cape: A package for the combined analysis of epistasis and pleiotropy
|
|
|
- James Pope
- 5 years ago
- Views:
Transcription
1 cape: A package for the combined analysis of epistasis and pleiotropy Anna L. Tyler 1, Wei Lu 1,2, Justin J. Hendrick 1, Vivek M. Philip 1, and Gregory W. Carter 1 June 9, The Jackson Laboratory, Bar Harbor, ME, Department of Electrical and Computer Engineering, Duke University, Durham, NC Contents Abstract 1 Loading Data 2 Manipulating the Data Object 4 Looking at the Data 5 Decomposing the Phenotypes 9 Kinship Corrections 11 Single-Variant Scan 12 Covariate Selection 14 Pairwise Scan 15 Combined Analysis for Detection of Interactions 18 Interpretation of Results 23 Tables of Functions 27 Abstract Here we present an R package for the Combined Analysis of Epistasis and Pleiotropy, or cape. This package implements a method, originally described in Carter et al. (2012), that infers directed interaction networks between genetic variants for predicting the influence 1
2 of genetic perturbations on phenotypes. This method takes advantage of complementary information in partially pleiotropic genetic variants to resolve directional influences between variants that interact epistatically. cape can be applied to a variety of genetic variants, such as single nucleotide polymorphisms (SNPs), copy number variations (CNVs) or structural variations (SVs). Here we demonstrate the functionality of cape by inferring a predictive network between quantitative trait loci (QTL) in a cross between the non-obese, non-diabetic (NON) mouse and the New Zealand obese (NZO) mouse (Reifsnyder, 2000). Loading Data For the purposes of demonstration, we will reanalyze a data set described in Reifsnyder (2000). This data set was established to find quantitative trait loci (QTL) for obesity and other risk factors of type II diabetes in a reciprocal back-cross of non-obese non-diabetic NON/Lt mice and diabetes-prone, New Zealand obese (NZO/HILt) mice. The study found multiple main-effect QTL influencing phenotypes associated with diabetes and obesity as well as multiple epistatic interactions. In addition, maternal environment (i.e. whether the mother was obese) was found to interact with several markers and epistatic pairs to influence the risk of obesity and diabetes of the offspring. The complex nature of diabetes and obesity, along with their complex and polygenic inheritance patterns, make this data set ideal for an analysis of epistasis and pleiotropy. Included in this dataset are 204 male mice genotyped at 85 markers across the genome. The phenotypes included are the body weight (g), insulin levels (ng/ml), and plasma levels (mg/dl), all measured at age 24 weeks. In addition, there is a variable called mom indicating whether the mother of each mouse was normal weight (0) or obese (1). After installing cape, load the package and the dataset type the following in the R command line. > library(cape) > data(obesity.cross) There are multiple functions that can be used to load your own data set. The primary function is called read.population(). This function is similar to the function read.cross() used in the R package qtl (Broman et al., 2003), and accepts the basic R/qtl CSV format. For more information about this function type?read.population. In this function the data file can be specified as an argument. > obesity.cross <- read.population("obesity.cross.csv") Alternatively, the filename can be left blank to choose a file through the file system. The following code will bring up a window for choosing the desired file: > obesity.cross <- read.population() If the phenotypes are not specified in read.population() the default behavior is to read in all phenotypes. The functions delete.pheno() and select.pheno() can be used after reading in the data to narrow down the phenotypes for analysis. For cape analysis there 2
3 must be at least two phenotypes. There must be at least two because the fundamental concept of cape uses partial pleiotropy to refine models of epistasis. However, too many phenotypes may reduce power to detect interactions. cape handles up to 12 phenotypes, and if you have more in your data set, such as in the case of gene expression data, we suggest some form of dimension reduction prior to analysis. We offer dimension reduction through singular value decomposition (SVD), which is discussed below. Upon being read in, the data are formatted and stored in a data object. This object is referred to as data.obj in the argument lists of most functions in cape. The main functions in this analysis return data.obj with any results appended to it. The structure of this object can be viewed at any time by using the core R function str(). For example, the data that we have just loaded is named obesity.cross, and its structure can be seen here: > str(obesity.cross) List of 6 $ pheno : num [1:204, 1:4] attr(*, "dimnames")=list of 2....$ : chr [1:204] "1" "2" "3" "4" $ : chr [1:4] "body_weight" "" "insulin" "mom" $ geno : num [1:204, 1:84] NA attr(*, "dimnames")=list of 2....$ : chr [1:204] "1" "2" "3" "4" $ : chr [1:84] "1" "2" "3" "4"... $ chromosome : chr [1:84] "1" "1" "1" "1"... $ marker.names : chr [1:84] "D1Mit296" "D1Mit211" "D1Mit411" "D1Mit123"... $ marker.num : int [1:84] $ marker.location: num [1:84] Individual elements of this list can be accessed by name by using the following syntax: data.obj$name When the cross is first loaded into cape it contains the following elements: $pheno - The phenotype matrix in which individuals are in rows and phenotype values are in columns. $geno - The genotype matrix in which individuals are in rows and genetic loci are in columns. Each cell of the genotype matrix contains the probability that individual i has the reference allele at that position. By default, the reference allele is the allele that is alphabetically first if the original data file used letter designations for genotypes, or 0 if the original data file used numeric genotype designations. In the example backcross, NON/Lt is the reference allele and the heterozygote state is coded as the perturbation. Thus cape will model the effects of NZO/HILt variants. $chromosome - The chromosome on which each genetic locus is situated 3
4 $marker.names - The alphanumeric names of markers. $marker.location - The position of each locus on each chromosome $marker.num - Numeric IDs for each marker Most functions performed on data.obj will add elements to this list containing the results of the analyses. Major exceptions to this are singlescan(), and pairscan(), which generate their own singlescan and pairscan objects. In the latest version of cape there are three additional functions for reading in data set. These are read.pheno(), read.geno(), and make.data.obj(). These are used when the genotype data are very large and are more practically stored in a separate object called the geno.obj. File formats for these functions can be found in their respective help files. The function read.pheno() initializes the data.obj. The function read.geno() initializes a separate geno.obj, which contains information about the genetic markers. This will be passed to cape functions separately from the data.obj. To complete the initialization of the data.obj, marker information must be transferred to the object generated by read.pheno(). This is done using make.data.obj(). After using these three functions, you will have a data.obj with phenotype data and genetic marker information, and a geno.obj with the genetic data. > obesity.cross <- read.pheno() > geno.data <- read.geno() > obesity.cross <- make.data.obj(obesity.cross, geno.data) Manipulating the Data Object There are a number of functions available for simple manipulation of the data object. For example, after the data have been read in, phenotypes may be removed from the phenotype matrix using delete.pheno(). To select specific phenotypes, use the function select.pheno(). > obesity.cross <- select.pheno(obesity.cross, + phenotypes = c("body_weight", "", "insulin", "mom")) In cape covariates are imported as phenotypes and must be reassigned covariates. To code a phenotypic variable, such as sex, experimental treatment, or another environmental variable, as a covariate, use the function pheno2covar(). Here we convert the factor maternal environment ( mom ) to a covariate. > obesity.cross <- pheno2covar(obesity.cross, "mom") Genetic markers can also be used as covariates using the function geno2covar(). The data object can also be subset by chromosome (select.by.chr()) or by individual (select.by.ind()) before the analysis. Individuals can be selected based either on phenotype 4
5 or genotype value. For example, to use only individuals with a plasma insulin level less than than 25 ng/ml: > obesity.cross <- select.by.ind(obesity.cross, "pheno", "insulin < 25") In general we recommend saving the data object after each major computation, as well as the singlescan and pairscan objects. This allows for easy re-analysis, and prevents having to repeat lengthy calculations if there is an unexpected problem. Saving can be done with either save() or saverds(). Examining the Data Before proceeding with an analysis it is recommended that the data be examined by eye. The R package qtl has sophisticated plotting tools for examining genetic cross data,which we do not try to duplicate here. It is straightforward enough, however, to examine phenotype distributions for normality, batch effects and other quality control issues. Using the function histpheno(), we can look as the distribution of each phenotype: body_weight insulin Frequency Frequency Frequency body_weight insulin While body weight looks relatively normally distributed, and insulin have obviously non-normal distributions. Examination of the Q-Q plots of pairs of phenotypes, using the function qqpheno(), can reveal phenotyping errors and other pathologies. Here we see a threshold and ceiling effect in the relationship between the distributions of insulin and the other two phenotypes. 5
6 body_weight body_weight insulin insulin We can also examine the phenotypes by individual using plotpheno() to get an idea of systematic bases in the data body_weight Individual body_weight non obese obese Individual non obese obese insulin Individual insulin non obese obese In general we recommend mean centering and normalizing all phenotypes before proceeding with the analysis. Phenotype normalization can be achieved through log transformation, quantile normalization, or another method before the analysis. The function norm.pheno() uses quantile normalization to fit the phenotypes to a normal distribution. Briefly, this process sorts the values of the phenotype and replaces each with a corresponding value drawn from a normal distribution with the same standard deviation and mean as the original distribution. Mean centering subtracts the mean phenotype value from each phenotype value yielding a distribution centered around 0. > obesity.cross <- norm.pheno(obesity.cross, mean.center = TRUE) Plotting the histograms of the normalized data confirms the normalization. Only insulin 6
7 still has a ceiling effect, which cannot be removed by normalization because rank cannot be determined among equal values. body_weight insulin Frequency Frequency Frequency body_weight insulin We can also see that the Q-Q plots from before show the example phenotypes have been converted to the same distribution. The ceiling effect is still visible in the insulin measurement, but this cannot be removed through normalization, and overall, the distribution is more similar to those of the other phenotypes than before normalization. Knowing that this ceiling effect is present will be important in interpreting the results of the analysis insulin insulin body_weight body_weight At this point, if we decide to exclude insulin from the analysis because its distribution cannot be normalized, we can simply remove it from the data object using delete.pheno(). > obesity.cross <- delete.pheno(obesity.cross, phenotypes = "insulin") For the example here, however, we will include it in the analysis. 7
8 A Note on Phenotype Selection Phenotype selection is an important component of the cape analysis and should be given considerable thought. This method relies on the selection of two or more phenotypes that have common genetic factors but are not identical across all individuals. Such phenotypes may describe multiple aspects of a single complex trait, such as obesity or diabetes, and may encompass a combination of molecular phenotypes, such as plasma levels, and phenotypes, such as body weight, that are measured at the organismal level. The central assumption of this method is that different genetic interactions found for a single gene pair in the context of different phenotypes represent multiple manifestations of a single underlying gene network. By measuring the interactions between genetic variants in different contexts we can gain a clearer picture of the network underlying statistical epistasis (Carter et al., 2012). The phenotypes in the Reifsnyder (2000) data set are ideal for the cape analysis. They measure different aspects of the diabetes and obesity, two complex traits that are known to be related biologically and are highly correlated. The phenotypes themselves range from molecular phenotypes to organismal phenotypes. By examining the correlations between phenotypes, we can see that the phenotypes measured in this experiment are correlated, but not identical across all individuals. Ideally, phenotypes used in cape should have a Pearson correlation coefficient r between 0.4 and
9 body_weight non obese obese R = = = non obese obese R = = = non obese obese non obese obese R = = = non obese obese non obese obese insulin Note that all mice in the example backcross are male. For multisex populations, sex is usually a covariate and correlation should be assessed for each sex separately. Decomposing the Phenotypes Although cape can find genetic variants associated with raw phenotypes, the analysis was designed to work on composite traits called eigentraits. Eigentraits are calculated by factoring the matrix of phenotypes by singular value decomposition (SVD): Y = U V W T Where Y is a matrix containing one column for each mean-centered, normalized phenotype and one row for each individual. If Y contains more individuals than phenotypes, the U 9
10 matrix has the same dimensions as Y with each column containing one eigentrait. V contains the singular values, and W T contains the right singular vectors in rows. See Carter et al. (2012) for more details. The SVD de-correlates the phenotypes concentrating phenotypic features into individual eigentraits. One benefit of this process is that variants that are weakly correlated to several phenotypes due to common underlying processes may be strongly correlated to one of the eigentraits. This eigentrait captures the information of the underlying process, making strong main effects distributed between phenotypes easier to detect and identify as potential interaction loci and/or covariates. Thus, analysis of eigentraits is recommended over the analysis of raw traits. To decompose the phenotypes to eigentraits use the function get.eigentraits() If the phenotypes have not already been mean centered and normalized, this step can be performed here. In this example, we do not carry out these steps, as they have already been performed. > obesity.cross <- get.eigentraits(obesity.cross, scale.pheno = FALSE, + normalize.pheno = FALSE) This function performs the SVD and returns the matrix of eigentraits in a new element of the list called ET. The result of the decomposition can be viewed with the function plotsvd(). plotsvd(obesity.cross, orientation = "vertical") Eigentrait Contributions to Phenotypes Percent Total Variance insulin body_weight ET1 ET2 ET3 In the example illustrated here, the first eigentrait captures more than 70% of the variance in the three phenotypes. This eigentrait describes the processes by which body weight, 10
11 levels, and insulin levels all vary together. The correlations between obesity and risk factors for obesity, such as elevated insulin and fasting levels are well known (Permutt et al., 2005; Das and Elbein, 2006; Haffner, 2003). The second eigentrait captures nearly 20% of the variance in the phenotypes. It captures the processes through which and body weight vary in opposite directions. This eigentrait may be important in distinguishing the genetic discordance between obesity and diabetes. While obesity is a strong risk factor for diabetes, not all those who are obese have diabetes, and not all those with diabetes are obese (Permutt et al., 2005; Burcelin et al., 2002). The third eigentrait is less interpretable biologically, as it describes the divergence of blood and insulin levels. It may represent a genetic link between and body weight that is non-insulin dependent. Because we are primarily interested in the connection between diabetes and insulin, we will use only the first two eigentraits for the analysis. In many cases in which more than two phenotypes are being analyzed, the first two or three eigentraits will capture the majority of the variance in the data and capture obvious features. Other eigentraits may capture noise or systematic bias in the data. Often the amount of total variance captured by such eigentraits is small, and they can be removed from the analysis. Ultimately, there is no universal recipe for selecting which eigentraits should be included in the analysis, and the decision will be based on how the eigentraits contribute to the original phenotypes and how much variance in the data they capture. To select eigentraits for the analysis, use the function select.eigentraits(). Here we select the first two eigentraits. > obesity.cross <- select.eigentraits(obesity.cross, traits.which = c(1,2)) Kinship Corrections cape includes a correction for relatedness among individuals called a kinship correction Kang et al. (2008), which can be applied in the single-variant scan and the pairwise scan (described below). In the singlescan, relatedness matrices among individuals are calculated using a leave-one-chromosome-out (LOCO) method. For each chromosome (C) being tested, the relatedness matrix (K C ) is calculated using the markers on the remaining chromosomes as follows: K C = G C G T C, n where G C is the genotype matrix with markers on chromosome C removed and n is the number of markers in G C. This matrix is then used to correct the genotype and phenotype matrices for kinship using hierarchical linear models Kang et al. (2008); Gelman and Hill (2006). In the pairwise scan, the same methods are performed except that relatedness matrices are calculated leaving two chromosomes out depending on where the two markers in the tested pair reside. 11
12 Relatedness matrices can be calculated using all markers or only the markers being tested. For very large genotype matrices, there is also the option of sampling the relatedness matrix. This is less accurate than calculating the relatedness matrix directly, but can save large amounts of computation time. Single-Variant Scan Once the eigentraits for the analysis have been selected, the single-locus scan is run to investigate how individual markers are associated with each eigentrait. Note that this scan performs a linear regression at each marker. A more sophisticated single-locus analysis can be performed by R/qtl (Broman et al., 2003). This single-marker scan in cape performs the following regression for each locus on each eigentrait: U j i = βj 0 +x iβ j +ǫ j i The index i runs from 1 to the number of individuals, and j runs from 1 to the number of eigentraits or phenotypes. x i is the probability of the presence of the reference allele for individual i at locus j. The purpose of this scan is two-fold: In large data sets the number of possible variant pairs may be too large to test exhaustively. The single-variant scan can be used as a filtering step to choose variants that will be included in the pair scan. Large main effects can obscure interactions. In sufficiently powered studies, conditioning on the large QTL can aid in the discovery of interactions, variants with large main-effects can be used as covariates in the pair scan. The function singlescan() runs the single-marker scan. > obesity.singlescan <- singlescan(obesity.cross, n.perm = 100, + covar = "mom", scan.what = "eigentraits", alpha = c(0.01, 0.05), + verbose = FALSE, use.kinship = FALSE, overwrite.alert = FALSE, + run.parallel = FALSE, n.cores = 2) 10%.20%.30%.40%.50%.60%.70%.80%.100%.10%.30%.50%.60%.80%.100%.20%.50%.70%.100%.20%.50%.70%.1 The argument alpha takes a vector of alpha values for which significance thresholds will be calculated. These values are used primarily for plotting purposes. Covariates to be used in the single-marker scan should be specified through the argument covar. These covariates should have already been moved to the marker matrix by pheno2covar() or marker2covar(). An additional argument scan.what determines whether eigentraits or raw.traits will be used as the dependent variable in the regression. The single-marker scan currently does not support markers on sex chromosomes. Because the X chromosome is hemizygous in males, sex differences in phenotype can lead to false associations, and markers on this chromosome require special consideration (Broman et al., 2006). Before the single-marker scan is performed, any markers on the X and Y chromosomes 12
13 are removed from the data object. The results of the single variant scan can be visualized with the function plotsinglescan(). > plotsinglescan(data.obj = obesity.cross, singlescan.obj = obesity.singlescan, + mark.chr = TRUE, mark.covar = FALSE) ET1 ET1 [ Eff /se] mom p = 0.01 p = 0.05 ET2 [ Eff /se] ET mom p = 0.01 p = 0.05 In this figure the t-statistic (β/σ) of each marker is plotted as a vertical line. Results for both eigentraits are shown here as ET1 and ET2, and chromosome numbers are written along the x axis. If filtering of markers for the pair-wise scan is desired, an effect size cutoff can be specified using select.markers.for.pairscan(). If a specific number of markers is desired, select.markers.for.pairscan() can determine the threshold at which this number of markers is included in the pair scan. Setting a threshold for inclusion is useful for large data sets for which it is impractical to test all marker pairs. Here we use all markers 13
14 in the pairwise scan. In this example we will not filter the markers. A kinship correction can be implemented in the singlescan simply by setting the argument use.kinship to TRUE. Covariate Selection As mentioned previously, conditioning on large main effects may aid in the discovery of interactions. Therefore cape allows specification of genetic markers as covariates. This specification is recommended if there are markers with particularly large effect sizes that may obfuscate weak interactions. In an organism in which variants segregate relatively independently, such as in Drosophila (Mackay et al., 2012), it may be useful to select individual markers with very high effect sizes as covariates. In a population with more correlation between adjacent markers, more care should be given to this process. In a mouse F2 intercross or backcross, on the other hand, markers on a single chromosome tend to be linked, and groups of markers may be significantly associated with a phenotype due to linkage with a causative locus. In this case, markers from a single linkage block carry partially redundant information and, if treated as independent covariates, will confound interactions between makers in the linkage block and other truly independent markers. Thus, if markers in the study population are linked, it is recommended that any marker covariates be set manually and that only one covariate should be selected from each linkage block. Setting covariates, as well as the filtering of markers for the pair-wise scan can be done with the function select.markers.for.pairscan(). When calling this function, if thresholding the markers based on effect size, set use.pairs.threshold to TRUE, and pairscan.thresh to the desired standardized effect size threshold. Alternatively, this function can be used to select a threshold based on a desired number of markers to test in the pair-wise scan. The target number of markers desired is set in num.markers. The function then searches through multiple thresholds to find a linearly independent matrix holding the number of desired markers. It starts the search at the threshold set in start.thresh and stops when it finds a threshold resulting in num.markers plus or minus the tolerance. This process is slow, and it is recommended that the starting threshold be set as close to the estimated correct threshold as possible and run only once to find the desired threshold. After the first run, the threshold can be set manually. In addition to selecting markers based on an effect size threshold, select.markers.for.pairscan() can also set specific markers as covariates through the argument specific.markers. This argument takes a vector of strings and sets each one as a covariate. > obesity.cross <- select.markers.for.pairscan(data.obj = obesity.cross, + singlescan.obj = obesity.singlescan) 82 markers were selected for the pairscan. This makes 3321 possible pairs. 14
15 Pairwise Scan The purpose of the pairwise scan is to find interactions, or epistasis, between variants. The epistatic models are then combined across phenotypes or eigentraits to infer a parsimonious network that takes data from all eigentraits into account. To find epistatic interactions pairscan() tests the following model for each variant 1 and 2: 2 U j i = βj 0 + x c,i βc j +x 1,i β j 1 +x 2,iβ j 2+x 1,i x 2,i β j 12 }{{}}{{} c=1 }{{} main effects interaction covariates The terms in this equation are the same as those in the equation for the single-variant scan except for the addition of the term for the interaction between the two variants being tested. This additional term brings further complications to the model, and restricts which markers can be tested. Because many markers, including covariates, may be included in the model, we need to be careful about including only markers that are linearly independent of each other. Linear dependence between markers occurs when two markers are fully linked and therefore perfectly correlated. The two markers provide the same genetic information, and one can be discarded without loss of information. Before running the pair scan, it is important that we reduce the genetic matrix to only markers that are linearly independent of one another. This step is performed by the function select.markers.for.pairscan(). It calculates the correlation between all pairs of markers. If any are found to be perfectly correlated, the first marker is removed. As mentioned above, select.markers.for.pairscan() also optionally filters markers for inclusion in the pair scan by standardized effect size. This is useful in large crosses in which it may be impossible to test all possible pairs of markers in a reasonable amount of time. obesity.cross <- select.markers.for.pairscan(data.obj = obesity.cross, singlescan.obj = obesity.singlescan) The number of markers removed is printed to the screen, and the names of these markers are written to the file markers.removed.txt in the current working directory. This filtering step adds one element to the data object: a filtered genotype matrix called geno.for.pairscan which holds the markers that will be used in the pairscan. The results of the filtering can be visualized with plotsinglscan(). This visualization can be helpful in seeing the locations of discarded markers, especially if many have been removed. To indicate which markers have been selected for the pair scan, set show.selected.markers to TRUE. To indicate which markers have been rejected from the pair scan, set show.rejected.markers to TRUE. > plotsinglescan(data.obj = obesity.cross, singlescan.obj = obesity.singlescan, + mark.chr = TRUE, show.rejected.markers = TRUE, standardized = TRUE) +ǫ j i 15
16 ET1 * rejected ET1 [ Eff /se] ** mom p = 0.01 p = 0.05 ET2 [ Eff /se] ET ** mom p = 0.01 p = 0.05 After ensuring that all markers are linearly independent and thresholded satisfactorily, the pairscan can be run using pairscan(). > obesity.pairscan <- pairscan(data.obj = obesity.cross, covar = "mom", + scan.what = "eigentraits", total.perm = 1000, min.per.genotype = 6, + verbose = FALSE, overwrite.alert = FALSE, n.cores = 2) The arguments here are familiar from singlescan(), with the exception of two additional arguments. min.per.genotype and max.pair.cor are thresholds that prevent highly correlated markers from being tested in pairs. Only one of these arguments can be set depending on whether the genotypes in the data set are continuous or discrete. If the genotypes are discrete, min.per.genotype can be set to prevent empty cells in genotype matrices. In an intercross, there are three possible genotypes at each marker, giving a total of nine possible genotypes for the pair. For example, for two markers each with the genotypes 16
17 AA, AB, and BB, the pairwise genotypes are the following: AA AB BB AA AB BB In a backcross each marker only has two possible genotypes, AA and AB, yielding four possible pairwise genotypes. In both cases, insufficient representative individuals of single genotypes indicates that the two markers in question are linked. This linkage leads to false associations in both the effects and permutations. False associations in permutations results in a heavy-tailed null distribution, which artificially inflates the threshold of significance and reduces power to find true interactions. Because of this effect, linked marker pairs are not tested. The threshold used to determine insufficient recombination between markers is given by min.per.genotype. The default behavior is to reject any marker pair for which there are fewer than six individuals in any of the genotype cells. This value can be adjusted, but caution should be used in interpreting the results if min.per.genotype is very low or 0. Alternatively, if genotypes are coded continuously, the argument max.pair.cor can be set to establish an upper threshold on the Pearson correlation between marker pairs. As in the case with discrete genotypes, testing highly correlated markers against each other can lead to false associations. Either max.pair.cor or min.per.genotype, but not both, must be set. The total.perm argument in pairscan() sets the total number of permutations performed over all marker pairs. To generate a null distribution for the pair-wise tests, CAPE performs permutation tests in the following way: First singlescan is run on the permuted phenotypes. Then the N markers with the largest effect sizes are selected, where N is the same number of markers being tested in the pairwise scan. This ensures that if there are very few markers being tested in the pairwise scan, we do not generate a biased null distribution by only performing permutation tests repeatedly on these markers. A pairwise scan is then performed on these top N markers using the permuted phenotype. To calculate empirical p values, the permutations are combined across all markers to result in one large null distribution. Thus, large null distributions can be achieved with relatively few permutations per pair. This is useful, as even with relatively sparse genotyping the number of tests performed can be large and take many hours to perform. In this demonstration we only run 1000 permutations, but in actual use, we recommend at least 500,000 permutations. Low numbers of permutations tend to result in many false positives. The results of the pair scan can be plotted with plotpairscan(). This plot shows the resulting β/σ for each pair of markers for ET1. Gray and white bars show the boundaries of the chromosomes. 17
18 > plotpairscan(data.obj = obesity.cross, pairscan.obj = obesity.pairscan, + phenotype = "ET1", pdf.label = "Pairscan_Regression.pdf") null device 1 ET1 marker c marker c Combined Analysis for Detection of Interactions From the pair scan, each pair of markers 1 and 2 receives a set of β coefficients describing the main effect of each marker on each eigentrait j (β j 1 and βj 2 ) as well as the interaction effect of both markers on each eigentrait (β j 12 ) (See figure below). The central idea of cape is that these coefficients can be combined across eigentraits and reparameterized to calculate how each pair of markers influences each other directly and independently of eigentrait. phenotype1 phenotype1 β 1 1 β 1 β var1 var2 β 2 12 β 2 1 β 2 2 phenotype2 β 1 1 var1 m 12 m 21 phenotype2 β 1 2 var2 β 2 1 β 2 2 The first step in this reparameterization is to define two new parameters (δ 1 and δ 2 ) in terms of the interaction coefficients. δ 1 can be thought of as the additional genetic activity 18
19 of marker 1 when marker 2 is present. Together the δ terms capture the interaction term, and are interpreted as the extent to which each marker influences the effect of the other on downstream phenotypes. For example, a negative δ 2 indicates that the presence of marker 2 represses the effect of marker 1 on the phenotypes or eigentraits. The δ terms are related to the main effects and interaction effects as follows: β 1 1 β2 1 β 1 2 β2 2 δ β = β δ In multiplying out this equation, it can be seen how the δ terms influence each main effect term to give rise to the interaction terms independent of phenotype. (1) β j 1 δ 1 +β j 2 δ 2 = β j 12 (2) If δ 1 = 0 and δ 2 = 0, there are no addition effects exerted by the markers when both are present. Substitution into the equations above shows that the interaction terms β j 12 are 0 and thus the interaction terms have no effect on the phenotype. Alternatively, consider the situation when δ 1 = 1 and δ 2 = 0. The positive δ 1 indicates that marker 1 should exert an additional effect when marker 2 is present. This can be seen again through substitution into equation 2: β 1 j = β j 12 These non-zero terms show that there is an interaction effect between marker 1 and marker 2. The positive δ 1 indicates that this interaction is driven through an enhanced effect of marker 1 in the presence of marker 2. The δs are calculated by solving for equation 1 using matrix inversion: δ β 1 1 β2 1 1 = β δ 1 2 β β 1 12 β This inversion is exact for two eigentraits, and cape implements pseudo-inversion for up to 12 eigentraits. The δ terms are then translated into directed variables defining the marker-to-marker influences m 12 and m 21. Whereas δ 2 described the change in activity of marker 2 in the presence of marker 1, m 12 can be thought of as the direct influence of marker 2 on marker 1, with negative values indicating repression and positive values indicating enhancement. The terms m 12 and m 21 are self-consistent and defined in terms of δ 1 and δ 2 : δ 1 = m 12 (1+δ 2 ), δ 2 = m 21 (1+δ 1 ) 19
20 Rearranging these equations yields the solutions: m 12 = δ 1 1+δ 2, m 21 = δ 2 1+δ 1. These directed influence variables provide a map of how each marker influences each other marker independent of phenotype. The significance of these influences is determined through standard error analysis on the regression parameters (Bevington, 1994; Carter et al., 2012). This step is particularly important as matrix inversion can lead to large values but larger standarderrors, yieldinginsignificantresults. Asanexample, thevarianceofm 12 iscalculated by differentiating with respect to all model parameters: σ 2 m 12 = ij σ 2 β j i ( m 12 β j i ) 2 +2 i<k,j<l σ 2 β j i βl k ( m 12 β j i )( m 12 β l k In this equation, the indices i and k run over regression parameters and j and l run from 1 to the number of traits. The calculations of the δ and the m terms, as well as the error propagation are performed by the function error.prop(). > obesity.cross <- error.prop(data.obj = obesity.cross, + pairscan.obj = obesity.pairscan, + perm = FALSE, verbose = FALSE, n.cores = 2) This function is applied to both the results from the pairwise scan, as well as the permutations of the pairwise scan for later calculation of p values. To apply the calculations to the pairwise scan permutations, set perm = TRUE. > obesity.cross <- error.prop(data.obj = obesity.cross, + pairscan.obj = obesity.pairscan, + perm = TRUE, verbose = FALSE, n.cores = 2) For large scans it might be desirable to observe the progress of the calculations. This can be done by setting verbose = TRUE. After these calculations have been performed the results of the permutation testing can be used to calculate empirical p values for each of the variant-to-variant effects. > obesity.cross <- calc.p(data.obj = obesity.cross, + pairscan.obj = obesity.pairscan, pval.correction = "fdr", + n.cores = 2) This function also adjusts the empirical p values for multiple testing. The default correction is Holm s stepdown procedure (Holm, 1979). Two other methods, false discovery rate (FDR) (Benjamini and Hochberg, 1995) and local false discovery rate (lfdr) (Liao et al., 2004) are also available. The latter methods of correction for multiple testing are less stringent than the Holm s step-down procedure. The function calc.p() adds two elements to the data object. The purpose and returned results of each function is summarized below. For tables of all functions, see the Tables of Functions section at end of this document. ) 20
21 Functions for Parsing Pair Scan Results Function Behavior Result Label Structure Columns error.prop propagates errors of coefficients calculated in the pair scan var.to.var.influences or var.to.var.influences.perm matrix marker1, marker2, m12, m21, m12 σ, m21 σ calc.p calculates empirical and corrected p values or fdr of interactions using permutations empirical.p list of 2 matrices (m12, m21) marker1, marker2, source, target, influence coefficient, influence σ, empirical p value, adjusted p value or fdr The final step in calculating the network of directed influences is to translate the effect of each marker on the eigentraits to effects on the original phenotypes. The effects of variants on eigentraits are defined in terms of the interaction coefficients m 12 and m 21 as well as the main effect of each variant β j i. To translate these effects to be in terms of the original phenotypes, the coefficient matrices are multiplied by the singular value matrices V W T. With two phenotypes and two eigentraits this conversion results in no loss of information. It should also be noted that the translation back to phenotype space does not affect the variant-to-variant influences. The translation to phenotype space is performed by the function direct.influence(). > obesity.cross <- direct.influence(data.obj = obesity.cross, + pairscan.obj = obesity.pairscan, pval.correction = "fdr") In these calculations each marker is assigned multiple direct influences on each phenotype dependent on the marker it was paired with. To reduce the results to a single direct influence of each marker on each phenotype, direct.influence() selects the maximum influence observed for each marker. These effects are stored in max.var.to.pheno.influence. All effects are accessible in the element variant.to.phenotype.influences. To save memory, data from the permutations are not saved by default; however, if save.permutations is set to TRUE all data are saved in the object permutation.data.rdata. For details on the elements added to obesity.cross and those saved in permutation.data.rdata, see the Tables of Functions section at end of this document. The function direct.influence() also performs correction for multiple testing for each of the influences. Holm step-down correction (Holm, 1979), FDR Benjamini and Hochberg (1995), and local FDR (Liao et al., 2004) are all available as correction methods. The marker-to-marker and marker-to-phenotype influences can be plotted with the function plotvariantinfluences(). This function plots the adjacency matrix of the final network. It shows all significant influences between variants and between variants and phenotypes. plotvariantinfluences(obesity.cross, p.or.q = 0.05, 21
22 standardize = TRUE, not.tested.col = "lightgray", pheno.width = 8) Variant Influences Source mom mom Target body_weight insulin not testable There are several arguments in this function to note. The argument p.or.q takes in the p, q, or local fdr value at which interactions are considered significant. Only significant interactions are plotted. Non-significant interactions are colored white. It should be noted that some of the white boxes have gray dots in them. These dots indicate that these pairs were not tested for interactions because they were filtered out due to insufficient representation of individual phenotypes. This usually occurs because the markers are linked and there is low recombination between them. These pairs are marked with dots to distinguish them from pairs that were tested but did not have significant interactions between them. The color of the dots can be changed with the argument not.tested.col. The default is lightgray. This can be changed to FALSE or NA if no distinction between not-tested and not-significant 22
23 is desired. Gray and white bars along the margins of the plot indicate the boundaries of the chromosomes. It should also be noted that not all of the original markers from the single-locus scan are represented in the final figure. This figure is showing only those markers that were included in the pair scan. The argument scale.effects allows application of a function, either log 10 or square root to the values in the adjacency matrix. These transformations can help increase the contrast between markers. And finally, the argument pheno.width specifies how wide the main effects should be relative to the interaction effects for better visualization. A table of significant influences can be written to a file using the function writevariantinfluences(). writevariantinfluences(obesity.cross, p.or.q = 0.05, filename = "Significant.Influences.txt") Interpretation of Results The table of variant-to-variant and variant-to-phenotype influences is the primary output of cape. The function for plotting results, plotvariantinfluences(), plots an asymmetric adjacency matrix, which is useful for identifying patterns in variant interactions. Another useful visualization of the network can be plotted using plotnetwork(). This function plots the chromosomes in a circle and shows interactions as arrows between regions on chromosomes (see figures below). Main effects are plotted in the circles around the chromosomes. Before plotting the network in this format, the function get.network() must be be run to create a network object. The function get.network() network offers the option of condensing markers based on their linkage. To group markers into linkage blocks cape calculates a correlation matrix for all markers on each chromosome. This similarity matrix is used as an adjacency matrix to construct a weighted network depicting the similarity between all pairs of markers on a single chromosome. Using the fastgreedy community detection algorithm (Clauset et al., 2004) in R/igraph (Csardi and Nepusz, 2006), cape then calculates the community membership of the vertices in the network. Adjacent markers in the same community are assigned to a single QTL region. Finally, the argument p.or.q determines the p, q, or local FDR value at which marker influences are determined significant. obesity.cross <- get.network(obesity.cross, p.or.q = 0.05, collapse.linked.markers = TRUE) plotnetwork(obesity.cross, collapsed.net = TRUE) 23
24 mom insulin body_weight Enhancing Suppressing If no collapse by linkage is desired, the argument collapse.linked.markers can be set to FALSE. obesity.cross <- get.network(obesity.cross, p.or.q = 0.05, collapse.linked.markers = TRUE) plotnetwork(obesity.cross, collapsed.net = FALSE) Both the network figure and the adjacency matrix show direct influences of markers on the phenotypes as well as interactions between markers. As expected, many NZO variants on multiple chromosomes show positive effects on plasma insulin and levels as well as on body weight. Two loci had no main effects themselves, but enhanced the effects of maternal obesity on levels. In an interaction plot (Figure??), we can see that animals with obese mothers had higher blood levels than those with non-obese mothers. However, those individuals with the heterozygous genotype at D13MIT54 showed a greater increase in with obese mothers than did the individuals with the NON genotype at this locus. Thus the NZO allele at this locus enhances the effect of maternal obesity on blood levels. 24
25 mom insulin body_weight Enhancing Suppressing NON 0 1 D13Mit54 Het Mom obese non-obese It can also be seen in this figure why the direction of action from the genetic locus to the maternal factor was inferred. The genetic locus enhances the phenotypic effect of maternal obesity, but the genetic marker itself does not have a direct effect on that can be enhanced or suppressed by maternal obesity. Furthermore, the phenotypic effects associated 25
26 with this locus on insulin and body weight are not influenced by maternal obesity. These findings illustrate how cape is designed to find interactions that simultaneously model all phenotypes under the assumption that interactions between variants across multiple contexts represent a single underlying interaction network. Thus we recommend users assess single-phenotype epistasis using functions in cape or in parallel analyses using tools such as R/qtl and R/qtlbim (Yandell et al., 2007). 26
27 Tables of Functions 27
28 Basic cape Functions read.population Reading and Writing Data read in population data writevariantinfluences write out a table of the variant influences to each other and the variant influences on traits. Data Manipulation delete.pheno get.covar marker2covar norm.pheno pheno2covar select.eigentraits select.pheno select.by.chr select.by.ind delete the specified phenotypes from the phenotype matrix retrieve information about set covariates designate a genetic marker as a covariate use rank Z normalization to normalize the phenotypes designate a phenotypic value as a covariate select a subset of eigentraits to use in the analysis select a subset of phenotypes to use in the analysis select a subset of chromosomes to use in the analysis subset the individuals in the population by either phenotypic or genotypic values Plotting Functions histpheno plotnetwork plotpheno plotphenocor plotsinglescan plotsinglescan.heat plotpairscan plotsvd plotvariantinfluences qqpheno plot histograms of phenotypes plot a network view of the significant influences of variants on each other and variants on traits plot phenotypes by individual with the option of coloring points according to a covariate plot correlations between phenotypes with the option of coloring points according to a covariate plot the results of the single-locus scan or markers selected for pairscan plot the results of the single-locus scan as a heatmap. plot the results of the pair scan plot the results of the singular value decomposition plot the adjacency matrix showing the significant influences of variants on each other and variants on traits plot qq plots for pairs of phenotypes. 28
29 Functions for cape Analysis Function Behavior Result Label Structure Internal elements Each Row Columns calc.p calculates empirical and corrected p values of interactions using permutations empirical.p list of 2 matrices m12, m21 marker pair marker1, marker2, source, target, influence coefficient, influence σ, empirical p value, adjusted p value direct.influence calculates the direct influence of each marker pair on each trait and the p value of each influence var.to.pheno.influence list of Nt matrices each matrix is named with its respective trait marker pair marker1, marker2, influence coefficients and standard errors for each marker on each trait var.to.pheno.influence.perm list of Nt matrices each matrix is named with its respective trait permutation of each marker pair marker1, marker2, permuted influence coefficients and standard errors for each marker on each trait var.to.pheno.test.stat list of Nt matrices each matrix is named with its respective trait one marker in one marker pair context marker, influence coefficient, σ, t statistic, t statistic var.to.pheno.test.stat.perm list of Nt matrices each matrix is named with its respective trait one permutation for each marker in one marker pair context marker, influence coefficient, σ, t statistic, t statistic max.var.to.pheno.influence list of Nt matrices each matrix is named with its respective trait marker marker, influence coefficient, σ, t statistic, t statistic, empirical p value, adjusted p value error.prop propagates errors of coefficients calculated in the pair scan var.to.var.influences or var.to.var.influences.perm matrix marker pair marker1, marker2, m12, m21, m12 σ, m21 σ get.eigentraits performs singular value decomposition on phenotype matrix ET matrix individual one column for each eigentrait maximum.influence calculates the maximum influence that each marker has on each trait across all pair contexts max.var.to.pheno.influence list of Nt matrices each matrix is named with its respective trait marker pair marker, influence coefficient, σ, t statistic, t statistic, empirical p value, adjusted p value singlescan performs all single-locus regressions for each marker on each trait singlescan.results list of Nt matrices each matrix is named with its respective trait marker marker coefficient, σ, t statistic, p value pairscan(n.perm 0) performs pair-wise marker regression for each trait pairscan.results list of Nt lists, each with 3 elements twod.effects marker pair marker labels and coefficients for each marker tested and each marker used as a covariate, including the interaction term twod.se marker pair marker labels and σ for each coefficient in twod.effects model.covariance marker pair the model covariance matrix for each regression pairscan(n.perm 1) performs pair-wise marker regression and permutations for each trait pairscan.perm list of Nt lists, each with 3 elements twod.effects.perm one permutation of marker pair marker labels and coefficients for each marker tested and each marker used as a covariate, including the interaction term for all permutations twod.se.perm one permutation of marker pair marker labels and σ for each coefficient in twod.effects.perm model.covariance.perm one permutation of marker pair the model covariance matrix for each permuted regression select.markers.for.pairscan selects markers for the pair scan based on which markers are linearly independent of the other markers in the data set geno.for.pairscan matrix individual one column for each independent marker selected for the pair scan N t = Number of traits (phenotypes or eigentraits) scanned These objects are not added to the data object, but saved in permutation.data.rdata if specified. covar.for.pairscan matrix marker each of the Nt columns indicates which markers are to be used as covariates in the pair scan of each trait 29
New Enhancements: GWAS Workflows with SVS
 New Enhancements: GWAS Workflows with SVS August 9 th, 2017 Gabe Rudy VP Product & Engineering 20 most promising Biotech Technology Providers Top 10 Analytics Solution Providers Hype Cycle for Life sciences
New Enhancements: GWAS Workflows with SVS August 9 th, 2017 Gabe Rudy VP Product & Engineering 20 most promising Biotech Technology Providers Top 10 Analytics Solution Providers Hype Cycle for Life sciences
An Introduction to Quantitative Genetics I. Heather A Lawson Advanced Genetics Spring2018
 An Introduction to Quantitative Genetics I Heather A Lawson Advanced Genetics Spring2018 Outline What is Quantitative Genetics? Genotypic Values and Genetic Effects Heritability Linkage Disequilibrium
An Introduction to Quantitative Genetics I Heather A Lawson Advanced Genetics Spring2018 Outline What is Quantitative Genetics? Genotypic Values and Genetic Effects Heritability Linkage Disequilibrium
Unit 1 Exploring and Understanding Data
 Unit 1 Exploring and Understanding Data Area Principle Bar Chart Boxplot Conditional Distribution Dotplot Empirical Rule Five Number Summary Frequency Distribution Frequency Polygon Histogram Interquartile
Unit 1 Exploring and Understanding Data Area Principle Bar Chart Boxplot Conditional Distribution Dotplot Empirical Rule Five Number Summary Frequency Distribution Frequency Polygon Histogram Interquartile
Discontinuous Traits. Chapter 22. Quantitative Traits. Types of Quantitative Traits. Few, distinct phenotypes. Also called discrete characters
 Discontinuous Traits Few, distinct phenotypes Chapter 22 Also called discrete characters Quantitative Genetics Examples: Pea shape, eye color in Drosophila, Flower color Quantitative Traits Phenotype is
Discontinuous Traits Few, distinct phenotypes Chapter 22 Also called discrete characters Quantitative Genetics Examples: Pea shape, eye color in Drosophila, Flower color Quantitative Traits Phenotype is
Tutorial on Genome-Wide Association Studies
 Tutorial on Genome-Wide Association Studies Assistant Professor Institute for Computational Biology Department of Epidemiology and Biostatistics Case Western Reserve University Acknowledgements Dana Crawford
Tutorial on Genome-Wide Association Studies Assistant Professor Institute for Computational Biology Department of Epidemiology and Biostatistics Case Western Reserve University Acknowledgements Dana Crawford
LTA Analysis of HapMap Genotype Data
 LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
The Loss of Heterozygosity (LOH) Algorithm in Genotyping Console 2.0
 The Loss of Heterozygosity (LOH) Algorithm in Genotyping Console 2.0 Introduction Loss of erozygosity (LOH) represents the loss of allelic differences. The SNP markers on the SNP Array 6.0 can be used
The Loss of Heterozygosity (LOH) Algorithm in Genotyping Console 2.0 Introduction Loss of erozygosity (LOH) represents the loss of allelic differences. The SNP markers on the SNP Array 6.0 can be used
Complex Traits Activity INSTRUCTION MANUAL. ANT 2110 Introduction to Physical Anthropology Professor Julie J. Lesnik
 Complex Traits Activity INSTRUCTION MANUAL ANT 2110 Introduction to Physical Anthropology Professor Julie J. Lesnik Introduction Human variation is complex. The simplest form of variation in a population
Complex Traits Activity INSTRUCTION MANUAL ANT 2110 Introduction to Physical Anthropology Professor Julie J. Lesnik Introduction Human variation is complex. The simplest form of variation in a population
Lab 5: Testing Hypotheses about Patterns of Inheritance
 Lab 5: Testing Hypotheses about Patterns of Inheritance How do we talk about genetic information? Each cell in living organisms contains DNA. DNA is made of nucleotide subunits arranged in very long strands.
Lab 5: Testing Hypotheses about Patterns of Inheritance How do we talk about genetic information? Each cell in living organisms contains DNA. DNA is made of nucleotide subunits arranged in very long strands.
CNV PCA Search Tutorial
 CNV PCA Search Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Data Preparation 2 A. Join Log Ratio Data with Phenotype Information.............................. 2 B. Activate only
CNV PCA Search Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Data Preparation 2 A. Join Log Ratio Data with Phenotype Information.............................. 2 B. Activate only
Abstract. Optimization strategy of Copy Number Variant calling using Multiplicom solutions APPLICATION NOTE. Introduction
 Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Figure S2. Distribution of acgh probes on all ten chromosomes of the RIL M0022
 96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
Intro to SPSS. Using SPSS through WebFAS
 Intro to SPSS Using SPSS through WebFAS http://www.yorku.ca/computing/students/labs/webfas/ Try it early (make sure it works from your computer) If you need help contact UIT Client Services Voice: 416-736-5800
Intro to SPSS Using SPSS through WebFAS http://www.yorku.ca/computing/students/labs/webfas/ Try it early (make sure it works from your computer) If you need help contact UIT Client Services Voice: 416-736-5800
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Complete Genomics, Inc.
 Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
Introduction of Genome wide Complex Trait Analysis (GCTA) Presenter: Yue Ming Chen Location: Stat Gen Workshop Date: 6/7/2013
 Introduction of Genome wide Complex Trait Analysis (GCTA) resenter: ue Ming Chen Location: Stat Gen Workshop Date: 6/7/013 Outline Brief review of quantitative genetics Overview of GCTA Ideas Main functions
Introduction of Genome wide Complex Trait Analysis (GCTA) resenter: ue Ming Chen Location: Stat Gen Workshop Date: 6/7/013 Outline Brief review of quantitative genetics Overview of GCTA Ideas Main functions
Estimating genetic variation within families
 Estimating genetic variation within families Peter M. Visscher Queensland Institute of Medical Research Brisbane, Australia peter.visscher@qimr.edu.au 1 Overview Estimation of genetic parameters Variation
Estimating genetic variation within families Peter M. Visscher Queensland Institute of Medical Research Brisbane, Australia peter.visscher@qimr.edu.au 1 Overview Estimation of genetic parameters Variation
Contents. 2 Statistics Static reference method Sampling reference set Statistics Sampling Types...
 Department of Medical Protein Research, VIB, B-9000 Ghent, Belgium Department of Biochemistry, Ghent University, B-9000 Ghent, Belgium http://www.computationalproteomics.com icelogo manual Niklaas Colaert
Department of Medical Protein Research, VIB, B-9000 Ghent, Belgium Department of Biochemistry, Ghent University, B-9000 Ghent, Belgium http://www.computationalproteomics.com icelogo manual Niklaas Colaert
The Association Design and a Continuous Phenotype
 PSYC 5102: Association Design & Continuous Phenotypes (4/4/07) 1 The Association Design and a Continuous Phenotype The purpose of this note is to demonstrate how to perform a population-based association
PSYC 5102: Association Design & Continuous Phenotypes (4/4/07) 1 The Association Design and a Continuous Phenotype The purpose of this note is to demonstrate how to perform a population-based association
Supplementary Figures
 Supplementary Figures Supplementary Fig 1. Comparison of sub-samples on the first two principal components of genetic variation. TheBritishsampleisplottedwithredpoints.The sub-samples of the diverse sample
Supplementary Figures Supplementary Fig 1. Comparison of sub-samples on the first two principal components of genetic variation. TheBritishsampleisplottedwithredpoints.The sub-samples of the diverse sample
Section 6: Analysing Relationships Between Variables
 6. 1 Analysing Relationships Between Variables Section 6: Analysing Relationships Between Variables Choosing a Technique The Crosstabs Procedure The Chi Square Test The Means Procedure The Correlations
6. 1 Analysing Relationships Between Variables Section 6: Analysing Relationships Between Variables Choosing a Technique The Crosstabs Procedure The Chi Square Test The Means Procedure The Correlations
GENETIC LINKAGE ANALYSIS
 Atlas of Genetics and Cytogenetics in Oncology and Haematology GENETIC LINKAGE ANALYSIS * I- Recombination fraction II- Definition of the "lod score" of a family III- Test for linkage IV- Estimation of
Atlas of Genetics and Cytogenetics in Oncology and Haematology GENETIC LINKAGE ANALYSIS * I- Recombination fraction II- Definition of the "lod score" of a family III- Test for linkage IV- Estimation of
bivariate analysis: The statistical analysis of the relationship between two variables.
 bivariate analysis: The statistical analysis of the relationship between two variables. cell frequency: The number of cases in a cell of a cross-tabulation (contingency table). chi-square (χ 2 ) test for
bivariate analysis: The statistical analysis of the relationship between two variables. cell frequency: The number of cases in a cell of a cross-tabulation (contingency table). chi-square (χ 2 ) test for
BIOL 458 BIOMETRY Lab 7 Multi-Factor ANOVA
 BIOL 458 BIOMETRY Lab 7 Multi-Factor ANOVA PART 1: Introduction to Factorial ANOVA ingle factor or One - Way Analysis of Variance can be used to test the null hypothesis that k or more treatment or group
BIOL 458 BIOMETRY Lab 7 Multi-Factor ANOVA PART 1: Introduction to Factorial ANOVA ingle factor or One - Way Analysis of Variance can be used to test the null hypothesis that k or more treatment or group
9 research designs likely for PSYC 2100
 9 research designs likely for PSYC 2100 1) 1 factor, 2 levels, 1 group (one group gets both treatment levels) related samples t-test (compare means of 2 levels only) 2) 1 factor, 2 levels, 2 groups (one
9 research designs likely for PSYC 2100 1) 1 factor, 2 levels, 1 group (one group gets both treatment levels) related samples t-test (compare means of 2 levels only) 2) 1 factor, 2 levels, 2 groups (one
MULTIPLE LINEAR REGRESSION 24.1 INTRODUCTION AND OBJECTIVES OBJECTIVES
 24 MULTIPLE LINEAR REGRESSION 24.1 INTRODUCTION AND OBJECTIVES In the previous chapter, simple linear regression was used when you have one independent variable and one dependent variable. This chapter
24 MULTIPLE LINEAR REGRESSION 24.1 INTRODUCTION AND OBJECTIVES In the previous chapter, simple linear regression was used when you have one independent variable and one dependent variable. This chapter
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Imaging Genetics: Heritability, Linkage & Association
 Imaging Genetics: Heritability, Linkage & Association David C. Glahn, PhD Olin Neuropsychiatry Research Center & Department of Psychiatry, Yale University July 17, 2011 Memory Activation & APOE ε4 Risk
Imaging Genetics: Heritability, Linkage & Association David C. Glahn, PhD Olin Neuropsychiatry Research Center & Department of Psychiatry, Yale University July 17, 2011 Memory Activation & APOE ε4 Risk
Chapter 1. Introduction
 Chapter 1 Introduction 1.1 Motivation and Goals The increasing availability and decreasing cost of high-throughput (HT) technologies coupled with the availability of computational tools and data form a
Chapter 1 Introduction 1.1 Motivation and Goals The increasing availability and decreasing cost of high-throughput (HT) technologies coupled with the availability of computational tools and data form a
Numerous hypothesis tests were performed in this study. To reduce the false positive due to
 Two alternative data-splitting Numerous hypothesis tests were performed in this study. To reduce the false positive due to multiple testing, we are not only seeking the results with extremely small p values
Two alternative data-splitting Numerous hypothesis tests were performed in this study. To reduce the false positive due to multiple testing, we are not only seeking the results with extremely small p values
MBG* Animal Breeding Methods Fall Final Exam
 MBG*4030 - Animal Breeding Methods Fall 2007 - Final Exam 1 Problem Questions Mick Dundee used his financial resources to purchase the Now That s A Croc crocodile farm that had been operating for a number
MBG*4030 - Animal Breeding Methods Fall 2007 - Final Exam 1 Problem Questions Mick Dundee used his financial resources to purchase the Now That s A Croc crocodile farm that had been operating for a number
Decomposition of the Genotypic Value
 Decomposition of the Genotypic Value 1 / 17 Partitioning of Phenotypic Values We introduced the general model of Y = G + E in the first lecture, where Y is the phenotypic value, G is the genotypic value,
Decomposition of the Genotypic Value 1 / 17 Partitioning of Phenotypic Values We introduced the general model of Y = G + E in the first lecture, where Y is the phenotypic value, G is the genotypic value,
White Paper Estimating Genotype-Specific Incidence for One or Several Loci
 White Paper 23-01 Estimating Genotype-Specific Incidence for One or Several Loci Authors: Mike Macpherson Brian Naughton Andro Hsu Joanna Mountain Created: September 5, 2007 Last Edited: November 18, 2007
White Paper 23-01 Estimating Genotype-Specific Incidence for One or Several Loci Authors: Mike Macpherson Brian Naughton Andro Hsu Joanna Mountain Created: September 5, 2007 Last Edited: November 18, 2007
Minimizing Uncertainty in Property Casualty Loss Reserve Estimates Chris G. Gross, ACAS, MAAA
 Minimizing Uncertainty in Property Casualty Loss Reserve Estimates Chris G. Gross, ACAS, MAAA The uncertain nature of property casualty loss reserves Property Casualty loss reserves are inherently uncertain.
Minimizing Uncertainty in Property Casualty Loss Reserve Estimates Chris G. Gross, ACAS, MAAA The uncertain nature of property casualty loss reserves Property Casualty loss reserves are inherently uncertain.
Non-parametric methods for linkage analysis
 BIOSTT516 Statistical Methods in Genetic Epidemiology utumn 005 Non-parametric methods for linkage analysis To this point, we have discussed model-based linkage analyses. These require one to specify a
BIOSTT516 Statistical Methods in Genetic Epidemiology utumn 005 Non-parametric methods for linkage analysis To this point, we have discussed model-based linkage analyses. These require one to specify a
Name: PS#: Biol 3301 Midterm 1 Spring 2012
 Name: PS#: Biol 3301 Midterm 1 Spring 2012 Multiple Choice. Circle the single best answer. (4 pts each) 1. Which of the following changes in the DNA sequence of a gene will produce a new allele? a) base
Name: PS#: Biol 3301 Midterm 1 Spring 2012 Multiple Choice. Circle the single best answer. (4 pts each) 1. Which of the following changes in the DNA sequence of a gene will produce a new allele? a) base
SUPPLEMENTAL MATERIAL
 1 SUPPLEMENTAL MATERIAL Response time and signal detection time distributions SM Fig. 1. Correct response time (thick solid green curve) and error response time densities (dashed red curve), averaged across
1 SUPPLEMENTAL MATERIAL Response time and signal detection time distributions SM Fig. 1. Correct response time (thick solid green curve) and error response time densities (dashed red curve), averaged across
Doing Quantitative Research 26E02900, 6 ECTS Lecture 6: Structural Equations Modeling. Olli-Pekka Kauppila Daria Kautto
 Doing Quantitative Research 26E02900, 6 ECTS Lecture 6: Structural Equations Modeling Olli-Pekka Kauppila Daria Kautto Session VI, September 20 2017 Learning objectives 1. Get familiar with the basic idea
Doing Quantitative Research 26E02900, 6 ECTS Lecture 6: Structural Equations Modeling Olli-Pekka Kauppila Daria Kautto Session VI, September 20 2017 Learning objectives 1. Get familiar with the basic idea
Welcome Back! 2/6/18. A. GGSS B. ggss C. ggss D. GgSs E. Ggss. 1. A species of mice can have gray or black fur
 Welcome Back! 2/6/18 1. A species of mice can have gray or black fur and long or short tails. A cross between blackfurred, long-tailed mice and gray-furred, shorttailed mice produce all black-furred, long-tailed
Welcome Back! 2/6/18 1. A species of mice can have gray or black fur and long or short tails. A cross between blackfurred, long-tailed mice and gray-furred, shorttailed mice produce all black-furred, long-tailed
For more information about how to cite these materials visit
 Author(s): Kerby Shedden, Ph.D., 2010 License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution Share Alike 3.0 License: http://creativecommons.org/licenses/by-sa/3.0/
Author(s): Kerby Shedden, Ph.D., 2010 License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution Share Alike 3.0 License: http://creativecommons.org/licenses/by-sa/3.0/
11/18/2013. Correlational Research. Correlational Designs. Why Use a Correlational Design? CORRELATIONAL RESEARCH STUDIES
 Correlational Research Correlational Designs Correlational research is used to describe the relationship between two or more naturally occurring variables. Is age related to political conservativism? Are
Correlational Research Correlational Designs Correlational research is used to describe the relationship between two or more naturally occurring variables. Is age related to political conservativism? Are
Nature Neuroscience: doi: /nn Supplementary Figure 1. Behavioral training.
 Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
White Paper Estimating Complex Phenotype Prevalence Using Predictive Models
 White Paper 23-12 Estimating Complex Phenotype Prevalence Using Predictive Models Authors: Nicholas A. Furlotte Aaron Kleinman Robin Smith David Hinds Created: September 25 th, 2015 September 25th, 2015
White Paper 23-12 Estimating Complex Phenotype Prevalence Using Predictive Models Authors: Nicholas A. Furlotte Aaron Kleinman Robin Smith David Hinds Created: September 25 th, 2015 September 25th, 2015
Performing. linkage analysis using MERLIN
 Performing linkage analysis using MERLIN David Duffy Queensland Institute of Medical Research Brisbane, Australia Overview MERLIN and associated programs Error checking Parametric linkage analysis Nonparametric
Performing linkage analysis using MERLIN David Duffy Queensland Institute of Medical Research Brisbane, Australia Overview MERLIN and associated programs Error checking Parametric linkage analysis Nonparametric
PSYCH-GA.2211/NEURL-GA.2201 Fall 2016 Mathematical Tools for Cognitive and Neural Science. Homework 5
 PSYCH-GA.2211/NEURL-GA.2201 Fall 2016 Mathematical Tools for Cognitive and Neural Science Homework 5 Due: 21 Dec 2016 (late homeworks penalized 10% per day) See the course web site for submission details.
PSYCH-GA.2211/NEURL-GA.2201 Fall 2016 Mathematical Tools for Cognitive and Neural Science Homework 5 Due: 21 Dec 2016 (late homeworks penalized 10% per day) See the course web site for submission details.
Expanded View Figures
 Solip Park & Ben Lehner Epistasis is cancer type specific Molecular Systems Biology Expanded View Figures A B G C D E F H Figure EV1. Epistatic interactions detected in a pan-cancer analysis and saturation
Solip Park & Ben Lehner Epistasis is cancer type specific Molecular Systems Biology Expanded View Figures A B G C D E F H Figure EV1. Epistatic interactions detected in a pan-cancer analysis and saturation
CHAPTER ONE CORRELATION
 CHAPTER ONE CORRELATION 1.0 Introduction The first chapter focuses on the nature of statistical data of correlation. The aim of the series of exercises is to ensure the students are able to use SPSS to
CHAPTER ONE CORRELATION 1.0 Introduction The first chapter focuses on the nature of statistical data of correlation. The aim of the series of exercises is to ensure the students are able to use SPSS to
WDHS Curriculum Map Probability and Statistics. What is Statistics and how does it relate to you?
 WDHS Curriculum Map Probability and Statistics Time Interval/ Unit 1: Introduction to Statistics 1.1-1.3 2 weeks S-IC-1: Understand statistics as a process for making inferences about population parameters
WDHS Curriculum Map Probability and Statistics Time Interval/ Unit 1: Introduction to Statistics 1.1-1.3 2 weeks S-IC-1: Understand statistics as a process for making inferences about population parameters
Selection at one locus with many alleles, fertility selection, and sexual selection
 Selection at one locus with many alleles, fertility selection, and sexual selection Introduction It s easy to extend the Hardy-Weinberg principle to multiple alleles at a single locus. In fact, we already
Selection at one locus with many alleles, fertility selection, and sexual selection Introduction It s easy to extend the Hardy-Weinberg principle to multiple alleles at a single locus. In fact, we already
Small Group Presentations
 Admin Assignment 1 due next Tuesday at 3pm in the Psychology course centre. Matrix Quiz during the first hour of next lecture. Assignment 2 due 13 May at 10am. I will upload and distribute these at the
Admin Assignment 1 due next Tuesday at 3pm in the Psychology course centre. Matrix Quiz during the first hour of next lecture. Assignment 2 due 13 May at 10am. I will upload and distribute these at the
OUTLIER SUBJECTS PROTOCOL (art_groupoutlier)
 OUTLIER SUBJECTS PROTOCOL (art_groupoutlier) Paul K. Mazaika 2/23/2009 Outlier subjects are a problem in fmri data sets for clinical populations. This protocol and program are a method to identify outlier
OUTLIER SUBJECTS PROTOCOL (art_groupoutlier) Paul K. Mazaika 2/23/2009 Outlier subjects are a problem in fmri data sets for clinical populations. This protocol and program are a method to identify outlier
Supplementary Information. Supplementary Figures
 Supplementary Information Supplementary Figures.8 57 essential gene density 2 1.5 LTR insert frequency diversity DEL.5 DUP.5 INV.5 TRA 1 2 3 4 5 1 2 3 4 1 2 Supplementary Figure 1. Locations and minor
Supplementary Information Supplementary Figures.8 57 essential gene density 2 1.5 LTR insert frequency diversity DEL.5 DUP.5 INV.5 TRA 1 2 3 4 5 1 2 3 4 1 2 Supplementary Figure 1. Locations and minor
An Introduction to Quantitative Genetics
 An Introduction to Quantitative Genetics Mohammad Keramatipour MD, PhD Keramatipour@tums.ac.ir ac ir 1 Mendel s work Laws of inheritance Basic Concepts Applications Predicting outcome of crosses Phenotype
An Introduction to Quantitative Genetics Mohammad Keramatipour MD, PhD Keramatipour@tums.ac.ir ac ir 1 Mendel s work Laws of inheritance Basic Concepts Applications Predicting outcome of crosses Phenotype
6. Unusual and Influential Data
 Sociology 740 John ox Lecture Notes 6. Unusual and Influential Data Copyright 2014 by John ox Unusual and Influential Data 1 1. Introduction I Linear statistical models make strong assumptions about the
Sociology 740 John ox Lecture Notes 6. Unusual and Influential Data Copyright 2014 by John ox Unusual and Influential Data 1 1. Introduction I Linear statistical models make strong assumptions about the
Application of Local Control Strategy in analyses of the effects of Radon on Lung Cancer Mortality for 2,881 US Counties
 Application of Local Control Strategy in analyses of the effects of Radon on Lung Cancer Mortality for 2,881 US Counties Bob Obenchain, Risk Benefit Statistics, August 2015 Our motivation for using a Cut-Point
Application of Local Control Strategy in analyses of the effects of Radon on Lung Cancer Mortality for 2,881 US Counties Bob Obenchain, Risk Benefit Statistics, August 2015 Our motivation for using a Cut-Point
Analysis of single gene effects 1. Quantitative analysis of single gene effects. Gregory Carey, Barbara J. Bowers, Jeanne M.
 Analysis of single gene effects 1 Quantitative analysis of single gene effects Gregory Carey, Barbara J. Bowers, Jeanne M. Wehner From the Department of Psychology (GC, JMW) and Institute for Behavioral
Analysis of single gene effects 1 Quantitative analysis of single gene effects Gregory Carey, Barbara J. Bowers, Jeanne M. Wehner From the Department of Psychology (GC, JMW) and Institute for Behavioral
THERE is broad interest in genetic loci (called quantitative
 Copyright Ó 2006 by the Genetics Society of America DOI: 10.1534/genetics.106.061176 The X Chromosome in Quantitative Trait Locus Mapping Karl W. Broman,*,1 Śaunak Sen, Sarah E. Owens,,2 Ani Manichaikul,*
Copyright Ó 2006 by the Genetics Society of America DOI: 10.1534/genetics.106.061176 The X Chromosome in Quantitative Trait Locus Mapping Karl W. Broman,*,1 Śaunak Sen, Sarah E. Owens,,2 Ani Manichaikul,*
Variant Classification. Author: Mike Thiesen, Golden Helix, Inc.
 Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Dan Koller, Ph.D. Medical and Molecular Genetics
 Design of Genetic Studies Dan Koller, Ph.D. Research Assistant Professor Medical and Molecular Genetics Genetics and Medicine Over the past decade, advances from genetics have permeated medicine Identification
Design of Genetic Studies Dan Koller, Ph.D. Research Assistant Professor Medical and Molecular Genetics Genetics and Medicine Over the past decade, advances from genetics have permeated medicine Identification
List of Figures. List of Tables. Preface to the Second Edition. Preface to the First Edition
 List of Figures List of Tables Preface to the Second Edition Preface to the First Edition xv xxv xxix xxxi 1 What Is R? 1 1.1 Introduction to R................................ 1 1.2 Downloading and Installing
List of Figures List of Tables Preface to the Second Edition Preface to the First Edition xv xxv xxix xxxi 1 What Is R? 1 1.1 Introduction to R................................ 1 1.2 Downloading and Installing
SPRING GROVE AREA SCHOOL DISTRICT. Course Description. Instructional Strategies, Learning Practices, Activities, and Experiences.
 SPRING GROVE AREA SCHOOL DISTRICT PLANNED COURSE OVERVIEW Course Title: Basic Introductory Statistics Grade Level(s): 11-12 Units of Credit: 1 Classification: Elective Length of Course: 30 cycles Periods
SPRING GROVE AREA SCHOOL DISTRICT PLANNED COURSE OVERVIEW Course Title: Basic Introductory Statistics Grade Level(s): 11-12 Units of Credit: 1 Classification: Elective Length of Course: 30 cycles Periods
Introduction to LOH and Allele Specific Copy Number User Forum
 Introduction to LOH and Allele Specific Copy Number User Forum Jonathan Gerstenhaber Introduction to LOH and ASCN User Forum Contents 1. Loss of heterozygosity Analysis procedure Types of baselines 2.
Introduction to LOH and Allele Specific Copy Number User Forum Jonathan Gerstenhaber Introduction to LOH and ASCN User Forum Contents 1. Loss of heterozygosity Analysis procedure Types of baselines 2.
25.1 QUANTITATIVE TRAITS
 CHAPTER OUTLINE 5.1 Quantitative Traits 5. Polygenic Inheritance 5.3 Heritability 5 QUANTITATIVE In this chapter, we will examine complex traits characteristics that are determined by several genes and
CHAPTER OUTLINE 5.1 Quantitative Traits 5. Polygenic Inheritance 5.3 Heritability 5 QUANTITATIVE In this chapter, we will examine complex traits characteristics that are determined by several genes and
HERITABILITY INTRODUCTION. Objectives
 36 HERITABILITY In collaboration with Mary Puterbaugh and Larry Lawson Objectives Understand the concept of heritability. Differentiate between broad-sense heritability and narrowsense heritability. Learn
36 HERITABILITY In collaboration with Mary Puterbaugh and Larry Lawson Objectives Understand the concept of heritability. Differentiate between broad-sense heritability and narrowsense heritability. Learn
CSE 255 Assignment 9
 CSE 255 Assignment 9 Alexander Asplund, William Fedus September 25, 2015 1 Introduction In this paper we train a logistic regression function for two forms of link prediction among a set of 244 suspected
CSE 255 Assignment 9 Alexander Asplund, William Fedus September 25, 2015 1 Introduction In this paper we train a logistic regression function for two forms of link prediction among a set of 244 suspected
Identification of Tissue Independent Cancer Driver Genes
 Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
Reveal Relationships in Categorical Data
 SPSS Categories 15.0 Specifications Reveal Relationships in Categorical Data Unleash the full potential of your data through perceptual mapping, optimal scaling, preference scaling, and dimension reduction
SPSS Categories 15.0 Specifications Reveal Relationships in Categorical Data Unleash the full potential of your data through perceptual mapping, optimal scaling, preference scaling, and dimension reduction
Single SNP/Gene Analysis. Typical Results of GWAS Analysis (Single SNP Approach) Typical Results of GWAS Analysis (Single SNP Approach)
 High-Throughput Sequencing Course Gene-Set Analysis Biostatistics and Bioinformatics Summer 28 Section Introduction What is Gene Set Analysis? Many names for gene set analysis: Pathway analysis Gene set
High-Throughput Sequencing Course Gene-Set Analysis Biostatistics and Bioinformatics Summer 28 Section Introduction What is Gene Set Analysis? Many names for gene set analysis: Pathway analysis Gene set
OECD QSAR Toolbox v.4.2. An example illustrating RAAF scenario 6 and related assessment elements
 OECD QSAR Toolbox v.4.2 An example illustrating RAAF scenario 6 and related assessment elements Outlook Background Objectives Specific Aims Read Across Assessment Framework (RAAF) The exercise Workflow
OECD QSAR Toolbox v.4.2 An example illustrating RAAF scenario 6 and related assessment elements Outlook Background Objectives Specific Aims Read Across Assessment Framework (RAAF) The exercise Workflow
Exercises: Differential Methylation
 Exercises: Differential Methylation Version 2018-04 Exercises: Differential Methylation 2 Licence This manual is 2014-18, Simon Andrews. This manual is distributed under the creative commons Attribution-Non-Commercial-Share
Exercises: Differential Methylation Version 2018-04 Exercises: Differential Methylation 2 Licence This manual is 2014-18, Simon Andrews. This manual is distributed under the creative commons Attribution-Non-Commercial-Share
Chapter 3 Software Packages to Install How to Set Up Python Eclipse How to Set Up Eclipse... 42
 Table of Contents Preface..... 21 About the Authors... 23 Acknowledgments... 24 How This Book is Organized... 24 Who Should Buy This Book?... 24 Where to Find Answers to Review Questions and Exercises...
Table of Contents Preface..... 21 About the Authors... 23 Acknowledgments... 24 How This Book is Organized... 24 Who Should Buy This Book?... 24 Where to Find Answers to Review Questions and Exercises...
Laboratory. Mendelian Genetics
 Laboratory 9 Mendelian Genetics Biology 171L FA17 Lab 9: Mendelian Genetics Student Learning Outcomes 1. Predict the phenotypic and genotypic ratios of a monohybrid cross. 2. Determine whether a gene is
Laboratory 9 Mendelian Genetics Biology 171L FA17 Lab 9: Mendelian Genetics Student Learning Outcomes 1. Predict the phenotypic and genotypic ratios of a monohybrid cross. 2. Determine whether a gene is
Name Class Date. KEY CONCEPT The chromosomes on which genes are located can affect the expression of traits.
 Section 1: Chromosomes and Phenotype KEY CONCEPT The chromosomes on which genes are located can affect the expression of traits. VOCABULARY carrier sex-linked gene X chromosome inactivation MAIN IDEA:
Section 1: Chromosomes and Phenotype KEY CONCEPT The chromosomes on which genes are located can affect the expression of traits. VOCABULARY carrier sex-linked gene X chromosome inactivation MAIN IDEA:
Supplementary Material Estimating the Distribution of Sensorimotor Synchronization Data: A Bayesian Hierarchical Modeling Approach Rasmus Bååth
 Supplementary Material Estimating the Distribution of Sensorimotor Synchronization Data: A Bayesian Hierarchical Modeling Approach Rasmus Bååth One advantage of working within a Bayesian statistical framework
Supplementary Material Estimating the Distribution of Sensorimotor Synchronization Data: A Bayesian Hierarchical Modeling Approach Rasmus Bååth One advantage of working within a Bayesian statistical framework
Early Learning vs Early Variability 1.5 r = p = Early Learning r = p = e 005. Early Learning 0.
 The temporal structure of motor variability is dynamically regulated and predicts individual differences in motor learning ability Howard Wu *, Yohsuke Miyamoto *, Luis Nicolas Gonzales-Castro, Bence P.
The temporal structure of motor variability is dynamically regulated and predicts individual differences in motor learning ability Howard Wu *, Yohsuke Miyamoto *, Luis Nicolas Gonzales-Castro, Bence P.
Overview of Animal Breeding
 Overview of Animal Breeding 1 Required Information Successful animal breeding requires 1. the collection and storage of data on individually identified animals; 2. complete pedigree information about the
Overview of Animal Breeding 1 Required Information Successful animal breeding requires 1. the collection and storage of data on individually identified animals; 2. complete pedigree information about the
Chapter 1: Introduction to Statistics
 Chapter 1: Introduction to Statistics Variables A variable is a characteristic or condition that can change or take on different values. Most research begins with a general question about the relationship
Chapter 1: Introduction to Statistics Variables A variable is a characteristic or condition that can change or take on different values. Most research begins with a general question about the relationship
Introduction to linkage and family based designs to study the genetic epidemiology of complex traits. Harold Snieder
 Introduction to linkage and family based designs to study the genetic epidemiology of complex traits Harold Snieder Overview of presentation Designs: population vs. family based Mendelian vs. complex diseases/traits
Introduction to linkage and family based designs to study the genetic epidemiology of complex traits Harold Snieder Overview of presentation Designs: population vs. family based Mendelian vs. complex diseases/traits
Chapter 1: Exploring Data
 Chapter 1: Exploring Data Key Vocabulary:! individual! variable! frequency table! relative frequency table! distribution! pie chart! bar graph! two-way table! marginal distributions! conditional distributions!
Chapter 1: Exploring Data Key Vocabulary:! individual! variable! frequency table! relative frequency table! distribution! pie chart! bar graph! two-way table! marginal distributions! conditional distributions!
Problem 3: Simulated Rheumatoid Arthritis Data
 Problem 3: Simulated Rheumatoid Arthritis Data Michael B Miller Michael Li Gregg Lind Soon-Young Jang The plan
Problem 3: Simulated Rheumatoid Arthritis Data Michael B Miller Michael Li Gregg Lind Soon-Young Jang The plan
Assignment #6. Chapter 10: 14, 15 Chapter 11: 14, 18. Due tomorrow Nov. 6 th by 2pm in your TA s homework box
 Assignment #6 Chapter 10: 14, 15 Chapter 11: 14, 18 Due tomorrow Nov. 6 th by 2pm in your TA s homework box Assignment #7 Chapter 12: 18, 24 Chapter 13: 28 Due next Friday Nov. 13 th by 2pm in your TA
Assignment #6 Chapter 10: 14, 15 Chapter 11: 14, 18 Due tomorrow Nov. 6 th by 2pm in your TA s homework box Assignment #7 Chapter 12: 18, 24 Chapter 13: 28 Due next Friday Nov. 13 th by 2pm in your TA
IN SILICO EVALUATION OF DNA-POOLED ALLELOTYPING VERSUS INDIVIDUAL GENOTYPING FOR GENOME-WIDE ASSOCIATION STUDIES OF COMPLEX DISEASE.
 IN SILICO EVALUATION OF DNA-POOLED ALLELOTYPING VERSUS INDIVIDUAL GENOTYPING FOR GENOME-WIDE ASSOCIATION STUDIES OF COMPLEX DISEASE By Siddharth Pratap Thesis Submitted to the Faculty of the Graduate School
IN SILICO EVALUATION OF DNA-POOLED ALLELOTYPING VERSUS INDIVIDUAL GENOTYPING FOR GENOME-WIDE ASSOCIATION STUDIES OF COMPLEX DISEASE By Siddharth Pratap Thesis Submitted to the Faculty of the Graduate School
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
Gregor Mendel and Genetics Worksheets
 Gregor Mendel and Genetics Worksheets Douglas Wilkin, Ph.D. (DWilkin) Say Thanks to the Authors Click http://www.ck12.org/saythanks (No sign in required) To access a customizable version of this book,
Gregor Mendel and Genetics Worksheets Douglas Wilkin, Ph.D. (DWilkin) Say Thanks to the Authors Click http://www.ck12.org/saythanks (No sign in required) To access a customizable version of this book,
Mendelian & Complex Traits. Quantitative Imaging Genomics. Genetics Terminology 2. Genetics Terminology 1. Human Genome. Genetics Terminology 3
 Mendelian & Complex Traits Quantitative Imaging Genomics David C. Glahn, PhD Olin Neuropsychiatry Research Center & Department of Psychiatry, Yale University July, 010 Mendelian Trait A trait influenced
Mendelian & Complex Traits Quantitative Imaging Genomics David C. Glahn, PhD Olin Neuropsychiatry Research Center & Department of Psychiatry, Yale University July, 010 Mendelian Trait A trait influenced
Confidence Intervals On Subsets May Be Misleading
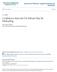 Journal of Modern Applied Statistical Methods Volume 3 Issue 2 Article 2 11-1-2004 Confidence Intervals On Subsets May Be Misleading Juliet Popper Shaffer University of California, Berkeley, shaffer@stat.berkeley.edu
Journal of Modern Applied Statistical Methods Volume 3 Issue 2 Article 2 11-1-2004 Confidence Intervals On Subsets May Be Misleading Juliet Popper Shaffer University of California, Berkeley, shaffer@stat.berkeley.edu
B-4.7 Summarize the chromosome theory of inheritance and relate that theory to Gregor Mendel s principles of genetics
 B-4.7 Summarize the chromosome theory of inheritance and relate that theory to Gregor Mendel s principles of genetics The Chromosome theory of inheritance is a basic principle in biology that states genes
B-4.7 Summarize the chromosome theory of inheritance and relate that theory to Gregor Mendel s principles of genetics The Chromosome theory of inheritance is a basic principle in biology that states genes
The North Carolina Health Data Explorer
 The North Carolina Health Data Explorer The Health Data Explorer provides access to health data for North Carolina counties in an interactive, user-friendly atlas of maps, tables, and charts. It allows
The North Carolina Health Data Explorer The Health Data Explorer provides access to health data for North Carolina counties in an interactive, user-friendly atlas of maps, tables, and charts. It allows
Package noia. February 20, 2015
 Type Package Package noia February 20, 2015 Title Implementation of the Natural and Orthogonal InterAction (NOIA) model Version 0.97.1 Date 2015-01-06 Author Arnaud Le Rouzic (2007-2014), Arne B. Gjuvsland
Type Package Package noia February 20, 2015 Title Implementation of the Natural and Orthogonal InterAction (NOIA) model Version 0.97.1 Date 2015-01-06 Author Arnaud Le Rouzic (2007-2014), Arne B. Gjuvsland
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
MMI 409 Spring 2009 Final Examination Gordon Bleil. 1. Is there a difference in depression as a function of group and drug?
 MMI 409 Spring 2009 Final Examination Gordon Bleil Table of Contents Research Scenario and General Assumptions Questions for Dataset (Questions are hyperlinked to detailed answers) 1. Is there a difference
MMI 409 Spring 2009 Final Examination Gordon Bleil Table of Contents Research Scenario and General Assumptions Questions for Dataset (Questions are hyperlinked to detailed answers) 1. Is there a difference
CS2220 Introduction to Computational Biology
 CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
Chapter 2. Linkage Analysis. JenniferH.BarrettandM.DawnTeare. Abstract. 1. Introduction
 Chapter 2 Linkage Analysis JenniferH.BarrettandM.DawnTeare Abstract Linkage analysis is used to map genetic loci using observations on relatives. It can be applied to both major gene disorders (parametric
Chapter 2 Linkage Analysis JenniferH.BarrettandM.DawnTeare Abstract Linkage analysis is used to map genetic loci using observations on relatives. It can be applied to both major gene disorders (parametric
Chapter 1: Explaining Behavior
 Chapter 1: Explaining Behavior GOAL OF SCIENCE is to generate explanations for various puzzling natural phenomenon. - Generate general laws of behavior (psychology) RESEARCH: principle method for acquiring
Chapter 1: Explaining Behavior GOAL OF SCIENCE is to generate explanations for various puzzling natural phenomenon. - Generate general laws of behavior (psychology) RESEARCH: principle method for acquiring
Chapter 15: The Chromosomal Basis of Inheritance
 Name Period Chapter 15: The Chromosomal Basis of Inheritance Concept 15.1 Mendelian inheritance has its physical basis in the behavior of chromosomes 1. What is the chromosome theory of inheritance? 2.
Name Period Chapter 15: The Chromosomal Basis of Inheritance Concept 15.1 Mendelian inheritance has its physical basis in the behavior of chromosomes 1. What is the chromosome theory of inheritance? 2.
Structural Variation and Medical Genomics
 Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
SNPrints: Defining SNP signatures for prediction of onset in complex diseases
 SNPrints: Defining SNP signatures for prediction of onset in complex diseases Linda Liu, Biomedical Informatics, Stanford University Daniel Newburger, Biomedical Informatics, Stanford University Grace
SNPrints: Defining SNP signatures for prediction of onset in complex diseases Linda Liu, Biomedical Informatics, Stanford University Daniel Newburger, Biomedical Informatics, Stanford University Grace
Bangor University Laboratory Exercise 1, June 2008
 Laboratory Exercise, June 2008 Classroom Exercise A forest land owner measures the outside bark diameters at.30 m above ground (called diameter at breast height or dbh) and total tree height from ground
Laboratory Exercise, June 2008 Classroom Exercise A forest land owner measures the outside bark diameters at.30 m above ground (called diameter at breast height or dbh) and total tree height from ground
Genetics of extreme body size evolution in mice from Gough Island
 Genetics of extreme body size evolution in mice from Gough Island Karl Broman Biostatistics & Medical Informatics, UW Madison kbroman.org github.com/kbroman @kwbroman bit.ly/apstats2017 This is a collaboration
Genetics of extreme body size evolution in mice from Gough Island Karl Broman Biostatistics & Medical Informatics, UW Madison kbroman.org github.com/kbroman @kwbroman bit.ly/apstats2017 This is a collaboration
Research Methods in Forest Sciences: Learning Diary. Yoko Lu December Research process
 Research Methods in Forest Sciences: Learning Diary Yoko Lu 285122 9 December 2016 1. Research process It is important to pursue and apply knowledge and understand the world under both natural and social
Research Methods in Forest Sciences: Learning Diary Yoko Lu 285122 9 December 2016 1. Research process It is important to pursue and apply knowledge and understand the world under both natural and social
