De novo replication of the influenza virus RNA genome is regulated by DNA replicative helicase, MCM
|
|
|
- Iris Hutchinson
- 5 years ago
- Views:
Transcription
1 The EMBO Journal (2007) 26, & 2007 European Molecular Biology Organization All Rights Reserved /07 De novo replication of the influenza virus RNA genome is regulated by DNA replicative helicase, MCM THE EMBO JOURNAL Atsushi Kawaguchi and Kyosuke Nagata* 1 Department of Infection Biology, Graduate School of Comprehensive Human Sciences and Institute of Basic Medical Sciences, University of Tsukuba, Tsukuba, Japan By dissecting and reconstituting a cell-free influenza virus genome replication system, we have purified and identified the minichromosome maintenance (MCM) complex, which is thought to be a DNA replicative helicase, as one of the host factors that regulate the virus genome replication. MCM interacted with the PA subunit of the viral RNAdependent RNA polymerase that is found to be involved in the replication genetically. The virus genome replication was decreased in MCM2 knockdown cells. The viral polymerase appeared to be a nonproductive complex, that is, it was capable of initiating replication but produced only abortive short RNA chains. MCM stimulated de novoinitiated replication reaction by stabilizing a replication complex during its transition from initiation to elongation. Based on the findings, including the result that the MCMmediated RNA replication reaction was competed with exogenously added RNA, we propose that MCM functions as a scaffold between the nascent RNA chains and the viral polymerase. The EMBO Journal (2007) 26, doi: / sj.emboj ; Published online 11 October 2007 Subject Categories: RNA; microbiology & pathogens Keywords: host factor; influenza virus; MCM; replication; RNA-dependent RNA polymerase Introduction Viruses are intracellular parasites. Since virus genomes are considerably small and can thus encode only a limited number of genes, viruses must use host factors and machineries to replicate. Therefore, the identification and functional characterization of host factors are indispensable to understand the mechanism of viral replication and pathogenicity, and provide with critical insights into the virus host interaction, which is shaped by unending adaptation and eradication between a virus and its host. Viruses often utilize critical regulatory processes of the host cell; therefore, studies on the processes subverted by viruses have often highlighted cellular regulatory mechanisms. This has indeed been the *Corresponding author. Department of Infection Biology, Graduate School of Comprehensive Human Sciences and Institute of Basic Medical Sciences, University of Tsukuba, Tennodai, Tsukuba , Japan. Tel.: þ ; Fax: þ ; knagata@md.tsukuba.ac.jp Received: 12 June 2007; accepted: 18 September 2007; published online: 11 October 2007 case, particularly for eukaryotic genome replication and transcription. The reconstitution of cell-free transcription and replication systems of the adenovirus and SV40 genomes, and the identification of host factors by biochemical fractionation and complementation have greatly contributed to our knowledge of the fundamental processes of eukaryotic replication and transcription. The genome of influenza type A viruses consists of eightsegmented and single-stranded RNAs of negative polarity. The viral RNA genome (vrna) is transcribed into mrna and replicated through crna (full-sized complementary copy of vrna) to produce a large number of progeny vrnas in the nucleus (reviewed in Engelhardt and Fodor, 2006). In viral particles and infected cells, the vrna exists as ribonucleoprotein (designated vrnp) complexes with viral RNA-dependent RNA polymerases consisting of three subunits PB1, PB2, and PA and nucleoprotein (NP). PB1 contains the conserved motifs characteristic of RNA polymerases and functions as a polymerase catalytic subunit for the sequential addition of nucleotides to the elongating RNA. PB1 binds to the and 3 0 -terminal sequences of vrna and crna, which are conserved in all segments and act as cis-acting elements for the viral RNA synthesis. Transcription is initiated using the oligonucleotide containing the cap-1 structure derived from cellular pre-mrnas as a primer. The capped oligonucleotide is generated by the recognition of the cap structure by PB2 and endonucleolytic cleavage by PB1. The elongation of the mrna chain proceeds until the polymerase reaches a polyadenylation signal, which consists of 5 7 U residues located near the 5 0 -terminal region of the vrna. In contrast, the genome replication is primer-independent and generates full-length vrna through crna synthesis. Genetic analyses suggest that PA participates in the replication process, although the precise function of PA is not well established (Sugiura et al, 1975; Ritchey and Palese, 1977). Recently, we found that PA is involved in the assembly of a functional polymerase (Kawaguchi et al, 2005). It was reported that NP is also important for the replication process (Shapiro and Krug, 1988; Medcalf et al, 1999). However, the precise function of NP in the replication remains uncertain. Since the replication and regulated transcription of the influenza virus genome do not occur only by the influenza viral components associated with virions, it has been thought that some factor(s) present in infected cells is required for the regulation of these processes. In fact, it has been reported that vrnp interacts with several cellular proteins (Wang et al, 1997; Digard et al, 1999; Huarte et al, 2001; Engelhardt et al, 2005; Garcia-Robles et al, 2005; Deng et al, 2006). In this regard, by dissecting a cell-free viral RNA synthesis system mimicking the viral transcription, we have identified RAF-1/ Hsp90 and RAF-2p48/UAP56/BAT1 as host factors that stimulate the viral RNA synthesis. RAF-1/Hsp90 regulates the assembly of viral RNA polymerase complexes and is also 4566 The EMBO Journal VOL 26 NO & 2007 European Molecular Biology Organization
2 involved in their stabilization during their transfer between templates (Momose et al, 2002; Naito et al, 2007). RAF-2p48/ UAP56/BAT1, which is thought to be involved in the RNA splicing of cellular mrna, interacts with NP and facilitates the NP RNA complex formation, thereby stimulating viral RNA synthesis by the viral RNA polymerase (Momose et al, 2001). Interestingly, it was reported that the viral RNA polymerase interacts with the serine 5-phosphorylated carboxyterminal domain of the largest subunit of the cellular RNA polymerase II (pol II), which is associated with a pol II transcription initiation complex and plays a role in the recruitment and stimulation of capping enzyme (Chan et al, 2006). However, the host factor(s) involved in the influenza virus genome replication has not been identified yet. Recently, it was suggested that the efficiency of transcription and replication by the viral RNA polymerase is a determinant of both host specificity and pathogenicity, and this may be regulated by the adaptation of the viral RNA polymerase to an unknown host factor(s) (Gabriel et al, 2005, 2007). Here, we reconstituted a cell-free virus genome replication system with virion-associated vrnp and nuclear extracts prepared from uninfected HeLa cells. By biochemical fractionation and complementation, we purified Influenza virus REplication Factor (IREF)-1 as a factor that is required for successful virus genome replication. TOF-MS analyses revealed that IREF-1 is a minichromosome maintenance (MCM) heterohexamer complex consisting of MCM2, 3, 4, 5, 6, and 7. MCM proteins were first identified for their roles in plasmid replication or cell cycle progression in yeast (Maine et al, 1984; Sinha et al, 1986). It is generally believed, although not completely proven, that MCM functions as a eukaryotic DNA replication fork helicase. In this report, we demonstrate that IREF-1/MCM stimulates virus genome replication by increasing the stability of replicating RNA polymerases, which produce only abortive short RNA chains in the absence of IREF-1/MCM. IREF-1/MCM interacted with vrnp through its contact with PA. Biochemical analyses strongly suggest that MCM stabilizes the replicating polymerase complexes by promoting the interaction between the viral polymerase and nascent crna. Taken together with the fact that virus genome replication was decreased in MCM knockdown (KD) cells, it is suggested that IREF-1/MCM is a host factor that regulates the influenza virus genome replication. Results Purification and characterization of IREF-1 The RNA-dependent RNA polymerase of influenza virus catalyzes both primer-dependent mrna synthesis and primer-independent virus genome replication in the nuclei of infected cells. It is demonstrated in cell-free RNA synthesis systems that vrnp isolated from virions catalyzes the primerdependent RNA synthesis (Plotch and Krug, 1977; Plotch et al, 1979), which is stimulated by host factors (Momose et al, 2001; Momose et al, 2002). De novo initiation of crna synthesis was also observed in a cell-free RNA synthesis system using vrnp or partially purified polymerase fractions as enzyme source (Lee et al, 2002; Vreede and Brownlee, 2007). However, in the system, only partial fragments of crna were detected, but the efficient full-length crna synthesis was not demonstrated. Therefore, it is possible that vrnp could be potentially but not highly active to synthesize crna. Figure 1 Purification of IREF-1. (A) General scheme for purification of IREF-1. (B) Stimulatory activity of IREF-1 for cell-free virus genome replication. RNA synthesis was carried out in the absence (lanes 1 4) or presence (lane 5) of ApG dinucleotide primer with 0 (lane 1), 0.5 (lane 2), 1 (lane 3), and 2 (lane 4) ml of purified IREF-1. (C) Identification of the polarity of newly synthesized RNA by using RNase T1 digestion. A band corresponding to segment 8 newly synthesized in the presence of [a- 32 P]UTP was excised from gel, and digested with RNase T1 (lane 3). Cleaved products were separated and visualized by autoradiography. In vitro synthesized vrna (lane 1) and crna (lane 2) of segment 8 using T7 RNA polymerase were also analyzed. Thus, it is reasonably hypothesized that some host protein(s) supports the primer-independent virus genome replication. Here, we first examined whether nuclear extracts prepared from uninfected cells promote vrnp to replicate the virus genome. This was indeed the case. The primer-independent replicative full-sized crna synthesis was promoted in the presence of nuclear extracts (see below), suggesting the presence of a host factor(s) involved in virus genome replication. We designated such an activity as Influenza virus REplication Factor-1 (IREF-1), and attempted to purify the factor(s) responsible for replication by the biochemical fractionation of nuclear extracts prepared from uninfected HeLa cells through sequential column chromatography and complementation of vrnp for replicative activity (Figure 1A). IREF-1 promoted the synthesis of full-length viral RNAs in a dose-dependent manner (Figure 1B). Since the influenza virus genome consists of eight-segmented and singlestranded RNAs, we detected eight segments of newly synthesized RNAs. The electrophoretic mobility of these RNAs synthesized in the presence of IREF-1 was similar to that of the virus genome RNAs prepared from purified virions (data not shown) and that of RNAs synthesized in the presence of the ApG dinucleotide primer complementary to the nucleotide positions 1 and 2 of the 3 0 -terminus of vrna (lane 5, and see Ritchey and Palese, 1977; Honda et al, 1986). & 2007 European Molecular Biology Organization The EMBO Journal VOL 26 NO
3 To determine the polarity of newly synthesized RNA, the RNA possibly synthesized from segment 8 was excised from the gel, and the isolated RNA was then digested with RNase T1. Since RNase T1 cleaves between the 3 0 -phosphate group of a guanine ribonucleotide and the 5 0 -hydroxyl group of the adjacent nucleotide, the digestion of vrna by RNase T1 yields RNA fragments of 42, 19, and 17 nt with various smaller fragments, while only small fragments are yielded for crna (Figure 1C, lanes 1 and 2). The cleavage pattern of the RNA synthesized in the presence of IREF-1 was found to be similar to that of the crna (Figure 1C, lane 3). These RNAs were digested with RNase H when hybridized with an oligonucleotide specific for the crna polarity (data not shown). Therefore, it is confirmed that the RNA synthesis mediated by IREF-1 generates full-sized crna in the absence of any primer. The RNA synthesis level in the presence of IREF-1 varied from segment to segment. The reason for this segment-specific efficiency of RNA synthesis is presently unknown. De novo initiation of the genome replication in the presence of IREF-1 One of the essential features of the RNA synthesized in a primer-independent manner is the presence of a triphosphate moiety at its 5 0 -end. In order to confirm that newly synthesized crna has the 5 0 -triphosphate end group, we compared the mobility of the crnas synthesized in the presence of IREF-1 (Figure 2A, lane 2) with that of ApG-primed products (Figure 2A, lane 3) in a limited elongation assay, in which UTP is omitted from the reaction mixture and the RNA polymerase pauses at the first adenine residue on the template (Momose et al, 2002). The expected lengths of limited elongation products are 12 nt for segments 1, 3, and 7; 13 nt for segments 5 and 8; 14 nt for segment 6; 18 nt for segment 4; and 19 nt for segment 2. The crnas synthesized in the presence of IREF-1 migrated differently from the ApG-primed products (Figure 2A, compare lane 2 with lane 3). The shortest limited elongation product in the presence of ApG is 12 nt, but the shortest product in the presence of IREF-1 migrated faster than the ApG-primed 12-nt-long product. To examine whether this difference in mobility is due to the absence or presence of 5 0 -triphosphate, the synthesized RNA bands indicated by asterisks were eluted from the gel, treated with Escherichia coli alkaline phosphatase, and then subjected to separation by 15% PAGE containing 8 M urea (Figure 2B). Following alkaline phosphatase treatment, the crna synthesized in the presence of IREF-1 migrated to the same position as the ApG-primed products (Figure 2B, lane 4). Next, we attempted to separate 5 0 -triphosphate by thinlayer chromatography after digestion with either RNase T2 or snake venom phosphodiesterase (SV-PDE) (Figure 2C). The first and second nucleotides of the crna are A and G residues, respectively. Digestion by RNase T2 of [a- 32 P]GTP-labeled unprimed crna would yield pppa 32 pif the 5 0 -triphosphate is present, whereas SV-PDE yields 32 pg. We detected pppa 32 p only from the crna synthesized in the presence of IREF-1 by RNase T2 digestion (Figure 2C, lane 4). Therefore, it was concluded that IREF-1 promotes de novoinitiated virus genome replication. Note that unprimed crnas in limited elongation assays were detected even in the absence of IREF-1 and ApG (Figure 2A, lane 1). This is in good agreement with the Figure 2 Detection of 5 0 triphosphate end group. (A) Limited elongation assays. Limited elongation assays were carried out in the absence (lanes 1 and 2) or presence (lane 3) of ApG with (lane 2) or without (lanes 1 and 3) IREF-1. (B) Alkaline phosphatase treatment of limited elongation products. Limited elongation products (indicated by asterisks in panel A) synthesized in the presence of ApG (lanes 1 and 2) or IREF-1 (lanes 3 and 4) were excised from gel, and treated with (lanes 1 and 3) or without (lanes 2 and 4) alkaline phosphatase (AP). (C) Detection of pppap using thin-layer chromatography. Twelve nucleotide long of limited elongation products in the presence of ApG (lanes 1 3) or IREF-1 (lanes 4 6) were excised from gel, and treated with RNase T2 (lanes 1, 2, 4, 5, and 7) or SV-PDE (lanes 3 and 6). Products were separated through thin-layer chromatography with 1.6 M LiCl, and visualized by autoradiography. Alkaline phosphatase-treated limited elongation products (AP; lanes 2 and 4) and RNA synthesized by T7 RNA polymerase in the presence of [g- 32 P]ATP (lane 7) were used as control products. unprimed RNA synthesis of short RNA of crna polarity by the viral RNA polymerase (Lee et al, 2002; Vreede and Brownlee, 2007). Thus, IREF-1 may be required for a step(s) post the initiation and the early elongation reactions, in which only short crnas are synthesized (Figure 2A, lane 2). Taken altogether, we assumed that the viral RNA polymerase encounters difficulty after a de novo initiation reaction, and that IREF-1 resolves this to promote the viral RNA polymerase to synthesize the de novo-initiated and full-sized crna (discussed later along with Figures 6 and 7). Identification of IREF-1 as MCM, a putative DNA replicative helicase Analyses of a Mono Q column fraction with MALDI-TOF MS indicated that human MCM proteins 2, 4, 5, 6, and 7 are major components of the IREF-1 fraction (Figure 3A, lanes 2 5, and Supplementary Table 1S). MCM possesses the DNA helicase activity and plays important roles in the regulation of genomic DNA replication (reviewed in Forsburg, 2004). To confirm the results obtained from MALDI-TOF MS analyses, western blotting assays were carried out with rabbit anti- MCM2 7 antibodies. All MCM2 7 proteins were detected in 4568 The EMBO Journal VOL 26 NO & 2007 European Molecular Biology Organization
4 Figure 3 Identification of functional components in IREF-1. (A) (Left panel) SDS PAGE analysis. The loaded amounts were adjusted to the equal level of the IREF-1 stimulatory activity attained in the cell-free virus genome replication assay. Lane 1, molecular size marker (Bio-Rad); lane 2, uninfected HeLa cell nuclear extracts; lane 3, 0.2 M KCl eluate from phosphocellulose column; lane 4, 0.25 M (NH 4 ) 2 SO 4 eluate from phenyl Sepharose column; lane 5, 0.33 M KCl eluate from Mono Q column (purified IREF-1 fraction). The gel was visualized by silver staining assay. (Right panel) Western blotting assay. The IREF-1 fraction of Mono Q column was separated through SDS PAGE, and subjected to western blotting assays with rabbit anti-mcm2, 3, 4, 5, 6, and 7 antibodies. (B) (Left panel) Purification of recombinant MCM complex from insect cells. The details of purification scheme and column chromatography are described in Supplementary data. Lane 1, molecular size marker (Bio-Rad); lane 2, purified IREF-1 fraction from uninfected HeLa nuclear extracts (Mono Q fraction); lanes 3 5, recombinant MCM heterohexamer complex containing 7.5 (lane 3), 15 (lane 4), 30 ng (lane 5) of MCM2 equivalent. The gel was visualized by silver staining assay. (Right panel) Cell-free virus genome replication assay. Equal amounts of IREF-1 and rmcm loaded in left panel were examined in the cell-free virus genome replication assay. Lane 6, purified IREF-1 fraction (Mono Q fraction); lanes 8 10, rmcm complex containing 7.5, 15, 30 ng of rmcm2. The assay in lane 7 was carried out in the absence of any proteins. the IREF-1 fraction (Figure 3A, lanes 6 12). To examine whether MCM proteins are responsible for the IREF-1 activity, unprimed RNA synthesis was performed in the presence of a recombinant MCM heterohexamer complex (rmcm) (Figure 3B). The rmcm was purified from insect cells expressing MCM proteins by recombinant baculovirus infection (Figure 3B, lanes 3 5). Since MCM 4, 5, and 7 were fused with histidine-tag, the mobility of these proteins was slightly different from that of authentic MCM proteins. Figure 3B shows that rmcm and authentic IREF-1 stimulate virus genome replication up to comparable levels (Figure 3B, lanes 6 10). These results indicate that MCM is responsible for IREF-1 activity. IREF-1/MCM complex interacts with PA polymerase subunit Because IREF-1/MCM was identified as a stimulatory factor in the cell-free virus genome replication system by using vrnp as the enzyme source, the target(s) of MCM must be one or more of the vrnp components, that is, PB1, PB2, PA, and NP. To determine the viral factor(s) that binds to MCM, we carried out immunoprecipitation assays with cell lysates prepared from cells expressing one of the FLAG- or Myc-tagged viral proteins (Figures 4A and B). MCM2 was immunoprecipitated with the PA subunit of the viral RNA polymerase (Figure 4A, lanes 10 12), but not with other viral proteins (lanes 4 9 and 16 18), suggesting that MCM interacts with the viral RNA polymerase through the contact with PA. To confirm whether PA interacts with MCM2 as a component of the MCM complex, we tried to detect other MCM proteins that co-immunoprecipitated with PA by performing western blotting analyses (Figure 4B). Not only MCM2 but also MCM3, 4, 5, and 7 were immunoprecipitated with PA, demonstrating that PA interacts with the MCM complex. We could not detect MCM6 since an anti-mcm6 antisera with high titer was not available. Next, to examine whether MCM also interacts with viral proteins in infected cells, the lysates were prepared from infected cells after crosslinking with DSP and formaldehyde and subjected to immunoprecipitation assays using an anti- MCM2 antibody (Figure 4C). We observed that PA, PB2, and NP co-immunoprecipitated with MCM2. Since Figure 4A reveals that MCM2 does not interact with PB2 and NP when singly expressed (Figure 4A, lanes 7 9 and 16 18) and the immunoprecipitation assay using rmcm and micrococcal nuclease-treated vrnp also showed that NP does not interact with MCM (Supplementary Figure 1S), it is quite likely that MCM interacts with PA in the vrnp complexes in infected cells. Further to confirm this, RNA was purified from immunoprecipitates and subjected to RT PCR with primers specific for the segment 5 vrna. The virus genome was immunoprecipitated with MCM2 (Figure 4D), indicating that MCM interacts with vrnp. There is a significant difference in the amount of proteins co-immunoprecipitated with MCM2 between polymerase proteins and NP (Figure 4C). This could be due to the fact that the amount of polymerase proteins in vrnp was significantly less than that of NP. One NP was shown to bind every 20 nucleotides (Yamanaka et al, 1990). Exactly the same patterns were observed with lysates prepared in the absence of crosslink reagents (data not shown). However, the amount of immunoprecipitated proteins was considerably less. Therefore, it is possible that MCM interacts with vrnp transiently or catalytically. Involvement of MCM in influenza virus genome replication in infected cells Since each MCM gene is essential for the cellular DNA replication, we cannot establish MCM gene knockout cells. It was reported that the nuclear transport of MCM complex is interdependent among MCM proteins (Pasion and Forsburg, 1999). Therefore, using sirna-mediated gene silencing of the MCM genes, we tried to examine whether MCM functions in the influenza virus genome replication in cultured cells (Figure 5A). After 60 h post transfection of duplex RNA oligonucleotides corresponding to the MCM2 and MCM3 genes, the expression level of MCM2 and MCM3 proteins in & 2007 European Molecular Biology Organization The EMBO Journal VOL 26 NO
5 Figure 4 MCM complex interacts with PA polymerase subunit. (A) The interaction of MCM2 with singly expressed viral proteins. HeLa cells were mock-transfected (lanes 1 3 and 13 15) or transfected with each plasmid encoding PB1-FLAG (lanes 4 6), PB2-FLAG (lanes 7 9), PA- FLAG (lanes 10 12), and NP-Myc (lanes 16 18). After 24 h post transfection, cell lysates were prepared and subjected to immunoprecipitation assays in the absence of antibody (lanes 2, 5, 8, 11, 14, and 17) or presence of either mouse anti-flag (lanes 3, 6, 9, and 12) or anti-myc antibody (lanes 15 and 18). Immunoprecipitated proteins were separated, and then visualized by western blotting assays with goat anti-mcm2 and rabbit anti-pb1, PB2, PA, and NP antibodies. (B) The interaction of MCM proteins with singly expressed PA. HeLa cells expressing PA (lanes 4 6) were lysed and subjected to immunoprecipitation assays in the absence (lanes 2 and 5) or presence (lanes 3 and 6) of anti-flag antibody. Lanes 1 3 represent mock experiments. Western blotting analyses were performed with rabbit anti-mcm2, 3, 4, 5, 7, and PA antibodies. (C) The interaction between MCM2 and viral proteins in infected cells. HeLa cells were infected with PR/8/34 influenza virus at an MOI ¼ 10. After 8 h post infection, cell lysates were prepared and subjected to immunoprecipitation assays with either goat control IgG (lanes 2 and 5) or anti-mcm2 antibody (lanes 3 and 6). Immunoprecipitated proteins were detected by western blotting analyses with rabbit anti-pa, PB2, NP, and MCM2 antibodies. (D) The interaction between MCM2 and the virus genome in infected cells. vrnas immunoprecipitated as described in (C) were purified, and then semiquantitatively analyzed by RT PCR with primers specific for segment 5 vrna. To quantitatively evaluate, 0.5% equivalents of mock-treated sample (lane 1) and 0.05, 0.15, and 0.5% equivalents of infected sample (lanes 2 4) were also subjected to RT PCR. transfected HeLa cells decreased to 20% of the cells transfected with random sequence sirna (Figure 5A, lanes 1 and 2). FACS analysis showed that the cell proliferation rate of MCM protein knockdown (KD) cells was found largely unchanged under this condition (data not shown). MCM proteins are expressed in excess over a number of replication origins (Lei et al, 1996; Mahbubani et al, 1997; Edwards et al, 2002), and the normal replication rate is maintained even when the number of MCM is reduced to 1 2 per origin (Mahbubani et al, 1997; Edwards et al, 2002; Cortez et al, 2004; Snyder et al, 2005). It is possible that KD of MCM proteins may cause some level of the G1 arrest and thereby affect the virus replication. We confirmed that the synthesis level of crna and viral mrna in G1 phase-synchronized cells was similar to that in S phase cells (Supplementary Figure 2S). Quantitative RT PCR assays with primer sets specific for segment 5 crna and NP mrna showed that the level of crna synthesis in MCM KD HeLa cells reduce to 70% of that in control cells (Figure 5B, left panel, Po0.01). In contrast, there was no significant difference in the level of mrna synthesis between control and MCM KD HeLa cells (Figure 5B, right panel, P40.05). To evaluate the viral transcription activity independent of the virus genome replication, we examined the level of viral transcription from infecting vrnp using cycloheximide (CHX), a potent protein synthesis inhibitor (Figure 5C). It is shown that CHX suppresses viral protein synthesis and thereby leads to degradation of replicated virus genome RNA but not viral mrna since newly vrnp formation was repressed (Vreede et al, 2004). As expected, the level of mrna synthesis from infecting vrnp in MCM KD cells was similar to that in control cells (Figure 5C). These results indicate that MCM is involved predominantly in virus genome replication in infected cells. These observations were confirmed by a plasmid-based replication system (Supplementary Figure 3S). It could be expected that the reduction of the vrna template causes concomitant reduction in mrna. It was reported that the level of mrna synthesis was not completely decreased by the defect of vrna amplification (Vreede et al, 2004; Kawaguchi et al, 2005; Hara et al, 2006). This was also the case in the plasmid-based replication system (Supplementary Figure 3S-D). The level of mrna synthesis remained unchanged at different levels of vrna synthesis. The amount of vrna responsible for mrna synthesis could be only a small portion of replicated vrna. Note that the viral genome replication in MCM KD cells was not abolished completely. This could be due to the fact that the rest of MCM proteins in KD HeLa cells is enough for the viral genome replication since MCM proteins are highly abundant in a cell (Lei et al, 1996; Mahbubani et al, 1997; Edwards et al, 2002). Then, we used human normal fibroblast WI-38 cells since the amount of MCM2-7 proteins is found to be 10 times less in WI-38 cells than in HeLa cells (Figure 5A, lanes 1 and 3, and see Ishimi et al, 2003). The expression level of the MCM proteins in MCM KD WI-38 cells decreased to 30% of that in control cells (Figure 5A, lanes 3 and 4). The synthesis level of crna in MCM KD WI-38 cells reduced to 40% of that in control cells (Figure 5D, left panel). In contrast, the synthesis level of mrna did not differ between control and MCM KD WI-38 cells (Figure 5D, right panels) The EMBO Journal VOL 26 NO & 2007 European Molecular Biology Organization
6 Figure 5 Involvement of MCM in influenza virus genome replication in infected cells. (A) Expression level of MCM2 and MCM3 in MCM knockdown (KD) cells. HeLa (lanes 1 and 2) and WI-38 cells (lanes 3 and 4) were transfected with random sirna (lanes 1 and 3) or both MCM2 and MCM3 sirnas (lanes 2 and 4). After 60 h post transfection, the amount of MCM2, MCM3, and b-actin proteins were determined by western blotting assays. (B) The level of virus genome replication in MCM KD HeLa cells. At 60 h post transfection of random or MCM sirna, cells were infected with influenza virus at an MOI ¼ 10. At 8 h post infection, total RNAs were purified, and then subjected to real-time quantitative RT PCR with primer sets specific for segment 5 crna (left panel, Po0.01) and NP mrna (right panel, P40.05). The details of the method employed are described in Supplementary data. (C) The level of viral mrna synthesis in MCM KD HeLa cells in the presence of cycloheximide. Control and MCM KD HeLa cells were infected with influenza virus in the presence of 100 mg/ml cycloheximide at an MOI ¼ 10 for 8 h. The real-time quantitative RT PCR assays were carried out with a primer set specific for NP mrna. (D) The level of virus genome replication in MCM KD WI-38 cells. Control and MCM KD WI-38 cells were infected with influenza virus at an MOI ¼ 10 for 8 h. The real-time quantitative RT PCR assays were carried out with primer sets specific for segment 5 crna (left panel) and NP mrna (right panel). b-actin mrna was used as an internal control for the whole procedure. IREF-1/MCM stabilizes RNA polymerase elongation complex As shown in Figures 1B and 2A, it is likely that IREF-1/MCM does not enhance the frequency of the replication initiation, but rather makes a nonproductive viral RNA polymerase to override the step for abortive synthesis. Both eukaryotic and bacterial DNA-dependent RNA polymerases proceed to the productive stage by passing through the abortive transcription stage, during which the polymerase releases short nascent RNA chains. After the synthesis of an approximately 10- nt-long RNA, the RNA synthesizing complex transits into a stable transcription elongation form. This transition from the initiation step, including a complex possibly undergoing abortive synthesis to the stable elongation step, is one of the important steps in the regulation of RNA synthesis by RNA polymerases. To clarify the role of MCM in virus genome replication, we examined the effect of MCM on this transition. The amount of the elongation complex should reflect the stability of the elongation complex. To assess the stability of the elongation complex, we tried to measure the amount of the nascent crna (Figure 6A, upper panel) and vrna template (Figure 6A, lower panel) associated with the RNA polymerase complexes stalled on the vrna template in limited elongation assays by immunoprecipitation using an anti-pb2 antibody. The [a- 32 P]GTP-labeled nascent crnas were detected by autoradiography, whereas the vrna templates were subjected to semiquantitative RT PCR with primers specific for the segment 5 vrna. The level of immunoprecipitated nascent crna was increased by the addition of MCM (Figure 6A, upper panel), whereas exactly the same amounts of the vrna template were immunoprecipitated in the absence and presence of MCM (Figure 6A, lower panel). These results indicate that MCM stabilizes the elongation complex possibly by preventing the release of nascent crna from RNA polymerase complexes. If the nascent crna tends to be dissociated from the elongation complex in the absence of MCM, full-length crna would not be synthesized even after allowing the elongation reaction to restart from limited elongation by the addition of MCM and nucleotides absent in the limited elongation assays. To examine whether MCM acts in the initiation or the elongation process of genome replication, unprimed limited elongation assays were performed to synthesize 2-nt- or 12-nt-long nascent crna from the segment 7 vrna in the absence of either CTP or UTP (Figures 6B and C). After the unprimed limited elongation, elongation reactions were restarted by the addition of either CTP and UTP or UTP, and MCM was also added either before or after the limited elongation. We detected the full-length crna from the limited elongation assays synthesizing 2-nt-long RNA irrespective of the presence or absence of MCM during the limited elongation reaction (Figure 6C, lanes 1 3). In contrast, the full-length crna was synthesized by restarting the limited elongation assay for 12-nt-long RNA in the presence of MCM, but not in the course of the addition of MCM after the limited elongation (Figure 6C, lanes 4 7). The hairpinloop and double-stranded promoter regions, which act as cis-elements essential for the interaction with the viral RNA polymerase, are located between nucleotide positions 1 12 of & 2007 European Molecular Biology Organization The EMBO Journal VOL 26 NO
7 Figure 6 MCM complex stabilizes RNA polymerase elongation complex. (A) Immunoprecipitation of replication elongation complex. After the limited elongation reaction performed in the absence (lanes 1 3 and 7 11) or presence (lanes 4 6 and 12 14) of rmcm, viral polymerase complexes were immunoprecipitated without (lanes 2, 5, 10, and 13) or with (lanes 3, 6, 11, and 14) anti-pb2 antibody. Immunoprecipitated [a- 32 P]GTP-labeled nascent crnas were separated and visualized by autoradiography (upper panel). Immunoprecipitated vrna templates were semiquantitatively analyzed by RT PCR as described in the legend of Figure 4 (lower panel). Inputs (2 (lane 7), 6 (lane 8), and 20% (lanes 9 and 12) equivalents) were also analyzed by RT PCR. (B) The structure of the influenza virus segment 7 vrna promoter. The figure is modified from one in Crow et al (2004). (C) MCM complex functions during transition from initiation to elongation. Limited elongation reactions were performed in the absence of CTP (lanes 1 3) or UTP (lanes 4 7) without (lanes 1, 3, 4, and 6) or with (lanes 2, 5, and 7) rmcm. After 1 h incubation, elongation reactions were restarted by the addition of CTP and UTP for lanes 1 3 and UTP for lanes 4 6. For lanes 3 and 6, rmcm was added at the restart of elongation reaction. (D) MCM functions as a scaffold between nascent crna and viral polymerase complexes. Limited elongation assays were performed with (lanes 1, and 3 5) or without (lane 2) rmcm. The reaction mixture was preincubated for 10 min in the absence (lane 1) or presence of 20 pmol of RNA-AG (5 0 -AGGGGAAAGGAGAAG-3 0, lanes 2 4) and/or 1 nmol RNA-CC (5 0 -AGGGGAAAGGAGACC-3 0, lanes 4 and 5). (E) ATPase dependency of the IREF-1 activity. The cell-free virus genome replication assays were performed in the presence of ATP (lanes 1 and 3) or ATPgS (lanes 2 and 4) without (lanes 1and 2) or with (lanes 3 and 4) rmcm. the 3 0 -terminal region of vrna (Figure 6B). It is possible that the viral RNA polymerase remains at and begins escaping from the promoter under the limited elongation condition synthesizing 2-nt- and 12-nt-long RNAs, respectively. It is quite likely that the RNA polymerase associated with the nascent 12-nt-long RNA synthesized in the presence of MCM can override the abortive destiny. Further, to avoid the abortive RNA synthesis, MCM is required prior to the synthesis of 12-nt-long RNA and possibly after the synthesis of 2-nt-long RNA. The same results were obtained for other segments (data not shown). Altogether, these findings suggest that MCM is involved in the stabilization of the elongation complex during the transition from the initiation to the productive elongation process around the promoter. It is possible that MCM functions as a scaffold protein between nascent crna and vrnp possibly through the PA subunit of the viral RNA polymerase since nascent crna tends to be released from the viral RNA polymerase in the absence of MCM (Figure 6A). To test this hypothesis, we examined whether a 15-nt-long RNA containing ApG at its terminus (RNA-AG) is recruited to the RNA polymerase by MCM (Figure 6D). Limited elongation assays were performed in the presence of 15 nt RNA-AG, the 3 0 -terminal ApG of which could be utilized as the ApG primer. By adding RNA- AG, limited elongation products of 25, 26, 27, 31, and 32 nt were synthesized as expected (Figure 6D, lane 2). MCM stimulated RNA-AG primed RNA synthesis (Figure 6D, lane 3). Furthermore, the effect of MCM was quenched by the addition of RNA-CC, which is 15-mer RNA with the terminal CpC and thus does not serve as a primer for RNA synthesis (Figure 6D, lanes 4 and 5). It could be interpreted that RNA-CC competes with RNA-AG for binding to MCM. These results indicate that RNA-AG is recruited to the viral RNA polymerase by the interaction with MCM. Thus, it is highly possible that MCM stabilizes the replication elongation complex through scaffolding between nascent crna and the viral RNA polymerase to make the polymerase competent for full-length crna synthesis. It should be noted that MCM did not facilitate the unprimed limited elongation reactions (Figure 2A). It is possible that the length of nascent crna in unprimed limited elongation reactions may not be enough for binding with MCM. Biochemical studies have shown that MCM possesses an ATP-dependent DNA helicase activity (You et al, 1999). Therefore, it is speculated that MCM acts as an RNA helicase in virus genome replication. However, full-length crna 4572 The EMBO Journal VOL 26 NO & 2007 European Molecular Biology Organization
8 synthesis occurred in the presence of ATPgS, which is a nonhydrolyzable analog of ATP but can be a substrate for the chain elongation of RNA synthesis (Figure 6E, compare lane 3 with lane 4). Discussion We have identified IREF-1 as a factor that stimulates the cellfree influenza virus genome replication activity. IREF-1 was found to be identical to the MCM complex consisting of MCM2, 3, 4, 5, 6, and 7. By the addition of IREF-1/MCM, the full-length crna was synthesized from each virus genome segment. The RNA synthesis level in the presence of MCM differed from segment to segment (Figure 1B, lanes 2 4). However, the amounts of replicated crnas are almost equal among eight segments in infected cells (Hatada et al, 1989). Therefore, it is possible that another host factor(s) is required for the quantitative control of the virus genome replication. However, this issue is still an open question. Previous reports showed that NP is important for virus genome replication (Shapiro and Krug, 1988; Medcalf et al, 1999). NP is required for elongation of RNA synthesis (Honda et al, 1988). Moreover, the in vitro crna synthesis using infected cell extracts as an enzyme source depends on a supply of non-rnp-associated NP (Shapiro and Krug, 1988). The latter finding may be interpreted as that NP functions in co-transcriptional encapsidation of nascent RNA for prevention of discrete RNA synthesis. Therefore, it is possible that MCM and NP free of vrna act in a similar fashion and cooperatively facilitate the virus genome replication. Further, it was reported that minimal crna synthesis occurs even in the absence of newly synthesized NP (Vreede et al, 2004). Thus, it is speculated that MCM activates and guarantees the virus genome replication from incoming vrnp at immediate early phases of infection, where newly synthesized NP is absent. MCM interacted with the viral RNA polymerase through its contact with PA (Figure 4). Previous genetic analyses suggested that PA participates in the replication process, although its precise role had not been well established (Sugiura et al, 1975; Ritchey and Palese, 1977; Kawaguchi et al, 2005). Recently, it has been reported that the influenza virus vrnp interacts with nucleosomes (Garcia-Robles et al, 2005). Both vrnp and NP that is free of RNA are capable of binding to core histones in vitro. It has also been reported that the viral RNA polymerase interact with the phosphoserine 5 form of the largest subunit of pol II (Engelhardt et al, 2005). MCM also interacts with pol II holoenzyme in addition to DNA replication origins in nucleosomes (Yankulov et al, 1999). Based on these, we tentatively hypothesize that vrnp binds to chromatin where MCM is present while interacting with chromatin proteins such as pol II and recruits MCM through the contact with PA. A functional form of the viral polymerase for the transcription and replication is thought to be a ternary complex consisting of PB1, PB2, and PA (Fodor et al, 2002; Gastaminza et al, 2003). However, the molecular basis of the switching mechanism between transcription and replication is unknown. In infected cells, MCM is involved predominantly in virus genome replication, but not transcription (Figure 5). MCM did not inhibit initiation of capped RNAprimed mrna synthesis in a cell-free RNA synthesis system Figure 7 A proposed model. MCM stabilizes replication elongation complexes through scaffolding between nascent crna and viral RNA polymerase complexes during its transition from initiation to elongation to allow viral RNA polymerase complexes to synthesize full-length crna. (data not shown). We speculate that another host and/or viral factor(s) other than MCM may be required for the switching mechanism of initiation reactions. A universal characteristic feature of RNA polymerases is the regulated conversion from an initiating form that holds nascent RNA weakly to an elongating form that holds RNA tightly during RNA synthesis. This transition occurs during promoter escape, and involves a complex series of molecular transformations in both prokaryotic and eukaryotic DNAdependent RNA polymerases (Zawel et al, 1995; Hsu, 2002). TFIIB and TFIIE, the general transcription factors for the cellular RNA polymerase II (pol II), are released from the elongation complex by the synthesis of an approximately 10- nt-long nascent RNA (Zawel et al, 1995). The nascent RNA forms a 9 bp RNA:DNA hybrid in the elongation complex (Gnatt et al, 2001), which is thought to be a primary stability determinant for the elongation complex (Kireeva et al, 2000). It is speculated that the hybridization between template DNA and nascent RNA chains with lengths less than 9 bp is less effective for the stabilization of the elongating complex. Our results showed that the influenza viral RNA polymerase catalyzes the initiation reaction, but cannot synthesize the full-length crna because the nascent crna was dissociated from the elongation complex in the absence of MCM (Figure 6). In contrast, in the presence of MCM, the elongation complex was stabilized and converted to the form competent for the processive RNA synthesis since MCM may act as a scaffold between nascent crna and the viral RNA polymerase without its helicase activity (Figures 6 and 7). It is possible that the association of nascent RNA with the elongation complex could be stabilized by MCM through its binding to nascent RNA. On this line, it was proposed that PA increases the interaction between PB1 and RNA (Lee et al, 2002). PA is UV-crosslinked to the 5 0 -terminal promoter of vrna (Fodor et al, 1994). Thus, it is also possible that MCM modulates the interaction between the viral polymerase and the promoter through its contact with PA. It was reported that MCM is associated with pol IImediated transcription. Antibodies against MCM2 inhibit pol II transcription in Xenopus oocytes (Yankulov et al, 1999); the interaction between MCM5 and the activation domain of STAT1a is essential for the expression of IFN-g & 2007 European Molecular Biology Organization The EMBO Journal VOL 26 NO
9 responsive genes (Snyder et al, 2005). The finding reported here may provide useful information to further understand the mechanism of cellular transcription regulated by MCM. Materials and methods Cell-free virus genome replication system vrnp was prepared from purified influenza A/PR/8/34 virus as previously described (Shimizu et al, 1994). A cell-free virus genome replication was carried out at 301C for 90 min in a final volume of 25 ml containing 50 mm HEPES-NaOH (ph 7.9), 3 mm MgCl 2, 50 mm KCl, 1.5 mm dithiothreitol, 500 mm each ATP, CTP, and UTP, 25 mm GTP,5mCi of [a- 32 P]GTP (3000 Ci/mmol), 8 U of RNase inhibitor, 25 mg of actinomycin D/ml, and vrnp (50 ng of NP equivalents) in the presence or absence of host factor fraction or purified proteins. RNA products were purified, subjected to 4% PAGE in the presence of 8 M urea, and visualized by autoradiography. For limited elongation assay, RNA synthesis was performed in the absence of UTP, and RNA products were separated through 15% PAGE containing 8 M urea. Supplementary data Supplementary data are available at The EMBO Journal Online ( Acknowledgements We thank F Momose (Kitasato University), Y Ishimi (Ibaraki University), H Nojima (Osaka University), and N Yabuta (Osaka University) for the generous gifts of pcaggs-np Myc (FM), UW31- hmcm4(his)-6, UW31-hMCM2-7(His), and pacab4-hmcm3-5(his) (YI), anti-mcm proteins antibodies (HN and NY). We also thank J Yanagisawa (University of Tsukuba) for his help for MALDI-TOF MS analyses. This research was supported in part by a grant-in-aid from the Ministry of Education, Culture, Sports, Science, and Technology of Japan (to KN) and Research Fellowships of the Japanese Society for the Promotion of Science (JSPS) (to AK). References Chan AY, Vreede FT, Smith M, Engelhardt OG, Fodor E (2006) Influenza virus inhibits RNA polymerase II elongation. Virology 351: Cortez D, Glick G, Elledge SJ (2004) Minichromosome maintenance proteins are direct targets of the ATM and ATR checkpoint kinases. Proc Natl Acad Sci USA 101: Crow M, Deng T, Addley M, Brownlee GG (2004) Mutational analysis of the influenza virus crna promoter and identification of nucleotides critical for replication. J Virol 78: Deng T, Engelhardt OG, Thomas B, Akoulitchev AV, Brownlee GG, Fodor E (2006) Role of ran binding protein 5 in nuclear import and assembly of the influenza virus RNA polymerase complex. J Virol 80: Digard P, Elton D, Bishop K, Medcalf E, Weeds A, Pope B (1999) Modulation of nuclear localization of the influenza virus nucleoprotein through interaction with actin filaments. JVirol73: Edwards MC, Tutter AV, Cvetic C, Gilbert CH, Prokhorova TA, Walter JC (2002) MCM2 7 complexes bind chromatin in a distributed pattern surrounding the origin recognition complex in Xenopus egg extracts. J Biol Chem 277: Engelhardt OG, Fodor E (2006) Functional association between viral and cellular transcription during influenza virus infection. Rev Med Virol 16: Engelhardt OG, Smith M, Fodor E (2005) Association of the influenza A virus RNA-dependent RNA polymerase with cellular RNA polymerase II. J Virol 79: Fodor E, Crow M, Mingay LJ, Deng T, Sharps J, Fechter P, Brownlee GG (2002) A single amino acid mutation in the PA subunit of the influenza virus RNA polymerase inhibits endonucleolytic cleavage of capped RNAs. J Virol 76: Fodor E, Pritlove DC, Brownlee GG (1994) The influenza virus panhandle is involved in the initiation of transcription. J Virol 68: Forsburg SL (2004) Eukaryotic MCM proteins: beyond replication initiation. Microbiol Mol Biol Rev 68: Gabriel G, Abram M, Keiner B, Wagner R, Klenk HD, Stech J (2007) Differential polymerase activity in avian and Mammalian cells determines host range of influenza virus. J Virol 81: Gabriel G, Dauber B, Wolff T, Planz O, Klenk HD, Stech J (2005) The viral polymerase mediates adaptation of an avian influenza virus to a mammalian host. Proc Natl Acad Sci USA 102: Garcia-Robles I, Akarsu H, Muller CW, Ruigrok RW, Baudin F (2005) Interaction of influenza virus proteins with nucleosomes. Virology 332: Gastaminza P, Perales B, Falcon AM, Ortin J (2003) Mutations in the N-terminal region of influenza virus PB2 protein affect virus RNA replication but not transcription. J Virol 77: Gnatt AL, Cramer P, Fu J, Bushnell DA, Kornberg RD (2001) Structural basis of transcription: an RNA polymerase II elongation complex at 3.3 A resolution. Science 292: Hara K, Schmidt FI, Crow M, Brownlee GG (2006) Amino acid residues in the N-terminal region of the PA subunit of influenza A virus RNA polymerase play a critical role in protein stability, endonuclease activity, cap binding, and virion RNA promoter binding. J Virol 80: Hatada E, Hasegawa M, Mukaigawa J, Shimizu K, Fukuda R (1989) Control of influenza virus gene expression: quantitative analysis of each viral RNA species in infected cells. J Biochem (Tokyo) 105: Honda A, Mizumoto K, Ishihama A (1986) RNA polymerase of influenza virus. Dinucleotide-primed initiation of transcription at specific positions on viral RNA. J Biol Chem 261: Honda A, Ueda K, Nagata K, Ishihama A (1988) RNA polymerase of influenza virus: role of NP in RNA chain elongation. J Biochem (Tokyo) 104: Hsu LM (2002) Promoter clearance and escape in prokaryotes. Biochim Biophys Acta 1577: Huarte M, Sanz-Ezquerro JJ, Roncal F, Ortin J, Nieto A (2001) PA subunit from influenza virus polymerase complex interacts with a cellular protein with homology to a family of transcriptional activators. J Virol 75: Ishimi Y, Okayasu I, Kato C, Kwon HJ, Kimura H, Yamada K, Song SY (2003) Enhanced expression of Mcm proteins in cancer cells derived from uterine cervix. Eur J Biochem 270: Kawaguchi A, Naito T, Nagata K (2005) Involvement of influenza virus PA subunit in assembly of functional RNA polymerase complexes. J Virol 79: Kireeva ML, Komissarova N, Waugh DS, Kashlev M (2000) The 8-nucleotide-long RNA:DNA hybrid is a primary stability determinant of the RNA polymerase II elongation complex. J Biol Chem 275: Lee MT, Bishop K, Medcalf L, Elton D, Digard P, Tiley L (2002) Definition of the minimal viral components required for the initiation of unprimed RNA synthesis by influenza virus RNA polymerase. Nucleic Acids Res 30: Lei M, Kawasaki Y, Tye BK (1996) Physical interactions among Mcm proteins and effects of Mcm dosage on DNA replication in Saccharomyces cerevisiae. Mol Cell Biol 16: Mahbubani HM, Chong JP, Chevalier S, Thommes P, Blow JJ (1997) Cell cycle regulation of the replication licensing system: involvement of a Cdk-dependent inhibitor. J Cell Biol 136: Maine GT, Sinha P, Tye BK (1984) Mutants of S. cerevisiae defective in the maintenance of minichromosomes. Genetics 106: Medcalf L, Poole E, Elton D, Digard P (1999) Temperature-sensitive lesions in two influenza A viruses defective for replicative transcription disrupt RNA binding by the nucleoprotein. J Virol 73: Momose F, Basler CF, O Neill RE, Iwamatsu A, Palese P, Nagata K (2001) Cellular splicing factor RAF-2p48/NPI-5/BAT1/UAP56 interacts with the influenza virus nucleoprotein and enhances viral RNA synthesis. J Virol 75: Momose F, Naito T, Yano K, Sugimoto S, Morikawa Y, Nagata K (2002) Identification of Hsp90 as a stimulatory host factor 4574 The EMBO Journal VOL 26 NO & 2007 European Molecular Biology Organization
The accumulation of influenza A virus segment 7 spliced mrnas is regulated by the NS1 protein
 Journal of General Virology (2012), 93, 113 118 DOI 10.1099/vir.0.035485-0 Short Communication Correspondence Ervin Fodor ervin.fodor@path.ox.ac.uk Received 24 June 2011 Accepted 12 September 2011 The
Journal of General Virology (2012), 93, 113 118 DOI 10.1099/vir.0.035485-0 Short Communication Correspondence Ervin Fodor ervin.fodor@path.ox.ac.uk Received 24 June 2011 Accepted 12 September 2011 The
The RNA Polymerase of Influenza Virus, Bound to the 5 End of Virion RNA, Acts in cis To Polyadenylate mrna
 JOURNAL OF VIROLOGY, Oct. 1998, p. 8214 8219 Vol. 72, No. 10 0022-538X/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. The RNA Polymerase of Influenza Virus, Bound to
JOURNAL OF VIROLOGY, Oct. 1998, p. 8214 8219 Vol. 72, No. 10 0022-538X/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. The RNA Polymerase of Influenza Virus, Bound to
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
influenza virus RNA synthesis Shirokane, Minato-ku, Tokyo , Japan Asahi-machi, Machida-shi, Tokyo , Japan
 JBC Papers in Press. Published on September 10, 2002 as Manuscript M206822200 Title: Identification of Hsp90 as a stimulatory host factor involved in influenza virus RNA synthesis Authors: Fumitaka Momose
JBC Papers in Press. Published on September 10, 2002 as Manuscript M206822200 Title: Identification of Hsp90 as a stimulatory host factor involved in influenza virus RNA synthesis Authors: Fumitaka Momose
Gag. Gag-Gag. TSG101 Nedd4. Two-Hybrid HIV Gag. Two-Hybrid. Two-Hybrid
 CytoTrap HIV Gag AnnexinA2 AUP1 AUP1 HIV AUP1 N Gag CytoTrap Gag-Gag TSG101 Nedd4 Two-Hybrid Two-Hybrid HIVGag Gag Two-Hybrid HIV Gag Gag HIV Two-Hybrid CytoTrap cdna Gag Gag CytoTrap Gag CytoTrap Gag-Gag
CytoTrap HIV Gag AnnexinA2 AUP1 AUP1 HIV AUP1 N Gag CytoTrap Gag-Gag TSG101 Nedd4 Two-Hybrid Two-Hybrid HIVGag Gag Two-Hybrid HIV Gag Gag HIV Two-Hybrid CytoTrap cdna Gag Gag CytoTrap Gag CytoTrap Gag-Gag
Different De Novo Initiation Strategies Are Used by Influenza Virus RNA Polymerase on Its crna and Viral RNA Promoters during Viral RNA Replication
 JOURNAL OF VIROLOGY, Mar. 2006, p. 2337 2348 Vol. 80, No. 5 0022-538X/06/$08.00 0 doi:10.1128/jvi.80.5.2337 2348.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Different De
JOURNAL OF VIROLOGY, Mar. 2006, p. 2337 2348 Vol. 80, No. 5 0022-538X/06/$08.00 0 doi:10.1128/jvi.80.5.2337 2348.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Different De
Mutational Analysis of the Influenza Virus crna Promoter and Identification of Nucleotides Critical for Replication
 JOURNAL OF VIROLOGY, June 2004, p. 6263 6270 Vol. 78, No. 12 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.12.6263 6270.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Mutational
JOURNAL OF VIROLOGY, June 2004, p. 6263 6270 Vol. 78, No. 12 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.12.6263 6270.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Mutational
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Supplementary Fig. S1. Schematic diagram of minigenome segments.
 open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
Influenza Virion-Derived Viral Ribonucleoproteins Synthesize both mrna and crna In Vitro
 JOURNAL OF VIROLOGY, Mar. 2007, p. 2196 2204 Vol. 81, No. 5 0022-538X/07/$08.00 0 doi:10.1128/jvi.02187-06 Copyright 2007, American Society for Microbiology. All Rights Reserved. Influenza Virion-Derived
JOURNAL OF VIROLOGY, Mar. 2007, p. 2196 2204 Vol. 81, No. 5 0022-538X/07/$08.00 0 doi:10.1128/jvi.02187-06 Copyright 2007, American Society for Microbiology. All Rights Reserved. Influenza Virion-Derived
L I F E S C I E N C E S
 1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 OCTOBER 31, 2006
1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 OCTOBER 31, 2006
1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics
 1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics to gain mechanistic insight! 4. Return to step 2, as
1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics to gain mechanistic insight! 4. Return to step 2, as
Life Sciences 1A Midterm Exam 2. November 13, 2006
 Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Materials and Methods , The two-hybrid principle.
 The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
Differential Roles of vrna and crna in Functional Modulation of the Influenza Virus RNA Polymerase *
 JBC Papers in Press. Published on May 23, 2001 as Manuscript M102856200 Differential Roles of vrna and crna in Functional Modulation of the Influenza Virus RNA Polymerase * Ayae Honda,1,2,3 Atsushi Endo,4
JBC Papers in Press. Published on May 23, 2001 as Manuscript M102856200 Differential Roles of vrna and crna in Functional Modulation of the Influenza Virus RNA Polymerase * Ayae Honda,1,2,3 Atsushi Endo,4
hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide gel electrophoresis/genetics)
 Proc. Natl. Acad. Sci. USA Vol. 73, No. 6, pp. 242-246, June 976 Microbiology Mapping of the influenza virus genome: Identification of the hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide
Proc. Natl. Acad. Sci. USA Vol. 73, No. 6, pp. 242-246, June 976 Microbiology Mapping of the influenza virus genome: Identification of the hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide
Structural vs. nonstructural proteins
 Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
October 26, Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
RECAP (1)! In eukaryotes, large primary transcripts are processed to smaller, mature mrnas.! What was first evidence for this precursorproduct
 RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
Supplementary Information
 Supplementary Information HBV maintains electrostatic homeostasis by modulating negative charges from phosphoserine and encapsidated nucleic acids Authors: Pei-Yi Su 1,2,3, Ching-Jen Yang 2, Tien-Hua Chu
Supplementary Information HBV maintains electrostatic homeostasis by modulating negative charges from phosphoserine and encapsidated nucleic acids Authors: Pei-Yi Su 1,2,3, Ching-Jen Yang 2, Tien-Hua Chu
Introduction retroposon
 17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
Ubiquitination and deubiquitination of NP protein regulates influenza A virus RNA replication
 Manuscript EMBO-2010-74756 Ubiquitination and deubiquitination of NP protein regulates influenza A virus RNA replication Tsai-Ling Liao, Chung-Yi Wu, Wen-Chi Su, King-Song Jeng and Michael Lai Corresponding
Manuscript EMBO-2010-74756 Ubiquitination and deubiquitination of NP protein regulates influenza A virus RNA replication Tsai-Ling Liao, Chung-Yi Wu, Wen-Chi Su, King-Song Jeng and Michael Lai Corresponding
Temperature-Sensitive Lesions in Two Influenza A Viruses Defective for Replicative Transcription Disrupt RNA Binding by the Nucleoprotein
 JOURNAL OF VIROLOGY, Sept. 1999, p. 7349 7356 Vol. 73, No. 9 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Temperature-Sensitive Lesions in Two Influenza
JOURNAL OF VIROLOGY, Sept. 1999, p. 7349 7356 Vol. 73, No. 9 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Temperature-Sensitive Lesions in Two Influenza
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature14008 Supplementary Figure 1. Sequence alignment of A/little yellow-shouldered bat/guatemala/060/2010 (H17N10) polymerase with that of human strain A/Victoria/3/75(H3N2). The secondary
doi:10.1038/nature14008 Supplementary Figure 1. Sequence alignment of A/little yellow-shouldered bat/guatemala/060/2010 (H17N10) polymerase with that of human strain A/Victoria/3/75(H3N2). The secondary
Cellular cap-binding proteins associate with influenza virus mrnas
 Journal of General Virology (2011), 92, 1627 1634 DOI 10.1099/vir.0.029231-0 Cellular cap-binding proteins associate with influenza virus mrnas Katja Bier, Ashley York and Ervin Fodor Correspondence Ervin
Journal of General Virology (2011), 92, 1627 1634 DOI 10.1099/vir.0.029231-0 Cellular cap-binding proteins associate with influenza virus mrnas Katja Bier, Ashley York and Ervin Fodor Correspondence Ervin
NS2/NEP protein regulates transcription and replication of the influenza virus RNA genome
 Journal of General Virology (2009), 90, 1398 1407 DOI 10.1099/vir.0.009639-0 NS2/NEP protein regulates transcription and replication of the influenza virus RNA genome Nicole C. Robb,3 Matt Smith,34 Frank
Journal of General Virology (2009), 90, 1398 1407 DOI 10.1099/vir.0.009639-0 NS2/NEP protein regulates transcription and replication of the influenza virus RNA genome Nicole C. Robb,3 Matt Smith,34 Frank
The U1 snrnp Base Pairs with the 5 Splice Site within a Penta-snRNP Complex
 MOLECULAR AND CELLULAR BIOLOGY, May 2003, p. 3442 3455 Vol. 23, No. 10 0270-7306/03/$08.00 0 DOI: 10.1128/MCB.23.10.3442 3455.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
MOLECULAR AND CELLULAR BIOLOGY, May 2003, p. 3442 3455 Vol. 23, No. 10 0270-7306/03/$08.00 0 DOI: 10.1128/MCB.23.10.3442 3455.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
Virology 396 (2010) Contents lists available at ScienceDirect. Virology. journal homepage:
 Virology 396 (2010) 125 134 Contents lists available at ScienceDirect Virology journal homepage: www.elsevier.com/locate/yviro Mechanisms and functional implications of the degradation of host RNA polymerase
Virology 396 (2010) 125 134 Contents lists available at ScienceDirect Virology journal homepage: www.elsevier.com/locate/yviro Mechanisms and functional implications of the degradation of host RNA polymerase
Increased amounts of the influenza virus nucleoprotein do not promote higher levels of viral genome replication
 Journal of General Virology (2004), 85, 3689 3698 DOI 10.1099/vir.0.80518-0 Increased amounts of the influenza virus nucleoprotein do not promote higher levels of viral genome replication Anne E. Mullin,3
Journal of General Virology (2004), 85, 3689 3698 DOI 10.1099/vir.0.80518-0 Increased amounts of the influenza virus nucleoprotein do not promote higher levels of viral genome replication Anne E. Mullin,3
Interaction of the Influenza A Virus Nucleocapsid Protein with the Viral RNA Polymerase Potentiates Unprimed Viral RNA Replication
 JOURNAL OF VIROLOGY, Jan. 2009, p. 29 36 Vol. 83, No. 1 0022-538X/09/$08.00 0 doi:10.1128/jvi.02293-07 Copyright 2009, American Society for Microbiology. All Rights Reserved. Interaction of the Influenza
JOURNAL OF VIROLOGY, Jan. 2009, p. 29 36 Vol. 83, No. 1 0022-538X/09/$08.00 0 doi:10.1128/jvi.02293-07 Copyright 2009, American Society for Microbiology. All Rights Reserved. Interaction of the Influenza
Lecture 2: Virology. I. Background
 Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Transcription and RNA processing
 Transcription and RNA processing Lecture 7 Biology 3310/4310 Virology Spring 2018 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at the
Transcription and RNA processing Lecture 7 Biology 3310/4310 Virology Spring 2018 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at the
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Section 6. Junaid Malek, M.D.
 Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
RNA Processing in Eukaryotes *
 OpenStax-CNX module: m44532 1 RNA Processing in Eukaryotes * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you
OpenStax-CNX module: m44532 1 RNA Processing in Eukaryotes * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004
 Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Influenza viruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
Biochemical Characterization of Enzyme Fidelity of Influenza A Virus RNA Polymerase Complex
 Biochemical Characterization of Enzyme Fidelity of Influenza A Virus RNA Polymerase Complex Shilpa Aggarwal, Birgit Bradel-Tretheway, Toru Takimoto, Stephen Dewhurst, Baek Kim* Department of Microbiology
Biochemical Characterization of Enzyme Fidelity of Influenza A Virus RNA Polymerase Complex Shilpa Aggarwal, Birgit Bradel-Tretheway, Toru Takimoto, Stephen Dewhurst, Baek Kim* Department of Microbiology
A Novel in Vitro Replication System for Dengue Virus
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 274, No. 47, Issue of November 19, pp. 33714 33722, 1999 1999 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. A Novel in
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 274, No. 47, Issue of November 19, pp. 33714 33722, 1999 1999 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. A Novel in
Regulation and Fidelity: A study of RNA and DNA polymerases
 University of Colorado, Boulder CU Scholar Chemistry & Biochemistry Graduate Theses & Dissertations Chemistry & Biochemistry Spring 1-1-2011 Regulation and Fidelity: A study of RNA and DNA polymerases
University of Colorado, Boulder CU Scholar Chemistry & Biochemistry Graduate Theses & Dissertations Chemistry & Biochemistry Spring 1-1-2011 Regulation and Fidelity: A study of RNA and DNA polymerases
The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein
 THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
Reverse Genetics of RNA Viruses
 Reverse Genetics of RNA Viruses Reverse Genetics (RG) he creation of a virus with a fulllength copy of the viral genome he most powerful tool in modern virology RG of RNA viruses Generation or recovery
Reverse Genetics of RNA Viruses Reverse Genetics (RG) he creation of a virus with a fulllength copy of the viral genome he most powerful tool in modern virology RG of RNA viruses Generation or recovery
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Virus and Prokaryotic Gene Regulation - 1
 Virus and Prokaryotic Gene Regulation - 1 We have discussed the molecular structure of DNA and its function in DNA duplication and in transcription and protein synthesis. We now turn to how cells regulate
Virus and Prokaryotic Gene Regulation - 1 We have discussed the molecular structure of DNA and its function in DNA duplication and in transcription and protein synthesis. We now turn to how cells regulate
Reverse transcription and integration
 Reverse transcription and integration Lecture 9 Biology 3310/4310 Virology Spring 2018 One can t believe impossible things, said Alice. I dare say you haven t had much practice, said the Queen. Why, sometimes
Reverse transcription and integration Lecture 9 Biology 3310/4310 Virology Spring 2018 One can t believe impossible things, said Alice. I dare say you haven t had much practice, said the Queen. Why, sometimes
Phenomena first observed in petunia
 Vectors for RNAi Phenomena first observed in petunia Attempted to overexpress chalone synthase (anthrocyanin pigment gene) in petunia. (trying to darken flower color) Caused the loss of pigment. Bill Douherty
Vectors for RNAi Phenomena first observed in petunia Attempted to overexpress chalone synthase (anthrocyanin pigment gene) in petunia. (trying to darken flower color) Caused the loss of pigment. Bill Douherty
reads observed in trnas from the analysis of RNAs carrying a 5 -OH ends isolated from cells induced to express
 Supplementary Figure 1. VapC-mt4 cleaves trna Ala2 in E. coli. Histograms representing the fold change in reads observed in trnas from the analysis of RNAs carrying a 5 -OH ends isolated from cells induced
Supplementary Figure 1. VapC-mt4 cleaves trna Ala2 in E. coli. Histograms representing the fold change in reads observed in trnas from the analysis of RNAs carrying a 5 -OH ends isolated from cells induced
SUPPLEMENTARY INFORMATION
 Supplementary Discussion The cell cycle machinery and the DNA damage response network are highly interconnected and co-regulated in assuring faithful duplication and partition of genetic materials into
Supplementary Discussion The cell cycle machinery and the DNA damage response network are highly interconnected and co-regulated in assuring faithful duplication and partition of genetic materials into
Supplementary information
 Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Plasmid-Driven Formation of Influenza Virus-Like Particles
 JOURNAL OF VIROLOGY, Jan. 2000, p. 547 551 Vol. 74, No. 1 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Plasmid-Driven Formation of Influenza Virus-Like
JOURNAL OF VIROLOGY, Jan. 2000, p. 547 551 Vol. 74, No. 1 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Plasmid-Driven Formation of Influenza Virus-Like
TRANSCRIPTION. DNA à mrna
 TRANSCRIPTION DNA à mrna Central Dogma Animation DNA: The Secret of Life (from PBS) http://www.youtube.com/watch? v=41_ne5ms2ls&list=pl2b2bd56e908da696&index=3 Transcription http://highered.mcgraw-hill.com/sites/0072507470/student_view0/
TRANSCRIPTION DNA à mrna Central Dogma Animation DNA: The Secret of Life (from PBS) http://www.youtube.com/watch? v=41_ne5ms2ls&list=pl2b2bd56e908da696&index=3 Transcription http://highered.mcgraw-hill.com/sites/0072507470/student_view0/
ISG15 sirna # Ctrl sirna+ifn+wt. Virus titer (Pfu/ml) hours post infection. d USP18 sirna #2 IFN
 a ISG15 sirna Ctrl #1 #2 ISG15 conjugates IB: ISG15 Virus titer (Pfu/ml) b ISG15 sirna #2 10 7 Ctrl sirna+ifn+wt Ctrl sirna+ifn+67 10 6 ISG15 sirna+ifn+wt ISG15 sirna+ifn+67 10 5 10 4 10 3 Free ISG15 10
a ISG15 sirna Ctrl #1 #2 ISG15 conjugates IB: ISG15 Virus titer (Pfu/ml) b ISG15 sirna #2 10 7 Ctrl sirna+ifn+wt Ctrl sirna+ifn+67 10 6 ISG15 sirna+ifn+wt ISG15 sirna+ifn+67 10 5 10 4 10 3 Free ISG15 10
PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland
 AD Award Number: W81XWH-05-1-0494 TITLE: Tumor Suppression by BRCA-1: A Critical Role at DNA Replication Forks PRINCIPAL INVESTIGATOR: Jean Gautier, Ph.D. CONTRACTING ORGANIZATION: Columbia University
AD Award Number: W81XWH-05-1-0494 TITLE: Tumor Suppression by BRCA-1: A Critical Role at DNA Replication Forks PRINCIPAL INVESTIGATOR: Jean Gautier, Ph.D. CONTRACTING ORGANIZATION: Columbia University
Last time we talked about the few steps in viral replication cycle and the un-coating stage:
 Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Mutagenic Analysis of the 5 Arm of the Influenza A Virus Virion RNA Promoter Defines the Sequence Requirements for Endonuclease Activity
 JOURNAL OF VIROLOGY, Jan. 2001, p. 134 142 Vol. 75, No. 1 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.1.134 142.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Mutagenic Analysis
JOURNAL OF VIROLOGY, Jan. 2001, p. 134 142 Vol. 75, No. 1 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.1.134 142.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Mutagenic Analysis
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk
 Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Tel: ; Fax: ;
 Tel.: +98 216 696 9291; Fax: +98 216 696 9291; E-mail: mrasadeghi@pasteur.ac.ir Tel: +98 916 113 7679; Fax: +98 613 333 6380; E-mail: abakhshi_e@ajums.ac.ir A Soluble Chromatin-bound MOI 0 1 5 0 1 5 HDAC2
Tel.: +98 216 696 9291; Fax: +98 216 696 9291; E-mail: mrasadeghi@pasteur.ac.ir Tel: +98 916 113 7679; Fax: +98 613 333 6380; E-mail: abakhshi_e@ajums.ac.ir A Soluble Chromatin-bound MOI 0 1 5 0 1 5 HDAC2
Transcription and RNA processing
 Transcription and RNA processing Lecture 7 Biology W3310/4310 Virology Spring 2016 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at
Transcription and RNA processing Lecture 7 Biology W3310/4310 Virology Spring 2016 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Recombinant Protein Expression Retroviral system
 Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
 http://nmhm.washingtondc.museum/collections/archives/agalleries/1918flu/ncp1603.jpg 1 https://assets-production-webvanta-com.s3-us-west- 2 2.amazonaws.com/000000/47/62/original/images/img_109_influenza/Spanish_flu_death_chart.jpg
http://nmhm.washingtondc.museum/collections/archives/agalleries/1918flu/ncp1603.jpg 1 https://assets-production-webvanta-com.s3-us-west- 2 2.amazonaws.com/000000/47/62/original/images/img_109_influenza/Spanish_flu_death_chart.jpg
Regulators of Cell Cycle Progression
 Regulators of Cell Cycle Progression Studies of Cdk s and cyclins in genetically modified mice reveal a high level of plasticity, allowing different cyclins and Cdk s to compensate for the loss of one
Regulators of Cell Cycle Progression Studies of Cdk s and cyclins in genetically modified mice reveal a high level of plasticity, allowing different cyclins and Cdk s to compensate for the loss of one
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer
 Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
7.012 Problem Set 6 Solutions
 Name Section 7.012 Problem Set 6 Solutions Question 1 The viral family Orthomyxoviridae contains the influenza A, B and C viruses. These viruses have a (-)ss RNA genome surrounded by a capsid composed
Name Section 7.012 Problem Set 6 Solutions Question 1 The viral family Orthomyxoviridae contains the influenza A, B and C viruses. These viruses have a (-)ss RNA genome surrounded by a capsid composed
Signal Transduction Cascades
 Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Figure mouse globin mrna PRECURSOR RNA hybridized to cloned gene (genomic). mouse globin MATURE mrna hybridized to cloned gene (genomic).
 Splicing Figure 14.3 mouse globin mrna PRECURSOR RNA hybridized to cloned gene (genomic). mouse globin MATURE mrna hybridized to cloned gene (genomic). mrna Splicing rrna and trna are also sometimes spliced;
Splicing Figure 14.3 mouse globin mrna PRECURSOR RNA hybridized to cloned gene (genomic). mouse globin MATURE mrna hybridized to cloned gene (genomic). mrna Splicing rrna and trna are also sometimes spliced;
RECAP (1)! In eukaryotes, large primary transcripts are processed to smaller, mature mrnas.! What was first evidence for this precursorproduct
 RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
Native Replication Intermediates of the Yeast 20 S RNA Virus Have a Single-stranded RNA Backbone*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 280, No. 8, Issue of February 25, pp. 7398 7406, 2005 2005 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Native Replication
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 280, No. 8, Issue of February 25, pp. 7398 7406, 2005 2005 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Native Replication
Cellular Human CLE/C14orf166 Protein Interacts with Influenza Virus Polymerase and Is Required for Viral Replication
 JOURNAL OF VIROLOGY, Nov. 2011, p. 12062 12066 Vol. 85, No. 22 0022-538X/11/$12.00 doi:10.1128/jvi.00684-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. Cellular Human CLE/C14orf166
JOURNAL OF VIROLOGY, Nov. 2011, p. 12062 12066 Vol. 85, No. 22 0022-538X/11/$12.00 doi:10.1128/jvi.00684-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. Cellular Human CLE/C14orf166
Mutational Analyses of the Influenza A Virus Polymerase Subunit PA Reveal Distinct Functions Related and Unrelated to RNA Polymerase Activity
 Mutational Analyses of the Influenza A Virus Polymerase Subunit PA Reveal Distinct Functions Related and Unrelated to RNA Polymerase Activity Yuhong Liang, Emory University Shamika Danzy, Emory University
Mutational Analyses of the Influenza A Virus Polymerase Subunit PA Reveal Distinct Functions Related and Unrelated to RNA Polymerase Activity Yuhong Liang, Emory University Shamika Danzy, Emory University
CHARACTERIZING THE ROLE OF N TERMINUS OF INFLUENZA A NUCLEOPROTEIN FOR LOCATION AND VIRAL RNP ACTIVITY
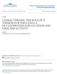 California State University, San Bernardino CSUSB ScholarWorks Electronic Theses, Projects, and Dissertations Office of Graduate Studies 6-2018 CHARACTERIZING THE ROLE OF N TERMINUS OF INFLUENZA A NUCLEOPROTEIN
California State University, San Bernardino CSUSB ScholarWorks Electronic Theses, Projects, and Dissertations Office of Graduate Studies 6-2018 CHARACTERIZING THE ROLE OF N TERMINUS OF INFLUENZA A NUCLEOPROTEIN
number Done by Corrected by Doctor Ashraf
 number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
LESSON 4.4 WORKBOOK. How viruses make us sick: Viral Replication
 DEFINITIONS OF TERMS Eukaryotic: Non-bacterial cell type (bacteria are prokaryotes).. LESSON 4.4 WORKBOOK How viruses make us sick: Viral Replication This lesson extends the principles we learned in Unit
DEFINITIONS OF TERMS Eukaryotic: Non-bacterial cell type (bacteria are prokaryotes).. LESSON 4.4 WORKBOOK How viruses make us sick: Viral Replication This lesson extends the principles we learned in Unit
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay
 Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Genetics. Instructor: Dr. Jihad Abdallah Transcription of DNA
 Genetics Instructor: Dr. Jihad Abdallah Transcription of DNA 1 3.4 A 2 Expression of Genetic information DNA Double stranded In the nucleus Transcription mrna Single stranded Translation In the cytoplasm
Genetics Instructor: Dr. Jihad Abdallah Transcription of DNA 1 3.4 A 2 Expression of Genetic information DNA Double stranded In the nucleus Transcription mrna Single stranded Translation In the cytoplasm
p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO
 Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Segment-specific and common nucleotide sequences in the
 Proc. Nati. Acad. Sci. USA Vol. 84, pp. 2703-2707, May 1987 Biochemistry Segment-specific and common nucleotide sequences in the noncoding regions of influenza B virus genome RNAs (viral transcription/viral
Proc. Nati. Acad. Sci. USA Vol. 84, pp. 2703-2707, May 1987 Biochemistry Segment-specific and common nucleotide sequences in the noncoding regions of influenza B virus genome RNAs (viral transcription/viral
Virology Journal. Open Access. Abstract. BioMed Central
 Virology Journal BioMed Central Research Stimulation of poliovirus RNA synthesis and virus maturation in a HeLa cell-free in vitro translation-rna replication system by viral protein 3CD pro David Franco
Virology Journal BioMed Central Research Stimulation of poliovirus RNA synthesis and virus maturation in a HeLa cell-free in vitro translation-rna replication system by viral protein 3CD pro David Franco
Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression
 Supplementary Figure 1 Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression. Quantitative real-time PCR of indicated mrnas in DCs stimulated with TLR2-Dectin-1 agonist zymosan
Supplementary Figure 1 Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression. Quantitative real-time PCR of indicated mrnas in DCs stimulated with TLR2-Dectin-1 agonist zymosan
The reovirus genome comprises 10 segments of doublestranded
 Reovirus reverse genetics: Incorporation of the CAT gene into the reovirus genome Michael R. Roner* and Wolfgang K. Joklik *Department of Biological Sciences, Center for Molecular Biology and Biotechnology,
Reovirus reverse genetics: Incorporation of the CAT gene into the reovirus genome Michael R. Roner* and Wolfgang K. Joklik *Department of Biological Sciences, Center for Molecular Biology and Biotechnology,
Unidirectional RNA polymerase I polymerase II transcription system for the generation of influenza A virus from eight plasmids
 Journal of General Virology (2000), 81, 2843 2847. Printed in Great Britain... SHORT COMMUNICATION Unidirectional RNA polymerase I polymerase II transcription system for the generation of influenza A virus
Journal of General Virology (2000), 81, 2843 2847. Printed in Great Britain... SHORT COMMUNICATION Unidirectional RNA polymerase I polymerase II transcription system for the generation of influenza A virus
Coronaviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
numbe r Done by Corrected by Doctor
 numbe r 5 Done by Mustafa Khader Corrected by Mahdi Sharawi Doctor Ashraf Khasawneh Viral Replication Mechanisms: (Protein Synthesis) 1. Monocistronic Method: All human cells practice the monocistronic
numbe r 5 Done by Mustafa Khader Corrected by Mahdi Sharawi Doctor Ashraf Khasawneh Viral Replication Mechanisms: (Protein Synthesis) 1. Monocistronic Method: All human cells practice the monocistronic
The influenza virus nucleoprotein: a multifunctional RNA-binding protein pivotal to virus replication
 Journal of General Virology (2002), 83, 723 734. Printed in Great Britain... The influenza virus nucleoprotein: a multifunctional RNA-binding protein pivotal to virus replication Agustı n Portela 1 and
Journal of General Virology (2002), 83, 723 734. Printed in Great Britain... The influenza virus nucleoprotein: a multifunctional RNA-binding protein pivotal to virus replication Agustı n Portela 1 and
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins.
 Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Reoviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Reoviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=13), diameter 60-85 nm Capsid consists of two or three concentric protein
Reoviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=13), diameter 60-85 nm Capsid consists of two or three concentric protein
Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP
 Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP 1 Learning Objectives Recognize hazards associated with viral vectors in research and animal
Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP 1 Learning Objectives Recognize hazards associated with viral vectors in research and animal
Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS
 Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS nucleotide sequences (a, b) or amino acid sequences (c) from
Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS nucleotide sequences (a, b) or amino acid sequences (c) from
Supplemental Information
 Supplemental Information Tobacco-specific Carcinogen Induces DNA Methyltransferases 1 Accumulation through AKT/GSK3β/βTrCP/hnRNP-U in Mice and Lung Cancer patients Ruo-Kai Lin, 1 Yi-Shuan Hsieh, 2 Pinpin
Supplemental Information Tobacco-specific Carcinogen Induces DNA Methyltransferases 1 Accumulation through AKT/GSK3β/βTrCP/hnRNP-U in Mice and Lung Cancer patients Ruo-Kai Lin, 1 Yi-Shuan Hsieh, 2 Pinpin
7.014 Problem Set 7 Solutions
 MIT Department of Biology 7.014 Introductory Biology, Spring 2005 7.014 Problem Set 7 Solutions Question 1 Part A Antigen binding site Antigen binding site Variable region Light chain Light chain Variable
MIT Department of Biology 7.014 Introductory Biology, Spring 2005 7.014 Problem Set 7 Solutions Question 1 Part A Antigen binding site Antigen binding site Variable region Light chain Light chain Variable
Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition
 Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition Andrew J Massey Supplementary Information Supplementary Figure S1. Related to Fig. 1. (a) HT29 or U2OS cells
Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition Andrew J Massey Supplementary Information Supplementary Figure S1. Related to Fig. 1. (a) HT29 or U2OS cells
SUPPLEMENTAL DATA RESULTS
 SUPPLEMENTAL DATA RESULTS Ded1-mediated ribosomal scanning is less leaky than scanning promoted by eifs 4A/4B/4F. The efficiency of leaky scanning in the presence of Ded1 or of eifs 4A/4B/4F was investigated
SUPPLEMENTAL DATA RESULTS Ded1-mediated ribosomal scanning is less leaky than scanning promoted by eifs 4A/4B/4F. The efficiency of leaky scanning in the presence of Ded1 or of eifs 4A/4B/4F was investigated
Role of Ran Binding Protein 5 in Nuclear Import and Assembly of the Influenza Virus RNA Polymerase Complex
 JOURNAL OF VIROLOGY, Dec. 2006, p. 11911 11919 Vol. 80, No. 24 0022-538X/06/$08.00 0 doi:10.1128/jvi.01565-06 Copyright 2006, American Society for Microbiology. All Rights Reserved. Role of Ran Binding
JOURNAL OF VIROLOGY, Dec. 2006, p. 11911 11919 Vol. 80, No. 24 0022-538X/06/$08.00 0 doi:10.1128/jvi.01565-06 Copyright 2006, American Society for Microbiology. All Rights Reserved. Role of Ran Binding
Protein Synthesis
 Protein Synthesis 10.6-10.16 Objectives - To explain the central dogma - To understand the steps of transcription and translation in order to explain how our genes create proteins necessary for survival.
Protein Synthesis 10.6-10.16 Objectives - To explain the central dogma - To understand the steps of transcription and translation in order to explain how our genes create proteins necessary for survival.
Fayth K. Yoshimura, Ph.D. September 7, of 7 RETROVIRUSES. 2. HTLV-II causes hairy T-cell leukemia
 1 of 7 I. Diseases Caused by Retroviruses RETROVIRUSES A. Human retroviruses that cause cancers 1. HTLV-I causes adult T-cell leukemia and tropical spastic paraparesis 2. HTLV-II causes hairy T-cell leukemia
1 of 7 I. Diseases Caused by Retroviruses RETROVIRUSES A. Human retroviruses that cause cancers 1. HTLV-I causes adult T-cell leukemia and tropical spastic paraparesis 2. HTLV-II causes hairy T-cell leukemia
Chapter 6- An Introduction to Viruses*
 Chapter 6- An Introduction to Viruses* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. 6.1 Overview of Viruses
Chapter 6- An Introduction to Viruses* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. 6.1 Overview of Viruses
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
