The Yeast Two-Hybrid System
|
|
|
- Pauline Manning
- 6 years ago
- Views:
Transcription
1 The Yeast Two-Hybrid System
2 What is the yeast two-hybrid system used for? Identifies novel protein-protein interactions Can identify protein cascades Identifies mutations that affect protein-protein binding Can identify interfering proteins in known interactions (Reverse Two-Hybrid System)
3 How does it work? Uses yeast as a model for eukaryotic protein interactions A library is screened or a protein is characterized using a bait construct Interactions are identified by the transcription of reporter genes Positives are selected using differential media
4 The Model Transcription Activating Region Bait Protein Prey Protein DNA-Binding Domain Reporter Gene DNA-Binding Site
5 Steps to Screen a Library Create the Bait Plasmid Construct from the gene of interest and the DNA binding domain of Gal4 or LexA or other suitable domain Transform with the bait construct a yeast strain lacking the promoter for the reporter genes and select for transformed yeast Transform the yeast again with the library plasmids Select for interaction
6 Sequence analysis Isolate plasmid from yeast and transform E. coli Purify plasmid from E. coli and sequence Blast sequence against database for known proteins or construct a possible protein sequence from the DNA sequence and compare to other proteins
7 Reporter Genes LacZ reporter - Blue/White Screening HIS3 reporter - Screen on His+ media (usually need to add 3AT to increase selectivity) LEU2 reporter - Screen on Leu+ media ADE2 reporter - Screen on Ade+ media URA3 reporter - Screen on Ura+ media (can do negative selection by adding FOA)
8 Plasmid Constructs Plasmids are constructed with the Gal4 DNA binding domain (or other suitable domain) in front of a Multiple Cloning Site (MCS) The plasmid contains genes that can be used for selection such as Amp, Leu2, Ura3, or Trp1
9 Sample Plasmid From Golemis Lab Homepage
10 False Positives False positives are the largest problem with the yeast two-hybrid system Can be caused by: Non-specific binding of the prey Ability to induce transcription without interaction with the bait (Majority of false positives)
11 Elimination of False Positives Sequence Analysis Plasmid Loss Assays Retransformation of both strain with bait plasmid and strain without bait plasmid Test for interaction with an unrelated protein as bait Two (or more) step selections
12 Advantages Immediate availability of the cloned gene of the interacting protein Only a single plasmid construction is required Interactions are detected in vivo Weak, transient interactions can be detected Can accumulate a weak signal over time
13 Examples of Uses of the Yeast Two- Hybrid System Identification of caspase substrates Interaction of Calmodulin and L-Isoaspartyl Methyltransferase Genetic characterization of mutations in E2F1 Peptide hormone-receptor interactions Pha-4 interactions in C. elegans
14 The Two-Hybrid System Bait! Protein! Binding! Domain! Activation! Domain! Prey! Protein! Reporter Gene
15 The Two-Hybrid System Interaction of bait and prey proteins localizes the activation domain to the reporter gene, thus activating transcription. Since the reporter gene typically codes for a survival factor, yeast colonies will grow only when an interaction occurs. Activation! Domain! Prey! Protein! Bait! Protein! Binding! Domain! Reporter Gene
16 Myriad s ProNet Two-Hybrid Process Industrial-scale application of the yeast two-hybrid system Roboticized bait creation process Custom activation domain libraries Efficient mating strategy Automated for quality and throughput Positive sample tracking Robots for media preparation, liquid handling, colony picking Statistical quality control
17 ProNet Process Flowchart 1 4! 3! 5! 7! 8! 0! 1! 2! 6! 9! 10! 23! 24! 2 22! 25! 29! 27! 26! 30! 28! 3 46! 40! 41! 44! 45! 48! 49! 42! 43! 47! 1 Bait Construction 2 Bait Verification 3 Two-Hybrid Screen 4 Identification and Confirmation of Interactions 12! 11! 34! 33! 31! 32! 50! 13! 35! 36! Seq! 51! 56! 14! 15! 16! 37! 52! 53! 57! 17! 38! 39! 54! 55! Seq! 18! 19! 20! 58! 59! 60! 21! 61! 62! 64! 4 63! Seq!
18 Bait Selection A number of criteria were taken into consideration in the selection of bait protein candidates: Proteins known to be expressed in B cells and/or cardiac myocytes. Proteins known or suspected to be involved in signaling pathways in B cells and/or cardiac myocytes. Proteins believed important in GPCR or BCR signaling leading to PIP3 generation. These were selected first in an attempt to define interactions within a focused network of signaling molecules. Preference was also given to proteins that had a higher likelihood of success in the yeast two-hybrid assay proteins with well-defined modular domains exhibiting secondary structure suggestive of involvement in protein-protein interactions.
19 Bait Design Fragments of proteins representing folded domains are often more effective than the full-length protein in identifying physiologically relevant interactions If the domain structure of a given bait protein was already established, the specific baits were designed to represent one or more folded domains. For cases in which domain structure was not available, a variety of secondary structure prediction algorithms were used to predict domains and thus direct bait design. were designed to cover the entire protein, with several overlapping fragments, as not all baits will work effectively. Phosphorylation sites Proline-rich SH3 binding BLNK Src homology 2 domain P! P! P! Pro! Pro! SH2! 110" 116" 134" 146" 350" 436" (P sites)" (linker)" (N-term)" (SH2)" (Pro regions)" (C-term)"
20 B Cell Receptor Pathway
21 B Cell Receptor Pathway
22 B Cell Receptor Pathway z Endophilin2 BAN-P PDE4B3 Nck Pak - Pix/Cool Ruk1 Git2 PP2C gamma
23 What can we learn from the larger data set? 170 interactions derived from just 16 bait proteins Limited interconnectivity We now have 473 interactions from 55 bait proteins How can the average biologist analyze this data set? With difficulty! All 473 interactions uploaded to Access database table Query: which prey IDs are correlated with more than one bait ID?
24 Interaction examples:1 AFCS ID Bait Protein Name Prey Nuc GI Prey ID Prey Protein Name A Grb A Blnk A Calcium-dependent protein kinase II alpha A Blnk A Grb A Blnk A Grb A Blnk A Grb A Blnk AFCS ID Bait Protein Name Prey Nuc GI Prey ID Prey Protein Name A cdc A Pak1 A PIX/COOL A Pak1 AFCS ID Bait Protein Name Prey Nuc GI Prey ID Prey Protein Name A Integrin beta A Nucleoside diphosphate kinase B A Integrin beta A Nucleoside diphosphate kinase B A Ryanodine receptor type I A Nucleoside diphosphate kinase B A PI3 kinase p85 regulatory subunit (alpha) A Nucleoside diphosphate kinase B A Dbl A Nucleoside diphosphate kinase B A Integrin beta A Nucleoside diphosphate kinase B AFCS ID Bait Protein Name Prey Nuc GI Prey ID Prey Protein Name A PI3 kinase p85 regulatory subunit (alpha) A Endophilin 2 A PI3K p85beta A Endophilin 2 A BLNK A Endophilin 2 AFCS ID Bait Protein Name Prey Nuc GI Prey ID Prey Protein Name A Nck A Bcap A Bcap A Bcap A Bcap A Bcap A Bcap A Bcap A Grb A Bcap A Grb A Bcap
25 Interaction examples: 2 AFCS ID Bait Protein Name Prey Nuc GI Prey ID A Lyn A CD AFCS ID Bait Protein Name Prey Nuc GI Prey ID A phospholipase C gamma A Rac AFCS ID Bait Bait Protein Name Prey Nuc GI Prey ID A phospholipase C gamma A phospholipase C gamma A phospholipase C gamma A phospholipase C gamma A phospholipase C gamma A Grb A Calcium-dependent protein kinase II alpha A Calcium-dependent protein kinase II alpha A phospholipase C gamma A phospholipase C gamma AFCS ID Bait Protein Name Prey Nuc GI Prey ID A vav A Grb A Grb A Grb A Nck AFCS ID Bait Protein Name Prey Nuc GI Prey ID A Wasp A Nck A Phosphatidyl-inositol 3'-kinase gamma A Nck A Wasp A Wasp
26
27 CamK II SOS2 CD19 CD22 Dbl Btk Fyn cdc42 CAP WISH NdkB PI3Kγ (p110) Actin Cytoskeleton PDK1
28 AIP Cbl-b CamK II SOS2 CD19 CD Dbl Btk Fyn cdc42 CAP WISH NdkB PI3Kγ (p110) Sam68 Actin Cytoskeleton PDK Protein 4.1G
29 Caveats! Y2H data requires validation by secondary assay Protein complementation Pull-downs Didn t observe source library in data analysis Didn t analyze all bait and prey coordinates to map sites of interaction Cant assume multi-protein complexes since proteins may be competing for same site of interaction This level of analysis requires more sophisticated computational approach
30 Bait Status Protein name Date Submitted AFCS ID Ordered Created Searched Status adenylyl cyclase V 11-Jul-01 A No Interactions N-Ras 12-Jul-02 A Screens in progress K-Ras 4-Apr-02 A Data Released p115rhogef 11-Jul-01 A Data Released Beta arrestin 1 11-Jul-01 A Data Released RGS Jul-02 A Screens in progress NFATc1 (AKA NFAT2) 8-Aug-01 A Data Released RGS 2 11-Jul-01 A No Interactions G alpha13 11-Jul-01 A No Interactions RGS Jul-02 A Screens in progress btk 25-Jul-01 A Data Released G alpha12 13-Jul-01 A No Interactions PKG 23-Jul-02 A No Interactions Protein kinase A RI alpha 10-Oct-02 A Screens in progress Protein kinase A RII beta 10-Oct-02 A No Interactions PAK1 11-Jul-01 A Data Released Acetylcholine (muscarinic) 10-Oct-02 A Screens in progress N-WASp 11-Jul-01 A Data Released RGS Sep-01 A Data Released I kappa B alpha 11-Jul-01 A No Interactions PDE4B3 11-Jul-01 A No Interactions G alpha i3 11-Jul-01 A Data Released Nck1 13-Sep-01 A Data Released Inositol (1,4,5,) P3 receptor 2 11-Jul-01 A No Interactions Wasp 4-Apr-02 A Data Released dock2 1-Feb-02 A Data Released RasGap p Oct-02 A Screens in progress Adrenergic receptor beta 1 10-Oct-02 A No Interactions Adrenergic receptor beta 3 10-Oct-02 A No Interactions AKAP12 11-Jul-01 A Data Released PKB-alpha (= Akt1) 8-Aug-01 A Data Released Akt2 1-Feb-02 A Data Released Akt3 10-Oct-02 A No Interactions AP-3 beta3a 10-Oct-02 A Screens in progress Bam32 13-Sep-01 A No Interactions Bcap 4-Apr-02 A Data Released Blk 12-Jul-02 A No Interactions BLNK 13-Jul-01 A Data Released calcineurin A-subunit 23-Jul-02 A No Interactions Calcineurin A beta 1-Feb-02 A No Interactions Calcium channel voltage- 1-Feb-02 A Data Released Calcium channel voltage- 8-Aug-01 A Screens in progress Calmodulin III 25-Jul-01 A No Interactions CaMKI 23-Jul-02 A No Interactions Calcium-dependent protein 1-Feb-02 A Data Released Calcium-dependent protein 1-Feb-02 A No Interactions caveolin3 13-Sep-01 A No Interactions Cbl 8-Aug-01 A No Interactions GRP-1 4-Apr-02 A No Interactions p70s6k 10-Oct-02 A No Interactions CD19 4-Apr-02 A Data Released CD21 12-Jul-02 A Screens in progress CD22 12-Jul-02 A Screens in progress CD32 4-Apr-02 A No Interactions cd44 1-Feb-02 A No Interactions Protein name Date Submitted AFCS ID Ordered Created Searched Status CD45 4-Apr-02 A No Interactions CD79a 4-Apr-02 A No Interactions CD79b 4-Apr-02 A Screens in progress CD81 12-Jul-02 A No Interactions cdc42 1-Feb-02 A Data Released CCR7 1-Feb-02 A No Interactions CXCR4 1-Feb-02 A No Interactions Chemokine receptor CXCR5 4-Apr-02 A Data Released PIX/COOL 13-Jul-01 A Data Released Dock1 12-Jul-02 A Screens in progress Dbl 13-Sep-01 A Data Released Dbs 10-Oct-02 A Screens in progress Dok2 12-Jul-02 A No Interactions CD150 4-Apr-02 A No Interactions Fyn 12-Jul-02 A Screens in progress G protein alpha i2 4-Apr-02 A No Interactions G protein beta 1 4-Apr-02 A No Interactions G protein beta 2 4-Apr-02 A No Interactions G protein gamma 2 4-Apr-02 A No Interactions GAB-1 23-Apr-02 A Data Released Emk 10-Oct-02 A Screens in progress Grb2 13-Sep-01 A Data Released GRK2 11-Jul-01 A No Interactions glycogen synthase kinase- 23-Jul-02 A No Interactions HDAC5 23-Jul-02 A No Interactions Integrin beta 2 4-Apr-02 A Data Released Integrin beta 1 4-Apr-02 A Data Released Kv Oct-02 A No Interactions Kv1.2 1-Feb-02 A Data Released Kv Oct-02 A No Interactions Kv Oct-02 A No Interactions Kv Oct-02 A No Interactions Lck 4-Apr-02 A Data Released Lyn 8-Aug-01 A Data Released Mcip-1 1-Feb-02 A Data Released MEF2A 23-Jul-02 A No Interactions MEF 2C 1-Feb-02 A Data Released MEF2D 23-Jul-02 A Screens in progress MEK5 23-Jul-02 A Screens in progress MKK6 23-Jul-02 A No Interactions Mitr 23-Jul-02 A Screens in progress Na/Ca exchanger 11-Jul-01 A No Interactions Nav Oct-02 A Screens in progress Neurofibromin 2 10-Oct-02 A Screens in progress NFAT2 8-Aug-01 A No Interactions NFAT3 23-Jul-02 A No Interactions NFkappaB p65 4-Apr-02 A Data Released P Jul-02 A Screens in progress p38 MAPK 23-Jul-02 A Screens in progress HDAC7 23-Jul-02 A Screens in progress G2A receptor 13-Sep-01 A No Interactions PDK1 25-Jul-01 A Data Released Phosphoinositide 3-kinase 4-Apr-02 A No Interactions Phosphatidyl-inositol 3'-kinase 8-Aug-01 A Data Released Protein name Date Submitted AFCS ID Ordered Created Searched Status p110 beta class Ia 8-Aug-01 A Data Released Phosphoinositide 3-kinase 12-Jul-02 A Screens in progress Phosphatidyl-inositol 3'-kinase 13-Jul-01 A Data Released PI3 kinase p85 regulatory 8-Aug-01 A Data Released PI3K p85beta 1-Feb-02 A Data Released Phosphoinositide 4-phosphate 10-Oct-02 A Screens in progress Phosphoinositide 4-phosphate 10-Oct-02 A No Interactions Phosphoinositide 4-phosphate 10-Oct-02 A No Interactions Phosphoinositide 5-phosphate 10-Oct-02 A Data Released Phospholamban 8-Aug-01 A No Interactions Phospholipase C beta 2 11-Jul-01 A Data Released phospholipase C gamma1 13-Jul-01 A Data Released phospholipase C gamma2 13-Jul-01 A Data Released PKA catalytic subunit alpha 8-Aug-01 A No Interactions Protein kinase A RI beta 10-Oct-02 A Screens in progress Protein kinase A RII alpha 10-Oct-02 A Screens in progress Protein kinase C beta1 11-Jul-01 A No Interactions PKC-epsilon 11-Jul-01 A No Interactions PKC zeta 25-Jul-01 A Data Released PTEN 13-Jul-01 A No Interactions Rac1 13-Sep-01 A Data Released Rac2 13-Jul-01 A No Interactions Rac3 12-Jul-02 A Screens in progress RasGEF1 10-Oct-02 A Screens in progress RasGEF2 10-Oct-02 A No Interactions RasGRF1 10-Oct-02 A Screens in progress RGS 1 11-Jul-01 A Data Released RGS Jul-02 A No Interactions RGS Jul-02 A No Interactions RGS 3 8-Aug-01 A No Interactions RhoA 1-Feb-02 A Data Released ROCK1 2-Aug-01 A No Interactions Rock2 10-Oct-02 A No Interactions p90rsk1 23-Apr-02 A Data Released Ryanodine receptor type I 13-Sep-01 A Data Released Ryanodine receptor Type II 1-Feb-02 A Data Released S100A1 1-Feb-02 A No Interactions SRF 23-Jul-02 A No Interactions Shc 4-Apr-02 A Data Released SHIP1 8-Aug-01 A No Interactions SHIP2 1-Feb-02 A No Interactions SHP-1 4-Apr-02 A Data Released Tec 23-Apr-02 A No Interactions Tumor necrosis factor 10-Oct-02 A Screens in progress Vav1 12-Jul-02 A Screens in progress vav2 1-Feb-02 A Data Released Vav3 12-Jul-02 A No Interactions RasGRF2 10-Oct-02 A Screens in progress Abi1 10-Oct-02 A Screens in progress Scar-2/WAVE-2 9-Aug-01 A Data Released HDAC4 23-Jul-02 A Screens in progress Gpr90 23-Jul-02 A No Interactions Fmr1 10-Oct-02 A Screens in progress PRex1 10-Oct-02 A No Interactions
31 Bait Status Protein name Date Submitted AFCS ID Ordered Created Searched Status K-Ras 4-Apr-02 A Data Released p115rhogef 11-Jul-01 A Data Released Beta arrestin 1 11-Jul-01 A Data Released NFATc1 (AKA NFAT2) 8-Aug-01 A Data Released btk 25-Jul-01 A Data Released PAK1 11-Jul-01 A Data Released N-WASp 11-Jul-01 A Data Released RGS Sep-01 A Data Released G alpha i3 11-Jul-01 A Data Released Nck1 13-Sep-01 A Data Released Wasp 4-Apr-02 A Data Released dock2 1-Feb-02 A Data Released AKAP12 11-Jul-01 A Data Released PKB-alpha (= Akt1) 8-Aug-01 A Data Released Akt2 1-Feb-02 A Data Released Bcap 4-Apr-02 A Data Released BLNK 13-Jul-01 A Data Released Calcium channel voltage- 1-Feb-02 A Data Released Calcium-dependent protein 1-Feb-02 A Data Released CD19 4-Apr-02 A Data Released cdc42 1-Feb-02 A Data Released Chemokine receptor CXCR5 4-Apr-02 A Data Released PIX/COOL 13-Jul-01 A Data Released Dbl 13-Sep-01 A Data Released GAB-1 23-Apr-02 A Data Released Grb2 13-Sep-01 A Data Released Integrin beta 2 4-Apr-02 A Data Released Integrin beta 1 4-Apr-02 A Data Released Kv1.2 1-Feb-02 A Data Released Lck 4-Apr-02 A Data Released Lyn 8-Aug-01 A Data Released Mcip-1 1-Feb-02 A Data Released MEF 2C 1-Feb-02 A Data Released NFkappaB p65 4-Apr-02 A Data Released PDK1 25-Jul-01 A Data Released Phosphatidyl-inositol 3'-kinase 8-Aug-01 A Data Released Phosphatidyl-inositol 3'-kinase 8-Aug-01 A Data Released Phosphatidyl-inositol 3'-kinase 13-Jul-01 A Data Released PI3 kinase p85 regulatory 8-Aug-01 A Data Released PI3K p85beta 1-Feb-02 A Data Released Phosphoinositide 5-phosphate 10-Oct-02 A Data Released Phospholipase C beta 2 11-Jul-01 A Data Released phospholipase C gamma1 13-Jul-01 A Data Released phospholipase C gamma2 13-Jul-01 A Data Released PKC zeta 25-Jul-01 A Data Released Rac1 13-Sep-01 A Data Released RGS 1 11-Jul-01 A Data Released RhoA 1-Feb-02 A Data Released p90rsk1 23-Apr-02 A Data Released Ryanodine receptor type I 13-Sep-01 A Data Released Ryanodine receptor Type II 1-Feb-02 A Data Released Shc 4-Apr-02 A Data Released SHP-1 4-Apr-02 A Data Released vav2 1-Feb-02 A Data Released Scar-2/WAVE-2 9-Aug-01 A Data Released Protein name Date Submitted AFCS ID Ordered Created Searched Status Abi1 10-Oct-02 A Screens in progress Acetylcholine (muscarinic) 10-Oct-02 A Screens in progress AP-3 beta3a 10-Oct-02 A Screens in progress Calcium channel voltage- 8-Aug-01 A Screens in progress CD21 12-Jul-02 A Screens in progress CD22 12-Jul-02 A Screens in progress CD79b 4-Apr-02 A Screens in progress Dbs 10-Oct-02 A Screens in progress Dock1 12-Jul-02 A Screens in progress Emk 10-Oct-02 A Screens in progress Fmr1 10-Oct-02 A Screens in progress Fyn 12-Jul-02 A Screens in progress HDAC4 23-Jul-02 A Screens in progress HDAC7 23-Jul-02 A Screens in progress MEF2D 23-Jul-02 A Screens in progress MEK5 23-Jul-02 A Screens in progress Mitr 23-Jul-02 A Screens in progress Nav Oct-02 A Screens in progress Neurofibromin 2 10-Oct-02 A Screens in progress N-Ras 12-Jul-02 A Screens in progress P Jul-02 A Screens in progress p38 MAPK 23-Jul-02 A Screens in progress Phosphoinositide 3-kinase 12-Jul-02 A Screens in progress Phosphoinositide 4-phosphate 10-Oct-02 A Screens in progress Protein kinase A RI alpha 10-Oct-02 A Screens in progress Protein kinase A RI beta 10-Oct-02 A Screens in progress Protein kinase A RII alpha 10-Oct-02 A Screens in progress Rac3 12-Jul-02 A Screens in progress RasGap p Oct-02 A Screens in progress RasGEF1 10-Oct-02 A Screens in progress RasGRF1 10-Oct-02 A Screens in progress RasGRF2 10-Oct-02 A Screens in progress RGS Jul-02 A Screens in progress RGS Jul-02 A Screens in progress Tumor necrosis factor 10-Oct-02 A Screens in progress Vav1 12-Jul-02 A Screens in progress adenylyl cyclase V 11-Jul-01 A No Interactions Adrenergic receptor beta 1 10-Oct-02 A No Interactions Adrenergic receptor beta 3 10-Oct-02 A No Interactions Akt3 10-Oct-02 A No Interactions Bam32 13-Sep-01 A No Interactions Blk 12-Jul-02 A No Interactions Calcineurin A beta 1-Feb-02 A No Interactions calcineurin A-subunit 23-Jul-02 A No Interactions Calcium-dependent protein 1-Feb-02 A No Interactions Calmodulin III 25-Jul-01 A No Interactions CaMKI 23-Jul-02 A No Interactions caveolin3 13-Sep-01 A No Interactions Cbl 8-Aug-01 A No Interactions CCR7 1-Feb-02 A No Interactions CD150 4-Apr-02 A No Interactions CD32 4-Apr-02 A No Interactions cd44 1-Feb-02 A No Interactions CD45 4-Apr-02 A No Interactions Protein name Date Submitted AFCS ID Ordered Created Searched Status CD79a 4-Apr-02 A No Interactions CD81 12-Jul-02 A No Interactions CXCR4 1-Feb-02 A No Interactions Dok2 12-Jul-02 A No Interactions G alpha12 13-Jul-01 A No Interactions G alpha13 11-Jul-01 A No Interactions G protein alpha i2 4-Apr-02 A No Interactions G protein beta 1 4-Apr-02 A No Interactions G protein beta 2 4-Apr-02 A No Interactions G protein gamma 2 4-Apr-02 A No Interactions G2A receptor 13-Sep-01 A No Interactions glycogen synthase kinase- 23-Jul-02 A No Interactions Gpr90 23-Jul-02 A No Interactions GRK2 11-Jul-01 A No Interactions GRP-1 4-Apr-02 A No Interactions HDAC5 23-Jul-02 A No Interactions I kappa B alpha 11-Jul-01 A No Interactions Inositol (1,4,5,) P3 receptor 2 11-Jul-01 A No Interactions Kv Oct-02 A No Interactions Kv Oct-02 A No Interactions Kv Oct-02 A No Interactions Kv Oct-02 A No Interactions MEF2A 23-Jul-02 A No Interactions MKK6 23-Jul-02 A No Interactions Na/Ca exchanger 11-Jul-01 A No Interactions NFAT2 8-Aug-01 A No Interactions NFAT3 23-Jul-02 A No Interactions p70s6k 10-Oct-02 A No Interactions PDE4B3 11-Jul-01 A No Interactions Phosphoinositide 3-kinase 4-Apr-02 A No Interactions Phosphoinositide 4-phosphate 10-Oct-02 A No Interactions Phosphoinositide 4-phosphate 10-Oct-02 A No Interactions Phospholamban 8-Aug-01 A No Interactions PKA catalytic subunit alpha 8-Aug-01 A No Interactions PKC-epsilon 11-Jul-01 A No Interactions PKG 23-Jul-02 A No Interactions PRex1 10-Oct-02 A No Interactions Protein kinase A RII beta 10-Oct-02 A No Interactions Protein kinase C beta1 11-Jul-01 A No Interactions PTEN 13-Jul-01 A No Interactions Rac2 13-Jul-01 A No Interactions RasGEF2 10-Oct-02 A No Interactions RGS Jul-02 A No Interactions RGS 2 11-Jul-01 A No Interactions RGS Jul-02 A No Interactions RGS 3 8-Aug-01 A No Interactions ROCK1 2-Aug-01 A No Interactions Rock2 10-Oct-02 A No Interactions S100A1 1-Feb-02 A No Interactions SHIP1 8-Aug-01 A No Interactions SHIP2 1-Feb-02 A No Interactions SRF 23-Jul-02 A No Interactions Tec 23-Apr-02 A No Interactions Vav3 12-Jul-02 A No Interactions
32 Pattern Among Negative? Of 72 negative baits, 45 are either membrane proteins (receptors, cell-surface antigens, channels) or are membrane associated (G proteins, GEFs, RGS etc.) May be overcome by generous and creative bait design but these proteins will always give lower return in Y2H assay Also several proteins that could be expected to do better (cytosolic protein kinases and some adaptors) Further baits advisable Future baits sets should also include some of novel preys identified in primary screen
33 References Bartel, Paul, C. Chien, R. Sternglanz, S. Fields. Elimination of False Positives that Arise in Using the Two- Hybrid System. Biotechniques (1993) Vol. 14, no. 6, p Chien, Cheng-ting, P. Bartel, R. Sternglanz, S. Fields. The two-hybrid system: A method to identify and clone genes for proteins that interact with a protein of interest. Proc. Natl. Acad. Sci. USA (1991) Vol. 88, p Fields, Stanley, O. Song. "A novel genetic system to detect protein-protein interactions." Nature (1989) Vol. 340, p James, Philip, J. Halladay, E. Craig. "Genomic Libraries and a Host Strain Designed for Highly Efficient Two-Hybrid Selection in Yeast." Genetics (1996) Vol. 144, p Kamada, S, H. Kusano, H. Fujita, M. Ohtsu, R. Koya, N. Kuzumaki, Y. Tsujimoto. "A cloning method for caspase substrates that uses the yeast two-hybrid system: Cloning of the antiapoptotic gene gelsolin." Proc. Natl. Acad. Sci. USA (1998) Vol 95, p O'Connor, Mirriam, C. O'Connor. "Complex Interactions of the Protein L-Isoaspartyl Methyltransferase and Calmodulin Revealed with the Yeast Two-hybrid System." The Journal of Biological Chemistry (1998) Vol. 273, p Staudinger, Jeff, J. Zhou, R. Burgess, S. Elledge, E. Olson. "PICK1: A Perinuclear Binding Protein and Substrate for Protein Kinase C Isolated by the Yeast Two-Hybrid System." The Journal of Cell Biology (1995) Vol. 128, p
34 References continued Vidal, Marc, P. Braun, E. Chen, J. Boeke, E. Harlow. "Genetic Characterization of a mammalian proteinprotein interaction domain by using a yeast reverse two-hybrid system." Proc. Natl. Acad. Sci. USA (1996) Vol. 93, p White, Michael. "The yeast two-hybrid system: Forward and reverse." Proc. Natl. Acad. Sci. USA (1996) Vol 93, p Zhu, Jianwei, C. Kahn. "Analysis of a peptide hormone-receptor interaction in the yeast two-hybrid system." Proc. Natl. Acad. Sci. USA (1997) Vol. 94, p Lab of Erica Golemis Special thanks to Dr. Susan Mango and the University of Utah
Signaling Through Immune System Receptors (Ch. 7)
 Signaling Through Immune System Receptors (Ch. 7) 1. General principles of signal transduction and propagation. 2. Antigen receptor signaling and lymphocyte activation. 3. Other receptors and signaling
Signaling Through Immune System Receptors (Ch. 7) 1. General principles of signal transduction and propagation. 2. Antigen receptor signaling and lymphocyte activation. 3. Other receptors and signaling
The Tissue Engineer s Toolkit
 The Tissue Engineer s Toolkit Stimuli Detection and Response Ken Webb, Ph. D. Assistant Professor Dept. of Bioengineering Clemson University Environmental Stimulus-Cellular Response Environmental Stimuli
The Tissue Engineer s Toolkit Stimuli Detection and Response Ken Webb, Ph. D. Assistant Professor Dept. of Bioengineering Clemson University Environmental Stimulus-Cellular Response Environmental Stimuli
Receptor mediated Signal Transduction
 Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
MCB*4010 Midterm Exam / Winter 2008
 MCB*4010 Midterm Exam / Winter 2008 Name: ID: Instructions: Answer all 4 questions. The number of marks for each question indicates how many points you need to provide. Write your answers in point form,
MCB*4010 Midterm Exam / Winter 2008 Name: ID: Instructions: Answer all 4 questions. The number of marks for each question indicates how many points you need to provide. Write your answers in point form,
Signal Transduction: G-Protein Coupled Receptors
 Signal Transduction: G-Protein Coupled Receptors Federle, M. (2017). Lectures 4-5: Signal Transduction parts 1&2: nuclear receptors and GPCRs. Lecture presented at PHAR 423 Lecture in UIC College of Pharmacy,
Signal Transduction: G-Protein Coupled Receptors Federle, M. (2017). Lectures 4-5: Signal Transduction parts 1&2: nuclear receptors and GPCRs. Lecture presented at PHAR 423 Lecture in UIC College of Pharmacy,
Cellular Signaling Pathways. Signaling Overview
 Cellular Signaling Pathways Signaling Overview Signaling steps Synthesis and release of signaling molecules (ligands) by the signaling cell. Transport of the signal to the target cell Detection of the
Cellular Signaling Pathways Signaling Overview Signaling steps Synthesis and release of signaling molecules (ligands) by the signaling cell. Transport of the signal to the target cell Detection of the
KEY CONCEPT QUESTIONS IN SIGNAL TRANSDUCTION
 Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Cell Signaling part 2
 15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
Biosignals, Chapter 8, rearranged, Part I
 Biosignals, Chapter 8, rearranged, Part I Nicotinic Acetylcholine Receptor: A Ligand-Binding Ion Channel Classes of Receptor Proteins in Eukaryotes, Heterotrimeric G Proteins Signaling View the Heterotrimeric
Biosignals, Chapter 8, rearranged, Part I Nicotinic Acetylcholine Receptor: A Ligand-Binding Ion Channel Classes of Receptor Proteins in Eukaryotes, Heterotrimeric G Proteins Signaling View the Heterotrimeric
G-Protein Signaling. Introduction to intracellular signaling. Dr. SARRAY Sameh, Ph.D
 G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
Revision. camp pathway
 االله الرحمن الرحيم بسم Revision camp pathway camp pathway Revision camp pathway Adenylate cyclase Adenylate Cyclase enzyme Adenylate cyclase catalyses the formation of camp from ATP. Stimulation or inhibition
االله الرحمن الرحيم بسم Revision camp pathway camp pathway Revision camp pathway Adenylate cyclase Adenylate Cyclase enzyme Adenylate cyclase catalyses the formation of camp from ATP. Stimulation or inhibition
Introduction! Introduction! Introduction! Chem Lecture 10 Signal Transduction & Sensory Systems Part 2
 Chem 452 - Lecture 10 Signal Transduction & Sensory Systems Part 2 Questions of the Day: How does the hormone insulin trigger the uptake of glucose in the cells that it targets. Introduction! Signal transduction
Chem 452 - Lecture 10 Signal Transduction & Sensory Systems Part 2 Questions of the Day: How does the hormone insulin trigger the uptake of glucose in the cells that it targets. Introduction! Signal transduction
The elements of G protein-coupled receptor systems
 The elements of G protein-coupled receptor systems Prostaglandines Sphingosine 1-phosphate a receptor that contains 7 membrane-spanning domains a coupled trimeric G protein which functions as a switch
The elements of G protein-coupled receptor systems Prostaglandines Sphingosine 1-phosphate a receptor that contains 7 membrane-spanning domains a coupled trimeric G protein which functions as a switch
Signal Transduction I
 Signal Transduction I Prof. Tianhua Zhou Department of Cell Biology Zhejiang University School of Medicine Office hours by appointment tzhou@zju.edu.cn Signal transduction: Key contents for signal transduction:
Signal Transduction I Prof. Tianhua Zhou Department of Cell Biology Zhejiang University School of Medicine Office hours by appointment tzhou@zju.edu.cn Signal transduction: Key contents for signal transduction:
Regulation of cell function by intracellular signaling
 Regulation of cell function by intracellular signaling Objectives: Regulation principle Allosteric and covalent mechanisms, Popular second messengers, Protein kinases, Kinase cascade and interaction. regulation
Regulation of cell function by intracellular signaling Objectives: Regulation principle Allosteric and covalent mechanisms, Popular second messengers, Protein kinases, Kinase cascade and interaction. regulation
Signal Transduction Pathways. Part 2
 Signal Transduction Pathways Part 2 GPCRs G-protein coupled receptors > 700 GPCRs in humans Mediate responses to senses taste, smell, sight ~ 1000 GPCRs mediate sense of smell in mouse Half of all known
Signal Transduction Pathways Part 2 GPCRs G-protein coupled receptors > 700 GPCRs in humans Mediate responses to senses taste, smell, sight ~ 1000 GPCRs mediate sense of smell in mouse Half of all known
T cell maturation. T-cell Maturation. What allows T cell maturation?
 T-cell Maturation What allows T cell maturation? Direct contact with thymic epithelial cells Influence of thymic hormones Growth factors (cytokines, CSF) T cell maturation T cell progenitor DN DP SP 2ry
T-cell Maturation What allows T cell maturation? Direct contact with thymic epithelial cells Influence of thymic hormones Growth factors (cytokines, CSF) T cell maturation T cell progenitor DN DP SP 2ry
Enzyme-coupled Receptors. Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors
 Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
INTERACTION DRUG BODY
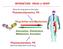 INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
Signal Transduction Cascades
 Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Chem Lecture 10 Signal Transduction
 Chem 452 - Lecture 10 Signal Transduction 111130 Here we look at the movement of a signal from the outside of a cell to its inside, where it elicits changes within the cell. These changes are usually mediated
Chem 452 - Lecture 10 Signal Transduction 111130 Here we look at the movement of a signal from the outside of a cell to its inside, where it elicits changes within the cell. These changes are usually mediated
Principles of Genetics and Molecular Biology
 Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
Chapter 15: Signal transduction
 Chapter 15: Signal transduction Know the terminology: Enzyme-linked receptor, G-protein linked receptor, nuclear hormone receptor, G-protein, adaptor protein, scaffolding protein, SH2 domain, MAPK, Ras,
Chapter 15: Signal transduction Know the terminology: Enzyme-linked receptor, G-protein linked receptor, nuclear hormone receptor, G-protein, adaptor protein, scaffolding protein, SH2 domain, MAPK, Ras,
Lecture: CHAPTER 13 Signal Transduction Pathways
 Lecture: 10 17 2016 CHAPTER 13 Signal Transduction Pathways Chapter 13 Outline Signal transduction cascades have many components in common: 1. Release of a primary message as a response to a physiological
Lecture: 10 17 2016 CHAPTER 13 Signal Transduction Pathways Chapter 13 Outline Signal transduction cascades have many components in common: 1. Release of a primary message as a response to a physiological
BL 424 Chapter 15: Cell Signaling; Signal Transduction
 BL 424 Chapter 15: Cell Signaling; Signal Transduction All cells receive and respond to signals from their environments. The behavior of each individual cell in multicellular plants and animals must be
BL 424 Chapter 15: Cell Signaling; Signal Transduction All cells receive and respond to signals from their environments. The behavior of each individual cell in multicellular plants and animals must be
2013 W. H. Freeman and Company. 12 Signal Transduction
 2013 W. H. Freeman and Company 12 Signal Transduction CHAPTER 12 Signal Transduction Key topics: General features of signal transduction Structure and function of G protein coupled receptors Structure
2013 W. H. Freeman and Company 12 Signal Transduction CHAPTER 12 Signal Transduction Key topics: General features of signal transduction Structure and function of G protein coupled receptors Structure
What would you observe if you fused a G1 cell with a S cell? A. Mitotic and pulverized chromosomes. B. Mitotic and compact G1 chromosomes.
 What would you observe if you fused a G1 cell with a S cell? A. Mitotic and pulverized chromosomes. B. Mitotic and compact G1 chromosomes. C. Mostly non-compact G1 chromosomes. D. Compact G1 and G2 chromosomes.
What would you observe if you fused a G1 cell with a S cell? A. Mitotic and pulverized chromosomes. B. Mitotic and compact G1 chromosomes. C. Mostly non-compact G1 chromosomes. D. Compact G1 and G2 chromosomes.
IMMUNORECEPTOR SIGNALING AS ATOPIC OF RESEARCH has a history of about 30 years. Though the effects of
 Preface IMMUNORECEPTOR SIGNALING AS ATOPIC OF RESEARCH has a history of about 30 years. Though the effects of plant lectins on lymphocyte activation were studied back in the 1960s, the modern era began
Preface IMMUNORECEPTOR SIGNALING AS ATOPIC OF RESEARCH has a history of about 30 years. Though the effects of plant lectins on lymphocyte activation were studied back in the 1960s, the modern era began
Supplementary Material. Part I: Sample Information. Part II: Pathway Information
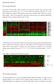 Supplementary Material Part I: Sample Information Three NPC cell lines, CNE1, CNE2, and HK1 were treated with CYC202. Gene expression of 380 selected genes were collected at 0, 2, 4, 6, 12 and 24 hours
Supplementary Material Part I: Sample Information Three NPC cell lines, CNE1, CNE2, and HK1 were treated with CYC202. Gene expression of 380 selected genes were collected at 0, 2, 4, 6, 12 and 24 hours
Protein kinases are enzymes that add a phosphate group to proteins according to the. ATP + protein OH > Protein OPO 3 + ADP
 Protein kinase Protein kinases are enzymes that add a phosphate group to proteins according to the following equation: 2 ATP + protein OH > Protein OPO 3 + ADP ATP represents adenosine trisphosphate, ADP
Protein kinase Protein kinases are enzymes that add a phosphate group to proteins according to the following equation: 2 ATP + protein OH > Protein OPO 3 + ADP ATP represents adenosine trisphosphate, ADP
Mechanisms of Hormone Action
 Mechanisms of Hormone Action General principles: 1. Signals act over different ranges. 2. Signals have different chemical natures. 3. The same signal can induce a different response in different cells.
Mechanisms of Hormone Action General principles: 1. Signals act over different ranges. 2. Signals have different chemical natures. 3. The same signal can induce a different response in different cells.
Signal-Transduction Pathways
 Signal-Transduction Pathways Pal Bauer 2014/2015 Copyright 2007 by W. H. Freeman and Company No men is an island entire of itself; every man Is a piece of the continent, a part of the main John Donne Introduction
Signal-Transduction Pathways Pal Bauer 2014/2015 Copyright 2007 by W. H. Freeman and Company No men is an island entire of itself; every man Is a piece of the continent, a part of the main John Donne Introduction
Signal Transduction: Information Metabolism. Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire
 Signal Transduction: Information Metabolism Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire Introduction Information Metabolism How cells receive, process and respond
Signal Transduction: Information Metabolism Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire Introduction Information Metabolism How cells receive, process and respond
Regulation of Enzymatic Activity. Lesson 4
 Regulation of Enzymatic Activity Lesson 4 Regulation of Enzymatic Activity no real regulation: - regulation of enzyme expression and turnover - control of enzyme trafficking - supply of cofactors real
Regulation of Enzymatic Activity Lesson 4 Regulation of Enzymatic Activity no real regulation: - regulation of enzyme expression and turnover - control of enzyme trafficking - supply of cofactors real
Receptors and Drug Action. Dr. Subasini Pharmacology Department Ishik University, Erbil
 Receptors and Drug Action Dr. Subasini Pharmacology Department Ishik University, Erbil Receptors and Drug Action Receptor Receptor is defined as a macromolecule or binding site located on the surface or
Receptors and Drug Action Dr. Subasini Pharmacology Department Ishik University, Erbil Receptors and Drug Action Receptor Receptor is defined as a macromolecule or binding site located on the surface or
Biochemie 4. Cell communication - GPCR
 Biochemie 4 Cell communication - GPCR 1 Lecture outline General principles - local and long-distance signaling - classes of receptors - molecular switches and second messengers Receptor tyrosine kinases
Biochemie 4 Cell communication - GPCR 1 Lecture outline General principles - local and long-distance signaling - classes of receptors - molecular switches and second messengers Receptor tyrosine kinases
Lecture 1, Fall 2014 The structure of the eukaryotic cell, as it relates to cells chemical and biological functions.
 Lecture 1, Fall 2014 The structure of the eukaryotic cell, as it relates to cells chemical and biological functions. There are several important themes that transcends just the chemistry and bring the
Lecture 1, Fall 2014 The structure of the eukaryotic cell, as it relates to cells chemical and biological functions. There are several important themes that transcends just the chemistry and bring the
Lecture 7: Signaling Through Lymphocyte Receptors
 Lecture 7: Signaling Through Lymphocyte Receptors Questions to Consider After recognition of its cognate MHC:peptide, how does the T cell receptor activate immune response genes? What are the structural
Lecture 7: Signaling Through Lymphocyte Receptors Questions to Consider After recognition of its cognate MHC:peptide, how does the T cell receptor activate immune response genes? What are the structural
Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction. Marker Sequence (5 3 ) Accession No.
 Supplementary Tables Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction Marker Sequence (5 3 ) Accession No. Angiopoietin 1, ANGPT1 A CCCTCCGGTGAATATTGGCTGG NM_001146.3
Supplementary Tables Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction Marker Sequence (5 3 ) Accession No. Angiopoietin 1, ANGPT1 A CCCTCCGGTGAATATTGGCTGG NM_001146.3
Zaid sarhan. Osama Al-Ghafri ... Dr.nayef karadsheh
 16 Zaid sarhan Osama Al-Ghafri... Dr.nayef karadsheh ALL THE FIGUERS IN THIS SHEET ARE VERY IMPORTANT AND USEFUL, PLEASE DON T SKIP THEM. Glycogen phosphorylase kinase = GPK // glycogen phosphorylase=gp
16 Zaid sarhan Osama Al-Ghafri... Dr.nayef karadsheh ALL THE FIGUERS IN THIS SHEET ARE VERY IMPORTANT AND USEFUL, PLEASE DON T SKIP THEM. Glycogen phosphorylase kinase = GPK // glycogen phosphorylase=gp
Protein kinases a CRASH course. Michael Freissmuth Institute of Pharmacology
 Protein kinases a CRASH course Michael Freissmuth Institute of Pharmacology Protein kinases Outline: Kinome - multitude of kinases (why, what for?) Specificity Regulation Significance for drug development
Protein kinases a CRASH course Michael Freissmuth Institute of Pharmacology Protein kinases Outline: Kinome - multitude of kinases (why, what for?) Specificity Regulation Significance for drug development
Table S7B- Biocarta functional annotation of celecoxib-modulated genes unique to COX-2 expressers
 Table S7B- Biocarta functional annotation of celecoxib-modulated genes unique to COX-2 expressers ListHits ListTotal PopulationHits PopulationTotal Terms 3 72 10 1429 Glycolysis Pathway 3 72 11 1429 Regulation
Table S7B- Biocarta functional annotation of celecoxib-modulated genes unique to COX-2 expressers ListHits ListTotal PopulationHits PopulationTotal Terms 3 72 10 1429 Glycolysis Pathway 3 72 11 1429 Regulation
PATHOGEN INNOCUOUS ANTIGEN. No Danger- very low expression of costimulatory ligands Signal One Only
 Harvard-MIT Division of Health Sciences and Technology HST.176: Cellular and Molecular Immunology Course Director: Dr. Shiv illai AICD Naive Activated Effector Memory Activated Effector Naive AICD Activated
Harvard-MIT Division of Health Sciences and Technology HST.176: Cellular and Molecular Immunology Course Director: Dr. Shiv illai AICD Naive Activated Effector Memory Activated Effector Naive AICD Activated
Cell responses to environment-- Signals
 Cell responses to environment-- Signals Signal transduction can coordinate: Development Formation of tissues Timing of cell division Direction of cell enlargement Size and shape of organs Responses to
Cell responses to environment-- Signals Signal transduction can coordinate: Development Formation of tissues Timing of cell division Direction of cell enlargement Size and shape of organs Responses to
Chapter 9: Biochemical Mechanisms for Information Storage at the Cellular Level. From Mechanisms of Memory, second edition By J. David Sweatt, Ph.D.
 Chapter 9: Biochemical Mechanisms for Information Storage at the Cellular Level From Mechanisms of Memory, second edition By J. David Sweatt, Ph.D. Chapter 9: Dendritic Spine Figure 1 Summary: Three Primary
Chapter 9: Biochemical Mechanisms for Information Storage at the Cellular Level From Mechanisms of Memory, second edition By J. David Sweatt, Ph.D. Chapter 9: Dendritic Spine Figure 1 Summary: Three Primary
CAN THE CD43 MOLECULE BE CONSIDERED AS A THERAPEUTIC TAGET? Yvonne Rosenstein Instituto de Biotecnologia, UNAM
 CAN THE CD43 MOLECULE BE CONSIDERED AS A THERAPEUTIC TAGET? Yvonne Rosenstein Instituto de Biotecnologia, UNAM CD43: the molecule and its signaling capacities favors a Th1 differentiation pattern and contributes
CAN THE CD43 MOLECULE BE CONSIDERED AS A THERAPEUTIC TAGET? Yvonne Rosenstein Instituto de Biotecnologia, UNAM CD43: the molecule and its signaling capacities favors a Th1 differentiation pattern and contributes
Chapter 11. B cell generation, Activation, and Differentiation. Pro-B cells. - B cells mature in the bone marrow.
 Chapter B cell generation, Activation, and Differentiation - B cells mature in the bone marrow. - B cells proceed through a number of distinct maturational stages: ) Pro-B cell ) Pre-B cell ) Immature
Chapter B cell generation, Activation, and Differentiation - B cells mature in the bone marrow. - B cells proceed through a number of distinct maturational stages: ) Pro-B cell ) Pre-B cell ) Immature
Lecture 15. Signal Transduction Pathways - Introduction
 Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Signal transduction and protein kinase inhibitors. Feng Qian ( 钱峰 )
 Signal transduction and protein kinase inhibitors Feng Qian ( 钱峰 ) fengqian@sjtu.edu.cn Protein Kinases in the Human Genome 518 kinases 1.7 % of human genome Lipid kinases Nucleotide kinases Cell Signaling
Signal transduction and protein kinase inhibitors Feng Qian ( 钱峰 ) fengqian@sjtu.edu.cn Protein Kinases in the Human Genome 518 kinases 1.7 % of human genome Lipid kinases Nucleotide kinases Cell Signaling
Chapter 11: Inherited Disorders of Human Memory Mental Retardation Syndromes. From Mechanisms of Memory, second edition By J. David Sweatt, Ph.D.
 Chapter 11: Inherited Disorders of Human Memory Mental Retardation Syndromes From Mechanisms of Memory, second edition By J. David Sweatt, Ph.D. Chapter 11: Mental Retardation Syndromes Table I: Mouse
Chapter 11: Inherited Disorders of Human Memory Mental Retardation Syndromes From Mechanisms of Memory, second edition By J. David Sweatt, Ph.D. Chapter 11: Mental Retardation Syndromes Table I: Mouse
Signal-Transduction Cascades - 2. The Phosphoinositide Cascade
 Signal-Transduction Cascades - 2 The Phosphoinositide Cascade Calcium ion as a second messenger Tyrosine kinase and receptor dimerization scribd.com Faisal Khatib JU The Phosphoinositide Cascade Used by
Signal-Transduction Cascades - 2 The Phosphoinositide Cascade Calcium ion as a second messenger Tyrosine kinase and receptor dimerization scribd.com Faisal Khatib JU The Phosphoinositide Cascade Used by
Chapter 9. Cellular Signaling
 Chapter 9 Cellular Signaling Cellular Messaging Page 215 Cells can signal to each other and interpret the signals they receive from other cells and the environment Signals are most often chemicals The
Chapter 9 Cellular Signaling Cellular Messaging Page 215 Cells can signal to each other and interpret the signals they receive from other cells and the environment Signals are most often chemicals The
Computational Biology I LSM5191
 Computational Biology I LSM5191 Aylwin Ng, D.Phil Lecture 6 Notes: Control Systems in Gene Expression Pulling it all together: coordinated control of transcriptional regulatory molecules Simple Control:
Computational Biology I LSM5191 Aylwin Ng, D.Phil Lecture 6 Notes: Control Systems in Gene Expression Pulling it all together: coordinated control of transcriptional regulatory molecules Simple Control:
MBG301. Class IV. Classification of GPCRs according to their effector function (according to Lodish)
 MBG301 Class IV Classification of GPCRs according to their effector function (according to Lodish) 1. Adenylcyclase activation by GPCRs 2. Ion channel regulation by GPCRs 3. Phospholipase C (PLC) activation
MBG301 Class IV Classification of GPCRs according to their effector function (according to Lodish) 1. Adenylcyclase activation by GPCRs 2. Ion channel regulation by GPCRs 3. Phospholipase C (PLC) activation
Sarah Jaar Marah Al-Darawsheh
 22 Sarah Jaar Marah Al-Darawsheh Faisal Mohammad Receptors can be membrane proteins (for water-soluble hormones/ligands) or intracellular (found in the cytosol or nucleus and bind to DNA, for lipid-soluble
22 Sarah Jaar Marah Al-Darawsheh Faisal Mohammad Receptors can be membrane proteins (for water-soluble hormones/ligands) or intracellular (found in the cytosol or nucleus and bind to DNA, for lipid-soluble
Supplementary Figure 1: Func8onal Network Analysis of Kinases Significantly Modulated by MERS CoV Infec8on and Conserved Across All Time Points
 A. B. 8 4 Supplementary Figure : Func8onal Network Analysis of Kinases Significantly Modulated by MERS CoV Infec8on and Conserved Across All Time Points Examined. A) Venn diagram analysis of kinases significantly
A. B. 8 4 Supplementary Figure : Func8onal Network Analysis of Kinases Significantly Modulated by MERS CoV Infec8on and Conserved Across All Time Points Examined. A) Venn diagram analysis of kinases significantly
Signaling. Dr. Sujata Persad Katz Group Centre for Pharmacy & Health research
 Signaling Dr. Sujata Persad 3-020 Katz Group Centre for Pharmacy & Health research E-mail:sujata.persad@ualberta.ca 1 Growth Factor Receptors and Other Signaling Pathways What we will cover today: How
Signaling Dr. Sujata Persad 3-020 Katz Group Centre for Pharmacy & Health research E-mail:sujata.persad@ualberta.ca 1 Growth Factor Receptors and Other Signaling Pathways What we will cover today: How
A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy se
 A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy serves as a defence mechanism that prevents or retards
A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy serves as a defence mechanism that prevents or retards
Chapter 15: Signal transduction
 Chapter 15: Signal transduction Know the terminology: Enzyme-linked receptor IP3+DAG G-protein linked receptor camp nuclear hormone receptor Ca 2+ G-protein adaptor protein protein kinase scaffolding protein
Chapter 15: Signal transduction Know the terminology: Enzyme-linked receptor IP3+DAG G-protein linked receptor camp nuclear hormone receptor Ca 2+ G-protein adaptor protein protein kinase scaffolding protein
Enzymes Part III: regulation II. Dr. Mamoun Ahram Summer, 2017
 Enzymes Part III: regulation II Dr. Mamoun Ahram Summer, 2017 Advantage This is a major mechanism for rapid and transient regulation of enzyme activity. A most common mechanism is enzyme phosphorylation
Enzymes Part III: regulation II Dr. Mamoun Ahram Summer, 2017 Advantage This is a major mechanism for rapid and transient regulation of enzyme activity. A most common mechanism is enzyme phosphorylation
Chapter 11. B cell generation, Activation, and Differentiation. Pro-B cells. - B cells mature in the bone marrow.
 Chapter B cell generation, Activation, and Differentiation - B cells mature in the bone marrow. - B cells proceed through a number of distinct maturational stages: ) Pro-B cell ) Pre-B cell ) Immature
Chapter B cell generation, Activation, and Differentiation - B cells mature in the bone marrow. - B cells proceed through a number of distinct maturational stages: ) Pro-B cell ) Pre-B cell ) Immature
Cell Communication. Cell Communication. Cell Communication. Cell Communication. Cell Communication. Chapter 9. Communication between cells requires:
 Chapter 9 Communication between cells requires: ligand: the signaling molecule receptor protein: the molecule to which the receptor binds -may be on the plasma membrane or within the cell 2 There are four
Chapter 9 Communication between cells requires: ligand: the signaling molecule receptor protein: the molecule to which the receptor binds -may be on the plasma membrane or within the cell 2 There are four
Cellular Physiology (PHSI3009) Contents:
 Cellular Physiology (PHSI3009) Contents: Cell membranes and communication 2 nd messenger systems G-coupled protein signalling Calcium signalling Small G-protein signalling o RAS o MAPK o PI3K RHO GTPases
Cellular Physiology (PHSI3009) Contents: Cell membranes and communication 2 nd messenger systems G-coupled protein signalling Calcium signalling Small G-protein signalling o RAS o MAPK o PI3K RHO GTPases
Designer Affinity Reagents. Brian Kay
 Designer Affinity Reagents Brian Kay bkay@uic.edu Types of Affinity Reagents Src SH3 domain Lysozyme Src SH3 domain FN3 monobody Peptide Ligand Antibody Fragment Scaffold M13 Bacteriophage 900 nm x 10
Designer Affinity Reagents Brian Kay bkay@uic.edu Types of Affinity Reagents Src SH3 domain Lysozyme Src SH3 domain FN3 monobody Peptide Ligand Antibody Fragment Scaffold M13 Bacteriophage 900 nm x 10
Cell Communication. Chapter 11. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 11 Cell Communication PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp
Chapter 11 Cell Communication PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp
I. Top scoring maps. 1.IFN gamma signaling
 I. Top scoring maps 1.IFN gamma signaling Interferon-gamma signaling Interferons (IFNs) are pleiotropic cytokines that mediate anti-viral responses, inhibit proliferation and participate in immune surveillance
I. Top scoring maps 1.IFN gamma signaling Interferon-gamma signaling Interferons (IFNs) are pleiotropic cytokines that mediate anti-viral responses, inhibit proliferation and participate in immune surveillance
Biol403 MAP kinase signalling
 Biol403 MAP kinase signalling The mitogen activated protein kinase (MAPK) pathway is a signalling cascade activated by a diverse range of effectors. The cascade regulates many cellular activities including
Biol403 MAP kinase signalling The mitogen activated protein kinase (MAPK) pathway is a signalling cascade activated by a diverse range of effectors. The cascade regulates many cellular activities including
Lipids and Membranes
 Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Membrane transport D. Endocytosis and Exocytosis
Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Membrane transport D. Endocytosis and Exocytosis
Materials and Methods , The two-hybrid principle.
 The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
G-Protein Coupled Receptors GPCRs. GPCRs
 2 type of ligands 1 G-Protein Coupled Receptors GPCRs One of the largest protein families: > 1000 type of GPCRs in mammals >3% of the human genes Major drug targets: ~ 60 % of approved drugs interact with
2 type of ligands 1 G-Protein Coupled Receptors GPCRs One of the largest protein families: > 1000 type of GPCRs in mammals >3% of the human genes Major drug targets: ~ 60 % of approved drugs interact with
Cell Communication. Local and Long Distance Signaling
 Cell Communication Cell to cell communication is essential for multicellular organisms Some universal mechanisms of cellular regulation providing more evidence for the evolutionary relatedness of all life
Cell Communication Cell to cell communication is essential for multicellular organisms Some universal mechanisms of cellular regulation providing more evidence for the evolutionary relatedness of all life
1. Activated receptor tyrosine kinases (RTKs) phosphorylates themselves
 Enzyme-coupled receptors Transmembrane proteins Ligand-binding domain on the outer surface Cytoplasmic domain acts as an enzyme itself or forms a complex with enzyme 1. Activated receptor tyrosine kinases
Enzyme-coupled receptors Transmembrane proteins Ligand-binding domain on the outer surface Cytoplasmic domain acts as an enzyme itself or forms a complex with enzyme 1. Activated receptor tyrosine kinases
Designer Affinity Reagents. Reagents. Types of Affinity. M13 Bacteriophage. 900 nm x 10 nm. Brian Kay Src SH3 domain.
 Designer Affinity Reagents Brian Kay bkay@uic.edu Types of Affinity Reagents Src SH3 domain Lysozyme Src SH3 domain FN3 monobody Peptide Ligand Antibody Fragment Scaffold M13 Bacteriophage 900 nm 10 nm
Designer Affinity Reagents Brian Kay bkay@uic.edu Types of Affinity Reagents Src SH3 domain Lysozyme Src SH3 domain FN3 monobody Peptide Ligand Antibody Fragment Scaffold M13 Bacteriophage 900 nm 10 nm
PCB 3023 Exam 4 - Form A First and Last Name
 PCB 3023 Exam 4 - Form A First and Last Name Student ID # (U Number) A Before beginning this exam, please complete the following instructions: 1) Write your name and U number on the first page of this
PCB 3023 Exam 4 - Form A First and Last Name Student ID # (U Number) A Before beginning this exam, please complete the following instructions: 1) Write your name and U number on the first page of this
INTRACELLULAR SIGNALLING
 INTRACELLULAR SIGNALLING R. Benacka,, MD, PhD Department of Pathophysiology Medical faculty, Safarik University 1. Long distance chemo A. Receptors without enzymatic activity c-amp IP3- dependent c-gmp/no
INTRACELLULAR SIGNALLING R. Benacka,, MD, PhD Department of Pathophysiology Medical faculty, Safarik University 1. Long distance chemo A. Receptors without enzymatic activity c-amp IP3- dependent c-gmp/no
Lecture 9: Cell Communication I
 02.05.10 Lecture 9: Cell Communication I Multicellular organisms need to coordinate cellular functions in different tissues Cell-to-cell communication is also used by single celled organisms to signal
02.05.10 Lecture 9: Cell Communication I Multicellular organisms need to coordinate cellular functions in different tissues Cell-to-cell communication is also used by single celled organisms to signal
PHSI3009 Frontiers in Cellular Physiology 2017
 Overview of PHSI3009 L2 Cell membrane and Principles of cell communication L3 Signalling via G protein-coupled receptor L4 Calcium Signalling L5 Signalling via Growth Factors L6 Signalling via small G-protein
Overview of PHSI3009 L2 Cell membrane and Principles of cell communication L3 Signalling via G protein-coupled receptor L4 Calcium Signalling L5 Signalling via Growth Factors L6 Signalling via small G-protein
Effects of Second Messengers
 Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Cell signaling. How do cells receive and respond to signals from their surroundings?
 Cell signaling How do cells receive and respond to signals from their surroundings? Prokaryotes and unicellular eukaryotes are largely independent and autonomous. In multicellular organisms there is a
Cell signaling How do cells receive and respond to signals from their surroundings? Prokaryotes and unicellular eukaryotes are largely independent and autonomous. In multicellular organisms there is a
Phospho-AKT Sampler Kit
 Phospho-AKT Sampler Kit E 0 5 1 0 0 3 Kits Includes Cat. Quantity Application Reactivity Source Akt (Ab-473) Antibody E021054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit Akt (Phospho-Ser473) Antibody
Phospho-AKT Sampler Kit E 0 5 1 0 0 3 Kits Includes Cat. Quantity Application Reactivity Source Akt (Ab-473) Antibody E021054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit Akt (Phospho-Ser473) Antibody
BCOR 011 Lecture 19 Oct 12, 2005 I. Cell Communication Signal Transduction Chapter 11
 BCOR 011 Lecture 19 Oct 12, 2005 I. Cell Communication Signal Transduction Chapter 11 External signal is received and converted to another form to elicit a response 1 Lecture Outline 1. Types of intercellular
BCOR 011 Lecture 19 Oct 12, 2005 I. Cell Communication Signal Transduction Chapter 11 External signal is received and converted to another form to elicit a response 1 Lecture Outline 1. Types of intercellular
Supplementary information S1 (table): Specifications of individual protein protein interactions within the Cbl and Cbl-b interactomes
 Supplementary information S1 (table): Specifications of individual protein protein interactions within the Cbl and Cbl-b interactomes Cbl/Cbl-b binding protein Cbl domain/residue Binding protein domain/residue
Supplementary information S1 (table): Specifications of individual protein protein interactions within the Cbl and Cbl-b interactomes Cbl/Cbl-b binding protein Cbl domain/residue Binding protein domain/residue
Chapter 20. Cell - Cell Signaling: Hormones and Receptors. Three general types of extracellular signaling. endocrine signaling. paracrine signaling
 Chapter 20 Cell - Cell Signaling: Hormones and Receptors Three general types of extracellular signaling endocrine signaling paracrine signaling autocrine signaling Endocrine Signaling - signaling molecules
Chapter 20 Cell - Cell Signaling: Hormones and Receptors Three general types of extracellular signaling endocrine signaling paracrine signaling autocrine signaling Endocrine Signaling - signaling molecules
Transduction of Receptor Signals by β-arrestins
 Science Supporting Online Material Transduction of Receptor Signals by β-arrestins Robert J. Lefkowitz and Sudha K. Shenoy Contents Figs. S1 Tables S1 to S3 Table S1. G protein-coupled receptor kinase
Science Supporting Online Material Transduction of Receptor Signals by β-arrestins Robert J. Lefkowitz and Sudha K. Shenoy Contents Figs. S1 Tables S1 to S3 Table S1. G protein-coupled receptor kinase
Growth and Differentiation Phosphorylation Sampler Kit
 Growth and Differentiation Phosphorylation Sampler Kit E 0 5 1 0 1 4 Kits Includes Cat. Quantity Application Reactivity Source Akt (Phospho-Ser473) E011054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit
Growth and Differentiation Phosphorylation Sampler Kit E 0 5 1 0 1 4 Kits Includes Cat. Quantity Application Reactivity Source Akt (Phospho-Ser473) E011054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit
Generation of the Immune Response
 Generation of the Immune Response Sheet 18 immunity I only added extra notes that were explained in the lecture, refer back to the slides. SLIDE 3: In the generation of Immune response whether by B or
Generation of the Immune Response Sheet 18 immunity I only added extra notes that were explained in the lecture, refer back to the slides. SLIDE 3: In the generation of Immune response whether by B or
Asma Karameh BAHAA NAJJAR. Ebaa' Alzayadneh
 26 Asma Karameh BAHAA NAJJAR Ebaa' Alzayadneh Generally speaking, all cells have been programmed during development to response to specific set of extracellular signals produced by other cells.these signals
26 Asma Karameh BAHAA NAJJAR Ebaa' Alzayadneh Generally speaking, all cells have been programmed during development to response to specific set of extracellular signals produced by other cells.these signals
Content Prioritization And Content Entry and Quality Control Process
 Content Prioritization And Content Entry and Quality Control Process The process of data capture begins with the definition of the content module or sub-module to be built (see figure 1). Broadly we define
Content Prioritization And Content Entry and Quality Control Process The process of data capture begins with the definition of the content module or sub-module to be built (see figure 1). Broadly we define
Signal Transduction SS Gerhild van Echten-Deckert
 Signal Transduction SS 2018 Gerhild van Echten-Deckert Tel. 73 2703 E-mail: g.echten.deckert@uni-bonn.de https://www.limes-institut-bonn.de/forschung/ Focus on 2 classes of cell-surface receptors (Growth
Signal Transduction SS 2018 Gerhild van Echten-Deckert Tel. 73 2703 E-mail: g.echten.deckert@uni-bonn.de https://www.limes-institut-bonn.de/forschung/ Focus on 2 classes of cell-surface receptors (Growth
Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION
 Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION 1. Feedback a. Negative feedback mechanisms maintain dynamic homeostasis for a particular condition (variable) by regulating physiological processes,
Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION 1. Feedback a. Negative feedback mechanisms maintain dynamic homeostasis for a particular condition (variable) by regulating physiological processes,
Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene
 Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene YUELIN ZHANG, WEIHUA FAN, MARK KINKEMA, XIN LI, AND
Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene YUELIN ZHANG, WEIHUA FAN, MARK KINKEMA, XIN LI, AND
Cell Signaling and Signal Transduction: Communication between Cells
 Cell Signaling and Signal Transduction: Communication between Cells - Cell signaling makes it possible for cells to respond in an appropriate manner to a specific environmental stimulus THE BASIC ELEMENTS
Cell Signaling and Signal Transduction: Communication between Cells - Cell signaling makes it possible for cells to respond in an appropriate manner to a specific environmental stimulus THE BASIC ELEMENTS
Hormones and Signal Transduction. Dr. Kevin Ahern
 Dr. Kevin Ahern Signaling Outline Signaling Outline Background Signaling Outline Background Membranes Signaling Outline Background Membranes Hormones & Receptors Signaling Outline Background Membranes
Dr. Kevin Ahern Signaling Outline Signaling Outline Background Signaling Outline Background Membranes Signaling Outline Background Membranes Hormones & Receptors Signaling Outline Background Membranes
About OMICS Group Conferences
 About OMICS Group OMICS Group International is an amalgamation of Open Access publications and worldwide international science conferences and events. Established in the year 2007 with the sole aim of
About OMICS Group OMICS Group International is an amalgamation of Open Access publications and worldwide international science conferences and events. Established in the year 2007 with the sole aim of
Cell Communication. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 11 Cell Communication PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp
Chapter 11 Cell Communication PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp
Cell Communication. Chapter 11. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 11 Cell Communication PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp
Chapter 11 Cell Communication PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp
Propagation of the Signal
 OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
Signal transduction by immunoglobulin Fc receptors
 Signal transduction by immunoglobulin Fc receptors Gabriela Sánchez-Mejorada and Carlos Rosales Immunology Department, Instituto de Investigaciones Biomédicas, Universidad Nacional Autónoma de México,
Signal transduction by immunoglobulin Fc receptors Gabriela Sánchez-Mejorada and Carlos Rosales Immunology Department, Instituto de Investigaciones Biomédicas, Universidad Nacional Autónoma de México,
Cell Communication. Chapter 11. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 11 Cell Communication PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp
Chapter 11 Cell Communication PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp
Cell Communication. Chapter 11. PowerPoint Lectures for Biology, Seventh Edition. Lectures by Chris Romero. Neil Campbell and Jane Reece
 Chapter 11 Cell Communication PowerPoint Lectures for Biology, Seventh Edition Neil Campbell and Jane Reece Lectures by Chris Romero Overview: The Cellular Internet Cell-to-cell communication Is absolutely
Chapter 11 Cell Communication PowerPoint Lectures for Biology, Seventh Edition Neil Campbell and Jane Reece Lectures by Chris Romero Overview: The Cellular Internet Cell-to-cell communication Is absolutely
