Supplementary Figure 1. Crystallography of PI4KIIα with ADP bound. a, A
|
|
|
- Gilbert Evans
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1. Crystallography of PI4KIIα with ADP bound. a, A stereo image of overall 2Fo-Fc electron density map of PI4KIIα contoured at 1.0 σ. b, 2FoFc electron density map (1.0 σ) around the palmitoylation insertion, colored in green. 1
2 Supplementary Figure 2. Analysis of the crystal packing of PI4KIIα. a, Superposition of the two PI4KIIα molecules (A, yellow; B, pink) in one asymmetric unit. The significantly different conformations at the P-loop region and N-terminal helix α1 are indicated and labeled. b, Overall crystal packing of PI4KIIα viewed along the b axis (left) and c axis (right). The unit cell is depicted and labeled. The two molecules of PI4KIIα in one asymmetric unit are colored in yellow, molecule A, and pink, molecule B. Octamers (A 8 and B 8 ) formed by each type of molecule are 2
3 indicated by respective rectangles. c and d, Overall structures of the octamers A 8 and B 8 are viewed from along (left) and perpendicular to (right) the membrane binding surface of PI4KIIα. The bound ADP in each molecule is shown as a colored sphere. Color schemes for the rectangles are the same as in b. 3
4 4
5 Supplementary Figure 3. Comparisons of the structures of PI4Ks and PI3Ks. a, Known crystal structures of class I PI3K (p110 α, β, γ and δ), class III PI3K (vps34) and PIP4KII β are shown as cartoon diagrams with the structure of PI4KIIα catalytic domain (colored in golden) superimposed, respectively. The views take the same angle as the crystal structure of PI4KIIα in Fig. 1c. The N- and C-lobes of the catalytic domain are colored in blue and green, respectively. The catalytic loop and activation loop (if present) are colored in purple and red, respectively. The corresponding PDB code is shown below each structure. b, Structure-based sequence alignment of the catalytic domains of PI4KIIα and PI3Ks. Structural superposition and the subsequent structure-based sequence alignment were performed using UCSF Chimera 1. Secondary structures of PI4KIIα are shown and labeled above the sequences, with blue for the N-lobe and green for the C-lobe; dashed lines represent untraced regions of the PI4KIIα structure. Identical residues in the alignment are colored in black and similar residues in grey. Residues that contribute to nucleotide binding in PI4KIIα are marked as solid circles with residues interacting with the phosphate moiety in red( ) and residues interacting with adenosinein cyan( ). GenBank accession numbers for the amino acid sequences of these proteins are Q9BTU6 for hpi4kiiα, P42336 for hpi3kα (p110α), P42338 for hpi3kβ (p110β), O00329 for hpi3kδ (p110δ), P48736 for hpi3kγ (p110γ) and Q9W1M7 for dvps34. The figure was prepared using Boxshade3.21 ( 5
6 Supplementary Figure 4. Sequence alignment of PI4KIIs around the G-loop region. Ser134 and Ser137 residues are indicated with red triangles. The GenBank accession numbers of the PI4KIIα sequences from different species are NP_ for Homo sapiens, NP_ for Daniorerio, ACN11482 for Salmosalar, XP_ for Susscrofa, NP_ for Bos Taurus, XP_ for Oryctolaguscuniculus, NP_ for Musmusculus, NP_ for Rattusnorvegicus, NP_ for Xenopuslaevis and XP_ for Trichinellaspiralis. The GenBank accession numbers of the PI4KII β sequences from different species are NP_ for Homo sapiens, NP_ for Bos Taurus, XP_ for Oryctolaguscuniculus, NP_ for Rattusnorvegicus, NP_ for Musmusculus, XP_ for Taeniopygiaguttata, NP_ for Gallus gallus, NP_ for Xenopuslaevis, NP_ for Daniorerio, EFN64759 for Camponotusfloridanus and XP_ for Trichinellaspiralis. The alignment was performed using ClustalW and the figure was prepared with Boxshade3.21 ( 6
7 Supplementary Figure 5. Blue shifts in the internal fluorescence of Trp residues validates the membrane binding surface of PI4KIIα. In addition to the untraced Trp332 on insertion I2 and three non-exposed residues, Trp273, Trp314 and Trp366, the remaining four Trp residues (Trp359, Trp368, Trp166, Trp169) are exposed at the putative membrane-binding surface of PI4KIIα. Internal tryptophan emission fluorescences of PI4KIIα variants (PI4KIIα SSPSSΔC and PI4KIIα FFPFFΔC ), which were in solution or reconstituted into liposomes, were measured at different protein concentrations, using the excitation wavelength at 295 nm. 7
8 Supplementary Figure 6. Kinase activity of PI4KIIα variants. a, Kinase activity of PI4KIIα SSPSSΔC increases when Triton X-100 is replaced by a zwitterionic detergent, dimethyl(3-sulfopropyl) tetradecyl-ammonium hydroxide (Anzergent3-14, AZ3-14, Anatrace ) since the negatively-charged head of AZ3-14 can enhance electrostatic interactions between PI4KIIα and the surface of the detergent micelle. b, Comparison of the kinase activities of PI4KIIα variants measured at conditions PI/0.2% TX-100 and PI/0.1% AZ3-14. Error bars represent the standard deviation (s.d.) from three independent experiments. 8
9 Supplementary Figure 7. Structural characterization of PI4KIIα in molecular dynamics simulations. a, Time evolution of RMSD of PI4KIIa relative to the crystal structure, with #1, #2 and #3 representing simulations of non-palmitoylated proteins, #4, #5 and #6 simulations of palmitoylated ones and #7 a simulation of membrane-free non-palmitoylated PI4KIIa. In simulations #1 and #2, the crystal structure of PI4KIIα was placed at t = 0 ns on top of the PI-containing membrane at a distance of 20 Å while in simulation #3 the distance was 10 Å. The average RMSD of non-palmitoylated and palmitoylated proteins relative to the crystal structure over the last 700 ns of 1-μs simulation were 3.59 Å and 2.64 Å, respectively. b, Time evolution of the relative height of the protein's center of mass with respect to upper lipid phosphate atoms' center of mass in different simulations. The choice of color is the same as in (a). c, Time evolution of the insertion depth of PI4KIIα's palmitoylation insertion (residue 166 to 179) center of 9
10 mass with respect to upper lipid phosphate atoms' center of mass. Negative values correspond to insertion of the loop into the membrane. The choice of color is the same as in (a). d, Conformational fluctuations of non-palmitoylated PI4KIIα in the absence of interactions with a membrane. The black curve represents the membrane-free simulation of PI4KIIα; blue and dark-green curves were obtained from the first 100 ns of non-palmitoylated PI4KIIα simulated in the presence of a membrane (#1 and #2 in a, b and c). The protein did not interact with the membrane during the first 100 ns of simulations #1 and #2., The RMSF was calculated using C α -atoms of each protein residue (residue 107 to 453) through the indicated simulation time. For this purpose, the proteins were aligned to the crystal structure using backbone atoms without three insertions. 10
11 Supplementary Figure 8. Enzymatic assays of PI4KIIα variants. a. Enzymatic V max and K m of PI4KIIα variants. Lipid kinase activity of purified PI4KIIα FFPFF and PI4KIIα CCPCC (0.5 μg) was determined with the ADP-Glo TM kinase kit (Promega). Effects of palmitoylation motif mutation on the kinetics of PI4KIIα catalysis were measured. V max and K m values for PI were calculated based on dose-dependent curves. Values are presented as means ± S.D. from three independent experiments (student's t-test). b, Lipid kinase activity of PI4KIIα variants on liposome with or without cholesterol. Proteoliposome of PI4KIIα variants were made according to METHODS with the lipid composition, 20% DOPE, 10% DOPS, 11
12 10% DOPA, 10% PI and 50% DOPC for the cholesterol-free system and 20% DOPE, 10% DOPS, 10% DOPA, 10% PI, 40% DOPC and 10% cholesterol for the cholesterol-containing system. The kinase reaction was initiated by adding 0.1 mm ATP and the kinase activity was plotted by monitoring ADP generation with the ADP-Glo TM kinase kit (Promega). The error bars represent means ± S.D. from three independent experiments. The binding efficiencies of PI4KIIα variants to the liposome membrane were evaluated from the co-sedimentation experiments below. W: the whole amount of the proteoliposome; S, the supernatant after centrifuge of the proteoliposome; P, the precipitate after centrifuging the proteoliposome. Uncropped images of gels are shown in Supplementary Fig
13 13
14 Supplementary Figure 9. Full scans of original Western blots and gels for data in Figure 3 and Supplementary Fig. 8. Panels corresponding to the figures in the paper are indicated. 14
15 Supplementary Table 1. Mutant constructs used in this study Region Name Fragment Mutation Plasmids Expression Control Palmitoylatio n motif and C terminal region ( ) Nucleotide binding site PI binding site Amphipathic α-helix of the palmitoylatio n insertion Membrane binding sites PI4KIIα E N none pfastbac Insect cell (Hi5) PI4KIIα CCPCC none pgex-6p-1 E. coli PI4KIIα SSPSS PI4KIIα SSPSS C PI4KIIα FFPFF PI4KIIα FFPFF C PI4KIIα F263A/I345A PI4KIIα D269A PI4KIIα K152A SSPSS 178 pgex-6p-1 E. coli 174 SSPSS 178 pgex-6p-1 E. coli 174 FFPFF 178 pgex-6p-1 E. coli 174 FFPFF 178 pgex-6p-1 E. coli 174 FFPFF 178 / F263A/I345A 174 FFPFF 178 / D269A 174 FFPFF 178 / K152A pgex-6p-1 pgex-6p-1 pgex-6p-1 E. coli E. coli E. coli PI4KIIα E157A/Y159A E157A/Y159A pgex-6p-1 E. coli PI4KIIα L184A/L349A L184A/L349A pgex-6p-1 E. coli PI4KIIα W166A W166A pgex-6p-1 E. coli PI4KIIα W169A W169A pgex-6p-1 E. coli PI4KIIα L170A L170A pgex-6p-1 E. coli PI4KIIα L173A L173A pgex-6p-1 E. coli PI4KIIα W166A/W169A/ L170A/L173A PI4KIIα K165A/K168A/ K172A PI4KIIα K165E/K168E/ K172E W166A/W169A/ L170A/L173A K165A/K168A/ K172A K165E/K168E/ K172E pgex-6p-1 pgex-6p-1 pgex-6p-1 E. coli E. coli E. coli PI4KIIα W359A W359A pgex-6p-1 E. coli PI4KIIα Y365A Y365A pgex-6p-1 E. coli PI4KIIα W368A W368A pgex-6p-1 E. coli W359A/ Y365A/ PI4KIIα W368A PI4KIIα R129A/R275A/ R276A PI4KIIα R129E/R275E/ R276E W359A/ Y365A/ W368A R129A/R275A/ R276A R129E/R275E/ R276E pgex-6p-1 pgex-6p-1 pgex-6p-1 E. coli E. coli E. coli PI4KIIα R129A R129A pgex-6p-1 E. coli 15
16 Supplementary Note 1 CHARMM36 force field 2-4 and the TIP3P water model 5 were used in all simulations. The simulations were carried out in an NPT ensemble; temperature was maintained at 310 K through a Langevin thermostat with a damping coefficient of 0.5 ps -1 ; pressure was maintained at 1 atm with a Langevin-piston barostat 6. Short-range non-bonded interactions were cut off smoothly between 1 and 1.2 nm; long-range electrostatics was computed with the PME algorithm 7 ; simulations were performed with an integration time step of 2 fs in NAMD For the force field of the palmitoylated cysteine residue, initial parameters were obtained from the amino acid cysteine and from the dioleoylphosphatidylcholine (DOPC) lipid tail, employing the corresponding CHARMM36 force field 4. The full cysteine residue and the first three carbons of the palmitoylation group were selected for further optimization by the force field toolkit (FFTK) 9 plugin of VMD 10. Partial atomic charges were optimized from water-interaction profiles by Gaussian 11 at a HF/6 31G(d) level of theory. Hydrogen atom charges were kept at 0.09e to be consistent with the CHARMM force field 12. Bond and angle parameters were optimized from distortions along internal coordinates by means of a QM Hessian matrix calculation at a MP2/6 31G(d) level of theory. Dihedral terms using torsion scans were optimized at a MP2/6 31G(d) level of theory. Final parameters for palmitoylated cysteine residue are shown below. Topology file: ATOM N NH ATOM HN H 0.31 ATOM CA CT ATOM HA HB1 0.09! HA-CA--CB--SG ATOM 1CB CT ATOM HB1 HA ATOM HB2 HA
17 ATOM 1SG SM ATOM C C 0.51 ATOM O O -0.51! beta4 ATOM C31 CL 0.67 ATOM O32 OBL ATOM C32 CTL ATOM H2X HAL ATOM H2Y HAL ATOM C33 CTL ATOM H3X HAL ATOM H3Y HAL ATOM C34 CTL ATOM H4X HAL ATOM H4Y HAL ATOM C35 CTL ATOM H5X HAL ATOM H5Y HAL ATOM C36 CTL ATOM H6X HAL ATOM H6Y HAL ATOM C37 CTL ATOM H7X HAL ATOM H7Y HAL ATOM C38 CTL ATOM H8X HAL ATOM H8Y HAL ATOM C39 CTL ATOM H9X HAL ATOM H9Y HAL ATOM C310 CTL ATOM H10X HAL ATOM H10Y HAL
18 ATOM C311 CTL ATOM H11X HAL ATOM H11Y HAL ATOM C312 CTL ATOM H12X HAL ATOM H12Y HAL ATOM C313 CTL ATOM H13X HAL ATOM H13Y HAL ATOM C314 CTL ATOM H14X HAL ATOM H14Y HAL ATOM C315 CTL ATOM H15X HAL ATOM H15Y HAL ATOM C316 CTL ATOM H16X HAL ATOM H16Y HAL ATOM H16Z HAL Parameter file: BONDS SM CL ANGLES CT2 SM CL SM CL OBL SM CL CTL CT1 CT2 SM CL CT2 SM CL OBL CT2 SM CL OBL CT2 SM CL CTL
19 CT2 SM CL CTL SM CL CTL2 HAL HA2 CT2 SM CL
20 Supplementary References 1. Pettersen, E.F. et al. UCSF Chimera--a visualization system for exploratory research and analysis. J Comput Chem 25, (2004). 2. Schlenkrich, M., Brickmann, J., MacKerell Jr, A.D. & Karplus, M. An empirical potential energy function for phospholipids: criteria for parameter optimization and applications. in Biological Membranes (Springer, 1996). 3. MacKerell, A.D. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. The Journal of Physical Chemistry B 102, (1998). 4. Klauda, J.B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. The Journal of Physical Chemistry B 114, (2010). 5. Jorgensen, W.L., Chandrasekhar, J., Madura, J.D., Impey, R.W. & Klein, M.L. Comparison of simple potential functions for simulating liquid water. The Journal of Chemical Physics 79, (1983). 6. Martyna, G.J., Tobias, D.J. & Klein, M.L. Constant pressure molecular dynamics algorithms. The Journal of Chemical Physics 101, 4177 (1994). 7. Darden, T., Toukmaji, A. & Pedersen, L. Long-range electrostatic effects in biomolecular simulations. Journal de chimie physique 94, (1997). 8. Phillips, J.C. et al. Scalable molecular dynamics with NAMD. Journal of Computational Chemistry 26, (2005). 9. Mayne, C.G., Saam, J., Schulten, K., Tajkhorshid, E. & Gumbart, J.C. Rapid parameterization of small molecules using the Force Field Toolkit. J Comput Chem 34, (2013). 10. Humphrey, W., Dalke, A. & Schulten, K. VMD: Visual molecular dynamics. Journal of Molecular Graphics 14, (1996). 11. Frisch, M. et al. Gaussian 09, revision A. 02; Gaussian. Inc., Wallingford, CT 270, 271 (2009). 12. Yin, D. & MacKerell, A.D. Combined ab initio/empirical approach for optimization of Lennard Jones parameters. Journal of Computational Chemistry 19, (1998). 20
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
SUPPLEMENTARY INFORMATION. Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity
 SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation
 www.bioinformation.net Hypothesis Volume 8(9) The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation Zahra Karimi 1 *,
www.bioinformation.net Hypothesis Volume 8(9) The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation Zahra Karimi 1 *,
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.
 Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
List of Figures. List of Tables
 Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular
 Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and in a
 Supplementary Figure S1. Distribution of atoms along the bilayer normal. Normalized density profiles obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and
Supplementary Figure S1. Distribution of atoms along the bilayer normal. Normalized density profiles obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 The UBL and RING1 interface remains associated in the complex structures of Parkin and pub. a) Asymmetric Unit of crystal structure of UBLR0RBR and pub complex showing UBL (green),
Supplementary Figure 1 The UBL and RING1 interface remains associated in the complex structures of Parkin and pub. a) Asymmetric Unit of crystal structure of UBLR0RBR and pub complex showing UBL (green),
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al.
 Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
Supplementary Information Janssen et al.
 Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Insights into the Giardia intestinalis Enolase and Human Plasminogen interaction
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Supplementary Information Insights into the Giardia intestinalis Enolase and Human
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Supplementary Information Insights into the Giardia intestinalis Enolase and Human
Supporting Information: Revisiting partition in. hydrated bilayer systems
 Supporting Information: Revisiting partition in hydrated bilayer systems João T. S. Coimbra, Pedro A. Fernandes, Maria J. Ramos* UCIBIO, REQUIMTE, Departamento de Química e Bioquímica, Faculdade de Ciências,
Supporting Information: Revisiting partition in hydrated bilayer systems João T. S. Coimbra, Pedro A. Fernandes, Maria J. Ramos* UCIBIO, REQUIMTE, Departamento de Química e Bioquímica, Faculdade de Ciências,
Supplementary Information
 Coarse-Grained Molecular Dynamics Simulations of Photoswitchable Assembly and Disassembly Xiaoyan Zheng, Dong Wang,* Zhigang Shuai,* MOE Key Laboratory of Organic Optoelectronics and Molecular Engineering,
Coarse-Grained Molecular Dynamics Simulations of Photoswitchable Assembly and Disassembly Xiaoyan Zheng, Dong Wang,* Zhigang Shuai,* MOE Key Laboratory of Organic Optoelectronics and Molecular Engineering,
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. Schematic of Ras biochemical coupled assay.
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. Schematic of Ras biochemical coupled assay.
Table S1. X-ray data collection and refinement statistics
 Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Supporting Information
 Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Supplementary Materials. High affinity binding of phosphatidylinositol-4-phosphate. by Legionella pneumophila DrrA
 Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Supplementary Figure 1. Overview of steps in the construction of photosynthetic protocellular systems
 Supplementary Figure 1 Overview of steps in the construction of photosynthetic protocellular systems (a) The small unilamellar vesicles were made with phospholipids. (b) Three types of small proteoliposomes
Supplementary Figure 1 Overview of steps in the construction of photosynthetic protocellular systems (a) The small unilamellar vesicles were made with phospholipids. (b) Three types of small proteoliposomes
Supplementary Information for Structural basis for the inhibition of Polo-like kinase 1
 Supplementary Information for Structural basis for inhibition of Polo-like kinase 1 Jun Xu, Chen Shen, Tao Wang & Junmin Quan Correspondence authors: Tao Wang, taowang@pkusz.edu.cn; Junmin Quan, quanjm@szpku.edu.cn
Supplementary Information for Structural basis for inhibition of Polo-like kinase 1 Jun Xu, Chen Shen, Tao Wang & Junmin Quan Correspondence authors: Tao Wang, taowang@pkusz.edu.cn; Junmin Quan, quanjm@szpku.edu.cn
Supplementary Information: A Critical. Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers
 Supplementary Information: A Critical Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers Kristyna Pluhackova,, Sonja A. Kirsch, Jing Han, Liping Sun,
Supplementary Information: A Critical Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers Kristyna Pluhackova,, Sonja A. Kirsch, Jing Han, Liping Sun,
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Cryo EM structure of the ribosome SecYE complex in the
 Supplementary Information Cryo EM structure of the ribosome SecYE complex in the membrane environment Jens Frauenfeld,, James Gumbart, Eli O. van der Sluis,, Soledad Funes, Marco Gartmann,, Birgitta Beatrix,,
Supplementary Information Cryo EM structure of the ribosome SecYE complex in the membrane environment Jens Frauenfeld,, James Gumbart, Eli O. van der Sluis,, Soledad Funes, Marco Gartmann,, Birgitta Beatrix,,
Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex
 John Clizer & Greg Ralph Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex The turkey ovomucoid third domain (OMTKY3) is considered to be one of the most studied protein inhibitors. 1 Ovomucin
John Clizer & Greg Ralph Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex The turkey ovomucoid third domain (OMTKY3) is considered to be one of the most studied protein inhibitors. 1 Ovomucin
STRUCTURE OF A UBIQUITIN-LOADED HECT LIGASE REVEALS THE MOLECULAR BASIS FOR CATALYTIC PRIMING SUPPLEMENTARY INFORMATION
 STRUCTURE OF A UBIQUITINLOADED ECT LIGASE REVEALS TE MOLECULAR BASIS FOR CATALYTIC PRIMING Elena Maspero 1, Eleonora Valentini 1, Sara Mari 1, Valentina Cecatiello 2, Paolo Soffientini 1, Sebastiano Pasqualato
STRUCTURE OF A UBIQUITINLOADED ECT LIGASE REVEALS TE MOLECULAR BASIS FOR CATALYTIC PRIMING Elena Maspero 1, Eleonora Valentini 1, Sara Mari 1, Valentina Cecatiello 2, Paolo Soffientini 1, Sebastiano Pasqualato
Interactions of Polyethylenimines with Zwitterionic and. Anionic Lipid Membranes
 Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Supporting Information for Origin of 1/f noise in hydration dynamics on lipid membrane surfaces
 Supporting Informati for Origin of /f noise in hydrati dynamics lipid membrane surfaces Eiji Yamamoto, Takuma kimoto, Masato Yasui, 2 and Kenji Yasuoka Department of Mechanical Engineering, Keio University,
Supporting Informati for Origin of /f noise in hydrati dynamics lipid membrane surfaces Eiji Yamamoto, Takuma kimoto, Masato Yasui, 2 and Kenji Yasuoka Department of Mechanical Engineering, Keio University,
Supplementary Information. A lithium-ion-active aerolysin nanopore for effectively trapping long. single-stranded DNA.
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2018 Supplementary Information A lithium-ion-active aerolysin nanopore for effectively trapping
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2018 Supplementary Information A lithium-ion-active aerolysin nanopore for effectively trapping
Structure of the measles virus hemagglutinin bound to the CD46 receptor. César Santiago, María L. Celma, Thilo Stehle and José M.
 Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
Biological Membranes. Lipid Membranes. Bilayer Permeability. Common Features of Biological Membranes. A highly selective permeability barrier
 Biological Membranes Structure Function Composition Physicochemical properties Self-assembly Molecular models Lipid Membranes Receptors, detecting the signals from outside: Light Odorant Taste Chemicals
Biological Membranes Structure Function Composition Physicochemical properties Self-assembly Molecular models Lipid Membranes Receptors, detecting the signals from outside: Light Odorant Taste Chemicals
Supplementary Information
 Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Adaptable Lipid Matrix Promotes Protein Protein Association in Membranes
 Supporting information Adaptable Lipid Matrix Promotes Protein Protein Association in Membranes Andrey S. Kuznetsov, Anton A. Polyansky,, Markus Fleck, Pavel E. Volynsky, and Roman G. Efremov *,, M. M.
Supporting information Adaptable Lipid Matrix Promotes Protein Protein Association in Membranes Andrey S. Kuznetsov, Anton A. Polyansky,, Markus Fleck, Pavel E. Volynsky, and Roman G. Efremov *,, M. M.
Supplementary Figure 1. Heavy chain sequences of 2G1 and 8M2 aligned with V H 1-69
 Supplementary Figure 1. Heavy chain sequences of 2G1 and 8M2 aligned with V H 1-69 germline gene sequence. 2G1 and 8M2 acquired 14 and 18 mutations from the germline gene sequence, respectively. In 2G1,
Supplementary Figure 1. Heavy chain sequences of 2G1 and 8M2 aligned with V H 1-69 germline gene sequence. 2G1 and 8M2 acquired 14 and 18 mutations from the germline gene sequence, respectively. In 2G1,
Membranes & Membrane Proteins
 School on Biomolecular Simulations Membranes & Membrane Proteins Vani Vemparala The Institute of Mathematical Sciences Chennai November 13 2007 JNCASR, Bangalore Cellular Environment Plasma membrane extracellular
School on Biomolecular Simulations Membranes & Membrane Proteins Vani Vemparala The Institute of Mathematical Sciences Chennai November 13 2007 JNCASR, Bangalore Cellular Environment Plasma membrane extracellular
BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached
 BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached 1). 20 points total T or F (2 points each; if false, briefly state why
BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached 1). 20 points total T or F (2 points each; if false, briefly state why
Data are contained in multiple tabs in Excel spreadsheets and in CSV files.
 Contents Overview Curves Methods Measuring enzymatic activity (figure 2) Enzyme characterisation (Figure S1, S2) Enzyme kinetics (Table 3) Effect of ph on activity (figure 3B) Effect of metals and inhibitors
Contents Overview Curves Methods Measuring enzymatic activity (figure 2) Enzyme characterisation (Figure S1, S2) Enzyme kinetics (Table 3) Effect of ph on activity (figure 3B) Effect of metals and inhibitors
Review II: The Molecules of Life
 Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
Table S1: Kinetic parameters of drug and substrate binding to wild type and HIV-1 protease variants. Data adapted from Ref. 6 in main text.
 Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Biochemistry - I. Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I
 Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
FCC2 5CY7 FCC1 5CY8. Actinonin 5CVQ
 Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
During the last half century, much effort has been devoted
 Membranes serve as allosteric activators of phospholipase A 2, enabling it to extract, bind, and hydrolyze phospholipid substrates Varnavas D. Mouchlis a,b,1, Denis Bucher b, J. Andrew McCammon a,b,c,1,
Membranes serve as allosteric activators of phospholipase A 2, enabling it to extract, bind, and hydrolyze phospholipid substrates Varnavas D. Mouchlis a,b,1, Denis Bucher b, J. Andrew McCammon a,b,c,1,
Molecular Dynamics Simulation Study Explaining Inhibitor Selectivity in Different Class of Histone Deacetylases
 Journal of Biomolecular Structure & Dynamics, ISSN 0739-1102 Volume 29, Issue Number 4, (2012) Adenine Press (2012) Abstract Molecular Dynamics Simulation Study Explaining Inhibitor Selectivity in Different
Journal of Biomolecular Structure & Dynamics, ISSN 0739-1102 Volume 29, Issue Number 4, (2012) Adenine Press (2012) Abstract Molecular Dynamics Simulation Study Explaining Inhibitor Selectivity in Different
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction
 Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Modeling and Molecular Dynamics of Membrane Proteins
 Modeling and Molecular Dynamics of Membrane Proteins Emad Tajkhorshid Department of Biochemistry, Center for Biophysics and Computational Biology, and Beckman Institute University of Illinois at Urbana-Champaign
Modeling and Molecular Dynamics of Membrane Proteins Emad Tajkhorshid Department of Biochemistry, Center for Biophysics and Computational Biology, and Beckman Institute University of Illinois at Urbana-Champaign
This PDF file includes: Supplementary Figures 1 to 6 Supplementary Tables 1 to 2 Supplementary Methods Supplementary References
 Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Introduction to Protein Structure Collection
 Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Secondary Structure North 72nd Street, Wauwatosa, WI Phone: (414) Fax: (414) dmoleculardesigns.com
 Secondary Structure In the previous protein folding activity, you created a generic or hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously
Secondary Structure In the previous protein folding activity, you created a generic or hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously
HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total. The ph at which a peptide has no net charge is its isoelectric point.
 BIOCHEMISTRY I HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total 1). 8 points total T or F (2 points each; if false, briefly state why it is false) The ph at which a peptide has no
BIOCHEMISTRY I HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total 1). 8 points total T or F (2 points each; if false, briefly state why it is false) The ph at which a peptide has no
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information.
 Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth 2 points.
 MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
7/11/17. Cell Function & Chemistry. Molecular and Cellular Biology. 2. Bio-Chemical Foundations & Key Molecules of a Cell
 Molecular and Cellular Biology Cell Function & Chemistry 2. Bio-Chemical Foundations & Key Molecules of a Cell Prof. Dr. Klaus Heese Interaction Molecular Bonds Define Cellular Functions Water H 2 O Interactions
Molecular and Cellular Biology Cell Function & Chemistry 2. Bio-Chemical Foundations & Key Molecules of a Cell Prof. Dr. Klaus Heese Interaction Molecular Bonds Define Cellular Functions Water H 2 O Interactions
John M. Leveritt III, Almudena Pino-Angeles, Themis Lazaridis*.
 The structure of a melittin-stabilized pore John M. Leveritt III, Almudena Pino-Angeles, Themis Lazaridis*. Department of Chemistry, The City College of New York, 160 Convent Ave, New York, NY, 10031,
The structure of a melittin-stabilized pore John M. Leveritt III, Almudena Pino-Angeles, Themis Lazaridis*. Department of Chemistry, The City College of New York, 160 Convent Ave, New York, NY, 10031,
Protein Secondary Structure
 Protein Secondary Structure Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 2, pp. 37-45 Problems in textbook: chapter 2, pp. 63-64, #1,5,9 Directory of Jmol structures of proteins: http://www.biochem.arizona.edu/classes/bioc462/462a/jmol/routines/routines.html
Protein Secondary Structure Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 2, pp. 37-45 Problems in textbook: chapter 2, pp. 63-64, #1,5,9 Directory of Jmol structures of proteins: http://www.biochem.arizona.edu/classes/bioc462/462a/jmol/routines/routines.html
Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes
 Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes Kevin Gasperich Advisor: Dmitry Kopelevich ABSTRACT The recent rapid development of applications for fullerenes has led to an increase
Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes Kevin Gasperich Advisor: Dmitry Kopelevich ABSTRACT The recent rapid development of applications for fullerenes has led to an increase
Supplementary Figure 1 Overall structure of the transmembrane pore of FraC. a, Crystal packing of the pore along the y-z, b, the z-x, and c, the y-x
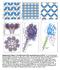 Supplementary Figure 1 Overall structure of the transmembrane pore of FraC. a, Crystal packing of the pore along the y-z, b, the z-x, and c, the y-x planes. The N-terminal and the β-core regions are depicted
Supplementary Figure 1 Overall structure of the transmembrane pore of FraC. a, Crystal packing of the pore along the y-z, b, the z-x, and c, the y-x planes. The N-terminal and the β-core regions are depicted
Membrane Sculpting by F-BAR Domains Studied by Molecular Dynamics Simulations
 Studied by Molecular Dynamics Simulations Hang Yu 1,2, Klaus Schulten 1,2,3 * 1 Beckman Institute, University of Illinois, Urbana, Illinois, United States of America, 2 Center of Biophysics and Computational
Studied by Molecular Dynamics Simulations Hang Yu 1,2, Klaus Schulten 1,2,3 * 1 Beckman Institute, University of Illinois, Urbana, Illinois, United States of America, 2 Center of Biophysics and Computational
Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector
 kda 1 2 3 4 5 26 14 1 7 Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector 5 4 35 25 15 1 Supplementary Figure 1. SDS-PAGE of acterially expressed and purified proteins.
kda 1 2 3 4 5 26 14 1 7 Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector 5 4 35 25 15 1 Supplementary Figure 1. SDS-PAGE of acterially expressed and purified proteins.
CHAPTER 4. Tryptophan fluorescence quenching by brominated lipids
 CHAPTER 4 Tryptophan fluorescence quenching by brominated lipids 102 4.1 INTRODUCTION The structure and dynamics of biological macromolecules have been widely studied with fluorescence quenching. The accessibility
CHAPTER 4 Tryptophan fluorescence quenching by brominated lipids 102 4.1 INTRODUCTION The structure and dynamics of biological macromolecules have been widely studied with fluorescence quenching. The accessibility
Structural analysis of fungus-derived FAD glucose dehydrogenase
 Structural analysis of fungus-derived FAD glucose dehydrogenase Hiromi Yoshida 1, Genki Sakai 2, Kazushige Mori 3, Katsuhiro Kojima 3, Shigehiro Kamitori 1, and Koji Sode 2,3,* 1 Life Science Research
Structural analysis of fungus-derived FAD glucose dehydrogenase Hiromi Yoshida 1, Genki Sakai 2, Kazushige Mori 3, Katsuhiro Kojima 3, Shigehiro Kamitori 1, and Koji Sode 2,3,* 1 Life Science Research
SUPPORTING INFORMATION FOR. A Computational Approach to Enzyme Design: Using Docking and MM- GBSA Scoring
 SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors
 Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
Bioinformatics for molecular biology
 Bioinformatics for molecular biology Structural bioinformatics tools, predictors, and 3D modeling Structural Biology Review Dr Research Scientist Department of Microbiology, Oslo University Hospital -
Bioinformatics for molecular biology Structural bioinformatics tools, predictors, and 3D modeling Structural Biology Review Dr Research Scientist Department of Microbiology, Oslo University Hospital -
Break Desired Compound into 3 Smaller Ones
 Break Desired Compound into 3 Smaller nes A B C Indole ydrazine Phenol When creating a covalent link between model compounds move the charge on the deleted into the carbon to maintain integer charge (i.e.
Break Desired Compound into 3 Smaller nes A B C Indole ydrazine Phenol When creating a covalent link between model compounds move the charge on the deleted into the carbon to maintain integer charge (i.e.
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
Rapid Characterization of Allosteric Networks. with Ensemble Normal Mode Analysis
 Supporting Information Rapid Characterization of Allosteric Networks with Ensemble Normal Mode Analysis Xin-Qiu Yao 1, Lars Skjærven 2, and Barry J. Grant 1* 1 Department of Computational Medicine and
Supporting Information Rapid Characterization of Allosteric Networks with Ensemble Normal Mode Analysis Xin-Qiu Yao 1, Lars Skjærven 2, and Barry J. Grant 1* 1 Department of Computational Medicine and
Life Sciences 1a. Practice Problems 4
 Life Sciences 1a Practice Problems 4 1. KcsA, a channel that allows K + ions to pass through the membrane, is a protein with four identical subunits that form a channel through the center of the tetramer.
Life Sciences 1a Practice Problems 4 1. KcsA, a channel that allows K + ions to pass through the membrane, is a protein with four identical subunits that form a channel through the center of the tetramer.
Coarse grained simulations of Lipid Bilayer Membranes
 Coarse grained simulations of Lipid Bilayer Membranes P. B. Sunil Kumar Department of Physics IIT Madras, Chennai 600036 sunil@iitm.ac.in Atomistic MD: time scales ~ 10 ns length scales ~100 nm 2 To study
Coarse grained simulations of Lipid Bilayer Membranes P. B. Sunil Kumar Department of Physics IIT Madras, Chennai 600036 sunil@iitm.ac.in Atomistic MD: time scales ~ 10 ns length scales ~100 nm 2 To study
Fluid Mozaic Model of Membranes
 Replacement for the 1935 Davson Danielli model Provided explanation for Gortner-Grendel lack of lipid and permitted the unit membrane model. Trans membrane protein by labelling Fry & Edidin showed that
Replacement for the 1935 Davson Danielli model Provided explanation for Gortner-Grendel lack of lipid and permitted the unit membrane model. Trans membrane protein by labelling Fry & Edidin showed that
Activities for the α-helix / β-sheet Construction Kit
 Activities for the α-helix / β-sheet Construction Kit The primary sequence of a protein, composed of amino acids, determines the organization of the sequence into the secondary structure. There are two
Activities for the α-helix / β-sheet Construction Kit The primary sequence of a protein, composed of amino acids, determines the organization of the sequence into the secondary structure. There are two
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids
 Supporting Information Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids Phansiri Boonnoy 1, Viwan Jarerattanachat 1,2, Mikko Karttunen 3*, and Jirasak Wongekkabut 1* 1 Department of
Supporting Information Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids Phansiri Boonnoy 1, Viwan Jarerattanachat 1,2, Mikko Karttunen 3*, and Jirasak Wongekkabut 1* 1 Department of
Supplementary materials
 Supplementary materials Chemical library from ChemBridge 50,240 structurally diverse small molecule compounds dissolved in DMSO Hits Controls: No virus added μ Primary screening at 20 g/ml of compounds
Supplementary materials Chemical library from ChemBridge 50,240 structurally diverse small molecule compounds dissolved in DMSO Hits Controls: No virus added μ Primary screening at 20 g/ml of compounds
HDL surface lipids mediate CETP binding as revealed by electron microscopy and molecular dynamics simulation
 HDL surface lipids mediate CETP binding as revealed by electron microscopy and molecular dynamics simulation Meng Zhang 1, River Charles 1, Huimin Tong 1, Lei Zhang 1, Mili Patel 2, Francis Wang 1, Matthew
HDL surface lipids mediate CETP binding as revealed by electron microscopy and molecular dynamics simulation Meng Zhang 1, River Charles 1, Huimin Tong 1, Lei Zhang 1, Mili Patel 2, Francis Wang 1, Matthew
Supplementary Information
 Supplementary Information Structural Basis of Sialidase in Complex with Geranylated Flavonoids as Potent Natural Inhibitors Youngjin Lee, a,b Young Bae Ryu, d Hyung-Seop Youn, a,b Jung Keun Cho, e Young
Supplementary Information Structural Basis of Sialidase in Complex with Geranylated Flavonoids as Potent Natural Inhibitors Youngjin Lee, a,b Young Bae Ryu, d Hyung-Seop Youn, a,b Jung Keun Cho, e Young
Levels of Protein Structure:
 Levels of Protein Structure: PRIMARY STRUCTURE (1 ) - Defined, non-random sequence of amino acids along the peptide backbone o Described in two ways: Amino acid composition Amino acid sequence M-L-D-G-C-G
Levels of Protein Structure: PRIMARY STRUCTURE (1 ) - Defined, non-random sequence of amino acids along the peptide backbone o Described in two ways: Amino acid composition Amino acid sequence M-L-D-G-C-G
We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field
 Parameterization of the fullerene coarse-grained model We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field for lipids 1 and proteins 2. In the MARTINI force
Parameterization of the fullerene coarse-grained model We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field for lipids 1 and proteins 2. In the MARTINI force
Molecular Dynamics Simulation of Transmembrane Polypeptide. Orientational Fluctuations
 Biophys J BioFAST, published on October 15, 2004 as doi:10.1529/biophysj.104.047506 Molecular Dynamics Simulation of Transmembrane Polypeptide Orientational Fluctuations David J. Goodyear*, Simon Sharpe,
Biophys J BioFAST, published on October 15, 2004 as doi:10.1529/biophysj.104.047506 Molecular Dynamics Simulation of Transmembrane Polypeptide Orientational Fluctuations David J. Goodyear*, Simon Sharpe,
Phospholipid Component Volumes: Determination and Application to Bilayer Structure Calculations
 734 Biophysical Journal Volume 75 August 1998 734 744 Phospholipid Component Volumes: Determination and Application to Bilayer Structure Calculations Roger S. Armen, Olivia D. Uitto, and Scott E. Feller
734 Biophysical Journal Volume 75 August 1998 734 744 Phospholipid Component Volumes: Determination and Application to Bilayer Structure Calculations Roger S. Armen, Olivia D. Uitto, and Scott E. Feller
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
Four-Scale Description of Membrane Sculpting by BAR Domains
 2806 Biophysical Journal Volume 95 September 2008 2806 2821 Four-Scale Description of Membrane Sculpting by BAR Domains Anton Arkhipov, Ying Yin, and Klaus Schulten Department of Physics and Beckman Institute,
2806 Biophysical Journal Volume 95 September 2008 2806 2821 Four-Scale Description of Membrane Sculpting by BAR Domains Anton Arkhipov, Ying Yin, and Klaus Schulten Department of Physics and Beckman Institute,
Supplementary Information
 Supplementary Information Structural basis of improved second generation 3-nitro-tyrosine trna synthetases Richard B. Cooley, Jessica L. Feldman, Camden M. Driggers, Taylor Bundy, Audrey L. Stokes, P.
Supplementary Information Structural basis of improved second generation 3-nitro-tyrosine trna synthetases Richard B. Cooley, Jessica L. Feldman, Camden M. Driggers, Taylor Bundy, Audrey L. Stokes, P.
Acta Crystallographica Section D
 Supporting information Acta Crystallographica Section D Volume 70 (2014) Supporting information for article: A conformational landscape for alginate secretion across the outer membrane of Pseudomonas aeruginosa
Supporting information Acta Crystallographica Section D Volume 70 (2014) Supporting information for article: A conformational landscape for alginate secretion across the outer membrane of Pseudomonas aeruginosa
Secondary Structure. by hydrogen bonds
 Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
Main Functions maintain homeostasis
 The Cell Membrane Main Functions The main goal is to maintain homeostasis. Regulates materials moving in and out of the cell. Provides a large surface area on which specific chemical reactions can occur.
The Cell Membrane Main Functions The main goal is to maintain homeostasis. Regulates materials moving in and out of the cell. Provides a large surface area on which specific chemical reactions can occur.
Biology 2E- Zimmer Protein structure- amino acid kit
 Biology 2E- Zimmer Protein structure- amino acid kit Name: This activity will use a physical model to investigate protein shape and develop key concepts that govern how proteins fold into their final three-dimensional
Biology 2E- Zimmer Protein structure- amino acid kit Name: This activity will use a physical model to investigate protein shape and develop key concepts that govern how proteins fold into their final three-dimensional
G protein coupled receptor interactions with cholesterol deep in the membrane
 G protein coupled receptor interactions with cholesterol deep in the membrane Samuel Genheden a,b, Jonathan W. Essex a,c, & Anthony G. Lee c,d * a School of Chemistry, University of Southampton, Southampton,
G protein coupled receptor interactions with cholesterol deep in the membrane Samuel Genheden a,b, Jonathan W. Essex a,c, & Anthony G. Lee c,d * a School of Chemistry, University of Southampton, Southampton,
