SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES
|
|
|
- Godwin Nelson
- 5 years ago
- Views:
Transcription
1 SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors: Ralph Adams 1,#, Rebecca J. Burnley 1,#, Chiara R. Valenzano 1,#, Omar Qureshi 1,CarlDoyle 1, Simon Lumb 1, Maria del Carmen Lopez 1,^, Robert Griffin 1, David McMillan 1, Richard D. Taylor 1,Chris Meier 1, Prashant Mori 1, Laura M. Griffin 1, Ulrich Wernery 2, Jörg Kinne 2, Stephen Rapecki 1, Terry S. Baker 1, Alastair D. G. Lawson 1, Michael Wright 1, and Anna Ettorre 1, * Affiliations: 1 New Medicines, UCB-Celltech, 208 Bath Road, SL1 3WE, Slough UK; 2 Central Veterinary Research Laboratory, P.O.Box 597, Dubai, United Arab Emirates. # These authors contributed equally to the work; * corresponding author.
2 2 SI Figure 1 S22 S22 Y27 Y27 Y32 Y32 SI Figure 1. Differential hydrogen bonding in the two copies of the crystallographic asymmetric unit. There are two copies of the VHH6 IL-6 gp80 6 complex in the crystallographic asymmetric unit. In one copy, IL-6 (green) residue Ser22 forms a hydrogen bond with VHH6 (orange) residue Tyr32 (left), and in the second copy, Ser22 forms a hydrogen bond with VHH6 residue Tyr27 (right).
3 SI Figure 2 IL 6 Site 1 3 gp130 VHH6 Site 2 gp80 SI Figure 2. Model of crystal structure of VHH6 IL-6 gp80 superimposed with gp130. Superimposition of IL-6 (green) and gp80 (blue) from the signaling complex, IL-6 gp80 gp130 (PDB code 1P9M), showed low r.m.s.d. values of 1.4±0.2 Å and 1.35±0.05 Å respectively, indicating that VHH6 (orange) holds IL-6 and gp80 together in a form able to bind gp130 (cyan).
4 4 SI Figure 3 SI Figure 3. Stabilization of the IL-6 gp80 complex by VHH6 promotes higher and sustained STAT3 phosphorylation signal in HUVECs comparable to FusionIL-6 fusion protein. HUVECs were treated with FusionIL-6, IL-6+gp80 and VHH6+IL-6+gp80 as described in Materials and Methods. STAT3 phosphorylation signal was quantified at different time points using the spot total intensity per object parameter as described in Materials and Methods. Statistical analysis of pstat3 signal from three replicates was performed for each time point (statistical significance: *=p 0.05; **=p 0.01 and ***=p 0.001). Unlike IL-6+gp80, in the presence of VHH6 or when FusionIL-6 was added to the culture, pstat3 was increased at earlier (30 min) and later time points (180 min and 360 min). y-axis: pstat3 fluorescence expressed as spot total intensity per object; x-axis: time (min).
5 SI Figure 4 VHH6+ IL 6+ gp80 IL 6+ gp SI Figure 4. Transcriptomic analysis of HUVECs treated with VHH6+IL-6+gp80 confirms selective up-regulation of proinflammatory genes. Three different batches of HUVECs were analyzed at 30, 180 and 360 min. VHH6+IL-6+gp80 (sample) were compared to IL-6+gp80 (control). Data were analyzed using the RT 2 Profiler PCR array web based data analysis template v3.5 ( pcr/arrayanalysis) and changes in gene expression changes were calculated using the C t method with normalization of the raw data to housekeeping genes. A heat map was generated using Genedata s Analyst software. Genes in grey-black are down-regulated, while genes in yellow are up-regulated. Gene regulation for each sample is expressed as fold change compared to control.
6 6 SI Table 1 Table 1A: Data collection and refinement statistics (molecular replacement) Junctional Epitope Antibody in complex with IL-6 and gp80 Data collection Space group C Cell dimensions a, b, c (Å) , 67.80, ( ) 90.00, , Resolution (Å) ( )* R meas (%) CC 1/2 (%) 7.2(50.1) 99.9(91.5) I / I 20.88(3.97) Completeness (%) 99.4(96.9) Redundancy 7.5(7.5) Refinement Resolution (Å) No. reflections 34,870 R work /R free / No. atoms Protein 7590 Water 111 B-factors Protein Water R.m.s. deviations Bond lengths (Å) Bond angles ( ) *Values in parentheses are for highest-resolution shell.
7 7 SI Table 1B: Single mutants of VHH6 kd (1/s) wtvhh6 N74A K113A Y32A S101A Y27A 3.6E E E E E E-04 SI Table 1C: Double mutants of VHH6 kd (1/s) wtvhh6 Y27A Y32A N74A S101A K113A S101A K113S S101A 3.6E E E E E-04 SI Table 1. Additional data supporting the junctional epitope nature of VHH6. (A) Data collection and refinement statistics (molecular replacement). To assess the contributions of the side chains to binding, alanine-scanning was carried out. The dissociation rate of single (B) and double mutants (C) were assessed using SPR.
8 8 SI Table 2: Peptides identified and quantified in IL-6 and gp80 by HDX-MS SI Table 2A: Peptide identified in IL-6 Start Residue End Residue Sequence 2 timepoints with p< APVPPGEDSKDV 3 23 PPGEDSKDVAAPHRQPLTSSE SSERIDKQ * SSERIDKQIRY * SERIDKQIRY * RIDKQIRYIL * * IDKQIRYILDGI * * IRYILDGISAL ALRKETCNKSNM # LRKETCNKSNM # RKETCNKSNM # RKETCNKSNMCE KETCNKSNMCESSKEALAENN # ETCNKSNMCESSKEA SSKEALAENNLNLPKMAEKDGCF Q SKEALAENNLN SKEALAENNLNLPKMAEKDGCFQ EALAENNLNLP AENNLNLPKM AENNLNLPKMAEKDGCF * * ENNLNLPKMAEKDGCFQSGFNE * NNLNLPKM NNLNLPKMAEK NNLNLPKMAEKDGCFQSGFNEET * AEKDGCF * * KDGCFQSGFNEE * QSGFNEETCL VKIITGLL VKIITGLLE IITGLLE LEYLQNRFESS EYLQNRFESSE EYLQNRFESSEE EYLQNRFESSEEQA 4 timepoints with p<0.01
9 EYLQNRFESSEEQARAVQMSTK YLQNRFESSEE YLQNRFESSEEQA RFESSEEQARA ESSEEQA ESSEEQARAVQ EEQARAVQ VQMSTKVL VQMSTKVL KVLIQFL IQFLQKKAKNLDA LQKKAKNL * LQKKAKNLDA * LQKKAKNLDAITTPDPTTN * LQKKAKNLDAITTPDPTTNA LQKKAKNLDAITTPDPTTNASL * KAKNLDAITTPDPTT ITTPDPTTNASL * * LTKLQAQN LTKLQAQN LTKLQAQNQWL LTKLQAQNQWLQD QAQNQWL QDMTTHL * QDMTTHLIL * MTTHLIL * ILRSFKE * * ILRSFKEF * * ILRSFKEFL * RSFKEFL * * RSFKEFLQSSL * * 9
10 10 SI Table 2B: SI Table 2A: Peptide identified in gp80 Start Residue End Sequence Residue HENLYFQGLAPRRCPAQEVAR GLAPRRCPAQ GLAPRRCPAQE * GLAPRRCPAQEVA * RCPAQEVARGAGAGDVPPE AQEVARGAGAGDVPPEEPQLSC * QEVARGAGAGDVPP EVARGAGAGDVP * VARGAGAGDVPPEEPQL * RGAGAGDVPPEEPQL * DVPPEEPQLSCFRKSPLSN * PPEEPQL FRKSPLSNV KSPLSNVVCEWGPRSTPSL VVCEWGPRSTPSL VCEWGPRSTPSL WGPRSTPSLTTKAVL TTKAVLL LVRKFQNSPAED VRKFQNSPAED QNSPAEDFQEP AEDFQEPCQYSQESQKFSCQ DFQEPCQ CQYSQESQKFSCQLAVPEG QYSQESQKFSCQLAVPEG YSQESQKFSCQ SQKFSCQ CQLAVPEGDSSFYI LAVPEGDSSF * PEGDSSF * CVASSVGSKF ASSVGSKF LQPDPPANITV * ANITVTAVAR TAVARNPRWLS * AVARNPRW VARNPRW VARNPRWLSV * 2 timepoints with p< timepoints with p<0.01
11 VARNPRWLSVT * TWQDPHSWNSSF * DPHSWNSSF YRLRFEL ELRYRAERSKT RYRAERSKT SKTFTTWMVKDLQH * * MVKDLQHHAVIHD * MVKDLQHHAVIHDAWSG * * MVKDLQHHAVIHDAWSGLRHVVQL * SGLRHVVQL LRHVVQL RHVVQLRA RAQEEFGQGE * * RAQEEFGQGEW * * RAQEEFGQGEWSE * * EEFGQGEWSEWSPEAMGTP GQGEWSEWSPEAMGTPW WSEWSPEA * WSEWSPEAMGTPWTE WSEWSPEAMGTPWTESRSPP SEWSPEA SEWSPEAMGTPWTE SEWSPEAMGTPWTESRSPP WSPEAMGTPWTE PEAMGTPWTESRSPP MGTPWTE SI Table 2: Additional data supporting HDX-MS. List of peptides identified in IL-6 (A) and gp80 (B) and quantified by HDX-MS. Asterisk (*) indicates deuterium uptake increasing the presence of VHH6; hash (#) indicates deuterium uptake decreasing in the presence of VHH6.
12 12 SI Table 3: SPR analysis IL-6 immobilized kd (1/s) SE (kd) KD (M) SE (KD) gp E E E-09 gp80+vhh6 (2 M) 1.94E E-07 NA gp80 immobilized kd (1/s) SE (kd) KD (M) SE (KD) IL E E E-09 IL-6+VHH6 (2 M) 2.10E E-07 NA VHH6 immobilized kd (1/s) SE (kd) KD (M) IL-6+gp80 (2 M) 3.09E E-07 NA gp80+il-6 (2 M) 3.75E E-07 NA gp130 immobilized ka (1/Ms) kd (1/s) gp80 [IL-6 (2 M)] ^ gp80 [IL-6 (2 M)+VHH6 (2 M)] ^ IL-6 [gp80 (2 M)] ^ IL-6 [gp80 (2 M) + VHH6 (2 M)] ^ SI Table 3: Additional SPR data analysis. During SPR studies binding of IL-6, gp80 and VHH6 were individually tested in a concentration series (0-250 nm, as two-fold serial dilution). When proteins were tested in combinations (IL-6 and gp80, IL-6 and VHH6, gp80 and VHH6, gp80 and IL-6 and VHH6) one of the proteins was titrated in a concentration series (0-250 nm), while the other was kept constant at an excess concentration of 2 µm (in brackets) to ensure complex formation. A total of four proteins were immobilized on the chip, from top to bottom: IL-6, gp80, VHH6 and gp130.
13 13 SI Table 4: In-house constructs Construct pnafl-8his- fusionil- 6(gp80D123) pnafl-gp80d123- hscfc Expression Host Promoter Affinity Tag Expressed Protein CHO CMV 8His gp80(l20- S320)(C211A, C277A)- IL-6(V30-M212) CHO CMV Single chain gp80(m1- human IgG Fc P322)(C211A, C277A)- fragment hscfc pnafl-8his-gp80v4 CHO CMV 8His gp80v4 ptrx-6his-hil-6 E.coli T7 Thioredoxin- 6His Trx-6His-IL-6(A28- M212) ptrx-6his-il-6(s21) E.coli T7 Thioredoxin- 6His Trx-6His-IL-6(S49- M212) pimms-6his-vhh6 Expi-HEK CMV 6His VHH6-6His SI Table 4: In-house designed constructs. All constructs listed in the table were designed in-house to support the immunization campaign, antibody screening, junctional antibody characterization and biophysics analysis.
Developments in NMR Fragment Screening at UCB. Richard J. Taylor CCPN Conference 13th.July.2017
 Developments in NMR Fragment Screening at UCB Richard J. Taylor CCPN Conference 13th.July.2017 High Quality, Soluble, Chemically Diverse Fragment Subset NMR Hardware AVIII HD 600 MHz. 5 mm QCI-F He Cryoprobe
Developments in NMR Fragment Screening at UCB Richard J. Taylor CCPN Conference 13th.July.2017 High Quality, Soluble, Chemically Diverse Fragment Subset NMR Hardware AVIII HD 600 MHz. 5 mm QCI-F He Cryoprobe
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Table S1. X-ray data collection and refinement statistics
 Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Information Janssen et al.
 Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supporting Information
 Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
SUPPLEMENTARY INFORMATION FOR. (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation. by COX-2
 SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
Crystallization-grade After D After V3 cocktail. Time (s) Time (s) Time (s) Time (s) Time (s) Time (s)
 Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Supplementary Information
 Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
FCC2 5CY7 FCC1 5CY8. Actinonin 5CVQ
 Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Supporting Information
 Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information.
 Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood
 Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Crystal structure of the neutralizing antibody HK20 in complex with its gp41 antigen
 Crystal structure of the neutralizing antibody HK20 in complex with its gp41 antigen David Lutje Hulsik Unit for Virus Host Cell Interaction UMI 3265 University Joseph Fourier-EMBL-CNRS, Grenoble Env catalyzed
Crystal structure of the neutralizing antibody HK20 in complex with its gp41 antigen David Lutje Hulsik Unit for Virus Host Cell Interaction UMI 3265 University Joseph Fourier-EMBL-CNRS, Grenoble Env catalyzed
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors
 Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Structure of the measles virus hemagglutinin bound to the CD46 receptor. César Santiago, María L. Celma, Thilo Stehle and José M.
 Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
SUPPORTING INFORMATION FOR. A Computational Approach to Enzyme Design: Using Docking and MM- GBSA Scoring
 SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 The UBL and RING1 interface remains associated in the complex structures of Parkin and pub. a) Asymmetric Unit of crystal structure of UBLR0RBR and pub complex showing UBL (green),
Supplementary Figure 1 The UBL and RING1 interface remains associated in the complex structures of Parkin and pub. a) Asymmetric Unit of crystal structure of UBLR0RBR and pub complex showing UBL (green),
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein
 Supplementary Methods Thermal shift assays Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein unfolding was examined by monitoring the fluorescence of ANS (1-anilinonaphthalene-8-
Supplementary Methods Thermal shift assays Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein unfolding was examined by monitoring the fluorescence of ANS (1-anilinonaphthalene-8-
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/283/ra57/dc1 Supplementary Materials for JNK3 Couples the Neuronal Stress Response to Inhibition of Secretory Trafficking Guang Yang,* Xun Zhou, Jingyan Zhu,
www.sciencesignaling.org/cgi/content/full/6/283/ra57/dc1 Supplementary Materials for JNK3 Couples the Neuronal Stress Response to Inhibition of Secretory Trafficking Guang Yang,* Xun Zhou, Jingyan Zhu,
Supplementary Figure 1. Properties of various IZUMO1 monoclonal antibodies and behavior of SPACA6. (a) (b) (c) (d) (e) (f) (g) .
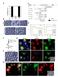 Supplementary Figure 1. Properties of various IZUMO1 monoclonal antibodies and behavior of SPACA6. (a) The inhibitory effects of new antibodies (Mab17 and Mab18). They were investigated in in vitro fertilization
Supplementary Figure 1. Properties of various IZUMO1 monoclonal antibodies and behavior of SPACA6. (a) The inhibitory effects of new antibodies (Mab17 and Mab18). They were investigated in in vitro fertilization
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism
 Supplementary Information The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism Address correspondence to Yong Li (yongli@xmu.edu.cn, Tel: 86-592-218151) GW464 CDCA Supplementary
Supplementary Information The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism Address correspondence to Yong Li (yongli@xmu.edu.cn, Tel: 86-592-218151) GW464 CDCA Supplementary
SUPPLEMENTARY INFORMATION. Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity
 SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTAL INFORMATION. Evolving Spindlin1 Small Molecule Inhibitors Using Protein Microarrays
 SUPPLEMETAL IFRMATI Evolving Spindlin1 Small Molecule Inhibitors Using Protein Microarrays arkhyun Bae 1, Monica Viviano 2, Xiaonan Su 3, Jie Lu 4, Donghang Cheng 1, Cari Sagum 1, Sabrina Castellano 2,6,
SUPPLEMETAL IFRMATI Evolving Spindlin1 Small Molecule Inhibitors Using Protein Microarrays arkhyun Bae 1, Monica Viviano 2, Xiaonan Su 3, Jie Lu 4, Donghang Cheng 1, Cari Sagum 1, Sabrina Castellano 2,6,
Supporting Information
 Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Supplementary Figure 1. BMS enhances human T cell activation in vitro in a
 Supplementary Figure 1. BMS98662 enhances human T cell activation in vitro in a concentration-dependent manner. Jurkat T cells were activated with anti-cd3 and anti-cd28 antibody in the presence of titrated
Supplementary Figure 1. BMS98662 enhances human T cell activation in vitro in a concentration-dependent manner. Jurkat T cells were activated with anti-cd3 and anti-cd28 antibody in the presence of titrated
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
SUPPLEMENTAL MATERIAL. UNC119 is required for G protein trafficking in sensory neurons
 1 SUPPLEMENTAL MATERIAL UNC119 is required for G protein trafficking in sensory neurons Houbin Zhang, Ryan N. Constantine, Sergey Vorobiev, Yang Chen, Jayaraman Seetharaman, Yuanpeng Janet Huang, Rong
1 SUPPLEMENTAL MATERIAL UNC119 is required for G protein trafficking in sensory neurons Houbin Zhang, Ryan N. Constantine, Sergey Vorobiev, Yang Chen, Jayaraman Seetharaman, Yuanpeng Janet Huang, Rong
Supporting Information
 Supporting Information Bauer et al. 10.1073/pnas.1610421113 SI Materials and Methods Immunofluorescence experiments were performed as described (8). Briefly, NUGC-3 cells were treated with 25 μm PK11007
Supporting Information Bauer et al. 10.1073/pnas.1610421113 SI Materials and Methods Immunofluorescence experiments were performed as described (8). Briefly, NUGC-3 cells were treated with 25 μm PK11007
ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP
 ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP Peera Jaru-Ampornpan 1, Kuang Shen 1,3, Vinh Q. Lam 1,3, Mona Ali 2, Sebastian Doniach 2, Tony Z. Jia 1, Shu-ou Shan 1 Supplementary
ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP Peera Jaru-Ampornpan 1, Kuang Shen 1,3, Vinh Q. Lam 1,3, Mona Ali 2, Sebastian Doniach 2, Tony Z. Jia 1, Shu-ou Shan 1 Supplementary
KDM2A. Reactions. containing. Reactions
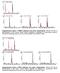 Supplementary Figure 1 KDM2A catalyses only lysine demethylation.. MALDI-TOF MS of demethylation of the shown variant histone peptides as catalysed by recombinant KDM2A. Reactions containing enzyme are
Supplementary Figure 1 KDM2A catalyses only lysine demethylation.. MALDI-TOF MS of demethylation of the shown variant histone peptides as catalysed by recombinant KDM2A. Reactions containing enzyme are
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. Schematic of Ras biochemical coupled assay.
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. Schematic of Ras biochemical coupled assay.
List of Figures. List of Tables
 Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
Supplementary Information for. Heavy chain-only IgG2b-llama antibody effects near-pan HIV-1 neutralization by
 Supplementary Information for Heavy chain-only IgG2b-llama antibody effects near-pan HIV-1 neutralization by recognizing a CD4-induced epitope that includes elements of co-receptor- and CD4-binding sites
Supplementary Information for Heavy chain-only IgG2b-llama antibody effects near-pan HIV-1 neutralization by recognizing a CD4-induced epitope that includes elements of co-receptor- and CD4-binding sites
Molecular Dynamics of HIV-1 Reverse Transcriptase
 Molecular Dynamics of HIV-1 Reverse Transcriptase Abderrahmane Benghanem Rensselaer Polytechnic Institute,Troy, NY Mentor: Dr. Maria Kurnikova Carnegie Mellon, Pittsburgh PA Outline HIV-1 Reverse Transcriptase
Molecular Dynamics of HIV-1 Reverse Transcriptase Abderrahmane Benghanem Rensselaer Polytechnic Institute,Troy, NY Mentor: Dr. Maria Kurnikova Carnegie Mellon, Pittsburgh PA Outline HIV-1 Reverse Transcriptase
BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached
 BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached 1). 20 points total T or F (2 points each; if false, briefly state why
BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached 1). 20 points total T or F (2 points each; if false, briefly state why
Supplementary Materials. High affinity binding of phosphatidylinositol-4-phosphate. by Legionella pneumophila DrrA
 Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Biology Open (2014) 000, 1 10 doi: /bio
 (2014) 000, 1 10 doi:10.1242/bio.201410041 Supplementary Material Michael Brauchle et al. doi: 10.1242/bio.201410041 Fig. S1. Alignment of GFP, sfgfp, egfp, eyfp, mcherry and mruby2. Sequence-based alignment
(2014) 000, 1 10 doi:10.1242/bio.201410041 Supplementary Material Michael Brauchle et al. doi: 10.1242/bio.201410041 Fig. S1. Alignment of GFP, sfgfp, egfp, eyfp, mcherry and mruby2. Sequence-based alignment
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Introduction to Protein Structure Collection
 Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Supplemental Information. An Atlas of b-glucuronidases. in the Human Intestinal Microbiome
 Structure, Volume 2 Supplemental Information An Atlas of b-glucuronidases in the Human Intestinal Microbiome Rebecca M. Pollet, Emma H. D'Agostino, William G. Walton, Yongmei Xu, Michael S. Little, Kristen
Structure, Volume 2 Supplemental Information An Atlas of b-glucuronidases in the Human Intestinal Microbiome Rebecca M. Pollet, Emma H. D'Agostino, William G. Walton, Yongmei Xu, Michael S. Little, Kristen
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/471/eaah5085/dc1 Supplementary Materials for Phosphorylation of the exocyst protein Exo84 by TBK1 promotes insulin-stimulated GLUT4 trafficking Maeran Uhm,
www.sciencesignaling.org/cgi/content/full/10/471/eaah5085/dc1 Supplementary Materials for Phosphorylation of the exocyst protein Exo84 by TBK1 promotes insulin-stimulated GLUT4 trafficking Maeran Uhm,
SUPPLEMENTARY INFORMATION
 Supplementary discussion of the role of C05 HCDR1 in antigen binding. Deletion of the 5 amino-acid insertion in HCDR1 del(fgest) and scanning mutagenesis of the tip of the loop had differing effects on
Supplementary discussion of the role of C05 HCDR1 in antigen binding. Deletion of the 5 amino-acid insertion in HCDR1 del(fgest) and scanning mutagenesis of the tip of the loop had differing effects on
T Cell Receptor Optimized Peptide Skewing of the T-Cell Repertoire Can Enhance Antigen Targeting
 T Cell Receptor Optimized Peptide Skewing of the T-Cell Repertoire Can Enhance Antigen Targeting Julia Ekeruche-Makinde 1*, Mathew Clement 1*, David K Cole 1*, Emily S J Edwards 1, Kristin Ladell 1, John
T Cell Receptor Optimized Peptide Skewing of the T-Cell Repertoire Can Enhance Antigen Targeting Julia Ekeruche-Makinde 1*, Mathew Clement 1*, David K Cole 1*, Emily S J Edwards 1, Kristin Ladell 1, John
Chapter VI. Increased affinity in the complex of yeast cytochrome c and cytochrome c peroxidase
 Chapter VI Increased affinity in the complex of yeast cytochrome c and cytochrome c peroxidase Chapter VI Abstract We report the study of T12A Cyt c CcP complex using isothermal titration calorimetry (ITC),
Chapter VI Increased affinity in the complex of yeast cytochrome c and cytochrome c peroxidase Chapter VI Abstract We report the study of T12A Cyt c CcP complex using isothermal titration calorimetry (ITC),
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction
 Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
This PDF file includes: Supplementary Figures 1 to 6 Supplementary Tables 1 to 2 Supplementary Methods Supplementary References
 Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Supplementary Figure 1. Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
 Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figures
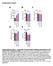 Supplementary Figures a b c d PDI activity in % ERp72 activity in % 4 3 2 1 1 1 ERp activity in % e ΔRFU min -1 1 1 ERp7 activity in % 1 1 Supplementary Figure 1. Selectivity of the bepristat-mediated
Supplementary Figures a b c d PDI activity in % ERp72 activity in % 4 3 2 1 1 1 ERp activity in % e ΔRFU min -1 1 1 ERp7 activity in % 1 1 Supplementary Figure 1. Selectivity of the bepristat-mediated
SUPPORTING INFORMATION. Lysine Carbonylation is a Previously Unrecognized Contributor. to Peroxidase Activation of Cytochrome c by Chloramine-T
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Crucial role for human Toll-like receptor 4 in the development of contact allergy to nickel
 Supplementary Figures 1-8 Crucial role for human Toll-like receptor 4 in the development of contact allergy to nickel Marc Schmidt 1,2, Badrinarayanan Raghavan 1,2, Verena Müller 1,2, Thomas Vogl 3, György
Supplementary Figures 1-8 Crucial role for human Toll-like receptor 4 in the development of contact allergy to nickel Marc Schmidt 1,2, Badrinarayanan Raghavan 1,2, Verena Müller 1,2, Thomas Vogl 3, György
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B
 3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
An esterase from anaerobic Clostridium hathewayi can hydrolyze aliphatic-aromatic polyesters
 Supporting Information An esterase from anaerobic Clostridium hathewayi can hydrolyze aliphatic-aromatic polyesters Veronika Perz a, Altijana Hromic b,c, Armin Baumschlager b, Georg Steinkellner b, Tea
Supporting Information An esterase from anaerobic Clostridium hathewayi can hydrolyze aliphatic-aromatic polyesters Veronika Perz a, Altijana Hromic b,c, Armin Baumschlager b, Georg Steinkellner b, Tea
Structural analysis of fungus-derived FAD glucose dehydrogenase
 Structural analysis of fungus-derived FAD glucose dehydrogenase Hiromi Yoshida 1, Genki Sakai 2, Kazushige Mori 3, Katsuhiro Kojima 3, Shigehiro Kamitori 1, and Koji Sode 2,3,* 1 Life Science Research
Structural analysis of fungus-derived FAD glucose dehydrogenase Hiromi Yoshida 1, Genki Sakai 2, Kazushige Mori 3, Katsuhiro Kojima 3, Shigehiro Kamitori 1, and Koji Sode 2,3,* 1 Life Science Research
Figure S1. HP1α localizes to centromeres in mitosis and interacts with INCENP. (A&B) HeLa
 SUPPLEMENTARY FIGURES Figure S1. HP1α localizes to centromeres in mitosis and interacts with INCENP. (A&B) HeLa tet-on cells that stably express HP1α-CFP, HP1β-CFP, or HP1γ-CFP were monitored with livecell
SUPPLEMENTARY FIGURES Figure S1. HP1α localizes to centromeres in mitosis and interacts with INCENP. (A&B) HeLa tet-on cells that stably express HP1α-CFP, HP1β-CFP, or HP1γ-CFP were monitored with livecell
172R 172K TAM-2/172R TAM-2/172K. AZT concentration [nm] AZT concentration [nm] MgCl 2 2.5K 2.5K 5K 2.5K 5K 2.5K K 5K 2.5K 5K 2.5K 50 2.
![172R 172K TAM-2/172R TAM-2/172K. AZT concentration [nm] AZT concentration [nm] MgCl 2 2.5K 2.5K 5K 2.5K 5K 2.5K K 5K 2.5K 5K 2.5K 50 2. 172R 172K TAM-2/172R TAM-2/172K. AZT concentration [nm] AZT concentration [nm] MgCl 2 2.5K 2.5K 5K 2.5K 5K 2.5K K 5K 2.5K 5K 2.5K 50 2.](/thumbs/82/85138523.jpg) 5 5 5 5 A MgCl 2 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] B 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] ATP + ATP - Supplemental Figure 1. Primer extension of HIV-1 RT polymorphisms
5 5 5 5 A MgCl 2 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] B 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] ATP + ATP - Supplemental Figure 1. Primer extension of HIV-1 RT polymorphisms
Proteins? Protein function. Protein folding. Protein folding diseases. Protein interactions. Macromolecular assemblies. The end product of Genes
 Proteins? Protein function Protein folding Protein folding diseases Protein interactions Macromolecular assemblies The end product of Genes Protein Unfolding DOD Acid Catalysis DOD HDOD + N H N D C N C
Proteins? Protein function Protein folding Protein folding diseases Protein interactions Macromolecular assemblies The end product of Genes Protein Unfolding DOD Acid Catalysis DOD HDOD + N H N D C N C
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Krenn et al., http://www.jcb.org/cgi/content/full/jcb.201110013/dc1 Figure S1. Levels of expressed proteins and demonstration that C-terminal
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Krenn et al., http://www.jcb.org/cgi/content/full/jcb.201110013/dc1 Figure S1. Levels of expressed proteins and demonstration that C-terminal
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Structural biology of viruses
 Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR
 Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR and LRRK2 WD40 GST fusion proteins (5 µg) were loaded
Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR and LRRK2 WD40 GST fusion proteins (5 µg) were loaded
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Information
 Supplementary Information Using the pimeloyl-coa synthetase adenylation fold to synthesise fatty acid thioesters Menglu Wang 1a ; Lucile Moynié 2a, Peter J. Harrison 1, Van Kelly 1, Andrew Piper 1, James
Supplementary Information Using the pimeloyl-coa synthetase adenylation fold to synthesise fatty acid thioesters Menglu Wang 1a ; Lucile Moynié 2a, Peter J. Harrison 1, Van Kelly 1, Andrew Piper 1, James
BMC Biology. Open Access. Abstract. BioMed Central
 BMC Biology BioMed Central Research article A modular interface of IL-4 allows for scalable affinity without affecting specificity for the IL-4 receptor Michael Kraich 1, Markus Klein 1, Edwin Patiño 1,
BMC Biology BioMed Central Research article A modular interface of IL-4 allows for scalable affinity without affecting specificity for the IL-4 receptor Michael Kraich 1, Markus Klein 1, Edwin Patiño 1,
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
SUPPLEMENTARY INFORMATION
 Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
on Non-Consensus Protein Motifs Analytical & Formulation Sciences, Amgen. Seattle, WA
 N-Linked Glycosylation on Non-Consensus Protein Motifs Alain Balland Analytical & Formulation Sciences, Amgen. Seattle, WA CASSS - Mass Spec 2010 Marina Del Rey, CA. September 8 th, 2010 Outline 2 Consensus
N-Linked Glycosylation on Non-Consensus Protein Motifs Alain Balland Analytical & Formulation Sciences, Amgen. Seattle, WA CASSS - Mass Spec 2010 Marina Del Rey, CA. September 8 th, 2010 Outline 2 Consensus
SUPPLEMENTARY INFORMATION
 Supplementary Discussion The HGGG sequence motif forms an oxyanion hole in HLSs. In the case of GID1, the third Gly residue of the sequence motif is replaced with Ser (Ser123 in OsGID1), i.e., the sequence
Supplementary Discussion The HGGG sequence motif forms an oxyanion hole in HLSs. In the case of GID1, the third Gly residue of the sequence motif is replaced with Ser (Ser123 in OsGID1), i.e., the sequence
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/9/e1600292/dc1 Supplementary Materials for Native phasing of x-ray free-electron laser data for a G protein coupled receptor Alexander Batyuk, Lorenzo Galli,
advances.sciencemag.org/cgi/content/full/2/9/e1600292/dc1 Supplementary Materials for Native phasing of x-ray free-electron laser data for a G protein coupled receptor Alexander Batyuk, Lorenzo Galli,
Insights into the Giardia intestinalis Enolase and Human Plasminogen interaction
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Supplementary Information Insights into the Giardia intestinalis Enolase and Human
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Supplementary Information Insights into the Giardia intestinalis Enolase and Human
Supporting Information. Lysine Propionylation to Boost Proteome Sequence. Coverage and Enable a Silent SILAC Strategy for
 Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Chapter 6. X-ray structure analysis of D30N tethered HIV-1 protease. dimer/saquinavir complex
 Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
Integrin v 3 targeted therapy for Kaposi s sarcoma with an in vitro evolved antibody 1
 Integrin v 3 targeted therapy for Kaposi s sarcoma with an in vitro evolved antibody 1 CHRISTOPH RADER, 2 MIKHAIL POPKOV, JOHN A. NEVES, AND CARLOS F. BARBAS III 2 Department of Molecular Biology and The
Integrin v 3 targeted therapy for Kaposi s sarcoma with an in vitro evolved antibody 1 CHRISTOPH RADER, 2 MIKHAIL POPKOV, JOHN A. NEVES, AND CARLOS F. BARBAS III 2 Department of Molecular Biology and The
The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation
 www.bioinformation.net Hypothesis Volume 8(9) The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation Zahra Karimi 1 *,
www.bioinformation.net Hypothesis Volume 8(9) The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation Zahra Karimi 1 *,
Supporting Information. Evolution of atomically precise silver clusters to superlattices
 Copyright WILEY-VCH Verlag GmbH & Co. KGaA, 69469 Weinheim, Germany, 2012. Supporting Information for Part. Part. Sys. Charact., DOI: 10.1002/ppsc.((please add manuscript number)) Evolution of atomically
Copyright WILEY-VCH Verlag GmbH & Co. KGaA, 69469 Weinheim, Germany, 2012. Supporting Information for Part. Part. Sys. Charact., DOI: 10.1002/ppsc.((please add manuscript number)) Evolution of atomically
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
Thesis by. Joshua Simon Klein. In Partial Fulfillment of the Requirements. for the Degree of. Doctor of Philosophy
 INVESTIGATIONS IN THE DESIGN AND CHARACTERIZATION OF HIV-1 NEUTRALIZING MOLECULES Thesis by Joshua Simon Klein In Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy California
INVESTIGATIONS IN THE DESIGN AND CHARACTERIZATION OF HIV-1 NEUTRALIZING MOLECULES Thesis by Joshua Simon Klein In Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy California
N α -Acetylation of yeast ribosomal proteins and its effect on protein synthesis
 JOURNAL OF PROTEOMICS 74 (2011) 431 441 available at www.sciencedirect.com www.elsevier.com/locate/jprot N α -Acetylation of yeast ribosomal proteins and its effect on protein synthesis Masahiro Kamita
JOURNAL OF PROTEOMICS 74 (2011) 431 441 available at www.sciencedirect.com www.elsevier.com/locate/jprot N α -Acetylation of yeast ribosomal proteins and its effect on protein synthesis Masahiro Kamita
Supplemental Data. Integrating omics and alternative splicing i reveals insights i into grape response to high temperature
 Supplemental Data Integrating omics and alternative splicing i reveals insights i into grape response to high temperature Jianfu Jiang 1, Xinna Liu 1, Guotian Liu, Chonghuih Liu*, Shaohuah Li*, and Lijun
Supplemental Data Integrating omics and alternative splicing i reveals insights i into grape response to high temperature Jianfu Jiang 1, Xinna Liu 1, Guotian Liu, Chonghuih Liu*, Shaohuah Li*, and Lijun
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH
 EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
Imaging energy status in live cells with a fluorescent biosensor of the intracellular ATP-to-ADP. ratio
 Imaging energy status in live cells with a fluorescent biosensor of the intracellular ATP-to-ADP ratio Mathew Tantama, Juan Ramón Martínez-François, Rebecca Mongeon, Gary Yellen* Department of Neurobiology,
Imaging energy status in live cells with a fluorescent biosensor of the intracellular ATP-to-ADP ratio Mathew Tantama, Juan Ramón Martínez-François, Rebecca Mongeon, Gary Yellen* Department of Neurobiology,
Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex
 John Clizer & Greg Ralph Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex The turkey ovomucoid third domain (OMTKY3) is considered to be one of the most studied protein inhibitors. 1 Ovomucin
John Clizer & Greg Ralph Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex The turkey ovomucoid third domain (OMTKY3) is considered to be one of the most studied protein inhibitors. 1 Ovomucin
Nature Immunology: doi: /ni.3631
 Supplementary Figure 1 SPT analyses of Zap70 at the T cell plasma membrane. (a) Total internal reflection fluorescent (TIRF) excitation at 64-68 degrees limits single molecule detection to 100-150 nm above
Supplementary Figure 1 SPT analyses of Zap70 at the T cell plasma membrane. (a) Total internal reflection fluorescent (TIRF) excitation at 64-68 degrees limits single molecule detection to 100-150 nm above
Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector
 kda 1 2 3 4 5 26 14 1 7 Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector 5 4 35 25 15 1 Supplementary Figure 1. SDS-PAGE of acterially expressed and purified proteins.
kda 1 2 3 4 5 26 14 1 7 Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector 5 4 35 25 15 1 Supplementary Figure 1. SDS-PAGE of acterially expressed and purified proteins.
Supplementary Information for Structural basis for the inhibition of Polo-like kinase 1
 Supplementary Information for Structural basis for inhibition of Polo-like kinase 1 Jun Xu, Chen Shen, Tao Wang & Junmin Quan Correspondence authors: Tao Wang, taowang@pkusz.edu.cn; Junmin Quan, quanjm@szpku.edu.cn
Supplementary Information for Structural basis for inhibition of Polo-like kinase 1 Jun Xu, Chen Shen, Tao Wang & Junmin Quan Correspondence authors: Tao Wang, taowang@pkusz.edu.cn; Junmin Quan, quanjm@szpku.edu.cn
Meaning-based guidance of attention in scenes as revealed by meaning maps
 SUPPLEMENTARY INFORMATION Letters DOI: 1.138/s41562-17-28- In the format provided by the authors and unedited. -based guidance of attention in scenes as revealed by meaning maps John M. Henderson 1,2 *
SUPPLEMENTARY INFORMATION Letters DOI: 1.138/s41562-17-28- In the format provided by the authors and unedited. -based guidance of attention in scenes as revealed by meaning maps John M. Henderson 1,2 *
Structural insights into xenobiotic and inhibitor binding to human Aldehyde Oxidase
 Suppementary Information Structura insights into xenobiotic and inhibitor binding to human Adehyde Oxidase Catarina Coeho 1, Aessandro Foti 2, Tobias Hartmann 2, Teresa Santos-Siva 1, Sike Leimküher 2
Suppementary Information Structura insights into xenobiotic and inhibitor binding to human Adehyde Oxidase Catarina Coeho 1, Aessandro Foti 2, Tobias Hartmann 2, Teresa Santos-Siva 1, Sike Leimküher 2
