Bacterial cellulose synthesis mechanism of facultative
|
|
|
- Sheila McKinney
- 5 years ago
- Views:
Transcription
1 Bacterial cellulose synthesis mechanism of facultative anaerobe Kaihua Ji 1 +, Wei Wang 2, 3 +, Bing Zeng 1, Sibin Chen 1, Qianqian Zhao 4, Yueqing Chen 1, Guoqiang Li 1* and Ting Ma 1*
2 5 6 7 Figure S1: Microscopic observations of E. coli transformant cells (A:without inducer; B: with inducer), (C) and Δhyp (D).
3 Figure S2: Comparison of transcriptome in identified COG categories of under aerobic and anaerobic conditions. S function unknown, R general function prediction only, Q secondary metabolites biosynthesis, P inorganic ion transport and metabolism, I lipid transport and metabolism, H coenzyme transport and metabolism, F nucleotide transport and metabolism, E amino acid transport and metabolism, G carbohydrate transport and metabolism, C energy production and conversion; X prophages, transposons, O post-translational modification, protein
4 turnover, chaperones, U intracellular trafficking and secretion, W extracellular structures, N cell motility, M cell wall/ membrane biogenesis, T signal transduction mechanisms, V defense mechanisms, D cell cycle control, mitosis, and meiosis, L replication, recombination, and repair, K transcription, J translation.
5 Figure S3: a: plasmid map of temperate-sensitive vector ptsk1 used for gene knockout experiment. b: design strategies of primers used for identification of gene knockout mutants.
6 25 26 Table S1: primers used for constructing of ptsk1 and constructing and identifying of gene knockout mutants primer name sequence(5'-3') purpose p46-1fw CAGACGAAGAATCCATGGGT construction of ptsk1 p46-1rv GTTCTCGAGATAAAGCGATGCAGGTGGC p18-1fw TATCCATGGCGGCTTCCATTCAGGTCG p18-1rv TATCTCGAGGGCACCCCAGGCTTTACA bcsi-1fw GATCAAGCTTTGTCGGTCGTTGAGCACATTG construction of gene bcsi-1rv CATTCACCCAGGCAGGCGTCCTTTCGGCGAAC AAACACAAA knockout vector ptsk-δbcsi bcsi-2fw TTTGTGTTTGTTCGCCGAAAGGACGCCTGCCT GGGTGAATG bcsi-2rv TACTAAGCTTATTCCGGCGCAGACTGCTC bcsi-1kfw CGGCCTGATGCTCGACCC verification of bcsi-1krv CGCCAGCCACACCAGCAT bcsi-2kfw GCCGGTGATGCAAATGCC ΔbcsI bcsi-2krv CGGTCCGGCCTTCTCGTT bcsa(i)-1fw TGTAGAATTCAGATCATGGCGGTGGTGGG construction of gene bcsa(i)-1rv GGTTTCGGCGGGTGAATCCTGGCTGTCCATCG GCGTAA knockout vector ptsk-δbcsa(i) bcsa(i)-2fw TTACGCCGATGGACAGCCAGGATTCACCCGCC GAAACC bcsa(i)-2rv TGTAAAGCTTCAGCGCGCCCTGCTTGTTA bcsa(i)-k1fw AGCGGCGCTACAATTTCAAC verification of bcsa(i)-k1rv GCGCGATGTCCAGGTCAT bcsa(i)-k2fw GGCACCCCCATGAAAAAGTT ΔbcsA(I) bcsa(i)-k2rv CAATGCGGCTTTTGATGTTGTTA bcsii-1fw AGCGGAATTCAATATGGTTCAGTGCGGTGTCG construction of gene bcsii-1rv CGATTAACCACCAGGAATTGTCTTTTAGTTTAA CCGATCCCTATGAATAAAAAGC knockout vector ptsk-δbcsii bcsii-2fw GCTTTTTATTCATAGGGATCGGTTAAACTAAAA GACAATTCCTGGTGGTTAATCG bcsii-2rv GCCCAAGCTTAAATCGCAAATTTTCCCTGCTCA bcsii-1kfw GCGGTGCCATCGTGTTCT verification of bcsii-1krv GGTACGATAAGCGCAGAGGAAA bcsii-2kfw AATATGGGGTCCACGATGTCC ΔbcsII bcsii-2krv AGGACTCTTCCAGCACCCAGT bcsa(ii)-1fw bcsa(ii)-1rv GCGCAAGCTTTTTTCCTGATTTGGCATGGTGAA C GAACAGGCTTTCCAGCGGTTTATGATCAGCAA construction of gene knockout vector ptsk-δbcsa(ii) CCAGGCAGACAGG
7 bcsa(ii)-2fw CCTGTCTGCCTGGTTGCTGATCATAAACCGCTG GAAAGCCTGTTC bcsa(ii)-2rv GCCGAAGCTTGGTTGATTGCACCTGCTCTTCAT A bcsa(ii)-k1fw ACCAGGCAACGTCATAAACATGT verification of bcsa(ii)-k1rv GCTGGCCCTTGCGTGAC bcsa(ii)-k2fw GCGTGCGGGCAGTGAG ΔbcsA(II) bcsa(ii)-k2rv CAGATCCGGCATGGCAATAAAGT bcsiii-1fw AGCTGAATTCTCATCGCCGCCAGGTTATCT construction of gene bcsiii-1rv GCCGGGGTACCTTTATTATTTTCTTTTCGCAAG GGCATTTCCAC knockout vector ptsk-δbcsiii bcsiii-2fw GTGGAAATGCCCTTGCGAAAAGAAAATAATAA AGGTACCCCGGC bcsiii-2rv GCTGAAGCTTGAAAGTTTAGTGACACGCTCCT GG bcsiii-1kfw TGTCTGACGGAGCCTCATC verification of bcsiii-1krv GACTAATTTGATGGCCAGCAGA bcsiii-2kfw TCATGGCTAAAACCGGTGAGC ΔbcsIII bcsiii-2krv GATGGCCACCAGCACGTTATC bcsiii-cfw TAAGGAATTCTACCGTTCAGCCATGTGGG complementation of bcsiii-crv GCAGTCTAGATTATTCCCCAATGGCGTAACG ΔbcsIII bcsa(iii)-1fw bcsa(iii)-1rv GGACAAGCTTAAAGTTTAGTGACACGCTCCTG GTC CTGAGGCCCAGTTCGATTTGTAATAAAGGTAC CCCGGCATAACAA construction of gene knockout vector ptsk-δbcsa(iii) bcsa(iii)-2fw TTGTTATGCCGGGGTACCTTTATTACAAATCGA ACTGGGCCTCAG bcsa(iii)-2rv GGCTGAATTCCGGAACGGCAAACGATTTT bcsa(iii)-k1f w bcsa(iii)-k1rv GAGCGCATCGTTAACGTCTTT GCGCGGGAGAGTACGAC verification of ΔbcsA(III) bcsa(iii)-k2f GTAACCGGAAACATTGTTATGCC w bcsa(iii)-k2rv CGGCGCCAGTAGACCTTCT bcsa(iii)-cfw TAAGGAATTCTACCGTTCAGCCATGTGGG complementation of bcsa(iii)-crv GCCGTCTAGATTAAAGCGCATTGTTAACCTCTT TC ΔbcsA(III) GFE-1Fw TATCAAGCTTGGCGATTCCGGTCTCGTTT construction of gene GFE-1Rv GGGCCACGATCTGCATAAAATT GCGAACGCGGTGGTTATTC knockout vector ptsk-δbcsgfe GFE-2Fw GAATAACCACCGCGTTCGC AATTTTATGCAGATCGTGGCCC GFE-2Rv AGCTAAGCTTGTCGAAAGGGCGTGGTTTG
8 GFE-k1Fw GCTGGTGTTCGGCGTTTTG verification of GFE-k1Rv GGCGGCGTCTGGTGGATAA GFE-k2Fw GGCGAGGGGAAGTCTGCAG ΔbcsGFE GFE-k2Rv TATGCGCCAGCTCTTTTTTGTG
9 27 28 Table S2: Overview of transcriptome sequencing results and QPCR results of differentially expressed genes Genebank number Product FPKM aerobic FPKM anaerobi c T-log2 (fold-change) anaerobic/aer obic Q-log2 (fold-change) anaerobic/aero bic AKI40_1504 Glucokinase AKI40_4682 Glucose-6-phosphate isomerase AKI40_ phosphofructokinase isozyme I AKI40_ phosphofructokinase II AKI40_0828 Fructose-bisphosphate aldolase class II AKI40_1706 Fructose-bisphosphate aldolase class 1 AKI40_4862 Triosephosphate isomerase AKI40_3256 Glyceraldehyde-3-phosph ate dehydrogenase AKI40_0827 Phosphoglycerate kinase AKI40_2201 Phosphoglycerate mutase family AKI40_0137 Phosphoglycerate mutase AKI40_4275 Probable phosphoglycerate mutase gpmb AKI40_4050 Enolase AKI40_2849 Pyruvate kinase I AKI40_3571 Pyruvate kinase II AKI40_4472 Fructose-1,6-bisphosphat ase class 1 AKI40_1324 Archaeal fructose-1,6-bisphosphata se and related enzymes of inositol monophosphatase family AKI40_4858 Fructose-1,6-bisphosphat ase class II AKI40_2388 Formate acetyltransferase AKI40_4164 Dihydrolipoyl dehydrogenase
10 AKI40_4165 Dihydrolipoyllysine-resid ue acetyltransferase component of pyruvate dehydrogenase c AKI40_4166 Pyruvate dehydrogenase E1 component AKI40_4167 Pyruvate dehydrogenase complex repressor (GntR-family transcriptional regulator) AKI40_1562 Phosphate acetyltransferase (Phosphotransacetylase) AKI40_1564 Acetate kinase A and propionate kinase 2 AKI40_2176 Citrate synthase AKI40_2177 Succinate dehydrogenase hydrophobic membrane anchor protein AKI40_2179 Succinate dehydrogenase flavoprotein AKI40_2180 Succinate dehydrogenase iron-sulfur protein AKI40_2181 Oxoglutarate dehydrogenase, E1 component AKI40_2182 Dihydrolipoyllysine-resid ue succinyltransferase, E2 component of oxoglutarate dehydrogenase complex AKI40_2183 Succinyl-CoA ligase beta AKI40_2184 Succinyl-CoA ligase alpha AKI40_3315 Aconitate hydratase AKI40_4162 Aconitate hydratase AKI40_2659 Isocitrate dehydrogenase AKI40_3457 Fumarate reductase, flavoprotein AKI40_4544 Fumarate reductase, flavoprotein ' AKI40_4545 Fumarate reductase iron-sulfur
11 AKI40_4546 Fumarate reductase C AKI40_4547 Fumarate reductase D AKI40_3459 Tartrate/fumarate subfamily Fe-S type hydro-lyase beta AKI40_0551 Malate dehydrogenase AKI40_3010 Malate dehydrogenase AKI40_1562 Phosphate acetyltransferase AKI40_1564 Acetate kinase A and propionate kinase 2 AKI40_2388 Formate acetyltransferase AKI40_3156 Formate dehydrogenase gamma AKI40_3157 Formate dehydrogenase, beta ' AKI40_3158 Formate dehydrogenase alpha AKI40_3242 Putative formate dehydrogenase AKI40_4598 Formate dehydrogenase H AKI40_4599 Formate dehydrogenase H alph AKI40_4203 Acetolactate synthase regulatory AKI40_4204 Acetolactate synthase isozyme 3 large AKI40_4818 acetolactate synthase catalytic AKI40_0023 acetolactate synthase catalytic AKI40_0024 Acetolactate synthase, isozyme I, small AKI40_0778 Alpha-acetolactate decarboxylase AKI40_0779 Acetolactate synthase, catabolic AKI40_2159 Phosphoglucomutase, alpha-d-glucose phosphate-specific
12 AKI40_3569 Glucose-6-phosphate dehydrogenase AKI40_ phosphogluconolactona se AKI40_ phosphogluconate dehydrogenase, decarboxylating AKI40_0415 Ribulose-phosphate epimerase AKI40_0833 Ribose-5-phosphate isomerase A AKI40_0825 Transketolase AKI40_1411 Transketolase AKI40_4259 transaldolase B AKI40_1412 transaldolase A AKI40_1025 Nitrate ABC superfamily ATP binding cassette transporter, binding protein AKI40_3367 Respiratory nitrate reductase gamma AKI40_3368 Nitrate reductase molybdenum cofactor assembly chaperone 1 AKI40_3369 Nitrate reductase 1, beta AKI40_3370 Nitrate reductase 1, alpha AKI40_0419 Probable nitrite transporter AKI40_0420 Nitrite reductase small AKI40_0421 Nitrite reductase, large AKI40_4912 Glutamine synthetase, type I AKI40_2718 Glutamate dehydrogenase AKI40_0561 Glutamate synthase, small AKI40_0563 Glutamate synthase large AKI40_3245 Alcohol dehydrogenase class III
13 AKI40_3351 Fused acetaldehyde-coa dehydrogenase iron-dependent alcohol dehydrogenase pyruvate-formate lyase deactivase AKI40_4427 Alcohol dehydrogenase GroES domain protein AKI40_1686 D-lactate dehydrogenase, membrane binding family AKI40_1574 NADH-quinone oxidoreductase B AKI40_1576 NADH-quinone oxidoreductase E AKI40_1578 NADH-quinone oxidoreductase G AKI40_1580 NADH-quinone oxidoreductase I AKI40_1582 NADH-quinone oxidoreductase K AKI40_1584 Proton-translocating NADH-quinone oxidoreductase, chain M AKI40_2908 NADP transhydrogenase alpha AKI40_2909 NAD/NADP transhydrogenase beta AKI40_0155 Predicted hydrogenase, 4Fe-4S ferredoxin-type component AKI40_0754 hydrogenase 2 small AKI40_0755 Hydrogenase 2 4Fe-4S ferredoxin-type component AKI40_0756 Ni/Fe-hydrogenase 2 b-type cytochrome AKI40_0757 Hydrogenase 2 large AKI40_0758 Maturation element for hydrogenase
14 AKI40_0759 AKI40_0760 AKI40_0761 AKI40_1736 AKI40_1739 AKI40_1745 AKI40_0889 AKI40_0890 AKI40_0894 AKI40_0893 AKI40_0892 Hydrogenase 2-specific chaperone Hydrogenase nickel incorporation protein HybF Hydrogenase maturation factor GDP-D-mannose dehydratase, NAD binding Glycosyl transferase, group 1 Colanic acid biosynthesis glycosyl transferase WcaL UTP-glucose-1-phosphate uridylyltransferase, GalU protein Diguanylate cyclase/phosphodiesteras e with PAS/PAC sensor glycosyl transferase, group 2 family protein Cyclic di-gmp-binding protein, Cellulose synthase operon protein B Cellulose synthase operon C domain protein AKI40_0891 Hypothetical protein AKI40_4699 Isocitrate lyase, putative AKI40_4700 Malate synthase A AKI40_1747 AKI40_3354 UTP-glucose-1-phosphate uridylyltransferase GalU UTP-glucose-1-phospha te uridylyltransferase GalU AKI40_0196 Hypothetical protein AKI40_0197 Cellulose synthase operon protein YhjQ -like protein AKI40_0198 Cellulose synthase
15 29 AKI40_0199 AKI40_0200 AKI40_0201 AKI40_0202 Cellulose synthase, B Cellulose synthase operon C domain protein Cellulose synthase operon protein D Glycosyl hydrolase, family AKI40_0206 Predicted protein AKI40_0207 Cellulose synthase operon protein YhjQ AKI40_0208 Cellulose synthase catalytic AKI40_0209 Cellulose synthase, B AKI40_0210 Endo-1,4-beta-glucanase AKI40_0211 Cellulose synthase operon C domain protein
incorporation into fructose-6-phosphate and glucose-6-phosphate (F6P/G6P); 2-
 Relative abundance (%) Relative abundance (%) Relative abundance (%) Relative abundance (%) 100% 80% 60% 40% F6P / G6P M+0 M+1 M+2 M+3 M+4 M+5 M+6 100% 80% 60% 40% 2PG / 3PG M+0 M+1 M+2 M+3 20% 20% 0%
Relative abundance (%) Relative abundance (%) Relative abundance (%) Relative abundance (%) 100% 80% 60% 40% F6P / G6P M+0 M+1 M+2 M+3 M+4 M+5 M+6 100% 80% 60% 40% 2PG / 3PG M+0 M+1 M+2 M+3 20% 20% 0%
Chem 109 C. Fall Armen Zakarian Office: Chemistry Bldn 2217
 Chem 109 C Fall 2014 Armen Zakarian ffice: Chemistry Bldn 2217 o Catabolism of carbohydrates: 10 reactions of glycolysis Chapter 25 C C 2 C 2 D-glucose α-d-glucopyranose aworth projection α-d-glucopyranose
Chem 109 C Fall 2014 Armen Zakarian ffice: Chemistry Bldn 2217 o Catabolism of carbohydrates: 10 reactions of glycolysis Chapter 25 C C 2 C 2 D-glucose α-d-glucopyranose aworth projection α-d-glucopyranose
Julia Vorholt Lecture 7:
 752-4001-00L Mikrobiologie Julia Vorholt Lecture 7: Chemoorganotrophy Nov 5, 2012 Brock Biology of Microorganisms, Twelfth Edition Madigan / Martinko / Dunlap / Clark Copyright 2009 Pearson Education Inc.,
752-4001-00L Mikrobiologie Julia Vorholt Lecture 7: Chemoorganotrophy Nov 5, 2012 Brock Biology of Microorganisms, Twelfth Edition Madigan / Martinko / Dunlap / Clark Copyright 2009 Pearson Education Inc.,
Photosynthesis in chloroplasts. Cellular respiration in mitochondria ATP. ATP powers most cellular work
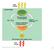 Light energy ECOSYSTEM CO + H O Photosynthesis in chloroplasts Cellular respiration in mitochondria Organic molecules + O powers most cellular work Heat energy 1 becomes oxidized (loses electron) becomes
Light energy ECOSYSTEM CO + H O Photosynthesis in chloroplasts Cellular respiration in mitochondria Organic molecules + O powers most cellular work Heat energy 1 becomes oxidized (loses electron) becomes
Supporting Information
 Supporting Information Bar-Even et al. 10.1073/pnas.0907176107 Fig. S1. A scheme of the reductive pentose phosphate cycle, as inspired from (81). Dashed line corresponds to RUBISCO's oxygenase reaction
Supporting Information Bar-Even et al. 10.1073/pnas.0907176107 Fig. S1. A scheme of the reductive pentose phosphate cycle, as inspired from (81). Dashed line corresponds to RUBISCO's oxygenase reaction
This is an example outline of 3 lectures in BSC (Thanks to Dr. Ellington for sharing this information.)
 This is an example outline of 3 lectures in BSC 2010. (Thanks to Dr. Ellington for sharing this information.) Topic 10: CELLULAR RESPIRATION (lectures 14-16) OBJECTIVES: 1. Know the basic reactions that
This is an example outline of 3 lectures in BSC 2010. (Thanks to Dr. Ellington for sharing this information.) Topic 10: CELLULAR RESPIRATION (lectures 14-16) OBJECTIVES: 1. Know the basic reactions that
Yield of energy from glucose
 Paper : Module : 05 Yield of Energy from Glucose Principal Investigator, Paper Coordinator and Content Writer Prof. Ramesh Kothari, Professor Dept. of Biosciences, Saurashtra University, Rajkot - 360005
Paper : Module : 05 Yield of Energy from Glucose Principal Investigator, Paper Coordinator and Content Writer Prof. Ramesh Kothari, Professor Dept. of Biosciences, Saurashtra University, Rajkot - 360005
Cellular Respiration: Harvesting Chemical Energy
 Chapter 9 Cellular Respiration: Harvesting Chemical Energy You should be able to: 1. Explain how redox reactions are involved in energy exchanges. Name and describe the three stages of cellular respiration;
Chapter 9 Cellular Respiration: Harvesting Chemical Energy You should be able to: 1. Explain how redox reactions are involved in energy exchanges. Name and describe the three stages of cellular respiration;
Pathway overview. Glucose + 2NAD + + 2ADP +2Pi 2NADH + 2pyruvate + 2ATP + 2H 2 O + 4H +
 Glycolysis Glycolysis The conversion of glucose to pyruvate to yield 2ATP molecules 10 enzymatic steps Chemical interconversion steps Mechanisms of enzyme conversion and intermediates Energetics of conversions
Glycolysis Glycolysis The conversion of glucose to pyruvate to yield 2ATP molecules 10 enzymatic steps Chemical interconversion steps Mechanisms of enzyme conversion and intermediates Energetics of conversions
Metabolism. Metabolic pathways. BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Metabolism Metabolism is the chemical change of
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Metabolism Metabolism is the chemical change of
Chapter 9. Cellular Respiration and Fermentation
 Chapter 9 Cellular Respiration and Fermentation Energy flows into an ecosystem as sunlight and leaves as heat Photosynthesis generates O 2 and organic molecules, which are used in cellular respiration
Chapter 9 Cellular Respiration and Fermentation Energy flows into an ecosystem as sunlight and leaves as heat Photosynthesis generates O 2 and organic molecules, which are used in cellular respiration
Respiration. Organisms can be classified based on how they obtain energy: Autotrophs
 Respiration rganisms can be classified based on how they obtain energy: Autotrophs Able to produce their own organic molecules through photosynthesis Heterotrophs Live on organic compounds produced by
Respiration rganisms can be classified based on how they obtain energy: Autotrophs Able to produce their own organic molecules through photosynthesis Heterotrophs Live on organic compounds produced by
Photosynthesis in chloroplasts CO2 + H2O. Cellular respiration in mitochondria ATP. powers most cellular work. Heat energy
 Figure 9-01 LE 9-2 Light energy ECOSYSTEM Photosynthesis in chloroplasts CO2 + H2O Cellular respiration in mitochondria Organic + O molecules 2 powers most cellular work Heat energy LE 9-UN161a becomes
Figure 9-01 LE 9-2 Light energy ECOSYSTEM Photosynthesis in chloroplasts CO2 + H2O Cellular respiration in mitochondria Organic + O molecules 2 powers most cellular work Heat energy LE 9-UN161a becomes
Comparison of catabolic and anabolic pathways
 Comparison of catabolic and anabolic pathways Three stages of catabolism Glucose Synthesis of compounds e.g. lactose glycolipids Glucose-6-P Pentosephosphate Pathway Glycolysis Glycogenesis Acetyl-CoA
Comparison of catabolic and anabolic pathways Three stages of catabolism Glucose Synthesis of compounds e.g. lactose glycolipids Glucose-6-P Pentosephosphate Pathway Glycolysis Glycogenesis Acetyl-CoA
Citric acid cycle. Tomáš Kučera.
 itric acid cycle Tomáš Kučera tomas.kucera@lfmotol.cuni.cz Department of Medical hemistry and linical Biochemistry 2nd Faculty of Medicine, harles University in Prague and Motol University Hospital 2017
itric acid cycle Tomáš Kučera tomas.kucera@lfmotol.cuni.cz Department of Medical hemistry and linical Biochemistry 2nd Faculty of Medicine, harles University in Prague and Motol University Hospital 2017
Metabolism III. Aim: understand gluconeogenesis, pentose phosphate pathway, photosynthesis and amino acid synthesis
 Metabolism III Aim: understand gluconeogenesis, pentose phosphate pathway, photosynthesis and amino acid synthesis Anabolism From a carbon source and inorganic molecules, microbes synthesize new organelles
Metabolism III Aim: understand gluconeogenesis, pentose phosphate pathway, photosynthesis and amino acid synthesis Anabolism From a carbon source and inorganic molecules, microbes synthesize new organelles
Chapter 9: Cellular Respiration Overview: Life Is Work. Living cells. Require transfusions of energy from outside sources to perform their many tasks
 Chapter 9: Cellular Respiration Overview: Life Is Work Living cells Require transfusions of energy from outside sources to perform their many tasks Biology, 7 th Edition Neil Campbell and Jane Reece The
Chapter 9: Cellular Respiration Overview: Life Is Work Living cells Require transfusions of energy from outside sources to perform their many tasks Biology, 7 th Edition Neil Campbell and Jane Reece The
Cellular Respiration: Harvesting Chemical Energy Chapter 9
 Cellular Respiration: Harvesting Chemical Energy Chapter 9 Assemble polymers, pump substances across membranes, move and reproduce The giant panda Obtains energy for its cells by eating plants which get
Cellular Respiration: Harvesting Chemical Energy Chapter 9 Assemble polymers, pump substances across membranes, move and reproduce The giant panda Obtains energy for its cells by eating plants which get
Biological processes. Mitochondrion Metabolic process Catalytic activity Oxidoreductase
 Full name Glyceraldehyde3 phosphate dehydrogenase Succinatesemialdehyde Glutamate dehydrogenase 1, Alcohol dehydrogenase [NADP+] 2',3'cyclicnucleotide 3' phosphodiesterase Dihydropyrimidinaserelated 2
Full name Glyceraldehyde3 phosphate dehydrogenase Succinatesemialdehyde Glutamate dehydrogenase 1, Alcohol dehydrogenase [NADP+] 2',3'cyclicnucleotide 3' phosphodiesterase Dihydropyrimidinaserelated 2
Chem Lecture 8 Carbohydrate Metabolism Part I: Glycolysis
 Chem 352 - Lecture 8 Carbohydrate Metabolism Part I: Glycolysis Introduction Carbohydrate metabolism involves a collection of pathways. Glycolysis Hexoses 3-Carbon molecules Gluconeogenesis 3-Carbon molecules
Chem 352 - Lecture 8 Carbohydrate Metabolism Part I: Glycolysis Introduction Carbohydrate metabolism involves a collection of pathways. Glycolysis Hexoses 3-Carbon molecules Gluconeogenesis 3-Carbon molecules
III. Metabolism The Citric Acid Cycle
 Department of Chemistry and Biochemistry University of Lethbridge III. Metabolism The Citric Acid Cycle Slide 1 The Eight Steps of the Citric Acid Cycle Enzymes: 4 dehydrogenases (2 decarboxylation) 3
Department of Chemistry and Biochemistry University of Lethbridge III. Metabolism The Citric Acid Cycle Slide 1 The Eight Steps of the Citric Acid Cycle Enzymes: 4 dehydrogenases (2 decarboxylation) 3
Cellular Respiration and Fermentation
 CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY REECE 7 Cellular Respiration and Fermentation Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge, Simon Fraser University SECOND EDITION
CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY REECE 7 Cellular Respiration and Fermentation Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge, Simon Fraser University SECOND EDITION
Cellular Respiration and Fermentation
 CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY REECE 7 Cellular Respiration and Fermentation Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge, Simon Fraser University SECOND EDITION
CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY REECE 7 Cellular Respiration and Fermentation Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge, Simon Fraser University SECOND EDITION
Chapter 13 Carbohydrate Metabolism
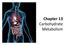 Chapter 13 Carbohydrate Metabolism Metabolism of Foods Food is broken down into carbohydrates, lipids, and proteins and sent through catabolic pathways to produce energy. Glycolysis glucose 2 P i 2 ADP
Chapter 13 Carbohydrate Metabolism Metabolism of Foods Food is broken down into carbohydrates, lipids, and proteins and sent through catabolic pathways to produce energy. Glycolysis glucose 2 P i 2 ADP
Aerobic Respiration. The four stages in the breakdown of glucose
 Aerobic Respiration The four stages in the breakdown of glucose 1 I. Aerobic Respiration Why can t we break down Glucose in one step? (Flaming Gummy Bear) Enzymes gently lower the potential energy until
Aerobic Respiration The four stages in the breakdown of glucose 1 I. Aerobic Respiration Why can t we break down Glucose in one step? (Flaming Gummy Bear) Enzymes gently lower the potential energy until
PRINT your Name Student (FAMILY, first name) Midterm 7:00 P.M.
 PRINT your Name Student No. (FAMILY, first name) BIOCHEMISTRY 311A VERSION 1 (ONE) Midterm 7:00 P.M. Examiners: Dr. R. E. MacKenzie (69%) Dr. A. Storer (18%) Dr. W. Mushynski (13%) READ THE QUESTIONS CAREFULLY!!
PRINT your Name Student No. (FAMILY, first name) BIOCHEMISTRY 311A VERSION 1 (ONE) Midterm 7:00 P.M. Examiners: Dr. R. E. MacKenzie (69%) Dr. A. Storer (18%) Dr. W. Mushynski (13%) READ THE QUESTIONS CAREFULLY!!
BIOLOGY. Cellular Respiration and Fermentation CAMPBELL. Photosynthesis in chloroplasts. Light energy ECOSYSTEM. Organic molecules CO 2 + H 2 O
 9 Cellular Respiration and Fermentation CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick Figure 9.1 Figure 9.2
9 Cellular Respiration and Fermentation CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick Figure 9.1 Figure 9.2
number Done by Corrected by Doctor Nayef Karadsheh
 number 11 Done by حسام أبو عوض Corrected by Moayyad Al-Shafei Doctor Nayef Karadsheh 1 P a g e General Regulatory Aspects in Metabolism: We can divide all pathways in metabolism to catabolicand anabolic.
number 11 Done by حسام أبو عوض Corrected by Moayyad Al-Shafei Doctor Nayef Karadsheh 1 P a g e General Regulatory Aspects in Metabolism: We can divide all pathways in metabolism to catabolicand anabolic.
BIOLOGY. Cellular Respiration and Fermentation CAMPBELL. Reece Urry Cain Wasserman Minorsky Jackson
 CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 9 Cellular Respiration and Fermentation Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick Figure 9.2 Light energy
CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 9 Cellular Respiration and Fermentation Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick Figure 9.2 Light energy
Carbohydrate Metabolism
 OpenStax-CNX module: m46451 1 Carbohydrate Metabolism OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
OpenStax-CNX module: m46451 1 Carbohydrate Metabolism OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
Microbial Metabolism. PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R
 PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R 5 Microbial Metabolism Big Picture: Metabolism Metabolism is the buildup and breakdown of nutrients
PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R 5 Microbial Metabolism Big Picture: Metabolism Metabolism is the buildup and breakdown of nutrients
CHE 242 Exam 3 Practice Questions
 CHE 242 Exam 3 Practice Questions Glucose metabolism 1. Below is depicted glucose catabolism. Indicate on the pathways the following: A) which reaction(s) of glycolysis are irreversible B) where energy
CHE 242 Exam 3 Practice Questions Glucose metabolism 1. Below is depicted glucose catabolism. Indicate on the pathways the following: A) which reaction(s) of glycolysis are irreversible B) where energy
Cellular Respiration: Harvesting Chemical Energy
 Chapter 9 Cellular Respiration: Harvesting Chemical Energy PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with
Chapter 9 Cellular Respiration: Harvesting Chemical Energy PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with
7 Cellular Respiration and Fermentation
 CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY REECE 7 Cellular Respiration and Fermentation Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge, Simon Fraser University SECOND EDITION
CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY REECE 7 Cellular Respiration and Fermentation Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge, Simon Fraser University SECOND EDITION
CHAPTER 16. Glycolysis
 CHAPTER 16 Glycolysis Net reaction of Glycolysis Converts: 1 Glucose Hexose stage 2 pyruvate - Two molecules of ATP are produced - Two molecules of NAD + are reduced to NADH Triose stage Glucose + 2 ADP
CHAPTER 16 Glycolysis Net reaction of Glycolysis Converts: 1 Glucose Hexose stage 2 pyruvate - Two molecules of ATP are produced - Two molecules of NAD + are reduced to NADH Triose stage Glucose + 2 ADP
7 Cellular Respiration and Fermentation
 CAMPBELL BIOLOGY IN FOCUS Urry Cain Wasserman Minorsky Jackson Reece 7 Cellular Respiration and Fermentation Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge Overview: Life Is Work Living
CAMPBELL BIOLOGY IN FOCUS Urry Cain Wasserman Minorsky Jackson Reece 7 Cellular Respiration and Fermentation Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge Overview: Life Is Work Living
Cellular Respiration: Harvesting Chemical Energy
 Chapter 9 Cellular Respiration: Harvesting Chemical Energy Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated
Chapter 9 Cellular Respiration: Harvesting Chemical Energy Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated
Glycolysis Part 2. BCH 340 lecture 4
 Glycolysis Part 2 BCH 340 lecture 4 Regulation of Glycolysis There are three steps in glycolysis that have enzymes which regulate the flux of glycolysis These enzymes catalyzes irreversible reactions of
Glycolysis Part 2 BCH 340 lecture 4 Regulation of Glycolysis There are three steps in glycolysis that have enzymes which regulate the flux of glycolysis These enzymes catalyzes irreversible reactions of
Points 1. Following is the overall reaction catalyzed by the Calvin-Benson cycle:
 BCH 4054 February 22, 2002 HOUR TEST 2 NAME_ Points 1. Following is the overall reaction catalyzed by the Calvin-Benson cycle: CO 2 + 3ATP + 2NADPH 1/3 glyceraldehyde-3-p + 3ADP + 2NADP + Give the structures
BCH 4054 February 22, 2002 HOUR TEST 2 NAME_ Points 1. Following is the overall reaction catalyzed by the Calvin-Benson cycle: CO 2 + 3ATP + 2NADPH 1/3 glyceraldehyde-3-p + 3ADP + 2NADP + Give the structures
respiration mitochondria mitochondria metabolic pathways reproduction can fuse or split DRP1 interacts with ER tubules chapter DRP1 ER tubule
 mitochondria respiration chapter 3-4 shape highly variable can fuse or split structure outer membrane inner membrane cristae intermembrane space mitochondrial matrix free ribosomes respiratory enzymes
mitochondria respiration chapter 3-4 shape highly variable can fuse or split structure outer membrane inner membrane cristae intermembrane space mitochondrial matrix free ribosomes respiratory enzymes
Campbell Biology 9. Chapter 9 Cellular Respiration and Fermentation. Chul-Su Yang, Ph.D., Lecture on General Biology 1
 Lecture on General Biology 1 Campbell Biology 9 th edition Chapter 9 Cellular Respiration and Fermentation Chul-Su Yang, Ph.D., chulsuyang@hanyang.ac.kr Infection Biology Lab., Dept. of Molecular & Life
Lecture on General Biology 1 Campbell Biology 9 th edition Chapter 9 Cellular Respiration and Fermentation Chul-Su Yang, Ph.D., chulsuyang@hanyang.ac.kr Infection Biology Lab., Dept. of Molecular & Life
7 Cellular Respiration and Fermentation
 CAMPBELL BIOLOGY IN FOCUS Urry Cain Wasserman Minorsky Jackson Reece 7 Cellular Respiration and Fermentation Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge Overview: Life Is Work Living
CAMPBELL BIOLOGY IN FOCUS Urry Cain Wasserman Minorsky Jackson Reece 7 Cellular Respiration and Fermentation Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge Overview: Life Is Work Living
BIOLOGY. Cellular Respiration and Fermentation CAMPBELL. Reece Urry Cain Wasserman Minorsky Jackson
 CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 9 Cellular Respiration and Fermentation Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick Life Is Work Living cells
CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 9 Cellular Respiration and Fermentation Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick Life Is Work Living cells
TCA CYCLE (Citric Acid Cycle)
 TCA CYCLE (Citric Acid Cycle) TCA CYCLE The Citric Acid Cycle is also known as: Kreb s cycle Sir Hans Krebs Nobel prize, 1953 TCA (tricarboxylic acid) cycle The citric acid cycle requires aerobic conditions!!!!
TCA CYCLE (Citric Acid Cycle) TCA CYCLE The Citric Acid Cycle is also known as: Kreb s cycle Sir Hans Krebs Nobel prize, 1953 TCA (tricarboxylic acid) cycle The citric acid cycle requires aerobic conditions!!!!
7/5/2014. Microbial. Metabolism. Basic Chemical Reactions Underlying. Metabolism. Metabolism: Overview
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University Basic Chemical Reactions Underlying Metabolism Metabolism C H A P T E R 5 Microbial Metabolism Collection
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University Basic Chemical Reactions Underlying Metabolism Metabolism C H A P T E R 5 Microbial Metabolism Collection
BIOLOGY. Cellular Respiration and Fermentation CAMPBELL. Reece Urry Cain Wasserman Minorsky Jackson
 CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 9 Cellular Respiration and Fermentation Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick Life Is Work Living cells
CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 9 Cellular Respiration and Fermentation Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick Life Is Work Living cells
Integration of Metabolism
 Integration of Metabolism Metabolism is a continuous process. Thousands of reactions occur simultaneously in order to maintain homeostasis. It ensures a supply of fuel, to tissues at all times, in fed
Integration of Metabolism Metabolism is a continuous process. Thousands of reactions occur simultaneously in order to maintain homeostasis. It ensures a supply of fuel, to tissues at all times, in fed
Ch 07. Microbial Metabolism
 Ch 07 Microbial Metabolism SLOs Differentiate between metabolism, catabolism, and anabolism. Fully describe the structure and function of enzymes. Differentiate between constitutive and regulated enzymes.
Ch 07 Microbial Metabolism SLOs Differentiate between metabolism, catabolism, and anabolism. Fully describe the structure and function of enzymes. Differentiate between constitutive and regulated enzymes.
Energetics of carbohydrate and lipid metabolism
 Energetics of carbohydrate and lipid metabolism 1 Metabolism: The sum of all the chemical transformations taking place in a cell or organism, occurs through a series of enzymecatalyzed reactions that constitute
Energetics of carbohydrate and lipid metabolism 1 Metabolism: The sum of all the chemical transformations taking place in a cell or organism, occurs through a series of enzymecatalyzed reactions that constitute
7 Cellular Respiration and Fermentation
 CAMPBELL BIOLOGY IN FOCUS Urry Cain Wasserman Minorsky Jackson Reece 7 Cellular Respiration and Fermentation Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge Overview: Life Is Work Living
CAMPBELL BIOLOGY IN FOCUS Urry Cain Wasserman Minorsky Jackson Reece 7 Cellular Respiration and Fermentation Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge Overview: Life Is Work Living
CH 7: Cell Respiration and Fermentation Overview. Concept 7.1: Catabolic pathways yield energy by oxidizing organic fuels
 CH 7: Cell Respiration and Fermentation Overview Living cells require energy from outside sources Some animals obtain energy by eating plants, and some animals feed on other organisms Energy flows into
CH 7: Cell Respiration and Fermentation Overview Living cells require energy from outside sources Some animals obtain energy by eating plants, and some animals feed on other organisms Energy flows into
Cellular Respiration and Fermentation
 LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 9 Cellular Respiration and Fermentation
LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 9 Cellular Respiration and Fermentation
Lecture 13 (10/13/17)
 Lecture 13 (10/13/17) Reading: Ch6; 187-189, 204-205 Problems: Ch4 (text); 2, 3 NXT (after xam 2) Reading: Ch6; 190-191, 194-195, 197-198 Problems: Ch6 (text); 5, 6, 7, 24 OUTLIN NZYMS: Binding & Catalysis
Lecture 13 (10/13/17) Reading: Ch6; 187-189, 204-205 Problems: Ch4 (text); 2, 3 NXT (after xam 2) Reading: Ch6; 190-191, 194-195, 197-198 Problems: Ch6 (text); 5, 6, 7, 24 OUTLIN NZYMS: Binding & Catalysis
4. Which step shows a split of one molecule into two smaller molecules? a. 2. d. 5
 1. Which of the following statements about NAD + is false? a. NAD + is reduced to NADH during both glycolysis and the citric acid cycle. b. NAD + has more chemical energy than NADH. c. NAD + is reduced
1. Which of the following statements about NAD + is false? a. NAD + is reduced to NADH during both glycolysis and the citric acid cycle. b. NAD + has more chemical energy than NADH. c. NAD + is reduced
Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function
 Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function 1 BPSL0024 26223 26621 + LrgA family BPSL0025 26690 27412 + hypothetical
Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function 1 BPSL0024 26223 26621 + LrgA family BPSL0025 26690 27412 + hypothetical
2. (12 pts) Given the following metabolic pathway (as it occurs in the cell):
 Answer Sheet 1 (Gold) 1. (1 pt) Write your exam ID (A) in the blank at the upper right of your answer sheet. 2. (12 pts) Given the following metabolic pathway (as it occurs in the cell): a. Would you expect
Answer Sheet 1 (Gold) 1. (1 pt) Write your exam ID (A) in the blank at the upper right of your answer sheet. 2. (12 pts) Given the following metabolic pathway (as it occurs in the cell): a. Would you expect
CITRIC ACID CYCLE ERT106 BIOCHEMISTRY SEM /19 BY: MOHAMAD FAHRURRAZI TOMPANG
 CITRIC ACID CYCLE ERT106 BIOCHEMISTRY SEM 1 2018/19 BY: MOHAMAD FAHRURRAZI TOMPANG Chapter Outline (19-1) The central role of the citric acid cycle in metabolism (19-2) The overall pathway of the citric
CITRIC ACID CYCLE ERT106 BIOCHEMISTRY SEM 1 2018/19 BY: MOHAMAD FAHRURRAZI TOMPANG Chapter Outline (19-1) The central role of the citric acid cycle in metabolism (19-2) The overall pathway of the citric
CELLULAR RESPIRATION SUMMARY EQUATION. C 6 H 12 O 6 + O 2 6CO2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION
 CELLULAR RESPIRATION SUMMARY EQUATION C 6 H 12 O 6 + O 2 6CO2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION Oxidation: partial or complete loss of electrons Reduction: partial or complete gain of electrons
CELLULAR RESPIRATION SUMMARY EQUATION C 6 H 12 O 6 + O 2 6CO2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION Oxidation: partial or complete loss of electrons Reduction: partial or complete gain of electrons
Module No. # 01 Lecture No. # 19 TCA Cycle
 Biochemical Engineering Prof. Dr. Rintu Banerjee Department of Agricultural and Food Engineering Asst. Prof. Dr. Saikat Chakraborty Department of Chemical Engineering Indian Institute of Technology, Kharagpur
Biochemical Engineering Prof. Dr. Rintu Banerjee Department of Agricultural and Food Engineering Asst. Prof. Dr. Saikat Chakraborty Department of Chemical Engineering Indian Institute of Technology, Kharagpur
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site.
 Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
MULTIPLE CHOICE QUESTIONS
 MULTIPLE CHOICE QUESTIONS 1. Which of the following statements concerning anabolic reactions is FALSE? A. They are generally endergonic. B. They usually require ATP. C. They are part of metabolism. D.
MULTIPLE CHOICE QUESTIONS 1. Which of the following statements concerning anabolic reactions is FALSE? A. They are generally endergonic. B. They usually require ATP. C. They are part of metabolism. D.
We must be able to make glucose
 Biosynthesis of Carbohydrates Synthesis of glucose (gluconeogenesis) Glycogen Formation of pentoses and NADPH Photosynthesis We must be able to make glucose Compulsory need for glucose (above all the brain)
Biosynthesis of Carbohydrates Synthesis of glucose (gluconeogenesis) Glycogen Formation of pentoses and NADPH Photosynthesis We must be able to make glucose Compulsory need for glucose (above all the brain)
Lecture METABOLISM OF CARBOHYDRATE
 Lecture 19-24 METABOLISM OF CARBOHYDRATE Introduction Carbohydrates are major sources of energy for living organisms. The chief source of carbohydrate in human food is starch, which is the storage form
Lecture 19-24 METABOLISM OF CARBOHYDRATE Introduction Carbohydrates are major sources of energy for living organisms. The chief source of carbohydrate in human food is starch, which is the storage form
Tutorial 27: Metabolism, Krebs Cycle and the Electron Transport Chain
 Tutorial 27: Metabolism, Krebs Cycle and the Electron Transport Chain Goals: To be able to describe the overall catabolic pathways for food molecules. To understand what bonds are hydrolyzed in the digestion
Tutorial 27: Metabolism, Krebs Cycle and the Electron Transport Chain Goals: To be able to describe the overall catabolic pathways for food molecules. To understand what bonds are hydrolyzed in the digestion
NAME KEY ID # EXAM 3a BIOC 460. Wednesday April 10, Please include your name and ID# on each page. Limit your answers to the space provided!
 EXAM 3a BIOC 460 Wednesday April 10, 2002 Please include your name and ID# on each page. Limit your answers to the space provided! 1 1. (5 pts.) Define the term energy charge: Energy charge refers to the
EXAM 3a BIOC 460 Wednesday April 10, 2002 Please include your name and ID# on each page. Limit your answers to the space provided! 1 1. (5 pts.) Define the term energy charge: Energy charge refers to the
Citric Acid Cycle: Central Role in Catabolism. Entry of Pyruvate into the TCA cycle
 Citric Acid Cycle: Central Role in Catabolism Stage II of catabolism involves the conversion of carbohydrates, fats and aminoacids into acetylcoa In aerobic organisms, citric acid cycle makes up the final
Citric Acid Cycle: Central Role in Catabolism Stage II of catabolism involves the conversion of carbohydrates, fats and aminoacids into acetylcoa In aerobic organisms, citric acid cycle makes up the final
Notes CELLULAR RESPIRATION SUMMARY EQUATION C 6 H 12 O 6 + O 2. 6CO 2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION
 AP BIOLOGY CELLULAR ENERGETICS ACTIVITY #2 Notes NAME DATE HOUR SUMMARY EQUATION CELLULAR RESPIRATION C 6 H 12 O 6 + O 2 6CO 2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION Oxidation: partial or complete
AP BIOLOGY CELLULAR ENERGETICS ACTIVITY #2 Notes NAME DATE HOUR SUMMARY EQUATION CELLULAR RESPIRATION C 6 H 12 O 6 + O 2 6CO 2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION Oxidation: partial or complete
Transport. Oxidation. Electron. which the en the ETC and. of NADH an. nd FADH 2 by ation. Both, Phosphorylation. Glycolysis Glucose.
 Electron Transport Chain and Oxidation Phosphorylation When one glucose molecule is oxidized to six CO 2 molecules by way of glycolysiss and TCA cycle, considerable amount of energy (ATP) is generated.
Electron Transport Chain and Oxidation Phosphorylation When one glucose molecule is oxidized to six CO 2 molecules by way of glycolysiss and TCA cycle, considerable amount of energy (ATP) is generated.
Carbohydrate Metabolism I
 Carbohydrate Metabolism I Outline Glycolysis Stages of glycolysis Regulation of Glycolysis Carbohydrate Metabolism Overview Enzyme Classification Dehydrogenase - oxidizes substrate using cofactors as
Carbohydrate Metabolism I Outline Glycolysis Stages of glycolysis Regulation of Glycolysis Carbohydrate Metabolism Overview Enzyme Classification Dehydrogenase - oxidizes substrate using cofactors as
BIOLOGY. Cellular Respiration and Fermentation. Concept 9.1: Catabolic pathways yield energy by oxidizing organic fuels
 9 Cellular Respiration and Fermentation CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson Energy flows into an ecosystem as sunlight and leaves as heat Photosynthesis generates
9 Cellular Respiration and Fermentation CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson Energy flows into an ecosystem as sunlight and leaves as heat Photosynthesis generates
Products List We have more enzymes other than listed here. Contact us freely.
 Glycolysis and Ethanol Fermentation Volume EC No. Thermostability Example of use product Glucokinase YK1 glucose 7.5~11.0 1 ml 2.7.1.2 phosphorylation Stable after incubation at 80 o C for 60 min GLK-97-01
Glycolysis and Ethanol Fermentation Volume EC No. Thermostability Example of use product Glucokinase YK1 glucose 7.5~11.0 1 ml 2.7.1.2 phosphorylation Stable after incubation at 80 o C for 60 min GLK-97-01
BY: RASAQ NURUDEEN OLAJIDE
 BY: RASAQ NURUDEEN OLAJIDE LECTURE CONTENT INTRODUCTION CITRIC ACID CYCLE (T.C.A) PRODUCTION OF ACETYL CoA REACTIONS OF THE CITIRC ACID CYCLE THE AMPHIBOLIC NATURE OF THE T.C.A CYCLE THE GLYOXYLATE CYCLE
BY: RASAQ NURUDEEN OLAJIDE LECTURE CONTENT INTRODUCTION CITRIC ACID CYCLE (T.C.A) PRODUCTION OF ACETYL CoA REACTIONS OF THE CITIRC ACID CYCLE THE AMPHIBOLIC NATURE OF THE T.C.A CYCLE THE GLYOXYLATE CYCLE
 Chapter-5 Respiration in Plants Very Short Answers Questions: 1. Different substrates get oxidized during respiration. How does respiratory quotient (RQ) indicate which type of substrate i.e. carbohydrate,
Chapter-5 Respiration in Plants Very Short Answers Questions: 1. Different substrates get oxidized during respiration. How does respiratory quotient (RQ) indicate which type of substrate i.e. carbohydrate,
Cellular Respiration: Harvesting Chemical Energy
 Chapter 9 Cellular Respiration: Harvesting Chemical Energy Overview: Life Is Work Living cells require energy from outside sources Some animals, such as the giant panda, obtain energy by eating plants,
Chapter 9 Cellular Respiration: Harvesting Chemical Energy Overview: Life Is Work Living cells require energy from outside sources Some animals, such as the giant panda, obtain energy by eating plants,
Anaerobic Protist Metabolism
 Anaerobic Protist Metabolism Giardia, Entamoeba and Trichomonads Review glycolysis and oxidative decarboxylation Parasite Biochemistry (in general) Parasitic life style = ADAPTATIONS Specific niche = diversity
Anaerobic Protist Metabolism Giardia, Entamoeba and Trichomonads Review glycolysis and oxidative decarboxylation Parasite Biochemistry (in general) Parasitic life style = ADAPTATIONS Specific niche = diversity
Biochemistry: A Short Course
 Tymoczko Berg Stryer Biochemistry: A Short Course Second Edition CHAPTER 16 Glycolysis 2013 W. H. Freeman and Company Chapter 16 Outline Why is glucose such a prominent fuel in all life forms? 1. Glucose
Tymoczko Berg Stryer Biochemistry: A Short Course Second Edition CHAPTER 16 Glycolysis 2013 W. H. Freeman and Company Chapter 16 Outline Why is glucose such a prominent fuel in all life forms? 1. Glucose
Chapter 9 Overview. Aerobic Metabolism I: The Citric Acid Cycle. Live processes - series of oxidation-reduction reactions. Aerobic metabolism I
 n n Chapter 9 Overview Aerobic Metabolism I: The Citric Acid Cycle Live processes - series of oxidation-reduction reactions Ingestion of proteins, carbohydrates, lipids Provide basic building blocks for
n n Chapter 9 Overview Aerobic Metabolism I: The Citric Acid Cycle Live processes - series of oxidation-reduction reactions Ingestion of proteins, carbohydrates, lipids Provide basic building blocks for
Notes CELLULAR RESPIRATION SUMMARY EQUATION C 6 H 12 O 6 + O 2. 6CO 2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION
 AP BIOLOGY CELLULAR ENERGETICS ACTIVITY #2 Notes NAME DATE HOUR SUMMARY EQUATION CELLULAR RESPIRATION C 6 H 12 O 6 + O 2 6CO 2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION Oxidation: partial or complete
AP BIOLOGY CELLULAR ENERGETICS ACTIVITY #2 Notes NAME DATE HOUR SUMMARY EQUATION CELLULAR RESPIRATION C 6 H 12 O 6 + O 2 6CO 2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION Oxidation: partial or complete
BIOLOGY - CLUTCH CH.9 - RESPIRATION.
 !! www.clutchprep.com CONCEPT: REDOX REACTIONS Redox reaction a chemical reaction that involves the transfer of electrons from one atom to another Oxidation loss of electrons Reduction gain of electrons
!! www.clutchprep.com CONCEPT: REDOX REACTIONS Redox reaction a chemical reaction that involves the transfer of electrons from one atom to another Oxidation loss of electrons Reduction gain of electrons
METABOLISM Biosynthetic Pathways
 METABOLISM Biosynthetic Pathways Metabolism Metabolism involves : Catabolic reactions that break down large, complex molecules to provide energy and smaller molecules. Anabolic reactions that use ATP energy
METABOLISM Biosynthetic Pathways Metabolism Metabolism involves : Catabolic reactions that break down large, complex molecules to provide energy and smaller molecules. Anabolic reactions that use ATP energy
E.coli Core Model: Metabolic Core
 1 E.coli Core Model: Metabolic Core 2 LEARNING OBJECTIVES Each student should be able to: Describe the glycolysis pathway in the core model. Describe the TCA cycle in the core model. Explain gluconeogenesis.
1 E.coli Core Model: Metabolic Core 2 LEARNING OBJECTIVES Each student should be able to: Describe the glycolysis pathway in the core model. Describe the TCA cycle in the core model. Explain gluconeogenesis.
Escherichia, Salmonella, Shigella, Proteus. Make lactic, acetic, succinic, formic acids
 Enterics Emphasize novel pyruvate enzymes Example of free radicals involved in C-C bond cleavage. Gram negative bacteria that ferment sugars to acids and gas. All use glycolysis Mixed acid group: Escherichia,
Enterics Emphasize novel pyruvate enzymes Example of free radicals involved in C-C bond cleavage. Gram negative bacteria that ferment sugars to acids and gas. All use glycolysis Mixed acid group: Escherichia,
III. 6. Test. Respiració cel lular
 III. 6. Test. Respiració cel lular Chapter Questions 1) What is the term for metabolic pathways that release stored energy by breaking down complex molecules? A) anabolic pathways B) catabolic pathways
III. 6. Test. Respiració cel lular Chapter Questions 1) What is the term for metabolic pathways that release stored energy by breaking down complex molecules? A) anabolic pathways B) catabolic pathways
BCH 4054 Chapter 19 Lecture Notes
 BCH 4054 Chapter 19 Lecture Notes 1 Chapter 19 Glycolysis 2 aka = also known as verview of Glycolysis aka The Embden-Meyerhoff Pathway First pathway discovered Common to almost all living cells ccurs in
BCH 4054 Chapter 19 Lecture Notes 1 Chapter 19 Glycolysis 2 aka = also known as verview of Glycolysis aka The Embden-Meyerhoff Pathway First pathway discovered Common to almost all living cells ccurs in
CHAPTER 24: Carbohydrate, Lipid, & Protein Metabolism. General, Organic, & Biological Chemistry Janice Gorzynski Smith
 CHAPTER 24: Carbohydrate, Lipid, & Protein Metabolism General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 24: Carbohydrate, Lipid, & Protein Metabolism Learning Objectives: q Role in
CHAPTER 24: Carbohydrate, Lipid, & Protein Metabolism General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 24: Carbohydrate, Lipid, & Protein Metabolism Learning Objectives: q Role in
CH395G FINAL (3 rd ) EXAM Kitto/Hackert - Fall 2003
 CH395G FINAL (3 rd ) EXAM Kitto/Hackert - Fall 2003 1. A cell in an active, catabolic state has a. a high (ATP/ADP) and a high (NADH/NAD + ) ratio b. a high (ATP/ADP) and a low (NADH/NAD + ) ratio c. a
CH395G FINAL (3 rd ) EXAM Kitto/Hackert - Fall 2003 1. A cell in an active, catabolic state has a. a high (ATP/ADP) and a high (NADH/NAD + ) ratio b. a high (ATP/ADP) and a low (NADH/NAD + ) ratio c. a
Chapter 9 Part A Lecture Notes: Metabolism Generation of Energy Metabolism is fundamental
 Chapter 9 Part A Lecture Notes: Metabolism Generation of Energy Metabolism is fundamental I. Introduction Metabolism is the total of all chemical reactions in the cell A. Catabolism = breakdown of complex
Chapter 9 Part A Lecture Notes: Metabolism Generation of Energy Metabolism is fundamental I. Introduction Metabolism is the total of all chemical reactions in the cell A. Catabolism = breakdown of complex
ANSC/NUTR 618 Lipids & Lipid Metabolism
 I. Overall concepts A. Definitions ANC/NUTR 618 Lipids & Lipid Metabolism 1. De novo synthesis = synthesis from non-fatty acid precursors a. Carbohydrate precursors (glucose, lactate, and pyruvate) b.
I. Overall concepts A. Definitions ANC/NUTR 618 Lipids & Lipid Metabolism 1. De novo synthesis = synthesis from non-fatty acid precursors a. Carbohydrate precursors (glucose, lactate, and pyruvate) b.
MITOCHONDRIA LECTURES OVERVIEW
 1 MITOCHONDRIA LECTURES OVERVIEW A. MITOCHONDRIA LECTURES OVERVIEW Mitochondrial Structure The arrangement of membranes: distinct inner and outer membranes, The location of ATPase, DNA and ribosomes The
1 MITOCHONDRIA LECTURES OVERVIEW A. MITOCHONDRIA LECTURES OVERVIEW Mitochondrial Structure The arrangement of membranes: distinct inner and outer membranes, The location of ATPase, DNA and ribosomes The
Medical Biochemistry Metabolism with Clinical Correlations CARBOHYDRATE METABOLISM
 Medical Biochemistry Metabolism with Clinical Correlations CARBOHYDRATE METABOLISM DIGESTIVE MECHANISM FOR CARBOHYDRATES 1. In the oral cavity - the salivary α-amylase - a single polypeptide chain, stabilized
Medical Biochemistry Metabolism with Clinical Correlations CARBOHYDRATE METABOLISM DIGESTIVE MECHANISM FOR CARBOHYDRATES 1. In the oral cavity - the salivary α-amylase - a single polypeptide chain, stabilized
Chem 109 C. Fall Armen Zakarian Office: Chemistry Bldn 2217
 Chem 109 C Fall 2014 Armen Zakarian ffice: Chemistry Bldn 2217 Chapter 25 o Glycolysis : fates of pyruvate NADH, H + H + C 2 NADH, H + H - (S)-lactic acid lactate dehydrogenase; anaerobic conditions -
Chem 109 C Fall 2014 Armen Zakarian ffice: Chemistry Bldn 2217 Chapter 25 o Glycolysis : fates of pyruvate NADH, H + H + C 2 NADH, H + H - (S)-lactic acid lactate dehydrogenase; anaerobic conditions -
Biology 638 Biochemistry II Exam-2
 Biology 638 Biochemistry II Exam-2 Biol 638, Exam-2 (Code-1) 1. Assume that 16 glucose molecules enter into a liver cell and are attached to a liner glycogen one by one. Later, this glycogen is broken-down
Biology 638 Biochemistry II Exam-2 Biol 638, Exam-2 (Code-1) 1. Assume that 16 glucose molecules enter into a liver cell and are attached to a liner glycogen one by one. Later, this glycogen is broken-down
Citrate Cycle Supplemental Reading
 Citrate Cycle Supplemental Reading Key Concepts - The Citrate Cycle captures energy using redox reactions - Eight enzymatic reactions of the Citrate Cycle - Key control points in the citrate cycle regulate
Citrate Cycle Supplemental Reading Key Concepts - The Citrate Cycle captures energy using redox reactions - Eight enzymatic reactions of the Citrate Cycle - Key control points in the citrate cycle regulate
Ch. 9 Cell Respiration. Title: Oct 15 3:24 PM (1 of 53)
 Ch. 9 Cell Respiration Title: Oct 15 3:24 PM (1 of 53) Essential question: How do cells use stored chemical energy in organic molecules and to generate ATP? Title: Oct 15 3:28 PM (2 of 53) Title: Oct 19
Ch. 9 Cell Respiration Title: Oct 15 3:24 PM (1 of 53) Essential question: How do cells use stored chemical energy in organic molecules and to generate ATP? Title: Oct 15 3:28 PM (2 of 53) Title: Oct 19
Answer three from questions 5, 6, 7, 8, and 9.
 BCH 4053 May 1, 2003 FINAL EXAM NAME There are 9 pages and 9 questions on the exam. nly five are to be answered, each worth 20 points. Answer two from questions 1, 2, 3, and 4 Answer three from questions
BCH 4053 May 1, 2003 FINAL EXAM NAME There are 9 pages and 9 questions on the exam. nly five are to be answered, each worth 20 points. Answer two from questions 1, 2, 3, and 4 Answer three from questions
Metabolic Pathways and Energy Metabolism
 Metabolic Pathways and Energy Metabolism Last Week Energy Metabolism - The first thing a living organism has got to be able to do is harness energy from the environment - Plants do it by absorbing sunlight
Metabolic Pathways and Energy Metabolism Last Week Energy Metabolism - The first thing a living organism has got to be able to do is harness energy from the environment - Plants do it by absorbing sunlight
2. What is molecular oxygen directly converted into? a. Carbon Dioxide b. Water c. Glucose d. None of the Above
 Biochem 1 Mock Exam 3 Chapter 11: 1. What is glucose completely oxidized into? a. Carbon Dioxide and Water b. Carbon Dioxide and Oxygen c. Oxygen and Water d. Water and Glycogen 2. What is molecular oxygen
Biochem 1 Mock Exam 3 Chapter 11: 1. What is glucose completely oxidized into? a. Carbon Dioxide and Water b. Carbon Dioxide and Oxygen c. Oxygen and Water d. Water and Glycogen 2. What is molecular oxygen
In glycolysis, glucose is converted to pyruvate. If the pyruvate is reduced to lactate, the pathway does not require O 2 and is called anaerobic
 Glycolysis 1 In glycolysis, glucose is converted to pyruvate. If the pyruvate is reduced to lactate, the pathway does not require O 2 and is called anaerobic glycolysis. If this pyruvate is converted instead
Glycolysis 1 In glycolysis, glucose is converted to pyruvate. If the pyruvate is reduced to lactate, the pathway does not require O 2 and is called anaerobic glycolysis. If this pyruvate is converted instead
Citrate Cycle. Lecture 28. Key Concepts. The Citrate Cycle captures energy using redox reactions
 Citrate Cycle Lecture 28 Key Concepts The Citrate Cycle captures energy using redox reactions Eight reactions of the Citrate Cycle Key control points in the Citrate Cycle regulate metabolic flux What role
Citrate Cycle Lecture 28 Key Concepts The Citrate Cycle captures energy using redox reactions Eight reactions of the Citrate Cycle Key control points in the Citrate Cycle regulate metabolic flux What role
