Comparative Analysis of Chromosomal Variation Altered New Region for Detecting Biomarker in Liver Cancer
|
|
|
- Aubrie O’Neal’
- 5 years ago
- Views:
Transcription
1 Computer Science and Engineering 2017, 7(1): DOI: /j.computer Comparative Analysis of Chromosomal Variation Altered New Region for Detecting Biomarker in Liver Cancer Esraa M. Hashem 1, Mai S. Mabrouk 1,*, Ayman M. Eldeib 2 1 Biomedical Engineering Department, Misr University for Science and Technology (MUST University), Egypt 2 Systems and Biomedical Engineering, Faculty of engineering, Cairo University, Giza, Egypt Abstract Liver cancer is one of the most widely recognized malignancies in adults, hepatocellular carcinoma (HCC) is an out face malignancy. Nearly instances of HCC are identified with a preceding stage, and, in this manner, medication choices are restricted. Most Hepatic carcinoma emerge from the accumulation of genetic abnormalities and potentially induced by exterior etiological factors. Genome-wide gene expression profiling is a generally utilized system for discovering new potential biomarkers for prediction and diagnosis of disease severity. In this work, two statistical techniques (circular binary segmentation and discrete stationary wavelet transform) are utilized for detection of copy number alterations to test single-nucleotide polymorphism array of ten cell lines hepatic tumor patients on certain chromosomes, which are frequently detected in hepatic cancer have been proposed. Results detect a new mutated chromosomal amplification region (4q22.1) has been identified through our investigation utilizing both methods. This region sets up, SPP1, and AFF1 tumor suppressor genes and has not observed previously, that opening new frontiers that can be involved in liver carcinogenesis. Keywords Hepatocellular carcinoma, Circular binary segmentation, Discrete stationary wavelet transform, Single-nucleotide polymorphism 1. Introduction Cancer has been over and again distinguished as a controlled disease. Certainly, the procedure of tumor growth from primary to malignant structures, including metastasis to different cells includes the conflict between various cell lineages contend to adjust to the changing conditions both inside and around the tumor. This complex operation is almost associated with genetic alterations and the incidence of mutations [1]. Greater than 80% of hepatic cancers are hepatocellular carcinoma (HCCs), It is once in a while identified early [2]. The end stage of many different chronic liver diseases is liver cirrhosis, it is normally diagnosed at an end-stage when effective treatment is impossible. Patient with cirrhosis have an over risk of liver tumor. Mostly cirrhosis created because of: hepatitis C (HCV), alcoholic liver disease (ALD), hepatitis B (HBV), auto-immune hepatitis (AIH), and other risk factors [3], as shown in Figure 1-1. As stated by those current studies, the large part of liver cancer patients contracted the disease from the accumulation the abnormalities of genetics, possibly rise by exterior etiological factors especially HCV and HBV contaminations [4]. So far, no significant proof has found that enhances * Corresponding author: msm_eng@yahoo.com (Mai S. Mabrouk) Published online at Copyright 2017 Scientific & Academic Publishing. All Rights Reserved survival utility with surveillance of high-risk patients. The diagnostic tools which utilized commonly to attempt to enhance patient survival are: biomarkers, liver biopsy, radiographic imaging and serum tumor marker alpha-fetoprotein (AFP) [5]. HCV PSC AIH ALD NASH HBV Liver PBC HCC Figure 1.1. Causes of liver cirrhosis Liver cirrhosis Mainly human malignancy cancers are genetic diseases raise by genome aberrations. Chromosomal instability is one of the characteristics in cancer, its outcomes in the structural aberration of DNA copy number alterations (CNAs). That alteration changes the scale of gene expression, which alter
2 Computer Science and Engineering 2017, 7(1): normal growth control and survival pathway [5]. Genetic variation between a similar tumor sort in various people may result in metabolic differences between one malignancy tissue and another that can influence disease progression or response to therapy. CNA is necessary for each of the major understanding of Hepatic carcinoma and its detection. Among such alterations, frequently detected of DNA copy number gains at chromosome 1, 8, 11, 17, and q [6-8] and losses at chromosome 4, 8, 9, 13, 16, and 17 have been specified in HCC [9-11] using traditional methodology. Single-nucleotide polymorphism array (SNP) are single base alterations between genomes within a species. SNPs is helpful markers for diseases in haplotype-based association studies and utilized to recognize chromosomal alterations to identify mutations [12]. Since the most recent decade, the entire genome SNP array has been the basic element in a highly productive synergistic relationship between advances in biological understanding and computational approaches. The Illumina Bead Array has steadily expanded limit throughout the years from SNPs to the current one million [13]. SNP array is a variation at one position in a DNA sequence among individuals. Recall that the DNA sequence is formed from a chain of nucleotide bases: A, C, G, and T. If more than 1% of a population does not carry the same nucleotide at a specific position in the DNA sequence, then this aberration can be classified as an SNP. It is needed to identify new biomarkers for early diagnosis of HCC and it is important to find new methods that can evaluate the prognosis of HCC patients. Numerous automated algorithms are used to define the characteristic of genomic profiles; Cooper et al. utilize SNP conditional mixture modeling (SCIMM) by using a mixture-likelihood clustering methodology within the R package to characterize deletions of CNA [14]. Franke et al. give a combined program focused on SNP interpretation [15]. A lot of automated algorithms to detect biomarkers in CNAs have a solid restriction it is the noise generated from signals [16]. To fill this obstacle, Circular Binary segmentation (CBS) and Discrete Stationary Wavelet transforms (DSWT) are utilized, that aids in the perception of the genomic basis of the disease progression and improve a method to find biomarkers for early identification of liver cancer. 2. Materials and Methods The most profuse genetic variants are SNPs, which supply the ultimate comprehensive resource for verification of genetic diversity [17]. The progress in high-throughput sequencing technology has made copy number discovery much easier, the application of known CNA information means that we can target structural variation in a sample using less expensive techniques such as the SNP array without a large reduction in genome-wide coverage Experimental Setup The data used in this study is SNP Genotyping Array, performed on Human Duov1 DNA investigation Bead Chip (HumanCNV370v1), includes the usual SNP content items on a HumanHap300-Duo Genotyping Bead Chip, with an extra 52,167 marker intended to specifically target close-by 14,000 CNA areas of the genome, for an aggregate of more than 370,000 markers. Ten HCC cell lines process according to the manufacturers' instructions. Raw data processed utilizing an in-house pipeline to take out a segment of copy number and gene-summarized copy number assessments. The data on the normalized log R ratios are obtainable in GEO [ with accession number [GSE38207] Statistical Methods Circular Binary Segmentation (CBS): Because of the complication of genetic variations in tumor cells, high density SNP array investigation is an extremely requesting method in the field of cancer research. The implementation of SNP arrays in combination with tissue array, PCR and functional analysis advances will finally prompt to the full revelation of the complex genetic alterations in human cancers. CBS can give a characteristic approach to segment a chromosome into contiguous regions iteratively searching for parts of chromosomal regions and bypasses parametric modeling of the data with its utilize of a variation reference division. Once the chromosome partitioned, a copy number can appraise from the segments and discover the change points on SNP array data with the help of assistance information such as the ploidy of the chromosome [18]. The algorithm stops when no more statistically significant change-points are situated, according to a heuristically chosen significance threshold. This information will give the locations of copy number aberrations, assigned by Z i as appeared in (1). Where SS ii = XX XX ii 1 ii nn are the partial sums and XX 1 XX nn is the log R ratios of the intensities. The probability proportion measurement for checking the null hypothesis that there is no change against the alternative that there is exactly one change at an unknown location i is given by (2) [19]. ZZ ii = ii nn ii 2 SS ii ii SS nn SS ii (1) nn ii ZZZZ = mmmmmm1 ii < nn (2) The CBS model makes losses or gains of CNVs discrete. These variations happen in adjacent regions of the chromosome that frequently hold different markers even entire chromosome regions. CBS precisely segment data by distinguishing change-points utilizing a maximal-t test [20]. The array information to be utilized for recognition change-point are the log R ratio of settled intensities listed by the marked positions. Note that there might be different change-points in a particular chromosome, everyone correlating to an adjustment in the copy number in the selected sample.
3 24 Esraa M. Hashem et al.: Comparative Analysis of Chromosomal Variation Altered New Region for Detecting Biomarker in Liver Cancer Discrete Stationary Wavelet transforms (DSWT): A discrete wavelet is any wavelet transform, for which the wavelets are discretely tested. The other form of representing the signal is the transform of a signal. It does not modify the information, content currently in the signal [21]. The Wavelet change supplies a time-frequency illustration of the signal. The wavelet transforms enable high compression ratios with good quality of reconstruction. Bases of wavelet are produced by the dilation and translation of a single wavelet role (x) called the Haar basis [22]. It is the simplest wavelet base, which produced by the Haar function in (3), it distinguishes statistically significant breakpoints in the data, utilizing the maxima of the Haar wavelet change, and segments accordingly [5] < xx 0 ψ(xx) = < xx 1 (3) 0 ooooheeeeeeeeeeee After using Haar function there is need to eliminate noise from the signal through decomposition the input signal into an approximation sub-bands and a set of detail sub-bands at different resolution scales utilizing a set of high pass filter and low pass filters [23]. A limit level assigned to each dyadic resolution level by the guideline of minimizing the Stein Unbiased Estimate of Risk (SURE) for threshold estimates. De-blurring increases the sharpness signal features by boosting the high frequencies, whereas denoising tries to remove whatever noise is present regardless of the spectral content of a noisy signal. Restoration is a kind of denoising that tries to retrieve the original signal with optimal balancing of de-blurring and smoothing Wavelet transforms enable us to represent signals with a high degree of sparsity. This is the principle behind a non-linear wavelet based signal estimation technique known as wavelet denoising. 3. Results HCC is a standout amongst the ultimate recurrent cancer entities worldwide, Hepatitis C and Hepatitis B are the main underlying basic causes of chronic liver disease which leads to cirrhosis. The tumor recurrence is the major restriction of long survival after liver transplantation. Most recent advancement of technologies for gene expression profiling and genome copy number investigation enable the comprehensive characterization of entire cancer genomes. This information will enhance our comprehension of the epigenetic and genetic aberrations leading to the evolution of HCC, as well as ultimately to provide information that helps in the treatment of patients [24]. By performing CBS algorithm in HCC cell lines. A region of 4q25-26 losses observed in 10 out of 10-HCC (100%), the allelic loss of chromosome 4q were significantly associated with HCC has elevated serum AFP [25]. A loss of 8p21.3 was observed in 10 out of 10-HCC (100%), loss of 9p was observed in 10 out of 10-HCC (100%) also loss of 16q11.2 and 17p13.1 observed in 10 out of 10-HCC (100%), all the results are summarized in table 3.1. While performing DSWT algorithm, it has been noticed that, the most commonly gained region of chromosome 1q in 4 out of 10-hepatocellular carcinoma (40%). Gains of 1q are the most frequent variation and occur early during tumor-genesis [26]. This region contains the tumor suppressors genes JTB, CCT3and COPA [27]. A loss of 4q was detected in 10 out of 10-HCC (100%). Loss for 9p additionally observed in 10 out of 10-HCC (100), the results summarized in table 3.1. This finding agrees with those of previous studies [28] leading to the assumption that these aberrations could have an essential part in the preparation of HCC, probably based on variations of genes lying on these chromosome segments. Table 3.1. Prediction accuracy of each chromosome for 10-HCC sample Chromosome CNAs type Percentage of CBS sensitivity Percentage of DSWT sensitivity Gain1q /10(30%) 4/10(40%) Gain4q /10(80%) 10/10(100%) Gain8q /10(40%) 2/10(20%) Gain11q /10(50%) 5/10(50%) Gain17q /10(20%) 5/10(50%) Gain20q12 5/10(50%) 1/10(10%) Loss4q /10(100%) 10/10(100%) Loss8p /10(100%) 6/10(60%) Loss9p /10(100%) 10/10(100%) Loss13q /10(60%) 4/10(40%) Loss16q /10(100%) 4/10(40%) Loss17p /10(100%) 9/10(90%) While performing CBS and DSWT the analysis shows that, a new altered region has been detected by amplification of chromosome 4 region 4q22.1 in 8 out of 10 HCC cell lines (80%) using CBS and 10 out of 10 HCC cell lines (100%) of SSP1 and AFF1 genes using DSWT as shown in figure (3-1,2). This observing has not previously reported being required in liver carcinogenesis before Esraa. M, et.al study [22]. 4. Discussions The liver has many identified functions related to metabolism, processing, and immunity, the liver disease indicates a different number of concerns for the delivery of medical consideration. Large-scale CN changes in the genomes of humans have been disregarded mostly because of technical challenges in scoring these changes genomes-wide with a favorable resolution. Gene expression profiling can prompt to discover new markers that may help enhance diagnostic for detect HCC, represent novel targets for therapeutic agents, and permit us to identify regions in the HCC genome that have undergone recurrent high-level focal amplifications or deletions. It
4 Computer Science and Engineering 2017, 7(1): should be considered for future investigation on detecting CNA regions. 5. Conclusions Hepatic carcinoma is a tumor of the liver, which is generally emerging in the background of chronic liver diseases. The assessment for patients with liver cancer is poor because of, the low chance of therapeutic treatment. The symptoms of HCC are usually absent. Further, the improvement of early detection markers is a vital missing component in methodologists to reduce HCC mortality. Chromosome aberrations in cancer cells can be resolved with a rising number of large-scale genomic and molecular genetics SNP array technology. However, the recent advancement of alteration of gene expression profiling and genome-wide copy number investigation have allowed the comprehensive selecting properties of whole cancer genomes. This finding enhances our comprehension of the epigenetic aberrations and genetic aberrations driving the development of liver cancer, instead of ultimately extend guidance to designed treatment of patients. SNP array is the most widely recognized strategy to examine the copy number of DNA sequences. It provides accurate information of CNAs which is essential to distinguish deletion and amplification in the chromosome. A better perception of gene expression in hepatic cancer can aid to comprehend the pathophysiology of HCC and enhanced prognostication. Figure 3.1. Gain of chromosome 4q22.1 using CBS algorithm Figure 3.2. Gain of chromosome 4q22.1 using DSWT
5 26 Esraa M. Hashem et al.: Comparative Analysis of Chromosomal Variation Altered New Region for Detecting Biomarker in Liver Cancer This work demonstrates that genomic high-density SNP-array test can be utilized to detect and characterize genome-wide CNAs on more than one thousand clones emerging from several chromosomes. By testing CBS and one-dimensional DSWT techniques on 10 HCC cell lines it has been found that there is a certain number of chromosome aberration including gains of 1q,8q,11q,17q,20q chromosomes and losses of 4q,8p,13q,16q,17p. A new altered region has been detected by amplification of chromosome 4 region 4q22.1. This gene expression profiling SSP1(secreted phosphoprotein 1) and AFF1(AF4/FMR2 family member 1) can prompt to: discover new biomarkers that may help enhance diagnostic for detecting HCC, represent novel objective for therapeutic agents, and allow us to identify regions in the HCC genome that have undergone recurrent high-level focal deletions or amplifications. REFERENCES [1] Alfonso. V and Manuel. H, Getting personalized cancer genome analysis into the clinic: the challenges in bioinformatics, Valencia and Hidalgo Genome Medicine, vol. 4, pp.1-17, [2] El-Serag. HB, Hepatocellular carcinoma, New England Journal of Medicine, vol.365, pp , [3] Guha. N. I,. Iredale. J.P, Clinical and diagnostic aspects of cirrhosis. In: Textbook of hepatology from basic science to clinical practice. 3rd edition. Edited by: Rodés J, Benhamou J-P, Blei A, Reichen J, Rizzetto M. Oxford, Blackwell Publishing, pp , [4] Esraa. M. H, Mai. S. M, A study of Support Vector Machine Algorithm for Liver Disease Diagnosis, American Journal of Intelligent Systems, vol. 4, pp.9-14, [5] Mai. S. M, Esraa. S. H, Amr. S, Discrete Stationary Wavelet Transform of Array CGH Data for Biomarkers Identification of Hepatocellular Carcinoma, Journal of Bioinformatics and Intelligent Control, vol. 1 (2), pp ,2012. [6] Beroukhim.R, Mermel. CH, Porter.D, Wei. G, Raychaudhuri. S, Donovan. J, et al. The landscape of somatic copy-number alteration across human cancers, Nature, vol.463, pp , [7] Lee. S.A, Ho. C, Roy. R, Kosinski. C, Patil. M.A, Tward. A.D, et al. Integration of genomic analysis and in vivo transfection to identify sprouty 2 as a candidate tumor suppressor in liver cancer, HEPATOLOGY, vol.47, pp , [8] Tanaka.Y, Kanai. F, Tada. M, Tateishi. R. M. Sanada. M, Nannya. Y, Gain of GRHL2 is associated with early recurance of hepatocellular carcinoma, J Hepatol., vol. 49, pp , [9] Midorikawa. Y, Tang. W, Sugiyama. Y, High-resolution mapping of copy number aberrations and identification of target genes in hepatocellular carcinoma, Biosci Trends, vol. 1, pp , [10] Patil. M. A, Gutgemann. I, Zhang. J, Ho. C, Cheung. S.T, Ginzinger. D, et al. Array-based comparative genomic hybridization reveals recurrent chromosomal aberrations and Jab1 as a potential target for 8q gain in hepatocellular carcinoma., Carcinogenesis,,vol.26, pp , [11] Chen T.C, Hsieh. L.L,. Kuo. T. T,. Ng. K.F,W.C. YH, et.al, p16ink4 gene mutation and allelic loss of chromosome 9p21-22 in Taiwanese hepatocellular carcinoma, Anticancer Res., vol.20, pp , [12] DE. REICH, M. CARGILL, S. BOLK, et al.: Linkage disequilibrium in the human genome. Nature, vol. 411, pp , [13] Framboise.T.L, Single nucleotide polymorphism arrays: a decade of biological, computational and technological advances, Nucleic Acids Research, Vol. 37, pp , [14] Kennedy. J, Efficient Algorithms for SNP Genotype Data Analysis using Hidden Markov Models of Haplotype Diversity, M.Sc., Rensselaer, Hartford, CT, USA, [15] Cooper. G.M, Zerr. T, Kidd. J.M, et al. Systematic assessment of copy number variant detection via genome-wide SNP genotyping, Nat Genet, vol.40 (10), pp , [16] Franke. L, Kovel. C.G, Aulchenko. Y.S, et al. Detection, imputation, and association analysis of small deletions and null alleles on oligonucleotide arrays, AmJ Hum Genet, vol. 82, pp , [17] Wiltshire. T, Pletcher. M. T, Batalov. S, Barnes. S. W, Tarantino. L. M, Genome-wide single-nucleotide polymorphism analysis defines haplotype patterns in mouse, PNAS, vol.100, pp , [18] Esraa. M.H, Mai. S. M, Amr. Sh, Circular Binary Segmentation Modeling of Array CGH Data on Hepatocellular carcinoma., Radio Science Conference (NRSC), 29th National, pp , [19] Sen. A and Srivastava. M. S, On tests for detecting a change in mean, Annals of Statistics, vol.3, pp , [20] Olshen. A.B and. Venkatraman. E.S. Circular binary segmentation for the analysis of array based DNA copy number data, Biostatistics, vol.5, pp , [21] Darshana. M, Asim. B, DISCRETE WAVELET TRANSFORM USING MATLAB, IGCET, Vol.4, pp , [22] Esraa. M. H, Mai. S. M, Ayman. M. El, Novel Altered Region for Biomarker Discovery in Hepatocellular Carcinoma (HCC) Using Whole Genome SNP Array,(IJACSA) International Journal of Advanced Computer Science and Applications, Vol. 7, No. 4,2016. [23] Esraa. S. H, Mai. S. M, Impact of parallel computing on identifying biomarkers of hepatocellular carcinoma, J. Med. Imaging Health Inf,, vol.4, pp. 1 5, [24] Esraa. M. H, Mai. S. M, Ayman. M. Eldeab, Clinical and Genomic strategies for Detecting Hepatocellular Carcinoma in Early Stages: A systematic review, American Journal of Biomedical Engineering, vol. 5, pp , [25] Singhal. A, Jayaraman. M, Dhanasekaran.D.N, Kohli. V,
6 Computer Science and Engineering 2017, 7(1): Molecular and serum markers in hepatocellular carcinoma: Predictive tools for prognosis and recurrence, Critical Reviews in Oncology/Hematology, vol. 82, pp , [26] Subramanian. A, Tamayo. P, Mootha. V. K, et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles, Proc Natl Acad Sci USA, vol.102, pp , [27] Wang. Y, Wu.M.C., Sham. J.S, Zhang. W, Wu. W. Q, and Guan. X.Y, Prognostic significance of c-myc and AIB1 amplification in hepatocellular carcinoma. A broad survey using high-throughput tissue microarray, Cancer, vol.95, pp , [28] Wong. N, Lai. P. Lee. S-W, et al. Assessment of genetic changes in hepatocellular carcinoma by comparative genomic hybridization analysis,. Am J Pathol, vol. 154, pp.37-43, View publication stats
Understanding DNA Copy Number Data
 Understanding DNA Copy Number Data Adam B. Olshen Department of Epidemiology and Biostatistics Helen Diller Family Comprehensive Cancer Center University of California, San Francisco http://cc.ucsf.edu/people/olshena_adam.php
Understanding DNA Copy Number Data Adam B. Olshen Department of Epidemiology and Biostatistics Helen Diller Family Comprehensive Cancer Center University of California, San Francisco http://cc.ucsf.edu/people/olshena_adam.php
Supplementary note: Comparison of deletion variants identified in this study and four earlier studies
 Supplementary note: Comparison of deletion variants identified in this study and four earlier studies Here we compare the results of this study to potentially overlapping results from four earlier studies
Supplementary note: Comparison of deletion variants identified in this study and four earlier studies Here we compare the results of this study to potentially overlapping results from four earlier studies
Statistical Analysis of Single Nucleotide Polymorphism Microarrays in Cancer Studies
 Statistical Analysis of Single Nucleotide Polymorphism Microarrays in Cancer Studies Stanford Biostatistics Workshop Pierre Neuvial with Henrik Bengtsson and Terry Speed Department of Statistics, UC Berkeley
Statistical Analysis of Single Nucleotide Polymorphism Microarrays in Cancer Studies Stanford Biostatistics Workshop Pierre Neuvial with Henrik Bengtsson and Terry Speed Department of Statistics, UC Berkeley
Clinical and Genomic Strategies for Detecting Hepatocellular Carcinoma in Early Stages: A Systematic Review
 American Journal of Biomedical Engineering 2015, 5(4): 101-115 DOI: 10.5923/j.ajbe.20150504.01 Clinical and Genomic Strategies for Detecting Hepatocellular Carcinoma in Early Stages: A Systematic Review
American Journal of Biomedical Engineering 2015, 5(4): 101-115 DOI: 10.5923/j.ajbe.20150504.01 Clinical and Genomic Strategies for Detecting Hepatocellular Carcinoma in Early Stages: A Systematic Review
Introduction to LOH and Allele Specific Copy Number User Forum
 Introduction to LOH and Allele Specific Copy Number User Forum Jonathan Gerstenhaber Introduction to LOH and ASCN User Forum Contents 1. Loss of heterozygosity Analysis procedure Types of baselines 2.
Introduction to LOH and Allele Specific Copy Number User Forum Jonathan Gerstenhaber Introduction to LOH and ASCN User Forum Contents 1. Loss of heterozygosity Analysis procedure Types of baselines 2.
Abstract. Optimization strategy of Copy Number Variant calling using Multiplicom solutions APPLICATION NOTE. Introduction
 Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Whole-genome detection of disease-associated deletions or excess homozygosity in a case control study of rheumatoid arthritis
 HMG Advance Access published December 21, 2012 Human Molecular Genetics, 2012 1 13 doi:10.1093/hmg/dds512 Whole-genome detection of disease-associated deletions or excess homozygosity in a case control
HMG Advance Access published December 21, 2012 Human Molecular Genetics, 2012 1 13 doi:10.1093/hmg/dds512 Whole-genome detection of disease-associated deletions or excess homozygosity in a case control
Computer Science, Biology, and Biomedical Informatics (CoSBBI) Outline. Molecular Biology of Cancer AND. Goals/Expectations. David Boone 7/1/2015
 Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
AVENIO ctdna Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB
 Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB Analysis Kits Next-generation performance in liquid biopsies 2 Accelerating clinical research From liquid biopsy to next-generation
Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB Analysis Kits Next-generation performance in liquid biopsies 2 Accelerating clinical research From liquid biopsy to next-generation
Early Detection of Lung Cancer
 Early Detection of Lung Cancer Aswathy N Iyer Dept Of Electronics And Communication Engineering Lymie Jose Dept Of Electronics And Communication Engineering Anumol Thomas Dept Of Electronics And Communication
Early Detection of Lung Cancer Aswathy N Iyer Dept Of Electronics And Communication Engineering Lymie Jose Dept Of Electronics And Communication Engineering Anumol Thomas Dept Of Electronics And Communication
DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK
 CHAPTER 6 DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK Genetic research aimed at the identification of new breast cancer susceptibility genes is at an interesting crossroad. On the one hand, the existence
CHAPTER 6 DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK Genetic research aimed at the identification of new breast cancer susceptibility genes is at an interesting crossroad. On the one hand, the existence
Structural Variation and Medical Genomics
 Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits
 AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
Genome-wide copy-number calling (CNAs not CNVs!) Dr Geoff Macintyre
 Genome-wide copy-number calling (CNAs not CNVs!) Dr Geoff Macintyre Structural variation (SVs) Copy-number variations C Deletion A B C Balanced rearrangements A B A B C B A C Duplication Inversion Causes
Genome-wide copy-number calling (CNAs not CNVs!) Dr Geoff Macintyre Structural variation (SVs) Copy-number variations C Deletion A B C Balanced rearrangements A B A B C B A C Duplication Inversion Causes
Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit
 APPLICATION NOTE Ion PGM System Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit Key findings The Ion PGM System, in concert with the Ion ReproSeq PGS View Kit and Ion Reporter
APPLICATION NOTE Ion PGM System Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit Key findings The Ion PGM System, in concert with the Ion ReproSeq PGS View Kit and Ion Reporter
HEPATOCELLULAR CARCINOMA: SCREENING, DIAGNOSIS, AND TREATMENT
 HEPATOCELLULAR CARCINOMA: SCREENING, DIAGNOSIS, AND TREATMENT INTRODUCTION: Hepatocellular carcinoma (HCC): Fifth most common cancer worldwide Third most common cause of cancer mortality In Egypt: 2.3%
HEPATOCELLULAR CARCINOMA: SCREENING, DIAGNOSIS, AND TREATMENT INTRODUCTION: Hepatocellular carcinoma (HCC): Fifth most common cancer worldwide Third most common cause of cancer mortality In Egypt: 2.3%
Compute-aided Differentiation of Focal Liver Disease in MR Imaging
 1063 Compute-aided Differentiation of Focal Liver Disease in MR Imaging Xuejun Zhang a, Masayuki Kanematsu b, Hiroshi Fujita c, Takeshi Hara c, Hiroshi Kondo b, Xiangrong Zhou c, Wenguang Li a and Hiroaki
1063 Compute-aided Differentiation of Focal Liver Disease in MR Imaging Xuejun Zhang a, Masayuki Kanematsu b, Hiroshi Fujita c, Takeshi Hara c, Hiroshi Kondo b, Xiangrong Zhou c, Wenguang Li a and Hiroaki
of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed.
 Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Computational Analysis of Genome-Wide DNA Copy Number Changes
 Computational Analysis of Genome-Wide DNA Copy Number Changes Lei Song Thesis submitted to the faculty of the Virginia Polytechnic Institute and State University in partial fulfillment of the requirements
Computational Analysis of Genome-Wide DNA Copy Number Changes Lei Song Thesis submitted to the faculty of the Virginia Polytechnic Institute and State University in partial fulfillment of the requirements
Cytogenetics 101: Clinical Research and Molecular Genetic Technologies
 Cytogenetics 101: Clinical Research and Molecular Genetic Technologies Topics for Today s Presentation 1 Classical vs Molecular Cytogenetics 2 What acgh? 3 What is FISH? 4 What is NGS? 5 How can these
Cytogenetics 101: Clinical Research and Molecular Genetic Technologies Topics for Today s Presentation 1 Classical vs Molecular Cytogenetics 2 What acgh? 3 What is FISH? 4 What is NGS? 5 How can these
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Nature Biotechnology: doi: /nbt.1904
 Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
DNA-seq Bioinformatics Analysis: Copy Number Variation
 DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
Microarray Analysis and Liver Diseases
 Microarray Analysis and Liver Diseases Snorri S. Thorgeirsson M.D., Ph.D. Laboratory of Experimental Carcinogenesis Center for Cancer Research, NCI, NIH Application of Microarrays to Cancer Research Identifying
Microarray Analysis and Liver Diseases Snorri S. Thorgeirsson M.D., Ph.D. Laboratory of Experimental Carcinogenesis Center for Cancer Research, NCI, NIH Application of Microarrays to Cancer Research Identifying
SNPrints: Defining SNP signatures for prediction of onset in complex diseases
 SNPrints: Defining SNP signatures for prediction of onset in complex diseases Linda Liu, Biomedical Informatics, Stanford University Daniel Newburger, Biomedical Informatics, Stanford University Grace
SNPrints: Defining SNP signatures for prediction of onset in complex diseases Linda Liu, Biomedical Informatics, Stanford University Daniel Newburger, Biomedical Informatics, Stanford University Grace
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research
 Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Mammogram Analysis: Tumor Classification
 Mammogram Analysis: Tumor Classification Literature Survey Report Geethapriya Raghavan geeragh@mail.utexas.edu EE 381K - Multidimensional Digital Signal Processing Spring 2005 Abstract Breast cancer is
Mammogram Analysis: Tumor Classification Literature Survey Report Geethapriya Raghavan geeragh@mail.utexas.edu EE 381K - Multidimensional Digital Signal Processing Spring 2005 Abstract Breast cancer is
T. R. Golub, D. K. Slonim & Others 1999
 T. R. Golub, D. K. Slonim & Others 1999 Big Picture in 1999 The Need for Cancer Classification Cancer classification very important for advances in cancer treatment. Cancers of Identical grade can have
T. R. Golub, D. K. Slonim & Others 1999 Big Picture in 1999 The Need for Cancer Classification Cancer classification very important for advances in cancer treatment. Cancers of Identical grade can have
Comparison of segmentation methods in cancer samples
 fig/logolille2. Comparison of segmentation methods in cancer samples Morgane Pierre-Jean, Guillem Rigaill, Pierre Neuvial Laboratoire Statistique et Génome Université d Évry Val d Éssonne UMR CNRS 8071
fig/logolille2. Comparison of segmentation methods in cancer samples Morgane Pierre-Jean, Guillem Rigaill, Pierre Neuvial Laboratoire Statistique et Génome Université d Évry Val d Éssonne UMR CNRS 8071
Contents. Just Classifier? Rules. Rules: example. Classification Rule Generation for Bioinformatics. Rule Extraction from a trained network
 Contents Classification Rule Generation for Bioinformatics Hyeoncheol Kim Rule Extraction from Neural Networks Algorithm Ex] Promoter Domain Hybrid Model of Knowledge and Learning Knowledge refinement
Contents Classification Rule Generation for Bioinformatics Hyeoncheol Kim Rule Extraction from Neural Networks Algorithm Ex] Promoter Domain Hybrid Model of Knowledge and Learning Knowledge refinement
Mammogram Analysis: Tumor Classification
 Mammogram Analysis: Tumor Classification Term Project Report Geethapriya Raghavan geeragh@mail.utexas.edu EE 381K - Multidimensional Digital Signal Processing Spring 2005 Abstract Breast cancer is the
Mammogram Analysis: Tumor Classification Term Project Report Geethapriya Raghavan geeragh@mail.utexas.edu EE 381K - Multidimensional Digital Signal Processing Spring 2005 Abstract Breast cancer is the
CNV Detection and Interpretation in Genomic Data
 CNV Detection and Interpretation in Genomic Data Benjamin W. Darbro, M.D., Ph.D. Assistant Professor of Pediatrics Director of the Shivanand R. Patil Cytogenetics and Molecular Laboratory Overview What
CNV Detection and Interpretation in Genomic Data Benjamin W. Darbro, M.D., Ph.D. Assistant Professor of Pediatrics Director of the Shivanand R. Patil Cytogenetics and Molecular Laboratory Overview What
Rare Variant Burden Tests. Biostatistics 666
 Rare Variant Burden Tests Biostatistics 666 Last Lecture Analysis of Short Read Sequence Data Low pass sequencing approaches Modeling haplotype sharing between individuals allows accurate variant calls
Rare Variant Burden Tests Biostatistics 666 Last Lecture Analysis of Short Read Sequence Data Low pass sequencing approaches Modeling haplotype sharing between individuals allows accurate variant calls
Molecular Markers. Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC
 Molecular Markers Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC Overview Testing methods Rationale for molecular testing
Molecular Markers Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC Overview Testing methods Rationale for molecular testing
Single SNP/Gene Analysis. Typical Results of GWAS Analysis (Single SNP Approach) Typical Results of GWAS Analysis (Single SNP Approach)
 High-Throughput Sequencing Course Gene-Set Analysis Biostatistics and Bioinformatics Summer 28 Section Introduction What is Gene Set Analysis? Many names for gene set analysis: Pathway analysis Gene set
High-Throughput Sequencing Course Gene-Set Analysis Biostatistics and Bioinformatics Summer 28 Section Introduction What is Gene Set Analysis? Many names for gene set analysis: Pathway analysis Gene set
Identification of regions with common copy-number variations using SNP array
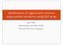 Identification of regions with common copy-number variations using SNP array Agus Salim Epidemiology and Public Health National University of Singapore Copy Number Variation (CNV) Copy number alteration
Identification of regions with common copy-number variations using SNP array Agus Salim Epidemiology and Public Health National University of Singapore Copy Number Variation (CNV) Copy number alteration
EARLY STAGE DIAGNOSIS OF LUNG CANCER USING CT-SCAN IMAGES BASED ON CELLULAR LEARNING AUTOMATE
 EARLY STAGE DIAGNOSIS OF LUNG CANCER USING CT-SCAN IMAGES BASED ON CELLULAR LEARNING AUTOMATE SAKTHI NEELA.P.K Department of M.E (Medical electronics) Sengunthar College of engineering Namakkal, Tamilnadu,
EARLY STAGE DIAGNOSIS OF LUNG CANCER USING CT-SCAN IMAGES BASED ON CELLULAR LEARNING AUTOMATE SAKTHI NEELA.P.K Department of M.E (Medical electronics) Sengunthar College of engineering Namakkal, Tamilnadu,
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data
 Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Improved Intelligent Classification Technique Based On Support Vector Machines
 Improved Intelligent Classification Technique Based On Support Vector Machines V.Vani Asst.Professor,Department of Computer Science,JJ College of Arts and Science,Pudukkottai. Abstract:An abnormal growth
Improved Intelligent Classification Technique Based On Support Vector Machines V.Vani Asst.Professor,Department of Computer Science,JJ College of Arts and Science,Pudukkottai. Abstract:An abnormal growth
Extraction of Unwanted Noise in Electrocardiogram (ECG) Signals Using Discrete Wavelet Transformation
 Extraction of Unwanted Noise in Electrocardiogram (ECG) Signals Using Discrete Wavelet Transformation Er. Manpreet Kaur 1, Er. Gagandeep Kaur 2 M.Tech (CSE), RIMT Institute of Engineering & Technology,
Extraction of Unwanted Noise in Electrocardiogram (ECG) Signals Using Discrete Wavelet Transformation Er. Manpreet Kaur 1, Er. Gagandeep Kaur 2 M.Tech (CSE), RIMT Institute of Engineering & Technology,
Understanding The Genetics of Diamond Blackfan Anemia
 Understanding The Genetics of Diamond Blackfan Anemia Jason Farrar, MD jefarrar@ About Me Assistant Professor of Pediatrics at University of Arkansas for Medical Sciences & Arkansas Children s Hospital
Understanding The Genetics of Diamond Blackfan Anemia Jason Farrar, MD jefarrar@ About Me Assistant Professor of Pediatrics at University of Arkansas for Medical Sciences & Arkansas Children s Hospital
cn.mops - Mixture of Poissons for CNV detection in NGS data Günter Klambauer Institute of Bioinformatics, Johannes Kepler University Linz
 Software Manual Institute of Bioinformatics, Johannes Kepler University Linz cn.mops - Mixture of Poissons for CNV detection in NGS data Günter Klambauer Institute of Bioinformatics, Johannes Kepler University
Software Manual Institute of Bioinformatics, Johannes Kepler University Linz cn.mops - Mixture of Poissons for CNV detection in NGS data Günter Klambauer Institute of Bioinformatics, Johannes Kepler University
Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University
 Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
OncoPhase: Quantification of somatic mutation cellular prevalence using phase information
 OncoPhase: Quantification of somatic mutation cellular prevalence using phase information Donatien Chedom-Fotso 1, 2, 3, Ahmed Ashour Ahmed 1, 2, and Christopher Yau 3, 4 1 Ovarian Cancer Cell Laboratory,
OncoPhase: Quantification of somatic mutation cellular prevalence using phase information Donatien Chedom-Fotso 1, 2, 3, Ahmed Ashour Ahmed 1, 2, and Christopher Yau 3, 4 1 Ovarian Cancer Cell Laboratory,
Development of novel algorithm by combining Wavelet based Enhanced Canny edge Detection and Adaptive Filtering Method for Human Emotion Recognition
 International Journal of Engineering Research and Development e-issn: 2278-067X, p-issn: 2278-800X, www.ijerd.com Volume 12, Issue 9 (September 2016), PP.67-72 Development of novel algorithm by combining
International Journal of Engineering Research and Development e-issn: 2278-067X, p-issn: 2278-800X, www.ijerd.com Volume 12, Issue 9 (September 2016), PP.67-72 Development of novel algorithm by combining
The role of cytogenomics in the diagnostic work-up of Chronic Lymphocytic Leukaemia
 The role of cytogenomics in the diagnostic work-up of Chronic Lymphocytic Leukaemia Adrian Zordan, Meaghan Wall, Ruth MacKinnon, Pina D Achille & Lynda Campbell Victorian Cancer Cytogenetics Service (VCCS)
The role of cytogenomics in the diagnostic work-up of Chronic Lymphocytic Leukaemia Adrian Zordan, Meaghan Wall, Ruth MacKinnon, Pina D Achille & Lynda Campbell Victorian Cancer Cytogenetics Service (VCCS)
SNP array-based analyses of unbalanced embryos as a reference to distinguish between balanced translocation carrier and normal blastocysts
 J Assist Reprod Genet (2016) 33:1115 1119 DOI 10.1007/s10815-016-0734-0 TECHNOLOGICAL INNOVATIONS SNP array-based analyses of unbalanced embryos as a reference to distinguish between balanced translocation
J Assist Reprod Genet (2016) 33:1115 1119 DOI 10.1007/s10815-016-0734-0 TECHNOLOGICAL INNOVATIONS SNP array-based analyses of unbalanced embryos as a reference to distinguish between balanced translocation
Challenges of CGH array testing in children with developmental delay. Dr Sally Davies 17 th September 2014
 Challenges of CGH array testing in children with developmental delay Dr Sally Davies 17 th September 2014 CGH array What is CGH array? Understanding the test Benefits Results to expect Consent issues Ethical
Challenges of CGH array testing in children with developmental delay Dr Sally Davies 17 th September 2014 CGH array What is CGH array? Understanding the test Benefits Results to expect Consent issues Ethical
Aspects of Statistical Modelling & Data Analysis in Gene Expression Genomics. Mike West Duke University
 Aspects of Statistical Modelling & Data Analysis in Gene Expression Genomics Mike West Duke University Papers, software, many links: www.isds.duke.edu/~mw ABS04 web site: Lecture slides, stats notes, papers,
Aspects of Statistical Modelling & Data Analysis in Gene Expression Genomics Mike West Duke University Papers, software, many links: www.isds.duke.edu/~mw ABS04 web site: Lecture slides, stats notes, papers,
The Hepatitis B Vaccine: What Went Wrong?
 The Hepatitis B Vaccine: What Went Wrong? By F. Edward Yazbak, MD, FAAP On Dec. 23, 2005, the Centers for Disease Control and Prevention (CDC) published a Morbidity and Mortality Weekly Report (1) in which
The Hepatitis B Vaccine: What Went Wrong? By F. Edward Yazbak, MD, FAAP On Dec. 23, 2005, the Centers for Disease Control and Prevention (CDC) published a Morbidity and Mortality Weekly Report (1) in which
Cancer Cells Detection using OTSU Threshold Algorithm
 Cancer Cells Detection using OTSU Threshold Algorithm Nalluri Sunny 1 Velagapudi Ramakrishna Siddhartha Engineering College Mithinti Srikanth 2 Velagapudi Ramakrishna Siddhartha Engineering College Kodali
Cancer Cells Detection using OTSU Threshold Algorithm Nalluri Sunny 1 Velagapudi Ramakrishna Siddhartha Engineering College Mithinti Srikanth 2 Velagapudi Ramakrishna Siddhartha Engineering College Kodali
Molecular Genetics of Paediatric Tumours. Gino Somers MBBS, BMedSci, PhD, FRCPA Pathologist-in-Chief Hospital for Sick Children, Toronto, ON, CANADA
 Molecular Genetics of Paediatric Tumours Gino Somers MBBS, BMedSci, PhD, FRCPA Pathologist-in-Chief Hospital for Sick Children, Toronto, ON, CANADA Financial Disclosure NanoString - conference costs for
Molecular Genetics of Paediatric Tumours Gino Somers MBBS, BMedSci, PhD, FRCPA Pathologist-in-Chief Hospital for Sick Children, Toronto, ON, CANADA Financial Disclosure NanoString - conference costs for
5/2/18. After this class students should be able to: Stephanie Moon, Ph.D. - GWAS. How do we distinguish Mendelian from non-mendelian traits?
 corebio II - genetics: WED 25 April 2018. 2018 Stephanie Moon, Ph.D. - GWAS After this class students should be able to: 1. Compare and contrast methods used to discover the genetic basis of traits or
corebio II - genetics: WED 25 April 2018. 2018 Stephanie Moon, Ph.D. - GWAS After this class students should be able to: 1. Compare and contrast methods used to discover the genetic basis of traits or
Human Cancer Genome Project. Bioinformatics/Genomics of Cancer:
 Bioinformatics/Genomics of Cancer: Professor of Computer Science, Mathematics and Cell Biology Courant Institute, NYU School of Medicine, Tata Institute of Fundamental Research, and Mt. Sinai School of
Bioinformatics/Genomics of Cancer: Professor of Computer Science, Mathematics and Cell Biology Courant Institute, NYU School of Medicine, Tata Institute of Fundamental Research, and Mt. Sinai School of
Comparing CNV detection methods for SNP arrays Laura Winchester, Christopher Yau and Jiannis Ragoussis
 BRIEFINGS IN FUNCTIONAL GENOMICS AND PROTEOMICS. VOL 8. NO 5. 353^366 doi:10.1093/bfgp/elp017 Comparing CNV detection methods for SNP arrays Laura Winchester, Christopher Yau and Jiannis Ragoussis Advance
BRIEFINGS IN FUNCTIONAL GENOMICS AND PROTEOMICS. VOL 8. NO 5. 353^366 doi:10.1093/bfgp/elp017 Comparing CNV detection methods for SNP arrays Laura Winchester, Christopher Yau and Jiannis Ragoussis Advance
Association mapping (qualitative) Association scan, quantitative. Office hours Wednesday 3-4pm 304A Stanley Hall. Association scan, qualitative
 Association mapping (qualitative) Office hours Wednesday 3-4pm 304A Stanley Hall Fig. 11.26 Association scan, qualitative Association scan, quantitative osteoarthritis controls χ 2 test C s G s 141 47
Association mapping (qualitative) Office hours Wednesday 3-4pm 304A Stanley Hall Fig. 11.26 Association scan, qualitative Association scan, quantitative osteoarthritis controls χ 2 test C s G s 141 47
Genetic alterations of histone lysine methyltransferases and their significance in breast cancer
 Genetic alterations of histone lysine methyltransferases and their significance in breast cancer Supplementary Materials and Methods Phylogenetic tree of the HMT superfamily The phylogeny outlined in the
Genetic alterations of histone lysine methyltransferases and their significance in breast cancer Supplementary Materials and Methods Phylogenetic tree of the HMT superfamily The phylogeny outlined in the
Diagnosis of Liver Tumor Using 3D Segmentation Method for Selective Internal Radiation Therapy
 Diagnosis of Liver Tumor Using 3D Segmentation Method for Selective Internal Radiation Therapy K. Geetha & S. Poonguzhali Department of Electronics and Communication Engineering, Campus of CEG Anna University,
Diagnosis of Liver Tumor Using 3D Segmentation Method for Selective Internal Radiation Therapy K. Geetha & S. Poonguzhali Department of Electronics and Communication Engineering, Campus of CEG Anna University,
Global variation in copy number in the human genome
 Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
Gene Selection for Tumor Classification Using Microarray Gene Expression Data
 Gene Selection for Tumor Classification Using Microarray Gene Expression Data K. Yendrapalli, R. Basnet, S. Mukkamala, A. H. Sung Department of Computer Science New Mexico Institute of Mining and Technology
Gene Selection for Tumor Classification Using Microarray Gene Expression Data K. Yendrapalli, R. Basnet, S. Mukkamala, A. H. Sung Department of Computer Science New Mexico Institute of Mining and Technology
Cytogenetics Technologies, Companies & Markets
 Cytogenetics Technologies, Companies & Markets By Prof. K. K. Jain MD, FRACS, FFPM Jain PharmaBiotech Basel, Switzerland November 2018 A Jain PharmaBiotech Report A U T H O R ' S B I O G R A P H Y Professor
Cytogenetics Technologies, Companies & Markets By Prof. K. K. Jain MD, FRACS, FFPM Jain PharmaBiotech Basel, Switzerland November 2018 A Jain PharmaBiotech Report A U T H O R ' S B I O G R A P H Y Professor
IJREAS Volume 2, Issue 2 (February 2012) ISSN: LUNG CANCER DETECTION USING DIGITAL IMAGE PROCESSING ABSTRACT
 LUNG CANCER DETECTION USING DIGITAL IMAGE PROCESSING Anita Chaudhary* Sonit Sukhraj Singh* ABSTRACT In recent years the image processing mechanisms are used widely in several medical areas for improving
LUNG CANCER DETECTION USING DIGITAL IMAGE PROCESSING Anita Chaudhary* Sonit Sukhraj Singh* ABSTRACT In recent years the image processing mechanisms are used widely in several medical areas for improving
Mosaic loss of chromosome Y in peripheral blood is associated with shorter survival and higher risk of cancer
 Supplementary Information Mosaic loss of chromosome Y in peripheral blood is associated with shorter survival and higher risk of cancer Lars A. Forsberg, Chiara Rasi, Niklas Malmqvist, Hanna Davies, Saichand
Supplementary Information Mosaic loss of chromosome Y in peripheral blood is associated with shorter survival and higher risk of cancer Lars A. Forsberg, Chiara Rasi, Niklas Malmqvist, Hanna Davies, Saichand
CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE. Dr. Bahar Naghavi
 2 CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE Dr. Bahar Naghavi Assistant professor of Basic Science Department, Shahid Beheshti University of Medical Sciences, Tehran,Iran 3 Introduction Over 4000
2 CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE Dr. Bahar Naghavi Assistant professor of Basic Science Department, Shahid Beheshti University of Medical Sciences, Tehran,Iran 3 Introduction Over 4000
Introduction to Discrimination in Microarray Data Analysis
 Introduction to Discrimination in Microarray Data Analysis Jane Fridlyand CBMB University of California, San Francisco Genentech Hall Auditorium, Mission Bay, UCSF October 23, 2004 1 Case Study: Van t
Introduction to Discrimination in Microarray Data Analysis Jane Fridlyand CBMB University of California, San Francisco Genentech Hall Auditorium, Mission Bay, UCSF October 23, 2004 1 Case Study: Van t
Integrated Analysis of Copy Number and Gene Expression
 Integrated Analysis of Copy Number and Gene Expression Nexus Copy Number provides user-friendly interface and functionalities to integrate copy number analysis with gene expression results for the purpose
Integrated Analysis of Copy Number and Gene Expression Nexus Copy Number provides user-friendly interface and functionalities to integrate copy number analysis with gene expression results for the purpose
Genomic complexity and arrays in CLL. Gian Matteo Rigolin, MD, PhD St. Anna University Hospital Ferrara, Italy
 Genomic complexity and arrays in CLL Gian Matteo Rigolin, MD, PhD St. Anna University Hospital Ferrara, Italy Clinical relevance of genomic complexity (GC) in CLL GC has been identified as a critical negative
Genomic complexity and arrays in CLL Gian Matteo Rigolin, MD, PhD St. Anna University Hospital Ferrara, Italy Clinical relevance of genomic complexity (GC) in CLL GC has been identified as a critical negative
Reading Assignments: Lecture 18: Visual Pre-Processing. Chapters TMB Brain Theory and Artificial Intelligence
 Brain Theory and Artificial Intelligence Lecture 18: Visual Pre-Processing. Reading Assignments: Chapters TMB2 3.3. 1 Low-Level Processing Remember: Vision as a change in representation. At the low-level,
Brain Theory and Artificial Intelligence Lecture 18: Visual Pre-Processing. Reading Assignments: Chapters TMB2 3.3. 1 Low-Level Processing Remember: Vision as a change in representation. At the low-level,
LTA Analysis of HapMap Genotype Data
 LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
Lung Cancer Diagnosis from CT Images Using Fuzzy Inference System
 Lung Cancer Diagnosis from CT Images Using Fuzzy Inference System T.Manikandan 1, Dr. N. Bharathi 2 1 Associate Professor, Rajalakshmi Engineering College, Chennai-602 105 2 Professor, Velammal Engineering
Lung Cancer Diagnosis from CT Images Using Fuzzy Inference System T.Manikandan 1, Dr. N. Bharathi 2 1 Associate Professor, Rajalakshmi Engineering College, Chennai-602 105 2 Professor, Velammal Engineering
The impact of the treatment of HCV in developing Hepatocellular Carcinoma
 The impact of the treatment of HCV in developing Hepatocellular Carcinoma Paul Y Kwo, MD Professor of Medicine Medical Director, Liver Transplantation Gastroenterology/Hepatology Division Indiana University
The impact of the treatment of HCV in developing Hepatocellular Carcinoma Paul Y Kwo, MD Professor of Medicine Medical Director, Liver Transplantation Gastroenterology/Hepatology Division Indiana University
Quick detection of QRS complexes and R-waves using a wavelet transform and K-means clustering
 Bio-Medical Materials and Engineering 26 (2015) S1059 S1065 DOI 10.3233/BME-151402 IOS Press S1059 Quick detection of QRS complexes and R-waves using a wavelet transform and K-means clustering Yong Xia
Bio-Medical Materials and Engineering 26 (2015) S1059 S1065 DOI 10.3233/BME-151402 IOS Press S1059 Quick detection of QRS complexes and R-waves using a wavelet transform and K-means clustering Yong Xia
2) Cases and controls were genotyped on different platforms. The comparability of the platforms should be discussed.
 Reviewers' Comments: Reviewer #1 (Remarks to the Author) The manuscript titled 'Association of variations in HLA-class II and other loci with susceptibility to lung adenocarcinoma with EGFR mutation' evaluated
Reviewers' Comments: Reviewer #1 (Remarks to the Author) The manuscript titled 'Association of variations in HLA-class II and other loci with susceptibility to lung adenocarcinoma with EGFR mutation' evaluated
CS2220 Introduction to Computational Biology
 CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
FONS Nové sekvenační technologie vklinickédiagnostice?
 FONS 2010 Nové sekvenační technologie vklinickédiagnostice? Sekvenování amplikonů Sequence capture Celogenomové sekvenování FONS 2010 Sekvenování amplikonů Amplicon sequencing - amplicon sequencing enables
FONS 2010 Nové sekvenační technologie vklinickédiagnostice? Sekvenování amplikonů Sequence capture Celogenomové sekvenování FONS 2010 Sekvenování amplikonů Amplicon sequencing - amplicon sequencing enables
Chapter 1 : Genetics 101
 Chapter 1 : Genetics 101 Understanding the underlying concepts of human genetics and the role of genes, behavior, and the environment will be important to appropriately collecting and applying genetic
Chapter 1 : Genetics 101 Understanding the underlying concepts of human genetics and the role of genes, behavior, and the environment will be important to appropriately collecting and applying genetic
White Paper Estimating Complex Phenotype Prevalence Using Predictive Models
 White Paper 23-12 Estimating Complex Phenotype Prevalence Using Predictive Models Authors: Nicholas A. Furlotte Aaron Kleinman Robin Smith David Hinds Created: September 25 th, 2015 September 25th, 2015
White Paper 23-12 Estimating Complex Phenotype Prevalence Using Predictive Models Authors: Nicholas A. Furlotte Aaron Kleinman Robin Smith David Hinds Created: September 25 th, 2015 September 25th, 2015
Are we adequately screening at-risk patients for hepatocellular carcinoma in the outpatient setting?
 Rajani Sharma, PGY1 Geriatrics CRC Project, 12/19/13 Are we adequately screening at-risk patients for hepatocellular carcinoma in the outpatient setting? A. Study Purpose and Rationale Hepatocellular carcinoma
Rajani Sharma, PGY1 Geriatrics CRC Project, 12/19/13 Are we adequately screening at-risk patients for hepatocellular carcinoma in the outpatient setting? A. Study Purpose and Rationale Hepatocellular carcinoma
Efficacy of the Extended Principal Orthogonal Decomposition Method on DNA Microarray Data in Cancer Detection
 202 4th International onference on Bioinformatics and Biomedical Technology IPBEE vol.29 (202) (202) IASIT Press, Singapore Efficacy of the Extended Principal Orthogonal Decomposition on DA Microarray
202 4th International onference on Bioinformatics and Biomedical Technology IPBEE vol.29 (202) (202) IASIT Press, Singapore Efficacy of the Extended Principal Orthogonal Decomposition on DA Microarray
Bayesian Prediction Tree Models
 Bayesian Prediction Tree Models Statistical Prediction Tree Modelling for Clinico-Genomics Clinical gene expression data - expression signatures, profiling Tree models for predictive sub-typing Combining
Bayesian Prediction Tree Models Statistical Prediction Tree Modelling for Clinico-Genomics Clinical gene expression data - expression signatures, profiling Tree models for predictive sub-typing Combining
Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD
 Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD Department of Biomedical Informatics Department of Computer Science and Engineering The Ohio State University Review
Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD Department of Biomedical Informatics Department of Computer Science and Engineering The Ohio State University Review
EXTRACT THE BREAST CANCER IN MAMMOGRAM IMAGES
 International Journal of Civil Engineering and Technology (IJCIET) Volume 10, Issue 02, February 2019, pp. 96-105, Article ID: IJCIET_10_02_012 Available online at http://www.iaeme.com/ijciet/issues.asp?jtype=ijciet&vtype=10&itype=02
International Journal of Civil Engineering and Technology (IJCIET) Volume 10, Issue 02, February 2019, pp. 96-105, Article ID: IJCIET_10_02_012 Available online at http://www.iaeme.com/ijciet/issues.asp?jtype=ijciet&vtype=10&itype=02
Human Genetics 542 Winter 2018 Syllabus
 Human Genetics 542 Winter 2018 Syllabus Monday, Wednesday, and Friday 9 10 a.m. 5915 Buhl Course Director: Tony Antonellis Jan 3 rd Wed Mapping disease genes I: inheritance patterns and linkage analysis
Human Genetics 542 Winter 2018 Syllabus Monday, Wednesday, and Friday 9 10 a.m. 5915 Buhl Course Director: Tony Antonellis Jan 3 rd Wed Mapping disease genes I: inheritance patterns and linkage analysis
Predicting Breast Cancer Survival Using Treatment and Patient Factors
 Predicting Breast Cancer Survival Using Treatment and Patient Factors William Chen wchen808@stanford.edu Henry Wang hwang9@stanford.edu 1. Introduction Breast cancer is the leading type of cancer in women
Predicting Breast Cancer Survival Using Treatment and Patient Factors William Chen wchen808@stanford.edu Henry Wang hwang9@stanford.edu 1. Introduction Breast cancer is the leading type of cancer in women
Targeted qpcr. Debate on PGS Technology: Targeted vs. Whole genome approach. Discolsure Stake shareholder of GENETYX S.R.L
 Antonio Capalbo, PhD Laboratory Director GENETYX, reproductive genetics laboratory, Italy PGT responsible GENERA centers for reproductive medicine, Italy Debate on PGS Technology: Targeted vs. Whole genome
Antonio Capalbo, PhD Laboratory Director GENETYX, reproductive genetics laboratory, Italy PGT responsible GENERA centers for reproductive medicine, Italy Debate on PGS Technology: Targeted vs. Whole genome
Research Strategy: 1. Background and Significance
 Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
Lung Tumour Detection by Applying Watershed Method
 International Journal of Computational Intelligence Research ISSN 0973-1873 Volume 13, Number 5 (2017), pp. 955-964 Research India Publications http://www.ripublication.com Lung Tumour Detection by Applying
International Journal of Computational Intelligence Research ISSN 0973-1873 Volume 13, Number 5 (2017), pp. 955-964 Research India Publications http://www.ripublication.com Lung Tumour Detection by Applying
EECS 433 Statistical Pattern Recognition
 EECS 433 Statistical Pattern Recognition Ying Wu Electrical Engineering and Computer Science Northwestern University Evanston, IL 60208 http://www.eecs.northwestern.edu/~yingwu 1 / 19 Outline What is Pattern
EECS 433 Statistical Pattern Recognition Ying Wu Electrical Engineering and Computer Science Northwestern University Evanston, IL 60208 http://www.eecs.northwestern.edu/~yingwu 1 / 19 Outline What is Pattern
Molecular Methods in the Diagnosis and Prognostication of Melanoma: Pros & Cons
 Molecular Methods in the Diagnosis and Prognostication of Melanoma: Pros & Cons Ben J. Friedman, MD Senior Staff Physician Department of Dermatology Department of Pathology and Laboratory Medicine Henry
Molecular Methods in the Diagnosis and Prognostication of Melanoma: Pros & Cons Ben J. Friedman, MD Senior Staff Physician Department of Dermatology Department of Pathology and Laboratory Medicine Henry
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Chromothripsis: A New Mechanism For Tumorigenesis? i Fellow s Conference Cheryl Carlson 6/10/2011
 Chromothripsis: A New Mechanism For Tumorigenesis? i Fellow s Conference Cheryl Carlson 6/10/2011 Massive Genomic Rearrangement Acquired in a Single Catastrophic Event during Cancer Development Cell 144,
Chromothripsis: A New Mechanism For Tumorigenesis? i Fellow s Conference Cheryl Carlson 6/10/2011 Massive Genomic Rearrangement Acquired in a Single Catastrophic Event during Cancer Development Cell 144,
Title: Pinpointing resilience in Bipolar Disorder
 Title: Pinpointing resilience in Bipolar Disorder 1. AIM OF THE RESEARCH AND BRIEF BACKGROUND Bipolar disorder (BD) is a mood disorder characterised by episodes of depression and mania. It ranks as one
Title: Pinpointing resilience in Bipolar Disorder 1. AIM OF THE RESEARCH AND BRIEF BACKGROUND Bipolar disorder (BD) is a mood disorder characterised by episodes of depression and mania. It ranks as one
Human Genetics 542 Winter 2017 Syllabus
 Human Genetics 542 Winter 2017 Syllabus Monday, Wednesday, and Friday 9 10 a.m. 5915 Buhl Course Director: Tony Antonellis Module I: Mapping and characterizing simple genetic diseases Jan 4 th Wed Mapping
Human Genetics 542 Winter 2017 Syllabus Monday, Wednesday, and Friday 9 10 a.m. 5915 Buhl Course Director: Tony Antonellis Module I: Mapping and characterizing simple genetic diseases Jan 4 th Wed Mapping
Hepatocellular Carcinoma: Epidemiology and Screening
 Hepatocellular Carcinoma: Epidemiology and Screening W. Ray Kim, MD Professor and Chief Gastroenterology and Hepatology Stanford University School of Medicine Case A 67 year old Filipino-American woman
Hepatocellular Carcinoma: Epidemiology and Screening W. Ray Kim, MD Professor and Chief Gastroenterology and Hepatology Stanford University School of Medicine Case A 67 year old Filipino-American woman
Hepatitis B virus X gene in the development of hepatocellular carcinoma. Citation Hong Kong Medical Journal, 2011, v. 17 n. 6, suppl. 6, p.
 Title Hepatitis B virus X gene in the development of hepatocellular carcinoma Author(s) Ma, NF; Lau, SH; Hu, L; Dong, SS; Guan, XY Citation Hong Kong Medical Journal, 2011, v. 17 n. 6, suppl. 6, p. 44-47
Title Hepatitis B virus X gene in the development of hepatocellular carcinoma Author(s) Ma, NF; Lau, SH; Hu, L; Dong, SS; Guan, XY Citation Hong Kong Medical Journal, 2011, v. 17 n. 6, suppl. 6, p. 44-47
How Does Analysis of Competing Hypotheses (ACH) Improve Intelligence Analysis?
 How Does Analysis of Competing Hypotheses (ACH) Improve Intelligence Analysis? Richards J. Heuer, Jr. Version 1.2, October 16, 2005 This document is from a collection of works by Richards J. Heuer, Jr.
How Does Analysis of Competing Hypotheses (ACH) Improve Intelligence Analysis? Richards J. Heuer, Jr. Version 1.2, October 16, 2005 This document is from a collection of works by Richards J. Heuer, Jr.
Nature Genetics: doi: /ng Supplementary Figure 1. Rates of different mutation types in CRC.
 Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
Autoantibodies as additive biomarkers to AFP for the detection of HCC
 Autoantibodies as additive biomarkers to AFP for the detection of HCC Christopher Welberry* 1, William Irving 2, Andrea Murray 3, Caroline Chapman 4 1 Division of Medical Sciences and Graduate Entry Medicine
Autoantibodies as additive biomarkers to AFP for the detection of HCC Christopher Welberry* 1, William Irving 2, Andrea Murray 3, Caroline Chapman 4 1 Division of Medical Sciences and Graduate Entry Medicine
Jose D Sollano, MD Professor of Medicine University of Santo Tomas Manila, Philippines. University of Santo Tomas
 Jose D Sollano, MD Professor of Medicine Manila, Philippines International Variation in Age-Standardized Liver Cancer Incidence Rates in Both Sexes, 2008 Global Age-Standardized Liver Cancer Incidence
Jose D Sollano, MD Professor of Medicine Manila, Philippines International Variation in Age-Standardized Liver Cancer Incidence Rates in Both Sexes, 2008 Global Age-Standardized Liver Cancer Incidence
Basics of Diagnostic DNA Microarray Technology. Case Study: Hepatocellular Carcinoma
 Review Basics of Diagnostic DNA Microarray Technology. Case Study: Hepatocellular Carcinoma ANAS EL-ANEED and JOSEPH BANOUB Memorial University of Newfoundland, Biochemistry Department, St. John s, NL,
Review Basics of Diagnostic DNA Microarray Technology. Case Study: Hepatocellular Carcinoma ANAS EL-ANEED and JOSEPH BANOUB Memorial University of Newfoundland, Biochemistry Department, St. John s, NL,
