of clinical findings in WWS patients carrying mutations in POMT1, POMT2, FKRP and FCMD Pat. Gene Brain findings Eye findings Muscle findings Other Cerebral Cortex Ventricles Cerebellum Tone CK (U/L) 1 POMT1 cobblestone enlarged small, dysmorphic bilateral glaucoma low 1545 and cataracts 2a POMT1 cobblestone enlarged small, dysmorphic retinal dysplasia, glaucoma 2b POMT1 cobblestone enlarged small hemispheres, 3 POMT1 lissencephaly, ctx too thin to assess enlarged, encephalocele small hemispheres, 4 POMT1 cobblestone enlarged small 5 POMT1 lissencephaly, ctx too thin to assess enlarged small hemispheres, L microphthalmia, retinal dysplasia, AC malformation, glaucoma microphthalmia shallow AC, retinal hypoplasia 6 POMT1 cobblestone enlarged small cataracts, AC malformation low 1565 low 787 low 5000 seizures, congenital hip dislocation seizures seizures, contractures, polydactyly contractures, bilateral kidney dysplasia, L hydronephrosis 7 POMT2 cobblestone enlarged small cataracts low elevated 8 POMT2 cobblestone enlarged small hemispheres, mild ON hypoplasia 9 POMT2 cobblestone enlarged small, dysmorphic retinal dysplasia, ON hypoplasia 2000 low 1666 tower skull deformity hydronephrosis 10 FKRP frontal PMG with cobblestone features enlarged small hemispheres, detached retina 11 FKRP cobblestone enlarged small, dysmorphic bilateral cataracts low 5941 seizures, arthrogryposis low seizures 12 FCMD cobblestone enlarged small hemispheres, ON hypoplasia low 2255 seizures 13 FCMD cobblestone enlarged small hemispheres, 14 FCMD cobblestone enlarged encephalocele ON hypoplasia microphthalmia, microcornea, cataracts low elevated 15 FCMD cobblestone enlarged small, dysmorphic microphthalmia, mild hydronephrosis retinal dysplasia 16 FCMD cobblestone enlarged small vermis glaucoma low 9900 seizures Abbreviations: R, right; L, left; AC, anterior chamber; ON, optic nerve; ctx, cortex; PMG, polymicrogyria Since genotype/phenotype correlation does not appear to allow for the prediction of the responsible mutated gene in this disorder, systematic mutation analysis and correlation with geographical and ethnic provenance may be useful to guide a genetic diagnosis. Our data suggests that POMT1 is more frequently mutated in Middle Eastern populations, while FCMD may be an additional candidate for testing in populations of European descent, in particular among Ashkenazi Jewish families. A reduced frequency of FCMD mutations in the Middle East may explain why FCMD was only identified twice by homozygosity mapping as a cause of WWS. In addition, because the exact involvement of FCMD, FKRP and LARGE in α-dystroglycan glycosylation is still largely unknown, comprehensive mutation analysis in patients and genotype/phenotype correlation across the spectrum of disease caused by these genes may provide clues to gene function. Only one mutation in the putative glycosyltransferase FKRP has previously been identified as a cause of WWS (Beltran-Valero de Bernabe, et al., 2004). We have identified one known and two novel alleles: p.a114g was present in heterozygosity in one patient and it was previously found to cause MEB in homozygosity (Mercuri, et al., 2003), p.e146a is a novel change, and the 9bp duplication in Family 11 is a novel mutation near the site of the only previously described WWS mutation (C318Y) (de Bernabe, et al., 2003). While all other known FKRP mutations usually cause congenital
10 E240 Manzini et al. muscular dystrophy with mild or no brain defects (Brockington, et al., 2001; Mercuri, et al., 2006), our results indicate that changes at certain residues in the protein are extremely deleterious for brain development. Functional analyses in animal models will determine how these mutations affect the FKRP protein. ACKNOWLEDGMENTS The authors would like to thank all the families for their participation in this study, and all the collaborators who contributed samples, where no mutation was identified, Adria Bodell and Kira Apse for their help in coordinating collection of patient samples in the past, Seung-Yun Yoo, Eric Morrow and Christina Austin for insightful critical discussion. This work was supported by National Institutes of Health grant R37 NS (CAW). M.C.M is supported by a Development Grant from the Muscular Dystrophy Association. B.S.C. is supported by the National Institutes of Health (K23 NS049159). CAW is an Investigator of the Howard Hughes Medical Institute. REFERENCES Akasaka-Manya K, Manya H, Nakajima A, Kawakita M, Endo T Physical and functional association of human protein O-mannosyltransferases 1 and 2. J Biol Chem 281(28): Barresi R, Campbell KP Dystroglycan: from biosynthesis to pathogenesis of human disease. J Cell Sci 119: Beltran-Valero de Bernabe D, Currier S, Steinbrecher A, Celli J, van Beusekom E, van der Zwaag B, Kayserili H, Merlini L, Chitayat D, Dobyns WB, Cormand B, Lehesjoki AE, Cruces J, Voit T, Walsh CA, van Bokhoven H, Brunner HG Mutations in the O-mannosyltransferase gene POMT1 give rise to the severe neuronal migration disorder Walker-Warburg syndrome. Am J Hum Genet 71(5): Beltran-Valero de Bernabe D, van Bokhoven H, van Beusekom E, Van den Akker W, Kant S, Dobyns WB, Cormand B, Currier S, Hamel B, Talim B, Topaloglu H, Brunner HG A homozygous nonsense mutation in the fukutin gene causes a Walker-Warburg syndrome phenotype. J Med Genet 40(11): Beltran-Valero de Bernabe D, Voit T, Longman C, Steinbrecher A, Straub V, Yuva Y, Herrmann R, Sperner J, Korenke C, Diesen C, Dobyns WB, Brunner HG, van Bokhoven H, Brockington M, Muntoni F Mutations in the FKRP gene can cause muscle-eye-brain disease and Walker-Warburg syndrome. J Med Genet 41(5):e61. Bouchet C, Gonzales M, Vuillaumier-Barrot S, Devisme L, Lebizec C, Alanio E, Bazin A, Bessieres-Grattagliano B, Bigi N, Blanchet P, Bonneau D, Bonnieres M, Carles D, Delahaye S, Fallet-Bianco C, Figarella-Branger D, Gaillard D, Gasser B, Guimiot F, Joubert M, Laurent N, Liprandi A, Loget P, Marcorelles P, Martinovic J, Menez F, Patrier S, Pelluard-Nehme F, Perez MJ, Rouleau-Dubois C, Triau S, Laquerriere A, Encha-Razavi F, Seta N Molecular heterogeneity in fetal forms of type II lissencephaly. Hum Mutat 28(10): Brockington M, Blake DJ, Prandini P, Brown SC, Torelli S, Benson MA, Ponting CP, Estournet B, Romero NB, Mercuri E, Voit T, Sewry CA, Guicheney P, Muntoni F Mutations in the fukutin-related protein gene (FKRP) cause a form of congenital muscular dystrophy with secondary laminin alpha2 deficiency and abnormal glycosylation of alpha-dystroglycan. Am J Hum Genet 69(6): Cartegni L, Chew SL, Krainer AR Listening to silence and understanding nonsense: exonic mutations that affect splicing. Nat Rev Genet 3(4): Cormand B, Pihko H, Bayes M, Valanne L, Santavuori P, Talim B, Gershoni-Baruch R, Ahmad A, van Bokhoven H, Brunner HG, Voit T, Topaloglu H, Dobyns WB, Lehesjoki AE Clinical and genetic distinction between Walker-Warburg syndrome and muscle-eye-brain disease. Neurology 56(8): Cotarelo RP, Valero MC, Prados B, Pena A, Rodriguez L, Fano O, Marco JJ, Martinez-Frias ML, Cruces J Two new patients bearing mutations in the fukutin gene confirm the relevance of this gene in Walker-Warburg syndrome. Clin Genet 73(2): Currier SC, Lee CK, Chang BS, Bodell AL, Pai GS, Job L, Lagae LG, Al-Gazali LI, Eyaid WM, Enns G, Dobyns WB, Walsh CA Mutations in POMT1 are found in a minority of patients with Walker-Warburg syndrome. Am J Med Genet A 133(1):53-7. Dobyns WB, Pagon RA, Armstrong D, Curry CJ, Greenberg F, Grix A, Holmes LB, Laxova R, Michels VV, Robinow M, Zimmerman RL Diagnostic criteria for Walker-Warburg syndrome. Am J Med Genet 32(2): Godfrey C, Clement E, Mein R, Brockington M, Smith J, Talim B, Straub V, Robb S, Quinlivan R, Feng L, Jimenez-Mallebrera C, Mercuri E, Manzur AY, Kinali M, Torelli S, Brown SC, Sewry CA, Bushby K, Topaloglu H, North K, Abbs S, Muntoni F Refining genotype phenotype correlations in muscular dystrophies with defective glycosylation of dystroglycan. Brain. 130:
11 Distribution of Mutations in Known WWS Genes E241 Kobayashi K, Nakahori Y, Miyake M, Matsumura K, Kondo-Iida E, Nomura Y, Segawa M, Yoshioka M, Saito K, Osawa M, Hamano K, Sakakihara Y, Nonaka I, Nakagome Y, Kanazawa I, Nakamura Y, Tokunaga K, Toda T An ancient retrotransposal insertion causes Fukuyama-type congenital muscular dystrophy. Nature 394(6691): Kondo-Iida E, Kobayashi K, Watanabe M, Sasaki J, Kumagai T, Koide H, Saito K, Osawa M, Nakamura Y, Toda T Novel mutations and genotype-phenotype relationships in 107 families with Fukuyama-type congenital muscular dystrophy (FCMD). Hum Mol Genet 8(12): Lin M, Wei LJ, Sellers WR, Lieberfarb M, Wong WH, Li C dchipsnp: significance curve and clustering of SNP-arraybased loss-of-heterozygosity data. Bioinformatics 20(8): Longman C, Brockington M, Torelli S, Jimenez-Mallebrera C, Kennedy C, Khalil N, Feng L, Saran RK, Voit T, Merlini L, Sewry CA, Brown SC, Muntoni F Mutations in the human LARGE gene cause MDC1D, a novel form of congenital muscular dystrophy with severe mental retardation and abnormal glycosylation of alpha-dystroglycan. Hum Mol Genet 12(21): Louhichi N, Triki C, Quijano-Roy S, Richard P, Makri S, Meziou M, Estournet B, Mrad S, Romero NB, Ayadi H, Guicheney P, Fakhfakh F New FKRP mutations causing congenital muscular dystrophy associated with mental retardation and central nervous system abnormalities. Identification of a founder mutation in Tunisian families. Neurogenetics 5(1): Mercuri E, Brockington M, Straub V, Quijano-Roy S, Yuva Y, Herrmann R, Brown SC, Torelli S, Dubowitz V, Blake DJ, Romero NB, Estournet B, Sewry CA, Guicheney P, Voit T, Muntoni F Phenotypic spectrum associated with mutations in the fukutin-related protein gene. Ann Neurol 53(4): Mercuri E, Topaloglu H, Brockington M, Berardinelli A, Pichiecchio A, Santorelli F, Rutherford M, Talim B, Ricci E, Voit T, Muntoni F Spectrum of brain changes in patients with congenital muscular dystrophy and FKRP gene mutations. Arch Neurol 63(2): Schouten JP, McElgunn CJ, Waaijer R, Zwijnenburg D, Diepvens F, Pals G Relative quantification of 40 nucleic acid sequences by multiplex ligation-dependent probe amplification. Nucleic Acids Res 30(12):e57. Silan F, Yoshioka M, Kobayashi K, Simsek E, Tunc M, Alper M, Cam M, Guven A, Fukuda Y, Kinoshita M, Kocabay K, Toda T A new mutation of the fukutin gene in a non-japanese patient. Ann Neurol 53(3): Thi Tran HT, Takeshima Y, Surono A, Yagi M, Wada H, Matsuo M A G-to-A transition at the fifth position of intron-32 of the dystrophin gene inactivates a splice-donor site both in vivo and in vitro. Mol Genet Metab 85(3): Vajsar J, Schachter H Walker-Warburg syndrome. Orphanet J Rare Dis 1:29. van Reeuwijk J, Brunner HG, van Bokhoven H. 2005a. Glyc-O-genetics of Walker-Warburg syndrome. Clin Genet 67(4): van Reeuwijk J, Janssen M, van den Elzen C, Beltran-Valero de Bernabe D, Sabatelli P, Merlini L, Boon M, Scheffer H, Brockington M, Muntoni F, Huynen MA, Verrips A, Walsh CA, Barth PG, Brunner HG, van Bokhoven H. 2005b. POMT2 mutations cause alpha-dystroglycan hypoglycosylation and Walker-Warburg syndrome. J Med Genet 42(12): van Reeuwijk J, Grewal PK, Salih MA, Beltran-Valero de Bernabe D, McLaughlan JM, Michielse CB, Herrmann R, Hewitt JE, Steinbrecher A, Seidahmed MZ, Shaheed MM, Abomelha A, Brunner HG, van Bokhoven H, Voit T Intragenic deletion in the LARGE gene causes Walker-Warburg syndrome. Hum Genet. 121(6): Woods CG, Cox J, Springell K, Hampshire DJ, Mohamed MD, McKibbin M, Stern R, Raymond FL, Sandford R, Malik Sharif S, Karbani G, Ahmed M, Bond J, Clayton D, Inglehearn CF Quantification of homozygosity in consanguineous individuals with autosomal recessive disease. Am J Hum Genet 78(5): Yoshida A, Kobayashi K, Manya H, Taniguchi K, Kano H, Mizuno M, Inazu T, Mitsuhashi H, Takahashi S, Takeuchi M, Herrmann R, Straub V, Talim B, Voit T, Topaloglu H, Toda T, Endo T Muscular dystrophy and neuronal migration disorder caused by mutations in a glycosyltransferase, POMGnT1. Developmental Cell 1(5):
T he hypoglycosylation of a-dystroglycan is a new disease
 1of5 ELECTRONIC LETTER Mutations in the FKRP gene can cause muscle-eye-brain disease and Walker Warburg syndrome D Beltran-Valero de Bernabé, T Voit, C Longman, A Steinbrecher, V Straub, Y Yuva, R Herrmann,
1of5 ELECTRONIC LETTER Mutations in the FKRP gene can cause muscle-eye-brain disease and Walker Warburg syndrome D Beltran-Valero de Bernabé, T Voit, C Longman, A Steinbrecher, V Straub, Y Yuva, R Herrmann,
A novel POMT2 mutation causes mild congenital muscular dystrophy with normal brain MRI
 Brain & Development 31 (2009) 465 468 Case report A novel POMT2 mutation causes mild congenital muscular dystrophy with normal brain MRI Terumi Murakami a,b, Yukiko K. Hayashi a, *, Megumu Ogawa a, Satoru
Brain & Development 31 (2009) 465 468 Case report A novel POMT2 mutation causes mild congenital muscular dystrophy with normal brain MRI Terumi Murakami a,b, Yukiko K. Hayashi a, *, Megumu Ogawa a, Satoru
Refining genotype ^phenotype correlations in muscular dystrophies with defective glycosylation of dystroglycan
 Brain Advance Access published September 18, 2007 doi:10.1093/brain/awm212 Brain (2007)Page1of11 Refining genotype ^phenotype correlations in muscular dystrophies with defective glycosylation of dystroglycan
Brain Advance Access published September 18, 2007 doi:10.1093/brain/awm212 Brain (2007)Page1of11 Refining genotype ^phenotype correlations in muscular dystrophies with defective glycosylation of dystroglycan
Genetic test for Bilateral frontoparietal polymicrogyria
 Genetic test for Bilateral frontoparietal polymicrogyria Daniela Pilz, Cardiff UKGTN Genetic testing for neurological conditions; London February 26 th 2013 Region-specific Polymicrogyria (PMG) bilateral
Genetic test for Bilateral frontoparietal polymicrogyria Daniela Pilz, Cardiff UKGTN Genetic testing for neurological conditions; London February 26 th 2013 Region-specific Polymicrogyria (PMG) bilateral
SALSA MLPA KIT P060-B2 SMA
 SALSA MLPA KIT P6-B2 SMA Lot 111, 511: As compared to the previous version B1 (lot 11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). Please note that, in contrast to the
SALSA MLPA KIT P6-B2 SMA Lot 111, 511: As compared to the previous version B1 (lot 11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). Please note that, in contrast to the
MRC-Holland MLPA. Description version 19;
 SALSA MLPA probemix P6-B2 SMA Lot B2-712, B2-312, B2-111, B2-511: As compared to the previous version B1 (lot B1-11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). SPINAL
SALSA MLPA probemix P6-B2 SMA Lot B2-712, B2-312, B2-111, B2-511: As compared to the previous version B1 (lot B1-11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). SPINAL
Test Information Sheet
 Prenatal Lissencephaly Panel Sequence Analysis and Exon-Level Deletion/Duplication Testing* of 24 Genes Panel Gene List: ACTB, ACTG1, X, ATP6V0A2, B3GALNT2*, B4GAT1*, DCX, FKRP*, FKTN, GMPPB*, ISPD, LAMB1,
Prenatal Lissencephaly Panel Sequence Analysis and Exon-Level Deletion/Duplication Testing* of 24 Genes Panel Gene List: ACTB, ACTG1, X, ATP6V0A2, B3GALNT2*, B4GAT1*, DCX, FKRP*, FKTN, GMPPB*, ISPD, LAMB1,
SALSA MLPA KIT P050-B2 CAH
 SALSA MLPA KIT P050-B2 CAH Lot 0510, 0909, 0408: Compared to lot 0107, extra control fragments have been added at 88, 96, 100 and 105 nt. The 274 nt probe gives a higher signal in lot 0510 compared to
SALSA MLPA KIT P050-B2 CAH Lot 0510, 0909, 0408: Compared to lot 0107, extra control fragments have been added at 88, 96, 100 and 105 nt. The 274 nt probe gives a higher signal in lot 0510 compared to
Supplemental Data: Detailed Characteristics of Patients with MKRN3. Patient 1 was born after an uneventful pregnancy. She presented in our
 1 2 Supplemental Data: Detailed Characteristics of Patients with MKRN3 Mutations 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 Patient 1 was born after an uneventful pregnancy. She presented
1 2 Supplemental Data: Detailed Characteristics of Patients with MKRN3 Mutations 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 Patient 1 was born after an uneventful pregnancy. She presented
MRC-Holland MLPA. Description version 12; 13 January 2017
 SALSA MLPA probemix P219-B3 PAX6 Lot B3-0915: Compared to version B2 (lot B2-1111) two reference probes have been replaced and one additional reference probe has been added. In addition, one flanking probe
SALSA MLPA probemix P219-B3 PAX6 Lot B3-0915: Compared to version B2 (lot B2-1111) two reference probes have been replaced and one additional reference probe has been added. In addition, one flanking probe
A rare case of muscular dystrophy with POMT2 and FKRP gene mutation. Present by : Ghasem Khazaei Supervisor :Dr Mina Mohammadi Sarband
 A rare case of muscular dystrophy with POMT2 and FKRP gene mutation Present by : Ghasem Khazaei Supervisor :Dr Mina Mohammadi Sarband Index : Congenital muscular dystrophy (CMD) Dystroglycanopathies Walker-Warburg
A rare case of muscular dystrophy with POMT2 and FKRP gene mutation Present by : Ghasem Khazaei Supervisor :Dr Mina Mohammadi Sarband Index : Congenital muscular dystrophy (CMD) Dystroglycanopathies Walker-Warburg
variant led to a premature stop codon p.k316* which resulted in nonsense-mediated mrna decay. Although the exact function of the C19L1 is still
 157 Neurological disorders primarily affect and impair the functioning of the brain and/or neurological system. Structural, electrical or metabolic abnormalities in the brain or neurological system can
157 Neurological disorders primarily affect and impair the functioning of the brain and/or neurological system. Structural, electrical or metabolic abnormalities in the brain or neurological system can
Mechanisms of Disease: congenital muscular dystrophies glycosylation takes center stage
 Mechanisms of Disease: congenital muscular dystrophies glycosylation takes center stage Paul T Martin SUMMARY Recent studies have defined a group of muscular dystrophies, now termed the dystroglycanopathies,
Mechanisms of Disease: congenital muscular dystrophies glycosylation takes center stage Paul T Martin SUMMARY Recent studies have defined a group of muscular dystrophies, now termed the dystroglycanopathies,
Mutational and phenotypical spectrum of phenylalanine hydroxylase deficiency in Denmark
 Clin Genet 2016: 90: 247 251 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12692 Mutational and
Clin Genet 2016: 90: 247 251 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12692 Mutational and
Iowa Wellstone Center Muscle Tissue and Cell Culture Repository
 Iowa Wellstone Center Muscle Tissue and Cell Culture Repository Steven A. Moore, M.D., Ph.D. The University of Iowa Department of Pathology and Iowa Wellstone Muscular Dystrophy Cooperative Research Center
Iowa Wellstone Center Muscle Tissue and Cell Culture Repository Steven A. Moore, M.D., Ph.D. The University of Iowa Department of Pathology and Iowa Wellstone Muscular Dystrophy Cooperative Research Center
Fukutin Gene Mutations Cause Dilated Cardiomyopathy with Minimal Muscle Weakness
 Fukutin Gene Mutations Cause Dilated Cardiomyopathy with Minimal Muscle Weakness Terumi Murakami, MD, 1,2 Yukiko K. Hayashi MD, PhD, 1 Satoru Noguchi, PhD 1,2 Megumu Ogawa, BS 1 Ikuya Nonaka, MD, PhD,
Fukutin Gene Mutations Cause Dilated Cardiomyopathy with Minimal Muscle Weakness Terumi Murakami, MD, 1,2 Yukiko K. Hayashi MD, PhD, 1 Satoru Noguchi, PhD 1,2 Megumu Ogawa, BS 1 Ikuya Nonaka, MD, PhD,
JBC Papers in Press. Published on February 24, 2009 as Manuscript C
 JBC Papers in Press. Published on February 24, 2009 as Manuscript C900007200 The latest version is at http://www.jbc.org/cgi/doi/10.1074/jbc.c900007200 LARGE repression in epithelium-derived cancer LOSS
JBC Papers in Press. Published on February 24, 2009 as Manuscript C900007200 The latest version is at http://www.jbc.org/cgi/doi/10.1074/jbc.c900007200 LARGE repression in epithelium-derived cancer LOSS
The congenital muscular dystrophies in 2004: a century of exciting progress
 Neuromuscular Disorders 14 (2004) 635 649 Review The congenital muscular dystrophies in 2004: a century of exciting progress Francesco Muntoni a, *, Thomas Voit b a Department of Paediatrics and Neonatal,
Neuromuscular Disorders 14 (2004) 635 649 Review The congenital muscular dystrophies in 2004: a century of exciting progress Francesco Muntoni a, *, Thomas Voit b a Department of Paediatrics and Neonatal,
MRC-Holland MLPA. Description version 29; 31 July 2015
 SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0114. As compared to the previous B2 version (lot 0813 and 0912), 11 target probes are replaced or added, and 10 new reference probes are included. P082
SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0114. As compared to the previous B2 version (lot 0813 and 0912), 11 target probes are replaced or added, and 10 new reference probes are included. P082
Single Gene (Monogenic) Disorders. Mendelian Inheritance: Definitions. Mendelian Inheritance: Definitions
 Single Gene (Monogenic) Disorders Mendelian Inheritance: Definitions A genetic locus is a specific position or location on a chromosome. Frequently, locus is used to refer to a specific gene. Alleles are
Single Gene (Monogenic) Disorders Mendelian Inheritance: Definitions A genetic locus is a specific position or location on a chromosome. Frequently, locus is used to refer to a specific gene. Alleles are
MRC-Holland MLPA. Description version 29;
 SALSA MLPA KIT P003-B1 MLH1/MSH2 Lot 1209, 0109. As compared to the previous lots 0307 and 1006, one MLH1 probe (exon 19) and four MSH2 probes have been replaced. In addition, one extra MSH2 exon 1 probe,
SALSA MLPA KIT P003-B1 MLH1/MSH2 Lot 1209, 0109. As compared to the previous lots 0307 and 1006, one MLH1 probe (exon 19) and four MSH2 probes have been replaced. In addition, one extra MSH2 exon 1 probe,
Introduction. Overview
 Congenital muscular dystrophies Emma Clement MD ( Dr. Clement of Great Ormond Street Children's Hospital has no relevant financial relationships to disclose. ) Heinz Jungbluth MD PhD ( Dr. Jungbluth of
Congenital muscular dystrophies Emma Clement MD ( Dr. Clement of Great Ormond Street Children's Hospital has no relevant financial relationships to disclose. ) Heinz Jungbluth MD PhD ( Dr. Jungbluth of
Molecular Characterization of the NF2 Gene in Korean Patients with Neurofibromatosis Type 2: A Report of Four Novel Mutations
 Korean J Lab Med 2010;30:190-4 DOI 10.3343/kjlm.2010.30.2.190 Original Article Diagnostic Genetics Molecular Characterization of the NF2 Gene in Korean Patients with Neurofibromatosis Type 2: A Report
Korean J Lab Med 2010;30:190-4 DOI 10.3343/kjlm.2010.30.2.190 Original Article Diagnostic Genetics Molecular Characterization of the NF2 Gene in Korean Patients with Neurofibromatosis Type 2: A Report
MRC-Holland MLPA. Description version 30; 06 June 2017
 SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0517, C1-0114. As compared to the previous B2 version (lot B2-0813, B2-0912), 11 target probes are replaced or added, and 10 new reference probes are
SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0517, C1-0114. As compared to the previous B2 version (lot B2-0813, B2-0912), 11 target probes are replaced or added, and 10 new reference probes are
Genotype Phenotype Analysis of Human Frontoparietal Polymicrogyria Syndromes
 Genotype Phenotype Analysis of Human Frontoparietal Polymicrogyria Syndromes Xianhua Piao, MD, PhD, 1 Bernard S. Chang, MD, 2 Adria Bodell, MS, 2 Katelyn Woods, BS, 1 Bruria BenZeev, MD, 3 Meral Topcu,
Genotype Phenotype Analysis of Human Frontoparietal Polymicrogyria Syndromes Xianhua Piao, MD, PhD, 1 Bernard S. Chang, MD, 2 Adria Bodell, MS, 2 Katelyn Woods, BS, 1 Bruria BenZeev, MD, 3 Meral Topcu,
MRC-Holland MLPA. Description version 08; 30 March 2015
 SALSA MLPA probemix P351-C1 / P352-D1 PKD1-PKD2 P351-C1 lot C1-0914: as compared to the previous version B2 lot B2-0511 one target probe has been removed and three reference probes have been replaced.
SALSA MLPA probemix P351-C1 / P352-D1 PKD1-PKD2 P351-C1 lot C1-0914: as compared to the previous version B2 lot B2-0511 one target probe has been removed and three reference probes have been replaced.
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/35456 holds various files of this Leiden University dissertation. Author: Hassan, Suha Mustafa Title: Toward prevention of Hemoglobinopathies in Oman Issue
Cover Page The handle http://hdl.handle.net/1887/35456 holds various files of this Leiden University dissertation. Author: Hassan, Suha Mustafa Title: Toward prevention of Hemoglobinopathies in Oman Issue
CHROMOSOMAL MICROARRAY (CGH+SNP)
 Chromosome imbalances are a significant cause of developmental delay, mental retardation, autism spectrum disorders, dysmorphic features and/or birth defects. The imbalance of genetic material may be due
Chromosome imbalances are a significant cause of developmental delay, mental retardation, autism spectrum disorders, dysmorphic features and/or birth defects. The imbalance of genetic material may be due
Advances in genetic diagnosis of neurological disorders
 Acta Neurol Scand 2014: 129 (Suppl. 198): 20 25 DOI: 10.1111/ane.12232 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd ACTA NEUROLOGICA SCANDINAVICA Review Article Advances in genetic diagnosis
Acta Neurol Scand 2014: 129 (Suppl. 198): 20 25 DOI: 10.1111/ane.12232 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd ACTA NEUROLOGICA SCANDINAVICA Review Article Advances in genetic diagnosis
READ ORPHA.NET WEBSITE ABOUT BETA-SARCOGLYOCANOPATHY LIMB-GIRDLE MUSCULAR DYSTROPHIES
 READ ORPHA.NET WEBSITE ABOUT BETA-SARCOGLYOCANOPATHY LIMB-GIRDLE MUSCULAR DYSTROPHIES (LGMD) Limb-girdle muscular dystrophies (LGMD) are a heterogeneous group of genetically determined disorders with a
READ ORPHA.NET WEBSITE ABOUT BETA-SARCOGLYOCANOPATHY LIMB-GIRDLE MUSCULAR DYSTROPHIES (LGMD) Limb-girdle muscular dystrophies (LGMD) are a heterogeneous group of genetically determined disorders with a
Low Proportion of Whole Exon Deletions Causing Phenylketonuria in Denmark and Germany
 HUMAN MUTATION Mutation in Brief #952 (2007) Online MUTATION IN BRIEF Low Proportion of Whole Exon Deletions Causing Phenylketonuria in Denmark and Germany Lisbeth Birk Møller 1 *, Anders O.H. Nygren 2,
HUMAN MUTATION Mutation in Brief #952 (2007) Online MUTATION IN BRIEF Low Proportion of Whole Exon Deletions Causing Phenylketonuria in Denmark and Germany Lisbeth Birk Møller 1 *, Anders O.H. Nygren 2,
Clinical Spectrum and Genetic Mechanism of GLUT1-DS. Yasushi ITO (Tokyo Women s Medical University, Japan)
 Clinical Spectrum and Genetic Mechanism of GLUT1-DS Yasushi ITO (Tokyo Women s Medical University, Japan) Glucose transporter type 1 (GLUT1) deficiency syndrome Mutation in the SLC2A1 / GLUT1 gene Deficiency
Clinical Spectrum and Genetic Mechanism of GLUT1-DS Yasushi ITO (Tokyo Women s Medical University, Japan) Glucose transporter type 1 (GLUT1) deficiency syndrome Mutation in the SLC2A1 / GLUT1 gene Deficiency
II. Characterization of the MMP activity that disrupts the DG complex
 Matrix Metalloproteinase Activity that Disrupts the Dystroglycan Complex: Its Role in the Molecular Pathogenesis of Muscular Dystrophies Kiichiro Matsumura, Ken Arai, Di Zhong, Fumiaki Saito, Hiroko Fukuta-Ohi,
Matrix Metalloproteinase Activity that Disrupts the Dystroglycan Complex: Its Role in the Molecular Pathogenesis of Muscular Dystrophies Kiichiro Matsumura, Ken Arai, Di Zhong, Fumiaki Saito, Hiroko Fukuta-Ohi,
SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY.
 SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: LOSS,
SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: LOSS,
SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY.
 SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: GAIN,
SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: GAIN,
Dubowitz Neuromuscular Centre, Institute of Child Health and Great Ormond Street Hospital for Children, UCL, London, UK. 2
 Brain Pathology ISSN 1015-6305 RESEARCH ARTICLE A Comparative Study of a-dystroglycan Glycosylation in Dystroglycanopathies Suggests that the Hypoglycosylation of a-dystroglycan Does Not Consistently Correlate
Brain Pathology ISSN 1015-6305 RESEARCH ARTICLE A Comparative Study of a-dystroglycan Glycosylation in Dystroglycanopathies Suggests that the Hypoglycosylation of a-dystroglycan Does Not Consistently Correlate
Systematizing in vivo modeling of pediatric disorders
 Systematizing in vivo modeling of pediatric disorders Nicholas Katsanis, Ph.D. Duke University Medical Center Center for Human Disease Modeling Rescindo Therapeutics www.dukegenes.org Task Force for
Systematizing in vivo modeling of pediatric disorders Nicholas Katsanis, Ph.D. Duke University Medical Center Center for Human Disease Modeling Rescindo Therapeutics www.dukegenes.org Task Force for
A Dystroglycan Mutation Associated with Limb-Girdle Muscular Dystrophy
 T h e n e w e ngl a nd j o u r na l o f m e dic i n e brief report A Dystroglycan Mutation Associated with Limb-Girdle Muscular Dystrophy Yuji Hara, Ph.D., Burcu Balci-Hayta, Ph.D., Takako Yoshida-Moriguchi,
T h e n e w e ngl a nd j o u r na l o f m e dic i n e brief report A Dystroglycan Mutation Associated with Limb-Girdle Muscular Dystrophy Yuji Hara, Ph.D., Burcu Balci-Hayta, Ph.D., Takako Yoshida-Moriguchi,
Identifying Mutations Responsible for Rare Disorders Using New Technologies
 Identifying Mutations Responsible for Rare Disorders Using New Technologies Jacek Majewski, Department of Human Genetics, McGill University, Montreal, QC Canada Mendelian Diseases Clear mode of inheritance
Identifying Mutations Responsible for Rare Disorders Using New Technologies Jacek Majewski, Department of Human Genetics, McGill University, Montreal, QC Canada Mendelian Diseases Clear mode of inheritance
Congenital muscular dystrophies: New aspects of an expanding group of disorders
 Congenital muscular dystrophies: New aspects of an expanding group of disorders Matthew T. Lisia, Ronald D. Cohn To cite this version: Matthew T. Lisia, Ronald D. Cohn. Congenital muscular dystrophies:
Congenital muscular dystrophies: New aspects of an expanding group of disorders Matthew T. Lisia, Ronald D. Cohn To cite this version: Matthew T. Lisia, Ronald D. Cohn. Congenital muscular dystrophies:
CHAPTER IV RESULTS Microcephaly General description
 47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
LARGE expression in different types of muscular dystrophies other than dystroglycanopathy
 Balci-Hayta et al. BMC Neurology (2018) 18:207 https://doi.org/10.1186/s12883-018-1207-0 RESEARCH ARTICLE LARGE expression in different types of muscular dystrophies other than dystroglycanopathy Burcu
Balci-Hayta et al. BMC Neurology (2018) 18:207 https://doi.org/10.1186/s12883-018-1207-0 RESEARCH ARTICLE LARGE expression in different types of muscular dystrophies other than dystroglycanopathy Burcu
Autism shares features with cerebellar syndromes
 NEWS Autism shares features with cerebellar syndromes BY KELLY RAE CHI 3 DECEMBER 2009 1 / 7 2 / 7 Misshapen structures: MRI scans show that, compared with controls (A), the cerebellum of a child with
NEWS Autism shares features with cerebellar syndromes BY KELLY RAE CHI 3 DECEMBER 2009 1 / 7 2 / 7 Misshapen structures: MRI scans show that, compared with controls (A), the cerebellum of a child with
Nature Genetics: doi: /ng Supplementary Figure 1. Country distribution of GME samples and designation of geographical subregions.
 Supplementary Figure 1 Country distribution of GME samples and designation of geographical subregions. GME samples collected across 20 countries and territories from the GME. Pie size corresponds to the
Supplementary Figure 1 Country distribution of GME samples and designation of geographical subregions. GME samples collected across 20 countries and territories from the GME. Pie size corresponds to the
ORIGINAL CONTRIBUTION. New Mutations, Prenatal Diagnosis, and Founder Effect
 ORIGINAL CONTRIBUTION LAMA2 Gene Analysis in Congenital Muscular Dystrophy New Mutations, Prenatal Diagnosis, and Founder Effect Claudia Di Blasi, PhD; Daniela Piga, PhD; Paolo Brioschi, PhD; Isabella
ORIGINAL CONTRIBUTION LAMA2 Gene Analysis in Congenital Muscular Dystrophy New Mutations, Prenatal Diagnosis, and Founder Effect Claudia Di Blasi, PhD; Daniela Piga, PhD; Paolo Brioschi, PhD; Isabella
Improving detection and genetic counseling in carriers of spinal muscular atrophy
 Clin Genet 2014: 85: 470 475 Printed in Singapore. All rights reserved Short Report 2013 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12222 Improving detection
Clin Genet 2014: 85: 470 475 Printed in Singapore. All rights reserved Short Report 2013 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12222 Improving detection
Beta Thalassemia Case Study Introduction to Bioinformatics
 Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
MRC-Holland MLPA. Description version 08; 18 November 2016
 SALSA MLPA probemix P122-D1 NF1 AREA Lot D1-1016. As compared to lot C2-0312, four probes in the NF1 area and one reference probe have been removed, four reference probes have been replaced and several
SALSA MLPA probemix P122-D1 NF1 AREA Lot D1-1016. As compared to lot C2-0312, four probes in the NF1 area and one reference probe have been removed, four reference probes have been replaced and several
Dr. Ayman Mohsen Mashi, MBBS Consultant Hematology & Blood Transfusion Department Head, Laboratory & Blood Bank King Fahad Central Hospital, Gazan,
 Dr. Ayman Mohsen Mashi, MBBS Consultant Hematology & Blood Transfusion Department Head, Laboratory & Blood Bank King Fahad Central Hospital, Gazan, KSA amashi@moh.gov.sa 24/02/2018 β-thalassemia syndromes
Dr. Ayman Mohsen Mashi, MBBS Consultant Hematology & Blood Transfusion Department Head, Laboratory & Blood Bank King Fahad Central Hospital, Gazan, KSA amashi@moh.gov.sa 24/02/2018 β-thalassemia syndromes
Hutterite brothers both affected with two forms of limb girdle muscular dystrophy: LGMD2H and LGMD2I
 (2005) 13, 978 982 & 2005 Nature Publishing Group All rights reserved 1018-4813/05 $30.00 ARTICLE www.nature.com/ejhg Hutterite brothers both with two forms of limb girdle muscular dystrophy: LGMD2H and
(2005) 13, 978 982 & 2005 Nature Publishing Group All rights reserved 1018-4813/05 $30.00 ARTICLE www.nature.com/ejhg Hutterite brothers both with two forms of limb girdle muscular dystrophy: LGMD2H and
MRC-Holland MLPA. Description version 14; 28 September 2016
 SALSA MLPA probemix P279-B3 CACNA1A Lot B3-0816. As compared to version B2 (lot B2-1012), one reference probe has been replaced and the length of several probes has been adjusted. Voltage-dependent calcium
SALSA MLPA probemix P279-B3 CACNA1A Lot B3-0816. As compared to version B2 (lot B2-1012), one reference probe has been replaced and the length of several probes has been adjusted. Voltage-dependent calcium
Most severely affected will be the probe for exon 15. Please keep an eye on the D-fragments (especially the 96 nt fragment).
 SALSA MLPA probemix P343-C3 Autism-1 Lot C3-1016. As compared to version C2 (lot C2-0312) five reference probes have been replaced, one reference probe added and several lengths have been adjusted. Warning:
SALSA MLPA probemix P343-C3 Autism-1 Lot C3-1016. As compared to version C2 (lot C2-0312) five reference probes have been replaced, one reference probe added and several lengths have been adjusted. Warning:
MEDICAL GENOMICS LABORATORY. Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG)
 Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG) Ordering Information Acceptable specimen types: Fresh blood sample (3-6 ml EDTA; no time limitations associated with receipt)
Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG) Ordering Information Acceptable specimen types: Fresh blood sample (3-6 ml EDTA; no time limitations associated with receipt)
JULY 21, Genetics 101: SCN1A. Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology
 JULY 21, 2018 Genetics 101: SCN1A Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology Disclosures: I have no financial interests or relationships to disclose. Objectives 1. Review genetic
JULY 21, 2018 Genetics 101: SCN1A Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology Disclosures: I have no financial interests or relationships to disclose. Objectives 1. Review genetic
Nature Genetics: doi: /ng Supplementary Figure 1. Brain magnetic resonance imaging in patient 9 at age 3.6 years.
 Supplementary Figure 1 Brain magnetic resonance imaging in patient 9 at age 3.6 years. (a d) Axial T2-weighted images show a marked degree of ventriculomegaly. Note the diencephalic mesencephalic junction
Supplementary Figure 1 Brain magnetic resonance imaging in patient 9 at age 3.6 years. (a d) Axial T2-weighted images show a marked degree of ventriculomegaly. Note the diencephalic mesencephalic junction
MRC-Holland MLPA. Description version 07; 26 November 2015
 SALSA MLPA probemix P266-B1 CLCNKB Lot B1-0415, B1-0911. As compared to version A1 (lot A1-0908), one target probe for CLCNKB (exon 11) has been replaced. In addition, one reference probe has been replaced
SALSA MLPA probemix P266-B1 CLCNKB Lot B1-0415, B1-0911. As compared to version A1 (lot A1-0908), one target probe for CLCNKB (exon 11) has been replaced. In addition, one reference probe has been replaced
The University of Chicago Genetic Services Laboratories LaboLaboratories. Molecular Testing for Lissencephaly
 The University of Chicago Genetic Services Laboratories LaboLaboratories 5841 S. Maryland Ave., Rm. G701, MC 0077, Chicago, Illinois 60637-9130 ucgslabs@genetics.uchicago.edu dnatesting.uchicago.edu CLIA
The University of Chicago Genetic Services Laboratories LaboLaboratories 5841 S. Maryland Ave., Rm. G701, MC 0077, Chicago, Illinois 60637-9130 ucgslabs@genetics.uchicago.edu dnatesting.uchicago.edu CLIA
Exploding Genetic Knowledge in Developmental Disabilities. Disclosures. The Genetic Principle
 Exploding Genetic Knowledge in Developmental Disabilities How to acquire the data and how to make use of it Elliott H. Sherr MD PhD Professor of Neurology & Pediatrics UCSF Disclosures InVitae: clinical
Exploding Genetic Knowledge in Developmental Disabilities How to acquire the data and how to make use of it Elliott H. Sherr MD PhD Professor of Neurology & Pediatrics UCSF Disclosures InVitae: clinical
SALSA MLPA probemix P185-C2 Intersex Lot C2-1015: As compared to the previous version C1 (lot C1-0611), the lengths of four probes have been adjusted.
 mix P185-C2 Intersex Lot C2-1015: As compared to the previous version C1 (lot C1-0611), the lengths of four s have been adjusted. The sex-determining region on chromosome Y (SRY) is the most important
mix P185-C2 Intersex Lot C2-1015: As compared to the previous version C1 (lot C1-0611), the lengths of four s have been adjusted. The sex-determining region on chromosome Y (SRY) is the most important
Supplementary Online Content
 Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
SALSA MLPA probemix P241-D2 MODY mix 1 Lot D As compared to version D1 (lot D1-0911), one reference probe has been replaced.
 mix P241-D2 MODY mix 1 Lot D2-0413. As compared to version D1 (lot D1-0911), one reference has been replaced. Maturity-Onset Diabetes of the Young (MODY) is a distinct form of non insulin-dependent diabetes
mix P241-D2 MODY mix 1 Lot D2-0413. As compared to version D1 (lot D1-0911), one reference has been replaced. Maturity-Onset Diabetes of the Young (MODY) is a distinct form of non insulin-dependent diabetes
What lies beneath: challenges in reporting SNP array results. Jonathan Waters
 What lies beneath: challenges in reporting SNP array results Jonathan Waters Challenges in variant interpretation Austin Court, Birmingham Monday 4 th July 2016 What lies beneath: challenges in reporting
What lies beneath: challenges in reporting SNP array results Jonathan Waters Challenges in variant interpretation Austin Court, Birmingham Monday 4 th July 2016 What lies beneath: challenges in reporting
Next-generation sequencing-based molecular diagnosis of neonatal hypotonia in Chinese
 Title page Next-generation sequencing-based molecular diagnosis of neonatal hypotonia in Chinese Population Yan Wang 1, #,Wei peng 1,#, Hong-Yan Guo 3,4, Hui Li 3,4, Jie Tian 3,4, Yu-Jing Shi 3,4, Xiao
Title page Next-generation sequencing-based molecular diagnosis of neonatal hypotonia in Chinese Population Yan Wang 1, #,Wei peng 1,#, Hong-Yan Guo 3,4, Hui Li 3,4, Jie Tian 3,4, Yu-Jing Shi 3,4, Xiao
Long-term follow-up of type 1 lissencephaly: survival is related to neuroimaging abnormalities
 DEVELOPMENTAL MEDICINE & CHILD NEUROLOGY ORIGINAL ARTICLE Long-term follow-up of type 1 lissencephaly: survival is related to neuroimaging abnormalities MARIE-CLAIRE Y DE WIT 1 JOJANNEKE DE RIJK-VAN ANDEL
DEVELOPMENTAL MEDICINE & CHILD NEUROLOGY ORIGINAL ARTICLE Long-term follow-up of type 1 lissencephaly: survival is related to neuroimaging abnormalities MARIE-CLAIRE Y DE WIT 1 JOJANNEKE DE RIJK-VAN ANDEL
SALSA MLPA probemix P241-D2 MODY mix 1 Lot D2-0716, D As compared to version D1 (lot D1-0911), one reference probe has been replaced.
 mix P241-D2 MODY mix 1 Lot D2-0716, D2-0413. As compared to version D1 (lot D1-0911), one reference has been replaced. Maturity-Onset Diabetes of the Young (MODY) is a distinct form of non insulin-dependent
mix P241-D2 MODY mix 1 Lot D2-0716, D2-0413. As compared to version D1 (lot D1-0911), one reference has been replaced. Maturity-Onset Diabetes of the Young (MODY) is a distinct form of non insulin-dependent
Introduction to genetic variation. He Zhang Bioinformatics Core Facility 6/22/2016
 Introduction to genetic variation He Zhang Bioinformatics Core Facility 6/22/2016 Outline Basic concepts of genetic variation Genetic variation in human populations Variation and genetic disorders Databases
Introduction to genetic variation He Zhang Bioinformatics Core Facility 6/22/2016 Outline Basic concepts of genetic variation Genetic variation in human populations Variation and genetic disorders Databases
SALSA MLPA probemix P315-B1 EGFR
 SALSA MLPA probemix P315-B1 EGFR Lot B1-0215 and B1-0112. As compared to the previous A1 version (lot 0208), two mutation-specific probes for the EGFR mutations L858R and T709M as well as one additional
SALSA MLPA probemix P315-B1 EGFR Lot B1-0215 and B1-0112. As compared to the previous A1 version (lot 0208), two mutation-specific probes for the EGFR mutations L858R and T709M as well as one additional
HST.161 Molecular Biology and Genetics in Modern Medicine Fall 2007
 MIT OpenCourseWare http://ocw.mit.edu HST.161 Molecular Biology and Genetics in Modern Medicine Fall 2007 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.161 Molecular Biology and Genetics in Modern Medicine Fall 2007 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Identification of a novel duplication mutation in the VHL gene in a large Chinese family with Von Hippel-Lindau (VHL) syndrome
 Identification of a novel duplication mutation in the VHL gene in a large Chinese family with Von Hippel-Lindau (VHL) syndrome L.H. Cao 1, B.H. Kuang 2, C. Chen 1, C. Hu 2, Z. Sun 1, H. Chen 2, S.S. Wang
Identification of a novel duplication mutation in the VHL gene in a large Chinese family with Von Hippel-Lindau (VHL) syndrome L.H. Cao 1, B.H. Kuang 2, C. Chen 1, C. Hu 2, Z. Sun 1, H. Chen 2, S.S. Wang
MRC-Holland MLPA. Description version 18; 09 September 2015
 SALSA MLPA probemix P090-A4 BRCA2 Lot A4-0715, A4-0714, A4-0314, A4-0813, A4-0712: Compared to lot A3-0710, the 88 and 96 nt control fragments have been replaced (QDX2). This product is identical to the
SALSA MLPA probemix P090-A4 BRCA2 Lot A4-0715, A4-0714, A4-0314, A4-0813, A4-0712: Compared to lot A3-0710, the 88 and 96 nt control fragments have been replaced (QDX2). This product is identical to the
Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016
 Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016 Marwan Tayeh, PhD, FACMG Director, MMGL Molecular Genetics Assistant Professor of Pediatrics Department of Pediatrics
Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016 Marwan Tayeh, PhD, FACMG Director, MMGL Molecular Genetics Assistant Professor of Pediatrics Department of Pediatrics
MEDICAL GENOMICS LABORATORY. Peripheral Nerve Sheath Tumor Panel by Next-Gen Sequencing (PNT-NG)
 Peripheral Nerve Sheath Tumor Panel by Next-Gen Sequencing (PNT-NG) Ordering Information Acceptable specimen types: Blood (3-6ml EDTA; no time limitations associated with receipt) Saliva (OGR-575 DNA Genotek;
Peripheral Nerve Sheath Tumor Panel by Next-Gen Sequencing (PNT-NG) Ordering Information Acceptable specimen types: Blood (3-6ml EDTA; no time limitations associated with receipt) Saliva (OGR-575 DNA Genotek;
Genotype-Phenotype in Egyptian Patients with Nephropathic Cystinosis. (December 2012 report)
 Genotype-Phenotype in Egyptian Patients with Nephropathic Cystinosis (December 2012 report) This is the first study of the genotype of Nephropathic Cystinosis (NC) patients in Egypt and the region of North
Genotype-Phenotype in Egyptian Patients with Nephropathic Cystinosis (December 2012 report) This is the first study of the genotype of Nephropathic Cystinosis (NC) patients in Egypt and the region of North
Investigating Seven Recently Identified Genes in 100 Iranian Families with Autosomal Recessive Non-syndromic Hearing Loss
 Iranian Rehabilitation Journal, Vol. 13, Issue 3, Autumn 2015 Original Article Investigating Seven Recently Identified Genes in 100 Iranian Families with Autosomal Recessive Non-syndromic Hearing Loss
Iranian Rehabilitation Journal, Vol. 13, Issue 3, Autumn 2015 Original Article Investigating Seven Recently Identified Genes in 100 Iranian Families with Autosomal Recessive Non-syndromic Hearing Loss
MRC-Holland MLPA. Related SALSA MLPA probemixes P190 CHEK2: Breast cancer susceptibility, genes included: CHEK2, ATM, PTEN, TP53.
 SALSA MLPA probemix P056-C1 TP53 Lot C1-0215 & lot C1-0214. As compared to version B1 (lot B1-1011) most of the reference and flanking probes have been replaced and several have been added. Furthermore,
SALSA MLPA probemix P056-C1 TP53 Lot C1-0215 & lot C1-0214. As compared to version B1 (lot B1-1011) most of the reference and flanking probes have been replaced and several have been added. Furthermore,
SALSA MS-MLPA KIT ME011-A1 Mismatch Repair genes (MMR) Lot 0609, 0408, 0807, 0407
 SALSA MS-MLPA KIT ME011-A1 Mismatch Repair genes (MMR) Lot 0609, 0408, 0807, 0407 The Mismatch Repair (MMR) system is critical for the maintenance of genomic stability. MMR increases the fidelity of DNA
SALSA MS-MLPA KIT ME011-A1 Mismatch Repair genes (MMR) Lot 0609, 0408, 0807, 0407 The Mismatch Repair (MMR) system is critical for the maintenance of genomic stability. MMR increases the fidelity of DNA
Genetic diagnosis of limb girdle muscular dystrophy type 2A, A Case Report
 Genetic diagnosis of limb girdle muscular dystrophy type 2A, A Case Report Roshanak Jazayeri, MD, PhD Assistant Professor of Medical Genetics Faculty of Medicine, Alborz University of Medical Sciences
Genetic diagnosis of limb girdle muscular dystrophy type 2A, A Case Report Roshanak Jazayeri, MD, PhD Assistant Professor of Medical Genetics Faculty of Medicine, Alborz University of Medical Sciences
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier
 Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name Leber congenital amaurosis OMIM number for disease 204000 Disease alternative
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name Leber congenital amaurosis OMIM number for disease 204000 Disease alternative
SALSA MLPA probemix P169-C2 HIRSCHSPRUNG-1 Lot C As compared to version C1 (lot C1-0612), the length of one probe has been adjusted.
 mix P169-C2 HIRSCHSPRUNG-1 Lot C2-0915. As compared to version C1 (lot C1-0612), the length of one has been adjusted. Hirschsprung disease (HSCR), or aganglionic megacolon, is a congenital disorder characterised
mix P169-C2 HIRSCHSPRUNG-1 Lot C2-0915. As compared to version C1 (lot C1-0612), the length of one has been adjusted. Hirschsprung disease (HSCR), or aganglionic megacolon, is a congenital disorder characterised
Challenges of CGH array testing in children with developmental delay. Dr Sally Davies 17 th September 2014
 Challenges of CGH array testing in children with developmental delay Dr Sally Davies 17 th September 2014 CGH array What is CGH array? Understanding the test Benefits Results to expect Consent issues Ethical
Challenges of CGH array testing in children with developmental delay Dr Sally Davies 17 th September 2014 CGH array What is CGH array? Understanding the test Benefits Results to expect Consent issues Ethical
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1171320/dc1 Supporting Online Material for A Frazzled/DCC-Dependent Transcriptional Switch Regulates Midline Axon Guidance Long Yang, David S. Garbe, Greg J. Bashaw*
www.sciencemag.org/cgi/content/full/1171320/dc1 Supporting Online Material for A Frazzled/DCC-Dependent Transcriptional Switch Regulates Midline Axon Guidance Long Yang, David S. Garbe, Greg J. Bashaw*
Psych 3102 Lecture 3. Mendelian Genetics
 Psych 3102 Lecture 3 Mendelian Genetics Gregor Mendel 1822 1884, paper read 1865-66 Augustinian monk genotype alleles present at a locus can we identify this? phenotype expressed trait/characteristic can
Psych 3102 Lecture 3 Mendelian Genetics Gregor Mendel 1822 1884, paper read 1865-66 Augustinian monk genotype alleles present at a locus can we identify this? phenotype expressed trait/characteristic can
Dysmorphology And The Paediatric Eye. Jill Clayton-Smith Manchester Centre For Genomic Medicine
 Dysmorphology And The Paediatric Eye Jill Clayton-Smith Manchester Centre For Genomic Medicine Why Make A Syndrome Diagnosis? Why did it happen? What does the future hold? How can you treat/manage it?
Dysmorphology And The Paediatric Eye Jill Clayton-Smith Manchester Centre For Genomic Medicine Why Make A Syndrome Diagnosis? Why did it happen? What does the future hold? How can you treat/manage it?
Neuroimaging Manifestations and Classification of Congenital Muscular Dystrophies
 AJNR Am J Neuroradiol 19:1389 1396, September 1998 Neuroimaging Manifestations and Classification of Congenital Muscular Dystrophies A. James Barkovich BACKGROUND AND PURPOSE: Recent work has shown that
AJNR Am J Neuroradiol 19:1389 1396, September 1998 Neuroimaging Manifestations and Classification of Congenital Muscular Dystrophies A. James Barkovich BACKGROUND AND PURPOSE: Recent work has shown that
MRC-Holland MLPA. Description version 08; 07 May 2015
 mix P185-C1 Intersex Lot C1-0611: As compared to the previous version B2 (lot B2-0311), s for CYP21A2 have been removed and s for the CXorf21 gene as well as additional s for NR0B1, NR5A1 and the Y chromosome
mix P185-C1 Intersex Lot C1-0611: As compared to the previous version B2 (lot B2-0311), s for CYP21A2 have been removed and s for the CXorf21 gene as well as additional s for NR0B1, NR5A1 and the Y chromosome
A Lawyer s Perspective on Genetic Screening Performed by Cryobanks
 A Lawyer s Perspective on Genetic Screening Performed by Cryobanks As a lawyer practicing in the area of sperm bank litigation, I have, unfortunately, represented too many couples that conceived a child
A Lawyer s Perspective on Genetic Screening Performed by Cryobanks As a lawyer practicing in the area of sperm bank litigation, I have, unfortunately, represented too many couples that conceived a child
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier
 Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name Parkinson disease 8, automsomal dominant OMIM number for disease 607060 Disease
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name Parkinson disease 8, automsomal dominant OMIM number for disease 607060 Disease
Identification of a novel in-frame de novo mutation in SPTAN1 in intellectual disability and pontocerebellar atrophy
 Hamdan et al., Identification of a novel in-frame de novo mutation in SPTAN1 in intellectual disability and pontocerebellar atrophy Supplementary Information Gene screening and bioinformatics PCR primers
Hamdan et al., Identification of a novel in-frame de novo mutation in SPTAN1 in intellectual disability and pontocerebellar atrophy Supplementary Information Gene screening and bioinformatics PCR primers
SHORT COMMUNICATION. of Medical Sciences, Isfahan, Iran 2 MRC Molecular Haematology Unit, Weatherall Institute of Molecular Medicine, John
 Hemoglobin, 34(1):115 120, (2010) Copyright Informa UK Ltd. ISSN: 0363-0269 print/1532-432x online DOI: 10.3109/03630260903554894 LHEM 0363-0269 1532-432X Hemoglobin, Vol. 34, No. 1, Dec 2009: pp. 0 0
Hemoglobin, 34(1):115 120, (2010) Copyright Informa UK Ltd. ISSN: 0363-0269 print/1532-432x online DOI: 10.3109/03630260903554894 LHEM 0363-0269 1532-432X Hemoglobin, Vol. 34, No. 1, Dec 2009: pp. 0 0
The genetic and molecular basis of muscular dystrophy: roles of cell matrix linkage in the pathogenesis
 J Hum Genet (2006) 51:915 926 DOI 10.1007/s10038-006-0056-7 MINIREVIEW The genetic and molecular basis of muscular dystrophy: roles of cell matrix linkage in the pathogenesis Motoi Kanagawa Æ Tatsushi
J Hum Genet (2006) 51:915 926 DOI 10.1007/s10038-006-0056-7 MINIREVIEW The genetic and molecular basis of muscular dystrophy: roles of cell matrix linkage in the pathogenesis Motoi Kanagawa Æ Tatsushi
Beta Thalassemia Sami Khuri Department of Computer Science San José State University Spring 2015
 Bioinformatics in Medical Product Development SMPD 287 Three Beta Thalassemia Sami Khuri Department of Computer Science San José State University Hemoglobin Outline Anatomy of a gene Hemoglobinopathies
Bioinformatics in Medical Product Development SMPD 287 Three Beta Thalassemia Sami Khuri Department of Computer Science San José State University Hemoglobin Outline Anatomy of a gene Hemoglobinopathies
Petri Net Representation and Analysis of Mannose Type O-Glycan Biosynthesis
 Institute of Experimental Morphology, Pathology and Anthropology with Museum Bulgarian Anatomical Society Acta morphologica et anthropologica, 25 (1-2) Sofia 2018 Petri Net Representation and Analysis
Institute of Experimental Morphology, Pathology and Anthropology with Museum Bulgarian Anatomical Society Acta morphologica et anthropologica, 25 (1-2) Sofia 2018 Petri Net Representation and Analysis
Limb-girdle Muscular Dystrophy with New Mutation in Sarcoglycan Beta Gene: A Case Report
 Iran J Public Health, Vol. 47, No.12, Dec 2018, pp.1953-1957 Case Report Limb-girdle Muscular Dystrophy with New Mutation in Sarcoglycan Beta Gene: A Case Report Eskandar TAGHIZADEH 1,2, Hamed ABDOLKARIMI
Iran J Public Health, Vol. 47, No.12, Dec 2018, pp.1953-1957 Case Report Limb-girdle Muscular Dystrophy with New Mutation in Sarcoglycan Beta Gene: A Case Report Eskandar TAGHIZADEH 1,2, Hamed ABDOLKARIMI
New: P077 BRCA2. This new probemix can be used to confirm results obtained with P045 BRCA2 probemix.
 SALSA MLPA KIT P045-B2 BRCA2/CHEK2 Lot 0410, 0609. As compared to version B1, four reference probes have been replaced and extra control fragments at 100 and 105 nt (X/Y specific) have been included. New:
SALSA MLPA KIT P045-B2 BRCA2/CHEK2 Lot 0410, 0609. As compared to version B1, four reference probes have been replaced and extra control fragments at 100 and 105 nt (X/Y specific) have been included. New:
Computational Systems Biology: Biology X
 Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA L#4:(October-0-4-2010) Cancer and Signals 1 2 1 2 Evidence in Favor Somatic mutations, Aneuploidy, Copy-number changes and LOH
Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA L#4:(October-0-4-2010) Cancer and Signals 1 2 1 2 Evidence in Favor Somatic mutations, Aneuploidy, Copy-number changes and LOH
Supplementary Document
 Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Introduction to LOH and Allele Specific Copy Number User Forum
 Introduction to LOH and Allele Specific Copy Number User Forum Jonathan Gerstenhaber Introduction to LOH and ASCN User Forum Contents 1. Loss of heterozygosity Analysis procedure Types of baselines 2.
Introduction to LOH and Allele Specific Copy Number User Forum Jonathan Gerstenhaber Introduction to LOH and ASCN User Forum Contents 1. Loss of heterozygosity Analysis procedure Types of baselines 2.
Molecular Testing for Lissencephaly
 Molecular Testing for Lissencephaly Clinical Features: Classic Lissencephaly (LIS) or Lissencephaly Type 1 is a smooth or nearly smooth cerebral surface caused by deficient neuronal migration. The spectrum
Molecular Testing for Lissencephaly Clinical Features: Classic Lissencephaly (LIS) or Lissencephaly Type 1 is a smooth or nearly smooth cerebral surface caused by deficient neuronal migration. The spectrum
Proteins. Length of protein varies from thousands of amino acids to only a few insulin only 51 amino acids
 Proteins Protein carbon, hydrogen, oxygen, nitrogen and often sulphur Length of protein varies from thousands of amino acids to only a few insulin only 51 amino acids During protein synthesis, amino acids
Proteins Protein carbon, hydrogen, oxygen, nitrogen and often sulphur Length of protein varies from thousands of amino acids to only a few insulin only 51 amino acids During protein synthesis, amino acids
SALSA MLPA probemix P360-A1 Y-Chromosome Microdeletions Lot A
 SALSA MLPA probemix P360-A1 Y-Chromosome Microdeletions Lot A1-1011. This SALSA MLPA probemix is for basic research and intended for experienced MLPA users only! This probemix enables you to quantify genes
SALSA MLPA probemix P360-A1 Y-Chromosome Microdeletions Lot A1-1011. This SALSA MLPA probemix is for basic research and intended for experienced MLPA users only! This probemix enables you to quantify genes
