A basic helix loop helix transcription factor controls cell growth
|
|
|
- Madeline Fields
- 5 years ago
- Views:
Transcription
1 A basic helix loop helix transcription factor controls cell growth and size in root hairs Keke Yi 1,2, Benoît Menand 1,3, Elizabeth Bell 1, Liam Dolan 1,4 Supplementary note Low soil phosphate availability (low phosphate stress) induces the elongation of root hairs 1 and enhances the capture of phosphate for plant growth 2. Plants grown in medium containing 10 µm phosphate are considered to be subject to low phosphate stress and root hairs are longer (698 ± 6 µm) than on plants grown with an adequate supply of phosphate (1000 µm phosphate) which develop shorter root hairs (338 ± 6 µm). To demonstrate a requirement for RSL4 in the response to low phosphate, we compared root hair growth of rhd6-3 rsl1-1 double mutants and rhd6-3 rsl1-1 rsl4-1 triple mutants in low phosphate. Low phosphate induces the development of a few hairs in the rhd6-3 rsl1-1 mutant indicating that hairs could respond to the low phosphate growth stimulus. In contrast no hairs developed on the rhd6-3 rsl1-1 rsl4-1 triple mutant (Supplementary Fig. 8a). This indicates that RSL4 is required for the increase in root hair growth that occurs when roots are grown in low phosphate. Further support for the role of RSL4 comes from the observation that low phosphate stress did not induce growth in rsl2-1 rsl4-1 double mutant (Supplementary Fig. 8a). Together, these data indicate that RSL4 activity is required for the root hair growth response induced by low phosphate stress. To confirm that low phosphate stress controls growth by modulating RSL4 activity, we determined the RSL4 transcript and protein levels in roots grown under phosphate stress and
2 control conditions. The levels of RSL4 transcript were 2 to 3 times higher in plants grown in conditions where phosphate was in short supply compared to those grown in high phosphate, while the transcript level of RSL2 did not change (Supplementary Fig. 8b). Furthermore, the steady state levels of GFP-RSL4 protein increased under the low phosphate stress, while the levels of GFP-RSL2 remain unchanged (Supplementary Fig. 8c). To verify that the induction of RSL4 was due to low phosphate and not another stress, we examined RSL4 induction under low phosphate stress in plants with defective low phosphate stress root responses 3. LPR1 and LPR2 are multicopper oxidases involved in triggering the root response to low phosphate stress signal 3. The root hair response to low phosphate is partially repressed in the lpr1 lpr2 double mutant (a gift from Thierry Desnos, CEA Cadarache, France); wild type root hairs grown in low phosphate stress are 100% longer than wild type hairs grown in control conditions. In contrast, the root hairs of lpr1 lpr2 double mutants are only 40% longer in low phosphate than in high phosphate (lpr1 lpr2 HP: 350 ± 6 μm; lpr1 lpr2 LP: 489 ± 6 μm). As expected, the induction of RSL4 mrna by low phosphate stress is defective in the lpr1 lpr2 double mutant background (Supplementary Fig. 6b). These data indicate that low phosphate stress controls root hair length by modulating steady state levels of RSL4 transcript and protein and requires the phosphate signalling activity of LPR1 and LPR2. 1. Bates, T.R. & Lynch, J.P. Stimulation of root hair elongation in Arabidopsis thaliana by low phosphorus availability. Plant, Cell & Environment 19, (1996). 2. Bates, T.R. & Lynch, J.P. The efficiency of Arabidopsis thaliana (Brassicaceae) root hairs in phosphorus acquisition. Am.J.Bot. 87, (2000). 3. Svistoonoff, S. et al. Root tip contact with low-phosphate media reprograms plant root architecture. Nat Genet 39, (2007).
3 Supplementary Fig. 1
4 Supplementary Fig. 1 Expression pattern of RSL4. (a) The distribution of RSL4 protein partially overlaps with RHD6. From left to right are GFP-RSL4, mcherry-rhd6, overlay and bright field; (b) RSL4 mrna was only present in the roots (R). No detectable mrna transcript was present in the rosette leaf (RL), cauline leaf (CL) stem (St) and flower (F); (c) GUS in an RSL4 gene trap line (CSHL_GT13756 from Cold Spring Harbor Laboratory) was detected only in the root hair cells, not in any other tissues including shoots, leaves and flowers.
5 Supplementary Fig. 2 Supplementary Fig. 2 qrt-pcr analysis of RSL4 relative levels in the RSL4 RNAi lines. Values represent mean±s.d., n=3.
6 Supplementary Fig. 3 Supplementary Fig. 3: Constitutive of RSL4 results in constitutive growth until root hair cells undergo programmed cell death. (a) Root hair phenotype of Col-0 (left side) and a plant transformed with 35S:RSL4 (right side) grown in half strength Johnson media. (b) Gray scale pictures are DIC images of root hairs of Col-0 (left side) and 35S:RSL4 (right side). Colour images above each DIC image are pseudo colour of root hairs grown in half strength Johnson media with 10 nm Rhodamine 123 showing the mitochondrial membrane potential ( ψm). The ruler above indicates the fluorescent intensity of the pseudo colour images. The absence of fluorescence intensity is an indicator of cell death.
7 Supplementary Fig. 4 Supplementary Fig. 4 Morphologies of 35S::RSL4 plants. Wild type (left) and 35S::RSL4 (right) plants are morphologically identical except that the root hairs are much longer in the 35S::RSL4 plants.
8 Supplementary Fig. 5 Supplementary Fig. 5: Nuclear volume is the same in wild type and in plants transformed with 35S::RSL4. DAPI staining of Col-0 and plants transformed with 35S::RSL4 root hairs indicates that the nuclear size is the same in these two genotypes.
9 Supplementary Fig. 6 Supplementary Fig. 6: RSL2 regulates root hair tip growth. (a) Root hair morphology of Col-0 (the left panel) and rsl2-1 (the right panel). (b) Root hair length distributions of Col-0 and rsl2-1. (c) GFP:RSL2 accumulates in initiating and growing root hair cells similarly to GFP:RSL4. (d) RSL2 is not a direct target of RHD6. qrt-pcr analysis of RNAs from rhd6-3 rsl1-1 harboring GR:RHD6 treated with DEX for 2h (open bar), 24h (black bar) and DEX/CHX for 2h (hatched bar).
10 Supplementary Fig. 7 Supplementary Fig. 7: RSL2 and RSL4 regulate root hair development. (a) Cryo-SEM images of Col-0 (left panel) and rsl2-1 rsl4-1 (right panel); (b) Genomic DNA fragments of RSL2 and RSL4 complement the rsl2 and rsl4 mutations.
11 Supplementary Fig. 8 Supplementary Fig. 8: Low phosphate stress modulates RSL4 to control hair cell growth. (a) Root hair morphologies of Col-0, rhd6-3 rsl1-1, rhd6-3 rsl1-1 rsl4-1 and rsl2-1 rsl4-1 under HP and LP treatments show that LP stress can induce root hair growth in Col-0 and rhd6-3 rsl1-1, but not in rhd6-3 rsl1-1 rsl4-1 and rsl2-1 rsl4-1. (b) qrt-pcr analysis of mrnas isolated from Col-0 and the lpr1 lpr2 double mutant under high phosphate and low phosphate treatments show that the induction of RSL4 transcripts by low phosphate stress is compromised in the lpr1 lpr2 double mutant background. Values represent mean±s.d., n=3. (c) Pseudo colour images of GFP:RSL2 (upper panel) and GFP:RSL4 (lower panel) under high phosphate and low phosphate treatments showing low phosphate stress induces the accumulation of RSL4 but not RSL2.
12 Supplementary Fig. 9 Supplementary Fig. 9 RSL4 positively regulates the of genes that promote growth in root hair cells. (a) Hierarchical clustering of the 83 genes positively regulated by RSL4. (b) qrt-pcr analysis of AtEXP7, MRH3 and MRH4 from mrnas isolated from plants with wild type levels of RSL4 (Col-0, rsl2-1), higher levels of RSL4 (35S::RSL4) or plants that lack RSL4 (rsl4-1 and rsl2-1 rsl4-1). These data show that RSL4 positively regulates AtEXP 7, MRH3 and MRH4. Values represent mean±s.d., n=3.
13 Supplementary Fig. 10 Supplementary Fig. 10 A working model showing that RSL4 integrates the internal and external signals to regulate root hair cell growth. The left diagram is a SEM picture of Arabidopsis primary root. The white dot line marks the area where RHD6 and RSL1 are expressed in the root hair cells, while the white line marks the area where RSL2 and RSL4 are expressed in the root hair cells. The partial overlapping of these two white lines indicates the slight overlapping of RHD6 with RSL2 and RSL4. The black arrows indicate the genetic relationships.
14 Supplementary Table 1 RSL4 positively regulated genes Probe Set ID Fold change (35S-RSL4 vs P.Value Fold change (Col-0 vs P.Value AGI Gene Symbol Related to root hair or not Col-0) rsl4-1) _at E E-08 AT1G27740 RSL _at E AT1G26420 FAD-binding domain-containing protein _at E-06 AT5G51870 AGL _at AT1G34330 putative peroxidase _at AT5G26080 proline-rich family protein _at AT4G28850 xyloglucan:xyloglucosyl transferase, putative _at AT1G34510 peroxidase, putative _at E AT1G12560 ATEXPA7 root hair _at E-05 AT1G30850 RHS4 root hair _at E AT1G34540 CYP94D _at AT5G22410 RHS18 root hair _at AT3G63380 calcium-transporting ATPase _at AT2G20520 FLA6
15 256352_at E-06 AT1G54970 RHS7/ATPRP1 root hair _at AT4G19680 IRT _at AT2G03980 GDSL-motif lipase/hydrolase family protein _at AT5G19790 RAP2.11 root hair _at AT5G67400 RHS19 root hair _at E AT4G25220 RHS15 root hair _at E-08 AT5G57530 xyloglucan:xyloglucosyl transferase, putative _at AT1G61080 proline-rich family protein _at AT3G10710 RHS12 root hair _s_at AT4G22080 /// AT4G22090 RHS14 root hair _s_at AT1G05240 /// peroxidase, putative
16 AT1G _at AT1G26250 proline-rich extensin, putative _at AT4G02270 RHS13 root hair _at E-05 AT4G25790 allergen V5/Tpx-1-related family protein _at AT3G60280 UCC _at AT5G61650 CYCP4; _at AT4G30320 allergen V5/Tpx-1-related family protein _at AT5G40860 unknown protein _at AT5G19800 hydroxyproline-rich glycoprotein family protein _at AT4G04900 RIC _at AT1G35330 zinc finger (C3HC4-type RING finger) family protein _at AT5G43580 Predicted to encode a PR peptide _at AT4G40090 AGP _at AT2G25240 serine-type endopeptidase inhibitor _at AT3G53150 UGT73D _at AT4G25820 XTR _at AT3G02240 unknown protein _at E-05 AT5G61350 protein kinase family protein _at AT1G05340 unknown protein
17 250683_x_at AT5G06640 proline-rich extensin-like family protein _at AT5G18910 protein kinase family protein _at AT1G16370 ATOCT _at E-06 AT5G57540 xyloglucan:xyloglucosyl transferase, putative _at AT2G29450 ATGSTU _at AT2G03720 MRH6 defective in root hair growth _at AT3G01175 unknown protein _at AT3G62680 PRP3 root hair _at AT4G03330 SYP123 root hair _at E AT1G62980 ATEXPA18 root hair _at AT5G24880 unknown protein _at AT1G59850 binding, biological process unknown _at AT5G23030 TET _at AT1G25240 epsin N-terminal homology (ENTH) domain-containing protein _at AT1G31750 proline-rich family protein _at AT1G34760 RHS5/GRF11 root hair
18 248441_at AT5G51270 protein kinase family protein _at AT4G34580 COW1 defective in root hair growth _at E-07 AT4G13390 proline-rich extensin-like family protein _s_at AT4G01820 PGP5 /// AT4G _at E-05 AT5G61550 protein kinase family protein _at E-05 AT2G46860 ATPPA _at AT1G14950 major latex protein-related / MLP-related _at AT5G03640 protein kinase family protein _at AT2G39690 unknown protein _at AT1G24030 protein kinase family protein _x_at AT4G08410 proline-rich extensin-like family protein _at AT3G12540 unknown protein _at AT1G21130 O-methyltransferase, putative _at AT1G30870 cationic peroxidase _at E-06 AT1G17180 ATGSTU _at AT1G63600 protein kinase-related _at AT1G22880 ATCEL _at AT1G51880 RHS6 root hair
19 255632_at AT4G00680 ADF8 root hair _at AT5G49700 DNA-binding protein-related _at E-05 AT3G07070 protein kinase family protein _at E-05 AT2G38500 unknown protein _at AT2G02990 RNS1 /// AT2G _at AT5G41730 protein kinase family protein _at E-05 AT5G07450 CYCP4; _at AT2G21880 AtRABG2
20 Supplementary Table 2 Primers used in this study rsl2-1 LB F: CTCGTCCCCAATGGAACAAAGGT rsl2-1 LB R: TAGCATCTGAATTTCATAACCAACTCGATACAC rsl2-1 homozygote F: CTCGTCCCCAATGGAACAAAGGT rsl2-1 homozygote R: CGAACGTGACTCTCCATTTCTCTCA rsl4-1 LB F: GGGAACCGGTATTTTTGTTCGG rsl4-1 LB R: TTGTAAGCCAATGGTGCGTACAT rsl4-1 homozygote F: GAAAGCTTCGGTCACAAGTGTTAAA rsl4-1 homozygote R: TTGTAAGCCAATGGTGCGTACAT 35SRSL4F(KpnI): cgg ggtacc ATGGACGTTTTTGTTGATGGT 35SRSL4R(BamHI): cgc ggatcc TCACATAAGCCGAGACAAAAG P RSL2 ::GFP:RSL2 GFPattB1F GGGG ACA AGT TTG TAC AAA AAA GCA GGC TCA ATG AGT AAA GGA GAA GAA CTT TTC
21 GFPattB2R GGGG AC CAC TTT GTA CAA GAA AGC TGG GTA TTT GTA TAG TTC ATC CAT GCC RSL2P_ATTB4F: GGGG ACA ACT TTG TAT AGA AAA GTT G tgc atg tca cct ttc ttt gc RSL2P _ATTB1R: GGGG AC TGC TTT TTT GTA CAA ACT TG a ttc tcc cat ggc ttc cat t RSL2:3 UTRATTB2F: GGGG ACA GCT TTC TTG TAC AAA GTG G gg agc aac aac ctc gga gga at RSL2:3 UTRATTB3R: GGGG AC AAC TTT GTA TAA TAA AGT TG g cca ctt tta aat gct ttg gac P RSL4 ::GFP:RSL4 RSL4P_ATTB4F: GGGG ACA ACT TTG TAT AGA AAA GTT G at acg cgt tgg gct taa atg RSL4P _ATTB1R: GGGG AC TGC TTT TTT GTA CAA ACT TG c aaa aac gtc cat cgc tct RSL4:3 UTRATTB2F: GGGG ACA GCT TTC TTG TAC AAA GTG G tg gac gtt ttt gtt gat ggt RSL4:3 UTRATTB3R: GGGG AC AAC TTT GTA TAA TAA AGT TG acgatacggtttggtttgactaat P RHD6 ::mcherry:rhd6 mcherryattb1f GGGG ACA AGT TTG TAC AAA AAA GCA GGC TTA atg gtg agc aag ggc gag ga mcherryattb2r GGGG AC CAC TTT GTA CAA GAA AGC TGG GT A CTT GTA CAG CTC GTC CAT GCC G
22 RHD6P_ATTB4F GGGG ACA ACT TTG TAT AGA AAA GTT Gtt ctc aaa gag gga caa gac caa agc cca tga c RHD6P_ATTB1R GGGG AC TGC TTT TTT GTA CAA ACT TGc tag aca cta ata agt ttg ata agt gat ttt ttg t RHD6:3 UTRATTB2F GGGG ACA GCT TTC TTG TAC AAA GTG Gcc atg gca ctc gtt aat gac cat ccc aac gag a RHD6:3 UTRATTB3R GGGG AC AAC TTT GTA TAA TAA AGT TGc tga taa atc gag atc tta ggt atg tcg tcc GR ATTB1F GGGG ACA AGT TTG TAC AAA AAA GCA GGC T CT ATG GAT CCT GAA GCT CGA AAA ACA AA GR ATTB2R GGGG AC CAC TTT GTA CAA GAA AGC TGG GTT TTT TTG ATG AAA CAG AAG CTT TTT G RSL2 genomic DNA RSL2genomicUATTB1: GGGG ACA AGT TTG TAC AAA AAA GCA GGC T tgcatgtcacctttctttgc RSL2genomicLATTB2: GGGG AC CAC TTT GTA CAA GAA AGC TGG GT gccacttttaaatgctttggac RSL4 genomic DNA RSL4genomicUATTB1: GGGG ACA AGT TTG TAC AAA AAA GCA GGC T at acg cgt tgg gct taa atg RSL4genomicLATTB2: GGGG AC CAC TTT GTA CAA GAA AGC TGG GT acgatacggtttggtttgactaat
23 qrt-pcr primers RSL2-F RSL2-R RSL4-F RSL4-R AtEXP7-F AtEXP7-R MRH3-pair1-F MRH3-pair1-R MRH4-pair1-F MRH4-pair1-R EF1α -F EF1α -R TCCCCAATGGAACAAAGGTC TCTCGGTGAGCTGAGACCAA GTGCCAAACGGGACAAAAGT TTGTGATGGAACCCCATGTC ATCCCAGTTGCATACCGAAG TATCCAATTCGTCCGGCTAC GATGACCTAGACCACCACTAT GCCTTCAATTCCAGGACTTGAC CGAGGGTTGGCTCTGTCC GGTGGTGATTGTTGTGTTGAC GGTGGTGGCATCCATCTTGTTACA TGAGCACGCTCTTCTTGCTTTCA
Supplementary Table 3. 3 UTR primer sequences. Primer sequences used to amplify and clone the 3 UTR of each indicated gene are listed.
 Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
c Tuj1(-) apoptotic live 1 DIV 2 DIV 1 DIV 2 DIV Tuj1(+) Tuj1/GFP/DAPI Tuj1 DAPI GFP
 Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplemental Data. Shin et al. Plant Cell. (2012) /tpc YFP N
 MYC YFP N PIF5 YFP C N-TIC TIC Supplemental Data. Shin et al. Plant Cell. ()..5/tpc..95 Supplemental Figure. TIC interacts with MYC in the nucleus. Bimolecular fluorescence complementation assay using
MYC YFP N PIF5 YFP C N-TIC TIC Supplemental Data. Shin et al. Plant Cell. ()..5/tpc..95 Supplemental Figure. TIC interacts with MYC in the nucleus. Bimolecular fluorescence complementation assay using
a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53
 1 2 3 4 5 6 7 8 9 10 Supplementary Figure 1. Induction of p53 LOH by MADM. a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53 mouse revealed increased p53 KO/KO (green,
1 2 3 4 5 6 7 8 9 10 Supplementary Figure 1. Induction of p53 LOH by MADM. a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53 mouse revealed increased p53 KO/KO (green,
Supplementary Document
 Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Figure 1. ROS induces rapid Sod1 nuclear localization in a dosagedependent manner. WT yeast cells (SZy1051) were treated with 4NQO at
 Supplementary Figure 1. ROS induces rapid Sod1 nuclear localization in a dosagedependent manner. WT yeast cells (SZy1051) were treated with 4NQO at different concentrations for 30 min and analyzed for
Supplementary Figure 1. ROS induces rapid Sod1 nuclear localization in a dosagedependent manner. WT yeast cells (SZy1051) were treated with 4NQO at different concentrations for 30 min and analyzed for
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Sherman SI, Wirth LJ, Droz J-P, et al. Motesanib diphosphate
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Sherman SI, Wirth LJ, Droz J-P, et al. Motesanib diphosphate
Supplementary Figure 1 a
 Supplementary Figure a Normalized expression/tbp (A.U.).6... Trip-br transcripts Trans Trans Trans b..5. Trip-br Ctrl LPS Normalized expression/tbp (A.U.) c Trip-br transcripts. adipocytes.... Trans Trans
Supplementary Figure a Normalized expression/tbp (A.U.).6... Trip-br transcripts Trans Trans Trans b..5. Trip-br Ctrl LPS Normalized expression/tbp (A.U.) c Trip-br transcripts. adipocytes.... Trans Trans
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most
 Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most differentially expressed between human synovial fibroblasts
Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most differentially expressed between human synovial fibroblasts
Figure S1. Analysis of genomic and cdna sequences of the targeted regions in WT-KI and
 Figure S1. Analysis of genomic and sequences of the targeted regions in and indicated mutant KI cells, with WT and corresponding mutant sequences underlined. (A) cells; (B) K21E-KI cells; (C) D33A-KI cells;
Figure S1. Analysis of genomic and sequences of the targeted regions in and indicated mutant KI cells, with WT and corresponding mutant sequences underlined. (A) cells; (B) K21E-KI cells; (C) D33A-KI cells;
Supplementary Materials
 Supplementary Materials 1 Supplementary Table 1. List of primers used for quantitative PCR analysis. Gene name Gene symbol Accession IDs Sequence range Product Primer sequences size (bp) β-actin Actb gi
Supplementary Materials 1 Supplementary Table 1. List of primers used for quantitative PCR analysis. Gene name Gene symbol Accession IDs Sequence range Product Primer sequences size (bp) β-actin Actb gi
Abbreviations: P- paraffin-embedded section; C, cryosection; Bio-SA, biotin-streptavidin-conjugated fluorescein amplification.
 Supplementary Table 1. Sequence of primers for real time PCR. Gene Forward primer Reverse primer S25 5 -GTG GTC CAC ACT ACT CTC TGA GTT TC-3 5 - GAC TTT CCG GCA TCC TTC TTC-3 Mafa cds 5 -CTT CAG CAA GGA
Supplementary Table 1. Sequence of primers for real time PCR. Gene Forward primer Reverse primer S25 5 -GTG GTC CAC ACT ACT CTC TGA GTT TC-3 5 - GAC TTT CCG GCA TCC TTC TTC-3 Mafa cds 5 -CTT CAG CAA GGA
Table S1. Oligonucleotides used for the in-house RT-PCR assays targeting the M, H7 or N9. Assay (s) Target Name Sequence (5 3 ) Comments
 SUPPLEMENTAL INFORMATION 2 3 Table S. Oligonucleotides used for the in-house RT-PCR assays targeting the M, H7 or N9 genes. Assay (s) Target Name Sequence (5 3 ) Comments CDC M InfA Forward (NS), CDC M
SUPPLEMENTAL INFORMATION 2 3 Table S. Oligonucleotides used for the in-house RT-PCR assays targeting the M, H7 or N9 genes. Assay (s) Target Name Sequence (5 3 ) Comments CDC M InfA Forward (NS), CDC M
CD31 5'-AGA GAC GGT CTT GTC GCA GT-3' 5 ' -TAC TGG GCT TCG AGA GCA GT-3'
 Table S1. The primer sets used for real-time RT-PCR analysis. Gene Forward Reverse VEGF PDGFB TGF-β MCP-1 5'-GTT GCA GCA TGA ATC TGA GG-3' 5'-GGA GAC TCT TCG AGG AGC ACT T-3' 5'-GAA TCA GGC ATC GAG AGA
Table S1. The primer sets used for real-time RT-PCR analysis. Gene Forward Reverse VEGF PDGFB TGF-β MCP-1 5'-GTT GCA GCA TGA ATC TGA GG-3' 5'-GGA GAC TCT TCG AGG AGC ACT T-3' 5'-GAA TCA GGC ATC GAG AGA
BIOLOGY 621 Identification of the Snorks
 Name: Date: Block: BIOLOGY 621 Identification of the Snorks INTRODUCTION: In this simulation activity, you will examine the DNA sequence of a fictitious organism - the Snork. Snorks were discovered on
Name: Date: Block: BIOLOGY 621 Identification of the Snorks INTRODUCTION: In this simulation activity, you will examine the DNA sequence of a fictitious organism - the Snork. Snorks were discovered on
Citation for published version (APA): Oosterveer, M. H. (2009). Control of metabolic flux by nutrient sensors Groningen: s.n.
 University of Groningen Control of metabolic flux by nutrient sensors Oosterveer, Maaike IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it.
University of Groningen Control of metabolic flux by nutrient sensors Oosterveer, Maaike IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it.
Toluidin-Staining of mast cells Ear tissue was fixed with Carnoy (60% ethanol, 30% chloroform, 10% acetic acid) overnight at 4 C, afterwards
 Toluidin-Staining of mast cells Ear tissue was fixed with Carnoy (60% ethanol, 30% chloroform, 10% acetic acid) overnight at 4 C, afterwards incubated in 100 % ethanol overnight at 4 C and embedded in
Toluidin-Staining of mast cells Ear tissue was fixed with Carnoy (60% ethanol, 30% chloroform, 10% acetic acid) overnight at 4 C, afterwards incubated in 100 % ethanol overnight at 4 C and embedded in
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. H3F3B expression in lung cancer. a. Comparison of H3F3B expression in relapsed and non-relapsed lung cancer patients. b. Prognosis of two groups of lung cancer
Supplementary Figures Supplementary Figure 1. H3F3B expression in lung cancer. a. Comparison of H3F3B expression in relapsed and non-relapsed lung cancer patients. b. Prognosis of two groups of lung cancer
Supplementary Table 2. Conserved regulatory elements in the promoters of CD36.
 Supplementary Table 1. RT-qPCR primers for CD3, PPARg and CEBP. Assay Forward Primer Reverse Primer 1A CAT TTG TGG CCT TGT GCT CTT TGA TGA GTC ACA GAA AGA ATC AAT TC 1B AGG AAA TGA ACT GAT GAG TCA CAG
Supplementary Table 1. RT-qPCR primers for CD3, PPARg and CEBP. Assay Forward Primer Reverse Primer 1A CAT TTG TGG CCT TGT GCT CTT TGA TGA GTC ACA GAA AGA ATC AAT TC 1B AGG AAA TGA ACT GAT GAG TCA CAG
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. Lats1/2 deleted ihbs and ihps showed decreased transcripts of hepatocyte related genes (a and b) Western blots (a) and recombination PCR (b) of control and
Supplementary Figure 1 Supplementary Figure 1. Lats1/2 deleted ihbs and ihps showed decreased transcripts of hepatocyte related genes (a and b) Western blots (a) and recombination PCR (b) of control and
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature05883 SUPPLEMENTARY INFORMATION Supplemental Figure 1 Prostaglandin agonists and antagonists alter runx1/cmyb expression. a-e, Embryos were exposed to (b) PGE2 and (c) PGI2 (20μM) and
doi: 10.1038/nature05883 SUPPLEMENTARY INFORMATION Supplemental Figure 1 Prostaglandin agonists and antagonists alter runx1/cmyb expression. a-e, Embryos were exposed to (b) PGE2 and (c) PGI2 (20μM) and
A smart acid nanosystem for ultrasensitive. live cell mrna imaging by the target-triggered intracellular self-assembly
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2017 A smart ZnO@polydopamine-nucleic acid nanosystem for ultrasensitive live cell mrna imaging
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2017 A smart ZnO@polydopamine-nucleic acid nanosystem for ultrasensitive live cell mrna imaging
Nature Immunology: doi: /ni.3836
 Supplementary Figure 1 Recombinant LIGHT-VTP induces pericyte contractility and endothelial cell activation. (a) Western blot showing purification steps for full length murine LIGHT-VTP (CGKRK) protein:
Supplementary Figure 1 Recombinant LIGHT-VTP induces pericyte contractility and endothelial cell activation. (a) Western blot showing purification steps for full length murine LIGHT-VTP (CGKRK) protein:
Culture Density (OD600) 0.1. Culture Density (OD600) Culture Density (OD600) Culture Density (OD600) Culture Density (OD600)
 A. B. C. D. E. PA JSRI JSRI 2 PA DSAM DSAM 2 DSAM 3 PA LNAP LNAP 2 LNAP 3 PAO Fcor Fcor 2 Fcor 3 PAO Wtho Wtho 2 Wtho 3 Wtho 4 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB
A. B. C. D. E. PA JSRI JSRI 2 PA DSAM DSAM 2 DSAM 3 PA LNAP LNAP 2 LNAP 3 PAO Fcor Fcor 2 Fcor 3 PAO Wtho Wtho 2 Wtho 3 Wtho 4 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB
Supplementary Figure 1a
 Supplementary Figure 1a Hours: E-cadherin TGF-β On TGF-β Off 0 12 24 36 48 24 48 72 Vimentin βactin Fig. S1a. Treatment of AML12 cells with TGF-β induces EMT. Treatment of AML12 cells with TGF-β results
Supplementary Figure 1a Hours: E-cadherin TGF-β On TGF-β Off 0 12 24 36 48 24 48 72 Vimentin βactin Fig. S1a. Treatment of AML12 cells with TGF-β induces EMT. Treatment of AML12 cells with TGF-β results
Astaxanthin prevents and reverses diet-induced insulin resistance and. steatohepatitis in mice: A comparison with vitamin E
 Supplementary Information Astaxanthin prevents and reverses diet-induced insulin resistance and steatohepatitis in mice: A comparison with vitamin E Yinhua Ni, 1,2 Mayumi Nagashimada, 1 Fen Zhuge, 1 Lili
Supplementary Information Astaxanthin prevents and reverses diet-induced insulin resistance and steatohepatitis in mice: A comparison with vitamin E Yinhua Ni, 1,2 Mayumi Nagashimada, 1 Fen Zhuge, 1 Lili
 www.lessonplansinc.com Topic: Protein Synthesis - Sentence Activity Summary: Students will simulate transcription and translation by building a sentence/polypeptide from words/amino acids. Goals & Objectives:
www.lessonplansinc.com Topic: Protein Synthesis - Sentence Activity Summary: Students will simulate transcription and translation by building a sentence/polypeptide from words/amino acids. Goals & Objectives:
SUPPLEMENTARY DATA. Supplementary Table 1. Primer sequences for qrt-pcr
 Supplementary Table 1. Primer sequences for qrt-pcr Gene PRDM16 UCP1 PGC1α Dio2 Elovl3 Cidea Cox8b PPARγ AP2 mttfam CyCs Nampt NRF1 16s-rRNA Hexokinase 2, intron 9 β-actin Primer Sequences 5'-CCA CCA GCG
Supplementary Table 1. Primer sequences for qrt-pcr Gene PRDM16 UCP1 PGC1α Dio2 Elovl3 Cidea Cox8b PPARγ AP2 mttfam CyCs Nampt NRF1 16s-rRNA Hexokinase 2, intron 9 β-actin Primer Sequences 5'-CCA CCA GCG
Supplemental Information. Th17 Lymphocytes Induce Neuronal. Cell Death in a Human ipsc-based. Model of Parkinson's Disease
 Cell Stem Cell, Volume 23 Supplemental Information Th17 Lymphocytes Induce Neuronal Cell Death in a Human ipsc-based Model of Parkinson's Disease Annika Sommer, Franz Maxreiter, Florian Krach, Tanja Fadler,
Cell Stem Cell, Volume 23 Supplemental Information Th17 Lymphocytes Induce Neuronal Cell Death in a Human ipsc-based Model of Parkinson's Disease Annika Sommer, Franz Maxreiter, Florian Krach, Tanja Fadler,
Single-Molecule Analysis of Gene Expression Using Two-Color RNA- Labeling in Live Yeast
 Supplemental Figures, Tables and Results Single-Molecule Analysis of Gene Expression Using Two-Color RNA- Labeling in Live Yeast Sami Hocine 1, Pascal Raymond 2, Daniel Zenklusen 2, Jeffrey A. Chao 1 &
Supplemental Figures, Tables and Results Single-Molecule Analysis of Gene Expression Using Two-Color RNA- Labeling in Live Yeast Sami Hocine 1, Pascal Raymond 2, Daniel Zenklusen 2, Jeffrey A. Chao 1 &
SUPPORTING INFORMATION
 SUPPORTING INFORMATION Biology is different in small volumes: endogenous signals shape phenotype of primary hepatocytes cultured in microfluidic channels Amranul Haque, Pantea Gheibi, Yandong Gao, Elena
SUPPORTING INFORMATION Biology is different in small volumes: endogenous signals shape phenotype of primary hepatocytes cultured in microfluidic channels Amranul Haque, Pantea Gheibi, Yandong Gao, Elena
Phylogenetic analysis of human and chicken importins. Only five of six importins were studied because
 Supplementary Figure S1 Phylogenetic analysis of human and chicken importins. Only five of six importins were studied because importin-α6 was shown to be testis-specific. Human and chicken importin protein
Supplementary Figure S1 Phylogenetic analysis of human and chicken importins. Only five of six importins were studied because importin-α6 was shown to be testis-specific. Human and chicken importin protein
Supplementary Information. Bamboo shoot fiber prevents obesity in mice by. modulating the gut microbiota
 Supplementary Information Bamboo shoot fiber prevents obesity in mice by modulating the gut microbiota Xiufen Li 1,2, Juan Guo 1, Kailong Ji 1,2, and Ping Zhang 1,* 1 Key Laboratory of Tropical Plant Resources
Supplementary Information Bamboo shoot fiber prevents obesity in mice by modulating the gut microbiota Xiufen Li 1,2, Juan Guo 1, Kailong Ji 1,2, and Ping Zhang 1,* 1 Key Laboratory of Tropical Plant Resources
Supporting Information
 Supporting Information Malapeira et al. 10.1073/pnas.1217022110 SI Materials and Methods Plant Material and Growth Conditions. A. thaliana seedlings were stratified at 4 C in the dark for 3 d on Murashige
Supporting Information Malapeira et al. 10.1073/pnas.1217022110 SI Materials and Methods Plant Material and Growth Conditions. A. thaliana seedlings were stratified at 4 C in the dark for 3 d on Murashige
Lezione 10. Sommario. Bioinformatica. Lezione 10: Sintesi proteica Synthesis of proteins Central dogma: DNA makes RNA makes proteins Genetic code
 Lezione 10 Bioinformatica Mauro Ceccanti e Alberto Paoluzzi Lezione 10: Sintesi proteica Synthesis of proteins Dip. Informatica e Automazione Università Roma Tre Dip. Medicina Clinica Università La Sapienza
Lezione 10 Bioinformatica Mauro Ceccanti e Alberto Paoluzzi Lezione 10: Sintesi proteica Synthesis of proteins Dip. Informatica e Automazione Università Roma Tre Dip. Medicina Clinica Università La Sapienza
Cross-talk between mineralocorticoid and angiotensin II signaling for cardiac
 ONLINE SUPPLEMENT TO Crosstalk between mineralocorticoid and angiotensin II signaling for cardiac remodeling An Di ZHANG,,3, Aurelie NGUYEN DINH CAT*,,3, Christelle SOUKASEUM *,,3, Brigitte ESCOUBET, 4,
ONLINE SUPPLEMENT TO Crosstalk between mineralocorticoid and angiotensin II signaling for cardiac remodeling An Di ZHANG,,3, Aurelie NGUYEN DINH CAT*,,3, Christelle SOUKASEUM *,,3, Brigitte ESCOUBET, 4,
Journal of Cell Science Supplementary information. Arl8b +/- Arl8b -/- Inset B. electron density. genotype
 J. Cell Sci. : doi:.4/jcs.59: Supplementary information E9. A Arl8b /- Arl8b -/- Arl8b Arl8b non-specific band Gapdh Tbp E7.5 HE Inset B D Control al am hf C E Arl8b -/- al am hf E8.5 F low middle high
J. Cell Sci. : doi:.4/jcs.59: Supplementary information E9. A Arl8b /- Arl8b -/- Arl8b Arl8b non-specific band Gapdh Tbp E7.5 HE Inset B D Control al am hf C E Arl8b -/- al am hf E8.5 F low middle high
Supplemental Information. Cancer-Associated Fibroblasts Neutralize. the Anti-tumor Effect of CSF1 Receptor Blockade
 Cancer Cell, Volume 32 Supplemental Information Cancer-Associated Fibroblasts Neutralize the Anti-tumor Effect of CSF1 Receptor Blockade by Inducing PMN-MDSC Infiltration of Tumors Vinit Kumar, Laxminarasimha
Cancer Cell, Volume 32 Supplemental Information Cancer-Associated Fibroblasts Neutralize the Anti-tumor Effect of CSF1 Receptor Blockade by Inducing PMN-MDSC Infiltration of Tumors Vinit Kumar, Laxminarasimha
Description of Supplementary Files. File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables
 Description of Supplementary Files File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables Supplementary Figure 1: (A), HCT116 IDH1-WT and IDH1-R132H cells were
Description of Supplementary Files File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables Supplementary Figure 1: (A), HCT116 IDH1-WT and IDH1-R132H cells were
Beta Thalassemia Sami Khuri Department of Computer Science San José State University Spring 2015
 Bioinformatics in Medical Product Development SMPD 287 Three Beta Thalassemia Sami Khuri Department of Computer Science San José State University Hemoglobin Outline Anatomy of a gene Hemoglobinopathies
Bioinformatics in Medical Product Development SMPD 287 Three Beta Thalassemia Sami Khuri Department of Computer Science San José State University Hemoglobin Outline Anatomy of a gene Hemoglobinopathies
Supplementary Figure 1
 Supplementary Figure 1 3 3 3 1 1 Bregma -1.6mm 3 : Bregma Ref) Http://www.mbl.org/atlas165/atlas165_start.html Bregma -.18mm Supplementary Figure 1 Schematic representation of the utilized brain slice
Supplementary Figure 1 3 3 3 1 1 Bregma -1.6mm 3 : Bregma Ref) Http://www.mbl.org/atlas165/atlas165_start.html Bregma -.18mm Supplementary Figure 1 Schematic representation of the utilized brain slice
Supplementary Materials and Methods
 DD2 suppresses tumorigenicity of ovarian cancer cells by limiting cancer stem cell population Chunhua Han et al. Supplementary Materials and Methods Analysis of publicly available datasets: To analyze
DD2 suppresses tumorigenicity of ovarian cancer cells by limiting cancer stem cell population Chunhua Han et al. Supplementary Materials and Methods Analysis of publicly available datasets: To analyze
Baseline clinical characteristics for the 81 CMML patients Routine diagnostic testing and statistical analyses... 3
 Next-Generation Sequencing Technology Reveals a Characteristic Pattern of Molecular Mutations in 72.8% of Chronic Myelomonocytic Leukemia (CMML) by Detecting Frequent Alterations in TET2, CBL, RAS, and
Next-Generation Sequencing Technology Reveals a Characteristic Pattern of Molecular Mutations in 72.8% of Chronic Myelomonocytic Leukemia (CMML) by Detecting Frequent Alterations in TET2, CBL, RAS, and
TetR repressor-based bioreporters for the detection of doxycycline using Escherichia
 Supplementary materials TetR repressor-based bioreporters for the detection of doxycycline using Escherichia coli and Acinetobacter oleivorans Hyerim Hong and Woojun Park * Department of Environmental
Supplementary materials TetR repressor-based bioreporters for the detection of doxycycline using Escherichia coli and Acinetobacter oleivorans Hyerim Hong and Woojun Park * Department of Environmental
BHP 2-7 and Nthy-ori 3-1 cells were grown in RPMI1640 medium (Hyclone) supplemented with 10% fetal bovine serum (Gibco), 2mM L-glutamine, and 100 U/mL
 1 2 3 4 Materials and Methods Cell culture BHP 2-7 and Nthy-ori 3-1 cells were grown in RPMI1640 medium (Hyclone) 5 supplemented with 10% fetal bovine serum (Gibco), 2mM L-glutamine, and 100 U/mL 6 penicillin-streptomycin.
1 2 3 4 Materials and Methods Cell culture BHP 2-7 and Nthy-ori 3-1 cells were grown in RPMI1640 medium (Hyclone) 5 supplemented with 10% fetal bovine serum (Gibco), 2mM L-glutamine, and 100 U/mL 6 penicillin-streptomycin.
Supplementary Figure 1
 Metastatic melanoma Primary melanoma Healthy human skin Supplementary Figure 1 CD22 IgG4 Supplementary Figure 1: Immunohisochemical analysis of CD22+ (left) and IgG4 (right), cells (shown in red and indicated
Metastatic melanoma Primary melanoma Healthy human skin Supplementary Figure 1 CD22 IgG4 Supplementary Figure 1: Immunohisochemical analysis of CD22+ (left) and IgG4 (right), cells (shown in red and indicated
Supporting Information. Mutational analysis of a phenazine biosynthetic gene cluster in
 Supporting Information for Mutational analysis of a phenazine biosynthetic gene cluster in Streptomyces anulatus 9663 Orwah Saleh 1, Katrin Flinspach 1, Lucia Westrich 1, Andreas Kulik 2, Bertolt Gust
Supporting Information for Mutational analysis of a phenazine biosynthetic gene cluster in Streptomyces anulatus 9663 Orwah Saleh 1, Katrin Flinspach 1, Lucia Westrich 1, Andreas Kulik 2, Bertolt Gust
Isolate Sexual Idiomorph Species
 SUPLEMENTARY TABLE 1. Isolate identification, sexual idiomorph and species of each isolate used for MAT locus distribution in Paracoccidioides species. Isolate Sexual Idiomorph Species Pb01 MAT1-1 P. lutzii
SUPLEMENTARY TABLE 1. Isolate identification, sexual idiomorph and species of each isolate used for MAT locus distribution in Paracoccidioides species. Isolate Sexual Idiomorph Species Pb01 MAT1-1 P. lutzii
CIRCRESAHA/2004/098145/R1 - ONLINE 1. Validation by Semi-quantitative Real-Time Reverse Transcription PCR
 CIRCRESAHA/2004/098145/R1 - ONLINE 1 Expanded Materials and Methods Validation by Semi-quantitative Real-Time Reverse Transcription PCR Expression patterns of 13 genes (Online Table 2), selected with respect
CIRCRESAHA/2004/098145/R1 - ONLINE 1 Expanded Materials and Methods Validation by Semi-quantitative Real-Time Reverse Transcription PCR Expression patterns of 13 genes (Online Table 2), selected with respect
Beta Thalassemia Case Study Introduction to Bioinformatics
 Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Relationship of the APOA5/A4/C3/A1 gene cluster and APOB gene polymorphisms with dyslipidemia
 elationship of the APOA5/A4/C3/A1 gene cluster and APOB gene polymorphisms with dyslipidemia H.J. Ou 1, G. Huang 2, W. Liu 3, X.L. Ma 2, Y. Wei 4, T. Zhou 5 and Z.M. Pan 3 1 Department of Neurology, The
elationship of the APOA5/A4/C3/A1 gene cluster and APOB gene polymorphisms with dyslipidemia H.J. Ou 1, G. Huang 2, W. Liu 3, X.L. Ma 2, Y. Wei 4, T. Zhou 5 and Z.M. Pan 3 1 Department of Neurology, The
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1: Cryopreservation alters CD62L expression by CD4 T cells. Freshly isolated (left) or cryopreserved PBMCs (right) were stained with the mix of antibodies described
Supplementary Figure 1 Supplementary Figure 1: Cryopreservation alters CD62L expression by CD4 T cells. Freshly isolated (left) or cryopreserved PBMCs (right) were stained with the mix of antibodies described
Supplementary information
 Supplementary information Unique polypharmacology nuclear receptor modulator blocks inflammatory signaling pathways Mi Ra Chang 1, Anthony Ciesla 1, Timothy S. Strutzenberg 1, Scott J. Novick 1, Yuanjun
Supplementary information Unique polypharmacology nuclear receptor modulator blocks inflammatory signaling pathways Mi Ra Chang 1, Anthony Ciesla 1, Timothy S. Strutzenberg 1, Scott J. Novick 1, Yuanjun
SUPPLEMENTAL METHODS Cell culture RNA extraction and analysis Immunohistochemical analysis and laser capture microdissection (LCM)
 SUPPLEMENTAL METHODS Cell culture Human peripheral blood mononuclear cells were isolated from healthy donors by Ficoll density gradient centrifugation. Monocyte differentiation to resting macrophages ()
SUPPLEMENTAL METHODS Cell culture Human peripheral blood mononuclear cells were isolated from healthy donors by Ficoll density gradient centrifugation. Monocyte differentiation to resting macrophages ()
Plasmids Western blot analysis and immunostaining Flow Cytometry Cell surface biotinylation RNA isolation and cdna synthesis
 Plasmids psuper-retro-s100a10 shrna1 was constructed by cloning the dsdna oligo 5 -GAT CCC CGT GGG CTT CCA GAG CTT CTT TCA AGA GAA GAA GCT CTG GAA GCC CAC TTT TTA-3 and 5 -AGC TTA AAA AGT GGG CTT CCA GAG
Plasmids psuper-retro-s100a10 shrna1 was constructed by cloning the dsdna oligo 5 -GAT CCC CGT GGG CTT CCA GAG CTT CTT TCA AGA GAA GAA GCT CTG GAA GCC CAC TTT TTA-3 and 5 -AGC TTA AAA AGT GGG CTT CCA GAG
Mutation analysis of a Chinese family with oculocutaneous albinism
 /, 2016, Vol. 7, (No. 51), pp: 84981-84988 Mutation analysis of a Chinese family with oculocutaneous albinism Xiong Wang 1, Yaowu Zhu 1, Na Shen 1, Jing Peng 1, Chunyu Wang 1, Haiyi Liu 2, Yanjun Lu 1
/, 2016, Vol. 7, (No. 51), pp: 84981-84988 Mutation analysis of a Chinese family with oculocutaneous albinism Xiong Wang 1, Yaowu Zhu 1, Na Shen 1, Jing Peng 1, Chunyu Wang 1, Haiyi Liu 2, Yanjun Lu 1
Nucleotide Sequence of the Australian Bluetongue Virus Serotype 1 RNA Segment 10
 J. gen. Virol. (1988), 69, 945-949. Printed in Great Britain 945 Key words: BTV/genome segment lo/nucleotide sequence Nucleotide Sequence of the Australian Bluetongue Virus Serotype 1 RNA Segment 10 By
J. gen. Virol. (1988), 69, 945-949. Printed in Great Britain 945 Key words: BTV/genome segment lo/nucleotide sequence Nucleotide Sequence of the Australian Bluetongue Virus Serotype 1 RNA Segment 10 By
SUPPLEMENTARY INFORMATION
 BASELINE ISCHAEMIA a b Phd2 +/- c d Collateral growth and maintenance SMC recruitment SMC proliferation Phd2 +/- NF- B off NF- B on NF- B on NF- B on Endothelial cell Smooth muscle cell Pro-arteriogenic
BASELINE ISCHAEMIA a b Phd2 +/- c d Collateral growth and maintenance SMC recruitment SMC proliferation Phd2 +/- NF- B off NF- B on NF- B on NF- B on Endothelial cell Smooth muscle cell Pro-arteriogenic
Supplementary Information
 Supplementary Information Remodeling of heterochromatin structure slows neuropathological progression and prolongs survival in an animal model of Huntington s disease Junghee Lee, Yu Jin Hwang, Yunha Kim,
Supplementary Information Remodeling of heterochromatin structure slows neuropathological progression and prolongs survival in an animal model of Huntington s disease Junghee Lee, Yu Jin Hwang, Yunha Kim,
Formylpeptide receptor2 contributes to colon epithelial homeostasis, inflammation, and tumorigenesis
 Supplementary Data Formylpeptide receptor2 contributes to colon epithelial homeostasis, inflammation, and tumorigenesis Keqiang Chen, Mingyong Liu, Ying Liu, Teizo Yoshimura, Wei Shen, Yingying Le, Scott
Supplementary Data Formylpeptide receptor2 contributes to colon epithelial homeostasis, inflammation, and tumorigenesis Keqiang Chen, Mingyong Liu, Ying Liu, Teizo Yoshimura, Wei Shen, Yingying Le, Scott
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10743 Supplementary Figures and Legends Supplementary Figure 1. CYP17A1 (red boxes) lies at the intersection of steroid hormone biosynthetic pathways. CYP17A1
SUPPLEMENTARY INFORMATION doi:10.1038/nature10743 Supplementary Figures and Legends Supplementary Figure 1. CYP17A1 (red boxes) lies at the intersection of steroid hormone biosynthetic pathways. CYP17A1
Mechanistic and functional insights into fatty acid activation in Mycobacterium tuberculosis SUPPLEMENTARY INFORMATION
 Mechanistic and functional insights into fatty acid activation in Mycobacterium tuberculosis Pooja Arora 1, Aneesh Goyal 2, Vivek T atarajan 1, Eerappa Rajakumara 2, Priyanka Verma 1, Radhika Gupta 3,
Mechanistic and functional insights into fatty acid activation in Mycobacterium tuberculosis Pooja Arora 1, Aneesh Goyal 2, Vivek T atarajan 1, Eerappa Rajakumara 2, Priyanka Verma 1, Radhika Gupta 3,
SUPPLEMENTAL FIGURE 1
 SUPPLEMENTL FIGURE 1 C Supplemental Figure 1. pproach for removal of snorns from Rpl13a gene. () Wild type Rpl13a exonintron structure is shown, with exo in black and intronic snorns in red rectangles.
SUPPLEMENTL FIGURE 1 C Supplemental Figure 1. pproach for removal of snorns from Rpl13a gene. () Wild type Rpl13a exonintron structure is shown, with exo in black and intronic snorns in red rectangles.
Loyer, et al. microrna-21 contributes to NASH Suppl 1/15
 Loyer, et al. microrna-21 contributes to NASH Suppl 1/15 SUPPLEMENTARY MATERIAL: Liver MicroRNA-21 is Overexpressed in Non Alcoholic Steatohepatitis and Contributes to the Disease in Experimental Models
Loyer, et al. microrna-21 contributes to NASH Suppl 1/15 SUPPLEMENTARY MATERIAL: Liver MicroRNA-21 is Overexpressed in Non Alcoholic Steatohepatitis and Contributes to the Disease in Experimental Models
University of Groningen. Vasoregression in incipient diabetic retinopathy Pfister, Frederick
 University of Groningen Vasoregression in incipient diabetic retinopathy Pfister, Frederick IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from
University of Groningen Vasoregression in incipient diabetic retinopathy Pfister, Frederick IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from
Resistance to Tetracycline Antibiotics by Wangrong Yang, Ian F. Moore, Kalinka P. Koteva, Donald W. Hughes, David C. Bareich and Gerard D. Wright.
 Supplementary Data for TetX is a Flavin-Dependent Monooxygenase Conferring Resistance to Tetracycline Antibiotics by Wangrong Yang, Ian F. Moore, Kalinka P. Koteva, Donald W. Hughes, David C. Bareich and
Supplementary Data for TetX is a Flavin-Dependent Monooxygenase Conferring Resistance to Tetracycline Antibiotics by Wangrong Yang, Ian F. Moore, Kalinka P. Koteva, Donald W. Hughes, David C. Bareich and
Mutation Screening and Association Studies of the Human UCP 3 Gene in Normoglycemic and NIDDM Morbidly Obese Patients
 Mutation Screening and Association Studies of the Human UCP 3 Gene in Normoglycemic and NIDDM Morbidly Obese Patients Shuichi OTABE, Karine CLEMENT, Séverine DUBOIS, Frederic LEPRETRE, Veronique PELLOUX,
Mutation Screening and Association Studies of the Human UCP 3 Gene in Normoglycemic and NIDDM Morbidly Obese Patients Shuichi OTABE, Karine CLEMENT, Séverine DUBOIS, Frederic LEPRETRE, Veronique PELLOUX,
SUPPLEMENTARY RESULTS
 SUPPLEMENTARY RESULTS Supplementary Table 1. hfpr1- Flpln-CHO hfpr2-flpln-cho pec 50 E max (%) Log( /K A) Log( /K A) N pec 50 E max (%) Log( /K A) Log( /K A) n ERK1/2 phosphorylation fmlp 9.0±0.6 80±7
SUPPLEMENTARY RESULTS Supplementary Table 1. hfpr1- Flpln-CHO hfpr2-flpln-cho pec 50 E max (%) Log( /K A) Log( /K A) N pec 50 E max (%) Log( /K A) Log( /K A) n ERK1/2 phosphorylation fmlp 9.0±0.6 80±7
The Clinical Performance of Primary HPV Screening, Primary HPV Screening Plus Cytology Cotesting, and Cytology Alone at a Tertiary Care Hospital
 The Clinical Performance of Primary HPV Screening, Primary HPV Screening Plus Cytology Cotesting, and Cytology Alone at a Tertiary Care Hospital Jung-Woo Choi MD, PhD; Younghye Kim MD, PhD; Ju-Han Lee
The Clinical Performance of Primary HPV Screening, Primary HPV Screening Plus Cytology Cotesting, and Cytology Alone at a Tertiary Care Hospital Jung-Woo Choi MD, PhD; Younghye Kim MD, PhD; Ju-Han Lee
Detection of 549 new HLA alleles in potential stem cell donors from the United States, Poland and Germany
 HLA ISSN 2059-2302 BRIEF COMMUNICATION Detection of 549 new HLA alleles in potential stem cell donors from the United States, Poland and Germany C. J. Hernández-Frederick 1, N. Cereb 2,A.S.Giani 1, J.
HLA ISSN 2059-2302 BRIEF COMMUNICATION Detection of 549 new HLA alleles in potential stem cell donors from the United States, Poland and Germany C. J. Hernández-Frederick 1, N. Cereb 2,A.S.Giani 1, J.
PATIENTS AND METHODS. Subjects
 PATIENTS AND METHODS Subjects Twenty-nine morbidly obese subjects involved in a gastric surgery program were enrolled in the study between October 25 and March 21. Bariatric surgery was performed in patients
PATIENTS AND METHODS Subjects Twenty-nine morbidly obese subjects involved in a gastric surgery program were enrolled in the study between October 25 and March 21. Bariatric surgery was performed in patients
Advanced Subsidiary Unit 1: Lifestyle, Transport, Genes and Health
 Write your name here Surname Other names Edexcel GCE Centre Number Candidate Number Biology Advanced Subsidiary Unit 1: Lifestyle, Transport, Genes and Health Thursday 8 January 2009 Morning Time: 1 hour
Write your name here Surname Other names Edexcel GCE Centre Number Candidate Number Biology Advanced Subsidiary Unit 1: Lifestyle, Transport, Genes and Health Thursday 8 January 2009 Morning Time: 1 hour
The animals were housed in standard cages with ad libitum access to both food and
 Supplementary data: Supplementary Material and Methods Animal preparation The animals were housed in standard cages with ad libitum access to both food and water. During the imaging sessions, the animals
Supplementary data: Supplementary Material and Methods Animal preparation The animals were housed in standard cages with ad libitum access to both food and water. During the imaging sessions, the animals
Enhanced detection and serotyping of Streptococcus pneumoniae using multiplex polymerase chain reaction
 Original article http://dx.doi.org/10.3345/kjp.2012.55.11.424 Korean J Pediatr 2012;55(11):424-429 eissn 1738-1061 pissn 2092-7258 Enhanced detection and serotyping of Streptococcus pneumoniae using multiplex
Original article http://dx.doi.org/10.3345/kjp.2012.55.11.424 Korean J Pediatr 2012;55(11):424-429 eissn 1738-1061 pissn 2092-7258 Enhanced detection and serotyping of Streptococcus pneumoniae using multiplex
McAlpine PERK-GSK3 regulates foam cell formation. Supplemental Material. Supplementary Table I. Sequences of real time PCR primers.
 Mclpine PERK-GSK3 regulates foam cell formation Supplemental Material Supplementary Table I. Sequences of real time PCR primers. Primer Name Primer Sequences (5-3 ) Product Size (bp) GRP78 (human) Fwd:
Mclpine PERK-GSK3 regulates foam cell formation Supplemental Material Supplementary Table I. Sequences of real time PCR primers. Primer Name Primer Sequences (5-3 ) Product Size (bp) GRP78 (human) Fwd:
Integration Solutions
 Integration Solutions (1) a) With no active glycosyltransferase of either type, an ii individual would not be able to add any sugars to the O form of the lipopolysaccharide. Thus, the only lipopolysaccharide
Integration Solutions (1) a) With no active glycosyltransferase of either type, an ii individual would not be able to add any sugars to the O form of the lipopolysaccharide. Thus, the only lipopolysaccharide
*To whom correspondence should be addressed. This PDF file includes:
 www.sciencemag.org/cgi/content/full/science.1212182/dc1 Supporting Online Material for Partial Retraction to Detection of an Infectious Retrovirus, XMRV, in Blood Cells of Patients with Chronic Fatigue
www.sciencemag.org/cgi/content/full/science.1212182/dc1 Supporting Online Material for Partial Retraction to Detection of an Infectious Retrovirus, XMRV, in Blood Cells of Patients with Chronic Fatigue
Supplementary Figure S1
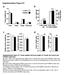 Supplementry Figure S Tissue weights (g).... Liver Hert Brin Pncres Len mss (g) 8 6 -% +% 8 6 Len mss Len mss (g) (% ody weight) Len mss (% ody weight) c Tiilis nterior weight (g).6...... Qudriceps weight
Supplementry Figure S Tissue weights (g).... Liver Hert Brin Pncres Len mss (g) 8 6 -% +% 8 6 Len mss Len mss (g) (% ody weight) Len mss (% ody weight) c Tiilis nterior weight (g).6...... Qudriceps weight
ice-cold 70% ethanol with gentle vortexing, incubated at -20 C for 4 hours, and washed with PBS.
 Cell cycle analysis For cell cycle analysis, single cell suspensions of E12.5 fetal liver cells were suspended in 4 ml ice-cold 7% ethanol with gentle vortexing, incubated at -2 C for 4 hours, and washed
Cell cycle analysis For cell cycle analysis, single cell suspensions of E12.5 fetal liver cells were suspended in 4 ml ice-cold 7% ethanol with gentle vortexing, incubated at -2 C for 4 hours, and washed
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/473/eaai7696/dc1 Supplementary Materials for Astrocyte-shed extracellular vesicles regulate the peripheral leukocyte response to inflammatory brain lesions
www.sciencesignaling.org/cgi/content/full/10/473/eaai7696/dc1 Supplementary Materials for Astrocyte-shed extracellular vesicles regulate the peripheral leukocyte response to inflammatory brain lesions
Cancer Genetics 204 (2011) 45e52
 Cancer Genetics 204 (2011) 45e52 Exon scanning by reverse transcriptaseepolymerase chain reaction for detection of known and novel EML4eALK fusion variants in nonesmall cell lung cancer Heather R. Sanders
Cancer Genetics 204 (2011) 45e52 Exon scanning by reverse transcriptaseepolymerase chain reaction for detection of known and novel EML4eALK fusion variants in nonesmall cell lung cancer Heather R. Sanders
3) Table_S1: Clinical Characteristics of Breast Cancer Patients. 5) Table_S3: Primer sequences used for qt-pcr of ChIP samples
 Supplemental Section: 1) Eight supplemental figures and legends 2) Supplemental Materials and Methods 3) Table_S1: Clinical Characteristics of Breast Cancer Patients 4) Table_S2: Oligonucleotide sequences
Supplemental Section: 1) Eight supplemental figures and legends 2) Supplemental Materials and Methods 3) Table_S1: Clinical Characteristics of Breast Cancer Patients 4) Table_S2: Oligonucleotide sequences
mir-1202: A Primate Specific and Brain Enriched mirna Involved in Major Depression and Antidepressant Treatment. Supplementary Information
 Title: mir-1202: A Primate Specific and Brain Enriched mirna Involved in Major Depression and Antidepressant Treatment. Authors: Juan Pablo Lopez 1, Raymond Lim 3, Cristiana Cruceanu 1, Liam Crapper 1,
Title: mir-1202: A Primate Specific and Brain Enriched mirna Involved in Major Depression and Antidepressant Treatment. Authors: Juan Pablo Lopez 1, Raymond Lim 3, Cristiana Cruceanu 1, Liam Crapper 1,
Expression of Selected Inflammatory Cytokine Genes in Bladder Biopsies
 Borneo Journal of Resource Science and Technology (2013) 3(2): 15-20 Expression of Selected Inflammatory Cytokine Genes in Bladder Biopsies EDMUND UI-HANG SIM *1, NUR DIANA ANUAR 2, TENG-AIK ONG 3, GUAN-
Borneo Journal of Resource Science and Technology (2013) 3(2): 15-20 Expression of Selected Inflammatory Cytokine Genes in Bladder Biopsies EDMUND UI-HANG SIM *1, NUR DIANA ANUAR 2, TENG-AIK ONG 3, GUAN-
Isolation and Genetic Characterization of New Reassortant H3N1 Swine Influenza Virus from Pigs in the Midwestern United States
 JOURNAL OF VIROLOGY, May 2006, p. 5092 5096 Vol. 80, No. 10 0022-538X/06/$08.00 0 doi:10.1128/jvi.80.10.5092 5096.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Isolation
JOURNAL OF VIROLOGY, May 2006, p. 5092 5096 Vol. 80, No. 10 0022-538X/06/$08.00 0 doi:10.1128/jvi.80.10.5092 5096.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Isolation
Supplementary Figure 1: GPCR profiling and G q signaling in murine brown adipocytes (BA). a, Number of GPCRs with 2-fold lower expression in mature
 Supplementary Figure 1: GPCR profiling and G q signaling in murine brown adipocytes (BA). a, Number of GPCRs with 2-fold lower expression in mature BA vs. preadipocytes. b, Number of GPCRs with 2-fold
Supplementary Figure 1: GPCR profiling and G q signaling in murine brown adipocytes (BA). a, Number of GPCRs with 2-fold lower expression in mature BA vs. preadipocytes. b, Number of GPCRs with 2-fold
Viral hepatitis, which affects half a billion people
 GASTROENTEROLOGY 2006;130:435 452 BASIC LIVER, PANCREAS, AND BILIARY TRACT Natural Killer Cells Ameliorate Liver Fibrosis by Killing Activated Stellate Cells in NKG2D-Dependent and Tumor Necrosis Factor
GASTROENTEROLOGY 2006;130:435 452 BASIC LIVER, PANCREAS, AND BILIARY TRACT Natural Killer Cells Ameliorate Liver Fibrosis by Killing Activated Stellate Cells in NKG2D-Dependent and Tumor Necrosis Factor
Supplementary Material Hofko M et al., Detection of carbapenemases by real-time PCR and melt-curve analysis on the BD MAX TM System
 Supplementary Material Hofko M et al., Detection of carbapenemases by real-time PCR and melt-curve analysis on the BD MAX TM System Supplementary Material and Methods Characterization of isolates by the
Supplementary Material Hofko M et al., Detection of carbapenemases by real-time PCR and melt-curve analysis on the BD MAX TM System Supplementary Material and Methods Characterization of isolates by the
Bacterial Gene Finding CMSC 423
 Bacterial Gene Finding CMSC 423 Finding Signals in DNA We just have a long string of A, C, G, Ts. How can we find the signals encoded in it? Suppose you encountered a language you didn t know. How would
Bacterial Gene Finding CMSC 423 Finding Signals in DNA We just have a long string of A, C, G, Ts. How can we find the signals encoded in it? Suppose you encountered a language you didn t know. How would
Supplemental Figures: Supplemental Figure 1
 Supplemental Figures: Supplemental Figure 1 Suppl. Figure 1. BM-DC infection with H. pylori does not induce cytotoxicity and treatment of BM-DCs with H. pylori sonicate, but not heat-inactivated bacteria,
Supplemental Figures: Supplemental Figure 1 Suppl. Figure 1. BM-DC infection with H. pylori does not induce cytotoxicity and treatment of BM-DCs with H. pylori sonicate, but not heat-inactivated bacteria,
Ho Young Jung Hye Seung Han Hyo Bin Kim 1 Seo Young Oh 1 Sun-Joo Lee 2 Wook Youn Kim
 Journal of Pathology and Translational Medicine 2016; 50: 138-146 ORIGINAL ARTICLE Comparison of Analytical and Clinical Performance of HPV 9G DNA Chip, PANArray HPV Genotyping Chip, and Hybrid-Capture
Journal of Pathology and Translational Medicine 2016; 50: 138-146 ORIGINAL ARTICLE Comparison of Analytical and Clinical Performance of HPV 9G DNA Chip, PANArray HPV Genotyping Chip, and Hybrid-Capture
HCV Persistence and Immune Evasion in the Absence of Memory T Cell Help.
 SOM Text HCV Persistence and Immune Evasion in the Absence of Memory T Cell Help. Arash Grakoui 1, Naglaa H. Shoukry 2, David J. Woollard 2, Jin-Hwan Han 1, Holly L. Hanson 1, John Ghrayeb 3, Krishna K.
SOM Text HCV Persistence and Immune Evasion in the Absence of Memory T Cell Help. Arash Grakoui 1, Naglaa H. Shoukry 2, David J. Woollard 2, Jin-Hwan Han 1, Holly L. Hanson 1, John Ghrayeb 3, Krishna K.
Nomenclature What is in a name? My name Joseˊ Jimenez = Bill Dana John L.C. Savony = Frank Fontaine
 Nomenclature What is in a name? My name Joseˊ Jimenez = Bill Dana John L.C. Savony = Frank Fontaine Three categories of Amyloid Terms 1. Amyloidologists, Amyloid Centers, Researchers official nomenclature
Nomenclature What is in a name? My name Joseˊ Jimenez = Bill Dana John L.C. Savony = Frank Fontaine Three categories of Amyloid Terms 1. Amyloidologists, Amyloid Centers, Researchers official nomenclature
without LOI phenotype by breeding female wild-type C57BL/6J and male H19 +/.
 Sakatani et al. 1 Supporting Online Material Materials and methods Mice and genotyping: H19 mutant mice with C57BL/6J background carrying a deletion in the structural H19 gene (3 kb) and 10 kb of 5 flanking
Sakatani et al. 1 Supporting Online Material Materials and methods Mice and genotyping: H19 mutant mice with C57BL/6J background carrying a deletion in the structural H19 gene (3 kb) and 10 kb of 5 flanking
What do you think of when you here the word genome?
 What do you think of when you here the word genome? What do you think of when you here the word genome? Personal Genomics Outline Review of pre-lab work Genomics and Medicine Case Overview & Assignment
What do you think of when you here the word genome? What do you think of when you here the word genome? Personal Genomics Outline Review of pre-lab work Genomics and Medicine Case Overview & Assignment
Development of RT-qPCR-based molecular diagnostic assays for therapeutic target selection of breast cancer patients
 Development of RT-qPCR-based molecular diagnostic assays for therapeutic target selection of breast cancer patients Sangjung Park The Graduate School Yonsei University Department of Biomedical Laboratory
Development of RT-qPCR-based molecular diagnostic assays for therapeutic target selection of breast cancer patients Sangjung Park The Graduate School Yonsei University Department of Biomedical Laboratory
Nucleotide diversity of the TNF gene region in an African village
 (2001) 2, 343 348 2001 Nature Publishing Group All rights reserved 1466-4879/01 $15.00 www.nature.com/gene Nucleotide diversity of the TNF gene region in an African village A Richardson 1, F Sisay-Joof
(2001) 2, 343 348 2001 Nature Publishing Group All rights reserved 1466-4879/01 $15.00 www.nature.com/gene Nucleotide diversity of the TNF gene region in an African village A Richardson 1, F Sisay-Joof
Genome-wide identification of TCF7L2/TCF4 target mirnas reveals a role for mir-21 in Wnt-driven epithelial cancer
 INTERNATIONAL JOURNAL OF ONCOLOGY 40: 519-526, 2012 Genome-wide identification of TCF7L2/TCF4 target mirnas reveals a role for mir-21 in Wnt-driven epithelial cancer FENGMING LAN 1-3*, XIAO YUE 1-3,5*,
INTERNATIONAL JOURNAL OF ONCOLOGY 40: 519-526, 2012 Genome-wide identification of TCF7L2/TCF4 target mirnas reveals a role for mir-21 in Wnt-driven epithelial cancer FENGMING LAN 1-3*, XIAO YUE 1-3,5*,
Supplementary Data. Clinical Setup. References. Program for Embryo Donation
 Supplementary Data Clinical Setup Notwithstanding their value in stem cell research, the source of supernumerary human blastocysts for the derivation of new stem cell lines and the modalities of their
Supplementary Data Clinical Setup Notwithstanding their value in stem cell research, the source of supernumerary human blastocysts for the derivation of new stem cell lines and the modalities of their
