Most genes in higher eukaryotes contain multiple introns
|
|
|
- Harold Lucas
- 5 years ago
- Views:
Transcription
1 Serine arginine-rich protein-dependent suppression of exon skipping by exonic splicing enhancers El Chérif Ibrahim*, Thomas D. Schaal, Klemens J. Hertel, Robin Reed*, and Tom Maniatis *Department of Cell Biology, Harvard Medical School, 240 Longwood Avenue, Boston, MA 02115; Department of Molecular and Cellular Biology, Harvard University, 7 Divinity Avenue, Cambridge, MA 02138; and Department of Microbiology and Molecular Genetics, University of California, Irvine, CA Contributed by Tom Maniatis, January 28, 2005 The 5 and 3 splice sites within an intron can, in principle, be joined to those within any other intron during pre-mrna splicing. However, exons are joined in a strict 5 to 3 linear order in constitutively spliced pre-mrnas. Thus, specific mechanisms must exist to prevent the random joining of exons. Here we report that insertion of exon sequences into an intron can inhibit splicing to the downstream 3 splice site and that this inhibition is independent of intron size. The exon sequences required for splicing inhibition were found to be exonic enhancer elements, and their inhibitory activity requires the binding of serine arginine-rich splicing factors. We conclude that exonic enhancers can act as barriers to prevent exon skipping and thereby may play a key role in ensuring the correct 5 to 3 linear order of exons in spliced mrna. intronic splicing silencers pre-mrna splicing accurate splice-site recognition and pairing Most genes in higher eukaryotes contain multiple introns and exons that are transcribed to generate pre-mrna. The production of mature mrna requires the precise removal of introns and joining of exons by splicing. In constitutively spliced pre-mrnas all of the introns are excised, and the exons are joined in the correct 5 to 3 order. This is a consequence of the fact that 5 splice sites are ligated to a 3 splice site exclusively within the same intron, thus avoiding the exclusion of exons ( exon skipping ). Two key problems in understanding the mechanisms of accurate pre-mrna splicing are how the splicing machinery recognizes exons and how exons are joined in the correct 5 to 3 order (i.e., how exon skipping is avoided). Significant advances have been made in understanding the mechanisms involved in accurate exon recognition (1 3). This process requires weakly conserved sequence elements located at the 5 and 3 splice sites and at the branchpoint sequence (BPS) within introns. In addition, exons contain sequence elements known as exonic splicing enhancers (ESEs), which function as binding sites for members of the serine arginine-rich (SR) family of splicing factors. The bound SR proteins promote the use of flanking 5 and 3 splice sites through both protein protein and protein RNA interactions (4 6). Although splicing enhancer SR protein complexes provide a mechanism for exon recognition, the mechanisms by which exon skipping is avoided are not understood. An important insight into this problem was provided by the observation that many human genetic diseases are the consequence of single base mutations in ESEs and that these mutations cause exon skipping (7). Further insights were provided by studies of regulated alternative splicing, where it was shown that the inclusion of an alternate exon requires an ESE and the presence of specific SR proteins (8). Thus, exonic enhancer SR protein complexes are required to avoid exon skipping in both constitutive and alternative splicing. Although these observations indicate that exon recognition is a necessary step in avoiding exon skipping, it is clearly not sufficient. Here we use pre-mrnas containing a single 5 splice site and tandemly duplicated 3 splice sites as a model system to study the mechanisms by which exon skipping is prevented. Our data reveal that splicing to the distal 3 splice site is suppressed by proximal exonic sequences and that SR proteins are required for this suppression. Thus, SR protein exonic enhancer complexes not only function in exon and splice-site recognition but also play a role in ensuring that 5 and 3 splice sites within the same intron are used, thus suppressing exon skipping. Materials and Methods Construction of Plasmids. The wild-type 3 duplication constructs 3 D-14, 3 D-55, 3 D-115, and 3 D-205 were previously described (9) and were renamed WT14, WT55, WT115, and WT205, respectively, after subcloning in pcdna3.1 vector. A 1,480-bp fragment from human thymidine kinase intron 2 (htki2) was cloned into the proximal intron of WT14, WT55, WT115, and WT205, generating WT14-long1, WT55-long1, WT115-long1, and WT205-long1, respectively. To create the mutated BPS (mbps) constructs, the wild-type BPS sequence CACUGAC was changed to CAGAGUG by site-directed mutagenesis (10). The 3 ss series has a naturally occurring thalassemia deletion, 3 (11), of the pyrimidine tract, 3 AG (italics), and the first two nucleotides of exon 2 (lowercase): (UUGGUCUAUUUUC- CCACCCUUAGgc). A 315-bp fragment from htki2 was cloned in the distal intron of WT55, WT205, 3 ss55, and 3 ss205, generating WT55-long2, WT205-long2, 3 ss55-long2, and 3 ss205-long2, respectively. A 212-bp fragment from htki3 was cloned into 3WT205 and 3 ss205 to produce WT205r-long2 and 3 ss205r-long2, respectively. SP64H - Bam-X is a modification of the SP64H - Bam vector containing human -globin exon 1, IVS 1 with an XhoI linker, and exon 2. The pcd vector is a modification of the pcdna3.1 myc-hisa vector with the myc-his portion replaced by an SP6 promoter site. The double duplication clone DD was described previously (9) and was subcloned in pcd vector. DD i was constructed by inserting the HindIII BamHI fragment of human -globin cdna in the XhoI linker of SP64H - Bam-X before subcloning into pcd vector. A 358-bp fragment from the human -globin IVS 2 (position ) was cloned in an XhoI-digested SP64H - Bam-X. A HindIII EcoRI fragment was then subcloned in a pcd vector to produce the control construct. The cis-competition constructs containing synthetic SR protein binding sites were constructed by inserting two copies of a strong 9G8 splicing enhancer (12) downstream of the proximal -globin 3 splice site and three multimerized copies of a strong SC35 splicing enhancer (12) downstream of the distal -globin 3 splice site. The appropriate oligonucleotides were subcloned into identical sites in each of the WT14 and 3 ss14 constructs to generate WT14 (distal SC35) and 3 ss14 (distal SC35). The 9G8 splicing enhancers were multimerized by annealing and Abbreviations: BPS, branchpoint sequence; ESE, exonic splicing enhancer; SR, serine arginine-rich; mbps, mutated BPS; hnrnp, heterogeneous nuclear ribonucleoprotein. See Commentary on page To whom correspondence should be addressed. maniatis@mcb.harvard.edu by The National Academy of Sciences of the USA PNAS April 5, 2005 vol. 102 no cgi doi pnas
2 Fig. 1. Proximal exon sequences suppress use of the distal 3 splice site. (Left) The structures of the pre-mrnas derived from the wild-type (A), mbps (B), and 3 ss (C) constructs are shown. Exonic sequences are represented by boxes; -globin intronic sequences are represented by a straight line. The numbers refer to the length of the duplicated portion of exon 2. (A) For the proximal spliced products (prox), the lengths of the internal exon sequences are indicated in parentheses. (B) *, The mbps. (C) The 3 ss pre-mrnas were presynthesized by using a T7 promoter and then spliced in vitro in HeLa nuclear extract (Left), transcribed by using a cytomegalovirus (CMV) promoter and concomitantly spliced in HeLa nuclear extract (Center), or presynthesized by using a T7 promoter and then microinjected in Xenopus oocyte nuclei to be spliced in vivo (Right). The deletion of the pyrimidine tract at the proximal 3 splice site is indicated by an X-marked 3. #, Pre-mRNA breakdown products; *, nonspecific signal. ligation, and two copies were ligated into the PstI sites of WT14 (distal SC35) and 3 ss14 (distal SC35) to generate WT14 (proximal 9G8 and distal SC35) and 3 ss14 (proximal 9G8 and distal SC35). Synthesis of Pre-mRNAs and in Vitro Splicing. All of the duplication constructs were linearized with BamHI except DD i and the control construct, which were linearized with EcoRI. Pre-mRNAs were synthesized by in vitro transcription with T7 RNA polymerase in the presence of [ - 32 P]UTP (NEN). HeLa-S3 (National Cell Culture Center, Minneapolis) nuclear extracts were prepared as described (13). Splicing assays were performed in 25- l reaction mixtures containing 50% nuclear extract, 0.5 mm ATP, 20 mm creatine phosphate, 3.2 mm MgCl 2, 60 mm KCl, 12 mm Tris HCl (ph 7.6), and ng of 32 P-labeled pre-mrna. Reactions were incubated at 30 C for 90 min. The extracted RNAs were resolved on 6.5% (19:1) 7 M urea polyacrylamide gel (or as indicated in the figure legends) and visualized by Molecular Imager (Bio-Rad). Complementation assays using HeLa cell S100 extracts and recombinant SR proteins were performed as described (12, 14), with the following modification. Reactions containing 40% cytoplasmic S100 extract were supplemented with 200 nm of SC35 and varying amounts of recombinant 9G8 (0 300 nm) before incubation at 30 C. Coupled RNA Polymerase II Transcription Splicing. An efficient system for coupled transcription and splicing has recently been established (R. Das and R.R., unpublished results). Briefly, 200 ng of linearized DNA template was incubated in 25- l reaction mixtures containing 50% HeLa nuclear extract, 0.5 mm ATP, 20 mm creatine phosphate, 3.2 mm MgCl 2, 60 mm KCl, 12 mm Tris HCl (ph 7.6), and 6 Ci of [ - 32 P]UTP (800 Ci mmol; 1 Ci 37 GBq) at 30 C for 90 min. Xenopus Oocyte Microinjection. Microinjection of 32 P-labeled RNAs into Xenopus laevis oocytes was performed as described (15). Oocytes were incubated for 40 min at 18 C, the nuclei were dissected, and nuclear RNAs were extracted and resolved by electrophoresis on 8% denaturing polyacrylamide gels. SR Proteins. Total SR proteins from HeLa-S3 cells were prepared according to Zahler (16). Recombinant SC35 and 9G8 were expressed and purified from baculovirus-infected cell lysates as described (17). Calculations for the Rate of Splicing. The spliced products were quantified by using QUANTITY ONE software (Bio-Rad), and the values obtained were normalized according to the number of uridines present in the RNAs. The splicing efficiency was calculated by using the following formula: [(normalized value of the proximal spliced product) (normalized value of the distal spliced product)] [(normalized value of the pre-mrna) (normalized value of the proximal spliced product) (normalized value of the distal spliced product)]. The ratio of proximal to distal 3 splice-site selection was calculated by using the following formula: (normalized value of the proximal spliced product) (normalized value of the distal spliced product). Results Suppression of Exon Skipping Requires Proximal Exon Sequences. Previous studies using model human -globin pre-mrnas containing a single 5 splice site and tandemly duplicated 3 splice sites showed that the proximal 3 splice site is selected when both BIOCHEMISTRY SEE COMMENTARY Ibrahim et al. PNAS April 5, 2005 vol. 102 no
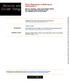
3 Fig. 3. Spliced exons within an intron suppress splicing. (A) Schematic of the double duplication construct and its derivatives. (B) Spliced pre-mrnas synthesized with T7 RNA polymerase. (C) Coupled transcription splicing system. Shaded boxes indicate full-length exon 1, and white boxes indicate full-length exon 2. Ovals indicate the position of the lariat exon intermediate or the excised intron. Coiled lines indicate a size-matched random sequence. Fig. 2. The repression of the use of the distal 3 splice site does not depend on the size of the proximal intron. (Left) The structures of the pre-mrnas derived from the WT-long1 (A), WT-long2 (B), and 3 ss-long2 (C) constructs are shown. A double-headed arrow indicates the insertion of a spacer within the proximal -globin intron. Thymidine kinase intronic sequences are represented by a coiled line. Pre-mRNA breakdown products are indicated by #. The transcripts were resolved on 4.5% denaturing polyacrylamide gels. For the proximal spliced products (prox) and some pre-mrnas, the lengths of the internal exon sequences are indicated in parentheses. the proximal and distal 3 splice sites are adjacent to a full-length exon (9). As the proximal exon is progressively truncated, use of the proximal 3 splice site decreases with a concomitant increase in use of the distal 3 splice site (Fig. 1A). This pattern of 3 splice-site selection results, in part, from the loss of exonic enhancers in the proximal exon, which can no longer bind to SR proteins and promote the use of the proximal 3 splice site (14). However, it is not known why only the proximal 3 splice site is used when both the proximal and distal 3 splice sites are adjacent to the full-length exon 2 (Fig. 1A, lane 1). We investigated the mechanism by which the full-length proximal exon inhibits the use of the distal 3 splice site. To eliminate the effects of competition between the proximal and distal 3 splice sites, we inactivated the proximal 3 splice site. We analyzed two mutants, a substitution mutant (mbps) that inactivates the BPS (10) and a deletion mutant ( 3 ss) lacking the 3 splice site (18). As expected, no splicing to the proximal 3 splice site was observed with the mbps or 3 ss mutant pre-mrnas (Fig. 1 B and C, lanes 1 4). Strikingly, however, when full-length exon 2 is adjacent to the inactivated proximal 3 splice site, the pre-mrnas largely remain unspliced (Fig. 1 B and C, lane 1). Thus, the full-length exon 2 alone inhibits use of the distal 3 splice site. In support of this conclusion, the distal 3 splice site is increasingly used in the pre-mrnas containing progressively larger deletions of proximal exon 2 (Fig. 1 B and C, lanes 2 4). To confirm that the validity of this result is not restricted to in vitro conditions under which presynthesized T7 pre-mrnas are used, we repeated the same experiments in a system where transcription and splicing are coupled. Significantly, we observed exactly the same pattern of 3 splice-site use (compare Fig. 1C Left with Center). The same pattern of 3 splice-site use was observed when the pre-mrnas were microinjected into Xenopus oocyte nuclei (Fig. 1C, lanes 9 12). Taken together, these results show that exon sequences function to suppress exon skipping in vitro and in vivo. Intron Size Does Not Affect Exon Skipping. The average size of introns in the human genome is 1,000 nt, whereas the first intron of the -globin gene is only 130 nt. To determine whether the failure to use the distal 3 splice site results from the small size of the proximal intron, we inserted a 1.5-kb intronic spacer into the proximal intron of the wild-type constructs (Fig. 2A Left). Splicing of these constructs showed the same pattern of cgi doi pnas Ibrahim et al.
4 BIOCHEMISTRY SEE COMMENTARY Fig. 4. SR proteins function in suppressing exon skipping. T7 precursors obtained from the wild-type series (A), the 3 ss series (B), or the double duplication construct and its derivative (C) were spliced in the absence ( ) or presence (filled triangle) of increasing amounts of purified HeLa SR proteins. (A and B) The rate of splicing efficiency (A and B) and the ratio of proximal to distal 3 splice-site selection mediated by the addition of SR proteins (A) are indicated below each lane. Pre-mRNA breakdown products are indicated by #. For the proximal spiced products (prox), the lengths of the internal exon sequences are indicated in parentheses. splice-site use as that observed with the 130-nt intron (Fig. 2A Right). We conclude that the length of the proximal intron is not a critical determinant in the suppression of exon skipping in the model -globin pre-mrnas. We also considered the possibility that the small size of the distal intron affects the accessibility of the distal 3 splice site to the splicing machinery. We therefore inserted a 0.3-kb intronic spacer just upstream of the distal BPS in the wild-type constructs (Fig. 2B Left). As a control, we also replaced the internal full-length exonic sequences with sizematched intronic sequences. As expected, when the internal exonic sequences were full-length, the proximal 3 splice site was preferentially selected (Fig. 2B, lane 2). In contrast, when the internal exonic sequences were shortened to 55 nt, splicing to the distal 3 splice site increased, whereas a proximal spliced product was almost undetectable (compare lanes 2 and 1 in Fig. 2B). Significantly, the replacement of internal exonic sequences with intronic sequences promoted the exclusive use of the distal 3 splice site (compare lanes 3 and 2 in Fig. 2B). This result indicates that even a 55-nt internal exon sequence is sufficient to partially suppress the use of the distal 3 splice site. We further confirmed, with constructs lacking a proximal 3 splice site and containing a 0.3-kb intronic insertion in the distal intron, that a full-size internal exon suppresses the use of the distal 3 splice site. A truncated exon had a weaker suppressor activity, and intronic sequences replacing the internal exon increased the use of the distal 3 splice site (Fig. 2C). Spliced Exons Within an Intron Suppress Splicing. The observation that exon sequences can suppress exon skipping in pre-mrnas containing duplicated 3 splice sites raises the possibility that, in general, splicing cannot occur over exon sequences. To further investigate this possibility, we examined splicing of a construct, Ibrahim et al. PNAS April 5, 2005 vol. 102 no
5 DD , containing duplicated 5 splice sites flanked by full-length exon 1 and duplicated 3 splice sites flanked by full-length exon 2 (Fig. 3A Top). In this context, only the internal pair of 5 and 3 splice sites is used (Fig. 3B, lanes 1 3). The resulting spliced product contains ligated exons 1 and 2 within the intron and is itself a potential splicing substrate (Fig. 3A Middle). Significantly, however, this substrate (DD i) is not spliced, not even when it is synthesized directly as a T7 transcript (Fig. 3B, lanes 4 6). To determine whether this splicing inhibition is due to the fused exons within the intron or to the presence of additional sequences, we examined the splicing of a control construct in which the internal spliced exons were replaced by a size-matched random sequence (Fig. 3A Bottom). In contrast to DD i, the control construct is spliced efficiently (Fig. 3B, lanes 7 9). The same results were obtained in a coupled transcription splicing system (Fig. 3C). Together, these data indicate that splicing cannot occur over exon sequences. SR Proteins Are Required to Suppress Exon Skipping. -Globin exon 2 contains several functional SR protein binding sites (14). Because SR proteins are usually limiting for splicing in nuclear extracts, we added increasing amounts of purified SR proteins to determine whether they affect the choice of proximal versus distal splice sites (Fig. 4). Consistent with previous studies, addition of SR proteins results in a dose-dependent stabilization of the pre-mrnas. For the pre-mrnas containing wild-type duplicated 3 splice sites, an overall increase in splicing efficiency was also observed (Fig. 4A). Moreover, a dose-dependent increase in use of the proximal, but not the distal, 3 splice site was observed for the pre-mrnas containing full-length proximal exons (Fig. 4A, lanes 1 4). This SR protein-dependent effect is progressively decreased as the proximal exon is shortened (115, 55, and 14 nt in lanes 8 11, 9 12, and 13 16, respectively, of Fig. 4A). Thus, suppression of the distal 3 splice site is both exon length-dependent and SR protein-dependent. This conclusion is further strengthened by the observation that SR proteins repress the distal 3 splice site in the 3 ss pre-mrna when the proximal exon 2 is either full-length or 115 nt long (Fig. 4B, lanes 1 4 and 5 8), whereas SR proteins activate the distal 3 splice site in the 3 ss pre-mrna when the proximal exon 2 is either 55 or 14 nt long (Fig. 4B, lanes 9 12 and 13 16). Finally, increasing SR protein levels enhances splicing of the internal intron in DD (Fig. 4C, lanes 1 4) but cannot overcome the block in splicing of DD i (Fig. 4C, lanes 5 8). Together, these data indicate that SR proteins bound to exon sequences can prevent exon skipping. SR proteins function by binding to loosely conserved exonic enhancer elements. To directly test the possibility that SR protein enhancer complexes play a role in preventing exon skipping, we examined pre-mrnas containing multimerized SR protein binding sites replacing both the proximal and distal exon 2. Two copies of a 9G8 binding site (12, 19) were substituted for proximal exon 2, and three copies of an SC35 binding site (12) were substituted for distal exon 2. These pre-mrnas were tested for splicing by using S100 extracts, which lack SR proteins. S100 extracts were complemented with a constant amount of recombinant SC35 and increasing amounts of recombinant 9G8 (Fig. 5). The addition of SC35 alone resulted in use of both the distal and proximal 3 splice sites (compare lanes 1 and 2 in Fig. 5). In contrast, addition of increasing amounts of 9G8 in the presence of a constant amount of SC35 resulted in a dramatic increase in the use of the proximal 3 splice site and a concomitant decrease in the use of the distal 3 splice site (compare lanes 3 5 and 2 in Fig. 5). When the same assay was performed with the 3 ss substrate, the addition of recombinant SC35 alone resulted in exclusive use of the distal 3 splice site as expected (compare lanes 6 and 7 in Fig. 5). Significantly, however, increasing Fig. 5. SR protein enhancer complexes suppress the distal 3 splice site. Splicing of wild-type (lanes 1 5) and mutant (lanes 6 10) pre-mrnas in S100 extract alone (lanes 1 and 6), S100 extract complemented with 200 nm SC35 (lanes 2 and 7), or S100 extract complemented with SC35 and increasing amounts (0 300 nm) of 9G8 (lanes 3 5 and lanes 8 10). amounts of 9G8 resulted in a dose-dependent decrease in use of the distal 3 splice site and in overall splicing efficiency (compare lanes 8 10 and 7 in Fig. 5). We conclude that SR proteinenhancer complexes are required not only to activate the proximal 3 splice site, but also to suppress exon skipping. Discussion We have used model pre-mrnas to investigate the mechanisms involved in preventing exon skipping during pre-mrna splicing. These substrates contain a single 5 splice site and two identical 3 splice sites, each of which are followed by exon sequences. By inactivating the proximal 3 splice site we have been able to uncouple the processes of exon recognition and prevention of exon skipping. We find that exon sequences downstream from the proximal 3 splice site can suppress the use of the distal 3 splice site and that SR proteins are required for this suppression. We also show that exon sequences within an intron can block splicing. We conclude that SR proteins bound to exonic enhancers can function to suppress exon skipping. In previous studies (20, 21) and in the results reported here, the activities of ESEs were shown to be context-specific. ESEs promote early steps of spliceosome assembly when located upstream or downstream of 5 or 3 splice sites, respectively, but they suppress splicing when located within the intron (4). When promoting splicing, SR proteins interact with spliceosomal proteins that bind to the 5 and 3 splice sites (22, 23). Recent studies have shown that the RS domain of SR proteins bound to the exon can also interact directly with the BPS (6). Finally, SR proteins not only function by binding to ESEs, but they also play a critical role in the basic splicing mechanism within the intron, either by facilitating interactions between proteins bound to the 5 and 3 splice sites within the intron (24 26) and or by contacting the 5 splice site in the mature spliceosome (27). The mechanism by which SR proteins that are bound to intron sequences suppress splicing is not understood. Computational analyses indicate that sequence motifs for ESEs are more frequent in exons than in introns (28). However, there are a number of examples in which SR protein binding sites are located within the intron and can function to promote splicing of upstream 5 splice sites (8, 29 32). At the same time, negative intronic regulatory elements with which SR proteins interact have also been found (20, 29, 33, 34). For example, a splicing suppressor element that binds to SR proteins, 3RE, has been identified in the adenovirus major late IIIa pre-mrna (20, 21). 3RE is located immediately adjacent to the BPS. The binding of SR proteins to 3RE seems to prevent the binding of U2 small nuclear ribonucleoprotein to the BPS. However, 3RE can function as a splicing silencer when moved 53 nt from the BPS and cgi doi pnas Ibrahim et al.
6 can also suppress splicing in a heterologous -globin intron (21). Thus, splicing enhancers can suppress splicing when located within the introns of different splicing substrates, and we have shown that this suppression can be observed both in vitro and in microinjected Xenopus oocytes. Thus, there is compelling evidence that SR proteins can suppress splicing when bound to sequences located within introns. How intronic SR protein binding sites can, on the one hand, promote and, on the other hand, inhibit splicing is not understood. It is important to note that the cases where intronic SR protein binding sites have been shown to promote splicing involve regulated alternative splicing and may therefore require the presence of specific regulatory proteins (in addition to SR proteins) that counteract the normally negative activity of the intronic SR protein binding site. There is evidence that the activities of ESEs may be determined by the ratio of heterogeneous nuclear ribonucleoprotein (hnrnp) proteins and SR proteins (35). When ESEs and hnrnp binding sites (exonic splicing silencers) are located adjacent to each other in exons, they can influence each other s activities. Thus, the overall activity of the exon sequence could be determined by the relative amounts of hnrnp proteins and SR proteins present. hnrnp proteins appear to spread from their initial binding sites in both directions, thus interfering with the binding of SR proteins to nearby sequences (36). Similarly, SR proteins are known to bind enhancer sequences cooperatively, to form highly stable complexes (37). These complexes could, in principle, interfere with the spread of hnrnp proteins from nearby binding sites. In the case of exons, positive splicing activity is determined by SR proteins, which promote spliceosome assembly (4), whereas hnrnp proteins block the activity of SR proteins through steric interference. The opposite may be true of constitutively spliced introns, where SR protein binding sites are rare and hnrnp proteins preferentially associate. Recent studies of the SR protein ASF SF2 provide evidence that the SR domain is required to promote splicing when bound 1. Berget, S. M. (1995) J. Biol. Chem. 270, Reed, R. (2000) Curr. Opin. Cell Biol. 12, Hastings, M. L. & Krainer, A. R. (2001) Curr. Opin. Cell Biol. 13, Graveley, B. R. (2000) RNA 6, Blencowe, B. J. (2000) Trends Biochem. Sci. 25, Shen, H., Kan, J. L. & Green, M. R. (2004) Mol. Cell 13, Cartegni, L., Chew, S. L. & Krainer, A. R. (2002) Nat. Rev. Genet. 3, Black, D. L. (2003) Annu. Rev. Biochem. 72, Reed, R. & Maniatis, T. (1986) Cell 46, Reed, R. & Maniatis, T. (1988) Genes Dev. 2, Reed, R. & Maniatis, T. (1985) Cell 41, Schaal, T. D. & Maniatis, T. (1999) Mol. Cell. Biol. 19, Krainer, A. R., Maniatis, T., Ruskin, B. & Green, M. R. (1984) Cell 36, Schaal, T. D. & Maniatis, T. (1999) Mol. Cell. Biol. 19, Luo, M. J. & Reed, R. (1999) Proc. Natl. Acad. Sci. USA 96, Zahler, A. M. (1999) Methods Mol. Biol. 118, Tian, M. & Maniatis, T. (1993) Cell 74, Treisman, R., Orkin, S. H. & Maniatis, T. (1983) Prog. Clin. Biol. Res. 134, Cavaloc, Y., Bourgeois, C. F., Kister, L. & Stevenin, J. (1999) RNA 5, Kanopka, A., Muhlemann, O. & Akusjarvi, G. (1996) Nature 381, Dauksaite, V. & Akusjarvi, G. (2002) J. Biol. Chem. 277, Wu, J. Y. & Maniatis, T. (1993) Cell 75, to sites downstream of a 3 splice site, whereas one of the two RNA binding domains, RBD2, is required to repress splicing when bound within the intron (21). These conclusions, based on studies of MS2 ASF SF2 fusion proteins, do not apply to all SR proteins. We have shown that 9G8, which lacks an RBD2 domain, is also capable of repressing splicing when bound within an intron (Fig. 5). However, the fact that at least two distinct SR protein domains can promote or repress splicing suggests the possibility that the activity of the domain involves interactions with other proteins. These interacting proteins could either promote spliceosome recruitment or interfere with spliceosome assembly. Regardless of the mechanisms involved, the data presented here strongly suggest that SR proteins bound to exons could play an important role in the prevention of exon skipping in constitutively spliced pre-mrnas. According to this view, SR proteins bound to unspliced exons would not only direct the splicing machinery to the nearest 5 or 3 splice sites, they would also suppress splicing between upstream 5 splice sites and downstream 3 splice sites. Similarly, once two internal exons are joined, SR proteins bound to the spliced exons would provide an even stronger barrier to exon skipping. Considering the importance of joining exons in the correct 5 to 3 order, it is likely that additional mechanisms are used to avoid exon skipping, especially for pre-mrnas containing very large introns. For example, the extensive coupling between the transcription and splicing machineries (38) has suggested models in which exon skipping would be avoided by immobilizing the 5 splice site on RNA polymerase until the 3 splice from the same intron is synthesized. We thank members of the R.R. and T.M. laboratories for helpful discussions; André Verdel, Claude Gazin, Jeanne Hsu, and Patricia Valencia for critical reading of the manuscript; and the National Cell Culture Center for providing the HeLa cells. This work was supported by National Institutes of Health Grants GM42231 (to T.M.), GM43375 (to R.R.), and GM62287 (to K.J.H.). 23. Kohtz, J. D., Jamison, S. F., Will, C. L., Zuo, P., Luhrmann, R., Garcia-Blanco, M. A. & Manley, J. L. (1994) Nature 368, Fu, X. D. & Maniatis, T. (1992) Proc. Natl. Acad. Sci. USA 89, Hertel, K. J. & Maniatis, T. (1999) Proc. Natl. Acad. Sci. USA 96, Boukis, L. A., Liu, N., Furuyama, S. & Bruzik, J. P. (2004) J. Biol. Chem. 279, Shen, H. & Green, M. R. (2004) Mol. Cell 16, Liu, H. X., Zhang, M. & Krainer, A. R. (1998) Genes Dev. 12, Gallego, M. E., Gattoni, R., Stevenin, J., Marie, J. & Expert-Bezancon, A. (1997) EMBO J. 16, Lou, H., Neugebauer, K. M., Gagel, R. F. & Berget, S. M. (1998) Mol. Cell. Biol. 18, Hastings, M. L., Wilson, C. M. & Munroe, S. H. (2001) RNA 7, Expert-Bezancon, A., Sureau, A., Durosay, P., Salesse, R., Groeneveld, H., Lecaer, J. P. & Marie, J. (2004) J. Biol. Chem. 279, Pagani, F., Buratti, E., Stuani, C., Romano, M., Zuccato, E., Niksic, M., Giglio, L., Faraguna, D. & Baralle, F. E. (2000) J. Biol. Chem. 275, Simard, M. J. & Chabot, B. (2002) Mol. Cell. Biol. 22, Black, D. L. & Grabowski, P. J. (2003) Prog. Mol. Subcell. Biol. 31, Zhu, J., Mayeda, A. & Krainer, A. R. (2001) Mol. Cell 8, Lynch, K. W. & Maniatis, T. (1995) Genes Dev. 9, Kornblihtt, A. R., De La Mata, M., Fededa, J. P., Munoz, M. J. & Nogues, G. (2004) RNA 10, BIOCHEMISTRY SEE COMMENTARY Ibrahim et al. PNAS April 5, 2005 vol. 102 no
RECAP (1)! In eukaryotes, large primary transcripts are processed to smaller, mature mrnas.! What was first evidence for this precursorproduct
 RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
SR Proteins. The SR Protein Family. Advanced article. Brenton R Graveley, University of Connecticut Health Center, Farmington, Connecticut, USA
 s s Brenton R Graveley, University of Connecticut Health Center, Farmington, Connecticut, USA Klemens J Hertel, University of California Irvine, Irvine, California, USA Members of the serine/arginine-rich
s s Brenton R Graveley, University of Connecticut Health Center, Farmington, Connecticut, USA Klemens J Hertel, University of California Irvine, Irvine, California, USA Members of the serine/arginine-rich
particles at the 3' splice site
 Proc. Nati. Acad. Sci. USA Vol. 89, pp. 1725-1729, March 1992 Biochemistry The 35-kDa mammalian splicing factor SC35 mediates specific interactions between Ul and U2 small nuclear ribonucleoprotein particles
Proc. Nati. Acad. Sci. USA Vol. 89, pp. 1725-1729, March 1992 Biochemistry The 35-kDa mammalian splicing factor SC35 mediates specific interactions between Ul and U2 small nuclear ribonucleoprotein particles
RECAP (1)! In eukaryotes, large primary transcripts are processed to smaller, mature mrnas.! What was first evidence for this precursorproduct
 RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics
 1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics to gain mechanistic insight! 4. Return to step 2, as
1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics to gain mechanistic insight! 4. Return to step 2, as
Exonic Splicing Enhancer Motif Recognized by Human SC35 under Splicing Conditions
 MOLECULAR AND CELLULAR BIOLOGY, Feb. 2000, p. 1063 1071 Vol. 20, No. 3 0270-7306/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Exonic Splicing Enhancer Motif Recognized
MOLECULAR AND CELLULAR BIOLOGY, Feb. 2000, p. 1063 1071 Vol. 20, No. 3 0270-7306/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Exonic Splicing Enhancer Motif Recognized
Received 26 January 1996/Returned for modification 28 February 1996/Accepted 15 March 1996
 MOLECULAR AND CELLULAR BIOLOGY, June 1996, p. 3012 3022 Vol. 16, No. 6 0270-7306/96/$04.00 0 Copyright 1996, American Society for Microbiology Base Pairing at the 5 Splice Site with U1 Small Nuclear RNA
MOLECULAR AND CELLULAR BIOLOGY, June 1996, p. 3012 3022 Vol. 16, No. 6 0270-7306/96/$04.00 0 Copyright 1996, American Society for Microbiology Base Pairing at the 5 Splice Site with U1 Small Nuclear RNA
The SR splicing factors ASF/SF2 and SC35 have antagonistic effects on intronic enhancer-dependent splicing of the β-tropomyosin alternative exon 6A
 The EMBO Journal Vol.16 No.7 pp.1772 1784, 1997 The SR splicing factors ASF/SF2 and SC35 have antagonistic effects on intronic enhancer-dependent splicing of the β-tropomyosin alternative exon 6A Maria
The EMBO Journal Vol.16 No.7 pp.1772 1784, 1997 The SR splicing factors ASF/SF2 and SC35 have antagonistic effects on intronic enhancer-dependent splicing of the β-tropomyosin alternative exon 6A Maria
A general role for splicing enhancers in exon definition
 RNA (2002), 8:1233 1241+ Cambridge University Press+ Printed in the USA+ Copyright 2002 RNA Society+ DOI: 10+1017/S1355838202028030 REPORT A general role for splicing enhancers in exon definition BIANCA
RNA (2002), 8:1233 1241+ Cambridge University Press+ Printed in the USA+ Copyright 2002 RNA Society+ DOI: 10+1017/S1355838202028030 REPORT A general role for splicing enhancers in exon definition BIANCA
Figure mouse globin mrna PRECURSOR RNA hybridized to cloned gene (genomic). mouse globin MATURE mrna hybridized to cloned gene (genomic).
 Splicing Figure 14.3 mouse globin mrna PRECURSOR RNA hybridized to cloned gene (genomic). mouse globin MATURE mrna hybridized to cloned gene (genomic). mrna Splicing rrna and trna are also sometimes spliced;
Splicing Figure 14.3 mouse globin mrna PRECURSOR RNA hybridized to cloned gene (genomic). mouse globin MATURE mrna hybridized to cloned gene (genomic). mrna Splicing rrna and trna are also sometimes spliced;
Function of a Bovine Papillomavirus Type 1 Exonic Splicing Suppressor Requires a Suboptimal Upstream 3 Splice Site
 JOURNAL OF VIROLOGY, Jan. 1999, p. 29 36 Vol. 73, No. 1 0022-538X/99/$04.00 0 Function of a Bovine Papillomavirus Type 1 Exonic Splicing Suppressor Requires a Suboptimal Upstream 3 Splice Site ZHI-MING
JOURNAL OF VIROLOGY, Jan. 1999, p. 29 36 Vol. 73, No. 1 0022-538X/99/$04.00 0 Function of a Bovine Papillomavirus Type 1 Exonic Splicing Suppressor Requires a Suboptimal Upstream 3 Splice Site ZHI-MING
Prediction and Statistical Analysis of Alternatively Spliced Exons
 Prediction and Statistical Analysis of Alternatively Spliced Exons T.A. Thanaraj 1 and S. Stamm 2 The completion of large genomic sequencing projects revealed that metazoan organisms abundantly use alternative
Prediction and Statistical Analysis of Alternatively Spliced Exons T.A. Thanaraj 1 and S. Stamm 2 The completion of large genomic sequencing projects revealed that metazoan organisms abundantly use alternative
The Protein Kinase Clk/Sty Directly Modulates SR Protein Activity: Both Hyper- and Hypophosphorylation Inhibit Splicing
 MOLECULAR AND CELLULAR BIOLOGY, Oct. 1999, p. 6991 7000 Vol. 19, No. 10 0270-7306/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. The Protein Kinase Clk/Sty Directly
MOLECULAR AND CELLULAR BIOLOGY, Oct. 1999, p. 6991 7000 Vol. 19, No. 10 0270-7306/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. The Protein Kinase Clk/Sty Directly
the basis of the disease is diverse (see ref. 16 and references within). One class of thalassemic lesions is single-base substitutions
 Proc. Natl. Acad. Sci. USA Vol. 87, pp. 6253-6257, August 1990 Biochemistry Mechanism for cryptic splice site activation during pre-mrna splicing (thalassemia/ul small nuclear ribonucleoprotein particle/5'
Proc. Natl. Acad. Sci. USA Vol. 87, pp. 6253-6257, August 1990 Biochemistry Mechanism for cryptic splice site activation during pre-mrna splicing (thalassemia/ul small nuclear ribonucleoprotein particle/5'
The role of the mammalian branchpoint sequence in pre-mrna splicing
 The role of the mammalian branchpoint sequence in pre-mrna splicing Robin Reed and Tom Maniatis Department of Biochemistry and Molecular Biology, Harvard University, Cambridge, Massachusetts 02138 USA
The role of the mammalian branchpoint sequence in pre-mrna splicing Robin Reed and Tom Maniatis Department of Biochemistry and Molecular Biology, Harvard University, Cambridge, Massachusetts 02138 USA
Alternative splicing control 2. The most significant 4 slides from last lecture:
 Alternative splicing control 2 The most significant 4 slides from last lecture: 1 2 Regulatory sequences are found primarily close to the 5 -ss and 3 -ss i.e. around exons. 3 Figure 1 Elementary alternative
Alternative splicing control 2 The most significant 4 slides from last lecture: 1 2 Regulatory sequences are found primarily close to the 5 -ss and 3 -ss i.e. around exons. 3 Figure 1 Elementary alternative
Determinants of Plant U12-Dependent Intron Splicing Efficiency
 This article is published in The Plant Cell Online, The Plant Cell Preview Section, which publishes manuscripts accepted for publication after they have been edited and the authors have corrected proofs,
This article is published in The Plant Cell Online, The Plant Cell Preview Section, which publishes manuscripts accepted for publication after they have been edited and the authors have corrected proofs,
Alternative pre-mrna splicing is a fundamental mechanism for
 Caspase-2 pre-mrna alternative splicing: Identification of an intronic element containing a decoy 3 acceptor site Jocelyn Côté*, Sophie Dupuis*, Zhi-Hong Jiang, and Jane Y. Wu Department of Pediatrics
Caspase-2 pre-mrna alternative splicing: Identification of an intronic element containing a decoy 3 acceptor site Jocelyn Côté*, Sophie Dupuis*, Zhi-Hong Jiang, and Jane Y. Wu Department of Pediatrics
ZHIHONG JIANG, 1 JOCELYN COTE, 1 JENNIFER M. KWON, 2 ALISON M. GOATE, 3
 MOLECULAR AND CELLULAR BIOLOGY, June 2000, p. 4036 4048 Vol. 20, No. 11 0270-7306/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Aberrant Splicing of tau Pre-mRNA Caused
MOLECULAR AND CELLULAR BIOLOGY, June 2000, p. 4036 4048 Vol. 20, No. 11 0270-7306/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Aberrant Splicing of tau Pre-mRNA Caused
Frank Rigo and Harold G. Martinson*
 MOLECULAR AND CELLULAR BIOLOGY, Jan. 2008, p. 849 862 Vol. 28, No. 2 0270-7306/08/$08.00 0 doi:10.1128/mcb.01410-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Functional Coupling
MOLECULAR AND CELLULAR BIOLOGY, Jan. 2008, p. 849 862 Vol. 28, No. 2 0270-7306/08/$08.00 0 doi:10.1128/mcb.01410-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Functional Coupling
Modulating alternative splicing by cotranscriptional cleavage of nascent intronic RNA
 Modulating alternative splicing by cotranscriptional cleavage of nascent intronic RNA NATALIA GROMAK, 1,3 GABRIELE TALOTTI, 2,3 NICHOLAS J. PROUDFOOT, 1 and FRANCO PAGANI 2 1 Sir William Dunn School of
Modulating alternative splicing by cotranscriptional cleavage of nascent intronic RNA NATALIA GROMAK, 1,3 GABRIELE TALOTTI, 2,3 NICHOLAS J. PROUDFOOT, 1 and FRANCO PAGANI 2 1 Sir William Dunn School of
Direct Repression of Splicing by transformer-2
 Direct Repression of Splicing by transformer-2 Dawn S. Chandler, Junlin Qi and William Mattox Mol. Cell. Biol. 2003, 23(15):5174. DOI: 10.1128/MCB.23.15.5174-5185.2003. REFERENCES CONTENT ALERTS Updated
Direct Repression of Splicing by transformer-2 Dawn S. Chandler, Junlin Qi and William Mattox Mol. Cell. Biol. 2003, 23(15):5174. DOI: 10.1128/MCB.23.15.5174-5185.2003. REFERENCES CONTENT ALERTS Updated
REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER
 REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER RNA Splicing Lecture 3, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice
REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER RNA Splicing Lecture 3, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice
The nuclear pre-mrna introns of eukaryotes are removed by
 U4 small nuclear RNA can function in both the major and minor spliceosomes Girish C. Shukla and Richard A. Padgett* Department of Molecular Biology, Lerner Research Institute, Cleveland Clinic Foundation,
U4 small nuclear RNA can function in both the major and minor spliceosomes Girish C. Shukla and Richard A. Padgett* Department of Molecular Biology, Lerner Research Institute, Cleveland Clinic Foundation,
In Vivo Regulation of Alternative Pre-mRNA Splicing by the Clk1 Protein Kinase
 MOLECULAR AND CELLULAR BIOLOGY, Oct. 1997, p. 5996 6001 Vol. 17, No. 10 0270-7306/97/$04.00 0 Copyright 1997, American Society for Microbiology In Vivo Regulation of Alternative Pre-mRNA Splicing by the
MOLECULAR AND CELLULAR BIOLOGY, Oct. 1997, p. 5996 6001 Vol. 17, No. 10 0270-7306/97/$04.00 0 Copyright 1997, American Society for Microbiology In Vivo Regulation of Alternative Pre-mRNA Splicing by the
Pyrimidine Tracts between the 5 Splice Site and Branch Point Facilitate Splicing and Recognition of a Small Drosophila Intron
 MOLECULAR AND CELLULAR BIOLOGY, May 1997, p. 2774 2780 Vol. 17, No. 5 0270-7306/97/$04.00 0 Copyright 1997, American Society for Microbiology Pyrimidine Tracts between the 5 Splice Site and Branch Point
MOLECULAR AND CELLULAR BIOLOGY, May 1997, p. 2774 2780 Vol. 17, No. 5 0270-7306/97/$04.00 0 Copyright 1997, American Society for Microbiology Pyrimidine Tracts between the 5 Splice Site and Branch Point
Genetics. Instructor: Dr. Jihad Abdallah Transcription of DNA
 Genetics Instructor: Dr. Jihad Abdallah Transcription of DNA 1 3.4 A 2 Expression of Genetic information DNA Double stranded In the nucleus Transcription mrna Single stranded Translation In the cytoplasm
Genetics Instructor: Dr. Jihad Abdallah Transcription of DNA 1 3.4 A 2 Expression of Genetic information DNA Double stranded In the nucleus Transcription mrna Single stranded Translation In the cytoplasm
Selection of Alternative 5 Splice Sites: Role of U1 snrnp and Models for the Antagonistic Effects of SF2/ASF and hnrnp A1
 MOLECULAR AND CELLULAR BIOLOGY, Nov. 2000, p. 8303 8318 Vol. 20, No. 22 0270-7306/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Selection of Alternative 5 Splice Sites:
MOLECULAR AND CELLULAR BIOLOGY, Nov. 2000, p. 8303 8318 Vol. 20, No. 22 0270-7306/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Selection of Alternative 5 Splice Sites:
Processing of RNA II Biochemistry 302. February 13, 2006
 Processing of RNA II Biochemistry 302 February 13, 2006 Precursor mrna: introns and exons Intron: Transcribed RNA sequence removed from precursor RNA during the process of maturation (for class II genes:
Processing of RNA II Biochemistry 302 February 13, 2006 Precursor mrna: introns and exons Intron: Transcribed RNA sequence removed from precursor RNA during the process of maturation (for class II genes:
A Purine-rich Intronic Element Enhances Alternative Splicing of Thyroid Hormone Receptor mrna
 Marquette University e-publications@marquette Biological Sciences Faculty Research and Publications Biological Sciences, Department of 1-1-2001 A Purine-rich Intronic Element Enhances Alternative Splicing
Marquette University e-publications@marquette Biological Sciences Faculty Research and Publications Biological Sciences, Department of 1-1-2001 A Purine-rich Intronic Element Enhances Alternative Splicing
REGULATED AND NONCANONICAL SPLICING
 REGULATED AND NONCANONICAL SPLICING RNA Processing Lecture 3, Biological Regulatory Mechanisms, Hiten Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice site consensus sequences do have
REGULATED AND NONCANONICAL SPLICING RNA Processing Lecture 3, Biological Regulatory Mechanisms, Hiten Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice site consensus sequences do have
Evidence that U5 snrnp recognizes the 3 splice site for catalytic step II in mammals
 The EMBO Journal Vol.16 No.15 pp.4746 4759, 1997 Evidence that U5 snrnp recognizes the 3 splice site for catalytic step II in mammals Maria Dolores Chiara 1, Leon Palandjian, sites in pre-mrna very near
The EMBO Journal Vol.16 No.15 pp.4746 4759, 1997 Evidence that U5 snrnp recognizes the 3 splice site for catalytic step II in mammals Maria Dolores Chiara 1, Leon Palandjian, sites in pre-mrna very near
Identification and characterization of multiple splice variants of Cdc2-like kinase 4 (Clk4)
 Identification and characterization of multiple splice variants of Cdc2-like kinase 4 (Clk4) Vahagn Stepanyan Department of Biological Sciences, Fordham University Abstract: Alternative splicing is an
Identification and characterization of multiple splice variants of Cdc2-like kinase 4 (Clk4) Vahagn Stepanyan Department of Biological Sciences, Fordham University Abstract: Alternative splicing is an
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10495 WWW.NATURE.COM/NATURE 1 2 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 3 4 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 5 6 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 7 8 WWW.NATURE.COM/NATURE
doi:10.1038/nature10495 WWW.NATURE.COM/NATURE 1 2 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 3 4 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 5 6 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 7 8 WWW.NATURE.COM/NATURE
Spliceosome Pathway. is required for a stable interaction between U2 snrnp and
 MOLECULAR AND CELLULAR BIOLOGY, Sept. 1988, p. 3755-3760 Vol. 8, No. 9 0270-7306/88/093755-06$02.00/0 Copyright 1988, American Society for Microbiology Early Commitment of Yeast Pre-mRNA to the Spliceosome
MOLECULAR AND CELLULAR BIOLOGY, Sept. 1988, p. 3755-3760 Vol. 8, No. 9 0270-7306/88/093755-06$02.00/0 Copyright 1988, American Society for Microbiology Early Commitment of Yeast Pre-mRNA to the Spliceosome
Processing of RNA II Biochemistry 302. February 14, 2005 Bob Kelm
 Processing of RNA II Biochemistry 302 February 14, 2005 Bob Kelm What s an intron? Transcribed sequence removed during the process of mrna maturation (non proteincoding sequence) Discovered by P. Sharp
Processing of RNA II Biochemistry 302 February 14, 2005 Bob Kelm What s an intron? Transcribed sequence removed during the process of mrna maturation (non proteincoding sequence) Discovered by P. Sharp
The organization of 3' splice-site sequences in mammalian introns
 The organization of 3' splice-site sequences in mammalian introns Robin Reed Department of Cellular and Molecular Physiology, Harvard Medical School, Boston, Massachusetts 02115 USA A model pre-mrna substrate
The organization of 3' splice-site sequences in mammalian introns Robin Reed Department of Cellular and Molecular Physiology, Harvard Medical School, Boston, Massachusetts 02115 USA A model pre-mrna substrate
Processing of RNA II Biochemistry 302. February 18, 2004 Bob Kelm
 Processing of RNA II Biochemistry 302 February 18, 2004 Bob Kelm What s an intron? Transcribed sequence removed during the process of mrna maturation Discovered by P. Sharp and R. Roberts in late 1970s
Processing of RNA II Biochemistry 302 February 18, 2004 Bob Kelm What s an intron? Transcribed sequence removed during the process of mrna maturation Discovered by P. Sharp and R. Roberts in late 1970s
In vitro DNase I foot printing. In vitro DNase I footprinting was performed as described
 Supplemental Methods In vitro DNase I foot printing. In vitro DNase I footprinting was performed as described previously 1 2 using 32P-labeled 211 bp fragment from 3 HS1. Footprinting reaction mixes contained
Supplemental Methods In vitro DNase I foot printing. In vitro DNase I footprinting was performed as described previously 1 2 using 32P-labeled 211 bp fragment from 3 HS1. Footprinting reaction mixes contained
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins.
 Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
Identification of cis-acting Intron and Exon Regions in Influenza
 MOLECULAR AND CELLULAR BIOLOGY, Mar. 1992, p. 962-970 0270-7306/92/030962-09$02.00/0 Copyright ) 1992, American Society for Microbiology Vol. 12, No. 3 Identification of cis-acting Intron and Exon Regions
MOLECULAR AND CELLULAR BIOLOGY, Mar. 1992, p. 962-970 0270-7306/92/030962-09$02.00/0 Copyright ) 1992, American Society for Microbiology Vol. 12, No. 3 Identification of cis-acting Intron and Exon Regions
A Novel Type of Splicing Enhancer Regulating Adenovirus Pre-mRNA Splicing
 A Novel Type of Splicing Enhancer Regulating Adenovirus Pre-mRNA Splicing Oliver Mühlemann, Bai-Gong Yue, Svend Petersen-Mahrt and Göran Akusjärvi Mol. Cell. Biol. 2000, 20(7):2317. DOI: 10.1128/MCB.20.7.2317-2325.2000.
A Novel Type of Splicing Enhancer Regulating Adenovirus Pre-mRNA Splicing Oliver Mühlemann, Bai-Gong Yue, Svend Petersen-Mahrt and Göran Akusjärvi Mol. Cell. Biol. 2000, 20(7):2317. DOI: 10.1128/MCB.20.7.2317-2325.2000.
MECHANISMS OF ALTERNATIVE PRE-MESSENGER RNA SPLICING
 Annu. Rev. Biochem. 2003. 72:291 336 doi: 10.1146/annurev.biochem.72.121801.161720 Copyright 2003 by Annual Reviews. All rights reserved First published online as a Review in Advance on February 27, 2003
Annu. Rev. Biochem. 2003. 72:291 336 doi: 10.1146/annurev.biochem.72.121801.161720 Copyright 2003 by Annual Reviews. All rights reserved First published online as a Review in Advance on February 27, 2003
The C-Terminal Region but Not the Arg-X-Pro Repeat of Epstein-Barr Virus Protein EB2 Is Required for Its Effect on RNA Splicing and Transport
 JOURNAL OF VIROLOGY, May 1999, p. 4090 4100 Vol. 73, No. 5 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. The C-Terminal Region but Not the Arg-X-Pro Repeat
JOURNAL OF VIROLOGY, May 1999, p. 4090 4100 Vol. 73, No. 5 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. The C-Terminal Region but Not the Arg-X-Pro Repeat
Where Splicing Joins Chromatin And Transcription. 9/11/2012 Dario Balestra
 Where Splicing Joins Chromatin And Transcription 9/11/2012 Dario Balestra Splicing process overview Splicing process overview Sequence context RNA secondary structure Tissue-specific Proteins Development
Where Splicing Joins Chromatin And Transcription 9/11/2012 Dario Balestra Splicing process overview Splicing process overview Sequence context RNA secondary structure Tissue-specific Proteins Development
Regulation of Alternative Splicing: More than Just the ABCs *
 MINIREVIEW This paper is available online at www.jbc.org THE JOURNA OF BIOOGICA CHEMISTRY VO. 283, NO. 3, pp. 1217 1221, January 18, 2008 2008 by The American Society for Biochemistry and Molecular Biology,
MINIREVIEW This paper is available online at www.jbc.org THE JOURNA OF BIOOGICA CHEMISTRY VO. 283, NO. 3, pp. 1217 1221, January 18, 2008 2008 by The American Society for Biochemistry and Molecular Biology,
Mechanism of splicing
 Outline of Splicing Mechanism of splicing Em visualization of precursor-spliced mrna in an R loop Kinetics of in vitro splicing Analysis of the splice lariat Lariat Branch Site Splice site sequence requirements
Outline of Splicing Mechanism of splicing Em visualization of precursor-spliced mrna in an R loop Kinetics of in vitro splicing Analysis of the splice lariat Lariat Branch Site Splice site sequence requirements
Eukaryotic Gene Regulation
 Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
An Exonic Splicing Enhancer Offsets the Atypical GU-rich 3 Splice Site of Human Apolipoprotein A-II Exon 3*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 279, No. 38, Issue of September 17, pp. 39331 39339, 2004 2004 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. An Exonic
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 279, No. 38, Issue of September 17, pp. 39331 39339, 2004 2004 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. An Exonic
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation
 Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Analyzing mechanisms of alternative pre-mrna splicing using in vitro splicing assays
 Methods 37 (2005) 306 313 www.elsevier.com/locate/ymeth Analyzing mechanisms of alternative pre-mrna splicing using in vitro splicing assays Martin J. Hicks, Bianca J. Lam, Klemens J. Hertel * Department
Methods 37 (2005) 306 313 www.elsevier.com/locate/ymeth Analyzing mechanisms of alternative pre-mrna splicing using in vitro splicing assays Martin J. Hicks, Bianca J. Lam, Klemens J. Hertel * Department
Chapter 10 - Post-transcriptional Gene Control
 Chapter 10 - Post-transcriptional Gene Control Chapter 10 - Post-transcriptional Gene Control 10.1 Processing of Eukaryotic Pre-mRNA 10.2 Regulation of Pre-mRNA Processing 10.3 Transport of mrna Across
Chapter 10 - Post-transcriptional Gene Control Chapter 10 - Post-transcriptional Gene Control 10.1 Processing of Eukaryotic Pre-mRNA 10.2 Regulation of Pre-mRNA Processing 10.3 Transport of mrna Across
MCB Chapter 11. Topic E. Splicing mechanism Nuclear Transport Alternative control modes. Reading :
 MCB Chapter 11 Topic E Splicing mechanism Nuclear Transport Alternative control modes Reading : 419-449 Topic E Michal Linial 14 Jan 2004 1 Self-splicing group I introns were the first examples of catalytic
MCB Chapter 11 Topic E Splicing mechanism Nuclear Transport Alternative control modes Reading : 419-449 Topic E Michal Linial 14 Jan 2004 1 Self-splicing group I introns were the first examples of catalytic
Purine-Rich Exon Sequences Are Not Necessarily Splicing Enhancer Sequence in the Dystrophin Gene
 Kobe J. Med. Sci. 47, 193/202 October 2001 Purine-Rich Exon Sequences Are Not Necessarily Splicing Enhancer Sequence in the Dystrophin Gene TOSHIYUKI ITO 1, YASUHIRO TAKESHIMA 2 *, HIROSHI SAKAMOTO 3,
Kobe J. Med. Sci. 47, 193/202 October 2001 Purine-Rich Exon Sequences Are Not Necessarily Splicing Enhancer Sequence in the Dystrophin Gene TOSHIYUKI ITO 1, YASUHIRO TAKESHIMA 2 *, HIROSHI SAKAMOTO 3,
U2 U6 base-pairing interaction (U2 nt 1-11 with U6 nt 87-97) occurs during the spliceosome cycle (7-9).
 Proc. Natl. Acad. Sci. USA Vol. 90, pp. 7139-7143, August 1993 iochemistry A base-pairing interaction between U2 and small nuclear RNAs occurs in >150S complexes in HeLa cell extracts: Implications for
Proc. Natl. Acad. Sci. USA Vol. 90, pp. 7139-7143, August 1993 iochemistry A base-pairing interaction between U2 and small nuclear RNAs occurs in >150S complexes in HeLa cell extracts: Implications for
Common themes in the function of transcription and splicing enhancers Klemens J Hertel, Kristen W Lynch and Tom Maniatis
 350 Common themes in the function of transcription and splicing enhancers Klemens J Hertel, Kristen W Lynch and Tom Maniatis Regulation of both transcription and RNA splicing requires enhancer elements,
350 Common themes in the function of transcription and splicing enhancers Klemens J Hertel, Kristen W Lynch and Tom Maniatis Regulation of both transcription and RNA splicing requires enhancer elements,
Splicing function of mammalian U6 small nuclear RNA: Conserved
 Proc. Nati. Acad. Sci. SA Vol. 91, pp. 903-907, February 1994 Biochemistry Splicing function of mammalian 6 small nuclear RNA: Conserved positions in central domain and helix I are essential during the
Proc. Nati. Acad. Sci. SA Vol. 91, pp. 903-907, February 1994 Biochemistry Splicing function of mammalian 6 small nuclear RNA: Conserved positions in central domain and helix I are essential during the
The U1 snrnp Base Pairs with the 5 Splice Site within a Penta-snRNP Complex
 MOLECULAR AND CELLULAR BIOLOGY, May 2003, p. 3442 3455 Vol. 23, No. 10 0270-7306/03/$08.00 0 DOI: 10.1128/MCB.23.10.3442 3455.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
MOLECULAR AND CELLULAR BIOLOGY, May 2003, p. 3442 3455 Vol. 23, No. 10 0270-7306/03/$08.00 0 DOI: 10.1128/MCB.23.10.3442 3455.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
Multifactorial Interplay Controls the Splicing Profile of Alu-Derived Exons
 MOLECULAR AND CELLULAR BIOLOGY, May 2008, p. 3513 3525 Vol. 28, No. 10 0270-7306/08/$08.00 0 doi:10.1128/mcb.02279-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Multifactorial
MOLECULAR AND CELLULAR BIOLOGY, May 2008, p. 3513 3525 Vol. 28, No. 10 0270-7306/08/$08.00 0 doi:10.1128/mcb.02279-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Multifactorial
Serine/Arginine-Rich Proteins Contribute to Negative Regulator of Splicing Element-Stimulated Polyadenylation in Rous Sarcoma Virus
 REFERENCES CONTENT ALERTS Serine/Arginine-Rich Proteins Contribute to Negative Regulator of Splicing Element-Stimulated Polyadenylation in Rous Sarcoma Virus Nicole L. Maciolek and Mark T. McNally J. Virol.
REFERENCES CONTENT ALERTS Serine/Arginine-Rich Proteins Contribute to Negative Regulator of Splicing Element-Stimulated Polyadenylation in Rous Sarcoma Virus Nicole L. Maciolek and Mark T. McNally J. Virol.
Genetics, State University of New York at Stony Brook, Stony Brook, NY 11790, USA
 2318-2325 Nucleic Acids Research, 1994, Vol. 22, No. 12 1994 Oxford University Press Alternative splicing of /3-tropomyosin pre-mrna: multiple cis-elements can contribute to the use of the 5'- and 3'-splice
2318-2325 Nucleic Acids Research, 1994, Vol. 22, No. 12 1994 Oxford University Press Alternative splicing of /3-tropomyosin pre-mrna: multiple cis-elements can contribute to the use of the 5'- and 3'-splice
The CUGBP2 Splicing Factor Regulates an Ensemble of Branchpoints from Perimeter Binding Sites with Implications for Autoregulation
 The CUGBP2 Splicing Factor Regulates an Ensemble of Branchpoints from Perimeter Binding Sites with Implications for Autoregulation Jill A. Dembowski, Paula J. Grabowski* Department of Biological Sciences,
The CUGBP2 Splicing Factor Regulates an Ensemble of Branchpoints from Perimeter Binding Sites with Implications for Autoregulation Jill A. Dembowski, Paula J. Grabowski* Department of Biological Sciences,
1) DNA unzips - hydrogen bonds between base pairs are broken by special enzymes.
 Biology 12 Cell Cycle To divide, a cell must complete several important tasks: it must grow, during which it performs protein synthesis (G1 phase) replicate its genetic material /DNA (S phase), and physically
Biology 12 Cell Cycle To divide, a cell must complete several important tasks: it must grow, during which it performs protein synthesis (G1 phase) replicate its genetic material /DNA (S phase), and physically
Deletion of the N-terminus of SF2/ASF Permits RS- Domain-Independent Pre-mRNA Splicing
 Deletion of the N-terminus of SF2/ASF Permits RS- Domain-Independent Pre-mRNA Splicing Stephanie D. Shaw 1,3, Sutapa Chakrabarti 2, Gourisankar Ghosh 2, Adrian R. Krainer 1 * 1 Cold Spring Harbor Laboratory,
Deletion of the N-terminus of SF2/ASF Permits RS- Domain-Independent Pre-mRNA Splicing Stephanie D. Shaw 1,3, Sutapa Chakrabarti 2, Gourisankar Ghosh 2, Adrian R. Krainer 1 * 1 Cold Spring Harbor Laboratory,
Bio 111 Study Guide Chapter 17 From Gene to Protein
 Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
An ATP-independent complex commits pre-mrna to the mammaiian spliceosome assembly pathway
 An ATP-independent complex commits pre-mrna to the mammaiian spliceosome assembly pathway Susan Michaud and Robin Reed Department of Cellular and Molecular Physiology and Program in Cellular and Developmental
An ATP-independent complex commits pre-mrna to the mammaiian spliceosome assembly pathway Susan Michaud and Robin Reed Department of Cellular and Molecular Physiology and Program in Cellular and Developmental
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project
 Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Association of U6 snrna with the 5'-splice site region of pre-mrna in the spliceosome
 Association of U6 snrna with the 5'-splice site region of pre-mrna in the spliceosome Hitoshi Sawa and Yoshiro Shimura^ Department of Biophysics, Faculty of Science, Kyoto University, Kyoto 606, Japan
Association of U6 snrna with the 5'-splice site region of pre-mrna in the spliceosome Hitoshi Sawa and Yoshiro Shimura^ Department of Biophysics, Faculty of Science, Kyoto University, Kyoto 606, Japan
Regulated by the Antagonistic Action of RBM4 and PTB
 Exon Selection in α-tropomyosin mrna Is Regulated by the Antagonistic Action of RBM4 and PTB Jung-Chun Lin and Woan-Yuh Tarn Mol. Cell. Biol. 2005, 25(22):10111. DOI: 10.1128/MCB.25.22.10111-10121.2005.
Exon Selection in α-tropomyosin mrna Is Regulated by the Antagonistic Action of RBM4 and PTB Jung-Chun Lin and Woan-Yuh Tarn Mol. Cell. Biol. 2005, 25(22):10111. DOI: 10.1128/MCB.25.22.10111-10121.2005.
Polypyrimidine Track-binding Protein Binding Downstream of Caspase-2 Alternative Exon 9 Represses Its Inclusion*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 276, No. 11, Issue of March 16, pp. 8535 8543, 2001 2001 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Polypyrimidine Track-binding
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 276, No. 11, Issue of March 16, pp. 8535 8543, 2001 2001 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Polypyrimidine Track-binding
L I F E S C I E N C E S
 1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 NOVEMBER 2, 2006
1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 NOVEMBER 2, 2006
A Bidirectional SF2/ASF- and SRp40-Dependent Splicing Enhancer Regulates Human Immunodeficiency Virus Type 1 rev, env, vpu, and nef Gene Expression
 JOURNAL OF VIROLOGY, June 2004, p. 6517 6526 Vol. 78, No. 12 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.12.6517 6526.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. A Bidirectional
JOURNAL OF VIROLOGY, June 2004, p. 6517 6526 Vol. 78, No. 12 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.12.6517 6526.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. A Bidirectional
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Stability determinants of Murine Cytomegalovirus long non-coding RNA7.2
 JVI Accepts, published online ahead of print on 23 July 2014 J. Virol. doi:10.1128/jvi.01695-14 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 3 Stability determinants of Murine
JVI Accepts, published online ahead of print on 23 July 2014 J. Virol. doi:10.1128/jvi.01695-14 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 3 Stability determinants of Murine
An Increased Specificity Score Matrix for the Prediction of. SF2/ASF-Specific Exonic Splicing Enhancers
 HMG Advance Access published July 6, 2006 1 An Increased Specificity Score Matrix for the Prediction of SF2/ASF-Specific Exonic Splicing Enhancers Philip J. Smith 1, Chaolin Zhang 1, Jinhua Wang 2, Shern
HMG Advance Access published July 6, 2006 1 An Increased Specificity Score Matrix for the Prediction of SF2/ASF-Specific Exonic Splicing Enhancers Philip J. Smith 1, Chaolin Zhang 1, Jinhua Wang 2, Shern
Control of Pre-mRNA Splicing by the General Splicing Factors PUF60 and U2AF 65
 Control of Pre-mRNA Splicing by the General Splicing Factors PUF60 and U2AF 65 Michelle L. Hastings, Eric Allemand, Dominik M. Duelli, Michael P. Myers, Adrian R. Krainer* Cold Spring Harbor Laboratory,
Control of Pre-mRNA Splicing by the General Splicing Factors PUF60 and U2AF 65 Michelle L. Hastings, Eric Allemand, Dominik M. Duelli, Michael P. Myers, Adrian R. Krainer* Cold Spring Harbor Laboratory,
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004
 Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
The Sex-lethal Early Splicing Pattern Uses a Default Mechanism Dependent on the Alternative 5 Splice Sites
 MOLECULAR AND CELLULAR BIOLOGY, Mar. 1997, p. 1674 1681 Vol. 17, No. 3 0270-7306/97/$04.00 0 Copyright 1997, American Society for Microbiology The Sex-lethal Early Splicing Pattern Uses a Default Mechanism
MOLECULAR AND CELLULAR BIOLOGY, Mar. 1997, p. 1674 1681 Vol. 17, No. 3 0270-7306/97/$04.00 0 Copyright 1997, American Society for Microbiology The Sex-lethal Early Splicing Pattern Uses a Default Mechanism
BIO360 Fall 2013 Quiz 1
 BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
UNDERSTANDING THE ROLE OF ATP HYDROLYSIS IN THE SPLICEOSOME
 UNDERSTANDING THE ROLE OF ATP HYDROLYSIS IN THE SPLICEOSOME RNA Splicing Lecture 2, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Eight essential ATPases
UNDERSTANDING THE ROLE OF ATP HYDROLYSIS IN THE SPLICEOSOME RNA Splicing Lecture 2, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Eight essential ATPases
8 Suppression Analysis
 Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 8 Suppression Analysis OVERVIEW Suppression
Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 8 Suppression Analysis OVERVIEW Suppression
Regulation of Alternative Splicing of CD45 by Antagonistic Effects of SR Protein Splicing Factors
 This information is current as of August 23, 2018. References Subscription Permissions Email Alerts Regulation of Alternative Splicing of CD45 by Antagonistic Effects of SR Protein Splicing Factors Gerdy
This information is current as of August 23, 2018. References Subscription Permissions Email Alerts Regulation of Alternative Splicing of CD45 by Antagonistic Effects of SR Protein Splicing Factors Gerdy
A CD45 Polymorphism Associated with Multiple Sclerosis Disrupts an Exonic Splicing Silencer*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 276, No. 26, Issue of June 29, pp. 24341 24347, 2001 2001 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. A CD45 Polymorphism
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 276, No. 26, Issue of June 29, pp. 24341 24347, 2001 2001 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. A CD45 Polymorphism
Negative feedback regulation among SR splicing factors encoded by Rbp1 and Rbp1-like in Drosophila
 The EMBO Journal (2005) 24, 2646 2655 & 2005 European Molecular Biology Organization All Rights Reserved 0261-4189/05 www.embojournal.org Negative feedback regulation among SR splicing factors encoded
The EMBO Journal (2005) 24, 2646 2655 & 2005 European Molecular Biology Organization All Rights Reserved 0261-4189/05 www.embojournal.org Negative feedback regulation among SR splicing factors encoded
TRANSCRIPTION CAPPING
 Messenger RNA Eucaryotic gene expression transcription START site exons upstream elements promoter elements TRANSCRIPTION CAPPING introns gene m 7 G pre-mrna m 7 G SPLICING POLYADENYLATION AAAAAAAAAn
Messenger RNA Eucaryotic gene expression transcription START site exons upstream elements promoter elements TRANSCRIPTION CAPPING introns gene m 7 G pre-mrna m 7 G SPLICING POLYADENYLATION AAAAAAAAAn
Genome-wide Analysis Reveals SR Protein Cooperation and Competition in Regulated Splicing
 Article Genome-wide Analysis Reveals SR Protein Cooperation and Competition in Regulated Splicing Shatakshi Pandit, 1,4 Yu Zhou, 1,4 Lily Shiue, 2,4 Gabriela Coutinho-Mansfield, 1 Hairi Li, 1 Jinsong Qiu,
Article Genome-wide Analysis Reveals SR Protein Cooperation and Competition in Regulated Splicing Shatakshi Pandit, 1,4 Yu Zhou, 1,4 Lily Shiue, 2,4 Gabriela Coutinho-Mansfield, 1 Hairi Li, 1 Jinsong Qiu,
Characterization of Regulatory Mechanisms for Alternative Splicing in Alpha Thyroid Hormone Receptor mrna
 Marquette University e-publications@marquette Master's Theses (2009 -) Dissertations, Theses, and Professional Projects Characterization of Regulatory Mechanisms for Alternative Splicing in Alpha Thyroid
Marquette University e-publications@marquette Master's Theses (2009 -) Dissertations, Theses, and Professional Projects Characterization of Regulatory Mechanisms for Alternative Splicing in Alpha Thyroid
Last time we talked about the few steps in viral replication cycle and the un-coating stage:
 Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Repressive Transcription
 Repressive Transcription The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Guenther, M. G., and R. A.
Repressive Transcription The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Guenther, M. G., and R. A.
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
MRC-Holland MLPA. Description version 29;
 SALSA MLPA KIT P003-B1 MLH1/MSH2 Lot 1209, 0109. As compared to the previous lots 0307 and 1006, one MLH1 probe (exon 19) and four MSH2 probes have been replaced. In addition, one extra MSH2 exon 1 probe,
SALSA MLPA KIT P003-B1 MLH1/MSH2 Lot 1209, 0109. As compared to the previous lots 0307 and 1006, one MLH1 probe (exon 19) and four MSH2 probes have been replaced. In addition, one extra MSH2 exon 1 probe,
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
TRANSCRIPTION. DNA à mrna
 TRANSCRIPTION DNA à mrna Central Dogma Animation DNA: The Secret of Life (from PBS) http://www.youtube.com/watch? v=41_ne5ms2ls&list=pl2b2bd56e908da696&index=3 Transcription http://highered.mcgraw-hill.com/sites/0072507470/student_view0/
TRANSCRIPTION DNA à mrna Central Dogma Animation DNA: The Secret of Life (from PBS) http://www.youtube.com/watch? v=41_ne5ms2ls&list=pl2b2bd56e908da696&index=3 Transcription http://highered.mcgraw-hill.com/sites/0072507470/student_view0/
Life Sciences 1A Midterm Exam 2. November 13, 2006
 Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Balanced splicing at the Tat-specific HIV-1 3 ss A3 is critical for HIV-1 replication
 Erkelenz et al. Retrovirology (2015) 12:29 DOI 10.1186/s12977-015-0154-8 RESEARCH Open Access Balanced splicing at the Tat-specific HIV-1 3 ss A3 is critical for HIV-1 replication Steffen Erkelenz 1, Frank
Erkelenz et al. Retrovirology (2015) 12:29 DOI 10.1186/s12977-015-0154-8 RESEARCH Open Access Balanced splicing at the Tat-specific HIV-1 3 ss A3 is critical for HIV-1 replication Steffen Erkelenz 1, Frank
Figure S1. Molecular confirmation of the precise insertion of the AsMCRkh2 cargo into the kh w locus.
 Supporting Information Appendix Table S1. Larval and adult phenotypes of G 2 progeny of lines 10.1 and 10.2 G 1 outcrosses to wild-type mosquitoes. Table S2. List of oligonucleotide primers. Table S3.
Supporting Information Appendix Table S1. Larval and adult phenotypes of G 2 progeny of lines 10.1 and 10.2 G 1 outcrosses to wild-type mosquitoes. Table S2. List of oligonucleotide primers. Table S3.
The Alternative Choice of Constitutive Exons throughout Evolution
 The Alternative Choice of Constitutive Exons throughout Evolution Galit Lev-Maor 1[, Amir Goren 1[, Noa Sela 1[, Eddo Kim 1, Hadas Keren 1, Adi Doron-Faigenboim 2, Shelly Leibman-Barak 3, Tal Pupko 2,
The Alternative Choice of Constitutive Exons throughout Evolution Galit Lev-Maor 1[, Amir Goren 1[, Noa Sela 1[, Eddo Kim 1, Hadas Keren 1, Adi Doron-Faigenboim 2, Shelly Leibman-Barak 3, Tal Pupko 2,
