Supplementary Information
|
|
|
- Adrian Baker
- 5 years ago
- Views:
Transcription
1 Supplementary Information Using the pimeloyl-coa synthetase adenylation fold to synthesise fatty acid thioesters Menglu Wang 1a ; Lucile Moynié 2a, Peter J. Harrison 1, Van Kelly 1, Andrew Piper 1, James H. Naismith 2, 3* and Dominic J. Campopiano 1* 1 EastChem School of Chemistry, David Brewster Road, University of Edinburgh, Edinburgh, EH9 3FJ. 2 Biomedical Sciences Research complex, University of St. Andrews, St Andrews, KY16 9ST 3 State Key Laboratory of Biotherapy and Cancer Center, West China Hospital of Sichuan University and Collaborative Innovation Center of Biotherapy and Cancer, Chengdu , China 1
2 Supplementary Results (a) MSYYHHHHHH DYDIPTTENL YFQGAMEEET FYSVRMRASM NGSHEDGGKH 27 ISGGERLIPF HEMKHTVNAL LEKGLSHSRG KPDFMQIQFE EVHESIKTIQ 77 PLPVHTNEVS CPEEGQKLAR LLLEKEGVSR DVIEKAYEQI PEWSDVRGAV 127 LFDIHTGKRM DQTKEKGVRV SRMDWPDANF EKWALHSHVP AHSRIKEALA 177 LASKVSRHPA VVAELCWSDD PDYITGYVAG KKMGYQRITA MKEYGTEEGC 227 RVFFIDGSND VNTYIHDLEK QPILIEWEED HDS 259 Supplementary Figure 1a. Amino acid sequence of B. subtilis 6His BioW. The sequence of the full length clone and recombinant BioW after TEV cleavage to remove the His tag is shown. The BioW numbering begins at Ala1 and ends with Ser259. Residues in the active site are in bold (Tyr199, Tyr211, Arg213 and Arg227) and the His tag and TEV cleavage site in blue and red respectively. (b) Supplementary Figure 1b. 15% SDS-PAGE gel showing the purity of purified BioW enzyme. M: LMW Marker, Lane 1: E. coli cell pellet, Lane 2: Cell lysate, Lane 3: Ni resin flow through, Lane 4: Ni resin wash, Lane 5: Ni resin elution, Lane 6: Pre-TEV protease cleavage, Lane 7: Second Ni resin flow through, Lane 8: Second Ni resin elution, Lane 9-12: Superdex 200 fractions A band at ~30 kda corresponds to the BioW molecular weight. 2
3 (c) Absorbance (280 nm) (mau) Retention Volume (ml) BioW V e = 73.4 ml Supplementary Figure 1c. Chromatogram of purified BioW. The purified BioW was applied to a calibrated Superdex S200HR gel filtration column (A280nm). BioW elutes at 73.4 ml which corresponds to a homodimer. 3
4 (a) (b) 4
5 (c) (d) 5
6 (e) Supplementary Figure. 2. Activity of recombinant B. subtilis BioW. (a) ATP (retention time 3.1 mins), ADP (4.0 mins) and AMP (4.2 mins) standards (Sigma) co-injected onto the C18 RP HPLC column. (b) Co-injection of ATP with Coenzyme A from Sigma (retention time 13.3 mins). (c) Synthetic pimeloyl-coa (retention time mins, see Methods). (d) ESI FT-ICR MS analysis of synthetic pimeloyl-coa (m/z range = ). Inset shows the ion with mass = Da consistent with [M+H] +. Predicted mass = Da (C 28 H 46 N 7 O 19 P 3 S). The red circles represent the isotopic distribution predicted from the chemical formula. (e) BioW-catalysed pimeloyl-coa synthesis. The incubation described in the Methods was analysed by HPLC and we observed peaks corresponding to pimeloyl-coa at 16.2 mins and AMP (4.2 mins). The inset show the ESI FT-ICR MS analysis of the peak at 16.2 mins which agrees well with the synthetic standard shown in (c). 6
7 Supplementary Figure 3. Basis of the coupled BioW activity assay. Pimelic is converted to pimeloyl-coa by BioW and the pyrophosphate (PPi) released is hydrolysed to two molecules of phosphate (Pi) by pyrophosphatase (PPase). Purine nucleoside phosphorylase (PNP) catalyses the reaction between the released P i and MESG to give the product 7-methyl- 6-thioguanine (which absorbs at 360 nm, ε 360 = 11,000 M -1 cm -1 ) and ribose phosphate. 7
8 (a) (b) (c) Supplementary Figure 4. Kinetic analysis of WT BioW using the coupled assay. Substrates (a) pimelic acid and (b) ATP. (c) Determination of the kinetic data for CoASH 8
9 used the 230nm UV assay. Data was fit using Michaelis-Menten model on GraphPad. Data represent mean values ± s.d. from multiple experiments. 9
10 10
11 11
12 h) 12
13 Supplementary Figure 5. Structures of BioW. (a) Binding sites of pimeloyl-adenylate and PPi. For more clarity, only the polar interactions have been represented and the figure has been split in two views, one for the AMP moiety (left) and the other for the pimeloyl group (right). Carbon atoms of the pimeloyl-adenylate are in cyan. Hydrogen bonds are shown as black dashes lines. Oxygen atoms are in red, nitrogen in blue, phosphate in orange and Mg 2+ ions are shown as green spheres. (b) Binding sites of AMP-PNP and pimelic acid. The colour scheme is the same than a) but the carbon atoms of AMP-PNP and pimelic acid are in yellow. (c) Fo-Fc electron density omit map at 3σ around AMP-PNP and pimelic acid. (d) Ligplot diagram of pimeloyl-adenylate/ppi. Covalent bonds of the ligands and protein residues are in purple and brown sticks, respectively. Hydrogen bonds are represented by green dashed line and hydrophobic contacts are shown as red semi-circles with radiating spokes. Figure prepared with Ligplot 1. (e) Ligplot diagram of AMP-PNP/pimelic acid. (f) Ligplot diagram of the adenosine 3', 5'-diphosphate group of CoA bound to BioW. (g) Fo-Fc electron density omit map at 3σ around the ordered portion of CoA. (h) Two views of the electrostatic surface of the BioW with the ordered portion of CoA shown as spheres. There are two possible routes to the active site denoted with dashed circle. Positively charged regions are shown in blue and negatively regions are in red. Oxygen atoms are in red, nitrogen in blue, phosphate in pink. 13
14 Supplementary Figure 6. Sequence alignment of bacterial BioW. The sequences used are from Bacillus subtilis (UNIPROT code: P53559, BioW_BACSU), Aquifex aeolicus (O67575, BioW_AQUAE), Lysinibacillus sphaericus (P22822, (IOW_LYSSH), and Staphylococcus aureus (P67549, BioW_STAAM). The residues targetted for engineering the fatty acid specificity (Y199, R211, R213, R227) are marked with blue triangles. Figure produced using ESPript
15 Supplementary Figure 7. The BioW Y211F mutant displays broadened fatty acid specificity. The turnover (k cat, s -1 ) of each BioW enzyme, WT and mutants Y199F, Y211F and R213A (3.0 µm) in the presence of different carboxylic acids (1.5 mm), ATP (1.0 mm) and CoASH (1.0 mm). Data represent mean values ± s.d. from multiple experiments. 15
16 Supplementary Figure 8. Kinetic analysis of the BioW R11A and R13A mutants with pimelic acid as a substrate. The data was obtained using the PPi assay and the WT activity is included for comparison. Data represent mean values ± s.d. from multiple experiments. 16
17 Supplementary Figure 9. Kinetic analysis of BioW Y211F. We used BioW Y211F (2.5 µm) and different concentrations of heptanoic acid (top panel) and ATP (lower panel) as substrates. The data were obtained using the PPi assay and the kinetic parameters for each substrate were determined. The data was fitted using the Michaelis-Menten model on GraphPad; heptanoic (k cat = s -1 ± , K m = ± 79.7) and ATP (k cat = ± s -1, K m = ± 81.5 µm). Data represent mean values ± s.d. from multiple experiments. 17
18 (a) Pimeloyl-CoA * WT Time (mins) * Y211F Base Peak Base Peak Intensity Time (mins) m/z (b) 7-Octenoyl- WT Base Peak Intensity Base Peak Intensity Time * Y211F Time (mins) m/z 18
19 (c) 7-Phenylheptanoyl-CoA WT SCoA Base Peak O 7-Phenylheptanoyl-CoA Time (mins) Y211F Base Peak * m/z Time (mins) (d) 6-Methylheptanoyl-CoA WT Base Peak Time (mins) * Y211F Base Peak Time (mins) m/z 19
20 (e) 7-Bromoheptanoyl-CoA WT Base Peak Intensity Base Peak Intensity Time (mins) * Y211F (f) 7-Aminoheptanoyl-CoA Time (mins) m/z WT Base Peak Intensity Time (mins) Y211F Expected [M+H] not observed Base Peak Time (mins) Supplementary Figure 10. Mass spectrometry analysis of acyl-coa products. Base peak intensity chromatograms are presented for LCMS analyses of reactions using six substrates; (a) pimelic acid, (b) 7-octenoic acid, (c) 7-phenyl-heptanoic acid, (d) 6-methylheptanoic acid, (e) 7-bromo-heptanoic acid and (f) 7-aminoheptanoic acid for both BioW WT and Y211F mutant enzymes. Chromatographic peaks corresponding to the expected acyl-coa thioester 20
21 product masses are annotated (*) and the mass spectrum of the [M+H] + ion is presented, if observed. The observed monoisotopic mass annotation was extracted from centroided data and all masses were observed within 3ppm from the expected mass. The acyl-coa thioester product was produced by the BioW Y211F mutant for all substrates tested, apart from F) 7- aminoheptanoic acid. 21
22 Supplementary Table 1: Kinetic parameters of BioW. The pimelic acid and ATP data were determined using the PPi coupled assay and the CoASH data using thioester bond formation at 230nm assay (*). Data represent mean values ± s.d. from multiple experiments. Pimelic acid ATP CoASH* K m (µm) 70.5 ± ± ± 25.4 k cat (s -1 ) 0.48 ± ± ± 0.05 k cat / K m (M -1 s -1 ) 8, , ,
23 Supplementary Table 2. Chain length specificity of WT BioW. Turnover values (k cat ) of the dicarboxylic and monocarboxylic acid substrates of different chain lengths. The values were determined using the PPi assay (see Methods). Common name Carbon Chain Length Turnover (s -1 ) Glutaric acid Adipic acid Pimelic acid Suberic acid Azelaic acid Heptanoic acid Octanoic acid
24 Supplementary Table 3: Crystallographic data and refinement statistics K 2 PtCl 4 Pimeloyladenylate/PPi complex (5fm0) AMP- PNP/pimelic acid complex (5flg) Pimeloyladenylate/PPi complex (5fll) Pimelic acid/coash complex (5g1f) Data collection Space group P P P P Cell dimensions a, b, c (Å) , 77.90, , 78.11, , 78.36, α, β, γ ( ) 90.0, 90.0, , 90.0, , 90.0, , 90.0, 90.0 Resolution (Å) ( )* ( ) ( ) ( ) R sym or R merge (0.681) (0.552) (0.419) (0.757) I / σi 23.3 (3.3) 17.3 (2.7) 21.4 (3.36) 18.8 (2.0) Completeness (%) 99.9 (99.9) 99.2 (92.2) 99.5 (94.2) 99.8 (99.7) Redundancy 15.2 (10.5) 6.4 (4.1) 4.2 (2.9) 6.4 (5.1) Refinement Resolution (Å) No. reflections R work / R free 0.200/ / / / No. atoms Protein Ligand/ion Water B-factors Protein Ligand/ion Water R.m.s deviations Bond lengths (Å) Bond angles ( ) Each dataset was collected from a single crystal. *Values in parentheses are for highest-resolution shell. 24
25 Supplementary References 1 Wallace, A. C., Laskowski, R. A. & Thornton, J. M. LIGPLOT: a program to generate schematic diagrams of protein-ligand interactions. Protein engineering 8, (1995). 2 Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic acids research 42, W , doi: /nar/gku316 (2014). 25
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
SUPPLEMENTARY INFORMATION FOR. (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation. by COX-2
 SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
Supporting Information
 Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Using the pimeloyl-coa synthetase adenylation fold to synthesise fatty acid thioesters
 Using the pimeloyl-coa synthetase adenylation fold to synthesise fatty acid thioesters Menglu Wang 1a ; Lucile Moynié 2a, Peter J. Harrison 1, Van Kelly 1, Andrew Piper 1, James H. Naismith 2, 3* and Dominic
Using the pimeloyl-coa synthetase adenylation fold to synthesise fatty acid thioesters Menglu Wang 1a ; Lucile Moynié 2a, Peter J. Harrison 1, Van Kelly 1, Andrew Piper 1, James H. Naismith 2, 3* and Dominic
Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants
 Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants Species UDP-2,3- diacylglucosamine hydrolase specific activity (nmol min -1 mg -1 ) Fold vectorcontrol specific
Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants Species UDP-2,3- diacylglucosamine hydrolase specific activity (nmol min -1 mg -1 ) Fold vectorcontrol specific
FCC2 5CY7 FCC1 5CY8. Actinonin 5CVQ
 Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Supplementary Information Janssen et al.
 Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Adenosine triphosphate (ATP)
 Adenosine triphosphate (ATP) 1 High energy bonds ATP adenosine triphosphate N NH 2 N -O O P O O P O- O- O O P O- O CH 2 H O H N N adenine phosphoanhydride bonds (~) H OH ribose H OH Phosphoanhydride bonds
Adenosine triphosphate (ATP) 1 High energy bonds ATP adenosine triphosphate N NH 2 N -O O P O O P O- O- O O P O- O CH 2 H O H N N adenine phosphoanhydride bonds (~) H OH ribose H OH Phosphoanhydride bonds
Fatty acid breakdown
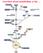 Fatty acids contain a long hydrocarbon chain and a terminal carboxylate group. Most contain between 14 and 24 carbon atoms. The chains may be saturated or contain double bonds. The complete oxidation of
Fatty acids contain a long hydrocarbon chain and a terminal carboxylate group. Most contain between 14 and 24 carbon atoms. The chains may be saturated or contain double bonds. The complete oxidation of
PAPER No. : 16 Bioorganic and biophysical chemistry MODULE No. : 25 Coenzyme-I Coenzyme A, TPP, B12 and biotin
 Subject Paper No and Title Module No and Title Module Tag 16, Bio organic and Bio physical chemistry 25, Coenzyme-I : Coenzyme A, TPP, B12 and CHE_P16_M25 TABLE OF CONTENTS 1. Learning Outcomes 2. Introduction
Subject Paper No and Title Module No and Title Module Tag 16, Bio organic and Bio physical chemistry 25, Coenzyme-I : Coenzyme A, TPP, B12 and CHE_P16_M25 TABLE OF CONTENTS 1. Learning Outcomes 2. Introduction
MBB 694:407, 115:511. Please use BLOCK CAPITAL letters like this --- A, B, C, D, E. Not lowercase!
 MBB 694:407, 115:511 First Test Severinov/Deis Tue. Sep. 30, 2003 Name Index number (not SSN) Row Letter Seat Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions
MBB 694:407, 115:511 First Test Severinov/Deis Tue. Sep. 30, 2003 Name Index number (not SSN) Row Letter Seat Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Tivadar Orban, Beata Jastrzebska, Sayan Gupta, Benlian Wang, Masaru Miyagi, Mark R. Chance, and Krzysztof Palczewski
 Structure, Volume Supplemental Information Conformational Dynamics of Activation for the Pentameric Complex of Dimeric G Protein-Coupled Receptor and Heterotrimeric G Protein Tivadar Orban, Beata Jastrzebska,
Structure, Volume Supplemental Information Conformational Dynamics of Activation for the Pentameric Complex of Dimeric G Protein-Coupled Receptor and Heterotrimeric G Protein Tivadar Orban, Beata Jastrzebska,
Supporting Information. Lysine Propionylation to Boost Proteome Sequence. Coverage and Enable a Silent SILAC Strategy for
 Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex
 John Clizer & Greg Ralph Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex The turkey ovomucoid third domain (OMTKY3) is considered to be one of the most studied protein inhibitors. 1 Ovomucin
John Clizer & Greg Ralph Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex The turkey ovomucoid third domain (OMTKY3) is considered to be one of the most studied protein inhibitors. 1 Ovomucin
SUPPLEMENTARY INFORMATION
 Supplementary Table 1. Cell sphingolipids and S1P bound to endogenous TRAF2. Sphingolipid Cell pmol/mg TRAF2 immunoprecipitate pmol/mg Sphingomyelin 4200 ± 250 Not detected Monohexosylceramide 311 ± 18
Supplementary Table 1. Cell sphingolipids and S1P bound to endogenous TRAF2. Sphingolipid Cell pmol/mg TRAF2 immunoprecipitate pmol/mg Sphingomyelin 4200 ± 250 Not detected Monohexosylceramide 311 ± 18
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.
 Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Structural analysis of fungus-derived FAD glucose dehydrogenase
 Structural analysis of fungus-derived FAD glucose dehydrogenase Hiromi Yoshida 1, Genki Sakai 2, Kazushige Mori 3, Katsuhiro Kojima 3, Shigehiro Kamitori 1, and Koji Sode 2,3,* 1 Life Science Research
Structural analysis of fungus-derived FAD glucose dehydrogenase Hiromi Yoshida 1, Genki Sakai 2, Kazushige Mori 3, Katsuhiro Kojima 3, Shigehiro Kamitori 1, and Koji Sode 2,3,* 1 Life Science Research
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Manja Henze, Dorothee Merker and Lothar Elling. 1. Characteristics of the Recombinant β-glycosidase from Pyrococcus
 S1 of S17 Supplementary Materials: Microwave-Assisted Synthesis of Glycoconjugates by Transgalactosylation with Recombinant Thermostable β-glycosidase from Pyrococcus Manja Henze, Dorothee Merker and Lothar
S1 of S17 Supplementary Materials: Microwave-Assisted Synthesis of Glycoconjugates by Transgalactosylation with Recombinant Thermostable β-glycosidase from Pyrococcus Manja Henze, Dorothee Merker and Lothar
Figure 1. A ribbon diagram of the aldolase (A) and a close up of the active site (B) including the bound substrate.
 Problem Set 4 (C-C bond formation, phosphoryl transfer reactions and the role of ATP) 1. Chemists can use the same strategies as nature to make new carbon-carbon bonds stereospecifically using enzymes
Problem Set 4 (C-C bond formation, phosphoryl transfer reactions and the role of ATP) 1. Chemists can use the same strategies as nature to make new carbon-carbon bonds stereospecifically using enzymes
PAPER No. : 16, Bioorganic and biophysical chemistry MODULE No. : 22, Mechanism of enzyme catalyst reaction (I) Chymotrypsin
 Subject Paper No and Title 16 Bio-organic and Biophysical Module No and Title 22 Mechanism of Enzyme Catalyzed reactions I Module Tag CHE_P16_M22 Chymotrypsin TABLE OF CONTENTS 1. Learning outcomes 2.
Subject Paper No and Title 16 Bio-organic and Biophysical Module No and Title 22 Mechanism of Enzyme Catalyzed reactions I Module Tag CHE_P16_M22 Chymotrypsin TABLE OF CONTENTS 1. Learning outcomes 2.
Tala Saleh. Ahmad Attari. Mamoun Ahram
 23 Tala Saleh Ahmad Attari Minna Mushtaha Mamoun Ahram In the previous lecture, we discussed the mechanisms of regulating enzymes through inhibitors. Now, we will start this lecture by discussing regulation
23 Tala Saleh Ahmad Attari Minna Mushtaha Mamoun Ahram In the previous lecture, we discussed the mechanisms of regulating enzymes through inhibitors. Now, we will start this lecture by discussing regulation
Minutes Figure S1. HPLC separation of nucleosides from LC/ESI-MS analysis of a total enzymatic Trp
 100 A % Relative Abundance m m Ø acp 5 m Øm 5 m m 7 m s * 6 ia m * 0 5 10 15 0 5 0 5 40 Minutes Figure S1. HPL separation of nucleosides from L/ESI-MS analysis of a total enzymatic digest of mt trna. V
100 A % Relative Abundance m m Ø acp 5 m Øm 5 m m 7 m s * 6 ia m * 0 5 10 15 0 5 0 5 40 Minutes Figure S1. HPL separation of nucleosides from L/ESI-MS analysis of a total enzymatic digest of mt trna. V
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular
 Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein
 Supplementary Methods Thermal shift assays Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein unfolding was examined by monitoring the fluorescence of ANS (1-anilinonaphthalene-8-
Supplementary Methods Thermal shift assays Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein unfolding was examined by monitoring the fluorescence of ANS (1-anilinonaphthalene-8-
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
Enzymes Part III: regulation II. Dr. Mamoun Ahram Summer, 2017
 Enzymes Part III: regulation II Dr. Mamoun Ahram Summer, 2017 Advantage This is a major mechanism for rapid and transient regulation of enzyme activity. A most common mechanism is enzyme phosphorylation
Enzymes Part III: regulation II Dr. Mamoun Ahram Summer, 2017 Advantage This is a major mechanism for rapid and transient regulation of enzyme activity. A most common mechanism is enzyme phosphorylation
Biological Mass Spectrometry. April 30, 2014
 Biological Mass Spectrometry April 30, 2014 Mass Spectrometry Has become the method of choice for precise protein and nucleic acid mass determination in a very wide mass range peptide and nucleotide sequencing
Biological Mass Spectrometry April 30, 2014 Mass Spectrometry Has become the method of choice for precise protein and nucleic acid mass determination in a very wide mass range peptide and nucleotide sequencing
Student Number: To form the polar phase when adsorption chromatography was used.
 Name: Student Number: April 14, 2001, 1:30 AM - 4:30 PM Page 1 (of 4) Biochemistry II Lab Section Final Examination Examiner: Dr. A. Scoot 1. Answer ALL questions in the space provided.. 2. The last page
Name: Student Number: April 14, 2001, 1:30 AM - 4:30 PM Page 1 (of 4) Biochemistry II Lab Section Final Examination Examiner: Dr. A. Scoot 1. Answer ALL questions in the space provided.. 2. The last page
Supplementary Information
 Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
The mechanism of patellamide macrocyclization revealed by study of the Prochloron sp PatG macrocyclase domain
 The mechanism of patellamide macrocyclization revealed by study of the Prochloron sp PatG macrocyclase domain Jesko Koehnke 1,2, Andrew Bent 1,2, Wael E. Houssen 2,3,4, David Zollman 1, Falk Morawitz 1,
The mechanism of patellamide macrocyclization revealed by study of the Prochloron sp PatG macrocyclase domain Jesko Koehnke 1,2, Andrew Bent 1,2, Wael E. Houssen 2,3,4, David Zollman 1, Falk Morawitz 1,
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB
 Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Chymotrypsin Lecture. Aims: to understand (1) the catalytic strategies used by enzymes and (2) the mechanism of chymotrypsin
 Chymotrypsin Lecture Aims: to understand (1) the catalytic strategies used by enzymes and (2) the mechanism of chymotrypsin What s so great about enzymes? They accomplish large rate accelerations (10 10-10
Chymotrypsin Lecture Aims: to understand (1) the catalytic strategies used by enzymes and (2) the mechanism of chymotrypsin What s so great about enzymes? They accomplish large rate accelerations (10 10-10
WHY IS THIS IMPORTANT?
 CHAPTER 2 FUNDAMENTAL CHEMISTRY FOR MICROBIOLOGY WHY IS THIS IMPORTANT? An understanding of chemistry is essential to understand cellular structure and function, which are paramount for your understanding
CHAPTER 2 FUNDAMENTAL CHEMISTRY FOR MICROBIOLOGY WHY IS THIS IMPORTANT? An understanding of chemistry is essential to understand cellular structure and function, which are paramount for your understanding
Lecture 34. Carbohydrate Metabolism 2. Glycogen. Key Concepts. Biochemistry and regulation of glycogen degradation
 Lecture 34 Carbohydrate Metabolism 2 Glycogen Key Concepts Overview of Glycogen Metabolism Biochemistry and regulation of glycogen degradation Biochemistry and regulation of glycogen synthesis What mechanisms
Lecture 34 Carbohydrate Metabolism 2 Glycogen Key Concepts Overview of Glycogen Metabolism Biochemistry and regulation of glycogen degradation Biochemistry and regulation of glycogen synthesis What mechanisms
Organic Chemistry Laboratory Fall Lecture 3 Gas Chromatography and Mass Spectrometry June
 344 Organic Chemistry Laboratory Fall 2013 Lecture 3 Gas Chromatography and Mass Spectrometry June 19 2013 Chromatography Chromatography separation of a mixture into individual components Paper, Column,
344 Organic Chemistry Laboratory Fall 2013 Lecture 3 Gas Chromatography and Mass Spectrometry June 19 2013 Chromatography Chromatography separation of a mixture into individual components Paper, Column,
(5) 1. List five unusual properties of water resulting from its hydrogen bonded structure
 BCH 4053 June 1, 2001 Points HOUR TEST 1 NAME (5) 1. List five unusual properties of water resulting from its hydrogen bonded structure. Page Points 1 2 3 4 5 Total (5) 2. Draw a diagram to show how water
BCH 4053 June 1, 2001 Points HOUR TEST 1 NAME (5) 1. List five unusual properties of water resulting from its hydrogen bonded structure. Page Points 1 2 3 4 5 Total (5) 2. Draw a diagram to show how water
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
6. The catalytic mechanism of arylsulfatase A and its theoretical investigation
 6. The catalytic mechanism of arylsulfatase A and its theoretical investigation When the crystal structure of arylsulfatase A was solved, a remarkable structural analogy to another hydrolytic enzyme, the
6. The catalytic mechanism of arylsulfatase A and its theoretical investigation When the crystal structure of arylsulfatase A was solved, a remarkable structural analogy to another hydrolytic enzyme, the
Will s Pre-Test. (4) A collection of cells that work together to perform a function is termed a(n): a) Organelle b) Organ c) Cell d) Tissue e) Prison
 Will s Pre-Test This is a representative of Exam I that you will take Tuesday September 18, 2007. The actual exam will be 50 multiple choice questions. (1) The basic structural and functional unit of the
Will s Pre-Test This is a representative of Exam I that you will take Tuesday September 18, 2007. The actual exam will be 50 multiple choice questions. (1) The basic structural and functional unit of the
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
KEY NAME (printed very legibly) UT-EID
 BIOLOGY 311C - Brand Spring 2007 KEY NAME (printed very legibly) UT-EID EXAMINATION II Before beginning, check to be sure that this exam contains 7 pages (including front and back) numbered consecutively,
BIOLOGY 311C - Brand Spring 2007 KEY NAME (printed very legibly) UT-EID EXAMINATION II Before beginning, check to be sure that this exam contains 7 pages (including front and back) numbered consecutively,
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Biochemistry 2 Recita0on Amino Acid Metabolism
 Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
Biochemistry I Professor S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture - 18 Vitamins and Coenzymes-I
 Biochemistry I Professor S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture - 18 Vitamins and Coenzymes-I We start our discussion on vitamins and coenzymes. We will have
Biochemistry I Professor S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture - 18 Vitamins and Coenzymes-I We start our discussion on vitamins and coenzymes. We will have
Final Exam Chemistry 391 Structural Biochemistry Fall Do not open the exam until ready to begin! Rules of the Game:
 Name Practice for 2018 Final Exam Chemistry 391 Structural Biochemistry Fall 2016 Do not open the exam until ready to begin! ules of the Game: This is a take-home Exam. The exam is due on Thursday, December
Name Practice for 2018 Final Exam Chemistry 391 Structural Biochemistry Fall 2016 Do not open the exam until ready to begin! ules of the Game: This is a take-home Exam. The exam is due on Thursday, December
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B
 3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
130327SCH4U_biochem April 09, 2013
 Option B: B1.1 ENERGY Human Biochemistry If more energy is taken in from food than is used up, weight gain will follow. Similarly if more energy is used than we supply our body with, weight loss will occur.
Option B: B1.1 ENERGY Human Biochemistry If more energy is taken in from food than is used up, weight gain will follow. Similarly if more energy is used than we supply our body with, weight loss will occur.
Supporting information
 Supporting information Figure legends Supplementary Table 1. Specific product ions obtained from fragmentation of lithium adducts in the positive ion mode comparing the different positional isomers of
Supporting information Figure legends Supplementary Table 1. Specific product ions obtained from fragmentation of lithium adducts in the positive ion mode comparing the different positional isomers of
SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES
 SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors:
SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors:
Chapter 3. Table of Contents. Section 1 Carbon Compounds. Section 2 Molecules of Life. Biochemistry
 Biochemistry Table of Contents Section 1 Carbon Compounds Section 2 Molecules of Life Section 1 Carbon Compounds Objectives Distinguish between organic and inorganic compounds. Explain the importance of
Biochemistry Table of Contents Section 1 Carbon Compounds Section 2 Molecules of Life Section 1 Carbon Compounds Objectives Distinguish between organic and inorganic compounds. Explain the importance of
Data are contained in multiple tabs in Excel spreadsheets and in CSV files.
 Contents Overview Curves Methods Measuring enzymatic activity (figure 2) Enzyme characterisation (Figure S1, S2) Enzyme kinetics (Table 3) Effect of ph on activity (figure 3B) Effect of metals and inhibitors
Contents Overview Curves Methods Measuring enzymatic activity (figure 2) Enzyme characterisation (Figure S1, S2) Enzyme kinetics (Table 3) Effect of ph on activity (figure 3B) Effect of metals and inhibitors
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/1/e1500678/dc1 Supplementary Materials for Chemical synthesis of erythropoietin glycoforms for insights into the relationship between glycosylation pattern and
advances.sciencemag.org/cgi/content/full/2/1/e1500678/dc1 Supplementary Materials for Chemical synthesis of erythropoietin glycoforms for insights into the relationship between glycosylation pattern and
MIDDLETOWN HIGH SCHOOL SOUTH BIOLOGY
 MIDDLETOWN HIGH SCHOOL SOUTH BIOLOGY BOOKLET 10 NAME: CLASS: 1 S.Tagore Middletown South High School March 2013 LEARNING OUTCOMES The role and production of ATP (a) Importance, role and structure of ATP
MIDDLETOWN HIGH SCHOOL SOUTH BIOLOGY BOOKLET 10 NAME: CLASS: 1 S.Tagore Middletown South High School March 2013 LEARNING OUTCOMES The role and production of ATP (a) Importance, role and structure of ATP
Biochemistry - I. Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I
 Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
KDM2A. Reactions. containing. Reactions
 Supplementary Figure 1 KDM2A catalyses only lysine demethylation.. MALDI-TOF MS of demethylation of the shown variant histone peptides as catalysed by recombinant KDM2A. Reactions containing enzyme are
Supplementary Figure 1 KDM2A catalyses only lysine demethylation.. MALDI-TOF MS of demethylation of the shown variant histone peptides as catalysed by recombinant KDM2A. Reactions containing enzyme are
Ultra Columns. also available. ordering note
 : 3μm or 5μm particles; 100Å pore size Our broadest selection of stationary phases, including unique phases. High density bondings, for maximum retention. High-purity, Type B silica gives excellent peak
: 3μm or 5μm particles; 100Å pore size Our broadest selection of stationary phases, including unique phases. High density bondings, for maximum retention. High-purity, Type B silica gives excellent peak
Supplementary Information for Structural basis for the inhibition of Polo-like kinase 1
 Supplementary Information for Structural basis for inhibition of Polo-like kinase 1 Jun Xu, Chen Shen, Tao Wang & Junmin Quan Correspondence authors: Tao Wang, taowang@pkusz.edu.cn; Junmin Quan, quanjm@szpku.edu.cn
Supplementary Information for Structural basis for inhibition of Polo-like kinase 1 Jun Xu, Chen Shen, Tao Wang & Junmin Quan Correspondence authors: Tao Wang, taowang@pkusz.edu.cn; Junmin Quan, quanjm@szpku.edu.cn
An Introduction to Carbohydrates
 An Introduction to Carbohydrates Carbohydrates are a large class of naturally occurring polyhydroxy aldehydes and ketones. Monosaccharides also known as simple sugars, are the simplest carbohydrates containing
An Introduction to Carbohydrates Carbohydrates are a large class of naturally occurring polyhydroxy aldehydes and ketones. Monosaccharides also known as simple sugars, are the simplest carbohydrates containing
Supplemental Information. An Atlas of b-glucuronidases. in the Human Intestinal Microbiome
 Structure, Volume 2 Supplemental Information An Atlas of b-glucuronidases in the Human Intestinal Microbiome Rebecca M. Pollet, Emma H. D'Agostino, William G. Walton, Yongmei Xu, Michael S. Little, Kristen
Structure, Volume 2 Supplemental Information An Atlas of b-glucuronidases in the Human Intestinal Microbiome Rebecca M. Pollet, Emma H. D'Agostino, William G. Walton, Yongmei Xu, Michael S. Little, Kristen
Self-organization of dipyridylcalix[4]pyrrole into a supramolecular cage for dicarboxylates
![Self-organization of dipyridylcalix[4]pyrrole into a supramolecular cage for dicarboxylates Self-organization of dipyridylcalix[4]pyrrole into a supramolecular cage for dicarboxylates](/thumbs/94/118164746.jpg) Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2016 Electronic Supplementary Information Self-organization of dipyridylcalix[4]pyrrole into a supramolecular
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2016 Electronic Supplementary Information Self-organization of dipyridylcalix[4]pyrrole into a supramolecular
A pillar[2]arene[3]hydroquinone which can self-assemble to a molecular zipper in the solid state
![A pillar[2]arene[3]hydroquinone which can self-assemble to a molecular zipper in the solid state A pillar[2]arene[3]hydroquinone which can self-assemble to a molecular zipper in the solid state](/thumbs/86/93673402.jpg) A pillar[2]arene[3]hydroquinone which can self-assemble to a molecular zipper in the solid state Mingguang Pan, Min Xue* Department of Chemistry, Zhejiang University, Hangzhou 310027, P. R. China Fax:
A pillar[2]arene[3]hydroquinone which can self-assemble to a molecular zipper in the solid state Mingguang Pan, Min Xue* Department of Chemistry, Zhejiang University, Hangzhou 310027, P. R. China Fax:
Metabolism Energy Pathways Biosynthesis. Catabolism Anabolism Enzymes
 Topics Microbial Metabolism Metabolism Energy Pathways Biosynthesis 2 Metabolism Catabolism Catabolism Anabolism Enzymes Breakdown of complex organic molecules in order to extract energy and dform simpler
Topics Microbial Metabolism Metabolism Energy Pathways Biosynthesis 2 Metabolism Catabolism Catabolism Anabolism Enzymes Breakdown of complex organic molecules in order to extract energy and dform simpler
Expression constructs
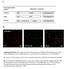 Gene expressed in bebe3 ZmBEa Expression constructs 35S ZmBEa Pnos:Hygromycin r 35S Pnos:Hygromycin r 35S ctp YFP Pnos:Hygromycin r B -1 Chl YFP- Merge Supplemental Figure S1: Constructs Used for the Expression
Gene expressed in bebe3 ZmBEa Expression constructs 35S ZmBEa Pnos:Hygromycin r 35S Pnos:Hygromycin r 35S ctp YFP Pnos:Hygromycin r B -1 Chl YFP- Merge Supplemental Figure S1: Constructs Used for the Expression
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Chem 109 C. Fall Armen Zakarian Office: Chemistry Bldn 2217
 Chem 109 C Fall 2014 Armen Zakarian ffice: Chemistry Bldn 2217 midterms Midterm 1 max 92.5, min 28.5 average 64; stdev 13.8 Midterm 2 max 90, min 15 average 54; stdev 16.3 utline o overview of catabolism
Chem 109 C Fall 2014 Armen Zakarian ffice: Chemistry Bldn 2217 midterms Midterm 1 max 92.5, min 28.5 average 64; stdev 13.8 Midterm 2 max 90, min 15 average 54; stdev 16.3 utline o overview of catabolism
Syllabus for BASIC METABOLIC PRINCIPLES
 Syllabus for BASIC METABOLIC PRINCIPLES The video lecture covers basic principles you will need to know for the lectures covering enzymes and metabolism in Principles of Metabolism and elsewhere in the
Syllabus for BASIC METABOLIC PRINCIPLES The video lecture covers basic principles you will need to know for the lectures covering enzymes and metabolism in Principles of Metabolism and elsewhere in the
Nature Structural & Molecular Biology: doi: /nsmb.1933
 The structural basis of open channel block in a prokaryotic pentameric ligand-gated ion channel Ricarda J. C. Hilf, Carlo Bertozzi, Iwan Zimmermann, Alwin Reiter, Dirk Trauner and Raimund Dutzler a GLIC
The structural basis of open channel block in a prokaryotic pentameric ligand-gated ion channel Ricarda J. C. Hilf, Carlo Bertozzi, Iwan Zimmermann, Alwin Reiter, Dirk Trauner and Raimund Dutzler a GLIC
SUPPLEMENTARY INFORMATION
 Supplementary Information Supplementary Figure 1. The serine paradox. Nature-designed AlaRSs pin-down the -NH3 + group of alanine with an acidic Asp that is critical for binding and activation of alanine.
Supplementary Information Supplementary Figure 1. The serine paradox. Nature-designed AlaRSs pin-down the -NH3 + group of alanine with an acidic Asp that is critical for binding and activation of alanine.
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Lecture 2: Glycogen metabolism (Chapter 15)
 Lecture 2: Glycogen metabolism (Chapter 15) First. Fig. 15.1 Review: Animals use glycogen for ENERGY STORAGE. Glycogen is a highly-branched polymer of glucose units: Basic structure is similar to that
Lecture 2: Glycogen metabolism (Chapter 15) First. Fig. 15.1 Review: Animals use glycogen for ENERGY STORAGE. Glycogen is a highly-branched polymer of glucose units: Basic structure is similar to that
CHAPTER 9: CATALYTIC STRATEGIES. Chess vs Enzymes King vs Substrate
 CHAPTER 9: CATALYTIC STRATEGIES Chess vs Enzymes King vs Substrate INTRODUCTION CHAPTER 9 What are the sources of the catalytic power and specificity of enzymes? Problems in reactions in cells Neutral
CHAPTER 9: CATALYTIC STRATEGIES Chess vs Enzymes King vs Substrate INTRODUCTION CHAPTER 9 What are the sources of the catalytic power and specificity of enzymes? Problems in reactions in cells Neutral
Protein Modification Overview DEFINITION The modification of selected residues in a protein and not as a component of synthesis
 Lecture Four: Protein Modification & Cleavage [Based on Chapters 2, 9, 10 & 11 Berg, Tymoczko & Stryer] (Figures in red are for the 7th Edition) (Figures in Blue are for the 8th Edition) Protein Modification
Lecture Four: Protein Modification & Cleavage [Based on Chapters 2, 9, 10 & 11 Berg, Tymoczko & Stryer] (Figures in red are for the 7th Edition) (Figures in Blue are for the 8th Edition) Protein Modification
This PDF file includes: Supplementary Figures 1 to 6 Supplementary Tables 1 to 2 Supplementary Methods Supplementary References
 Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
The Chemical Building Blocks of Life. Chapter 3
 The Chemical Building Blocks of Life Chapter 3 Biological Molecules Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent
The Chemical Building Blocks of Life Chapter 3 Biological Molecules Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information.
 Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached
 BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached 1). 20 points total T or F (2 points each; if false, briefly state why
BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached 1). 20 points total T or F (2 points each; if false, briefly state why
Biological Molecules
 The Chemical Building Blocks of Life Chapter 3 Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent bonds. Carbon may
The Chemical Building Blocks of Life Chapter 3 Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent bonds. Carbon may
Enzymes. Enzyme. Aim: understanding the basic concepts of enzyme catalysis and enzyme kinetics
 Enzymes Substrate Enzyme Product Aim: understanding the basic concepts of enzyme catalysis and enzyme kinetics Enzymes are efficient Enzyme Reaction Uncatalysed (k uncat s -1 ) Catalysed (k cat s -1 )
Enzymes Substrate Enzyme Product Aim: understanding the basic concepts of enzyme catalysis and enzyme kinetics Enzymes are efficient Enzyme Reaction Uncatalysed (k uncat s -1 ) Catalysed (k cat s -1 )
Supporting Information
 Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Practice Problems 3. a. What is the name of the bond formed between two amino acids? Are these bonds free to rotate?
 Life Sciences 1a Practice Problems 3 1. Draw the oligopeptide for Ala-Phe-Gly-Thr-Asp. You do not need to indicate the stereochemistry of the sidechains. Denote with arrows the bonds formed between the
Life Sciences 1a Practice Problems 3 1. Draw the oligopeptide for Ala-Phe-Gly-Thr-Asp. You do not need to indicate the stereochemistry of the sidechains. Denote with arrows the bonds formed between the
B i o c h e m i s t r y N o t e s
 14 P a g e Carbon Hydrogen Nitrogen Oxygen Phosphorus Sulfur ~Major ~Found in all ~Found in most ~Found in all component of all organic organic molecules. molecules. ~Major structural atom in all organic
14 P a g e Carbon Hydrogen Nitrogen Oxygen Phosphorus Sulfur ~Major ~Found in all ~Found in most ~Found in all component of all organic organic molecules. molecules. ~Major structural atom in all organic
This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth 2 points.
 MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
SUPPORTING INFORMATION. Lysine Carbonylation is a Previously Unrecognized Contributor. to Peroxidase Activation of Cytochrome c by Chloramine-T
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Supporting Information for:
 Supporting Information for: Methylerythritol Cyclodiphosphate (MEcPP) in Deoxyxylulose Phosphate Pathway: Synthesis from an Epoxide and Mechanisms Youli Xiao, a Rodney L. Nyland II, b Caren L. Freel Meyers
Supporting Information for: Methylerythritol Cyclodiphosphate (MEcPP) in Deoxyxylulose Phosphate Pathway: Synthesis from an Epoxide and Mechanisms Youli Xiao, a Rodney L. Nyland II, b Caren L. Freel Meyers
Biology 2E- Zimmer Protein structure- amino acid kit
 Biology 2E- Zimmer Protein structure- amino acid kit Name: This activity will use a physical model to investigate protein shape and develop key concepts that govern how proteins fold into their final three-dimensional
Biology 2E- Zimmer Protein structure- amino acid kit Name: This activity will use a physical model to investigate protein shape and develop key concepts that govern how proteins fold into their final three-dimensional
Supporting Information
 Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Background knowledge
 Background knowledge This is the required background knowledge: State three uses of energy in living things Give an example of an energy conversion in a living organism State that fats and oils contain
Background knowledge This is the required background knowledge: State three uses of energy in living things Give an example of an energy conversion in a living organism State that fats and oils contain
Review Session 1. Control Systems and Homeostasis. Figure 1.8 A simple control system. Biol 219 Review Sessiono 1 Fall 2016
 Control Systems and Homeostasis Review Session 1 Regulated variables are kept within normal range by control mechanisms Keeps near set point, or optimum value Control systems local and reflex Input signal
Control Systems and Homeostasis Review Session 1 Regulated variables are kept within normal range by control mechanisms Keeps near set point, or optimum value Control systems local and reflex Input signal
SUPPLEMENTARY INFORMATION
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2919 Direct observation of the influence of cardiolipin and antibiotics on lipid II binding to MurJ Jani
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2919 Direct observation of the influence of cardiolipin and antibiotics on lipid II binding to MurJ Jani
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site.
 Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
