The Saccharomyces cerevisiae YBR159w Gene Encodes the 3-Ketoreductase of the Microsomal Fatty Acid Elongase*
|
|
|
- Darrell Stanley
- 6 years ago
- Views:
Transcription
1 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 277, No. 38, Issue of September 20, pp , 2002 Printed in U.S.A. The Saccharomyces cerevisiae YBR159w Gene Encodes the 3-Ketoreductase of the Microsomal Fatty Acid Elongase* Received for publication, June 6, 2002, and in revised form, June 25, 2002 Published, JBC Papers in Press, June 26, 2002, DOI /jbc.M Gongshe Han, Ken Gable, Sepp D. Kohlwein, Frédéric Beaudoin, Johnathan A. Napier, and Teresa M. Dunn From the Department of Biochemistry and Molecular Biology, Uniformed Services University of the Health Sciences, Bethesda, Maryland 20184, Institute of Arable Crops-Long Ashton Research Station, Long Ashton, Bristol BS41 9AF, United Kingdom, and the Department of Molecular Biology, Biochemistry, and Microbiology, SFB Biomembrane Research Center, University Graz, 1 Schubertstrasse, A8010 Graz, Austria The YBR159w gene encodes the major 3-ketoreductase activity of the elongase system of enzymes required for very long-chain fatty acid (VLCFA) synthesis. Mutants lacking the YBR159w gene display many of the phenotypes that have previously been described for mutants with defects in fatty acid elongation. These phenotypes include reduced VLCFA synthesis, accumulation of high levels of dihydrosphingosine and phytosphingosine, and accumulation of medium-chain ceramides. In vitro elongation assays confirm that the ybr159 mutant is deficient in the reduction of the 3-ketoacyl intermediates of fatty acid elongation. The ybr159 mutant also displays reduced dehydration of the 3-OH acyl intermediates of fatty acid elongation, suggesting that Ybr159p is required for the stability or function of the dehydratase activity of the elongase system. Green fluorescent protein-tagged Ybr159p co-localizes and co-immunoprecipitates with other elongating enzymes, Elo3p and Tsc13p. Whereas VLCFA synthesis is essential for viability, the ybr159 mutant cells are viable (albeit very slowly growing) and do synthesize some VLCFA. This suggested that a functional ortholog of Ybr159p exists that is responsible for the residual 3-ketoreductase activity. By disrupting the orthologs of Ybr159w in the ybr159 mutant we found that the ybr159 ayr1 double mutant was inviable, suggesting that Ayr1p is responsible for the residual 3-ketoreductase activity. The distinct fatty acid compositions of different cellular membranes highlight the importance of lipid structure and composition for membrane function. The fatty acids are not only structural components of membranes; they are also bioactive metabolites that regulate various cellular processes. Cytosolic fatty acid synthase catalyzes the de novo synthesis of the majority of cellular fatty acids, which have 16 or 18 carbons. In contrast, the membrane-associated fatty acid chain-elongating systems synthesize the very long-chain fatty * This work was supported by National Science Foundation Grant G171FL and Uniformed Services University of the Health Sciences Grant C071FT (to T. M. D.) and Austrian Science Fund Grant F706 (to S. D. K.). IACR-Long Ashton Research Station receives grant-aided support from the Biotechnology and Biological Sciences Research Council of the United Kingdom. The costs of publication of this article were defrayed in part by the payment of page charges. This article must therefore be hereby marked advertisement in accordance with 18 U.S.C. Section 1734 solely to indicate this fact. To whom correspondence should be addressed: Dept. of Biochemistry and Molecular Biology, Uniformed Services University of the Health Sciences, 4301 Jones Bridge Rd., Bethesda, MD Tel.: ; Fax: ; tdunn@usuhs.mil acids (VLCFAs 1 ; 18 carbons). The VLCFAs confer unique physical properties upon the lipids into which they are incorporated. Furthermore, different tissues contain structurally distinct VLCFAs, suggesting that they confer tissue-specific functions. For example, the VLCFAs in brain are predominantly saturated or monounsaturated with chain lengths of 24 or 26 carbons; those in the stomach, kidney, and brain are -hydroxylated; and those in testes, epidermis, and retina are polyunsaturated with chain lengths of up to 34 carbons. Fatty acid elongation, beyond C16/C18 carbon atoms synthesized by fatty acid synthase in conjunction with Elo1p, is catalyzed by the microsomal elongase that consists of four distinct enzymatic reactions. In sequential order, these are a condensation reaction between a CoA-esterified fatty acyl substrate and malonyl-coa, a 3-ketoacyl-CoA reduction, a 3-hydroxyacyl-CoA dehydration, and a final enoyl-coa reduction (Fig. 1). Early biochemical studies suggested that the enzymatic activities reside in distinct proteins but that they may associate into a complex that catalyzes fatty acid elongation. In fact, the evidence suggests that there are multiple complexes that elongate different FA substrates. For example, inhibition studies indicated that different proteins are responsible for the elongation of unsaturated and saturated fatty acids and for the elongation of fatty acids of different chain lengths (1). The genetic studies in yeast support these findings, since Elo2p and Elo3p are homologous proteins believed to catalyze the same reaction but with preferences for substrates of different chain lengths. It has been hypothesized that the specificity of each elongation reaction is conferred through the selectivity of the first and rate-limiting step, the condensation reaction. Investigators working with Arabidopsis found that the FAE1 gene is required for VLCFA synthesis (2 6). The FAE1 gene was predicted to encode a condensing enzyme (3-ketoacyl-CoA synthase) based on sequence homology with other condensing enzymes (2). Biochemical characterization of the FAE1 gene from both Arabidopsis and jojoba (7) supports the conclusion that Fae1p is involved in VLCFA synthesis. Surprisingly, there is no gene in Saccharomyces cerevisiae with significant homology to FAE1. On the other hand, the ELO homologs comprise a gene family conserved from yeast to humans that are candidates for a novel class of condensing enzymes. Two lines of experiments identified the S. cerevisiae ELO2 and ELO3 genes as being required for VLCFA synthesis. Based on their homol- 1 The abbreviations used are: VLCFAs, very long-chain fatty acids; LCB, long-chain base; ER, endoplasmic reticulum; FAMES, fatty acid methyl esters; GCMS, gas chromatography mass spectrometry; SPT, serine palmitoyltransferase; IPC, inositolphosphoceramide; PHS, phytosphingosine; HA, hemagglutinin; GFP, green fluorescent protein. This paper is available on line at
2 Ybr159p Is the Major 3-Ketoreductase of Fatty Acid Elongation ogy to ELO1, a gene encoding a protein required to convert myristate to palmitate (8, 9), the ELO2 and ELO3 genes were determined to be required for VLCFA synthesis (10). We identified a role for the ELO2 and ELO3 genes in VLCFA synthesis through the analysis of sphingolipid synthesis mutants (discussed below). Elo2p is involved in elongation of fatty acids up to C22 or C24, whereas Elo3p has a broader substrate specificity and is essential for conversion of C24 to C26 (10). The hypothesis that the specificity of elongation is conferred by the condensing enzymes is further supported by the observation that heterologous expression in yeast of the plant Fae1p condensing enzyme confers the ability to elongate C18:1 to C20:1 and C22:1, although yeast does not normally elongate monounsaturated fatty acids longer than C16 or C18 fatty acids (11). Similarly, heterologous expression of the Caenorhabditis elegans ELO-like PEA1 gene in yeast confers the ability to elongate exogenously supplied polyunsaturated C18 fatty acids, although yeast has only very limited endogenous capacity for synthesizing or elongating polyunsaturated fatty acids (12, 13). We have previously shown that mutants defective in the only yeast fatty acid desaturase, Ole1p, survive if supplemented with unsaturated C14:1 9 fatty acids that become elongated to C16:1 11 by the Elo1p gene product (14). In contrast to the Elo enzymes, which are believed to display substrate specificity, it may be that the other elongating enzymes (two reductases and a dehydratase; see Fig. 1) are common components of the microsomal fatty acyl elongases that function to lengthen all of the fatty acid elongase substrates. We recently identified the TSC13 gene in a screen for temperature-sensitive (ts) mutants with defects in sphingolipid synthesis (15 17) and provided evidence that the TSC13 gene encodes the enoyl reductase component of the microsomal elongase. Tsc13p was shown to coimmunoprecipitate with the presumptive condensing enzymes, Elo2p and Elo3p. Interestingly, a growth state-dependent enrichment of Tsc13p to sites of vacuole-nuclear envelope interaction was observed, but the significance of this is unclear, and Elo2p and Elo3p do not display this localization. TSC13 is essential, as is expected for a gene encoding a nonredundant fatty acid-elongating activity. In recent studies aimed at identifying other enzymatic components of the fatty acid elongase, we found that the YBR159w gene is required for reconstituting heterologous (Fae1p- or Pea1p-mediated) elongase activity in yeast. Here we report analysis of the mutants lacking Ybr159p and demonstrate that these mutant cells have many of the phenotypes that we have previously described for the elo2, elo3, and tsc13-1 (elongasedefective) mutant cells. These phenotypes indicate that Ybr159p is the major 3-ketoreductase activity required for endogenous VLCFA synthesis in yeast. We report that Ybr159p is a nuclear/endoplasmic reticulum (ER)-localized protein that associates in complexes with Elo3p and Tsc13p. Mutants that are devoid of elongase activity (the elo2 elo3 double mutant or the tsc13 mutant) are inviable, indicating that the VLCFAs are essential. Although it grows poorly, the ybr159 mutant is viable, indicating that there is another gene that encodes a 3-ketoreductase activity that can function in fatty acid elongation. We provide evidence that the AYR1 gene, previously reported to encode 1-acyldihydroxyacetone-phosphate reductase activity (18), is responsible for the residual fatty acid elongation activity. FIG. 1.Pathway of fatty acid elongation. Each cycle of elongation requires four successive reactions and lengthens the growing fatty acid by two carbon units: condensation of malonyl-coa with the acyl-coa substrate, reduction of the 3-ketoacyl-CoA, dehydration of the 3-hydroxy acyl-coa, and reduction of the trans-2,3-acyl-coa. Although the intermediates and the product of the elongation cycle are shown as CoA derivatives, this has not yet been experimentally confirmed. EXPERIMENTAL PROCEDURES Media, Strains, and Genetic Manipulations Yeast media were prepared, and cells were grown according to standard procedures (19). The majority of the yeast strains used in this study are listed in Table I. A set of deletion mutants disrupted for putative oxidoreductases (Table II) was purchased from Research Genetics. To construct the ybr159::trp1- and ybr159::ura3-disrupting alleles, a SalI/XbaI-ended PCR fragment extending from 200 bp upstream of the start codon of YBR159w to 200 bp past the stop codon was generated using primer 159F (SalI-ended 5 -ACAGCTGTCGACTATA- GAGATAATTGTAGG-3 ) and 159R (XbaI-ended 5 -AGAGCTCTAGAT- GATATTCTGGAAAAGCA-3 ). The fragment was ligated between the XhoI and XbaI sites of a Bluescript plasmid. The resulting plasmid was digested with XhoI to remove a 779-bp fragment extending from codon 93 past the end of the YBR159w coding sequence. A SalI-ended TRP1 (or URA3) fragment, generated by PCR, was ligated into the XhoI site at the deletion junction to generate the disrupting allele. The disrupting allele was liberated from the plasmid by digestion with KpnI and NotI. The ybr159::trp1 elo3::ura3 double mutant was generated by transforming the KpnI/NotI-ended ybr159w::trp1-disrupting fragment into strain TDY 2054 (elo3::ura3) (16) and selecting tryptophan prototrophs. The replacement of the YBR159w allele by the ybr159::trp1 allele was verified by PCR analysis. The ybr159w::trp1 tsc13-1 double mutant was generated by crossing a ybr159w::trp1 haploid mutant (TDY 1590, generated by disrupting YBR159w in TDY 2039) with TDY 2051a (tsc13-1) (16). The resulting diploid was sporulated, and following tetrad dissection, the products of meiosis that were the ybr159w::trp1 tsc13-1 double mutants were identified as tryptophan prototrophs that were unable to grow at 37 C. The ypc1::kanmx-disrupting allele was generated by PCR using
3 35442 Ybr159p Is the Major 3-Ketoreductase of Fatty Acid Elongation TABLE I Strain list Strain name TDY 2037 TDY 2053 TDY 2054 TDY 2057 TDY 2058 TDY 1590 TDY 2051a TDY 1591 TDY 1593 TDY 2302 TDY 1592 TDY 5999 TDY 6000 TDY 6001 TDY 6002 TDY 6003 TDY 6004 TDY 6005 TDY 6006 TDY 6007 TDY 6008 Genotype Mat ura3 52 leu2 trp1 lys2 Mata his4 ura3 trp1 leu2 elo2 URA3 Mata elo3 ura3 his4 619 ura3 52 trp1 leu2 Mat ade2 ura3 trp1 leu2 lys2 elo2 TRP1 tsc13 1 Mat ade2 ura3 trp1 leu2 lys2 elo3 TRP1 tsc13 1 Mat ybr159 TRP1 ura3 52 trp1 leu2 lys2 Mata tsc13 1 ura3 52 trp1 leu2 Mat ybr159 TRP1 tsc13 1 ura3 52 trp1 leu2 lys2 Mat ybr159 TRP1 elo3 URA3 ura3 52 trp1 leu2 lys2 Mat ura3 52 trp1 lys2 csg2 URA3 Mata ura3 52 trp1 leu2 lys2 his4 ybr159 TRP1 csg2 LEU2 Mata ura3 52 leu2 trp1 ade2 elo2 TRP1 Mat ura3 52 leu2 trp1 lys2 ypc1 kanmx ydc1 URA3 Mat ura3 52 leu2 trp1 lys2 Mat ura3 52 leu2 trp1 lys2 ade2 ypc1 kanmx Mat ura3 52 leu2 trp1 lys2 ydc1 URA3 Mat ura3 52 leu2 trp1 ade2 ypc1 kanmx ydc1 URA3 Mata ura3 52 leu2 trp1 lys2 elo2 TRP1 Mata ura3 52 leu2 trp1 ade2 elo2 TRP1 ypc1 kanmx Mata ura3 52 leu2 trp1 elo2 TRP1 ydc1 URA3 Mata ura3 52 leu2 trp1 ade2 elo2 TRP ypc1 kanmx ydc1 URA3 TABLE II Candidate genes responsible for residual 3-ketoreductase activity in the ybr159 mutant a Ref. 18. b Ref. 34. Gene Homology to Ybr159p YMR226c Score 171 Identities 69/212 (32%), Positives 102/212 (48%) YIL124w/AYR1 a Score 118 Identities 44/183 (24%), Positives 85/183 (46%) YIR036c Score 108 Identities 51/212 (24%), Positives 92/212 (43%) YKL055c/OAR1 b Score 70 Identities 25/106 (23%), Positives 53/106 (50%) YDL114w Score 91 Identities 34/125 (27%), Positives 60/125 (48%) genomic DNA prepared from the Research Genetics strain harboring the disruption and the EcoRI-ended YPC1F (5 -CCCGGGGATCCGC- CGATTAGATCCGGCCC) and XbaI-ended YPC1R (5 -CCGGGCTC- GAGCATGTCCCGAATTAGCTA) primers. Construction of the ydc1::ura3-disrupting allele (kindly provided by the Obeid laboratory) was described earlier (20). The YPC1 and YDC1 genes were sequentially disrupted in TDY 2037 to generate TDY 6000, which was crossed to TDY 2053 (elo2 ). This diploid was sporulated, and the tetrads were dissected to generate the set of haploid strains (TDY ) having the various combinations of the ypc1::kanmx, ydc1::ura3, and elo2 mutations (Table I). Construction of Ybr159p-GFP by Chromosomal Fusion Chromosomal C-terminal GFP fusion to Ybr159p was performed by the short flanking homology method (21). In brief, coding sequences for GFP and the kanmx6 selection marker were amplified from template plasmid pfa6a-gfps65t-kanmx6 (provided by J. Hegemann, Heinrich-Heine- Universität, Düsseldorf, Germany) using hybrid primers homologous to 30 or 26 bp 5 or 3 of the template plasmid, respectively, and 50 nucleotides upstream or downstream of the chromosomal integration sites, flanking the stop codon. The following primers were used (letters in boldface type mark the vector sequences): YBR159 upstream primer, 5 -CTATCAGAATTAGAGCCTTAAAAAAAGCCGCAAGACAGGTTAA- AAAGGAAGGAGCAGGTGCTGGTGCTGGTGCTGGAGCA-3 ; YBR159 downstream primer, 5 -ATATATATATATATATGTATTGTAT- AAAAACTATCTCGAGAACGATAATTATCGATGAATTCGAGCTC- GTTTAAAC-3. After transformation of the linear fragments into FY1679 (MATa/ ura3 52/ura3 52 trp1 /TRP1 leu2 /LEU2 his3 / HIS3 GAL2/GAL2) and selection of Geneticin-resistant colonies, cells were sporulated, and haploid strains containing the GFP-tagged alleles were selected. Growth tests in the presence of inhibitors of fatty acid synthesis/elongation (cerulenin, soraphen A (22), and calcium) and at elevated or reduced temperatures did not unveil any changes compared with wild type, demonstrating that the GFP-tagged fusion allele of YBR159w was fully functional. The pyes-at-159 Plasmid An Arabidopsis thaliana expressed sequence tag (GenBank TM accession number AA10E08; kindly provided by Prof. H. J. Bohnert, University of Arizona) derived from gene F12A21.31 was used as template DNA for PCR amplification using primers YAt159.for (5 -GCGGGTACCACCATGGAGATCTGCACTTAC- 3 ) and YAt159.rev (5 -GCGGAGCTCTCATTCTTTCTTCATGGA-3 ). The amplified sequence was then restricted using KpnI and SacI (underlined in the forward and reverse primers, respectively), purified using the Qiagen PCR purification kit, and ligated into the KpnI and SacI sites of the pyes2 plasmid (Invitrogen). Ceramide, Long-chain Base, and Fatty Acid Analyses Ceramides were extracted and analyzed by thin layer chromatography (TLC) as previously described (23). Ceramides were purified by preparative silica gel TLC and subjected to acid methanolysis, and the fatty acid methyl esters (FAMEs) and long-chain bases (LCBs) were recovered as described earlier (23). LCBs were extracted, separated by TLC, and visualized using ninhydrin as described (15). Fatty acids were extracted, and the FAMEs were prepared as previously described (16). Gas chromatography/mass spectrometry (GCMS) was performed using an HP 6890 Series GC system with a Supelcowax 10 column, coupled to an HP 5973 Mass Selective Detector. Elongase Assays Microsomes were prepared from the wild-type or mutant cells as has been previously described (24). Total elongase activity was measured in a volume of 200 l containing 50 mm Tris, ph 7.5, 1 mm MgCl 2, 150 M Triton X-100, 1 mm NADPH, 1 mm NADH, 10 mm -mercaptoethanol, 40 M palmitoyl-coa, and 60 M [2-14 C]malonyl-CoA (0.05 Ci/ml) at 37 C. The reaction was initiated by the addition of mg of microsomal protein. Protein concentrations were determined using the Bio-Rad protein assay reagent. For assays of only the condensing activity, the NADPH and NADH were omitted. At the indicated times, the reaction was terminated by adding 200 l of5 M KOH, 10% MeOH and heating at 80 C for 1 h. Following the addition of 200 l of 10 N H 2 SO 4, fatty acids were recovered by two 1.5-ml extractions into hexane. The extracted fatty acids were resolved by silica gel TLC using hexane/diethyl ether/acetic acid (30:70:1) as the developing solvent. The radiolabeled fatty acids were detected and
4 Ybr159p Is the Major 3-Ketoreductase of Fatty Acid Elongation quantified using a PhosphorImager SI (Amersham Biosciences). Serine Palmitoyltransferase (SPT) Assays and Immunoblotting of Lcb1p and Lcb2p The SPT assays were preformed as previously described (24), except that 0.4 mg of microsomal protein and 75 M palmitoyl-coa were used. Each assay was conducted in quadruplicate, and the average SPT activity is reported. The methods used for immunoblotting and for detection of Lcb1p and Lcb2p with the anti-lcb1p and anti-lcb2p antibodies were also previously described (24). Immunoprecipitation Microsomes were prepared from strains containing Ybr159p-GFP and Tsc13p-MYC or containing Ybr159p-GFP, Tsc13p-MYC, and Elo3p-HA. The microsomes were solubilized at 1 mg/ml with 2 mm sucrose monolaurate (Roche Molecular Biochemicals) for 10 min, and the high speed ( g, 30 min) supernatant was collected. The supernatant (150 l) was incubated with 25 l ofthe precipitating antibody (0.5 mg/ml) coupled to Sepharose (from Babco, Berkeley, CA) for 2 h. The precipitates were washed three times with 600 l of50mm HEPES, ph 7.5, and resuspended in 150 l ofsds loading buffer, and a 10- l sample was subjected to 8% SDS-PAGE. Following transfer of the separated proteins to nitrocellulose, the blots were blocked in 0.1 M Tris, ph 7.5, 0.15 M NaCl, 0.1% Tween 20, 5% dry milk. The Tsc13p-Myc was detected with horseradish peroxidase-conjugated monoclonal anti-myc antibodies (from Invitrogen) at 1:5000. Elo3p-HA was detected using horseradish peroxidase-conjugated monoclonal anti-ha antibodies (from Roche Molecular Biochemicals) at 1:1000. The bound antibodies were detected by the ECL Western blotting detection system (Amersham Biosciences). Immunofluorescence and Fluorescence Microscopy Immunostaining of protein in yeast cells was performed as described (25) with the following modifications. The cells harboring the Ybr159p-GFP chromosomal fusion were transformed with plasmids expressing Elo3p-HA, Elo2p-HA, or Tsc13-MYC (the construction of these plasmids was described previously (16)). The cells were fixed with 4% methanol-free formaldehyde in PBS for 2 h at room temperature. Cells were permeabilized in phosphate-buffered saline containing 0.1% Triton X-100, 0.05% saponin, 10 mm glycine, and 0.1% bovine serum albumin. Before adding antibody, the digested cells were blocked by incubating in 10% of bovine serum albumin for 30 min. For detecting Elo3p-HA or Elo2p- HA, the cells were incubated with anti-ha rhodamine antibody (Roche Molecular Biochemicals) at 2 g/ml in phosphate-buffered saline containing 1.0% bovine serum albumin, 0.1% Tween 20 for 2 h. For detecting Tsc13p-MYC, cells were incubated with a monoclonal anti-myc antibody (from Invitrogen; 1:200 dilution) followed by the Cy3-conjugated anti-mouse IgG secondary antibody (Sigma; 1:5000). Fluorescence microscopy was performed on an IX70 inverted fluorescence microscope (Olympus) equipped with a HiQ fluorescein filter set (for GFP, excitation/emission was nm/510 nm; for rhodamine and Cy3, excitation/emission was nm/590 nm) and a Planapochromatic 100 oil immersion objective lens and a 100-watt mercury lamp. Images were collected with a Princeton Instruments 5-MHz MicroMax cooled CCD camera, a shutter and controller unit, and IPLab software (version 3.5; Scanalytic Co.). Signals from the fluorescein isothiocyanate (Ybr159-GFP) and rhodamine and Cy3 fluorescence channels were merged to unveil colocalization of signals. RESULTS Deletion of the YBR159w Gene Causes Slow Growth, Suppression of the Ca 2 -sensitive Phenotype Associated with the csg2 Mutation, and Synthetic Lethality with the elo2 Mutation The Ybr159p protein was previously shown to be required for C2 elongation of polyunsaturated fatty acids mediated by heterologous expression of a C. elegans polyunsaturated fatty acid-elongating activity F56H11.4 activity in S. cerevisiae (11). The Ybr159p requirement for polyunsaturated fatty acid elongation along with its homology to oxidoreductases suggested that it is the 3-ketoreductase activity of the endogenous elongase system in yeast. However, whereas the role of Ybr159p in heterologous fatty acid elongation has been demonstrated, whether or not it functions in endogenous elongation remained to be determined. The role of Ybr159p in the synthesis of endogenous VLCFAs is addressed in this study. The ybr159 mutant cells grow very slowly, especially upon germination from spores (Fig. 2A) and when growing at elevated temperature (e.g. 37 C) (11) (Fig. 2B). The presumptive Arabidopsis thaliana homolog (F12A21.31; designated At- FIG. 2. The Ybr159 mutation causes slow growth, synthetic lethality with the elo2 mutation, and suppression of the Ca 2 - sensitive phenotype caused by the csg2 mutation. A, a heterozygous ybr159::trp1/ybr159 diploid that had been transformed with a p-yes-ura3 plasmid carrying the Arabidopsis thaliana YBR159w homolog (F12A21.31) was sporulated and dissected on YPD agar plates. The plates were incubated at 26 C for 7 days and then photographed. Subsequent analysis revealed which spores had the wild-type YBR159 gene, which had the ybr159::trp1-disrupted gene, and which harbored the plasmid. The spore colonies from two representative tetrads are shown. B, the growth phenotypes of the elongase single and double mutants are shown. The indicated strains were grown in YPD medium to an A 600 of 0.1 and were diluted into the wells of a microtiter plate. The cells were transferred to either YPD or SD plates, and the plates were incubated at 26 C for 3 days or 37 C for 2 days. The ybr159 elo2 double mutant is missing because it is inviable. C, the ybr159 mutation suppresses the calcium sensitivity of the csg2 mutant. The indicated strains were streaked onto YPD agar plates with or without 50 mm CaCl 2, and the plates were incubated at 26 C for 3 days. The ybr159 csg2 double mutant is far less sensitive to calcium than the csg2 single mutant. YBR159) of the S. cerevisiae YBR159w gene complemented the deficiency in the heterologous microsomal elongation activity associated with the ybr159 mutation (11). Heterologous expression of the At-Ybr159p also complemented the slow growth
5 35444 Ybr159p Is the Major 3-Ketoreductase of Fatty Acid Elongation phenotype of the ybr159 mutant spores (Fig. 2A), demonstrating that the Arabidopsis gene also compensated for the endogenous (presumed VLCFA synthesis) defect that conferred slow growth. Mutations in the other known elongase genes (ELO2, ELO3, and TSC13) suppress the Ca 2 -sensitive phenotype of the csg2 mutation (16). The csg2 mutant accumulates high levels of inositolphosphoceramide (IPC) due to failure to mannosylate IPC (17, 26). The accumulation of IPC confers Ca 2 sensitivity, and mutants that reduce IPC levels suppress the Ca 2 -sensitive phenotype (15). In wild-type yeast, all ceramides and sphingolipids contain C26-VLCFA; therefore, mutations that reduce fatty acid elongation reduce the levels of IPC and thereby suppress the Ca 2 sensitivity conferred by the lack of Csg2p. As would be expected if Ybr159p is required for endogenous VLCFA synthesis, the ybr159 csg2 double mutant was far less Ca 2 -sensitive than the csg2 single mutant (Fig. 2C). We also investigated whether combining the ybr159 mutation with other elongase mutations would result in synthetic growth phenotypes. For these studies, we crossed a ybr159 mutant with a mutant harboring a second elongase mutation (the elo2, the elo3, orthetsc13-1 mutation) to generate the doubly heterozygous diploids. The products of meiosis were analyzed following sporulation and tetrad dissection. We found that the combination of the ybr159 mutation with an elo2 mutation was synthetically lethal, since all of the spores (from 16 tetrads) that could be deduced to have inherited both mutations (the ybr159 elo2 double mutant spores) failed to germinate (data not shown). The ybr159 elo3 and the ybr159 tsc13-1 double mutants were both viable, but they grew more slowly, especially at elevated temperatures, than the single mutants (Fig. 2B). The ybr159 Mutant, Like Other Elongase Mutants, Accumulates Long-chain Bases and Medium-chain Ceramides In previous studies, we found that mutants with defects in VLCFA synthesis accumulated very high levels of the free LCBs, phytosphingosine (PHS) and dihydrosphingosine (16). This phenotype is also observed for the ybr159 mutant (Fig. 3A). The accumulation of high levels of free LCBs in the elongase mutants does not reflect reduced partitioning of the LCBs into ceramides due to the VLCFA deficiency. Rather, the elongase mutants accumulate significant levels of medium-chain ceramides containing PHS (discussed below), and thus the total LCB levels (free LCBs and LCBs incorporated into ceramides and sphingolipids) are increased in the mutants. The elevated LCB pool suggested an increased rate of synthesis of the LCBs in the elongase mutants. Therefore, we tested whether serine palmitoyltransferase (SPT), the committed and presumed rate-limiting enzyme of LCB synthesis, was up-regulated in the elongase mutants. The abundance of the Lcb1p and Lcb2p subunits of SPT was similar in microsomes prepared from wild-type and elongase mutant cells (Fig. 3B). We also measured SPT activity in microsomes prepared from wild-type and elongase mutant microsomes and found no increase in the in vitro SPT activity for the elongase mutants (Fig. 3C). These experiments indicate that the elevated LCBs do not result from increased expression of the SPT enzyme. It is possible that the reduced partitioning of palmitoyl-coa (a common substrate for SPT and the condensing enzyme of the elongase system) into the elongase pathway results in enhanced LCB synthesis by providing higher substrate pools for SPT. However, this is not likely, because a similar LCB-accumulating phenotype is reported for the lac1 lagl mutant cells, but in this case the VLCFA levels are also elevated (27, 28) (see Discussion ). FIG. 3. Similar to the other elongase mutants, the ybr159 mutant accumulates high levels of LCBs. A, LCBs were extracted from 10 A 600 units of the indicated cells, separated by TLC, and visualized by ninhydrin staining. The LCB standards sphingosine (SPH), 3-ketosphingosine (3-KS), dihydrosphingosine (DHS), and phytosphingosine (PHS) were spotted in lanes 1 4 as indicated. The ninhydrinreactive species that migrate just above PHS and near the origin are phosphatidylethanolamine (PE) and phosphatidylserine (PS) (see standards, lanes 14 and 15). B, the elevated LCB levels in the elongase mutants do not result from increased expression of the Lcb1p or Lcb2p subunits of SPT. Ten g of microsomal proteins from the indicated strains were resolved by SDS-PAGE electrophoresis, transferred to nitrocellulose, and subjected to Western blot analysis using affinitypurified polyclonal anti-lcb1p and anti-lcb2p antibodies as previously described (33). C, the in vitro SPT activity, measured using microsomes from the indicated elongase mutants (33), is similar to the activity in microsomes from the wild-type (wt) cells. Whereas the ceramide (C-ceramide) in wild-type cells contains PHS and -OH-C26 fatty acids, the elongase mutants also accumulated significant levels of ceramides with shorter chain fatty acids, and these fatty acids were also -hydroxylated (16). These medium-chain (relatively hydrophilic) ceramides also accumulated in the ybr159 mutant (Fig. 4A). The presumptive medium-chain ceramides were eluted from the TLC plate and subjected to acid methanolysis, and the resultant LCBs (by TLC) and FAMEs (by GCMS) were analyzed. The ceramides were composed of PHS and -OH-C16 fatty acids
6 Ybr159p Is the Major 3-Ketoreductase of Fatty Acid Elongation FIG. 4. The elongase mutants accumulate medium-chain ceramides: Synthesis of the medium-chain ceramides is not mediated by the Ypc1p and/or Ydc1p ceramidases. A, ceramides were extracted from 10 A 600 units of the indicated cells, separated by TLC, and visualized by charring (23). Bovine ceramide type III (consisting of sphingosine and C16-FA) and type IV (consisting of sphingosine and -OH-C16-FA) standards run at the indicated positions. The major ceramide in wild-type cells (C-ceramide) consists of PHS and -OH- C26-FA. The elongase mutants have reduced C-ceramide, and they accumulate high levels of relatively hydrophilic ceramides that are composed of PHS and -OH-C16-FA (the medium-chain ceramides). B, ceramides from 12.5 A 600 units of the indicated strains were extracted and analyzed by TLC. In this case, the ceramides were analyzed by UV light after spraying the plates with 1% 8-anilino-1-napthalenesulfonic acid. The deletion of YPC1 (lane 3), YDC1 (lane 4), or both YPC1 and YDC1 (lane 9) did not prevent the medium-chain ceramides from accumulating in the elo2 mutant. The medium-chain ceramides from the bracketed region were eluted from the TLC plate and subjected to acid methanolysis, and the FAMEs and LCBs that were produced were recovered and analyzed. (data not shown). A small amount of ceramide that was unhydroxylated on the C16 fatty acid (labeled B-ceramide in Fig. 4A) was also present and migrated in the TLC between the type III (unhydroxylated) and type IV (hydroxylated) bovine ceramide standards. In addition, a novel species that migrated similarly to the B-ceramide accumulated in the ybr159 mutant (Fig. 4A). This species was found to be a 3-OH fatty acid (discussed below). These medium-chain ceramides might be synthesized by the acyl-coa-dependent ceramide synthase activity; however, in wild-type cells, this enzyme appears to have high selectivity for C26 fatty acyl CoA, despite the relatively high intracellular level of C16 fatty acids. Alternatively, the medium-chain ceramides could arise from the reversal of a ceramidase activity, possibly driven by the high LCB levels in the elongase mutants. Two ceramidase-encoding genes, YPC1 and YDC1, with specificity toward PHS-containing and dihydrosphingosine-containing ceramides respectively, have been identified in yeast (20, 29). To determine whether the accumulation of the medium-chain ceramides in the elongase mutants depends on Ypc1p and/or Ydc1p, the ceramides from an elo2 mutant, an elo2 ypc1 or an elo2 ydc1 double mutant, or an elo2 ypc1 ydc1 triple mutant were compared. The mediumchain ceramide accumulation caused by the elo2 mutation was not blocked by eliminating the ceramidase genes (Fig. 4B). The medium-chain ceramides from these strains were purified and hydrolyzed by acid methanolysis, and again the LCB moiety of these ceramides was found to be PHS, and the fatty acid moiety was found to be predominantly -OH-C16 (data not shown). Since deleting either YPC1 and/or YDC1 did not prevent the accumulation of the medium-chain ceramides, they are most likely synthesized by the acyl-coa-dependent ceramide synthase. This raises the question of why ceramide synthase, which normally displays high selectivity for C26- acyl-coa, uses palmitoyl-coa in the elongase mutants (see Discussion ). These results show that the LCB- and medium-chain ceramide-accumulating phenotypes previously found to be characteristic of elongase mutants are also observed for the ybr159 mutant and support a role for Ybr159p in endogenous VLCFA synthesis. In addition, they demonstrate that the high levels of LCBs in the mutants do not result from increased expression of the Lcb1p and/or Lcb2p subunit of SPT. Finally, the mediumchain ceramides that accumulate in the elongase mutants do not depend on the presence of the ceramidases, Ypc1p or Ydc1p. In Vivo and in Vitro Assays of VLCFA Synthesis Confirm a Deficiency in the ybr159 Mutant Fatty acids from wild-type, ybr159, ybr159 tsc13-1, and ybr159 elo3 cells were extracted and analyzed by GCMS (see Experimental Procedures ). As shown in Fig. 5, there was a significant reduction in the levels of the C26 fatty acids in cells harboring the ybr159 mutation. In addition, several fatty acid species were observed in the mutants that were not present in wild-type cells, including -OH-C16 fatty acid. As discussed above, the elongase mutants synthesize ceramides that have C16 fatty acids, whereas in wild-type cells the ceramides have exclusively C26 fatty acids. These medium-chain ceramides are substrates for the -hydroxylating enzyme, Scs7p, and thereby generate the -OH-C16 fatty acids (16, 23). Thus, the accumulation of the -OH-C16 fatty acids in the ybr159 mutant reflects the accumulation of the medium-chain ceramides, a phenotype diagnostic of a VLCFA synthesis deficiency. In addition to the -OH-C16 fatty acids, the ybr159 mutant cells also accumulated 3-OH fatty acids of different chain lengths (C16, C18, and C20). There are no known enzymes that hydroxylate fatty acids or ceramides at C3; rather, these fatty acids are presumed to be intermediates of the elongation pathway (Fig. 1). As mentioned above, the 3-OH fatty acids from the ybr159 mutant were also observed on the ceramide TLC plates (see Fig. 4A). The accumulation of these 3-OH fatty acid species suggests that the ybr159 mutant is deficient in the dehydratase activity as well as in the 3-ketoreductase activity of the elongation system (discussed further below). We previously assayed microsomes prepared from the ybr159 mutant for in vitro elongase activity (11). The first step of the elongation cycle is the condensation of malonyl-coa with an acyl-coa (e.g. palmitoyl-coa) to form a 3-keto-acyl- CoA intermediate (Fig. 1). Omitting pyridine nucleotide from the assay mix prevents the reduction of the 3-keto-acyl-CoA intermediate and thereby allows the first step of elongation (condensation) to be measured (Fig. 6, lane 1 of each panel). A time course of the overall elongation was measured in the presence of NADH/NADPH (Fig. 6, lanes 2 4 of each panel).
7 35446 Ybr159p Is the Major 3-Ketoreductase of Fatty Acid Elongation FIG. 5. The ybr159 mutant cells are deficient in C26 fatty acids and accumulate 2-OH-C16 and 3-OH-C16, -C18, and -C20 Fatty Acids. FAMEs were derived from the indicated strains and were analyzed by GCMS. The profile spanning retention times from 9 to 21 min is shown. An internal C17 standard (retention time, 8.8 min), added to the cells prior to the extraction, was used to normalize the data. A small percentage of the fatty acids were not methylated (labeled free fatty acids (FFA)). Elongation intermediates did not accumulate during the elongation reactions catalyzed by the wild-type microsomes (Fig. 6a), which is consistent with previous studies of the rat microsomal elongating systems demonstrating that the condensation reaction is rate-limiting (30). However, when microsomes prepared from the ybr159 mutant were used in the assay, accumulation of the 3-keto-acyl and 3-hydroxyacyl intermediates was observed (11) (Fig. 6b). This is consistent with the defect in Ybr159p causing reduced activity of both the 3-ketoreductase and the dehydratase activity of the elongase system. We had also previously assayed in vitro elongation using microsomes prepared from the tsc13-1 mutant. In this case, accumulation of the trans-2,3-stearoyl and the 3-hydroxystearoyl intermediates was observed (16) (Fig. 6e). This is consistent with the defect in Tsc13p causing reduced activity of the trans-2,3-enoyl-coa reductase leading to accumulation of the trans-2,3-stearoyl-coa. Because the dehydratase step of elongation is reversible (31), accumulation of trans-2,3-stearoyl- CoA resulted in the observed accumulation of 3-hydroxystearoyl-CoA as well. These previous studies were extended by comparing the elongase activity of microsomes prepared from mutants with various combinations of the elongase mutations (Fig. 6, f i). These experiments confirmed our previous finding that when Ybr159p was missing, the 3-keto-acyl intermediate accumulated, consistent with Ybr159p being the major 3-ketoreductase activity of the microsomal yeast fatty acid elongase system. In addition, when Ybr159p was missing, the 3-OH-acyl intermediate of fatty acid elongation accumulated, indicating that the dehydratase activity was also compromised in the absence of Ybr159p. Significantly, when Tsc13p activity was deficient, the 3-OH-acyl intermediate accumulated in proportion to the trans-2,3-acyl intermediate, as would be expected if it arose from reversal of the dehydratase activity. On the other hand, when Ybr159p was missing, there was a large increase in the 3-OH-acyl intermediate with no concomitant accumulation of the trans-2,3-acyl intermediate. This indicates that when Ybr159p is missing, the forward dehydratase activity is decreased to cause the 3-OH-acyl intermediate to accumulate, whereas when Tsc13p is missing, the reverse dehydratase activity is increased to cause the 3-OH-acyl intermediate to accumulate. The in vitro elongase activity measured using the microsomes from the elo2 (Fig. 6c) and elo3 (Fig. 6d) single mutants appeared very similar to that with the wild-type microsomes, in that no intermediates accumulated. However, the condensation activity in the elo2 mutant was reduced about 3-fold (Fig. 6c), consistent with Elo2p being required for efficient condensation of palmitoyl-coa with malonyl-coa. The reduced overall elongation in both the elo2 and the ybr159 mutants is also consistent with the observation that the elo2 ybr159 double mutant is inviable. Although the elo3 ybr159 double mutant accumulates both the 3-keto and the 3-OH acyl intermediates, less 3-keto intermediate forms than in the ybr159 single mutant (Fig. 6f). This suggests that the 3-keto intermediate that is formed by Elo2p (since Elo3p is missing) is reduced more efficiently by an alternative 3-ketoreductase than the 3-keto intermediate formed by Elo3p, but this requires further investigation. The ybr159 mutation does not prevent the accumulation of the trans-2,3-enoyl intermediate in the ybr159 tsc13-1 double mutant; nor does the tsc13-1 mutation prevent the high 3-OH acyl intermediate associated with the ybr159 mutation from accumulating (Fig. 6g). In summary, the accumulation of the 3-ketoacyl intermediate (in vitro) and the 3-OH acyl intermediate (both in vivo and in vitro) indicate that both the 3-ketoreductase and the dehydratase activities are compromised in the ybr159 mutant. The accumulation of these intermediates is not blocked by the elo3 mutation or by the tsc13-1 mutation. The synthetic lethal phenotype of the elo2 ybr159 double mutant indicates that the reduced overall elongation observed by either single mutant is additive, making the double mutant unable to synthesize sufficient VLCFAs for viability. The Ybr159p Protein Resides in the Endoplasmic Reticulum, Co-localizes with Tsc13p and Elo3p, and Also Co-immunoprecipitates with Tsc13p and Elo3p Tsc13p, Elo3p, and Elo2p all displayed a perinuclear and peripheral staining consistent with localization to the ER (16, 32). Furthermore, Tsc13p was found to co-immunoprecipitate with Elo2p and with Elo3p, indicating that the elongase proteins are organized in a complex. We next addressed whether Ybr159p co-localizes with and associates with the other elongase proteins. Ybr159p has a canonical dilysine ER retention motif, which along with its homology to the oxidoreductases, was the criterion used to identify it as a potential 3-ketoreductase of the elongase system (11). In addition, all enzymes involved in fatty acid elongation
8 Ybr159p Is the Major 3-Ketoreductase of Fatty Acid Elongation FIG. 6.The ybr159 mutant cells have normal condensation activity but are deficient in 3-ketoreductase and dehydratase activity. Fatty acid elongation activity in microsomes prepared from the indicated strains was compared using C16-CoA as a substrate by measuring the incorporation of radiolabeled malonyl-coa into hexane-extractable fatty acids. The assays were conducted in the absence of NADPH/NADH for 5 min to measure condensation activity (lane 1 of each set). NADPH/NADH was included in the reactions measuring total elongation for 0.2 (lane 2), 1.0 (lane 3), or 5.0 (lane 4) min. The reactions were stopped at the indicated times, and the fatty acids were extracted and separated by TLC. The positions of the 3-ketostearate (3-Keto), stearate (FA), trans-2,3-stearate (Trans-2,3), and 3-hydroxystearate (3-Hydroxy) intermediates were determined by running the standards on the TLC plate and charring after exposure to PhosphorImager screens. identified so far have very basic pi values ( 9) and several transmembrane domains, which is also characteristic for Ybr159p. To analyze subcellular localization, Ybr159p was C-terminally tagged with GFP by chromosomal fusion (see Experimental Procedures ). The Ybr159p-GFP tagged protein was functional because the strain grew normally, and the 2-OH and 3-OH fatty acids (diagnostic of an elongase defect) did not accumulate. Indeed, localization studies of Ybr159p-GFP confirmed that it localizes to the nuclear/peripheral ER membrane (Fig. 7A, panel a). The Ybr159-GFP protein co-localizes with Elo3p-HA (Fig. 7A, panel b), Elo2p-HA (Fig. 7A, panel c), and Tsc13p-Myc (Fig. 7A, panel d). The epitope-tagged proteins were detected by immunofluorescence using either rhodamineconjugated anti-ha (for Elo3p and Elo2p) or Cy3-conjugated anti-myc (for Tsc13p-Myc) antibodies. Detailed microscopic analyses of GFP-tagged Ybr159p unveiled exclusive ER localization throughout various growth phases and no enrichment in the nuclear-vacuolar junctions (not shown). Thus, the physiological relevance of an enrichment of one component of the microsomal fatty acid complex, Tsc13p, to nuclear-vacuolar junctions remains obscure (16). For the immunoprecipitation experiments, solubilized microsomes were prepared from cells that were co-expressing Tsc13p-Myc and Ybr159p-GFP either with or without Elo3p-HA (Fig. 7B). As we reported previously, the anti-ha antibodies pulled down Tsc13p-Myc, and the anti-myc antibodies pulled down Elo3p-HA when Tsc13p-Myc and Elo3p-HA were co-expressed (Fig. 7B). These experiments also indicated that Ybr159p-GFP associates with the Elo3p-HA/Tsc13p-Myccontaining complexes, since anti-gfp antibodies pulled down Tsc13p-Myc and Elo3p-HA (Fig. 7B). Whether or not Ybr159p co-immunoprecipitates with Elo2p has not yet been tested. The Residual 3-Ketoreductase Activity in the Ybr159 Mutant Is Likely to Depend on the AYR1 Gene Product The results reported above indicate that the YBR159w gene encodes the major 3-ketoreductase activity of the yeast elongase system of enzymes. However, it cannot be the only gene that encodes such an activity, because there is residual VLCFA synthesis in the ybr159 mutant (Fig. 5). Furthermore, other mutants that fail to synthesize VLCFAs (the tsc13 mutant and the elo2 elo3 double mutant) are inviable, whereas the ybr159 mutant grows, albeit slowly. Therefore, we investigated whether any of the genes most closely related to YBR159w was likely to encode the residual 3-ketoreductase activity. We reasoned that disruption of the gene that encodes the residual 3-ketoreductase activity would be lethal in combination with the ybr159 mutation. There are several other putative oxidoreductase-encoding genes in yeast that display homologies to the YBR159w gene (11) (Table II). We obtained the set of haploid disruptant mutants in which each of these genes had been replaced with the kanmx resistance marker (from Research Genetics) and crossed them to the haploid ybr159::ura3 mutant. For four of five candidates, Geneticinresistant uracil-prototrophic segregants of the heterozygous diploid strains were recovered, indicating that the double mutants were viable. However, for the diploid that was heterozygous for the yil124w/ayr1::kanmx and the ybr159::ura3 mutations, no viable Geneticin-resistant uracil-prototrophic meiotic segregants were recovered. Therefore, disruption of the AYR1 gene was demonstrated to be synthetically lethal with the ybr159 mutation. In a previous study (18), spores harboring the ayr1 mutation were found to be unable to germinate, raising the possibility that the AYR1 gene was not disrupted in the strain purchased from Research Genetics. However, we confirmed the ayr1::kanmx disruption in this strain by PCR. In our studies, the ayr1::kanmx single mutant spores germinated as well as wild type; this phenomenon may be related to differences in the construction of the disrupting allele or in the yeast strains. To further investigate the possibility that Ayr1p catalyzes 3-ketoreductase activity in the ybr159 mutant, we addressed whether overexpression of Ayr1p would suppress the slow growth phenotype of ybr159 mutant spore colonies. However, a high copy number (prs424-trp1-based) AYR1-containing plasmid did not suppress the slow growth of ybr159 mutant spore colonies. The ayr1::kanmx mutant was also analyzed for elongase phenotypes. However, the mutant did not accumulate LCBs or medium-chain ceramides. Furthermore, in vivo fatty acid analyses and in vitro elongase assays did not reveal any evidence of a VLCFA synthesis defect in the ayr1 mutant. Finally, the ayr1 mutation was not synthetically lethal with the elo2 or the elo3 mutation. Therefore, although the synthetic lethality of the ayr1 ybr159 double mutant suggests that Ayr1p provides the residual 3-ketoreductase activity in the ybr159 mutant, the lack of elongase phenotypes in the
9 35448 Ybr159p Is the Major 3-Ketoreductase of Fatty Acid Elongation ayr1 single mutant indicates that it is a minor activity. This is consistent with Ybr159p being the major 3-ketoreductase of the microsomal elongase. DISCUSSION The main conclusion from this study is that Ybr159p is the major 3-ketoreductase activity of the endogenous yeast elongase system of enzymes required for VLCFA synthesis. This conclusion is based on several observations. The ybr159 mutant shares many of the phenotypes that are characteristic of previously studied elongase mutants, including high levels of LCBs and of medium-chain ceramides as well as low levels of VLCFAs. The ybr159 mutation is synthetically lethal in combination with an elo2 mutation, and the ybr159 mutation is a suppressor of the csg2 mutation. The in vitro elongation assays reveal a defect in the 3-ketoreductase activity in the mutant. These phenotypes, combined with the homology of Ybr159p to other members of the oxidoreductases, support our conclusion that this protein is a 3-ketoreductase. Although Ybr159p is a major 3-ketoreductase responsible for VLCFA synthesis, it is not the only enzyme in yeast responsible for this activity. We provide evidence that Ayr1p is responsible for the residual 3-ketoreductase activity in the ybr159 mutant. Ayr1p was previously reported to be responsible for all of the 1-acyldihydroxyacetone-phosphate reductase activity of lipid particles as well as a significant fraction of this activity in microsomes (18). That this enzyme apparently reduces both 1-acyldihydroxyacetone phosphate and 3-keto-acyl elongation intermediates suggests that it is quite promiscuous. Whereas our data suggest that Ayr1p has 3-ketoreductase activity, it is possible that Ayr1p is essential in a ybr159 mutant for another reason. For example, its role in phosphatidic acid biosynthesis through the dihydroxyacetone pathway (18) may alter glycerophospholipid metabolism, and the combination of defects in VLCFA synthesis and glycerophospholipid metabolism may result in the synthetic lethal phenotype of the ayr1 ybr159 double mutant. We note, however, that deletion of the ayr1 gene in other elongase mutants (elo2 and elo3 ) does not confer synthetic lethality. In addition to reduced 3-ketoreductase activity, the ybr159 mutant is also deficient in the dehydratase activity that converts the 3-hydroxy intermediates formed during elongation to the trans-2,3-enoyl intermediates. The basis of the dehydratase deficiency in the ybr159 mutant is not understood, but several possibilities are suggested. For example, the Ybr159p enzyme could be bifunctional and catalyze both the 3-keto reduction and the subsequent dehydration reaction. This seems unlikely, because the protein is homologous to characterized oxidoreductases over its entire length. Another possibility is that the stability and or proper localization of the dehydratase depends on the presence of Ybr159p. There are many examples in which the loss of one component of a multienzyme complex results in the instability of other components of the complex. A third possibility is that the lack of Ybr159p, while not affecting the stability of the dehydratase, precludes formation of the elongase complex. Clearly, the condensation step is not affected by FIG. 7.Ybr159p-GFP resides in the nuclear/er membrane; colocalizes with Elo3p-HA, Elo2p-HA, and Tsc13p-Myc; and coimmunoprecipitates with Elo3p-HA and Tsc13p-Myc. A, C-terminally tagged Ybr159p-GFP shows a typical ER localization pattern (i.e. around the nucleus and cell periphery). Ybr159p-GFP co-localizes with Elo3p, Elo2p, and Tsc13p. a, GFP fluorescence of Ybr159p-GFP; b, GFP fluorescence of Ybr159p-GFP and immunofluorescence of Elo3p-HA with rhodamine-conjugated anti-ha; c, GFP fluorescence of Ybr159p- GFP and immunofluorescence of Elo2p-HA with rhodamine-conjugated anti-ha; d, GFP fluorescence of Ybr159p-GFP and immunofluorescence of Tsc13p-Myc with Cy3-conjugated anti-myc. B, anti-gfp antibodies coimmunoprecipitate Elo3p-HA and Tsc13p-Myc with Ybr159p-GFP. Microsomes were prepared from cells containing Ybr159-GFP with Tsc13p-Myc alone (on left) or with Tsc13p-Myc and Elo3p-HA. The microsomes were solubilized, and the 100,000 g supernatant was used for immunoprecipitation with Sephadex beads (lane 1) or Sephadex beads conjugated to anti-gfp (lane 2), anti-ha (lane 3), or anti- Myc (lane 4) antibodies. The immunoprecipitated proteins were separated by SDS-PAGE and analyzed by immunoblotting with horseradish peroxidase-conjugated anti-myc or anti-ha antibodies as indicated.
Synthesis and elongation of fatty acids
 Synthesis and elongation of fatty acids A molecular caliper mechanism for determining very long-chain fatty acid length Vladimir Denic and Jonathan S. Weissman (2007) Cell 130, 663-677 February 28, 2008
Synthesis and elongation of fatty acids A molecular caliper mechanism for determining very long-chain fatty acid length Vladimir Denic and Jonathan S. Weissman (2007) Cell 130, 663-677 February 28, 2008
Very-Long Chain Fatty Acid Biosynthesis
 Very-Long Chain Fatty Acid Biosynthesis Objectives: 1. Review information on the isolation of mutants deficient in VLCFA biosynthesis 2. Generate hypotheses to explain the absence of mutants with lesions
Very-Long Chain Fatty Acid Biosynthesis Objectives: 1. Review information on the isolation of mutants deficient in VLCFA biosynthesis 2. Generate hypotheses to explain the absence of mutants with lesions
Very-Long Chain Fatty Acid Biosynthesis
 Very-Long Chain Fatty Acid Biosynthesis Objectives: 1. Review information on the isolation of mutants deficient in VLCFA biosynthesis 2. Generate hypotheses to explain the absence of mutants with lesions
Very-Long Chain Fatty Acid Biosynthesis Objectives: 1. Review information on the isolation of mutants deficient in VLCFA biosynthesis 2. Generate hypotheses to explain the absence of mutants with lesions
Very-Long Chain Fatty Acid Biosynthesis
 Very-Long Chain Fatty Acid Biosynthesis Objectives: 1. Review information on the isolation of mutants deficient in VLCFA biosynthesis 2. Generate hypotheses to explain the absence of mutants with lesions
Very-Long Chain Fatty Acid Biosynthesis Objectives: 1. Review information on the isolation of mutants deficient in VLCFA biosynthesis 2. Generate hypotheses to explain the absence of mutants with lesions
Very-Long Chain Fatty Acid Biosynthesis
 Very-Long Chain Fatty Acid Biosynthesis Objectives: 1. Review information on the isolation of mutants deficient in VLCFA biosynthesis 2. Generate hypotheses to explain the absence of mutants with lesions
Very-Long Chain Fatty Acid Biosynthesis Objectives: 1. Review information on the isolation of mutants deficient in VLCFA biosynthesis 2. Generate hypotheses to explain the absence of mutants with lesions
TFEB-mediated increase in peripheral lysosomes regulates. Store Operated Calcium Entry
 TFEB-mediated increase in peripheral lysosomes regulates Store Operated Calcium Entry Luigi Sbano, Massimo Bonora, Saverio Marchi, Federica Baldassari, Diego L. Medina, Andrea Ballabio, Carlotta Giorgi
TFEB-mediated increase in peripheral lysosomes regulates Store Operated Calcium Entry Luigi Sbano, Massimo Bonora, Saverio Marchi, Federica Baldassari, Diego L. Medina, Andrea Ballabio, Carlotta Giorgi
ANSC/NUTR 618 LIPIDS & LIPID METABOLISM. Fatty Acid Elongation and Desaturation
 ANSC/NUTR 618 LIPIDS & LIPID METABOLISM I. Fatty acid elongation A. General 1. At least 60% of fatty acids in triacylglycerols are C18. 2. Free palmitic acid (16:0) synthesized in cytoplasm is elongated
ANSC/NUTR 618 LIPIDS & LIPID METABOLISM I. Fatty acid elongation A. General 1. At least 60% of fatty acids in triacylglycerols are C18. 2. Free palmitic acid (16:0) synthesized in cytoplasm is elongated
Protocol for Gene Transfection & Western Blotting
 The schedule and the manual of basic techniques for cell culture Advanced Protocol for Gene Transfection & Western Blotting Schedule Day 1 26/07/2008 Transfection Day 3 28/07/2008 Cell lysis Immunoprecipitation
The schedule and the manual of basic techniques for cell culture Advanced Protocol for Gene Transfection & Western Blotting Schedule Day 1 26/07/2008 Transfection Day 3 28/07/2008 Cell lysis Immunoprecipitation
LIPID METABOLISM
 LIPID METABOLISM LIPOGENESIS LIPOGENESIS LIPOGENESIS FATTY ACID SYNTHESIS DE NOVO FFA in the blood come from :- (a) Dietary fat (b) Dietary carbohydrate/protein in excess of need FA TAG Site of synthesis:-
LIPID METABOLISM LIPOGENESIS LIPOGENESIS LIPOGENESIS FATTY ACID SYNTHESIS DE NOVO FFA in the blood come from :- (a) Dietary fat (b) Dietary carbohydrate/protein in excess of need FA TAG Site of synthesis:-
Unsaturated fatty acids are essential components required for
 Heterologous reconstitution in yeast of the polyunsaturated fatty acid biosynthetic pathway Frédéric Beaudoin, Louise V. Michaelson, Sandra J. Hey, Mervyn J. Lewis, Peter R. Shewry, Olga Sayanova, and
Heterologous reconstitution in yeast of the polyunsaturated fatty acid biosynthetic pathway Frédéric Beaudoin, Louise V. Michaelson, Sandra J. Hey, Mervyn J. Lewis, Peter R. Shewry, Olga Sayanova, and
Supplemental Figure S1
 Supplemental Figure S1 GC-MS profile of total FA of BY-2 purified PM IS: h14 IS: 17: Glycerolipids 16: GIPCs 18:1,2,3 18: h22 h24 Sterols GluCER h16: 2: h2 22: h21 c23 24: h23 h25 h26 Supplemental Figure
Supplemental Figure S1 GC-MS profile of total FA of BY-2 purified PM IS: h14 IS: 17: Glycerolipids 16: GIPCs 18:1,2,3 18: h22 h24 Sterols GluCER h16: 2: h2 22: h21 c23 24: h23 h25 h26 Supplemental Figure
SUPPLEMENTARY INFORMATION
 Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
BCM 221 LECTURES OJEMEKELE O.
 BCM 221 LECTURES BY OJEMEKELE O. OUTLINE INTRODUCTION TO LIPID CHEMISTRY STORAGE OF ENERGY IN ADIPOCYTES MOBILIZATION OF ENERGY STORES IN ADIPOCYTES KETONE BODIES AND KETOSIS PYRUVATE DEHYDROGENASE COMPLEX
BCM 221 LECTURES BY OJEMEKELE O. OUTLINE INTRODUCTION TO LIPID CHEMISTRY STORAGE OF ENERGY IN ADIPOCYTES MOBILIZATION OF ENERGY STORES IN ADIPOCYTES KETONE BODIES AND KETOSIS PYRUVATE DEHYDROGENASE COMPLEX
The Schedule and the Manual of Basic Techniques for Cell Culture
 The Schedule and the Manual of Basic Techniques for Cell Culture 1 Materials Calcium Phosphate Transfection Kit: Invitrogen Cat.No.K2780-01 Falcon tube (Cat No.35-2054:12 x 75 mm, 5 ml tube) Cell: 293
The Schedule and the Manual of Basic Techniques for Cell Culture 1 Materials Calcium Phosphate Transfection Kit: Invitrogen Cat.No.K2780-01 Falcon tube (Cat No.35-2054:12 x 75 mm, 5 ml tube) Cell: 293
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
UNIFORMED SERVICES UNIVERSITY OF THE HEALTH SCIENCES F. EDWARD HEBERT SCHOOL OF MEDICINE 4301 JONES BRIDGE ROAD BETHESDA, MARYLAND
 UNIFORMED SERVICES UNIVERSITY OF THE HEALTH SCIENCES F. EDWARD HEBERT SCHOOL OF MEDICINE 4301 JONES BRIDGE ROAD BETHESDA, MARYLAND 20814-4799 March 6, 2007 APPROVAL SHEET Title ofdissertation: Elongation"
UNIFORMED SERVICES UNIVERSITY OF THE HEALTH SCIENCES F. EDWARD HEBERT SCHOOL OF MEDICINE 4301 JONES BRIDGE ROAD BETHESDA, MARYLAND 20814-4799 March 6, 2007 APPROVAL SHEET Title ofdissertation: Elongation"
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
BIOSYNTHESIS OF FATTY ACIDS. doc. Ing. Zenóbia Chavková, CSc.
 BIOSYNTHESIS OF FATTY ACIDS doc. Ing. Zenóbia Chavková, CSc. The pathway for the of FAs is not the reversal of the oxidation pathway Both pathways are separated within different cellular compartments In
BIOSYNTHESIS OF FATTY ACIDS doc. Ing. Zenóbia Chavková, CSc. The pathway for the of FAs is not the reversal of the oxidation pathway Both pathways are separated within different cellular compartments In
Biosynthesis of Fatty Acids
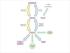 Biosynthesis of Fatty Acids Fatty acid biosynthesis takes place in the cytosol rather than the mitochondria and requires a different activation mechanism and different enzymes and coenzymes than fatty
Biosynthesis of Fatty Acids Fatty acid biosynthesis takes place in the cytosol rather than the mitochondria and requires a different activation mechanism and different enzymes and coenzymes than fatty
Fatty Acid Desaturation
 Fatty Acid Desaturation Objectives: 1. Isolation of desaturase mutants 2. Substrates for fatty acid desaturation 3. ellular localization of desaturases References: Buchanan et al. 2000. Biochemistry and
Fatty Acid Desaturation Objectives: 1. Isolation of desaturase mutants 2. Substrates for fatty acid desaturation 3. ellular localization of desaturases References: Buchanan et al. 2000. Biochemistry and
Biosynthesis of Fatty Acids. By Dr.QUTAIBA A. QASIM
 Biosynthesis of Fatty Acids By Dr.QUTAIBA A. QASIM Fatty Acids Definition Fatty acids are comprised of hydrocarbon chains terminating with carboxylic acid groups. Fatty acids and their associated derivatives
Biosynthesis of Fatty Acids By Dr.QUTAIBA A. QASIM Fatty Acids Definition Fatty acids are comprised of hydrocarbon chains terminating with carboxylic acid groups. Fatty acids and their associated derivatives
number Done by Corrected by Doctor Faisal Al-Khatibe
 number 24 Done by Mohammed tarabieh Corrected by Doctor Faisal Al-Khatibe 1 P a g e *Please look over the previous sheet about fatty acid synthesis **Oxidation(degradation) of fatty acids, occurs in the
number 24 Done by Mohammed tarabieh Corrected by Doctor Faisal Al-Khatibe 1 P a g e *Please look over the previous sheet about fatty acid synthesis **Oxidation(degradation) of fatty acids, occurs in the
Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants
 Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants Species UDP-2,3- diacylglucosamine hydrolase specific activity (nmol min -1 mg -1 ) Fold vectorcontrol specific
Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants Species UDP-2,3- diacylglucosamine hydrolase specific activity (nmol min -1 mg -1 ) Fold vectorcontrol specific
Prerequisites Protein purification techniques and protein analytical methods. Basic enzyme kinetics.
 Case 19 Purification of Rat Kidney Sphingosine Kinase Focus concept The purification and kinetic analysis of an enzyme that produces a product important in cell survival is the focus of this study. Prerequisites
Case 19 Purification of Rat Kidney Sphingosine Kinase Focus concept The purification and kinetic analysis of an enzyme that produces a product important in cell survival is the focus of this study. Prerequisites
A mutant in Arabidopsis Lacking a Chloroplast Specific Lipid. Lewis Kurschner and Karen Thulasi Masters in Botany
 A mutant in Arabidopsis Lacking a Chloroplast Specific Lipid Lewis Kurschner and Karen Thulasi Masters in Botany Fatty acid nomenclature Fatty acyl composition Chain length Degree of unsaturation and position
A mutant in Arabidopsis Lacking a Chloroplast Specific Lipid Lewis Kurschner and Karen Thulasi Masters in Botany Fatty acid nomenclature Fatty acyl composition Chain length Degree of unsaturation and position
ANSC/NUTR 618 Lipids & Lipid Metabolism
 I. Overall concepts A. Definitions ANC/NUTR 618 Lipids & Lipid Metabolism 1. De novo synthesis = synthesis from non-fatty acid precursors a. Carbohydrate precursors (glucose, lactate, and pyruvate) b.
I. Overall concepts A. Definitions ANC/NUTR 618 Lipids & Lipid Metabolism 1. De novo synthesis = synthesis from non-fatty acid precursors a. Carbohydrate precursors (glucose, lactate, and pyruvate) b.
Overview on the identification of different classes of. lipids by HPTLC (High Performance Thin Layer. Chromatography) and ITLC (Immuno Thin Layer
 Overview on the identification of different classes of lipids by HPTLC (High Performance Thin Layer Chromatography) and ITLC (Immuno Thin Layer Chromatography) Iuliana Popa 1, Marie-Jeanne David 2, Daniel
Overview on the identification of different classes of lipids by HPTLC (High Performance Thin Layer Chromatography) and ITLC (Immuno Thin Layer Chromatography) Iuliana Popa 1, Marie-Jeanne David 2, Daniel
Lecture: 26 OXIDATION OF FATTY ACIDS
 Lecture: 26 OXIDATION OF FATTY ACIDS Fatty acids obtained by hydrolysis of fats undergo different oxidative pathways designated as alpha ( ), beta ( ) and omega ( ) pathways. -oxidation -Oxidation of fatty
Lecture: 26 OXIDATION OF FATTY ACIDS Fatty acids obtained by hydrolysis of fats undergo different oxidative pathways designated as alpha ( ), beta ( ) and omega ( ) pathways. -oxidation -Oxidation of fatty
Fatty acids synthesis
 Fatty acids synthesis The synthesis start from Acetyl COA the first step requires ATP + reducing power NADPH! even though the oxidation and synthesis are different pathways but from chemical part of view
Fatty acids synthesis The synthesis start from Acetyl COA the first step requires ATP + reducing power NADPH! even though the oxidation and synthesis are different pathways but from chemical part of view
Isolation and Characterization of Sphingolipids in Arabidopsis thaliana
 Isolation and Characterization of Sphingolipids in Arabidopsis thaliana A thesis submitted to the Miami University Honors Program in partial fulfillment of the requirements for University Honors by Marc
Isolation and Characterization of Sphingolipids in Arabidopsis thaliana A thesis submitted to the Miami University Honors Program in partial fulfillment of the requirements for University Honors by Marc
Tsc10p and FVT1: topologically distinct short-chain reductases required for long-chain base synthesis in yeast and mammals
 Tsc10p and FVT1: topologically distinct short-chain reductases required for long-chain base synthesis in yeast and mammals Sita D. Gupta,* Kenneth Gable,* Gongshe Han,* Anna Borovitskaya, Luke Selby,*
Tsc10p and FVT1: topologically distinct short-chain reductases required for long-chain base synthesis in yeast and mammals Sita D. Gupta,* Kenneth Gable,* Gongshe Han,* Anna Borovitskaya, Luke Selby,*
JBC Papers in Press. Published on January 14, 2002 as Manuscript M
 JBC Papers in Press. Published on January 14, 2002 as Manuscript M109976200 Pig Liver Carnitine Palmitoyltransferase: Chimera Studies Show That Both the N- and C-Terminal Regions of the Enzyme Are Important
JBC Papers in Press. Published on January 14, 2002 as Manuscript M109976200 Pig Liver Carnitine Palmitoyltransferase: Chimera Studies Show That Both the N- and C-Terminal Regions of the Enzyme Are Important
Luminescent platforms for monitoring changes in the solubility of amylin and huntingtin in living cells
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2016 Contents Supporting Information Luminescent platforms for monitoring changes in the
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2016 Contents Supporting Information Luminescent platforms for monitoring changes in the
Producing Wax Esters in Transgenic Plants by Expression of Genes Derived from Jojoba
 Reprinted from: Perspectives on new crops and new uses. 1999. J. Janick (ed.), ASHS Press, Alexandria, VA. Producing Wax Esters in Transgenic Plants by Expression of Genes Derived from Jojoba Michael W.
Reprinted from: Perspectives on new crops and new uses. 1999. J. Janick (ed.), ASHS Press, Alexandria, VA. Producing Wax Esters in Transgenic Plants by Expression of Genes Derived from Jojoba Michael W.
MCB130 Midterm. GSI s Name:
 1. Peroxisomes are small, membrane-enclosed organelles that function in the degradation of fatty acids and in the degradation of H 2 O 2. Peroxisomes are not part of the secretory pathway and peroxisomal
1. Peroxisomes are small, membrane-enclosed organelles that function in the degradation of fatty acids and in the degradation of H 2 O 2. Peroxisomes are not part of the secretory pathway and peroxisomal
Phospholipids Metabolism
 Chapter VI: Phospholipids Metabolism Dr. Sameh Sarray Hlaoui Phospholipids Features: Amphipatic: - Hydrophobic head: fatty acids - Hydropholic head: P group+ alcohol Composed of alcohol attached by a phosphodiester
Chapter VI: Phospholipids Metabolism Dr. Sameh Sarray Hlaoui Phospholipids Features: Amphipatic: - Hydrophobic head: fatty acids - Hydropholic head: P group+ alcohol Composed of alcohol attached by a phosphodiester
Biotage Microwave Symposium
 Biotage Microwave Symposium 3-Aryl-4-hydroxyquinolin-2(1)-one Derivatives as Type I Fatty Acid Synthase Inhibitors: Development of a Solvent-Free Microwave Synthesis for Rapid SAR Studies Alexey Rivkin,
Biotage Microwave Symposium 3-Aryl-4-hydroxyquinolin-2(1)-one Derivatives as Type I Fatty Acid Synthase Inhibitors: Development of a Solvent-Free Microwave Synthesis for Rapid SAR Studies Alexey Rivkin,
Mitochondrial Trifunctional Protein (TFP) Protein Quantity Microplate Assay Kit
 PROTOCOL Mitochondrial Trifunctional Protein (TFP) Protein Quantity Microplate Assay Kit DESCRIPTION Mitochondrial Trifunctional Protein (TFP) Protein Quantity Microplate Assay Kit Sufficient materials
PROTOCOL Mitochondrial Trifunctional Protein (TFP) Protein Quantity Microplate Assay Kit DESCRIPTION Mitochondrial Trifunctional Protein (TFP) Protein Quantity Microplate Assay Kit Sufficient materials
Zool 3200: Cell Biology Exam 4 Part II 2/3/15
 Name:Key Trask Zool 3200: Cell Biology Exam 4 Part II 2/3/15 Answer each of the following questions in the space provided, explaining your answers when asked to do so; circle the correct answer or answers
Name:Key Trask Zool 3200: Cell Biology Exam 4 Part II 2/3/15 Answer each of the following questions in the space provided, explaining your answers when asked to do so; circle the correct answer or answers
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Stelter et al., http://www.jcb.org/cgi/content/full/jcb.201105042/dc1 S1 Figure S1. Dyn2 recruitment to edid-labeled FG domain Nups.
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Stelter et al., http://www.jcb.org/cgi/content/full/jcb.201105042/dc1 S1 Figure S1. Dyn2 recruitment to edid-labeled FG domain Nups.
Chromatin IP (Isw2) Fix soln: 11% formaldehyde, 0.1 M NaCl, 1 mm EDTA, 50 mm Hepes-KOH ph 7.6. Freshly prepared. Do not store in glass bottles.
 Chromatin IP (Isw2) 7/01 Toshi last update: 06/15 Reagents Fix soln: 11% formaldehyde, 0.1 M NaCl, 1 mm EDTA, 50 mm Hepes-KOH ph 7.6. Freshly prepared. Do not store in glass bottles. 2.5 M glycine. TBS:
Chromatin IP (Isw2) 7/01 Toshi last update: 06/15 Reagents Fix soln: 11% formaldehyde, 0.1 M NaCl, 1 mm EDTA, 50 mm Hepes-KOH ph 7.6. Freshly prepared. Do not store in glass bottles. 2.5 M glycine. TBS:
Case 19 Purification of Rat Kidney Sphingosine Kinase
 Case 19 Purification of Rat Kidney Sphingosine Kinase Focus concept The purification and kinetic analysis of an enzyme that produces a product important in cell survival is the focus of this study. Prerequisites
Case 19 Purification of Rat Kidney Sphingosine Kinase Focus concept The purification and kinetic analysis of an enzyme that produces a product important in cell survival is the focus of this study. Prerequisites
Fatty acid breakdown
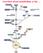 Fatty acids contain a long hydrocarbon chain and a terminal carboxylate group. Most contain between 14 and 24 carbon atoms. The chains may be saturated or contain double bonds. The complete oxidation of
Fatty acids contain a long hydrocarbon chain and a terminal carboxylate group. Most contain between 14 and 24 carbon atoms. The chains may be saturated or contain double bonds. The complete oxidation of
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
LS1a Fall 2014 Problem Set #4 Due Monday 11/3 at 6 pm in the drop boxes on the Science Center 2 nd Floor
 LS1a Fall 2014 Problem Set #4 Due Monday 11/3 at 6 pm in the drop boxes on the Science Center 2 nd Floor Note: Adequate space is given for each answer. Questions that require a brief explanation should
LS1a Fall 2014 Problem Set #4 Due Monday 11/3 at 6 pm in the drop boxes on the Science Center 2 nd Floor Note: Adequate space is given for each answer. Questions that require a brief explanation should
Chapter 26 Biochemistry 5th edition. phospholipids. Sphingolipids. Cholesterol. db=books&itool=toolbar
 http://www.ncbi.nlm.nih.gov/sites/entrez? db=books&itool=toolbar 1 The surface of a soap bubble is a bilayer formed by detergent molecules 2 Chapter 26 Biochemistry 5th edition phospholipids Sphingolipids
http://www.ncbi.nlm.nih.gov/sites/entrez? db=books&itool=toolbar 1 The surface of a soap bubble is a bilayer formed by detergent molecules 2 Chapter 26 Biochemistry 5th edition phospholipids Sphingolipids
Mammalian Membrane Protein Extraction Kit
 Mammalian Membrane Protein Extraction Kit Catalog number: AR0155 Boster s Mammalian Membrane Protein Extraction Kit is a simple, rapid and reproducible method to prepare cellular protein fractions highly
Mammalian Membrane Protein Extraction Kit Catalog number: AR0155 Boster s Mammalian Membrane Protein Extraction Kit is a simple, rapid and reproducible method to prepare cellular protein fractions highly
Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene
 Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene YUELIN ZHANG, WEIHUA FAN, MARK KINKEMA, XIN LI, AND
Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene YUELIN ZHANG, WEIHUA FAN, MARK KINKEMA, XIN LI, AND
Communication. Identification of Methionine N -Acetyltransferase from Saccharomyces cerevisiae
 Communication THE JOURNAL OP BIOLOGICAL CHEMISTRY Vol. 265, No. 7, Issue of March 5, pp. 3603-3606,lSSO 0 1990 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U. S. A. Identification
Communication THE JOURNAL OP BIOLOGICAL CHEMISTRY Vol. 265, No. 7, Issue of March 5, pp. 3603-3606,lSSO 0 1990 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U. S. A. Identification
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Synthesis and degradation of fatty acids Martina Srbová
 Synthesis and degradation of fatty acids Martina Srbová martina.srbova@lfmotol.cuni.cz Fatty acids (FA) mostly an even number of carbon atoms and linear chain in esterified form as component of lipids
Synthesis and degradation of fatty acids Martina Srbová martina.srbova@lfmotol.cuni.cz Fatty acids (FA) mostly an even number of carbon atoms and linear chain in esterified form as component of lipids
Synthesis of Fatty Acids and Triacylglycerol
 Fatty Acid Synthesis Synthesis of Fatty Acids and Triacylglycerol Requires Carbon Source: Reducing Power: NADPH Energy Input: ATP Why Energy? Why Energy? Fatty Acid Fatty Acid + n(atp) ΔG o : -ve Fatty
Fatty Acid Synthesis Synthesis of Fatty Acids and Triacylglycerol Requires Carbon Source: Reducing Power: NADPH Energy Input: ATP Why Energy? Why Energy? Fatty Acid Fatty Acid + n(atp) ΔG o : -ve Fatty
Biosynthesis and secretion of plant cuticular wax
 Progress in Lipid Research 42 (2003) 51 80 www.elsevier.com/locate/plipres Review Biosynthesis and secretion of plant cuticular wax L. Kunst, A.L. Samuels* Department of Botany, UBC, 6270 University Boulevard,
Progress in Lipid Research 42 (2003) 51 80 www.elsevier.com/locate/plipres Review Biosynthesis and secretion of plant cuticular wax L. Kunst, A.L. Samuels* Department of Botany, UBC, 6270 University Boulevard,
Acetyl-CoA or BC acyl-coa. Int1-ACP. FabH. FabF. Anti-FabF Platensimycin ACP. Fak
 Initiation Acetyl-CoA AccBCDA FabZ Int-ACP Elongation [N+] FabG Int1-ACP Acetyl-CoA or BC acyl-coa FabH FabD CoA Malonyl-ACP Malonyl CoA ACP Anti-FabI Triclosan AFN-15 FabI acyl-acp PlsX PlsY Pls C Lipoic
Initiation Acetyl-CoA AccBCDA FabZ Int-ACP Elongation [N+] FabG Int1-ACP Acetyl-CoA or BC acyl-coa FabH FabD CoA Malonyl-ACP Malonyl CoA ACP Anti-FabI Triclosan AFN-15 FabI acyl-acp PlsX PlsY Pls C Lipoic
MILK BIOSYNTHESIS PART 3: FAT
 MILK BIOSYNTHESIS PART 3: FAT KEY ENZYMES (FROM ALL BIOSYNTHESIS LECTURES) FDPase = fructose diphosphatase Citrate lyase Isocitrate dehydrogenase Fatty acid synthetase Acetyl CoA carboxylase Fatty acyl
MILK BIOSYNTHESIS PART 3: FAT KEY ENZYMES (FROM ALL BIOSYNTHESIS LECTURES) FDPase = fructose diphosphatase Citrate lyase Isocitrate dehydrogenase Fatty acid synthetase Acetyl CoA carboxylase Fatty acyl
Biochemical characterization of the very long-chain fatty acid elongase ELOVL7
 FEBS Letters 585 (2011) 3337 3341 journal homepage: www.febsletters.org Biochemical characterization of the very long-chain fatty acid elongase ELOVL7 Tatsuro Naganuma 1, Yuichiro Sato 1, Takayuki Sassa,
FEBS Letters 585 (2011) 3337 3341 journal homepage: www.febsletters.org Biochemical characterization of the very long-chain fatty acid elongase ELOVL7 Tatsuro Naganuma 1, Yuichiro Sato 1, Takayuki Sassa,
Synthesis of Fatty Acids and Triacylglycerol
 Synthesis of Fatty Acids and Triacylglycerol Lippincott s Chapter 16 Fatty Acid Synthesis Mainly in the Liver Requires Carbon Source: Acetyl CoA Reducing Power: NADPH 8 CH 3 COO C 15 H 33 COO Energy Input:
Synthesis of Fatty Acids and Triacylglycerol Lippincott s Chapter 16 Fatty Acid Synthesis Mainly in the Liver Requires Carbon Source: Acetyl CoA Reducing Power: NADPH 8 CH 3 COO C 15 H 33 COO Energy Input:
Materials and Methods , The two-hybrid principle.
 The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
CA Tel.: ; Fax: ; ucsd.edu.
 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 283, NO. 10, pp. 6300 6311, March 7, 2008 2008 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. The Phosphatase PHLPP
THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 283, NO. 10, pp. 6300 6311, March 7, 2008 2008 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. The Phosphatase PHLPP
A Molecular Caliper Mechanism for Determining Very Long-Chain Fatty Acid Length
 A Molecular Caliper Mechanism for Determining Very Long-Chain Fatty Acid Length Vladimir Denic 1 and Jonathan S. Weissman 1, * 1 Howard Hughes Medical Institute, Department of Cellular and Molecular Pharmacology,
A Molecular Caliper Mechanism for Determining Very Long-Chain Fatty Acid Length Vladimir Denic 1 and Jonathan S. Weissman 1, * 1 Howard Hughes Medical Institute, Department of Cellular and Molecular Pharmacology,
ab E3 Ligase Auto- Ubiquitilylation Assay Kit
 ab139469 E3 Ligase Auto- Ubiquitilylation Assay Kit Instructions for Use For testing ubiquitin E3 ligase activity through assessment of their ability to undergo auto-ubiquitinylation This product is for
ab139469 E3 Ligase Auto- Ubiquitilylation Assay Kit Instructions for Use For testing ubiquitin E3 ligase activity through assessment of their ability to undergo auto-ubiquitinylation This product is for
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation
 Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
The Role of Lipids in Flowering Development of Arabidopsis Enhanced pah1pah2 Plants. Toshiro Ito 1 & Lee Lishi 2
 The Role of Lipids in Flowering Development of Arabidopsis Enhanced pah1pah2 Plants Toshiro Ito 1 & Lee Lishi 2 Department of Biological Sciences, Faculty of Science, National University of Singapore,
The Role of Lipids in Flowering Development of Arabidopsis Enhanced pah1pah2 Plants Toshiro Ito 1 & Lee Lishi 2 Department of Biological Sciences, Faculty of Science, National University of Singapore,
Small Subunits of Serine Palmitoyltransferase (ssspts) and their. physiological roles
 Small Subunits of Serine Palmitoyltransferase (ssspts) and their physiological roles by Saurav Majumder Thesis submitted to the Faculty of the Molecular and Cell biology Graduate Program Uniformed Services
Small Subunits of Serine Palmitoyltransferase (ssspts) and their physiological roles by Saurav Majumder Thesis submitted to the Faculty of the Molecular and Cell biology Graduate Program Uniformed Services
Instructions for Use. APO-AB Annexin V-Biotin Apoptosis Detection Kit 100 tests
 3URGXFW,QIRUPDWLRQ Sigma TACS Annexin V Apoptosis Detection Kits Instructions for Use APO-AB Annexin V-Biotin Apoptosis Detection Kit 100 tests For Research Use Only. Not for use in diagnostic procedures.
3URGXFW,QIRUPDWLRQ Sigma TACS Annexin V Apoptosis Detection Kits Instructions for Use APO-AB Annexin V-Biotin Apoptosis Detection Kit 100 tests For Research Use Only. Not for use in diagnostic procedures.
Biochemistry: A Short Course
 Tymoczko Berg Stryer Biochemistry: A Short Course Second Edition CHAPTER 28 Fatty Acid Synthesis 2013 W. H. Freeman and Company Chapter 28 Outline 1. The first stage of fatty acid synthesis is transfer
Tymoczko Berg Stryer Biochemistry: A Short Course Second Edition CHAPTER 28 Fatty Acid Synthesis 2013 W. H. Freeman and Company Chapter 28 Outline 1. The first stage of fatty acid synthesis is transfer
Sphingosine-1-phosphate signaling and cardiac fibrosis. Department of Physiology, Kanazawa University School of Medicine, Kanazawa, Japan
 96 Special Issue: Cellular and Molecular Bases for Fibrotic Diseases Review Article Sphingosine-1-phosphate signaling and cardiac fibrosis Noriko Takuwa 1, 2, ), Yasuo Okamoto 1), Kazuaki Yoshioka 1) and
96 Special Issue: Cellular and Molecular Bases for Fibrotic Diseases Review Article Sphingosine-1-phosphate signaling and cardiac fibrosis Noriko Takuwa 1, 2, ), Yasuo Okamoto 1), Kazuaki Yoshioka 1) and
Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.
 Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.129-Gt(ROSA)26Sor tm1(cre/ert2)tyj /J mice. To induce deletion of the Pten locus,
Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.129-Gt(ROSA)26Sor tm1(cre/ert2)tyj /J mice. To induce deletion of the Pten locus,
Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION
 Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION X. Shawn Liu 1, 3, Bing Song 2, 3, Bennett D. Elzey 3, 4, Timothy L. Ratliff 3, 4, Stephen F. Konieczny
Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION X. Shawn Liu 1, 3, Bing Song 2, 3, Bennett D. Elzey 3, 4, Timothy L. Ratliff 3, 4, Stephen F. Konieczny
ab SREBP-2 Translocation Assay Kit (Cell-Based)
 ab133114 SREBP-2 Translocation Assay Kit (Cell-Based) Instructions for Use For analysis of translocation of SREBP-2 into nuclei. This product is for research use only and is not intended for diagnostic
ab133114 SREBP-2 Translocation Assay Kit (Cell-Based) Instructions for Use For analysis of translocation of SREBP-2 into nuclei. This product is for research use only and is not intended for diagnostic
Growth of the tps1 strain and tps1 suppressor strains in rich liquid medium with
 OD 6 YP Galactose.8.6.4..8.6 5 5 5 3 35 4 45 5 Time (h) tps tps ras tps + ppde OD 6 YP Glucose.8.6.4..8.6 5 5 5 3 35 4 45 5 Time (h) Supplementary Fig. Growth of the tps strain and tps suppressor strains
OD 6 YP Galactose.8.6.4..8.6 5 5 5 3 35 4 45 5 Time (h) tps tps ras tps + ppde OD 6 YP Glucose.8.6.4..8.6 5 5 5 3 35 4 45 5 Time (h) Supplementary Fig. Growth of the tps strain and tps suppressor strains
Student Number: To form the polar phase when adsorption chromatography was used.
 Name: Student Number: April 14, 2001, 1:30 AM - 4:30 PM Page 1 (of 4) Biochemistry II Lab Section Final Examination Examiner: Dr. A. Scoot 1. Answer ALL questions in the space provided.. 2. The last page
Name: Student Number: April 14, 2001, 1:30 AM - 4:30 PM Page 1 (of 4) Biochemistry II Lab Section Final Examination Examiner: Dr. A. Scoot 1. Answer ALL questions in the space provided.. 2. The last page
Test Bank for Lehninger Principles of Biochemistry 5th Edition by Nelson
 Test Bank for Lehninger Principles of Biochemistry 5th Edition by Nelson Link download full: http://testbankair.com/download/test-bank-forlehninger-principles-of-biochemistry-5th-edition-by-nelson/ Chapter
Test Bank for Lehninger Principles of Biochemistry 5th Edition by Nelson Link download full: http://testbankair.com/download/test-bank-forlehninger-principles-of-biochemistry-5th-edition-by-nelson/ Chapter
Fatty Acid Methylation Kits
 Methyl esterification kit for fatty acids analysis Fatty Acid Methylation Kits Below are two methods for efficiently preparing fatty acid samples for GC analysis. Neither method requires high temperatures,
Methyl esterification kit for fatty acids analysis Fatty Acid Methylation Kits Below are two methods for efficiently preparing fatty acid samples for GC analysis. Neither method requires high temperatures,
In Vivo Action of the HRD Ubiquitin Ligase Complex: Mechanisms of Endoplasmic Reticulum Quality Control and Sterol Regulation
 MOLECULAR AND CELLULAR BIOLOGY, July 2001, p. 4276 4291 Vol. 21, No. 13 0270-7306/01/$04.00 0 DOI: 10.1128/MCB.21.13.4276 4291.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved.
MOLECULAR AND CELLULAR BIOLOGY, July 2001, p. 4276 4291 Vol. 21, No. 13 0270-7306/01/$04.00 0 DOI: 10.1128/MCB.21.13.4276 4291.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved.
MITOCW watch?v=ddt1kusdoog
 MITOCW watch?v=ddt1kusdoog The following content is provided under a Creative Commons license. Your support will help MIT OpenCourseWare continue to offer high-quality educational resources for free. To
MITOCW watch?v=ddt1kusdoog The following content is provided under a Creative Commons license. Your support will help MIT OpenCourseWare continue to offer high-quality educational resources for free. To
TSH Receptor Monoclonal Antibody (49) Catalog Number MA3-218 Product data sheet
 Website: thermofisher.com Customer Service (US): 1 800 955 6288 ext. 1 Technical Support (US): 1 800 955 6288 ext. 441 TSH Receptor Monoclonal Antibody (49) Catalog Number MA3-218 Product data sheet Details
Website: thermofisher.com Customer Service (US): 1 800 955 6288 ext. 1 Technical Support (US): 1 800 955 6288 ext. 441 TSH Receptor Monoclonal Antibody (49) Catalog Number MA3-218 Product data sheet Details
Chapter 3. Expression of α5-megfp in Mouse Cortical Neurons. on the β subunit. Signal sequences in the M3-M4 loop of β nachrs bind protein factors to
 22 Chapter 3 Expression of α5-megfp in Mouse Cortical Neurons Subcellular localization of the neuronal nachr subtypes α4β2 and α4β4 depends on the β subunit. Signal sequences in the M3-M4 loop of β nachrs
22 Chapter 3 Expression of α5-megfp in Mouse Cortical Neurons Subcellular localization of the neuronal nachr subtypes α4β2 and α4β4 depends on the β subunit. Signal sequences in the M3-M4 loop of β nachrs
Expression of acid base transporters in the kidney collecting duct in Slc2a7 -/-
 Supplemental Material Results. Expression of acid base transporters in the kidney collecting duct in Slc2a7 -/- and Slc2a7 -/- mice. The expression of AE1 in the kidney was examined in Slc26a7 KO mice.
Supplemental Material Results. Expression of acid base transporters in the kidney collecting duct in Slc2a7 -/- and Slc2a7 -/- mice. The expression of AE1 in the kidney was examined in Slc26a7 KO mice.
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/283/ra57/dc1 Supplementary Materials for JNK3 Couples the Neuronal Stress Response to Inhibition of Secretory Trafficking Guang Yang,* Xun Zhou, Jingyan Zhu,
www.sciencesignaling.org/cgi/content/full/6/283/ra57/dc1 Supplementary Materials for JNK3 Couples the Neuronal Stress Response to Inhibition of Secretory Trafficking Guang Yang,* Xun Zhou, Jingyan Zhu,
Supplementary data Supplementary Figure 1 Supplementary Figure 2
 Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supporting Information
 Supporting Information Valkenburg et al. 10.1073/pnas.1403684111 SI Materials and Methods ELISA and Microneutralization. Sera were treated with Receptor Destroying Enzyme II (RDE II, Accurate) before ELISA
Supporting Information Valkenburg et al. 10.1073/pnas.1403684111 SI Materials and Methods ELISA and Microneutralization. Sera were treated with Receptor Destroying Enzyme II (RDE II, Accurate) before ELISA
(Stratagene, La Jolla, CA) (Supplemental Fig. 1A). A 5.4-kb EcoRI fragment
 SUPPLEMENTAL INFORMATION Supplemental Methods Generation of RyR2-S2808D Mice Murine genomic RyR2 clones were isolated from a 129/SvEvTacfBR λ-phage library (Stratagene, La Jolla, CA) (Supplemental Fig.
SUPPLEMENTAL INFORMATION Supplemental Methods Generation of RyR2-S2808D Mice Murine genomic RyR2 clones were isolated from a 129/SvEvTacfBR λ-phage library (Stratagene, La Jolla, CA) (Supplemental Fig.
Mannosylinositol phosphorylceramides and ergosterol coordinately maintain cell wall integrity in the yeast Saccharomyces cerevisiae
 EDITOR S CHOICE Mannosylinositol phosphorylceramides and ergosterol coordinately maintain cell wall integrity in the yeast Saccharomyces cerevisiae Seiya Tanaka and Motohiro Tani Department of Chemistry,
EDITOR S CHOICE Mannosylinositol phosphorylceramides and ergosterol coordinately maintain cell wall integrity in the yeast Saccharomyces cerevisiae Seiya Tanaka and Motohiro Tani Department of Chemistry,
Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in
 Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in Supplementary Fig. 2 Substitution Sequence Position variant Sequence original APNCYGNIPL original APNCYGNIPL
Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in Supplementary Fig. 2 Substitution Sequence Position variant Sequence original APNCYGNIPL original APNCYGNIPL
Lipids and Classification:
 Lipids and Classification: Lipids: Biological lipids are a chemically diverse group of organic compounds which are insoluble or only poorly soluble in water. They are readily soluble in non-polar solvents
Lipids and Classification: Lipids: Biological lipids are a chemically diverse group of organic compounds which are insoluble or only poorly soluble in water. They are readily soluble in non-polar solvents
Problem Set #5 4/3/ Spring 02
 Question 1 Chloroplasts contain six compartments outer membrane, intermembrane space, inner membrane, stroma, thylakoid membrane, and thylakoid lumen each of which is populated by specific sets of proteins.
Question 1 Chloroplasts contain six compartments outer membrane, intermembrane space, inner membrane, stroma, thylakoid membrane, and thylakoid lumen each of which is populated by specific sets of proteins.
1. endoplasmic reticulum This is the location where N-linked oligosaccharide is initially synthesized and attached to glycoproteins.
 Biology 4410 Name Spring 2006 Exam 2 A. Multiple Choice, 2 pt each Pick the best choice from the list of choices, and write it in the space provided. Some choices may be used more than once, and other
Biology 4410 Name Spring 2006 Exam 2 A. Multiple Choice, 2 pt each Pick the best choice from the list of choices, and write it in the space provided. Some choices may be used more than once, and other
Formation of the Peroxisome Lumen Is Abolished by Loss of Pichia pastoris Pas7p, a Zinc-Binding Integral Membrane Protein of the Peroxisome
 MOLECULAR AND CELLULAR BIOLOGY, Nov. 1995, p. 6406 6419 Vol. 15, No. 11 0270-7306/95/$04.00 0 Copyright 1995, American Society for Microbiology Formation of the Peroxisome Lumen Is Abolished by Loss of
MOLECULAR AND CELLULAR BIOLOGY, Nov. 1995, p. 6406 6419 Vol. 15, No. 11 0270-7306/95/$04.00 0 Copyright 1995, American Society for Microbiology Formation of the Peroxisome Lumen Is Abolished by Loss of
Energy storage in cells
 Energy storage in cells Josef Fontana EC - 58 Overview of the lecture Introduction to the storage substances of human body Overview of storage compounds in the body Glycogen metabolism Structure of glycogen
Energy storage in cells Josef Fontana EC - 58 Overview of the lecture Introduction to the storage substances of human body Overview of storage compounds in the body Glycogen metabolism Structure of glycogen
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2
 Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
The rabbit femoral artery was prepared and each arterial ring was permeabilized
 Online Supplement Nakmura et al. cgmp-dependent relaxation of smooth muscle Materials and Methods Measurement of tension The rabbit femoral artery was prepared and each arterial ring was permeabilized
Online Supplement Nakmura et al. cgmp-dependent relaxation of smooth muscle Materials and Methods Measurement of tension The rabbit femoral artery was prepared and each arterial ring was permeabilized
Distribution of molecular species of sphingomyelins in different parts of bovine digestive tract
 Distribution of molecular species of sphingomyelins in different parts of bovine digestive tract M. E. Breimer Membrane Biochemistry Group, Department of Medical Biochemistry, University of Giiteborg,
Distribution of molecular species of sphingomyelins in different parts of bovine digestive tract M. E. Breimer Membrane Biochemistry Group, Department of Medical Biochemistry, University of Giiteborg,
ANSC (NUTR) 618 LIPIDS & LIPID METABOLISM Membrane Lipids and Sphingolipidsd
 ANSC (NUTR) 618 LIPIDS & LIPID METABOLISM Membrane Lipids and Sphingolipidsd I. Classes of membrane lipids A. Glycerolipids (quantitatively the most important of the three membrane lipids) B. Shingolipids
ANSC (NUTR) 618 LIPIDS & LIPID METABOLISM Membrane Lipids and Sphingolipidsd I. Classes of membrane lipids A. Glycerolipids (quantitatively the most important of the three membrane lipids) B. Shingolipids
PRODUCT INFORMATION & MANUAL
 PRODUCT INFORMATION & MANUAL 0.4 micron for Overall Exosome Isolation (Cell Media) NBP2-49826 For research use only. Not for diagnostic or therapeutic procedures. www.novusbio.com - P: 303.730.1950 - P:
PRODUCT INFORMATION & MANUAL 0.4 micron for Overall Exosome Isolation (Cell Media) NBP2-49826 For research use only. Not for diagnostic or therapeutic procedures. www.novusbio.com - P: 303.730.1950 - P:
TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer
 Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
Bio 401 Sp2014. Multiple Choice (Circle the letter corresponding to the correct answer)
 MIDTERM EXAM KEY (60 pts) You should have 5 pages. Please put your name on every page. You will have 70 minutes to complete the exam. You may begin working as soon as you receive the exam. NOTE: the RED
MIDTERM EXAM KEY (60 pts) You should have 5 pages. Please put your name on every page. You will have 70 minutes to complete the exam. You may begin working as soon as you receive the exam. NOTE: the RED
BIOCATALYTIC CONVERSION OF OTHER LIPIDS
 Chapter 6 BIOCATALYTIC CONVERSION OF OTHER LIPIDS In Chapters 4 and 5, lipase-mediated conversion of acylglycerols were presented. The Chapter 6 deals with biocatalytic modification of phospholipids, sphingolipids,
Chapter 6 BIOCATALYTIC CONVERSION OF OTHER LIPIDS In Chapters 4 and 5, lipase-mediated conversion of acylglycerols were presented. The Chapter 6 deals with biocatalytic modification of phospholipids, sphingolipids,
Supplementary Figure 1. Chemical structures of activity-based probes (ABPs) and of click reagents used in this study.
 Supplementary Figure 1. Chemical structures of activity-based probes (ABPs) and of click reagents used in this study. In this study, one fluorophosphonate (FP, 1), three nitrophenol phosphonate probes
Supplementary Figure 1. Chemical structures of activity-based probes (ABPs) and of click reagents used in this study. In this study, one fluorophosphonate (FP, 1), three nitrophenol phosphonate probes
