Chromatin structure and epigenetic regulation of eukaryotic gene expression. Giacomo Cavalli
|
|
|
- Frederick Nichols
- 5 years ago
- Views:
Transcription
1 Chromatin structure and epigenetic regulation of eukaryotic gene expression Giacomo Cavalli
2 10,000 nm DNA compaction in a human nucleus 11 nm 30nm 1bp (0.3nm) compact size DNA length compaction nucleus (human) 2 x 23 = 46 chromosomes 92 DNA molecules 10 µ m ball 12,000 Mbp 4 m DNA 400,000 x mitotic chromosome 2 chromatids, 1 µ m thick 2 DNA molecules 10 µ m long X 2x 130 Mbp 2x 43 mm DNA 10,000 x DNA domain anchored DNA loop 1 replicon? 60 nm x 0.5 µ m 60 kbp 20 µ m DNA 35 x chromatin fiber approx. 6 nucleosomes per turn of 11 nm 30 nm diameter 1200 bp 400 nm DNA 35 x nucleosome disk 1? turn of DNA (146 bp) + linker DNA 6 x 11 nm 200 bp 66 nm DNA 6-11 x base pair 0.33 x 1.1 nm 1 bp 0.33 nm DNA 1 x Compaction of DNA by histones Compaction by chromosome scaffold / nuclear matrix JHW 3-99 Copyright 1999, J.H.Waterborg, UMKC
3 H1 Linker histone H2A HISTONES are highly conserved, small, basic proteins H2B Core histones JHW H3 H4 N helix variable conserved Histone acetylation is a reversible modification of lysines in the N-termini of the core histones. Result: reduced binding to DNA destabilization of chromatin 3-99 Copyright 1999, J.H.Waterborg, UMKC
4 Core Histones The histone-fold JHW The basic structure of ALL core histones is the same: 1 long hydrophobic alpha-helix, bordered by 2 short hydrophobic alpha helices that form pairs H2A - H2B and H3 - H4 which interact. References: Moudrianakis et al. PNAS 88, (1991); PNAS 90, (1993); PNAS 92, (1995) 3-99 Copyright 1999, J.H.Waterborg, UMKC
5 Histone octamer assembly JHW H3-H4 tetramer Histone octamer H2A-H2B dimer 3-99 Copyright 1999, J.H.Waterborg, UMKC
6 Nucleosome assembly Histone octamer bp of DNA An octamer (8 units) of histones: 4 core histones H2A, H2B, H3, H4 (X2) Nucleosome --> The histone octamer organizes 145 bp of DNA in 1 3/4 helical turn of DNA: 48 nm of DNA packaged in a disc of 6 x 11nm
7 The Nucleosome as the fundamental chromatin unit JHW 3-99 Copyright 1999, J.H.Waterborg, UMKC
8 bp DNA in a 1.65 turns of a flat, left-handed superhelix one pseudo twofold axis centered at the dyad (reference: 0 helical turns) one base-pair precisely at the dyad sharp bends at and turns Histone-fold domains organize 121 bp of DNA. The DNA is bound at 10 bp intervals through many contacts, including penetration of arginines at all 14 minor grooves facing the protein core The grooves from neighboring DNA turns line up; forming channels H3 and H2B N-termini exit one of these channels every 20bp. The H4 tail establishes contacts with the next core particle. H2A JHW H2B H3 H Luger, Mader, Richmond, Sargent & Richmond Copyright 1999, J.H.Waterborg, UMKC Nucleosome features
9 Chromatosome C Core histone octamer + 1 Linker Histone + 2 full turns of DNA (168 bp) N H1 H3 H3 1 mm JHW Zhou, Gerchman, Ramakrishnan, Travers, Muyldermans Nature 395, 402 (1998) 5 mm Linker Histone and histone termini control linker DNA entry/exit of chromatosome in chromatin fiber. An, Leuba, van Holde, Zlatanova PNAS 95, 3396 (1998) 3-99 Copyright 1999, J.H.Waterborg, UMKC
10 Chromatin fibers 30 nm chromatin fiber 11 nm (beads) + charged N termini (bind DNA on neigboring nucleosomes) HIGH level of histone H1 NO gene transcription JHW highly acetylated core histones (especially H3 and H4) Reduced level of histone H1 Gene transcription possible 3-99 Copyright 1999, J.H.Waterborg, UMKC
11 Alternative chromatin fiber models Schalch, Duda, Sargent & Richmond, Nature 2005 Zigzag fiber
12 Molecular basis of epigenetics - Chromatin regulation via post-translational modifications of histones and the action of chromatin proteins such as HP1, Polycomb et Trithorax - Histone variants - DNA methylation - Noncoding RNAs - Links between DNA methylation, histone modifications and the function of ncrnas
13
14 Post translational histone modifications
15 Histone modifications 27 Cell (2002) 111,
16 Various PTMs Review from Gardner, Allis and Strahl et al, J.Mol.Biol. 2011
17 Acetylation of conserved lysines The N-termini of histones H4 and H3, and their acetylation patterns, are absolutely conserved Ac Ac Ac Ac 9 Ac 18 Ac 14 Ac Acetyl-CoA 23 Ac - O 27 Ac or Me P O DNA P backbone binding Histone Deacetylase reversible reactions CoA O O C C γ C ε N+ O C α β C δ C HAT (Histone Acetyl-Transferase) C C O C C ε C C C N C O no DNA binding P N ε-n-acetyl-lysine JHW Ac or Me A-R-T-K-Q-T-A-R-K-S-T-G-G-K-A-P-R-K-Q-L-A-T-K-A-A-R-K-S-A-P N Lysine 20 Ac-S-G-R-G-K-G-G-K-G-L-G-K-G-G-A-K-R-H-R-K-V-L-R-D Me H3 N-terminus Copyright 1999, J.H.Waterborg, UMKC H4 N-terminus 5
18 Histone acetylation versus deacetylation Histone acetylation Decondensed chromatin: Active chromatin state Histone deacetylation Condensed chromatin: repressed transcriptional state
19 Histone Methylation / Demethylation Histones can be methylated at lysines or arginines. Example: H3 K4 methylation S-adenosylmethyionine Lysine O N C C C α β γ C C δ ε C N+ HMT (Histone Methyl-Transferase) ε-n-monomethyl-lysine S-adenosylmethyionine HMT (Histone Methyl-Transferase) ε-n-dimethyl-lysine S-adenosylmethyionine HMT (Histone Methyl-Transferase) ε-n-trimethyl-lysine O O O N C C C N C C C N C C C C C C C C C ε C ε C ε C C N+ C N+ C C N+ C C Histone demethylase Histone demethylase Histone demethylase DNA backbone binding may not be strongly affected, but specific proteins may recognize these modifications Histone demethylases found for K4, K9, K27 and K36
20 Histone methylation versus demethylation Methylation of lysines H3K4, H3K36 et H3K79 Decondensed chromatin: Active transcriptional state Demethylation of lysines H3K9, H3K27 et H4K20 Methylation of lysines H3K9, H3K27 et H4K20 Condensed chromatin: Repressed transcriptional state Demethylation of lysines H3K4, H3K36 et H3K79
21 Is there a histone code?
22 Histone modifications can be recognized by chromatin proteins in order to elicit specific functions: The concept of «writer», «reader» and «eraser»
23 «writers» and «readers» Writer: HAT: Histone acetyl-transferases HMT: Histone methyl-transferases Reader: Specific factors or macromolecular complexes Eraser: HDAC: Histone deacetylases KDM: Histone demethylases Review from Gardner, Allis and Strahl et al, J.Mol.Biol. 2011
24 Targeting specific multiprotein chromatin complexes via histone modifications Review from Gardner, Allis and Strahl et al, J.Mol.Biol. 2011
25 H3-K9me and H3-K27me recognition by the chromodomain proteins HP1 and Polycomb SUV39H1/2 (HMT) HP1 EZH1/2 (HMT) Polycomb H3-K9me2/3 H3-K27me3 PRC1 PC Constitutive Heterochromatin Polycomb silencing (Facultative Heterochromatin)
26 H3-K4me recognition by the PHD domain protein NURF-301 MLL1/2/3 TRX/ASH1 (HMT) NURF-301 H3-K4me3 Wysocka et al., Nature, May 2006 Li et al., Nature, May 2006 ISWI NURF 301 Trithorax activation ISWI-dependent Chromatin remodeling leading to Chromatin opening and activation
27 Histone Marks and Their Readers Readers identified by SILAC (Stable Isotope Labeling by Amino acids in Cell culture). Vermeulen et al., Cell 2010
28 Chromatin Immunoprecipitation (ChIP) Chromosome Histone modifica*ons Various chroma*n proteins (writers, readers, and transcrip*on factors) Principle 1- Cross- link protein- DNA interac*ons with formaldehyde 2- chroma*n fragmenta*on by sonica*on 3- immunoprecipitate using a specific an*body against protein or histone modifica,on of interest DNA amplifica,on Liga*on of Illumina linkers Forma*on of DNA clusters in a flow cell High-throughput sequencing: ChIP-seq A T C G Sequence one base at a time
29 High-resolution profiling of Histone Methylations in the Human Genome Data from Barski et al., Cell 2007
30 Same, but in Drosophila Sg4 cells Abd-B active H3K27me3 Pc Trx-C Trx-N Ash1 H3K27ac PolII H3K4me3 Abd-B Figure 1 from Schwartz et al, PlosGenetics 2010
31 H3K4me3 and H3K36me3 annotate genes and non-coding RNA The FoxP1 gene The Neat non-coding RNA Data (ChIP-seq) from Mikkelsen et al., Nature 2007
32 Polycomb and Trithorax proteins
33 PcG and trxg proteins regulate cell memory and dynamic patterns of gene expression stem cell maintenance and plasticity proliferation differentiation cell fate Determination Hox gene regulation (cell memory) PcG and trxg proteins Cancer Genomic imprinting X-chromosome inactivation Vertebrates Copyright 1999, J.H.Waterborg, UMKC
34 Opposing functions of PcG (Polycomb group) and trxg (trithorax group) complexes on chromatin Histone acetylation and Methylation (TAC1 and ASH1 complexes) H4-Ac H3-K4me3 H3-K36me3 Nucleosome remodeling (BRM/Nurf complexes) ON trxg Maintenance of active states (open chromatin) PRE/TRE Target gene Deacetylation and Methylation (ESC-E(Z) PRC2 complex) H3-K27me3 OFF PcG Maintenance of repressed states (compact chromatin) - Chromatin compaction - H2A Ubiquitination (PRC1 complex) H2A Ub
35 Methylase vs. Demethylase Polycomb vs. Trithorax H3-K27me3 vs. H3-K4me3 «The Ying and the Yang» on chromatin From Agger et al., Nature, August 2007
36 Illustration of the epigenetic memory by the PcG proteins for the regulation of the Hox Ubx gene in Drosophila Larval Discs Construction transgénique Early embryo Late embryo head wing haltere P A Polycomb Response Element Legend: The correct expression pattern is created in the embryo and epigenetically maintained throughout development when all three elements are combined: the embryonic enhancer sets the pattern, the PRE maintains the repressed or non-repressed state and the imaginal disc now remains active only posterior to parasegment 6, not in the head or wings. Illustration from a Review of Schwartz and Pirrotta
37 Data (ChIP-seq in Drosophila embryos) from Bowman et al., elife 2014
38 Hox gene regulation in mouse development Relative H3K27me From a Review of Lyons and Lomvardas, BBA 2014
39 Chromatin marks during temporal collinearity at the HoxD cluster Data (ChIP-chip in mouse embryos) from Soshnikova and Duboule, Science 2009
40 Histone variants
41 General remarks The incorporation of histone variants is an alternative way to mark chromatin. H2A and H3 variants are known since decades and they have been shown to regulate and to maintain differential chromatin states. H2A, H2B, H3 and H4 core histones are coded by gene cluster sont codées par des gènes en clusters fortement exprimé lors that are strongly expressed during the S phase of DNA replication, whereas histone variants are coded by separate genes, which are expressed throughout the cell cycle. Therefore, histone variants can be incorporated in chromatin all the time.
42 Notable examples of histone variants: CENP-A: H3-like protein found at centromeres in mammalian species. It is highly conserved throughout evolution and generally called CenH3 H3.3: differs by just 4 amino acids from canonical H3. Outside S-phase, H3.3-H4 dimers may replace H3-H4 dimers during gene activation («replacement variant») Moreover, H3.3 is enriched 2 to5 fold in activating post translational modifications (K9, K14, K18 and K23 acetylation, K4 and K79 methylation) H3.3 is an important mark of active chromatin states H2A.Z: nucleosomes containing H2A.Z are biochemically less stable than those with H2A. This variant is thus associated to active chromatin MacroH2A: specific mark of X chromosome inactivation in mammals See for reviews: 1. Zink and Hake: «Histone variants, nuclear function and disease», Curr Opin Genet Dev , 82-9 and 2. Harr, Gonzalez-Sandoval, and Gasser: «Histones and histone modifications in perinuclear chromatin anchoring: from yeast to man». EMBO Reports 2016, 17,
43 H3.3 Variant and transcriptional activation Old H3/H4 dimer evicted H2A/H2B dimer Replacement with a new H3.3/H4 dimer Ø Lead to the inheritance of an active chromatin state. Review by Henikoff, Furuyama and Ahmad: «Histone variants, nucleosome, assembly and epigenetic inheritance», TRENDS in Genetics 2004, 20:
44 MacroH2A variant and X chromosome inactivation Review by Banaszynski, Allis and Lewis: «Histone variants in Metazoan Development, Developmental Cell, 2010, 19:
45 DNA methylation
46 Ø DNA methylation: DNA modification which is not a mutation (it does not modify the base pairing code) Ø In mammals, the most frequent DNA methylation is observed at cytosines to produce 5-methylcytosine - DNA methylation is generally observed in CG dinucleotides and is symmetric: CG GC - CH3 groups are exposed in the large groove - NOTE: recently 6mA DNA methylation has been observed in multiple species. Its biological significance is currently under investigation
47 - CpGs, the methylation targets, are most concentrated in gene promoters (CpG islands), and on repetitive elements The enzymes for DNA methylation are called DNA methyltransferases (DNMTs. In human: DNMT1 is a maintenance methylase, whereas DNMT3a and DNMT3b, can methylate DNA de novo) - Specific transcriptional repressors can «read» the methyl mark and bind 5mCpG MeCP2 MBD1 MBD2
48 Epigenetic regulation involving DNA methylation - Some genes inactivated by methylation during development (IL4, but this is not a major mechanism) - X-chromosome inactivation in mammals - Polycomb-dependent silencing in mammals - Genomic imprinting - DNA methylation is frequently involved in the malignant transformation
49 X chromosome inactivation Sperm Xp a Ooocyte Xm a PREFERENTIAL INACTIVATION OF PATERNAL X (In mouse, not in human) Xm a Xp a Reactivation of Xi in the germ line X a X a 4-8 cells Xm a Xp i Morula Xm a Xp i Germ line X a X i Inner cell mass Xm a Xp i Inner cell mass Xm a Xp a ICM ICM Trophectoderme Xm a Xp i Endoderme primitif Xm a Xp i Xm a Xp i or Xm i Xp a Foetus Embryo (Epiblast) Stade blastocyste Reactivation of Xp in the ICM Epigenetic reprogramming RANDOM INACTIVATION OF MATERNAL OR PATERNAL X Régulation d ordre Epigénétique Chaumeil and Heard, Biofutur, avril 2004
50 Molecular mechanisms of X chromosome inactivation HDACs PRC2 PRC1 DNMTs Xist Ac Ac Ac Xist Me K9/27 Me K9/27 Me K9/27 Me Me PRC1 Me Me Me Ac Ac Ac Me K9/27 Me K9/27 Me K9/27 Me Me Me Me Me Active gene Reversible Xist-dependent gene repression - Establishment of histone modifications - Stable and irreversible X Inactivation (Xist independent) - DNA methylation, histone modification and histone variant deposition lock the inactive state - Me K9/27 Ac Methylation of lysines 9 and 27 of histone H3 Acetylation of histones H3 et H4 DNA methylation Chromatin = ADN + nucleosomes Adapted from Chaumeil and Heard, Biofutur, avril 2004
51 Like histone modifications, DNA methylation is reversible Sébastien A. Smallwood, Gavin Kelsey, De novo DNA methylation: a germ cell perspective, TIG, Volume 28, Issue 1, 2012, 33 42, dx.doi.org/ /j.tig
52 Like histone modifications, DNA methylation is reversible Figure 4. Routes for ten eleven translocation (Tet)-mediated DNA demethylation. Methylated cytosine (5mC) can be converted into 5-hydroxymethylcytosine (5hmC) then to 5-formylcytosine (5fC), then 5-carboxylcytosine (5caC). Alternatively, Aid/Apobec deaminases can transform 5hmC into 5-hydroxymethylUracyl (5hmU). 5caC and 5hmU can be removed by the DNA glycosylase Tdg and then by the base excision repair to finally restor a C instead of 5mC Francesco M. Piccolo, Amanda G. Fisher, Getting rid of DNA methylation, Trends Cell Biol., Volume 24, Issue 2, 2014,
53 Roles of DNA methylation regulation - The discovery of molecular pathways capable of erasing DNA methylation is recent - genome-wide detection of hmc is difficult because the technique most widely used, bisulfite sequencing, detects 5mC as well as its oxydized forms - nevertheless, it is clear that DNA methylation regulation is important in normal physiology - Furthermore, DNA methylation plays an important role in cancer, where it is frequently altered. Furthermore Dnmts and Tet genes are frequently mutated or misexpressed in cancer
54 DNA methylation and cancer ü Several diseases have genetic determinants, which may be modified by epigenetic processes, such as DNA methylation ü DNA methylation can be inhibited. In general, epigenetic phenomena are good targets for therapy because they are reversible, whereas DNA mutation can generally not be reversed ü In most cancer cells, a fraction of the CpG islands is hypermethylated whereas the rest of the genome is globally hypomethylated This implies that DNA methylation profiles of specific genes can be diagnostic at least for certain cancers
55 Methylations, Polycomb and cancer. - Stem cell Polycomb group targets are up to 12-fold more likely to have cancer-specific promoter DNA hypermethylation than non-targets. - Reversible gene repression is replaced by permanent silencing, locking the cell into a perpetual state of selfrenewal and thereby predisposing to subsequent malignant transformation. From Epigenetic stem cell signature in cancer, Nature Genetics 2006 Martin Widschwendter, Heidi Fiegl, Daniel Egle, Elisabeth Mueller-Holzner, Gilbert Spizzo, Christian Marth, Daniel J Weisenberger4, Mihaela Campan4, Joanne Young5, Ian Jacobs & Peter W Laird
56 For more info EpigeneSys network of excellence: Labex EpiGenMed: Chromatin movie: Website to download this course:
Histones modifications and variants
 Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Epigenetics DNA methylation. Biosciences 741: Genomics Fall, 2013 Week 13. DNA Methylation
 Epigenetics DNA methylation Biosciences 741: Genomics Fall, 2013 Week 13 DNA Methylation Most methylated cytosines are found in the dinucleotide sequence CG, denoted mcpg. The restriction enzyme HpaII
Epigenetics DNA methylation Biosciences 741: Genomics Fall, 2013 Week 13 DNA Methylation Most methylated cytosines are found in the dinucleotide sequence CG, denoted mcpg. The restriction enzyme HpaII
Genetics and Genomics in Medicine Chapter 6 Questions
 Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Epigenetics. Lyle Armstrong. UJ Taylor & Francis Group. f'ci Garland Science NEW YORK AND LONDON
 ... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
Gene Expression DNA RNA. Protein. Metabolites, stress, environment
 Gene Expression DNA RNA Protein Metabolites, stress, environment 1 EPIGENETICS The study of alterations in gene function that cannot be explained by changes in DNA sequence. Epigenetic gene regulatory
Gene Expression DNA RNA Protein Metabolites, stress, environment 1 EPIGENETICS The study of alterations in gene function that cannot be explained by changes in DNA sequence. Epigenetic gene regulatory
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3
 Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
Lecture 8. Eukaryotic gene regulation: post translational modifications of histones
 Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
Stem Cell Epigenetics
 Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Epigenetics Armstrong_Prelims.indd 1 04/11/2013 3:28 pm
 Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
Transcriptional repression of Xi
 Transcriptional repression of Xi Xist Transcription of Xist Xist RNA Spreading of Xist Recruitment of repression factors. Stable repression Translocated Xic cannot efficiently silence autosome regions.
Transcriptional repression of Xi Xist Transcription of Xist Xist RNA Spreading of Xist Recruitment of repression factors. Stable repression Translocated Xic cannot efficiently silence autosome regions.
R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS
 R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS EPIGENETICS THE STUDY OF CHANGES IN GENE EXPRESSION THAT ARE POTENTIALLY HERITABLE AND THAT DO NOT ENTAIL A
R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS EPIGENETICS THE STUDY OF CHANGES IN GENE EXPRESSION THAT ARE POTENTIALLY HERITABLE AND THAT DO NOT ENTAIL A
DNA methylation & demethylation
 DNA methylation & demethylation Lars Schomacher (Group Christof Niehrs) What is Epigenetics? Epigenetics is the study of heritable changes in gene expression (active versus inactive genes) that do not
DNA methylation & demethylation Lars Schomacher (Group Christof Niehrs) What is Epigenetics? Epigenetics is the study of heritable changes in gene expression (active versus inactive genes) that do not
Repressive Transcription
 Repressive Transcription The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Guenther, M. G., and R. A.
Repressive Transcription The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Guenther, M. G., and R. A.
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Epigenetic Mechanisms
 RCPA Lecture Epigenetic chanisms Jeff Craig Early Life Epigenetics Group, MCRI Dept. of Paediatrics Overview What is epigenetics? Chromatin The epigenetic code What is epigenetics? the interactions of
RCPA Lecture Epigenetic chanisms Jeff Craig Early Life Epigenetics Group, MCRI Dept. of Paediatrics Overview What is epigenetics? Chromatin The epigenetic code What is epigenetics? the interactions of
Lecture 10. Eukaryotic gene regulation: chromatin remodelling
 Lecture 10 Eukaryotic gene regulation: chromatin remodelling Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
Lecture 10 Eukaryotic gene regulation: chromatin remodelling Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
Today. Genomic Imprinting & X-Inactivation
 Today 1. Quiz (~12 min) 2. Genomic imprinting in mammals 3. X-chromosome inactivation in mammals Note that readings on Dosage Compensation and Genomic Imprinting in Mammals are on our web site. Genomic
Today 1. Quiz (~12 min) 2. Genomic imprinting in mammals 3. X-chromosome inactivation in mammals Note that readings on Dosage Compensation and Genomic Imprinting in Mammals are on our web site. Genomic
Eukaryotic Gene Regulation
 Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Are you the way you are because of the
 EPIGENETICS Are you the way you are because of the It s my fault!! Nurture Genes you inherited from your parents? Nature Experiences during your life? Similar DNA Asthma, Autism, TWINS Bipolar Disorders
EPIGENETICS Are you the way you are because of the It s my fault!! Nurture Genes you inherited from your parents? Nature Experiences during your life? Similar DNA Asthma, Autism, TWINS Bipolar Disorders
Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure
 Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure Frederick E. Domann, Ph.D. Associate Professor of Radiation Oncology The University of Iowa Iowa City,
Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure Frederick E. Domann, Ph.D. Associate Professor of Radiation Oncology The University of Iowa Iowa City,
Lecture 27. Epigenetic regulation of gene expression during development
 Lecture 27 Epigenetic regulation of gene expression during development Development of a multicellular organism is not only determined by the DNA sequence but also epigenetically through DNA methylation
Lecture 27 Epigenetic regulation of gene expression during development Development of a multicellular organism is not only determined by the DNA sequence but also epigenetically through DNA methylation
DNA Methylation and Cancer
 DNA Methylation and Cancer October 25, 2016 Dominic Smiraglia, Ph.D. Department of Cancer Genetics From Alan Wolffe, Science and Medicine, 1999 Vital Statistics Human genome contains 3 billion bp ~ 50,000
DNA Methylation and Cancer October 25, 2016 Dominic Smiraglia, Ph.D. Department of Cancer Genetics From Alan Wolffe, Science and Medicine, 1999 Vital Statistics Human genome contains 3 billion bp ~ 50,000
Hox genes. Discovered in Drosophila in 1923 by Bridges and Morgan Antennapaedia complex ANT-C Bithorax complex BX-C
 Hox genes Establish body plan during development Specify head to tail axis of animal embryos Head Hox genes, abdomen hox genes. Mutations can cause one body part to transform to another 39 transcription
Hox genes Establish body plan during development Specify head to tail axis of animal embryos Head Hox genes, abdomen hox genes. Mutations can cause one body part to transform to another 39 transcription
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development
 Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Epigenetics: The Future of Psychology & Neuroscience. Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1
 Epigenetics: The Future of Psychology & Neuroscience Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1 Nature versus Nurture Despite the belief that the Nature vs. Nurture
Epigenetics: The Future of Psychology & Neuroscience Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1 Nature versus Nurture Despite the belief that the Nature vs. Nurture
Epigenomics. Ivana de la Serna Block Health Science
 Epigenomics Ivana de la Serna Block Health Science 388 383-4111 ivana.delaserna@utoledo.edu Outline 1. Epigenetics-definition and overview 2. DNA methylation/hydroxymethylation 3. Histone modifications
Epigenomics Ivana de la Serna Block Health Science 388 383-4111 ivana.delaserna@utoledo.edu Outline 1. Epigenetics-definition and overview 2. DNA methylation/hydroxymethylation 3. Histone modifications
EPIGENOMICS PROFILING SERVICES
 EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
Molecular Cell Biology. Prof. D. Karunagaran. Department of Biotechnology. Indian Institute of Technology Madras
 Molecular Cell Biology Prof. D. Karunagaran Department of Biotechnology Indian Institute of Technology Madras Module-9 Molecular Basis of Cancer, Oncogenes and Tumor Suppressor Genes Lecture 6 Epigenetics
Molecular Cell Biology Prof. D. Karunagaran Department of Biotechnology Indian Institute of Technology Madras Module-9 Molecular Basis of Cancer, Oncogenes and Tumor Suppressor Genes Lecture 6 Epigenetics
Epigenetics and Chromatin
 Progress in Molecular and Subcellular Biology 38 Epigenetics and Chromatin Bearbeitet von Philippe Jeanteur 1. Auflage 2008. Taschenbuch. xiii, 266 S. Paperback ISBN 978 3 540 85236 0 Format (B x L): 15,5
Progress in Molecular and Subcellular Biology 38 Epigenetics and Chromatin Bearbeitet von Philippe Jeanteur 1. Auflage 2008. Taschenbuch. xiii, 266 S. Paperback ISBN 978 3 540 85236 0 Format (B x L): 15,5
Genetics and Genomics in Medicine Chapter 6. Questions & Answers
 Genetics and Genomics in Medicine Chapter 6 Multiple Choice Questions Questions & Answers Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the
Genetics and Genomics in Medicine Chapter 6 Multiple Choice Questions Questions & Answers Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the
Chromatin Modifications by Methylation and Ubiquitination: Implications in the Regulation of Gene Expression
 Annu. Rev. Biochem. 2006. 75:243 69 First published online as a Review in Advance on February 22, 2006 The Annual Review of Biochemistry is online at biochem.annualreviews.org doi: 10.1146/ annurev.biochem.75.103004.142422
Annu. Rev. Biochem. 2006. 75:243 69 First published online as a Review in Advance on February 22, 2006 The Annual Review of Biochemistry is online at biochem.annualreviews.org doi: 10.1146/ annurev.biochem.75.103004.142422
Chromatin-Based Regulation of Gene Expression
 Chromatin-Based Regulation of Gene Expression.George J. Quellhorst, Jr., PhD.Associate Director, R&D.Biological Content Development Topics to be Discussed Importance of Chromatin-Based Regulation Mechanism
Chromatin-Based Regulation of Gene Expression.George J. Quellhorst, Jr., PhD.Associate Director, R&D.Biological Content Development Topics to be Discussed Importance of Chromatin-Based Regulation Mechanism
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Oncology 520 Epigene0cs and Cancer. February 16, 2011
 Oncology 520 Epigene0cs and Cancer February 16, 2011 Outline What is epigene0cs? What are examples of epigene0c processes? What is the epigenome and how is it decoded? Are epigene0c mechanisms important
Oncology 520 Epigene0cs and Cancer February 16, 2011 Outline What is epigene0cs? What are examples of epigene0c processes? What is the epigenome and how is it decoded? Are epigene0c mechanisms important
Alpha thalassemia mental retardation X-linked. Acquired alpha-thalassemia myelodysplastic syndrome
 Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
An epigenetic approach to understanding (and predicting?) environmental effects on gene expression
 www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
I) Development: tissue differentiation and timing II) Whole Chromosome Regulation
 Epigenesis: Gene Regulation Epigenesis : Gene Regulation I) Development: tissue differentiation and timing II) Whole Chromosome Regulation (X chromosome inactivation or Lyonization) III) Regulation during
Epigenesis: Gene Regulation Epigenesis : Gene Regulation I) Development: tissue differentiation and timing II) Whole Chromosome Regulation (X chromosome inactivation or Lyonization) III) Regulation during
Chromatin Structure & Gene activity part 2
 Chromatin Structure & Gene activity part 2 Figure 13.30 Make sure that you understand it. It is good practice for identifying the purpose for various controls. Chromatin remodeling Acetylation helps to
Chromatin Structure & Gene activity part 2 Figure 13.30 Make sure that you understand it. It is good practice for identifying the purpose for various controls. Chromatin remodeling Acetylation helps to
9/3/2009 DNA i DNA n euk euk yotes Organizatio Organ izatio n of o f gen ge e n tic Locati t on: In n ucleu e s material mater in e ial
 DNA in eukaryotes Organization of genetic material in eukaryotes Location: In nucleus In mitochondria DNA in eukaryotes Nuclear DNA: Long, linear molecules; Chromatin chromosomes; 10% of DNA in genes,
DNA in eukaryotes Organization of genetic material in eukaryotes Location: In nucleus In mitochondria DNA in eukaryotes Nuclear DNA: Long, linear molecules; Chromatin chromosomes; 10% of DNA in genes,
HGD 5502 EPIGENETICS. Dr. Abhi Veerakumarasivam (2011)
 HGD 5502 EPIGENETICS Dr. Abhi Veerakumarasivam (2011) Outline History Definition Mechanisms of Epigenetics Epigenetic Processes Review History Conrad Hall Waddington (1905 1975) Developmental biologist,
HGD 5502 EPIGENETICS Dr. Abhi Veerakumarasivam (2011) Outline History Definition Mechanisms of Epigenetics Epigenetic Processes Review History Conrad Hall Waddington (1905 1975) Developmental biologist,
DNA methylation: a potential clinical biomarker for the detection of human cancers
 DNA methylation: a potential clinical biomarker for the detection of human cancers Name: Tong Samuel Supervisor: Zigui CHEN Date: 1 st December 2016 Department: Microbiology Source: cited from Jakubowski,
DNA methylation: a potential clinical biomarker for the detection of human cancers Name: Tong Samuel Supervisor: Zigui CHEN Date: 1 st December 2016 Department: Microbiology Source: cited from Jakubowski,
Epigenetics: A historical overview Dr. Robin Holliday
 Epigenetics 1 Rival hypotheses Epigenisis - The embryo is initially undifferentiated. As development proceeds, increasing levels of complexity emerge giving rise to the larval stage or to the adult organism.
Epigenetics 1 Rival hypotheses Epigenisis - The embryo is initially undifferentiated. As development proceeds, increasing levels of complexity emerge giving rise to the larval stage or to the adult organism.
Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc
 Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc P. Matthias, 16 April 2014 DNA methylation basics Acetylation Acetyltransferases/Deacetylases
Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc P. Matthias, 16 April 2014 DNA methylation basics Acetylation Acetyltransferases/Deacetylases
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Transcription and chromatin. General Transcription Factors + Promoter-specific factors + Co-activators
 Transcription and chromatin General Transcription Factors + Promoter-specific factors + Co-activators Cofactor or Coactivator 1. work with DNA specific transcription factors to make them more effective
Transcription and chromatin General Transcription Factors + Promoter-specific factors + Co-activators Cofactor or Coactivator 1. work with DNA specific transcription factors to make them more effective
The Epigenome Tools 2: ChIP-Seq and Data Analysis
 The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
An Overview of the Molecular Basis of Epigenetics
 C H A P T E R 1 An Overview of the Molecular Basis of Epigenetics J. David Sweatt, 1 Eric J. Nestler, 2 Michael J. Meaney 3 and Schahram Akbarian 4 1 McKnight Brain Institute, Department of Neurobiology,
C H A P T E R 1 An Overview of the Molecular Basis of Epigenetics J. David Sweatt, 1 Eric J. Nestler, 2 Michael J. Meaney 3 and Schahram Akbarian 4 1 McKnight Brain Institute, Department of Neurobiology,
Protein methylation CH 3
 Protein methylation CH 3 methionine S-adenosylmethionine (SAM or adomet) Methyl group of the methionine is activated by the + charge of the adjacent sulfur atom. SAM S-adenosylhomocysteine homocysteine
Protein methylation CH 3 methionine S-adenosylmethionine (SAM or adomet) Methyl group of the methionine is activated by the + charge of the adjacent sulfur atom. SAM S-adenosylhomocysteine homocysteine
Epigenetics ~from mechanism to therapy~ Literature seminar January 31, 2012 Soichi Ito (B4)
 Epigenetics ~from mechanism to therapy~ Literature seminar January 31, 2012 Soichi Ito (B4) 1 Contents Introduction Central dogma What is the Epigenetics? Topics 1. DNA methylation 2. Histone modification
Epigenetics ~from mechanism to therapy~ Literature seminar January 31, 2012 Soichi Ito (B4) 1 Contents Introduction Central dogma What is the Epigenetics? Topics 1. DNA methylation 2. Histone modification
Epigenetics & cancer. Present by : Sanaz Zebardast Under supervision : Dr. Gheibi. 31 December 2016
 Epigenetics & cancer Present by : Sanaz Zebardast Under supervision : Dr. Gheibi 31 December 2016 1 contents Introduction Epigenetic & signaling pathways Epigenetic & integral protein Epigenetic & apoptosis
Epigenetics & cancer Present by : Sanaz Zebardast Under supervision : Dr. Gheibi 31 December 2016 1 contents Introduction Epigenetic & signaling pathways Epigenetic & integral protein Epigenetic & apoptosis
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression. Chromatin Array of nuc
 Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature23267 Discussion Our findings reveal unique roles for the methylation states of histone H3K9 in RNAi-dependent and - independent heterochromatin formation. Clr4 is the sole S. pombe enzyme
doi:10.1038/nature23267 Discussion Our findings reveal unique roles for the methylation states of histone H3K9 in RNAi-dependent and - independent heterochromatin formation. Clr4 is the sole S. pombe enzyme
Epigenetic Inheritance
 (2) The role of Epigenetic Inheritance Lamarck Revisited Lamarck was incorrect in thinking that the inheritance of acquired characters is the main mechanism of evolution (Natural Selection more common)
(2) The role of Epigenetic Inheritance Lamarck Revisited Lamarck was incorrect in thinking that the inheritance of acquired characters is the main mechanism of evolution (Natural Selection more common)
'''''''''''''''''Fundamental'Biology' BI'1101' ' an'interdisciplinary'approach'to'introductory'biology' Five'Levels'of'Organiza-on' Molecular'
 '''''''''''''''''Fundamental'Biology' BI'1101' ' an'interdisciplinary'approach'to'introductory'biology' Anggraini'Barlian,' Iriawa-' Tjandra'Anggraeni' SITH4ITB' Five'Levels'of'Organiza-on' Molecular'
'''''''''''''''''Fundamental'Biology' BI'1101' ' an'interdisciplinary'approach'to'introductory'biology' Anggraini'Barlian,' Iriawa-' Tjandra'Anggraeni' SITH4ITB' Five'Levels'of'Organiza-on' Molecular'
The interplay of histone modifications writers that read
 Review Histones and Chromatin Review series The interplay of histone modifications writers that read Tianyi Zhang, Sarah Cooper *, & Neil Brockdorff Abstract Histones are subject to a vast array of posttranslational
Review Histones and Chromatin Review series The interplay of histone modifications writers that read Tianyi Zhang, Sarah Cooper *, & Neil Brockdorff Abstract Histones are subject to a vast array of posttranslational
Control of nuclear receptor function by local chromatin structure
 MINIREVIEW Control of nuclear receptor function by local chromatin structure Malgorzata Wiench, Tina B. Miranda and Gordon L. Hager Laboratory of Receptor Biology and Gene Expression, National Cancer Institute,
MINIREVIEW Control of nuclear receptor function by local chromatin structure Malgorzata Wiench, Tina B. Miranda and Gordon L. Hager Laboratory of Receptor Biology and Gene Expression, National Cancer Institute,
Epigenetics q&more 01.11
 Laurie. Knight, istockphoto.com Epigenetics 6 Bookmarks About the reading of genes in the Book of Life Prof. Dr. Manfred Jung, Julia M. Wagner, Institute for Pharmaceutical Sciences, Albert-Ludwig-University
Laurie. Knight, istockphoto.com Epigenetics 6 Bookmarks About the reading of genes in the Book of Life Prof. Dr. Manfred Jung, Julia M. Wagner, Institute for Pharmaceutical Sciences, Albert-Ludwig-University
DNA Methylation and Demethylation as Targets for Anticancer Therapy
 Biochemistry (Moscow), Vol. 70, No. 5, 2005, pp. 533-549. Translated from Biokhimiya, Vol. 70, No. 5, 2005, pp. 651-669. Original Russian Text Copyright 2005 by Szyf. DNA Methylation and Demethylation
Biochemistry (Moscow), Vol. 70, No. 5, 2005, pp. 533-549. Translated from Biokhimiya, Vol. 70, No. 5, 2005, pp. 651-669. Original Russian Text Copyright 2005 by Szyf. DNA Methylation and Demethylation
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf. Rachel Schwope EPIGEN May 24-27, 2016
 Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf Rachel Schwope EPIGEN May 24-27, 2016 What does H3K4 methylation do? Plant of interest: Vitis vinifera Culturally important
Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf Rachel Schwope EPIGEN May 24-27, 2016 What does H3K4 methylation do? Plant of interest: Vitis vinifera Culturally important
Epigenetic mechanisms behind cellular sensitivity to DNA damage
 www.cell-stress.com Review Epigenetic mechanisms behind cellular sensitivity to DNA damage Amanda K. Williamson 1, Zijing Zhu 1 and Zhi-Min Yuan 1, * 1 Department of Environmental Health, John B. Little
www.cell-stress.com Review Epigenetic mechanisms behind cellular sensitivity to DNA damage Amanda K. Williamson 1, Zijing Zhu 1 and Zhi-Min Yuan 1, * 1 Department of Environmental Health, John B. Little
A balancing act: heterochromatin protein 1a and the Polycomb group coordinate their levels to silence chromatin in Drosophila
 Cabrera et al. Epigenetics & Chromatin (2015) 8:17 DOI 10.1186/s13072-015-0010-z RESEARCH Open Access A balancing act: heterochromatin protein 1a and the Polycomb group coordinate their levels to silence
Cabrera et al. Epigenetics & Chromatin (2015) 8:17 DOI 10.1186/s13072-015-0010-z RESEARCH Open Access A balancing act: heterochromatin protein 1a and the Polycomb group coordinate their levels to silence
Organization of genetic material in eukaryotes
 Organization of genetic material in eukaryotes biologiemoleculara.usmf.md pass.: bmgu e.usmf.md 1 DNA in eukaryotes Location: In nucleus In mitochondria biologiemoleculara.usmf.md e.usmf.md pass.: bmgu
Organization of genetic material in eukaryotes biologiemoleculara.usmf.md pass.: bmgu e.usmf.md 1 DNA in eukaryotes Location: In nucleus In mitochondria biologiemoleculara.usmf.md e.usmf.md pass.: bmgu
Epigenetics: The New Science of Genetics
 CHAPTER 1 Epigenetics: The New Science of Genetics Trygve O. Tollefsbol 1,2,3,4,5 1 Department of Biology, University of Alabama at Birmingham, AL 35294 2 Center for Aging, University of Alabama at Birmingham,
CHAPTER 1 Epigenetics: The New Science of Genetics Trygve O. Tollefsbol 1,2,3,4,5 1 Department of Biology, University of Alabama at Birmingham, AL 35294 2 Center for Aging, University of Alabama at Birmingham,
Chapter 11 How Genes Are Controlled
 Chapter 11 How Genes Are Controlled PowerPoint Lectures for Biology: Concepts & Connections, Sixth Edition Campbell, Reece, Taylor, Simon, and Dickey Copyright 2009 Pearson Education, Inc. Lecture by Mary
Chapter 11 How Genes Are Controlled PowerPoint Lectures for Biology: Concepts & Connections, Sixth Edition Campbell, Reece, Taylor, Simon, and Dickey Copyright 2009 Pearson Education, Inc. Lecture by Mary
Changes in DNA methylation and histone modification during epigenetic transitions
 Oregon Health & Science University OHSU Digital Commons Scholar Archive May 2009 Changes in DNA methylation and histone modification during epigenetic transitions Jon Andrew Oyer Follow this and additional
Oregon Health & Science University OHSU Digital Commons Scholar Archive May 2009 Changes in DNA methylation and histone modification during epigenetic transitions Jon Andrew Oyer Follow this and additional
6. Heterochromatin allows for silencing of large domains of the genome and appears to utilize a combination of mechanisms to do so.
 Biochemistry 201 Biological Regulatory Mechanisms Lecturer: Geeta Narlikar February 18, 2014 Chromatin Structure and Its Regulation 2 Key Points 1. There are at least two ways in which arrays of chromatin
Biochemistry 201 Biological Regulatory Mechanisms Lecturer: Geeta Narlikar February 18, 2014 Chromatin Structure and Its Regulation 2 Key Points 1. There are at least two ways in which arrays of chromatin
Dedicated to my family
 Dedicated to my family List of Papers This thesis is based on the following papers, which are referred to in the text by their Roman numerals. I Mohammad F*, Pandey RR*, Nagano T, Chakalova L, Mondal
Dedicated to my family List of Papers This thesis is based on the following papers, which are referred to in the text by their Roman numerals. I Mohammad F*, Pandey RR*, Nagano T, Chakalova L, Mondal
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
STEM CELL GENETICS AND GENOMICS
 STEM CELL GENETICS AND GENOMICS Concise Review: Roles of Polycomb Group Proteins in Development and Disease: A Stem Cell Perspective VINAGOLU K. RAJASEKHAR, a MARTIN BEGEMANN b a Memorial Sloan-Kettering
STEM CELL GENETICS AND GENOMICS Concise Review: Roles of Polycomb Group Proteins in Development and Disease: A Stem Cell Perspective VINAGOLU K. RAJASEKHAR, a MARTIN BEGEMANN b a Memorial Sloan-Kettering
CHAIRE ÉPIGÉNÉTIQUE ET MÉMOIRE CELLULAIRE. Année : Reprogrammations développementales, induites et pathologiques
 CHAIRE ÉPIGÉNÉTIQUE ET MÉMOIRE CELLULAIRE Année 2013-2014 : Reprogrammations développementales, induites et pathologiques Cours II Etapes de la reprogrammation au cours du développement chez les mammifères
CHAIRE ÉPIGÉNÉTIQUE ET MÉMOIRE CELLULAIRE Année 2013-2014 : Reprogrammations développementales, induites et pathologiques Cours II Etapes de la reprogrammation au cours du développement chez les mammifères
Long noncoding RNAs in DNA methylation: new players stepping into the old game
 DOI 10.1186/s13578-016-0109-3 Cell & Bioscience REVIEW Long noncoding RNAs in DNA methylation: new players stepping into the old game Yu Zhao 1, Hao Sun 1,2* and Huating Wang 1,3* Open Access Abstract
DOI 10.1186/s13578-016-0109-3 Cell & Bioscience REVIEW Long noncoding RNAs in DNA methylation: new players stepping into the old game Yu Zhao 1, Hao Sun 1,2* and Huating Wang 1,3* Open Access Abstract
Session 2: Biomarkers of epigenetic changes and their applicability to genetic toxicology
 Session 2: Biomarkers of epigenetic changes and their applicability to genetic toxicology Bhaskar Gollapudi, Ph.D The Dow Chemical Company Workshop: Genetic Toxicology: Opportunities to Integrate New Approaches
Session 2: Biomarkers of epigenetic changes and their applicability to genetic toxicology Bhaskar Gollapudi, Ph.D The Dow Chemical Company Workshop: Genetic Toxicology: Opportunities to Integrate New Approaches
Constitutive heterochromatin formation and transcription in mammals
 Saksouk et al. Epigenetics & Chromatin 2015, 8:3 REVIEW Constitutive heterochromatin formation and transcription in mammals Nehmé Saksouk, Elisabeth Simboeck and Jérôme Déjardin * Open Access Abstract
Saksouk et al. Epigenetics & Chromatin 2015, 8:3 REVIEW Constitutive heterochromatin formation and transcription in mammals Nehmé Saksouk, Elisabeth Simboeck and Jérôme Déjardin * Open Access Abstract
Xist function: bridging chromatin and stem cells
 Review TRENDS in Genetics Vol.23 No.9 Xist function: bridging chromatin and stem cells Anton Wutz Research Institute of Molecular Pathology, Dr. Bohr-Gasse 7, 1030 Vienna, Austria In mammals, dosage compensation
Review TRENDS in Genetics Vol.23 No.9 Xist function: bridging chromatin and stem cells Anton Wutz Research Institute of Molecular Pathology, Dr. Bohr-Gasse 7, 1030 Vienna, Austria In mammals, dosage compensation
Structure and function of active chromatin and DNase I hypersensitive sites
 MINIREVIEW Structure and function of active chromatin and DNase I hypersensitive sites Peter N. Cockerill Experimental Haematology, Leeds Institute of Molecular Medicine, University of Leeds, UK Keywords
MINIREVIEW Structure and function of active chromatin and DNase I hypersensitive sites Peter N. Cockerill Experimental Haematology, Leeds Institute of Molecular Medicine, University of Leeds, UK Keywords
Fragile X Syndrome. Genetics, Epigenetics & the Role of Unprogrammed Events in the expression of a Phenotype
 Fragile X Syndrome Genetics, Epigenetics & the Role of Unprogrammed Events in the expression of a Phenotype A loss of function of the FMR-1 gene results in severe learning problems, intellectual disability
Fragile X Syndrome Genetics, Epigenetics & the Role of Unprogrammed Events in the expression of a Phenotype A loss of function of the FMR-1 gene results in severe learning problems, intellectual disability
The genetics of heterochromatin. in metazoa. mutations by means of X-ray irradiation" "for the discovery of the production of
 The genetics of heterochromatin in metazoa 1 Hermann Joseph Muller 1946 Nobel Prize in Medicine: "for the discovery of the production of mutations by means of X-ray irradiation" 3 4 The true meaning of
The genetics of heterochromatin in metazoa 1 Hermann Joseph Muller 1946 Nobel Prize in Medicine: "for the discovery of the production of mutations by means of X-ray irradiation" 3 4 The true meaning of
Imprinting. Joyce Ohm Cancer Genetics and Genomics CGP-L2-319 x8821
 Imprinting Joyce Ohm Cancer Genetics and Genomics CGP-L2-319 x8821 Learning Objectives 1. To understand the basic concepts of genomic imprinting Genomic imprinting is an epigenetic phenomenon that causes
Imprinting Joyce Ohm Cancer Genetics and Genomics CGP-L2-319 x8821 Learning Objectives 1. To understand the basic concepts of genomic imprinting Genomic imprinting is an epigenetic phenomenon that causes
Expert Intelligence for Better Decisions Epigenetics:
 Expert Intelligence for Better Decisions Epigenetics: Emerging Targets, Available Technologies, Expert Interviews, and an Epigenetic Community Perspective Using This Document Insight Pharma Reports are
Expert Intelligence for Better Decisions Epigenetics: Emerging Targets, Available Technologies, Expert Interviews, and an Epigenetic Community Perspective Using This Document Insight Pharma Reports are
Epigenetics: Basic Principals and role in health and disease
 Epigenetics: Basic Principals and role in health and disease Cambridge Masterclass Workshop on Epigenetics in GI Health and Disease 3 rd September 2013 Matt Zilbauer Overview Basic principals of Epigenetics
Epigenetics: Basic Principals and role in health and disease Cambridge Masterclass Workshop on Epigenetics in GI Health and Disease 3 rd September 2013 Matt Zilbauer Overview Basic principals of Epigenetics
Histone octamer assembly
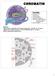 CHROMATIN Chromatin Histone structure Nucleosome Chromatin fiber Histone acetylation Chromatin transcription Regulation of gene expression: -Cell cycle control (+/-) -Cell differentiation Genome integrity
CHROMATIN Chromatin Histone structure Nucleosome Chromatin fiber Histone acetylation Chromatin transcription Regulation of gene expression: -Cell cycle control (+/-) -Cell differentiation Genome integrity
FOLLICULAR LYMPHOMA- ILLUMINA METHYLATION. Jude Fitzgibbon
 FOLLICULAR LYMPHOMA- ILLUMINA METHYLATION Jude Fitzgibbon j.fitzgibbon@qmul.ac.uk Molecular Predictors of Clinical outcome Responders Alive Non-responders Dead TUMOUR GENETICS Targeted therapy, Prognostic
FOLLICULAR LYMPHOMA- ILLUMINA METHYLATION Jude Fitzgibbon j.fitzgibbon@qmul.ac.uk Molecular Predictors of Clinical outcome Responders Alive Non-responders Dead TUMOUR GENETICS Targeted therapy, Prognostic
H19 Chromatin Insulator in Development and DiseaseEpigenetic Regulation of the
 Comprehensive Summaries of Uppsala Dissertations from the Faculty of Science and Technology 825 H19 Chromatin Insulator in Development and DiseaseEpigenetic Regulation of the BY CLAES HOLMGREN ACTA UNIVERSITATIS
Comprehensive Summaries of Uppsala Dissertations from the Faculty of Science and Technology 825 H19 Chromatin Insulator in Development and DiseaseEpigenetic Regulation of the BY CLAES HOLMGREN ACTA UNIVERSITATIS
5. Heterochromatin allows for silencing of large domains of the genome and utilizes a combination of mechanisms to do so.
 Biochemistry 201 Biological Regulatory Mechanisms Lecturer: Geeta Narlikar February 1, 2016 Chromatin Structure and Its Regulation 2 Key Points 1. Histones provide a versatile regulatory platform through
Biochemistry 201 Biological Regulatory Mechanisms Lecturer: Geeta Narlikar February 1, 2016 Chromatin Structure and Its Regulation 2 Key Points 1. Histones provide a versatile regulatory platform through
AGCGGCTGGTGTATGAGG AGCTCGAGCTGCAGGACT AAAGGTGCACTGGTAGCC AGGGCACAAGTGCATGGC CCCGCTGTGCAGAAGTAG
 Supplementary Table S1. Primer sequence information Gene Primer sequence 1 st round PCR 2 nd round PCR Accession# Dnmt3a TCAAGGAGATCATTGATG TGTTCCCACACATGAGCA AGCGGCTGGTGTATGAGG AGCAGATGGTGCAATAGGAC NM_007872.4
Supplementary Table S1. Primer sequence information Gene Primer sequence 1 st round PCR 2 nd round PCR Accession# Dnmt3a TCAAGGAGATCATTGATG TGTTCCCACACATGAGCA AGCGGCTGGTGTATGAGG AGCAGATGGTGCAATAGGAC NM_007872.4
A Genetic Program for Embryonic Development
 Concept 18.4: A program of differential gene expression leads to the different cell types in a multicellular organism During embryonic development, a fertilized egg gives rise to many different cell types
Concept 18.4: A program of differential gene expression leads to the different cell types in a multicellular organism During embryonic development, a fertilized egg gives rise to many different cell types
Epigenetics and Toxicology
 Epigenetics and Toxicology Aline.deconti@fda.hhs.gov Division of Biochemical Toxicology National Center for Toxicology Research U.S.-Food and Drug Administration The views expressed in this presentation
Epigenetics and Toxicology Aline.deconti@fda.hhs.gov Division of Biochemical Toxicology National Center for Toxicology Research U.S.-Food and Drug Administration The views expressed in this presentation
HDAC1 Inhibitor Screening Assay Kit
 HDAC1 Inhibitor Screening Assay Kit Catalog Number KA1320 96 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
HDAC1 Inhibitor Screening Assay Kit Catalog Number KA1320 96 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
OVERVIEW OF EPIGENETICS
 OVERVIEW OF EIENETICS Date: * Time: 9:00 am - 9:50 am * Room: Berryhill 103 Lecturer: Terry Magnuson 4312 MBRB trm4@med.unc.edu 843-6475 *lease consult the online schedule for this course for the definitive
OVERVIEW OF EIENETICS Date: * Time: 9:00 am - 9:50 am * Room: Berryhill 103 Lecturer: Terry Magnuson 4312 MBRB trm4@med.unc.edu 843-6475 *lease consult the online schedule for this course for the definitive
Transgenerational epigenetics in the germline cycle of Caenorhabditis elegans
 Transgenerational epigenetics in the germline cycle of Caenorhabditis elegans William G Kelly, Emory University Journal Title: Epigenetics and Chromatin Volume: Volume 7, Number 6 Publisher: BioMed Central
Transgenerational epigenetics in the germline cycle of Caenorhabditis elegans William G Kelly, Emory University Journal Title: Epigenetics and Chromatin Volume: Volume 7, Number 6 Publisher: BioMed Central
Session 6: Integration of epigenetic data. Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016
 Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE
 Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE If looking for the book Systems Analysis of Chromatin-Related Protein Complexes in Cancer in pdf format, then you have come
Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE If looking for the book Systems Analysis of Chromatin-Related Protein Complexes in Cancer in pdf format, then you have come
Host cell activation
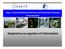 Dept. of Internal Medicine/Infectious and Respiratory Diseases Stefan Hippenstiel Epigenetics as regulator of inflammation Host cell activation LPS TLR NOD2 MDP TRAF IKK NF-κB IL-x, TNFα,... Chromatin
Dept. of Internal Medicine/Infectious and Respiratory Diseases Stefan Hippenstiel Epigenetics as regulator of inflammation Host cell activation LPS TLR NOD2 MDP TRAF IKK NF-κB IL-x, TNFα,... Chromatin
Supplemental Figure 1: Asymmetric chromatin maturation leads to epigenetic asymmetries on sister chromatids.
 Supplemental Material: Annu. Rev. Cell Dev. Biol. 2017. 33:291 318 https://doi.org/10.1146/annurev-cellbio-100616-060447 The Inherent Asymmetry of DNA Replication Snedeker, Wooten, and Chen Supplemental
Supplemental Material: Annu. Rev. Cell Dev. Biol. 2017. 33:291 318 https://doi.org/10.1146/annurev-cellbio-100616-060447 The Inherent Asymmetry of DNA Replication Snedeker, Wooten, and Chen Supplemental
Epigenetic regulation of germ cell and early embryonic development by Polycomb group proteins. Inauguraldissertation
 Epigenetic regulation of germ cell and early embryonic development by Polycomb group proteins Inauguraldissertation zur Erlangung der Würde eines Doktors der Philosophie vorgelegt der Philosophisch Naturwissenschaftlichen
Epigenetic regulation of germ cell and early embryonic development by Polycomb group proteins Inauguraldissertation zur Erlangung der Würde eines Doktors der Philosophie vorgelegt der Philosophisch Naturwissenschaftlichen
Dissecting gene regulation network in human early embryos. at single-cell and single-base resolution
 Dissecting gene regulation network in human early embryos at single-cell and single-base resolution Fuchou Tang BIOPIC, College of Life Sciences Peking University 07/10/2015 Cockburn and Rossant, 2010
Dissecting gene regulation network in human early embryos at single-cell and single-base resolution Fuchou Tang BIOPIC, College of Life Sciences Peking University 07/10/2015 Cockburn and Rossant, 2010
Nuclear reprogramming of sperm and somatic nuclei in eggs and oocytes
 Reprod Med Biol (2013) 12:133 149 DOI 10.1007/s12522-013-0155-z REVIEW ARTICLE Nuclear reprogramming of sperm and somatic nuclei in eggs and oocytes Marta Teperek Kei Miyamoto Received: 27 March 2013 /
Reprod Med Biol (2013) 12:133 149 DOI 10.1007/s12522-013-0155-z REVIEW ARTICLE Nuclear reprogramming of sperm and somatic nuclei in eggs and oocytes Marta Teperek Kei Miyamoto Received: 27 March 2013 /
