Correction Notice: Structural insight into mammalian sialyltransferases
|
|
|
- Dorothy Ball
- 5 years ago
- Views:
Transcription
1 CORRECTION NOTICE Correction Notice: Structural insight into mammalian sialyltransferases Francesco V Rao, Jamie R Rich, Bojana Rakić, Sai Buddai, Marc F Schwartz, Karl Johnson, Caryn Bowe, Warren W Wakarchuk, Shawn DeFrees, Stephen G Withers & Natalie C J Strynadka Nat. Struct. Mol. Biol. 16, (2009) In the version of the supplementary file originally posted online, the acknowledgements were incomplete. The error has been corrected in this file as of 18 November 2009.
2 Supporting Online Material for Structural Insight into Mammalian Sialyltransferases Francesco V. Rao, Jamie R. Rich, Bojana Rakić, Sai Buddai, Marc F. Schwartz, Karl Johnson, Caryn Bowe, Warren W. Wakarchuk, Shawn DeFrees, Stephen G. Withers, Natalie C. J. Strynadka * * To whom correspondence should be addressed. natalie@byron.biochem.ubc.ca; Tel.: ; Fax: This PDF file includes: Supplementary Figure 1-4 Supplementary Table 1-2 Materials and Methods References for Supporting Online Material Acknowledgments Author Contributions S1
3 Supplementary Figure 1 Supplementary Fig. 1 Comparison of pst3gal-i (GT-29) with CstII (GT-42). The structures are shown in a ribbon representation, with red helices and blue strands. The lid domain of CstII (PDB entry 1RO7 1 ), is represented in magenta (b) and is modeled as a dashed line on pst3gal-i (a). The ligands are shown in a stick representation with yellow carbon atoms, and the catalytic base in a stick representation with cyan carbon atoms. The disulphide-linked cysteines in pst3gal-i are represented in green and labeled C1 to C3. Residues predicted to be glycosylated are shown in stick representation with black carbon atoms. (c) depicts a structure-based sequence alignment of pst3gal-i (GT- 29) and CstII (GT-42). Sequences were aligned using the program LSQMAN using the Needleman-Wunsch alignment algorithm and applying a 5 Å cut-off 2. Secondary structure and location of the lid domain for pst3gal-i and CstII are shown above and below each sequence respectively following the color scheme in (a). Identical residues and those that align structurally are depicted in black and grey respectively. The catalytic base is represented with a cyan colored box. Residues from the lid domain of pst3gal-i have not been included in the alignment as they are disordered in the crystal structure. (d) shows topology diagrams of GT-2, GT-42 and GT-29. The color scheme for secondary structure elements follows that of (a). Figures were created using TopDraw 3. S2
4 Supplementary Figure 2 Supplementary Fig. 2 Active site comparisons and reaction scheme. The pst3gal-1- disaccharide-cmp complex (a) is compared to the CstII-CMP3FNeuAc complex (GT- 42) (PDB entry 1RO7 1 ) (b), Δ24PmST1-lactose-CMP3FNeuAc (GT-80) (c) (PDB entry 2IHZ 4 ) and C2GnT-I-Gal β1,3galnac (GT-14) (d) (PDB entry 2GAM 5 ). The first domain of Δ24PmST1 (GT-B fold) is shown with pink carbon atoms (b). The proposed catalytic base for each structure is shown with cyan colored carbon atoms. Hydrogen bond interactions are shown with dashed lines. Fig. 2e. Reaction scheme of pst3gal-i. His319 deprotonates the acceptor sugar at the O3 position, resulting in nucleophilic attack on the anomeric (C2) carbon of Neu5Ac. This leads to breakage of the C2-O bond linking Neu5Ac with CMP. The oxocarbenium-ion like transition state is defined within the large brackets. S3
5 Supplementary Figure 3 Supplementary Fig. 3. Sequence alignment of pst3gal-i with other ST3 and ST6Gal-I. The pst3gal-i secondary structure and location of the lid domain are shown using the color scheme in Fig. 1a. Cysteines involved in a disulphide bond are indicated with a green triangle. Residues interacting with CMP or the disaccharide acceptor are indicated by purple and yellow diamonds respectively. Three of the predicted glycosylation sites are indicated with yellow squares. The first glycosylation site is predicted to be located on the stem region (not shown). The proposed catalytic base is represented with a cyan colored star. Sialyl motifs are represented in labeled colored boxes following the color scheme in Fig. 1b. Numbering of the above sequences is according to the UniProt accession numbers; Q02745, Q11201, Q16842, Q11203, Q11206, Q9UNP4, Q9Y274, P15907, respectively. Amino acids completely conserved or greater than 50% conserved are depicted in black and grey respectively. The sequence alignment was performed using the program ALINE 6. S4
6 Supplementary Figure 4 Supplementary Fig. 4. Surface sequence conservation. The protein surface of pst3gal-i-disaccharide-cmp complex is shown in three different colors to reflect sequence conservation with hst6gal-i, based on the sequence alignment in Fig. 3. Dark blue denotes identical surface residues, with light blue and grey indicating similar and no sequence conservation, respectively. The location of the catalytic residue is depicted in orange. The disaccharide acceptor and CMP are shown in stick representations with yellow carbon atoms. A polar cleft (depicted in green) of suitable dimensions to bind a potential peptide or lipid (found in natural acceptors of GT-29) is shown extending appropriately from the terminal sugar of the disaccharide acceptor towards solvent. S5
7 Supplementary table 1. Kinetic parameters Substrate varied (reaction monitored) protein k cat (min -1 ) K m (mm) k cat / K m (mm -1 min -1 ) CMPNeu5Ac (hydrolase) a pst3gal-i n.d n.d n.d CMPNeu5Ac (transferase) b pst3gal-i Gal-β-1,3GalNAc-α-OBn c pst3gal-i CMPNeu5Ac (hydrolase) a Cst II CMPNeu5Ac (transferase) d Cst II Lactose f Cst II '-Sialyl Lactose f Cst II *- error range in data is between 5-15% n.d no significant value was detected. a- determined in the absence of acceptor substrate b- determined at a constant Gal-β-1,3GalNac-α-OBn concentration of 0.5 mm c- determined at a constant CMPNeu5Ac concentration of 0.3 mm d- determined at a constant lactose concentration of 160 mm f- determined at a constant CMPNeu5Ac concentration of 1 mm Comparison with the bacterial CstII sialyltransferase reveals that pst3gal-i is a much more efficient enzyme with K m values for CMP-Neu5Ac five times lower and k cat values thirty-five times higher. Interestingly, whereas CstII has been shown to additionally catalyze the hydrolysis of CMP-Neu5Ac at a significant rate in the absence of acceptor substrate, pst3gal-i has no such activity. S6
8 Supplementary table 2. Data collection, phasing and refinement statistics pst3gal-i-semet pst3gal-i-disaccharide pst3gal-i-cmp-disaccharide Data collection Space group I222 I222 I222 Cell dimensions a, b, c (Å) 73.38, 78.01, , 78.91, , 78.43, α, β, γ ( ) 90.00, 90.00, , 90.00, , 90.00, Peak Inflection Remote Wavelength Resolution (Å) ( ) ( ( ) ( ) ( ) * 1.8) R merge (0.333) (0.332) (0.504) (0.399) (0.252) I / σi 18.2 (3.4) 22.6 (3.8) 16.6 (3.3) 23.4 (2.8) 19.3 (3.5) Completeness (%) 98.3 (92.2) 98.8 (89.5) 99.3 (99.0) 99.8 (98.3) 85.0 (83.0) Redundancy 6.2 (4.5) 6.2 (4.4) 6.4 (4.5) 6.9 (4.9) 1.9 (1.7) Refinement Resolution (Å) No. reflections R work / R free 0.18/ / /0.20 No. atoms Protein Ligand/ion Water B-factors Protein Ligand/ion Water R.m.s deviations Bond lengths (Å) Bond angles ( ) *Values in parentheses are for highest-resolution shell. S7
9 Materials & Methods Cloning, expression and crystallization DNA encoding porcine ST3Gal-I (Genbank accession NP ) starting at Glu46 was synthesized with codon selection optimized for bacterial expression (Blue Heron Biotechnology). The pst3gal-i open reading frame was cloned by PCR, restriction digest, and ligation into the BamHI and XhoI sites of pcwin2-mbp 11. For protein expression, pcwin2-mbp-pst3gal-i was transformed into Origami- 2 cells (Novagen). Cells were grown in rich media (typically LB or 2BN (2% martone B- 1 (Marcor), 2% yeast extract, trace metals 12, 1% NaCl, and 1 mm MgSO 4 ), with kanamycin at 50 µg ml -1 ) to mid-logarithmic phase, shifted to 20 C for at least 30 minutes, and induced overnight with 0.2 mm IPTG. Induced cells were harvested by centrifugation and stored at -80 C. Induced cells were thawed and resuspended in lysis buffer (10 mm Tris ph 8.2, 5 mm NaCl) at 6-10 ml g -1 wet cells, and lysed with two to four passes through a microfluidizer. The lysate was clarified by centrifugation for one hour at 27,000 x g, 4 C, concentrated approximately five-fold, and diafiltered against lysis buffer to 1.4 ms cm -1 using two Pellicon-2 mini ultrafiltration cassettes (Millipore). The lysate was applied to a Source 30Q column (GE Healthcare) pre-equilibrated with 20 mm Tris ph 8.2, 10 mm NaCl, and eluted with a gradient to 100 mm NaCl. Fractions containing active, full length MBP-tagged ST3Gal-I were pooled and diluted three-fold with 20 mm Tris ph 8.2. The diluted material was then applied to a Source 15Q column (GE Healthcare) preequilibrated with Q column buffer, and eluted with a gradient to 80 mm NaCl. Fractions containing active, full length MBP-tagged pst3gal-i were pooled and buffer exchanged to 10 mm sodium phosphate, ph 8.0, 10 mm NaCl by centrifugal ultrafiltration (Centricon Plus 70, Millipore). The material was then applied to a ceramic hydroxyapatite column (20 µm, Type-I, Bio-Rad) pre-equilibrated with 10 mm sodium phosphate, ph 8.0, 10 mm NaCl, and eluted with a gradient to 130 mm sodium phosphate. Fractions containing active, full length MBP-tagged pst3gal-i were pooled, concentrated, buffer exchanged to 20 mm Tris ph 8.2, 10 mm NaCl by centrifugal ultrafiltration, and mixed with 1 volume of glycerol prior to storage at -20 C. For the expression of selenomethionine (Se-Met)-labeled MBP-pST3Gal-I, cells were grown and induced under similar conditions using synthetic complete media (50 mm KH 2 PO 4 and Na 2 HPO 4, 25 mm (NH 4 ) 2 SO 4, 0.5% glycerol, 0.05% glucose, 1 mm MgSO 4, trace metals, 500 mg l -1 adenine, guanosine, thymine, and uracil, 15 mg l -1 p- aminobenzoic acid, 1 mg l -1 thiamine, 1 mg l -1 biotin, 304 mg l -1 leucine, 150 mg l -1 myoinositol, alanine, arginine, asparagine, aspartate, cysteine, glutamine, glutamate, glycine, histidine, isoleucine, lysine, phenylalanine, proline, serine, tryptophan, tyrosine, valine, and selenomethionine (media recipe modified from 12,13. Se-Met-labeled MBP-tagged pst3gal-i was purified in smaller lots using methods similar to those described above. Purified MBP-tagged pst3gal-i was digested with 25 Units of Factor Xa (Novagen) and dialyzed overnight into 25 mm Tris ph 8.2, 150 mm NaCl, at 4 C. Dialyzed protein sample was applied to a gravity column (Bio-Rad) containing 10 ml of pre-equilibrated (25 mm Tris ph 8.2, 150 mm NaCl) amylose resin (NEB) and washed with 3 column volumes of buffer. The flow-through was applied to a gravity column (Bio-Rad) containing 2 ml of pre-equilibrated Xarrest agarose (Novagen) and washed S8
10 with 3 column volumes of buffer. The flow-through containing pure pst3gal-i was spin concentrated to approximately 5 mg ml -1. Microbatch crystallization experiments were set-up by mixing 0.5 µl of protein and 0.5 µl of mother liquor (30% PEG 4000 and 0.1 M MES, ph 6.8), which was overlaid with Al s oil. Thin plate-shaped crystals would appear after 6 days, growing to a maximum size of approximately 0.2 x 0.2 x 0.1 mm. Data Collection, structure solution and refinement The crystals were cryoprotected by a five second immersion in a solution containing 27% PEG 4000, 0.09 M MES, ph 6.8 and 10% glycerol, and frozen in liquid nitrogen. A complex with a synthetic acceptor was obtained by adding 10 mm Galβ1,3GalNAc α-onitrophenyl to the protein and mother liquor drop. A complex with CMP was obtained by adding 0.2 µl of 10 mm CMP to the crystals for 3 minutes, followed by cryoprotection. Co-crystallization and soaking experiments with the inert donor substrate analog CMP- 3FNeu5Ac were unsuccessful. A six-fold redundant 1.8 Å MAD data set was collected from Se-Met labeled protein crystals, at beamline at the Advanced Light Source (Berkeley, CA). Data were processed using MOSFLM and scaled with SCALA 14. Initial phases were calculated with SOLVE 15 (figure of merit = 0.51), which located 3 Se sites (from a possible 4). Solvent flattening and initial model building with RESOLVE 16 yielded an interpretable 1.8 Å experimental map. This was used as input for ArpWarp 17, which was able to build 281 out of 298 possible residues. Iterative model building with COOT 18 and refinement with REFMAC 19 and PHENIX 20, yielded the final model with the statistics shown in Table S1. Topologies for the ligands were generated with PRODRG 21. Enzymology A continuous coupled assay 22, in which the release of CMP is coupled to the oxidation of NADH ( λ = 340nm, ε = 6.22 mm -1 cm -1 ), was used to monitor the activity of pst3gal-i. Absorbance measurements were obtained using a Cary 300 UV-Vis spectrophotometer equipped with a circulating water bath. GraFit version (Erithacus) was used to calculate kinetic parameters by direct fit of initial rates to the Michaelis-Menten equation. The assay conditions in the 200 µl quartz cuvette are as follows: 20 mm HEPES (ph 7.5), 50 mm KCl, 10 mm MgCl 2, 10 mm MnCl 2, 0.7 mm PEP, 0.25 mm or 0.3 mm NADH, 2 mm ATP, 1.0 mg ml -1 BSA, 5.9U of PK, 8.4U of LDH, 7.5U of NMPK, and varying concentrations of donor and acceptor substrates. The cuvettes were left at 37 C until a stable rate of decreasing absorbance at 340 nm was established. This period is required to deplete the CMP present in the donor solution and any NDPs present in the ATP or NMPK solutions. The stable rate of change in absorbance is the result of the spontaneous hydrolysis of the relatively labile CMP Neu5Ac substrate. Once a stable background rate is established (20-30 min), 4 µl of transferase enzyme solution was added to each cuvette and the solutions were mixed thoroughly to initiate the assay. The concentration of enzyme was chosen such that the rate of chance in absorbance resulting from transferase activity is constant for a period of more than 10 minutes. The initial rate of transferase activity (µm min. -1 ) is calculated as follows (((Slope Background) /2) / 6220 M -1 cm -1 ) x A simple TLC assay was used to monitor the sialylation of bodipy-labeled acceptor substrates. Direct comparison of reactions carried out in the presence of EDTA S9
11 with those carried out in the presence of Mn 2+ or Mg 2+ revealed no significant rate differences. Synthesis 2-Nitrophenyl 2-acetamido-2-deoxy-(b-D-galactopyranosyl)-(1 3)-a-Dgalactopyranoside. 2-Nitrophenyl 2-acetamido-2-deoxy-(β-D-galactopyranosyl)-(1 3)-α-Dgalactopyranoside 2-nitrophenyl 2-acetamido-2-deoxy-α-D-galactopyranoside (5.8 mg; 17 µmol; 5.6 mm in reaction) and UDP-galactopyranose, disodium salt (11.8 mg; 19 µmol; 6.3 mm in reaction) were combined as solids and dissolved in 2290 µl of deionized water. To this solution were added sequentially NaOAc, ph 6.0 (300 µl of 500 mm; 50 mm in reaction), MnCl 2 (300 µl of 100 mm; 10 mm in reaction), DTT (9 µl of 325 mm; 1 mm in reaction), recombinant β1,3-galactosyltransferase (CgtB OH4384 ΔC30- C-term MalE; 100 µl of 5 mg ml -1 in 50 mm NaOAc, ph 6.0), 23 and alkaline phosphatase (Sigma, 0.5 µl; 65 units). After 24 hours, the reaction mixture was applied to a plus tc18 SepPak cartridge (1g; Waters), prewashed with MeOH (8 ml) and equilibrated with water (15 ml). Elution was performed with water (20 ml), followed by a gradient of methanol in water. The title compound (6.9 mg; 14 µmol; 81%) eluted at 25% MeOH. 1 H NMR (300MHz, D 2 O, δ) 7.84 (dd, 1H, Ar, J = 1.6, 8.2 Hz), 7.54 (ddd, 1H, Ar, J = 1.5, 7.6, 8.5 Hz), 7.35 (d, 1H, Ar, J = 8.3 Hz), 7.13 (dt, 1H, Ar, J = 0.7, 7.6 Hz), 5.75 (d, 1H, H-1, J 1,2 = 3.7 Hz), 4.46 (d, 1H, H-1, J 1,2 = 7.5 Hz), 4.43 (dd, 1H, H-2, J 2,3 = 11.3 Hz), 4.23 (d, 1H, H-4, J 4,3 = 2.9 Hz), 4.12 (dd, 1H, H-3), 3.98 (dd, 1H, H-5 or H-5, J = 4.6, 6.1 Hz), 3.82 (d, 1H, H-4, J 4,3 = 3.2 Hz), (m, 5H, H-6a, H-6b, H-6a, H-6b, H-5 or H-5 ), 3.54 (dd, 1H, H-3, J 3,2 = 9.9 Hz), 3.43 (dd, 1H, H-2 ), 1.91 (s, 3H, NHAc). 13 C NMR (75 MHz, D 2 O, δ) (C=O), 148.9, 140.2, 135.1, 125.8, 123.1, 118.3, 104.8, 97.0, 77.2, 75.1, 72.7, 72.4, 70.8, 68.7, 68.5, 61.08, 61.06, 48.5, HRMS calc d for C 20 H 28 N 2 O 13 Na: Found: References for supporting online material 1. Chiu, C.P. et al. Nat Struct Mol Biol 11, (2004). 2. Kleywegt, G.J. Acta Crystallogr D Biol Crystallogr 55, (1999). 3. Bond, C.S. Bioinformatics 19, (2003). 4. Ni, L. et al. Biochemistry 46, (2007). 5. Pak, J.E. et al. J Biol Chem 281, (2006). 6. Bond, C.S. & Schuttelkopf, A.W. Acta Crystallogr D Biol Crystallogr 65, (2009). 7. Jeanneau, C. et al. J Biol Chem 279, (2004). 8. Vallejo-Ruiz, V. et al. Biochim Biophys Acta 1549, (2001). 9. Kono, M. et al. Glycobiology 7, (1997). 10. Rearick, J.I., Sadler, J.E., Paulson, J.C. & Hill, R.L. J Biol Chem 254, (1979). 11. Johnson, K.F., Bezila, D., Ngo, W., and Hakes, D PCT patent application publication WO2005/ Studier, F.W. Protein Expr Purif 41, (2005). 13. Doublie, S. Methods Mol Biol 363, (2007). 14. The CCP4 suite: programs for protein crystallography. Acta Crystallogr D Biol Crystallogr 50, (1994). 15. Terwilliger, T.C.. Acta Crystallogr D Biol Crystallogr 55, (1999). 16. Terwilliger, T.C. Acta Crystallogr D Biol Crystallogr 59, (2003). 17. Perrakis, A., Morris, R. & Lamzin, V.S. Nat Struct Biol 6, (1999). 18. Emsley, P. & Cowtan, K. Acta Crystallogr D Biol Crystallogr 60, (2004). 19. Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Acta Crystallogr D Biol Crystallogr 53, (1997). 20. Adams, P.D. et al. Acta Crystallogr D Biol Crystallogr 58, (2002). 21. Schuttelkopf, A.W. & van Aalten, D.M. Acta Crystallogr D Biol Crystallogr 60, (2004). 22. Gosselin, S., Alhussaini, M., Streiff, M.B., Takabayashi, K. & Palcic, M.M. Anal Biochem 220, 92-7 (1994). 23. Bernatchez, S. et al. Glycobiology 17, (2007). S10
12 Acknowledgments We thank the staff at the Advanced Light Source beamline and the Canadian Light Source Inc. ID08-1 for data collection time and assistance. We thank the Canadian Institutes for Health Research (CIHR) for financial support. N.C.J.S. also thanks the Howard Hughes Medical Institute (HHMI) International Scholar program and the Michael Smith Foundation for Health Research (MSFHR) for operational and salary support, as well as the Canada Foundation for Innovation (CFI), the MSFHR and the British Columbia Knowledge Development Fund (BCKDF) for infrastructure funding. S.G.W. thanks the Canada Research Chairs Program for salary support. F.V.R. is supported by long-term European Molecular Biology Organization (EMBO) and MSHFR fellowships. J.R.R is supported by a MSHFR fellowship. Author contributions F.V.R. performed purification, crystallization and structure determination; J.R.R. synthesized acceptor sugars; B.R. performed the enzymology; S.B., M.F.S., K.J., C.B. and S.D performed construct design and purification; F.V.R., J.R.R., S.G.W. and N.C.J.S. analyzed the data and wrote the manuscript; F.V.R. made the figures; W.W.W. provided feedback on the manuscript. S11
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
<Supplemental information>
 The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
Midterm 1 Last, First
 Midterm 1 BIS 105 Prof. T. Murphy April 23, 2014 There should be 6 pages in this exam. Exam instructions (1) Please write your name on the top of every page of the exam (2) Show all work for full credit
Midterm 1 BIS 105 Prof. T. Murphy April 23, 2014 There should be 6 pages in this exam. Exam instructions (1) Please write your name on the top of every page of the exam (2) Show all work for full credit
Data are contained in multiple tabs in Excel spreadsheets and in CSV files.
 Contents Overview Curves Methods Measuring enzymatic activity (figure 2) Enzyme characterisation (Figure S1, S2) Enzyme kinetics (Table 3) Effect of ph on activity (figure 3B) Effect of metals and inhibitors
Contents Overview Curves Methods Measuring enzymatic activity (figure 2) Enzyme characterisation (Figure S1, S2) Enzyme kinetics (Table 3) Effect of ph on activity (figure 3B) Effect of metals and inhibitors
Biochemistry - I. Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I
 Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Constructs expressing recombinant proteins were generated by PCR; mutant variants were
 Cell, Volume 136 Supplemental Data The Ste5 Scaffold Directs Mating Signaling by Catalytically Unlocking the Fus3 MAP Kinase for Activation Matthew Good, Grace Tang, Julie Singleton, Attila Remenyi, and
Cell, Volume 136 Supplemental Data The Ste5 Scaffold Directs Mating Signaling by Catalytically Unlocking the Fus3 MAP Kinase for Activation Matthew Good, Grace Tang, Julie Singleton, Attila Remenyi, and
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Activities for the α-helix / β-sheet Construction Kit
 Activities for the α-helix / β-sheet Construction Kit The primary sequence of a protein, composed of amino acids, determines the organization of the sequence into the secondary structure. There are two
Activities for the α-helix / β-sheet Construction Kit The primary sequence of a protein, composed of amino acids, determines the organization of the sequence into the secondary structure. There are two
Introduction to Protein Structure Collection
 Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Manja Henze, Dorothee Merker and Lothar Elling. 1. Characteristics of the Recombinant β-glycosidase from Pyrococcus
 S1 of S17 Supplementary Materials: Microwave-Assisted Synthesis of Glycoconjugates by Transgalactosylation with Recombinant Thermostable β-glycosidase from Pyrococcus Manja Henze, Dorothee Merker and Lothar
S1 of S17 Supplementary Materials: Microwave-Assisted Synthesis of Glycoconjugates by Transgalactosylation with Recombinant Thermostable β-glycosidase from Pyrococcus Manja Henze, Dorothee Merker and Lothar
CHEM 160A Final Exam. 1. (5 points) What factors influence an enzyme s substrate specificity?
 CHEM 160A Final Exam December 17, 2004 Name (1 point) 1. (5 points) What factors influence an enzyme s substrate specificity? 2. (4 points) Why are cofactors required for some enzymatic reactions? 3. (5
CHEM 160A Final Exam December 17, 2004 Name (1 point) 1. (5 points) What factors influence an enzyme s substrate specificity? 2. (4 points) Why are cofactors required for some enzymatic reactions? 3. (5
Introduction to Biochemistry Midterm exam )ومن أحياها(
 Introduction to Biochemistry Midterm exam 2016-2017 )ومن أحياها( 1. Which of the following amino (in a peptide chain) would probably be found at a beta bend or turn? a. lysine * b. Gly c. arg d. asn 2.
Introduction to Biochemistry Midterm exam 2016-2017 )ومن أحياها( 1. Which of the following amino (in a peptide chain) would probably be found at a beta bend or turn? a. lysine * b. Gly c. arg d. asn 2.
Supplementary Information
 Supplementary Information Structural basis of improved second generation 3-nitro-tyrosine trna synthetases Richard B. Cooley, Jessica L. Feldman, Camden M. Driggers, Taylor Bundy, Audrey L. Stokes, P.
Supplementary Information Structural basis of improved second generation 3-nitro-tyrosine trna synthetases Richard B. Cooley, Jessica L. Feldman, Camden M. Driggers, Taylor Bundy, Audrey L. Stokes, P.
Supporting Information for:
 Supporting Information for: Methylerythritol Cyclodiphosphate (MEcPP) in Deoxyxylulose Phosphate Pathway: Synthesis from an Epoxide and Mechanisms Youli Xiao, a Rodney L. Nyland II, b Caren L. Freel Meyers
Supporting Information for: Methylerythritol Cyclodiphosphate (MEcPP) in Deoxyxylulose Phosphate Pathway: Synthesis from an Epoxide and Mechanisms Youli Xiao, a Rodney L. Nyland II, b Caren L. Freel Meyers
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Glycosyltransferase Activity Kit
 Glycosyltransferase Activity Kit Catalog Number EA001 This package insert must be read in its entirety before using this product. For research use only. Not for use in diagnostic procedures. TABLE OF CONTENTS
Glycosyltransferase Activity Kit Catalog Number EA001 This package insert must be read in its entirety before using this product. For research use only. Not for use in diagnostic procedures. TABLE OF CONTENTS
Amino acids. (Foundation Block) Dr. Essa Sabi
 Amino acids (Foundation Block) Dr. Essa Sabi Learning outcomes What are the amino acids? General structure. Classification of amino acids. Optical properties. Amino acid configuration. Non-standard amino
Amino acids (Foundation Block) Dr. Essa Sabi Learning outcomes What are the amino acids? General structure. Classification of amino acids. Optical properties. Amino acid configuration. Non-standard amino
Properties of amino acids in proteins
 Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
Objective: You will be able to explain how the subcomponents of
 Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Supporting information (protein purification, kinetic characterization, product isolation, and characterization by NMR and mass spectrometry):
 Supporting Information Mechanistic studies of a novel C-S lyase in ergothioneine biosynthesis: the involvement of a sulfenic acid intermediate Heng Song, 1 Wen Hu, 1,2 Nathchar Naowarojna, 1 Ampon Sae
Supporting Information Mechanistic studies of a novel C-S lyase in ergothioneine biosynthesis: the involvement of a sulfenic acid intermediate Heng Song, 1 Wen Hu, 1,2 Nathchar Naowarojna, 1 Ampon Sae
Biochemistry Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology Kharagpur. Lecture -02 Amino Acids II
 Biochemistry Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology Kharagpur Lecture -02 Amino Acids II Ok, we start off with the discussion on amino acids. (Refer Slide Time: 00:48)
Biochemistry Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology Kharagpur Lecture -02 Amino Acids II Ok, we start off with the discussion on amino acids. (Refer Slide Time: 00:48)
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION DOI: 10.1038/NNANO.2012.80 Protein-Inorganic Hybrid Nanoflowers Jun Ge, Jiandu Lei, and Richard N. Zare Supporting Online Material Materials Proteins including albumin from bovine
SUPPLEMENTARY INFORMATION DOI: 10.1038/NNANO.2012.80 Protein-Inorganic Hybrid Nanoflowers Jun Ge, Jiandu Lei, and Richard N. Zare Supporting Online Material Materials Proteins including albumin from bovine
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Lecture 3: 8/24. CHAPTER 3 Amino Acids
 Lecture 3: 8/24 CHAPTER 3 Amino Acids 1 Chapter 3 Outline 2 Amino Acid Are Biomolecules and their Atoms Can Be Visualized by Two Different Ways 1) Fischer projections: Two dimensional representation of
Lecture 3: 8/24 CHAPTER 3 Amino Acids 1 Chapter 3 Outline 2 Amino Acid Are Biomolecules and their Atoms Can Be Visualized by Two Different Ways 1) Fischer projections: Two dimensional representation of
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
Tivadar Orban, Beata Jastrzebska, Sayan Gupta, Benlian Wang, Masaru Miyagi, Mark R. Chance, and Krzysztof Palczewski
 Structure, Volume Supplemental Information Conformational Dynamics of Activation for the Pentameric Complex of Dimeric G Protein-Coupled Receptor and Heterotrimeric G Protein Tivadar Orban, Beata Jastrzebska,
Structure, Volume Supplemental Information Conformational Dynamics of Activation for the Pentameric Complex of Dimeric G Protein-Coupled Receptor and Heterotrimeric G Protein Tivadar Orban, Beata Jastrzebska,
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A
 Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Europium Labeling Kit
 Europium Labeling Kit Catalog Number KA2096 100ug *1 Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 Principle of the Assay...
Europium Labeling Kit Catalog Number KA2096 100ug *1 Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 Principle of the Assay...
9/16/15. Properties of Water. Benefits of Water. More properties of water
 Properties of Water Solid/Liquid Density Water is densest at 4⁰C Ice floats Allows life under the ice Hydrogen bond Ice Hydrogen bonds are stable Liquid water Hydrogen bonds break and re-form Benefits
Properties of Water Solid/Liquid Density Water is densest at 4⁰C Ice floats Allows life under the ice Hydrogen bond Ice Hydrogen bonds are stable Liquid water Hydrogen bonds break and re-form Benefits
UV Tracer TM Maleimide NHS ester
 UV Tracer TM Maleimide HS ester Product o.: 1020 Product ame: UV-Tracer TM Maleimide-HS ester Chemical Structure: Chemical Composition: C 41 H 67 5 18 Molecular Weight: 1014.08 Appearance: Storage: Yellow
UV Tracer TM Maleimide HS ester Product o.: 1020 Product ame: UV-Tracer TM Maleimide-HS ester Chemical Structure: Chemical Composition: C 41 H 67 5 18 Molecular Weight: 1014.08 Appearance: Storage: Yellow
Biomolecules: amino acids
 Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
LANCE Eu-W1024 ITC Chelate & Europium Standard AD0013 Development grade
 AD0017P-4 (en) 1 LANCE Eu-W1024 ITC Chelate & Europium Standard AD0013 Development grade INTRODUCTION Fluorescent isothiocyanato-activated (ITC-activated) Eu-W1024 chelate is optimized for labelling proteins
AD0017P-4 (en) 1 LANCE Eu-W1024 ITC Chelate & Europium Standard AD0013 Development grade INTRODUCTION Fluorescent isothiocyanato-activated (ITC-activated) Eu-W1024 chelate is optimized for labelling proteins
THE UNIVERSITY OF MANITOBA. DATE: Oct. 22, 2002 Midterm EXAMINATION. PAPER NO.: PAGE NO.: 1of 6 DEPARTMENT & COURSE NO.: 2.277/60.
 PAPER NO.: PAGE NO.: 1of 6 GENERAL INSTRUCTIONS You must mark the answer sheet with pencil (not pen). Put your name and enter your student number on the answer sheet. The examination consists of multiple
PAPER NO.: PAGE NO.: 1of 6 GENERAL INSTRUCTIONS You must mark the answer sheet with pencil (not pen). Put your name and enter your student number on the answer sheet. The examination consists of multiple
So where were we? But what does the order mean? OK, so what's a protein? 4/1/11
 So where were we? We know that DNA is responsible for heredity Chromosomes are long pieces of DNA DNA turned out to be the transforming principle We know that DNA is shaped like a long double helix, with
So where were we? We know that DNA is responsible for heredity Chromosomes are long pieces of DNA DNA turned out to be the transforming principle We know that DNA is shaped like a long double helix, with
Biochemistry 2 Recita0on Amino Acid Metabolism
 Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
Fundamentals of Organic Chemistry CHEM 109 For Students of Health Colleges
 Fundamentals of Organic Chemistry CHEM 109 For Students of Health Colleges Credit hrs.: (2+1) King Saud University College of Science, Chemistry Department CHEM 109 CHAPTER 9. AMINO ACIDS, PEPTIDES AND
Fundamentals of Organic Chemistry CHEM 109 For Students of Health Colleges Credit hrs.: (2+1) King Saud University College of Science, Chemistry Department CHEM 109 CHAPTER 9. AMINO ACIDS, PEPTIDES AND
Protocol for purification of recombinant protein from 300 ml yeast culture
 Protocol for purification of recombinant protein from 300 ml yeast culture Equipment and reagents needed: Zirconia beads (0.5 mm diameter from BSP, Germany) Paint Shaker (at 4 C) Tube rotator for 15 ml
Protocol for purification of recombinant protein from 300 ml yeast culture Equipment and reagents needed: Zirconia beads (0.5 mm diameter from BSP, Germany) Paint Shaker (at 4 C) Tube rotator for 15 ml
Supplementary Information
 Supplementary Information ne-step Synthesis of Novel Glycosyltransferase Inhibitors Andrew Evitt [a], Lauren M. Tedaldi [a] and Gerd K. Wagner* [a,b] [a] Institute of Pharmaceutical Science, School of
Supplementary Information ne-step Synthesis of Novel Glycosyltransferase Inhibitors Andrew Evitt [a], Lauren M. Tedaldi [a] and Gerd K. Wagner* [a,b] [a] Institute of Pharmaceutical Science, School of
Controlled biomimetic crystallization of ZIF-8 particles by amino acids
 Electronic Supplementary Material (ESI) for CrystEngComm. This journal is The Royal Society of Chemistry 2016 Supporting information for Controlled biomimetic crystallization of ZIF-8 particles by amino
Electronic Supplementary Material (ESI) for CrystEngComm. This journal is The Royal Society of Chemistry 2016 Supporting information for Controlled biomimetic crystallization of ZIF-8 particles by amino
Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants
 Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants Species UDP-2,3- diacylglucosamine hydrolase specific activity (nmol min -1 mg -1 ) Fold vectorcontrol specific
Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants Species UDP-2,3- diacylglucosamine hydrolase specific activity (nmol min -1 mg -1 ) Fold vectorcontrol specific
Supplementary Material (ESI) for Chemical Communications This journal is (c) The Royal Society of Chemistry 2008
 Experimental Details Unless otherwise noted, all chemicals were purchased from Sigma-Aldrich Chemical Company and were used as received. 2-DOS and neamine were kindly provided by Dr. F. Huang. Paromamine
Experimental Details Unless otherwise noted, all chemicals were purchased from Sigma-Aldrich Chemical Company and were used as received. 2-DOS and neamine were kindly provided by Dr. F. Huang. Paromamine
Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin
 SUPPORTING INFORMATION Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin Anna R. Arnold, Andy Zhou, and Jacqueline K. Barton Division of Chemistry and Chemical Engineering, California
SUPPORTING INFORMATION Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin Anna R. Arnold, Andy Zhou, and Jacqueline K. Barton Division of Chemistry and Chemical Engineering, California
PAPER No. : 16, Bioorganic and biophysical chemistry MODULE No. : 22, Mechanism of enzyme catalyst reaction (I) Chymotrypsin
 Subject Paper No and Title 16 Bio-organic and Biophysical Module No and Title 22 Mechanism of Enzyme Catalyzed reactions I Module Tag CHE_P16_M22 Chymotrypsin TABLE OF CONTENTS 1. Learning outcomes 2.
Subject Paper No and Title 16 Bio-organic and Biophysical Module No and Title 22 Mechanism of Enzyme Catalyzed reactions I Module Tag CHE_P16_M22 Chymotrypsin TABLE OF CONTENTS 1. Learning outcomes 2.
Proteins consist in whole or large part of amino acids. Simple proteins consist only of amino acids.
 Today we begin our discussion of the structure and properties of proteins. Proteins consist in whole or large part of amino acids. Simple proteins consist only of amino acids. Conjugated proteins contain
Today we begin our discussion of the structure and properties of proteins. Proteins consist in whole or large part of amino acids. Simple proteins consist only of amino acids. Conjugated proteins contain
Short polymer. Dehydration removes a water molecule, forming a new bond. Longer polymer (a) Dehydration reaction in the synthesis of a polymer
 HO 1 2 3 H HO H Short polymer Dehydration removes a water molecule, forming a new bond Unlinked monomer H 2 O HO 1 2 3 4 H Longer polymer (a) Dehydration reaction in the synthesis of a polymer HO 1 2 3
HO 1 2 3 H HO H Short polymer Dehydration removes a water molecule, forming a new bond Unlinked monomer H 2 O HO 1 2 3 4 H Longer polymer (a) Dehydration reaction in the synthesis of a polymer HO 1 2 3
Amino acids. Ing. Petrová Jaroslava. Workshop on Official Controls of Feed AGR 46230, , Ankara. Turkey ÚKZÚZ - NRL RO Praha 1
 Amino acids Ing. Petrová Jaroslava Workshop on Official Controls of Feed AGR 46230, 6. 7. 12. 2011, Ankara. Turkey 6.12.2011 ÚKZÚZ - NRL RO Praha 1 Content of this presentation 1. Function of amino acids
Amino acids Ing. Petrová Jaroslava Workshop on Official Controls of Feed AGR 46230, 6. 7. 12. 2011, Ankara. Turkey 6.12.2011 ÚKZÚZ - NRL RO Praha 1 Content of this presentation 1. Function of amino acids
Supporting Information
 Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked. amino-modification products by acrolein
![2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked. amino-modification products by acrolein 2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked. amino-modification products by acrolein](/thumbs/86/94743397.jpg) Supplementary Information 2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked amino-modification products by acrolein Ayumi Tsutsui and Katsunori Tanaka* Biofunctional Synthetic Chemistry Laboratory, RIKEN
Supplementary Information 2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked amino-modification products by acrolein Ayumi Tsutsui and Katsunori Tanaka* Biofunctional Synthetic Chemistry Laboratory, RIKEN
Chapter 5: Structure and Function of Macromolecules AP Biology 2011
 Chapter 5: Structure and Function of Macromolecules AP Biology 2011 1 Macromolecules Fig. 5.1 Carbohydrates Lipids Proteins Nucleic Acids Polymer - large molecule consisting of many similar building blocks
Chapter 5: Structure and Function of Macromolecules AP Biology 2011 1 Macromolecules Fig. 5.1 Carbohydrates Lipids Proteins Nucleic Acids Polymer - large molecule consisting of many similar building blocks
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Fatty acids and phospholipids
 PYS 4xx Intro 2 1 PYS 4xx Intro 2 - Molecular building blocks We now describe in more detail the nomenclature and composition of several classes of compounds of relevance to the cell, including: membrane
PYS 4xx Intro 2 1 PYS 4xx Intro 2 - Molecular building blocks We now describe in more detail the nomenclature and composition of several classes of compounds of relevance to the cell, including: membrane
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Biomimetic Chirality Sensing with Pyridoxal-5 -phosphate
 Supporting Information Biomimetic Chirality Sensing with Pyridoxal-5 -phosphate Samantha L. Pilicer, Pegah R. Bakhshi, Keith W. Bentley, Christian Wolf Department of Chemistry, Georgetown University, Washington,
Supporting Information Biomimetic Chirality Sensing with Pyridoxal-5 -phosphate Samantha L. Pilicer, Pegah R. Bakhshi, Keith W. Bentley, Christian Wolf Department of Chemistry, Georgetown University, Washington,
Enzyme Catalytic Mechanisms. Dr. Kevin Ahern
 Enzyme Catalytic Mechanisms Dr. Kevin Ahern Cleave Peptide Bonds Specificity of Cutting Common Active Site Composition/Structure Mechanistically Well Studied Chymotrypsin Chymotrypsin Catalysis H2O Chymotrypsin
Enzyme Catalytic Mechanisms Dr. Kevin Ahern Cleave Peptide Bonds Specificity of Cutting Common Active Site Composition/Structure Mechanistically Well Studied Chymotrypsin Chymotrypsin Catalysis H2O Chymotrypsin
AMINO ACIDS STRUCTURE, CLASSIFICATION, PROPERTIES. PRIMARY STRUCTURE OF PROTEINS
 AMINO ACIDS STRUCTURE, CLASSIFICATION, PROPERTIES. PRIMARY STRUCTURE OF PROTEINS Elena Rivneac PhD, Associate Professor Department of Biochemistry and Clinical Biochemistry State University of Medicine
AMINO ACIDS STRUCTURE, CLASSIFICATION, PROPERTIES. PRIMARY STRUCTURE OF PROTEINS Elena Rivneac PhD, Associate Professor Department of Biochemistry and Clinical Biochemistry State University of Medicine
Student name ID # 2. (4 pts) What is the terminal electron acceptor in respiration? In photosynthesis?
 1. Membrane transport. A. (4 pts) What ion couples primary and secondary active transport in animal cells? What ion serves the same function in plant cells? 2. (4 pts) What is the terminal electron acceptor
1. Membrane transport. A. (4 pts) What ion couples primary and secondary active transport in animal cells? What ion serves the same function in plant cells? 2. (4 pts) What is the terminal electron acceptor
Caution: For Laboratory Use. A product for research purposes only. Eu-W1284 Iodoacetamido Chelate & Europium Standard. Product Number: AD0014
 TECHNICAL DATA SHEET Lance Caution: For Laboratory Use. A product for research purposes only. Eu-W1284 Iodoacetamido Chelate & Europium Standard Product Number: AD0014 INTRODUCTION: Iodoacetamido-activated
TECHNICAL DATA SHEET Lance Caution: For Laboratory Use. A product for research purposes only. Eu-W1284 Iodoacetamido Chelate & Europium Standard Product Number: AD0014 INTRODUCTION: Iodoacetamido-activated
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth 2 points.
 MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
BIOLOGY 103 Spring 2001 MIDTERM LAB SECTION
 BIOLOGY 103 Spring 2001 MIDTERM NAME KEY LAB SECTION ID# (last four digits of SS#) STUDENT PLEASE READ. Do not put yourself at a disadvantage by revealing the content of this exam to your classmates. Your
BIOLOGY 103 Spring 2001 MIDTERM NAME KEY LAB SECTION ID# (last four digits of SS#) STUDENT PLEASE READ. Do not put yourself at a disadvantage by revealing the content of this exam to your classmates. Your
20X Buffer (Tube1) 96-well microplate (12 strips) 1
 PROTOCOL MitoProfile Rapid Microplate Assay Kit for PDH Activity and Quantity (Combines Kit MSP18 & MSP19) 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 MSP20 Rev.1 DESCRIPTION MitoProfile Rapid Microplate
PROTOCOL MitoProfile Rapid Microplate Assay Kit for PDH Activity and Quantity (Combines Kit MSP18 & MSP19) 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 MSP20 Rev.1 DESCRIPTION MitoProfile Rapid Microplate
Chapter 3: Amino Acids and Peptides
 Chapter 3: Amino Acids and Peptides BINF 6101/8101, Spring 2018 Outline 1. Overall amino acid structure 2. Amino acid stereochemistry 3. Amino acid sidechain structure & classification 4. Non-standard
Chapter 3: Amino Acids and Peptides BINF 6101/8101, Spring 2018 Outline 1. Overall amino acid structure 2. Amino acid stereochemistry 3. Amino acid sidechain structure & classification 4. Non-standard
PHAR3316 Pharmacy biochemistry Exam #2 Fall 2010 KEY
 1. How many protons is(are) lost when the amino acid Asparagine is titrated from its fully protonated state to a fully deprotonated state? A. 0 B. 1 * C. 2 D. 3 E. none Correct Answer: C (this question
1. How many protons is(are) lost when the amino acid Asparagine is titrated from its fully protonated state to a fully deprotonated state? A. 0 B. 1 * C. 2 D. 3 E. none Correct Answer: C (this question
Kit for assay of thioredoxin
 FkTRX-02-V2 Kit for assay of thioredoxin The thioredoxin system is the major protein disulfide reductase in cells and comprises thioredoxin, thioredoxin reductase and NADPH (1). Thioredoxin systems are
FkTRX-02-V2 Kit for assay of thioredoxin The thioredoxin system is the major protein disulfide reductase in cells and comprises thioredoxin, thioredoxin reductase and NADPH (1). Thioredoxin systems are
Towards a New Paradigm in Scientific Notation Patterns of Periodicity among Proteinogenic Amino Acids [Abridged Version]
![Towards a New Paradigm in Scientific Notation Patterns of Periodicity among Proteinogenic Amino Acids [Abridged Version] Towards a New Paradigm in Scientific Notation Patterns of Periodicity among Proteinogenic Amino Acids [Abridged Version]](/thumbs/84/90818314.jpg) Earth/matriX: SCIENCE TODAY Towards a New Paradigm in Scientific Notation Patterns of Periodicity among Proteinogenic Amino Acids [Abridged Version] By Charles William Johnson Earth/matriX Editions P.O.
Earth/matriX: SCIENCE TODAY Towards a New Paradigm in Scientific Notation Patterns of Periodicity among Proteinogenic Amino Acids [Abridged Version] By Charles William Johnson Earth/matriX Editions P.O.
Chapter 2: Biochemistry
 Chapter 2: Biochemistry Biochemistry Biochemistry is the study of chemical makeup and reactions of living matter All chemicals in the body are either organic & inorganic Organic compounds contain carbon
Chapter 2: Biochemistry Biochemistry Biochemistry is the study of chemical makeup and reactions of living matter All chemicals in the body are either organic & inorganic Organic compounds contain carbon
Cells. Variation and Function of Cells
 Cells Variation and Function of Cells Plasma Membrane= the skin of a cell, it protects and nourishes the cell while communicating with other cells at the same time. Lipid means fat and they are hydrophobic
Cells Variation and Function of Cells Plasma Membrane= the skin of a cell, it protects and nourishes the cell while communicating with other cells at the same time. Lipid means fat and they are hydrophobic
Analysis of Amino Acids Derived Online Using an Agilent AdvanceBio AAA Column
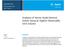 Application Note Pharmaceutical and Food Testing Analysis of Amino Acids Derived Online Using an Agilent AdvanceBio AAA Column Author Lu Yufei Agilent Technologies, Inc. Abstract A liquid chromatographic
Application Note Pharmaceutical and Food Testing Analysis of Amino Acids Derived Online Using an Agilent AdvanceBio AAA Column Author Lu Yufei Agilent Technologies, Inc. Abstract A liquid chromatographic
CELL CULTURE MEDIA AND REAGENTS
 CELL CULTURE MEDIA AND REAGENTS Classical Media Buffered Salt Solutions Reagents & Supplements CELL CULTURE MEDIA AND REAGENTS MEDIA DULBECCO S MODIFIED EAGLE S MEDIUM (DMEM) A modification of Basal Medium
CELL CULTURE MEDIA AND REAGENTS Classical Media Buffered Salt Solutions Reagents & Supplements CELL CULTURE MEDIA AND REAGENTS MEDIA DULBECCO S MODIFIED EAGLE S MEDIUM (DMEM) A modification of Basal Medium
Analysis of Cancer Proteins: Pattern Identification on Escherichia coli Species
 Research Article imedpub Journals http://www.imedpub.com Colorectal Cancer: Open Access DOI: 10.21767/2471-9943.100005 Abstract Analysis of Cancer Proteins: Pattern Identification on Escherichia coli Species
Research Article imedpub Journals http://www.imedpub.com Colorectal Cancer: Open Access DOI: 10.21767/2471-9943.100005 Abstract Analysis of Cancer Proteins: Pattern Identification on Escherichia coli Species
SUPPLEMENTARY INFORMATION FOR. (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation. by COX-2
 SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
For questions 1-4, match the carbohydrate with its size/functional group name:
 Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the free response
Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the free response
Affinity Purification of Photosystem I from Chlamydomonas reinhardtii using a Polyhistidine Tag
 Affinity Purification of Photosystem I from Chlamydomonas reinhardtii using a Polyhistidine Tag Jonathan A. Brain Galina Gulis, Ph.D. 1 Kevin E. Redding, Ph.D. 2 Associate Professor of Chemistry Adjunct
Affinity Purification of Photosystem I from Chlamydomonas reinhardtii using a Polyhistidine Tag Jonathan A. Brain Galina Gulis, Ph.D. 1 Kevin E. Redding, Ph.D. 2 Associate Professor of Chemistry Adjunct
Proteins are sometimes only produced in one cell type or cell compartment (brain has 15,000 expressed proteins, gut has 2,000).
 Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
PhosFree TM Phosphate Assay Biochem Kit
 PhosFree TM Phosphate Assay Biochem Kit (Cat. # BK050) ORDERING INFORMATION To order by phone: (303) - 322-2254 To order by Fax: (303) - 322-2257 To order by e-mail: cservice@cytoskeleton.com Technical
PhosFree TM Phosphate Assay Biochem Kit (Cat. # BK050) ORDERING INFORMATION To order by phone: (303) - 322-2254 To order by Fax: (303) - 322-2257 To order by e-mail: cservice@cytoskeleton.com Technical
Review II: The Molecules of Life
 Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
2. Which of the following amino acids is most likely to be found on the outer surface of a properly folded protein?
 Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
The Structure and Function of Macromolecules
 The Structure and Function of Macromolecules Macromolecules are polymers Polymer long molecule consisting of many similar building blocks. Monomer the small building block molecules. Carbohydrates, proteins
The Structure and Function of Macromolecules Macromolecules are polymers Polymer long molecule consisting of many similar building blocks. Monomer the small building block molecules. Carbohydrates, proteins
Chapter 8. Interaction between the phosphatidylinositol 3- kinase SH3 domain and a photocleavable cyclic peptide
 Interaction between the phosphatidylinositol 3- kinase SH3 domain and a photocleavable cyclic peptide 129 Abstract The interaction of the PI3K SH3 domain with a cyclic photocleavable peptide and the linear
Interaction between the phosphatidylinositol 3- kinase SH3 domain and a photocleavable cyclic peptide 129 Abstract The interaction of the PI3K SH3 domain with a cyclic photocleavable peptide and the linear
Lecture 10 - Protein Turnover and Amino Acid Catabolism
 Lecture 10 - Protein Turnover and Amino Acid Catabolism Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire 1 Introduction 2 Proteins are degraded into amino acids. Protein
Lecture 10 - Protein Turnover and Amino Acid Catabolism Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire 1 Introduction 2 Proteins are degraded into amino acids. Protein
Solutions to Quiz I
 Solutions to 7.014 Quiz I lass Average = 75 Median = 75 Grade Range % A 87-100 18 B 74-86 39 62-73 31 D 48-61 8 F 0-47 4 Question 1 (26 points) The protein complex, LETMEGTHRUIN (LMGT for short), acts
Solutions to 7.014 Quiz I lass Average = 75 Median = 75 Grade Range % A 87-100 18 B 74-86 39 62-73 31 D 48-61 8 F 0-47 4 Question 1 (26 points) The protein complex, LETMEGTHRUIN (LMGT for short), acts
Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast
 Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast Application note Food supplements Authors Juliusz Bianga and Joanna Szpunar
Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast Application note Food supplements Authors Juliusz Bianga and Joanna Szpunar
For questions 1-4, match the carbohydrate with its size/functional group name:
 Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 ANSWERS For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the
Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 ANSWERS For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the
From sugar unit A From sugar unit B From sugar unit C
 ame 215 F12-Exam o. Page 2 I. (26 points) Raffinose is a trisaccharide occurring in cottonseed meal. Answer the following questions as directed in the boxes. (1) ( points) Label each of the glycosidic
ame 215 F12-Exam o. Page 2 I. (26 points) Raffinose is a trisaccharide occurring in cottonseed meal. Answer the following questions as directed in the boxes. (1) ( points) Label each of the glycosidic
Supplementary Materials: An NMR Guided Screening Method for Selective Fragment Docking and Synthesis of a Warhead Inhibitor
 Molecules 216, 21, 846; doi:1.339/molecules217846 S1 of S14 Supplementary Materials: An NMR Guided Screening Method for Selective Fragment Docking and Synthesis of a Warhead Inhibitor Ram B. Khattri, Daniel
Molecules 216, 21, 846; doi:1.339/molecules217846 S1 of S14 Supplementary Materials: An NMR Guided Screening Method for Selective Fragment Docking and Synthesis of a Warhead Inhibitor Ram B. Khattri, Daniel
I) Choose the best answer: 1- All of the following amino acids are neutral except: a) glycine. b) threonine. c) lysine. d) proline. e) leucine.
 1- All of the following amino acids are neutral except: a) glycine. b) threonine. c) lysine. d) proline. e) leucine. 2- The egg white protein, ovalbumin, is denatured in a hard-boiled egg. Which of the
1- All of the following amino acids are neutral except: a) glycine. b) threonine. c) lysine. d) proline. e) leucine. 2- The egg white protein, ovalbumin, is denatured in a hard-boiled egg. Which of the
Glycolysis. Cellular Respiration
 Glucose is the preferred carbohydrate of cells. In solution, it can change from a linear chain to a ring. Energy is stored in the bonds of the carbohydrates. Breaking these bonds releases that energy.
Glucose is the preferred carbohydrate of cells. In solution, it can change from a linear chain to a ring. Energy is stored in the bonds of the carbohydrates. Breaking these bonds releases that energy.
BabyBio IMAC columns DATA SHEET DS
 BabyBio IMAC columns DATA SHEET DS 45 655 010 BabyBio columns for Immobilized Metal Ion Affinity Chromatography (IMAC) are ready-to-use for quick and easy purification of polyhistidine-tagged (His-tagged)
BabyBio IMAC columns DATA SHEET DS 45 655 010 BabyBio columns for Immobilized Metal Ion Affinity Chromatography (IMAC) are ready-to-use for quick and easy purification of polyhistidine-tagged (His-tagged)
Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein
 Supplementary Methods Thermal shift assays Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein unfolding was examined by monitoring the fluorescence of ANS (1-anilinonaphthalene-8-
Supplementary Methods Thermal shift assays Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein unfolding was examined by monitoring the fluorescence of ANS (1-anilinonaphthalene-8-
Supplemental Information. Structural Basis for the Regulation. of Cysteine-Protease Activity by a New Class. of Protease Inhibitors in Plasmodium
 Structure 19 Supplemental Information Structural Basis for the Regulation of Cysteine-Protease Activity by a New Class of Protease Inhibitors in Plasmodium Guido Hansen, Anna Heitmann, Tina Witt, Honglin
Structure 19 Supplemental Information Structural Basis for the Regulation of Cysteine-Protease Activity by a New Class of Protease Inhibitors in Plasmodium Guido Hansen, Anna Heitmann, Tina Witt, Honglin
Chemical Nature of the Amino Acids. Table of a-amino Acids Found in Proteins
 Chemical Nature of the Amino Acids All peptides and polypeptides are polymers of alpha-amino acids. There are 20 a- amino acids that are relevant to the make-up of mammalian proteins (see below). Several
Chemical Nature of the Amino Acids All peptides and polypeptides are polymers of alpha-amino acids. There are 20 a- amino acids that are relevant to the make-up of mammalian proteins (see below). Several
The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism
 Supplementary Information The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism Address correspondence to Yong Li (yongli@xmu.edu.cn, Tel: 86-592-218151) GW464 CDCA Supplementary
Supplementary Information The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism Address correspondence to Yong Li (yongli@xmu.edu.cn, Tel: 86-592-218151) GW464 CDCA Supplementary
Previous Class. Today. Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism. Protein Kinase Catalytic Properties
 Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
Macromolecules of Life -3 Amino Acids & Proteins
 Macromolecules of Life -3 Amino Acids & Proteins Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering E-mail: splin@dragon.nchu.edu.tw Website: http://web.nchu.edu.tw/pweb/users/splin/ Amino Acids Proteins
Macromolecules of Life -3 Amino Acids & Proteins Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering E-mail: splin@dragon.nchu.edu.tw Website: http://web.nchu.edu.tw/pweb/users/splin/ Amino Acids Proteins
Extracellular Enzymes Lab
 Biochemistry Extracellular Enzymes Lab All organisms convert small organic compounds, such as glucose, into monomers required for the production of macromolecules; e.g., Building Blocs Monomers Macromolecules
Biochemistry Extracellular Enzymes Lab All organisms convert small organic compounds, such as glucose, into monomers required for the production of macromolecules; e.g., Building Blocs Monomers Macromolecules
STORE AT 4 o C Version 3
 Instruction Manual Advanced Protein Assay Reagent (Cat. # ADV01) ORDERING INFORMATION To order by phone: (303) - 322-2254 To order by Fax: (303) - 322-2257 To order by e-mail: cserve@cytoskeleton.com Technical
Instruction Manual Advanced Protein Assay Reagent (Cat. # ADV01) ORDERING INFORMATION To order by phone: (303) - 322-2254 To order by Fax: (303) - 322-2257 To order by e-mail: cserve@cytoskeleton.com Technical
