Supplemental Information. An Atlas of b-glucuronidases. in the Human Intestinal Microbiome
|
|
|
- Amberly Stevens
- 5 years ago
- Views:
Transcription
1 Structure, Volume 2 Supplemental Information An Atlas of b-glucuronidases in the Human Intestinal Microbiome Rebecca M. Pollet, Emma H. D'Agostino, William G. Walton, Yongmei Xu, Michael S. Little, Kristen A. Biernat, Samuel J. Pellock, Loraine M. Patterson, Benjamin C. Creekmore, Hanna N. Isenberg, Rohini R. Bahethi, Aadra P. Bhatt, Jian Liu, Raad Z. Gharaibeh, and Matthew R. Redinbo
2 Figure S1A. Loop Category Variability. Related to Table 1, Table % Zero L1 GUS Genes 66.2% 1 or more L1 GUS Genes 33.8% % of individuals have 1 L1 GUS 46 Number Individuals with this Number of L1 GUS Genes % % 1 Average 1.2 +/- 1.2 L1 GUS Genes 2.8% 2.2% Number L1 GUS Genes
3 Figure S1B. Loop Category Variability. Related to Table 1, Table % Zero ml1 GUS Genes 98.6% 1 or more ml1 GUS Genes 2.1% of individuals have 3 ml1 GUS 28 Number Individuals with this Numbe rof ml1 Genes % 2 6.% 9 1.8% 1 1.8% % % % Number ml1 GUS Genes Average 4.4 +/- 2.2 ml1 GUS Genes 3.6% 2.9% 4 1.4% 2.1% 1
4 Figure S1C. Loop Category Variability. Related to Table 1, Table % Zero L2 GUS Genes 92.1% 1 or more L2 GUS Genes Number Individuals with this Number of L2 GUS Genes % % % % of individuals have 3 L2 GUS % Number L2 GUS Genes Average 2. +/- 1.6 L2 GUS Genes 7.2% 1 2.2% 3.7% 1
5 Figure S1D. Loop Category Variability. Related to Table 1, Table % Zero ml2 GUS Genes 8.6% 1 or more ml2 GUS Genes Number Individuals with this Number of ml2 GUS Genes % % of individuals have 1 ml2 GUS % 39 Average 1. +/- 1.1 ml2 GUS Genes 17.3% % Number ml2 GUS Genes
6 Figure S1E. Loop Category Variability. Related to Table 1, Table 2. 76% Zero ml1ml2 GUS Genes 76% 1 24% 1 or more ml1ml2 GUS Genes Number Individuals with this Number of ml1,2 GUS Genes Average.2 +/-. ml1,2 GUS Genes 23% of individuals have 1 ml1ml2 GUS % Number ml1,2 GUS Genes x
7 Figure S1F. Loop Category Variability. Related to Table 1, Table 2. 1% 1 or more nl GUS Genes 14 1% of individuals have 1 nl GUS 14 Number Individuals with this Number of NL GUS Genes % 2 3.6% 7.9% 11 Average 1.7 +/- 4.6 nl GUS Genes 2.2% 3.72% Number NL GUS Genes
8 Figure S1G. Loop Category Variability. Related to Table 1, Table 2. 41% Zero nc GUS Genes 7 9% 1 or more nc GUS Genes Number Individuals with this Number of NL GUS Genes % 4 1.1% of individuals have 2 nc GUS 21 6.% Number NL GUS Genes 3.6% Average 1. +/- 1.2 nc GUS Genes 1.4% 2
9 Figure S2. Summary of Loop Variability. Related to Table 2. Average Percent Composition of Abundance Reads % 21% 14% 7.1% 3.1% 4.6% 1.% L1 ml1 L2 ml2 ml1ml2 NL NC Loop Category
10 Figure S3. GUS Panel Sequence Identities. Related to Figure. Pairwise Sequence Identities for GUS Panel Proteins Ec Ee Fp Bf Bu Pm Bo Bd Ec Ee Fp Bf Bu Pm Bo 31 Bd
11 Figure S4A. GUS Panel ph Optima. Related to Figure. 3 3 E. coli GUS Optimal ph 7.4 PNP Generated (nmoles) ph 4 ph 4. ph ph. ph 6 ph 6. ph 7 ph Time (minutes)
12 Figure S4B. GUS Panel ph Optima. Related to Figure. 3 Eubacterium eligens GUS Optimal ph 6. 2 PNP Generated (nmoles) ph 4 ph 4. ph ph. ph 6 ph 6. ph 7 ph Time (minutes)
13 Figure S4C. GUS Panel ph Optima. Related to Figure. 4 Facaelibacterium prausnitzii GUS 3 Optimal ph 6 PNP Generated (nmoles) ph 4. ph 4. ph. ph. ph 6. ph 6. ph 7. ph Time (minutes)
14 Figure S4D. GUS Panel ph Optima. Related to Figure Bacteroides fragilis GUS Optimal ph PNP Generated (nmoles) ph 4 ph 4. ph ph. ph 6 ph 6. ph 7 ph Time (minutes)
15 Figure S4E. GUS Panel ph Optima. Related to Figure. 2 2 Bacteroides uniformis GUS Optimal ph 6 PNP Generated (nmoles) 1 1 ph 4 ph 4. ph ph. ph 6 ph 6. ph 7 ph Time (minutes)
16 Figure S4F. GUS Panel ph Optima. Related to Figure Parabacteroides merdae GUS Optimal ph. PNP Generated (nmoles) ph4. ph 4. ph. ph. ph 6. ph 6. ph 7. ph Time (minutes)
17 Figure S4G. GUS Panel ph Optima. Related to Figure. 6 Bacteroides ovatus GUS Optimal ph PNP Generated (nmoles) ph 4. ph 4. ph. ph. ph 6. ph 6. ph 7. ph Time (minutes)
18 Figure S4H. GUS Panel ph Optima. Related to Figure. 3 2 Bacteroides dorei GUS Optimal ph 6 PNP Generated (nmoles) ph 4. ph 4. ph. ph. ph 6. ph 6. ph 7. ph Time (min)
19 Figure SA. Heparan Carbohydrate Cleavage by GUS Panel. Related to Figure. GlcUA-GlcNAc-GlcUA-GlcNAc-GlcUA-GlcNAc-GlcUA-GlcNAc-GlcUA-PNP, 9-mer (MW = 1832.) GlcUA b-glucuronidase (GUS) GlcNAc-GlcUA-GlcNAc-GlcUA-GlcNAc-GlcUA-GlcNAc-GlcUA-PNP, 8-mer (MW = 166.4)
20 Figure SB. Heparan Carbohydrate Cleavage by GUS Panel. Related to Figure. Representative Representative HPLC chromatograms and ESI-MS spectra of nonasaccharide digestion HPLC chromatograms and ESI-MS spectra of 9-mer substrate digested by β-glucuronidases Representative HPLC chromatograms and ESI-MS spectra of of 9-mer substrate digested by by β-glucuronidases epresentative HPLC chromatograms and ESI-MS spectra of 9-mer substrate digested by β-glucuronidases 4e+ 4e+ 4e+ 4e+ 3e+ O.D. 31 nm O.D. 31 nm O.D. O.D nm nm 3e+ 3e+ 3e+ 2e+ 2e+ 2e+ 2e+ 1e+ 1e+ 1e+ 1e+ HPLC of uncleaved reaction 9-mer (e.g., Bo, Ec, Ee, Fp) 9-mer 9-mer 9-mer 9-mer Retention time (min) Retention time time (min) ESI-MS of uncleaved reactions (e.g., Bo,Ec,Ee,Fp) O.D. O.D. 31nm 31nm O.D. 31nm O.D. O.D. 31nm 31nm HPLC of partially cleaved reaction (e.g., Bd, Bf, Bu) 9-mer Relative abundance (%) Relative abundance (%) Relative Relative abundance abundance (%) (%) HPLC of fully cleaved reaction 8-mer (e.g., Pm) Retention time (min) Retention Retention time time (min) (min) Retention time (min) HPLC of uncleaved reaction (Bo,Ec,Ee,Fp) HPLC of partially cleaved reaction (Bd,Bf,Bu) HPLC of cleaved reaction (Pm) HPLC of of of uncleaved uncleavedreaction (Bo,Ec,Ee,Fp) HPLC HPLC of of partially cleaved reaction (Bd,Bf,Bu) HPLC of of cleaved reaction (Pm) 9mer 1 z= mer 9mer 1 9mer mer z=3 z=3 z=3 z= mer 86 69mer9mer z=4 z=4 z= Relative abundance (%) Relative abundance (%) Relative abundance (%) Relative Relative abundance abundance (%) (%) mer z= mer 9mer 9mer z=2 z=2 z= m/z Mass of uncleaved m/z reaction m/z (Bo,Ec,Ee,Fp) m/z Mass of uncleaved reaction (Bo,Ec,Ee,Fp) 2.e+ 2.e+ 2.e+ 2.e+ 2.e+ 2.e+ 2.e+ 2.e+ 1.e+ 1.e+ 1.e+ 1.e+ 1.e+ 1.e+ 1.e+ 1.e+.e+4.e+4.e Retention time (min) Retention time (min) 1 9mer 9mer z=3 z= mer mer 9mer 9mer z=3 z=4 z=3 z= z= z= z=3 z= z=4 1.2 z=4 z= Relative abundance (%) Relative abundance (%) Relative Relative abundance abundance (%) (%) 9-mer 8-mer 9-mer 9-mer 9-mer 8-mer 8-mer 8-mer ESI-MS of partially cleaved reactions (e.g., Bd, Bf, Bu) z=2 9mer z= z=2 9mer z=2 9mer z= z= Mass of partially m/zcleaved reaction m/z m/z (Bd,Bf,Bu) 4e+ 4e+ 4e+ 4e+ 3e+ 3e+ 3e+ 3e+ 2e+ O.D 31 nm O.D O.D nm nm O.D O.D nm nm 2e+ 2e+ 2e+ 1e+ 1e+ 1e+ 1e m/z Retention time (min) Retention time time (min) (min) 6 z= z=4 4 z= mer 8-mer 8-mer 8-mer ESI-MS of fully cleaved reaction (e.g., Pm) z=3 1.2 z=3 z= z= z=2 z= m/z m/z Mass of cleaved m/zreaction (Pm)
21 Supplemental Table 3. Crystallographic Statistics for B. uniformis GUS Resolution (highest shell; Å) ( ) Space group P Unit cell dimensions (a, b, c; Å) 74.7, 142.9, Total reflections (F>) 1,668,9 Unique reflections 12,73 Multiplicity 1.9 (11.) Completeness, % (highest shell) 99.4 (98.7) Mean I/sigma(I) (highest shell) 21.1 (.) Wilson B-factor (Å 2 ) 22.1 R merge (highest shell).68 (.448) R.144 R free.18 Number of protein, metal, ligand, and water atoms per asymmetric unit 13,6, 4, 36, 177 rms bonds (Å).11 rms angles ( ) 1.28 Ramachandran favored (%) 97. Ramachandran outliers (%) Clash score Average B-factor (Å 2 ) 3.2 RCSB ID UJ6
22 Table S4. Primer Sequences Primer Name Sequence EeGUS Fwd EeGUS Rev FpGUS Fwd FpGUS Rev BfGUS Fwd BfGUS Rev BuGUS Fwd BuGUS Rev PmGUS Fwd TACTTCCAATCCAATGCGATGCTGTACCCTGTCCTGACTCAGTCTC G TTATCCACTTCCAATGCGCTATTTGGTTTTGTAGCCAAATTCCGGA ATGGTGGACCAG TACTTCCAATCCAATGCGATGAACCGTAGCCTGCTGTACCCTCGTG TATCCACTTCCAATGCGCTTTTTTTGCGTTTTTTGAAGTCAACCG TACTTCCAATCCAATGCGCGTGAGGATATTCTACTCAACAATAACT GGAATTTCCGTTTT TTATCCACTTCCAATGCGCTATAGGGGTCTGTCTTCTTTATATTCTA CTGTCACTTCGTC TACTTCCAATCCAATGCGCAACG TCAAACTCAAACCATCAACGACTCCTGGAAG TTATCCACTTCCAATGCGCTAATAAATGTTGCGCAGTTTAATGCCAT TCAGGAAACACG TACTTCCAATCCAATGCGATGAAATACCTGTTCGTCGCTTGCCTGC TGTG PmGUS Rev TTATCCACTTCCAATGCGCTATTTCGGGCTTTTCAGTTCCAGAAAC GCGGTC BoGUS Fwd BoGUS Rev BdGUS Fwd BdGUS Rev TACTTCCAATCCAATGCGCAGGAAACTTCTCCGCGTACTATCTTCT CTCTGAAC TTATCCACTTCCAATGCGCTATTTAATGCAGTACCATTCGCAGGTG TCGGACAG TACTTCCAATCCAATGCGAGCGAGATCAGCATCACTGACTCTTGGA AGTATAAG TTATCCACTTCCAATGCGCTAGCGAATTTTTTTAACTTTCAGGCCG CTAATTACGCCCTG
23 SUPPLEMENTAL FIGURE, TABLE, AND DATA FILE LEGENDS Figure S1. Loop Category Variability. Related to Table 1, Table 2. For the HMGI313 GUS analysis, the number of individuals of each loop category is presented, with annotations to indicate averages and percent of individuals with no GUS enzymes of that category, and the averages over all individuals. Panels A-F present this information for the six formal GUS categories (Loop 1, Mini-Loop 1, Loop 2, Mini- Loop 2, Mini-Loop 1,2, No Loop, respectively), while panel G presents this information for the no content GUS proteins. Figure S2. Summary of Loop Variability. Related to Table 2. Bar graph of the average percent composition of abundance reads for the HMGI313 dataset within each loop category, with standard deviations of the averages indicated. Figure S3. GUS Panel Sequence Identities. Related to Figure. Pairwise sequence identities in percent for the GUS panel proteins from E. coli (Ec), Eubacterium eligens (Ee), Faecalibacterium prausnitzii (Fp), Bacteroides fragilis (Bf), B. uniformis (Bu), Parabacteroides merdae (Pm), B. ovatus (Bo), and B. dorei (Bd). Each enzyme s identifier is colored by loop category: red, Loop 1, green, Mini-Loop 1; blue, Loop 2; orange, Mini-Loop 2; magenta, Mini-Loop 1,2; purple, No Loop. Figure S4. GUS Panel ph Optima. Related to Figure. Results for amount of PNP produced from PNPG per minute at the eight indicated ph values are presented for each of the eight enzymes in the GUS panel, and colored by loop category as outlined in Figure S3 above. Amount of PNP Generated is shown as the average and standard deviation over three replicates. The ph optimum selected for each GUS is also indicated. Figure S. Heparan Carbohydrate Cleavage by GUS Panel. Related to Figure. A. Chemical structures and molecular weights of heparan nonasaccharaide (9-mer) assay substrate and octasaccharide (8-mer) product detected by both HPLC and ESI- MS, where GlcUA is glucuronic acid, GlcNAc is N-acetylglucosamine, and PNP is p- nitrophenol. B. Representative HPLC chromatograms and ESI-MS spectra for heparan digestion assays with the GUS panel. HPLC results (top) are shown for uncleaved (left), partially cleaved (middle), and fully cleaved (right) reactions. Electrospray ionization mass spectrometry (ESI-MS) spectra are shown for the same types of reactions (left, middle, center). GUS enzymes from the following organisms were examined: Bo, B. ovatus, Ec, E. coli, Ee, Eu. eligens, Fp, F. prausnitzii, Bd, B. dorei, Bf, B. fragilis, Bu, B. uniformis, Pm, P. merdae. Table S1. HMGI313 Characteristics. Related to Table 1, Table 2. Detailed information is provided for all 313 GUS proteins identified in the HMGI database, including the sequences of their Loop 1 and Loop 2 regions, protein length in amino acid residues, read abundances, and taxonomy along with BLAST hit characteristics.
24 Table S2. HMGC279 Characteristics. Related to Table 1, Figure 3. Detailed information is provided for the 279 GUS proteins identified in the HMGC Clustered Gene Indices database of the HMP, including the sequences of their Loop 1 and Loop 2 regions, protein length in amino acid residues, taxonomy along with BLAST hit characteristics, and signal peptide search results. Table S3. BuGUS Crystallographic Analysis. Related to Figure. X-ray data collection, structure determination, and structure refinement information are provided for the crystal structure of the Loop 2 GUS from B. uniformis (BuGUS). Table S4. Primer Sequences. Related to Figure. Forward and reverse primers employed in this study. Data File S1. HMGI313 Sequences. Related to Figure 1. FASTA sequences for the 313 total GUS proteins identified in the HMGI Gene Indices database, part of the Stool Sample Catalog of the Human Microbiome Project (HMP). Entries are labeled by their HMP identifier. Data File S2. HMGC279 Sequences. Related to Figure 1, Figure 3, Table 1. FASTA sequences for the 279 unique GUS proteins identified in the HMGC Clustered Gene Indices database, part of the Stool Sample Catalog of the Human Microbiome Project (HMP). Entries are labeled by their HMP identifier, along with the GUS category it belongs to (L1, ml1, L2, ml2, ml1,2, NL, nc), and taxonomy when identified. Data File S3. Representative GUS Loop Alignment. Related to Figure 2. Multiple sequence alignment of the amino acid sequence of twelve representative GUS enzymes, with residue numbers indicated at right, and loop regions colored as follows: red, Loop 1; green, Mini-Loop 1; blue, Loop 2; orange, Mini-Loop 2; magenta, Mini-Loop 1, 2; purpe, No Loop. The twelve GUS enzymes are from Homo sapiens (HsGUS), Faecalibacterium prausnitzii (FpGUS), E. coli (EcGUS), Streptococcus agalactiae (SaGUS), Clostridium perfringens (CpGUS), Eubacterium eligens (EeGUS), Parabacteroides merdae (PmGUS), a Firmicutes GUS described from human fecal metagenomic sequencing (H11G11-BG), Bacteroides fragilis (BfGUS), B. ovatus (BoGUS), B. uniformis (BuGUS), and B. dorei (BdGUS). Identical amino acid positions are indicated with an asterisk (*), while largely conserved and conserved amino acid positions are indicated with a colon (:) or period (.), respectively.
Table S1. X-ray data collection and refinement statistics
 Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Supporting Information
 Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Supporting Information
 Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
FCC2 5CY7 FCC1 5CY8. Actinonin 5CVQ
 Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/9/e1600292/dc1 Supplementary Materials for Native phasing of x-ray free-electron laser data for a G protein coupled receptor Alexander Batyuk, Lorenzo Galli,
advances.sciencemag.org/cgi/content/full/2/9/e1600292/dc1 Supplementary Materials for Native phasing of x-ray free-electron laser data for a G protein coupled receptor Alexander Batyuk, Lorenzo Galli,
Minutes Figure S1. HPLC separation of nucleosides from LC/ESI-MS analysis of a total enzymatic Trp
 100 A % Relative Abundance m m Ø acp 5 m Øm 5 m m 7 m s * 6 ia m * 0 5 10 15 0 5 0 5 40 Minutes Figure S1. HPL separation of nucleosides from L/ESI-MS analysis of a total enzymatic digest of mt trna. V
100 A % Relative Abundance m m Ø acp 5 m Øm 5 m m 7 m s * 6 ia m * 0 5 10 15 0 5 0 5 40 Minutes Figure S1. HPL separation of nucleosides from L/ESI-MS analysis of a total enzymatic digest of mt trna. V
An esterase from anaerobic Clostridium hathewayi can hydrolyze aliphatic-aromatic polyesters
 Supporting Information An esterase from anaerobic Clostridium hathewayi can hydrolyze aliphatic-aromatic polyesters Veronika Perz a, Altijana Hromic b,c, Armin Baumschlager b, Georg Steinkellner b, Tea
Supporting Information An esterase from anaerobic Clostridium hathewayi can hydrolyze aliphatic-aromatic polyesters Veronika Perz a, Altijana Hromic b,c, Armin Baumschlager b, Georg Steinkellner b, Tea
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12198 1. Supplementary results 1.1. Associations between gut microbiota, glucose control and medication Women with T2D who used metformin had increased levels of Enterobacteriaceae (i.e.
doi:10.1038/nature12198 1. Supplementary results 1.1. Associations between gut microbiota, glucose control and medication Women with T2D who used metformin had increased levels of Enterobacteriaceae (i.e.
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
High-yield novel leech hyaluronidase to expedite the. preparation of specific hyaluronan oligomers
 High-yield novel leech hyaluronidase to expedite the preparation of specific hyaluronan oligomers Peng Jin 1,2,3, Zhen Kang 1,2,3,6, Na Zhang 1,2,3, Guocheng Du 1,3,5,6 & Jian Chen 1,3,4,6 1 Synergetic
High-yield novel leech hyaluronidase to expedite the preparation of specific hyaluronan oligomers Peng Jin 1,2,3, Zhen Kang 1,2,3,6, Na Zhang 1,2,3, Guocheng Du 1,3,5,6 & Jian Chen 1,3,4,6 1 Synergetic
SUPPLEMENTARY INFORMATION FOR. (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation. by COX-2
 SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
Supporting Information
 Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
-Supplementary Information-
 -Supplementary Information- Probiotic Bifidobacterium longum alters gut luminal metabolism through modification of the gut microbial community Hirosuke Sugahara,2,3*, Toshitaka Odamaki, Shinji Fukuda 2,4,
-Supplementary Information- Probiotic Bifidobacterium longum alters gut luminal metabolism through modification of the gut microbial community Hirosuke Sugahara,2,3*, Toshitaka Odamaki, Shinji Fukuda 2,4,
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
This PDF file includes: Supplementary Figures 1 to 6 Supplementary Tables 1 to 2 Supplementary Methods Supplementary References
 Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Overview. The Microbiome: A New Lens for Human Disease. HMP Overarching Conclusions 10/2/2015. Human Microbiome Project
 Overview The Microbiome: A New Lens for Human Disease Matthew R Redinbo University of North Carolina at Chapel Hill Human Microbiome Project 1. Nasal Microbiota 2. Pulmonary Microbiota 3. Intestinal Microbiota
Overview The Microbiome: A New Lens for Human Disease Matthew R Redinbo University of North Carolina at Chapel Hill Human Microbiome Project 1. Nasal Microbiota 2. Pulmonary Microbiota 3. Intestinal Microbiota
The N-terminal loop of IRAK-4 death domain regulates ordered assembly of the Myddosome signalling scaffold
 The N-terminal loop of IRAK-4 death domain regulates ordered assembly of the Myddosome signalling scaffold Anthony C.G. Dossang * 1, Precious G. Motshwene * 1, Yang Yang* 1, Martyn F. Symmons*, Clare E.
The N-terminal loop of IRAK-4 death domain regulates ordered assembly of the Myddosome signalling scaffold Anthony C.G. Dossang * 1, Precious G. Motshwene * 1, Yang Yang* 1, Martyn F. Symmons*, Clare E.
Supplementary Information
 Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Supporting Information. Lysine Propionylation to Boost Proteome Sequence. Coverage and Enable a Silent SILAC Strategy for
 Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein
 Supplementary Methods Thermal shift assays Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein unfolding was examined by monitoring the fluorescence of ANS (1-anilinonaphthalene-8-
Supplementary Methods Thermal shift assays Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein unfolding was examined by monitoring the fluorescence of ANS (1-anilinonaphthalene-8-
Supplementary Information
 Supplementary Information Using the pimeloyl-coa synthetase adenylation fold to synthesise fatty acid thioesters Menglu Wang 1a ; Lucile Moynié 2a, Peter J. Harrison 1, Van Kelly 1, Andrew Piper 1, James
Supplementary Information Using the pimeloyl-coa synthetase adenylation fold to synthesise fatty acid thioesters Menglu Wang 1a ; Lucile Moynié 2a, Peter J. Harrison 1, Van Kelly 1, Andrew Piper 1, James
biochem480 [Spring 2018] Enzyme Bio-informatics project
![biochem480 [Spring 2018] Enzyme Bio-informatics project biochem480 [Spring 2018] Enzyme Bio-informatics project](/thumbs/95/122795322.jpg) biochem480 [Spring 2018] Enzyme Bio-informatics project Student Name: Alissa Burbridge Enzyme Name: Fructose-bisphosphate aldolase # of PDB entries for this enzyme: 129 PDB code: 4ALD E.C. # 4.1.2.13 Authors
biochem480 [Spring 2018] Enzyme Bio-informatics project Student Name: Alissa Burbridge Enzyme Name: Fructose-bisphosphate aldolase # of PDB entries for this enzyme: 129 PDB code: 4ALD E.C. # 4.1.2.13 Authors
MSSimulator. Simulation of Mass Spectrometry Data. Chris Bielow, Stephan Aiche, Sandro Andreotti, Knut Reinert FU Berlin, Germany
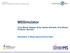 Chris Bielow Algorithmic Bioinformatics, Institute for Computer Science MSSimulator Chris Bielow, Stephan Aiche, Sandro Andreotti, Knut Reinert FU Berlin, Germany Simulation of Mass Spectrometry Data Motivation
Chris Bielow Algorithmic Bioinformatics, Institute for Computer Science MSSimulator Chris Bielow, Stephan Aiche, Sandro Andreotti, Knut Reinert FU Berlin, Germany Simulation of Mass Spectrometry Data Motivation
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
KDM2A. Reactions. containing. Reactions
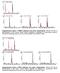 Supplementary Figure 1 KDM2A catalyses only lysine demethylation.. MALDI-TOF MS of demethylation of the shown variant histone peptides as catalysed by recombinant KDM2A. Reactions containing enzyme are
Supplementary Figure 1 KDM2A catalyses only lysine demethylation.. MALDI-TOF MS of demethylation of the shown variant histone peptides as catalysed by recombinant KDM2A. Reactions containing enzyme are
Figure S1. HP1α localizes to centromeres in mitosis and interacts with INCENP. (A&B) HeLa
 SUPPLEMENTARY FIGURES Figure S1. HP1α localizes to centromeres in mitosis and interacts with INCENP. (A&B) HeLa tet-on cells that stably express HP1α-CFP, HP1β-CFP, or HP1γ-CFP were monitored with livecell
SUPPLEMENTARY FIGURES Figure S1. HP1α localizes to centromeres in mitosis and interacts with INCENP. (A&B) HeLa tet-on cells that stably express HP1α-CFP, HP1β-CFP, or HP1γ-CFP were monitored with livecell
SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES
 SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors:
SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors:
Supplementary Information
 Supplementary Information Structural Basis of Sialidase in Complex with Geranylated Flavonoids as Potent Natural Inhibitors Youngjin Lee, a,b Young Bae Ryu, d Hyung-Seop Youn, a,b Jung Keun Cho, e Young
Supplementary Information Structural Basis of Sialidase in Complex with Geranylated Flavonoids as Potent Natural Inhibitors Youngjin Lee, a,b Young Bae Ryu, d Hyung-Seop Youn, a,b Jung Keun Cho, e Young
ect of Apple Pectin Oligosaccharide on Intestinal Disorders
 23 Nippon Shokuhin Kagaku Kogaku Kaishi Vol. //, No. +*,.//.0* (,**2) 455 E# ect of Apple Pectin Oligosaccharide on Intestinal Disorders Yoshinori Takahashi, Yasuyuki Masuda, Masahiro Sugimoto and Hiroyuki
23 Nippon Shokuhin Kagaku Kogaku Kaishi Vol. //, No. +*,.//.0* (,**2) 455 E# ect of Apple Pectin Oligosaccharide on Intestinal Disorders Yoshinori Takahashi, Yasuyuki Masuda, Masahiro Sugimoto and Hiroyuki
Shotgun metaproteomics of the human distal gut microbiota. Present by Lei Chen
 Shotgun metaproteomics of the human distal gut microbiota Present by Lei Chen (lc6@indana.edu) Outline Background What are the goals? Materials and Methods Results Discussion Background The human gastrointestinal
Shotgun metaproteomics of the human distal gut microbiota Present by Lei Chen (lc6@indana.edu) Outline Background What are the goals? Materials and Methods Results Discussion Background The human gastrointestinal
Manja Henze, Dorothee Merker and Lothar Elling. 1. Characteristics of the Recombinant β-glycosidase from Pyrococcus
 S1 of S17 Supplementary Materials: Microwave-Assisted Synthesis of Glycoconjugates by Transgalactosylation with Recombinant Thermostable β-glycosidase from Pyrococcus Manja Henze, Dorothee Merker and Lothar
S1 of S17 Supplementary Materials: Microwave-Assisted Synthesis of Glycoconjugates by Transgalactosylation with Recombinant Thermostable β-glycosidase from Pyrococcus Manja Henze, Dorothee Merker and Lothar
Gut Microbiomes of Malawian Twin Pairs Discordant for Kwashiorkor
 Gut Microbiomes of Malawian Twin Pairs Discordant for Kwashiorkor Michelle I. Smith et al. Science 339, 548 (2013) Dept Meeting, 28 May 2013, M. UMEZAKI ABSTRACT. Kwashiorkor, an enigmatic form of severe
Gut Microbiomes of Malawian Twin Pairs Discordant for Kwashiorkor Michelle I. Smith et al. Science 339, 548 (2013) Dept Meeting, 28 May 2013, M. UMEZAKI ABSTRACT. Kwashiorkor, an enigmatic form of severe
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
172R 172K TAM-2/172R TAM-2/172K. AZT concentration [nm] AZT concentration [nm] MgCl 2 2.5K 2.5K 5K 2.5K 5K 2.5K K 5K 2.5K 5K 2.5K 50 2.
![172R 172K TAM-2/172R TAM-2/172K. AZT concentration [nm] AZT concentration [nm] MgCl 2 2.5K 2.5K 5K 2.5K 5K 2.5K K 5K 2.5K 5K 2.5K 50 2. 172R 172K TAM-2/172R TAM-2/172K. AZT concentration [nm] AZT concentration [nm] MgCl 2 2.5K 2.5K 5K 2.5K 5K 2.5K K 5K 2.5K 5K 2.5K 50 2.](/thumbs/82/85138523.jpg) 5 5 5 5 A MgCl 2 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] B 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] ATP + ATP - Supplemental Figure 1. Primer extension of HIV-1 RT polymorphisms
5 5 5 5 A MgCl 2 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] B 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] ATP + ATP - Supplemental Figure 1. Primer extension of HIV-1 RT polymorphisms
SUPPORTING INFORMATION. Lysine Carbonylation is a Previously Unrecognized Contributor. to Peroxidase Activation of Cytochrome c by Chloramine-T
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Molecular Graphics Perspective of Protein Structure and Function
 Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
High-Throughput Analysis of Oligonucleotides using Automated Electrospray Ionization Mass Spectrometry
 High-Throughput Analysis of Oligonucleotides using Automated Electrospray Ionization Mass Spectrometry Mark E. Hail 1, Brian Elliott 2, Kerry Nugent 3, Jeffrey L. Whitney 1, and David J. Detlefsen 1 1
High-Throughput Analysis of Oligonucleotides using Automated Electrospray Ionization Mass Spectrometry Mark E. Hail 1, Brian Elliott 2, Kerry Nugent 3, Jeffrey L. Whitney 1, and David J. Detlefsen 1 1
Catalysis & specificity: Proteins at work
 Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
T Cell Receptor Optimized Peptide Skewing of the T-Cell Repertoire Can Enhance Antigen Targeting
 T Cell Receptor Optimized Peptide Skewing of the T-Cell Repertoire Can Enhance Antigen Targeting Julia Ekeruche-Makinde 1*, Mathew Clement 1*, David K Cole 1*, Emily S J Edwards 1, Kristin Ladell 1, John
T Cell Receptor Optimized Peptide Skewing of the T-Cell Repertoire Can Enhance Antigen Targeting Julia Ekeruche-Makinde 1*, Mathew Clement 1*, David K Cole 1*, Emily S J Edwards 1, Kristin Ladell 1, John
Microbial DNA qpcr Array Metabolic Disorders
 Microbial DNA qpcr Array Metabolic Disorders Cat. no. 330261 BAID-1406ZRA For real-time PCR-based, application-specific microbial identification or profiling The Metabolic Disorders Microbial DNA qpcr
Microbial DNA qpcr Array Metabolic Disorders Cat. no. 330261 BAID-1406ZRA For real-time PCR-based, application-specific microbial identification or profiling The Metabolic Disorders Microbial DNA qpcr
Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev
 Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev Institute for Biochemical Physics, Rus. Acad. Sci., Moscow, Russia. Institute for Energy Problems
Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev Institute for Biochemical Physics, Rus. Acad. Sci., Moscow, Russia. Institute for Energy Problems
Final Report. Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD. Wendy Fong ( 方美珍 ) CNIC; Beijing, China
 Final Report Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD Wendy Fong ( 方美珍 ) CNIC; Beijing, China Influenza s Viral Life Cycle 1. HA on virus binds to sialic
Final Report Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD Wendy Fong ( 方美珍 ) CNIC; Beijing, China Influenza s Viral Life Cycle 1. HA on virus binds to sialic
Crystallization-grade After D After V3 cocktail. Time (s) Time (s) Time (s) Time (s) Time (s) Time (s)
 Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Insights into the Giardia intestinalis Enolase and Human Plasminogen interaction
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Supplementary Information Insights into the Giardia intestinalis Enolase and Human
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Supplementary Information Insights into the Giardia intestinalis Enolase and Human
[ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS
![[ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS [ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS](/thumbs/79/80328542.jpg) Yun Wang Alelyunas, Henry Shion, Mark Wrona Waters Corporation, Milford, MA, USA APPLICATION BENEFITS mab LC-MS method which enables users to achieve highly sensitive bioanalysis of intact trastuzumab
Yun Wang Alelyunas, Henry Shion, Mark Wrona Waters Corporation, Milford, MA, USA APPLICATION BENEFITS mab LC-MS method which enables users to achieve highly sensitive bioanalysis of intact trastuzumab
Supporting InformationCathepsin-Mediated Cleavage of Peptides from Peptide Amphiphiles Leads to Enhanced Intracellular Peptide Accumulation
 Supporting InformationCathepsin-Mediated Cleavage of Peptides from Peptide Amphiphiles Leads to Enhanced Intracellular Peptide Accumulation Handan Acar, PhD,, Ravand Samaeekia, MS,, Mathew R. Schnorenberg,,
Supporting InformationCathepsin-Mediated Cleavage of Peptides from Peptide Amphiphiles Leads to Enhanced Intracellular Peptide Accumulation Handan Acar, PhD,, Ravand Samaeekia, MS,, Mathew R. Schnorenberg,,
Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid
 Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid Joanne ( 乔安妮 ) Mather Senior Scientist Waters Corporation Data courtesy of Erin Chambers and Mary
Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid Joanne ( 乔安妮 ) Mather Senior Scientist Waters Corporation Data courtesy of Erin Chambers and Mary
2200 GI Effects Comprehensive Profile Stool Interpretation At-a-Glance
 P: 1300 688 522 E: info@nutripath.com.au A: PO Box 442 Ashburton VIC 3142 TEST PATIENT Sample Test Name Sex : F Date Collected : 00-00-0000 111 TEST ROAD TEST SUBURB LAB ID: 00000000 UR#:0000000 TEST PHYSICIAN
P: 1300 688 522 E: info@nutripath.com.au A: PO Box 442 Ashburton VIC 3142 TEST PATIENT Sample Test Name Sex : F Date Collected : 00-00-0000 111 TEST ROAD TEST SUBURB LAB ID: 00000000 UR#:0000000 TEST PHYSICIAN
Nature Methods: doi: /nmeth.3177
 Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism
 Supplementary Information The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism Address correspondence to Yong Li (yongli@xmu.edu.cn, Tel: 86-592-218151) GW464 CDCA Supplementary
Supplementary Information The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism Address correspondence to Yong Li (yongli@xmu.edu.cn, Tel: 86-592-218151) GW464 CDCA Supplementary
User Guide. Protein Clpper. Statistical scoring of protease cleavage sites. 1. Introduction Protein Clpper Analysis Procedure...
 User Guide Protein Clpper Statistical scoring of protease cleavage sites Content 1. Introduction... 2 2. Protein Clpper Analysis Procedure... 3 3. Input and Output Files... 9 4. Contact Information...
User Guide Protein Clpper Statistical scoring of protease cleavage sites Content 1. Introduction... 2 2. Protein Clpper Analysis Procedure... 3 3. Input and Output Files... 9 4. Contact Information...
SMPD 287 Spring 2015 Bioinformatics in Medical Product Development. Final Examination
 Final Examination You have a choice between A, B, or C. Please email your solutions, as a pdf attachment, by May 13, 2015. In the subject of the email, please use the following format: firstname_lastname_x
Final Examination You have a choice between A, B, or C. Please email your solutions, as a pdf attachment, by May 13, 2015. In the subject of the email, please use the following format: firstname_lastname_x
CHM 424L Organic Laboratory, Dr. Laurie S. Starkey Introduction to Mass Spectrometry
 CM 424L rganic Laboratory, Dr. Laurie S. Starkey Introduction to Mass Spectrometry Mass spectrometry is used to determine a sample's molecular mass and molecular formula. Some structural information can
CM 424L rganic Laboratory, Dr. Laurie S. Starkey Introduction to Mass Spectrometry Mass spectrometry is used to determine a sample's molecular mass and molecular formula. Some structural information can
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry. Supporting Information
 Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan
 PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/0/493/eaal54/dc Supplementary Materials for A systems approach for discovering linoleic acid derivatives that potentially mediate pain and itch Christopher E.
www.sciencesignaling.org/cgi/content/full/0/493/eaal54/dc Supplementary Materials for A systems approach for discovering linoleic acid derivatives that potentially mediate pain and itch Christopher E.
Carnelian uncovers hidden functional patterns across diverse study populations from whole metagenome sequencing reads
 Carnelian uncovers hidden functional patterns across diverse study populations from whole metagenome sequencing reads Sumaiya Nazeen 1 and Bonnie Berger 1,2* 1 Computer Science and Artificial Intelligence
Carnelian uncovers hidden functional patterns across diverse study populations from whole metagenome sequencing reads Sumaiya Nazeen 1 and Bonnie Berger 1,2* 1 Computer Science and Artificial Intelligence
Stool Testing for the Microbiome - Ready for Primetime?
 Stool Testing for the Microbiome - Ready for Primetime? Gillian M. Barlow, PhD Project Scientist, Medically Associated Science and Technology (MAST) Program Cedars-Sinai Medical Center Take Charge of
Stool Testing for the Microbiome - Ready for Primetime? Gillian M. Barlow, PhD Project Scientist, Medically Associated Science and Technology (MAST) Program Cedars-Sinai Medical Center Take Charge of
Microbe-Host Interactions in Inflammatory Bowel Diseases. Hera Vlamakis Oct 3, 2018
 Microbe-Host Interactions in Inflammatory Bowel Diseases Hera Vlamakis Oct 3, 2018 Most of the bacteria in your body are in your gut HEALTH BENEFITS Breakdown of polysaccharides Synthesis of vitamins Colonization
Microbe-Host Interactions in Inflammatory Bowel Diseases Hera Vlamakis Oct 3, 2018 Most of the bacteria in your body are in your gut HEALTH BENEFITS Breakdown of polysaccharides Synthesis of vitamins Colonization
Fused-Core Particles:
 Fused-Core Particles: Varying Shell Thickness and Pore Size Stephanie A. Schuster; Joseph J. Kirkland; Brian M. Wagner; Barry E. Boyes; William L. Johnson; Timothy J. Langlois; Joseph J. DeStefano Advanced
Fused-Core Particles: Varying Shell Thickness and Pore Size Stephanie A. Schuster; Joseph J. Kirkland; Brian M. Wagner; Barry E. Boyes; William L. Johnson; Timothy J. Langlois; Joseph J. DeStefano Advanced
Mouse Clec9a ORF sequence
 Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
List of Figures. List of Tables
 Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
biochem480 [Autumn2014] Enzyme bio-informatics project
![biochem480 [Autumn2014] Enzyme bio-informatics project biochem480 [Autumn2014] Enzyme bio-informatics project](/thumbs/91/104514013.jpg) biochem480 [Autumn2014] Enzyme bio-informatics project Edit out any stuff in italics for the final version Student Name: IUBSystematic Name: 3-hydroxy-3-methylglutaryl-CoA reductase Other protein Names:
biochem480 [Autumn2014] Enzyme bio-informatics project Edit out any stuff in italics for the final version Student Name: IUBSystematic Name: 3-hydroxy-3-methylglutaryl-CoA reductase Other protein Names:
Supporting Information
 Supporting Information Identification, characterization and molecular adaptation of class I redox systems for the production of multi-functionalized diterpenoids Christian Görner a,, Patrick Schrepfer
Supporting Information Identification, characterization and molecular adaptation of class I redox systems for the production of multi-functionalized diterpenoids Christian Görner a,, Patrick Schrepfer
bifidobacteria Genus Bacteroides fragilis Non-specific longum, B. breve and B. longum bv. Infantis CGACGATCCCAGAGATGG Bifidobacterium longum and
 Additional file 3. Target organisms for 16-21-mer probes and actual tested microarray specificity. Probe name Full name a Target organism(s) Sequence (5'-3') Predicted probe specificity m Hybridisation
Additional file 3. Target organisms for 16-21-mer probes and actual tested microarray specificity. Probe name Full name a Target organism(s) Sequence (5'-3') Predicted probe specificity m Hybridisation
Protein Reports CPTAC Common Data Analysis Pipeline (CDAP)
 Protein Reports CPTAC Common Data Analysis Pipeline (CDAP) v. 05/03/2016 Summary The purpose of this document is to describe the protein reports generated as part of the CPTAC Common Data Analysis Pipeline
Protein Reports CPTAC Common Data Analysis Pipeline (CDAP) v. 05/03/2016 Summary The purpose of this document is to describe the protein reports generated as part of the CPTAC Common Data Analysis Pipeline
Lecture 3. Tandem MS & Protein Sequencing
 Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
The role of Bacteroides fragilis in the development of colorectal cancer. Jacqui Keenan Department of Surgery
 The role of Bacteroides fragilis in the development of colorectal cancer Jacqui Keenan Department of Surgery Microbial dysbiosis is observed in CRC patients Sobhani et al. PLoS ONE, 2011 Are there strains
The role of Bacteroides fragilis in the development of colorectal cancer Jacqui Keenan Department of Surgery Microbial dysbiosis is observed in CRC patients Sobhani et al. PLoS ONE, 2011 Are there strains
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 A β-strand positions consistently places the residues at CDR3β P6 and P7 within human and mouse TCR-peptide-MHC interfaces. (a) E8 TCR containing V β 13*06 carrying with an 11mer
Supplementary Figure 1 A β-strand positions consistently places the residues at CDR3β P6 and P7 within human and mouse TCR-peptide-MHC interfaces. (a) E8 TCR containing V β 13*06 carrying with an 11mer
Fig.S1 ESI-MS spectrum of reaction of ApA and THPTb after 16 h.
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Experiment Cleavage of dinucleotides Dinucleotides (ApA, CpC, GpG, UpU) were purchased from
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Experiment Cleavage of dinucleotides Dinucleotides (ApA, CpC, GpG, UpU) were purchased from
Chapter 6. X-ray structure analysis of D30N tethered HIV-1 protease. dimer/saquinavir complex
 Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Dr. Erin E. Chambers Waters Corporation. Presented by Dr. Diego Rodriguez Cabaleiro Waters Europe Waters Corporation 1
 Development of an SPE-LC/MS/MS Assay for the Simultaneous Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid in Support of Alzheimer s Research Dr. Erin E. Chambers Waters Corporation Presented
Development of an SPE-LC/MS/MS Assay for the Simultaneous Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid in Support of Alzheimer s Research Dr. Erin E. Chambers Waters Corporation Presented
NON TARGETED SEARCHING FOR FOOD
 NON TARGETED SEARCHING FOR FOOD CONTAMINANTS USING ORBITRAP HIGH RESOLUTION MASS SPECTROMETRY Michal Godula 1, Adrian Charlton 2 and Klaus Mittendorf 1 1 Thermo Fisher Scientific, Dreieich, Germany 2 Food
NON TARGETED SEARCHING FOR FOOD CONTAMINANTS USING ORBITRAP HIGH RESOLUTION MASS SPECTROMETRY Michal Godula 1, Adrian Charlton 2 and Klaus Mittendorf 1 1 Thermo Fisher Scientific, Dreieich, Germany 2 Food
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB
 Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Nature Structural & Molecular Biology: doi: /nsmb.3218
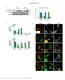 Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.
 Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Consistent and Reproducible Production of a Microbiota-based Drug for Recurrent C. difficile
 Consistent and Reproducible Production of a Microbiota-based Drug for Recurrent C. difficile Infection: Application of a Novel Diagnostic for Dysbiosis Courtney Jones, BS Rebiotix Inc. Roseville, MN USA
Consistent and Reproducible Production of a Microbiota-based Drug for Recurrent C. difficile Infection: Application of a Novel Diagnostic for Dysbiosis Courtney Jones, BS Rebiotix Inc. Roseville, MN USA
BBSI Lab Assignment for 6/15/2005. ** Turn in the answers to the questions on a separate piece of paper.
 BBSI Lab Assignment for 6/15/2005 ** Turn in the answers to the questions on a separate piece of paper. 1. Ramachandran Plot and Molecular Dynamics of Alanine Dipeptide In this Molecular Dynamics section,
BBSI Lab Assignment for 6/15/2005 ** Turn in the answers to the questions on a separate piece of paper. 1. Ramachandran Plot and Molecular Dynamics of Alanine Dipeptide In this Molecular Dynamics section,
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Time (min) Supplementary Figure 1: Gas decomposition products of irradiated DMC.
 200000 C 2 CH 3 CH 3 DMC 180000 160000 140000 Intensity 120000 100000 80000 60000 40000 C 2 H 6 CH 3 CH 2 CH 3 CH 3 CCH 3 EMC DEC 20000 C 3 H 8 HCCH 3 5 10 15 20 25 Time (min) Supplementary Figure 1: Gas
200000 C 2 CH 3 CH 3 DMC 180000 160000 140000 Intensity 120000 100000 80000 60000 40000 C 2 H 6 CH 3 CH 2 CH 3 CH 3 CCH 3 EMC DEC 20000 C 3 H 8 HCCH 3 5 10 15 20 25 Time (min) Supplementary Figure 1: Gas
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Biomolecular Mass Spectrometry
 Lipids ot different than other organic small molecules Carbohydrates Polymers of monosaccharides linked via glycosidic bonds (acetals/ ketals) many different combinationsvery interesting no time ucleic
Lipids ot different than other organic small molecules Carbohydrates Polymers of monosaccharides linked via glycosidic bonds (acetals/ ketals) many different combinationsvery interesting no time ucleic
Analysis of Uncomplexed and Copper-complexed Methanobactin with UV/Visible Spectrophotometry, Mass Spectrometry and NMR Spectrometry
 Analysis of Uncomplexed and Copper-complexed Methanobactin with UV/Visible Spectrophotometry, Mass Spectrometry and NMR Spectrometry Lee Behling, Alan DiSpirito, Scott Hartsel, Larry Masterson, Gianluigi
Analysis of Uncomplexed and Copper-complexed Methanobactin with UV/Visible Spectrophotometry, Mass Spectrometry and NMR Spectrometry Lee Behling, Alan DiSpirito, Scott Hartsel, Larry Masterson, Gianluigi
Mechanisms of Enzymes
 Mechanisms of Enzymes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy How enzymes work * Chemical reactions have an energy
Mechanisms of Enzymes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy How enzymes work * Chemical reactions have an energy
Phospholipid characterization by a TQ-MS data based identification scheme
 P-CN1716E Phospholipid characterization by a TQ-MS data based identification scheme ASMS 2017 MP-406 Tsuyoshi Nakanishi 1, Masaki Yamada 1, Ningombam Sanjib Meitei 2, 3 1 Shimadzu Corporation, Kyoto, Japan,
P-CN1716E Phospholipid characterization by a TQ-MS data based identification scheme ASMS 2017 MP-406 Tsuyoshi Nakanishi 1, Masaki Yamada 1, Ningombam Sanjib Meitei 2, 3 1 Shimadzu Corporation, Kyoto, Japan,
Microbe-managing: Manipulating the Human Gut Microbial Ecosystem to Enhance Health
 Microbe-managing: Manipulating the Human Gut Microbial Ecosystem to Enhance Health Emma Allen-Vercoe AMMI CANADA CACMID ANNUAL CONFERENCE April 5 th 2014 Conflict of Interest Disclosure Slide In the past
Microbe-managing: Manipulating the Human Gut Microbial Ecosystem to Enhance Health Emma Allen-Vercoe AMMI CANADA CACMID ANNUAL CONFERENCE April 5 th 2014 Conflict of Interest Disclosure Slide In the past
Rapid Characterization of Allosteric Networks. with Ensemble Normal Mode Analysis
 Supporting Information Rapid Characterization of Allosteric Networks with Ensemble Normal Mode Analysis Xin-Qiu Yao 1, Lars Skjærven 2, and Barry J. Grant 1* 1 Department of Computational Medicine and
Supporting Information Rapid Characterization of Allosteric Networks with Ensemble Normal Mode Analysis Xin-Qiu Yao 1, Lars Skjærven 2, and Barry J. Grant 1* 1 Department of Computational Medicine and
Supplemental Data. Candat et al. Plant Cell (2014) /tpc Cytosol. Nucleus. Mitochondria. Plastid. Peroxisome. Endomembrane system
 Cytosol Nucleus 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 PSORT MultiLoc YLoc SubLoc BaCelLo WoLF PSORT
Cytosol Nucleus 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 PSORT MultiLoc YLoc SubLoc BaCelLo WoLF PSORT
Biological Mass Spectrometry. April 30, 2014
 Biological Mass Spectrometry April 30, 2014 Mass Spectrometry Has become the method of choice for precise protein and nucleic acid mass determination in a very wide mass range peptide and nucleotide sequencing
Biological Mass Spectrometry April 30, 2014 Mass Spectrometry Has become the method of choice for precise protein and nucleic acid mass determination in a very wide mass range peptide and nucleotide sequencing
Figure S6. A-J) Annotated UVPD mass spectra for top ten peptides found among the peptides identified by Byonic but not SEQUEST + Percolator.
 Extending Proteome Coverage by Combining MS/MS Methods and a Modified Bioinformatics Platform adapted for Database Searching of Positive and Negative Polarity 193 nm Ultraviolet Photodissociation Mass
Extending Proteome Coverage by Combining MS/MS Methods and a Modified Bioinformatics Platform adapted for Database Searching of Positive and Negative Polarity 193 nm Ultraviolet Photodissociation Mass
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
SUPPLEMENTARY INFORMATION In format provided by Javier DeFelipe et al. (MARCH 2013)
 Supplementary Online Information S2 Analysis of raw data Forty-two out of the 48 experts finished the experiment, and only data from these 42 experts are considered in the remainder of the analysis. We
Supplementary Online Information S2 Analysis of raw data Forty-two out of the 48 experts finished the experiment, and only data from these 42 experts are considered in the remainder of the analysis. We
Unsupervised Identification of Isotope-Labeled Peptides
 Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
