Supplementary Figures. Supplementary Figure 1. Dinucleotide variant proportions. These are described and quantitated. for each lesion type.
|
|
|
- Mercy Harrison
- 5 years ago
- Views:
Transcription
1 Supplementary Figures Supplementary Figure 1. Dinucleotide variant proportions. These are described and quantitated for each lesion type.
2 a b Supplementary Figure 2. Non-negative matrix factorization-derived orthogonal mutational profiles. (a) Heatmap of NMF-derived orthogonal mutational profiles derived from over 6,000 human cancers for the human exome samples. The scale bar refers to the proportion of mutations attributable to a given signature. The clustering confirms a strong enrichment for CpG-associated C T transitions classically associated with UVB-exposure (signature 14), particularly for AK and cuscc, with segregation of most of the NS samples away from AK and cuscc, with the exception of 1-NS, which has the highest mutational load in that class. Two other profiles dominated by C T transitions are significantly represented in the mutational data and relatively enriched in NS (signatures 10, 15), including one first described in the context of temozolamide exposure (signature 15). (b) Table of NMF Cluster Definitions. Our strategy employed non-smooth NMF, a variant which approximates the data using the basis and
3 coefficient matrices as above with the addition of a third smoothing matrix which serves to absorb noise within the data, thus driving the coefficient and basis matrices to increase sparseness. The resulting basis matrix generated k=21 signatures from a diverse set of over 6000 cancers 39, reducing correlations between signatures originally derived by Alexandrov et al. 72 thereby increasing orthogonality.
4 Supplementary Figure 3. Histograms of variant allele frequencies of samples grouped by patient.
5 Supplementary Figure 4. Enumeration of overlaps between mutated genes and variant positions within mutated genes within samples. Overlaps in non-silent SNVs at the gene and position level are enumerated here.
6 a b c
7 Supplementary Figure 5. Analysis of mrna expression in human and mouse lesions. (a) Principal component analysis of unsupervised mrna expression. Using principal component analysis across three dimensions as an alternative means of organizing the transcriptional profiles, the results of the unsupervised clustering are largely recapitulated: cuscc and AK/PAP segregate away from NS most prominently. (b) Clustering of recurrently differentially expressed human genes in at least 2 of 3 possible pairwise comparisons between NS, AK, and cuscc shows that AKs cluster predominantly with cuscc, consistent with the notion that they are largely transcriptomically more similar to cuscc. (c) Clustering of recurrently differentially expressed mouse genes in at least 2 of 3 possible pairwise comparisons between CHR, PAP, and cuscc shows that PAP cluster predominantly with cuscc, consistent with the notion that they are largely transcriptomically more similar to cuscc.
8 a b c d e f g h i
9 Supplementary Figure 6. TRANSFAC based analysis of transcription factor regulators. (a) ETS2 targets and (b) SP1 target are upregulated across the continuum of samples. (c) TCF3 targets that are overall downregulated have greater significance across the continuum than those that are upregulated in the late stage transition (AK/PAP to cuscc). (d) LEF1 targets also exhibit a similar mixed pattern, with one subset of target genes with significant downregulation early (NS/CHR to AK/PAP) and a distinct subset showing upregulation late (AK/PAP to cuscc). The majority of NFAT targets (e) also exhibit downregulation early. (f) AP1 and (g) FREAC2 (FOXF2) targets are downregulated across the continuum of cuscc development. Targets of (h) E2F and (i) NFY are significantly upregulated early (NS to AK/PAP) and these were also highlighted in the linear mixed-effects (LME) model of progression which are highlighted in red in panel (Fig. 5a).
10 Supplementary Figure 7. Principal component analysis of unsupervised microrna expression. Using principal component analysis across three dimensions as an alternative means of organizing the microrna profiles, the results of the unsupervised clustering are largely recapitulated: while NS/CHR are very distinctly separated from cuscc, segregation of AK/PAP as a distinct group is more clearly seen here as compared to the principal component analysis of mrna profiles, particularly in mouse.
11 a b Supplementary Figure 8. Survival analysis for head & neck squamous cell carcinoma. (a) All of the individual signatures for all significant pairwise comparisons in human and mouse were probed for their ability to predict survival in TP53-mutant HNSCC in the top and bottom 25% of ranked signatures; none was significant. (b) The remaining late transition and stepwise (both
12 early and late) signatures from the LME model data were probed for their ability to predict survival in HNSCC; none was significant.
13 Supplementary Tables Gene # Patients Removed as common false positive genes: GPR98 4 PT1,PT2,PT3,PT4 Gene # Patients MYO7B 4 PT1,PT3,PT4,PT5 TTN 7 PT1,PT2,PT3,PT4,PT5,PT6,PT8 RELN 4 PT1,PT3,PT4,PT5 MUC16 6 PT1,PT2,PT3,PT4,PT5,PT6 TP53 4 PT1,PT3,PT4,PT5 CSMD3 3 PT2,PT3,PT5 CRB1 3 PT2,PT3,PT6 DNAH6 3 PT2,PT3,PT6 FAT1 3 PT3,PT4,PT5 DNAH7 3 PT3,PT5,PT6 FSIP2 3 PT2,PT3,PT5 HYDIN 3 PT3,PT5,PT6 IGFN1 3 PT2,PT3,PT5 LRP1B 3 PT1,PT3,PT5 MLL3 3 PT2,PT5,PT6 MUC4 3 PT2,PT3,PT5 RYR1 3 PT2,PT4,PT5 SYNE1 3 PT3,PT4,PT5 VWDE 3 PT2,PT3,PT5 ADAMTS20 2 PT2,PT4 ADAM29 2 PT3,PT4 CSMD1 2 PT3,PT5 ADGB 2 PT2,PT3 CTTNBP2 2 PT5,PT6 ANKRD31 2 PT3,PT6 DNAH10 2 PT1,PT5 ATP8A1 2 PT3,PT5 DNAH14 2 PT3,PT5 BTBD9 2 PT3,PT5 DNAH5 2 PT2,PT4 C16orf62 2 PT3,PT4 DNAH8 2 PT3,PT5 C2orf16 2 PT4,PT5 KALRN 2 PT3,PT5 CCDC168 2 PT3,PT5 OR4K17 2 PT3,PT5 CHD8 2 PT1,PT4 PCLO 2 PT1,PT5 CLTCL1 2 PT3,PT6 CMYA5 2 PT4,PT5 COL11A1 2 PT4,PT5 COL1A2 2 PT3,PT5 COL24A1 2 PT3,PT5 COL27A1 2 PT4,PT5 COL4A2 2 PT2,PT3 COL4A3 2 PT3,PT4 COL6A5 2 PT3,PT5 COL6A6 2 PT2,PT5 CSF3R 2 PT3,PT5 CXorf22 2 PT3,PT4 DDX60L 2 PT3,PT4 DENND4A 2 PT4,PT5 DGKH 2 PT4,PT5 DNHD1 2 PT2,PT5 DPP3 2 PT5,PT6 DYNC1I1 2 PT3,PT5 ENPP3 2 PT2,PT5 FAM135A 2 PT5,PT8 FER1L6 2 PT3,PT5 FGA 2 PT3,PT5 FREM3 2 PT3,PT5 FRY 2 PT3,PT5 GLDN 2 PT5,PT6 HMCN1 2 PT3,PT5 KIAA1324L 2 PT3,PT5 KIAA PT2,PT3 LAMA1 2 PT3,PT4 LAMC3 2 PT3,PT6 LGI1 2 PT1,PT3 LRP2 2 PT3,PT5 MLL2 2 PT2,PT5 MYH2 2 PT3,PT5 MYH4 2 PT3,PT5 MYO18B 2 PT3,PT4 NAV3 2 PT2,PT5 NHSL2 2 PT3,PT5 NPAP1 2 PT3,PT5 PALM2-AKAP2 2 PT2,PT3 PAPPA2 2 PT3,PT5 POLR3A 2 PT4,PT5 PRB4 2 PT1,PT5 PRDM9 2 PT4,PT5 PREX1 2 PT2,PT4 PRKD1 2 PT3,PT4 PRR16 2 PT3,PT5 PRRC2A 2 PT5,PT6 PRUNE2 2 PT2,PT3 PTPRQ 2 PT3,PT5 ROS1 2 PT3,PT5 RP1 2 PT3,PT5 SCAND3 2 PT3,PT6 SLC12A3 2 PT2,PT5 SLC26A7 2 PT4,PT6 SPAG17 2 PT3,PT5 STARD9 2 PT1,PT3 TCF20 2 PT5,PT6 THSD7A 2 PT2,PT4 TMEM131 2 PT2,PT5 TRDN 2 PT1,PT3 TRIM58 2 PT3,PT4 TRIM66 2 PT2,PT4 TTC40 2 PT3,PT6 USH2A 2 PT2,PT6 VWF 2 PT3,PT5 WBP2NL 2 PT3,PT4 WBSCR17 2 PT3,PT5 WDR87 2 PT3,PT4 XIRP2 2 PT2,PT3 ZFHX4 2 PT2,PT3 Supplementary Table 1. Table of overlapping mutated genes across patients.
14 MOUSE HUMAN Supplementary Table 2. Ingenuity Pathway Analysis of differentially expressed genes in the early (NS/CHR to AK/PAP) transition. Using the significantly differentially expressed genes in the early transition from NS/CHR to AK/PAP in the linear mixed effects model, IPA analysis was conducted separately for human and mouse genes. There is striking similarity in the key canonical pathways and upstream regulators identified.
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first
 Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10860 Supplementary Discussion It remains unclear why H3K9 demethylation appeared to be more sensitive to suppression than at least some other histone methylation marks as a result of
doi:10.1038/nature10860 Supplementary Discussion It remains unclear why H3K9 demethylation appeared to be more sensitive to suppression than at least some other histone methylation marks as a result of
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
Nature Medicine: doi: /nm.4439
 Figure S1. Overview of the variant calling and verification process. This figure expands on Fig. 1c with details of verified variants identification in 547 additional validation samples. Somatic variants
Figure S1. Overview of the variant calling and verification process. This figure expands on Fig. 1c with details of verified variants identification in 547 additional validation samples. Somatic variants
SUPPLEMENTARY APPENDIX
 SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
Nature Neuroscience: doi: /nn Supplementary Figure 1. Missense damaging predictions as a function of allele frequency
 Supplementary Figure 1 Missense damaging predictions as a function of allele frequency Percentage of missense variants classified as damaging by eight different classifiers and a classifier consisting
Supplementary Figure 1 Missense damaging predictions as a function of allele frequency Percentage of missense variants classified as damaging by eight different classifiers and a classifier consisting
stomach p63/krt5 CDH17 CEACAM5 CEACAM7 ANXA13 ANXA10 CLDN18 VSIG1 TFF2 VIL1 GDA
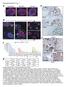 Supplementary Fig. 1 a esophagus Barrett s e FABP1 Ecad stomach TFF3 b p63/krt5 CDH17 REG4 CDH17 f c signal intensity p63/krt5 EsoSC 5000 4000 3000 2000 CDH17 1500 1000 Barrett s KRT14 KRT6A KRT13 KRT5
Supplementary Fig. 1 a esophagus Barrett s e FABP1 Ecad stomach TFF3 b p63/krt5 CDH17 REG4 CDH17 f c signal intensity p63/krt5 EsoSC 5000 4000 3000 2000 CDH17 1500 1000 Barrett s KRT14 KRT6A KRT13 KRT5
Expanded View Figures
 EMO Molecular Medicine Proteomic map of squamous cell carcinomas Hanibal ohnenberger et al Expanded View Figures Figure EV1. Technical reproducibility. Pearson s correlation analysis of normalised SILC
EMO Molecular Medicine Proteomic map of squamous cell carcinomas Hanibal ohnenberger et al Expanded View Figures Figure EV1. Technical reproducibility. Pearson s correlation analysis of normalised SILC
ARTICLE RESEARCH. Macmillan Publishers Limited. All rights reserved
 Extended Data Figure 6 Annotation of drivers based on clinical characteristics and co-occurrence patterns. a, Putative drivers affecting greater than 10 patients were assessed for enrichment in IGHV mutated
Extended Data Figure 6 Annotation of drivers based on clinical characteristics and co-occurrence patterns. a, Putative drivers affecting greater than 10 patients were assessed for enrichment in IGHV mutated
The epigenetic landscape of T cell subsets in SLE identifies known and potential novel drivers of the autoimmune response
 Abstract # 319030 Poster # F.9 The epigenetic landscape of T cell subsets in SLE identifies known and potential novel drivers of the autoimmune response Jozsef Karman, Brian Johnston, Sofija Miljovska,
Abstract # 319030 Poster # F.9 The epigenetic landscape of T cell subsets in SLE identifies known and potential novel drivers of the autoimmune response Jozsef Karman, Brian Johnston, Sofija Miljovska,
Supplementary. properties of. network types. randomly sampled. subsets (75%
 Supplementary Information Gene co-expression network analysis reveals common system-level prognostic genes across cancer types properties of Supplementary Figure 1 The robustness and overlap of prognostic
Supplementary Information Gene co-expression network analysis reveals common system-level prognostic genes across cancer types properties of Supplementary Figure 1 The robustness and overlap of prognostic
Supplementary Table 1. Information on the 174 single nucleotide variants identified by whole-exome sequencing
 Supplementary Table 1. Information on the 174 single nucleotide s identified by whole-exome sequencing Chr Position Type AF exomes Database 1 10163148 A/G UBE4B missense 0.000326 1534 2 98 1 4.44 Damaging
Supplementary Table 1. Information on the 174 single nucleotide s identified by whole-exome sequencing Chr Position Type AF exomes Database 1 10163148 A/G UBE4B missense 0.000326 1534 2 98 1 4.44 Damaging
Nature Immunology: doi: /ni Supplementary Figure 1. Transcriptional program of the TE and MP CD8 + T cell subsets.
 Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Nature Immunology: doi: /ni Supplementary Figure 1. RNA-Seq analysis of CD8 + TILs and N-TILs.
 Supplementary Figure 1 RNA-Seq analysis of CD8 + TILs and N-TILs. (a) Schematic representation of the tumor and cell types used for the study. HNSCC, head and neck squamous cell cancer; NSCLC, non-small
Supplementary Figure 1 RNA-Seq analysis of CD8 + TILs and N-TILs. (a) Schematic representation of the tumor and cell types used for the study. HNSCC, head and neck squamous cell cancer; NSCLC, non-small
Supplemental Figure legends
 Supplemental Figure legends Supplemental Figure S1 Frequently mutated genes. Frequently mutated genes (mutated in at least four patients) with information about mutation frequency, RNA-expression and copy-number.
Supplemental Figure legends Supplemental Figure S1 Frequently mutated genes. Frequently mutated genes (mutated in at least four patients) with information about mutation frequency, RNA-expression and copy-number.
Nature Neuroscience: doi: /nn Supplementary Figure 1. Behavioral training.
 Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Nature Medicine: doi: /nm.3967
 Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
Supplementary Tables. Supplementary Figures
 Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Multiple mutations of lung squamous cell carcinoma shared common mechanisms
 /, Vol. 7, No. 48 Multiple mutations of lung squamous cell carcinoma shared common mechanisms Qianping Li 1,*, Junyi Hou 2,*, Zhaoyan Hu 3, Biao Gu 4, Yan Shi 5 1 Department of Cardiothoracic Surgery,
/, Vol. 7, No. 48 Multiple mutations of lung squamous cell carcinoma shared common mechanisms Qianping Li 1,*, Junyi Hou 2,*, Zhaoyan Hu 3, Biao Gu 4, Yan Shi 5 1 Department of Cardiothoracic Surgery,
(A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC,
 Supplemental Figure Legends Figure S1. Cell line characterization (A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC, green) and co-stained with DAPI to visualize the nuclei.
Supplemental Figure Legends Figure S1. Cell line characterization (A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC, green) and co-stained with DAPI to visualize the nuclei.
Variant prioritization
 Variant prioritization University of Cambridge Marta Bleda Latorre Cambridge, UK mb2033@cam.ac.uk 30th September 2014 Research Assistant at the Department of Medicine University of Cambridge Cambridge,
Variant prioritization University of Cambridge Marta Bleda Latorre Cambridge, UK mb2033@cam.ac.uk 30th September 2014 Research Assistant at the Department of Medicine University of Cambridge Cambridge,
Supplementary Materials for
 www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
Nature Genetics: doi: /ng Supplementary Figure 1. PCA for ancestry in SNV data.
 Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
BCR ABL1 like ALL: molekuliniai mechanizmai ir klinikinė reikšmė. IKAROS delecija: molekulinė biologija, prognostinė reikšmė. ASH 2015 naujienos
 BCR ABL1 like ALL: molekuliniai mechanizmai ir klinikinė reikšmė. IKAROS delecija: molekulinė biologija, prognostinė reikšmė. ASH 2015 naujienos Ph like ALL BCR ABL1 like acute lymphoblastic leukemia (ALL)
BCR ABL1 like ALL: molekuliniai mechanizmai ir klinikinė reikšmė. IKAROS delecija: molekulinė biologija, prognostinė reikšmė. ASH 2015 naujienos Ph like ALL BCR ABL1 like acute lymphoblastic leukemia (ALL)
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
HepaRG LX2. HepaRG HepaRG LX2 LX2
 C Supporting Figure 1. Experimental design of s between and cells. (A) -hepatocytes were isolated from a 30 days of -progenitors. Differentiation into mature hepatocytes was achieved following a 2-weeks
C Supporting Figure 1. Experimental design of s between and cells. (A) -hepatocytes were isolated from a 30 days of -progenitors. Differentiation into mature hepatocytes was achieved following a 2-weeks
DYSREGULATION OF WT1 (-KTS) IS ASSOCIATED WITH THE KIDNEY-SPECIFIC EFFECTS OF THE LMX1B R246Q MUTATION
 DYSREGULATION OF WT1 (-KTS) IS ASSOCIATED WITH THE KIDNEY-SPECIFIC EFFECTS OF THE LMX1B R246Q MUTATION Gentzon Hall 1,2#, Brandon Lane 1,3#, Megan Chryst-Ladd 1,3, Guanghong Wu 1,3, Jen-Jar Lin 4, XueJun
DYSREGULATION OF WT1 (-KTS) IS ASSOCIATED WITH THE KIDNEY-SPECIFIC EFFECTS OF THE LMX1B R246Q MUTATION Gentzon Hall 1,2#, Brandon Lane 1,3#, Megan Chryst-Ladd 1,3, Guanghong Wu 1,3, Jen-Jar Lin 4, XueJun
Supplementary Figure 1. Estimation of tumour content
 Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded
 Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Paternal exposure and effects on microrna and mrna expression in developing embryo. Department of Chemical and Radiation Nur Duale
 Paternal exposure and effects on microrna and mrna expression in developing embryo Department of Chemical and Radiation Nur Duale Our research question Can paternal preconceptional exposure to environmental
Paternal exposure and effects on microrna and mrna expression in developing embryo Department of Chemical and Radiation Nur Duale Our research question Can paternal preconceptional exposure to environmental
SUPPLEMENTARY INFORMATION
 Supplementary text Collectively, we were able to detect ~14,000 expressed genes with RPKM (reads per kilobase per million) > 1 or ~16,000 with RPKM > 0.1 in at least one cell type from oocyte to the morula
Supplementary text Collectively, we were able to detect ~14,000 expressed genes with RPKM (reads per kilobase per million) > 1 or ~16,000 with RPKM > 0.1 in at least one cell type from oocyte to the morula
RNA SEQUENCING AND DATA ANALYSIS
 RNA SEQUENCING AND DATA ANALYSIS Download slides and package http://odin.mdacc.tmc.edu/~rverhaak/package.zip http://odin.mdacc.tmc.edu/~rverhaak/rna-seqlecture.zip Overview Introduction into the topic
RNA SEQUENCING AND DATA ANALYSIS Download slides and package http://odin.mdacc.tmc.edu/~rverhaak/package.zip http://odin.mdacc.tmc.edu/~rverhaak/rna-seqlecture.zip Overview Introduction into the topic
Expanded View Figures
 Solip Park & Ben Lehner Epistasis is cancer type specific Molecular Systems Biology Expanded View Figures A B G C D E F H Figure EV1. Epistatic interactions detected in a pan-cancer analysis and saturation
Solip Park & Ben Lehner Epistasis is cancer type specific Molecular Systems Biology Expanded View Figures A B G C D E F H Figure EV1. Epistatic interactions detected in a pan-cancer analysis and saturation
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature15260 Supplementary Data 1: Gene expression in individual basal/stem, luminal, and luminal progenitor cells. Box plots show expression levels for each gene from the 49-gene differentiation
doi:10.1038/nature15260 Supplementary Data 1: Gene expression in individual basal/stem, luminal, and luminal progenitor cells. Box plots show expression levels for each gene from the 49-gene differentiation
SUPPLEMENTARY INFORMATION
 Somatic ERCC2 Mutations Are Associated with a Distinct Genomic Signature in Urothelial Tumors Jaegil Kim, Kent W Mouw, Paz Polak, Lior Z Braunstein, Atanas Kamburov, Grace Tiao, David J Kwiatkowski, Jonathan
Somatic ERCC2 Mutations Are Associated with a Distinct Genomic Signature in Urothelial Tumors Jaegil Kim, Kent W Mouw, Paz Polak, Lior Z Braunstein, Atanas Kamburov, Grace Tiao, David J Kwiatkowski, Jonathan
POT1 mutations cause telomere dysfunction in chronic lymphocytic leukemia
 POT1 mutations cause telomere dysfunction in chronic lymphocytic leukemia Andrew J. Ramsay, Víctor Quesada, Miguel Foronda, Laura Conde, Alejandra Martínez-Trillos, Neus Villamor, David Rodríguez, Agnieszka
POT1 mutations cause telomere dysfunction in chronic lymphocytic leukemia Andrew J. Ramsay, Víctor Quesada, Miguel Foronda, Laura Conde, Alejandra Martínez-Trillos, Neus Villamor, David Rodríguez, Agnieszka
SUPPLEMENTARY FIGURES: Supplementary Figure 1
 SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
Nature Getetics: doi: /ng.3471
 Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Journal: Nature Methods
 Journal: Nature Methods Article Title: Network-based stratification of tumor mutations Corresponding Author: Trey Ideker Supplementary Item Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure
Journal: Nature Methods Article Title: Network-based stratification of tumor mutations Corresponding Author: Trey Ideker Supplementary Item Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure
Supplementary Figure 1
 Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure 1: Classification scheme for non-synonymous and nonsense germline MC1R variants. The common variants with previously established
 Supplementary Figure 1: Classification scheme for nonsynonymous and nonsense germline MC1R variants. The common variants with previously established classifications 1 3 are shown. The effect of novel missense
Supplementary Figure 1: Classification scheme for nonsynonymous and nonsense germline MC1R variants. The common variants with previously established classifications 1 3 are shown. The effect of novel missense
SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer.
 Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Supplemental Table 1. List of genes and mature shrna sequences identified from the shrna screen in BRCA2-mutant PEO1 cells.
 Guillemette_Supplemental Table 1 Gene Mature Sequence Gene Mature Sequence ATTCATAGGATGTCAGCAG LOC157193 TATGCGTAAGTAATAAATTGCT TTAGTTCTGTCTTGGAGG LOC152225 CGCATATATGTTCTGACTT ALDH2 CACTTCAGTGTATGCCT
Guillemette_Supplemental Table 1 Gene Mature Sequence Gene Mature Sequence ATTCATAGGATGTCAGCAG LOC157193 TATGCGTAAGTAATAAATTGCT TTAGTTCTGTCTTGGAGG LOC152225 CGCATATATGTTCTGACTT ALDH2 CACTTCAGTGTATGCCT
Expanded View Figures
 Molecular Systems iology Tumor CNs reflect metabolic selection Nicholas Graham et al Expanded View Figures Human primary tumors CN CN characterization by unsupervised PC Human Signature Human Signature
Molecular Systems iology Tumor CNs reflect metabolic selection Nicholas Graham et al Expanded View Figures Human primary tumors CN CN characterization by unsupervised PC Human Signature Human Signature
BWA alignment to reference transcriptome and genome. Convert transcriptome mappings back to genome space
 Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Supplement Results, Figures and Tables
 Supplement Results, Figures and Tables Results PET imaging The SUV was significantly higher in tumours of the parental HSC-3 cell line than tumours of the EV and the MMP-8 overexpressing cell lines (Figure
Supplement Results, Figures and Tables Results PET imaging The SUV was significantly higher in tumours of the parental HSC-3 cell line than tumours of the EV and the MMP-8 overexpressing cell lines (Figure
Research Strategy: 1. Background and Significance
 Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
A pediatric patient with acute leukemia of ambiguous lineage with a NUP98-NSD1 rearrangement SH
 A pediatric patient with acute leukemia of ambiguous lineage with a NUP98NSD1 rearrangement SH20170203 Rebecca LeemanNeill, Ronald Rice, Anita Malek, Patricia Raciti, Susan Hsiao, Mahesh Mansukhani, Bachir
A pediatric patient with acute leukemia of ambiguous lineage with a NUP98NSD1 rearrangement SH20170203 Rebecca LeemanNeill, Ronald Rice, Anita Malek, Patricia Raciti, Susan Hsiao, Mahesh Mansukhani, Bachir
Supplementary Methods
 Supplementary Methods Short Read Preprocessing Reads are preprocessed differently according to how they will be used: detection of the variant in the tumor, discovery of an artifact in the normal or for
Supplementary Methods Short Read Preprocessing Reads are preprocessed differently according to how they will be used: detection of the variant in the tumor, discovery of an artifact in the normal or for
RNA SEQUENCING AND DATA ANALYSIS
 RNA SEQUENCING AND DATA ANALYSIS Length of mrna transcripts in the human genome 5,000 5,000 4,000 3,000 2,000 4,000 1,000 0 0 200 400 600 800 3,000 2,000 1,000 0 0 2,000 4,000 6,000 8,000 10,000 Length
RNA SEQUENCING AND DATA ANALYSIS Length of mrna transcripts in the human genome 5,000 5,000 4,000 3,000 2,000 4,000 1,000 0 0 200 400 600 800 3,000 2,000 1,000 0 0 2,000 4,000 6,000 8,000 10,000 Length
Nature Genetics: doi: /ng Supplementary Figure 1. Rates of different mutation types in CRC.
 Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
Supplementary Information to
 Supplementary Information to Exome sequencing to identify de novo mutations in sporadic ALS trios Alessandra Chesi 1,11, Brett T. Staahl 2, Ana Jovicic 1, Julien Couthouis 1, Maria Fasolino 3, Alya R.
Supplementary Information to Exome sequencing to identify de novo mutations in sporadic ALS trios Alessandra Chesi 1,11, Brett T. Staahl 2, Ana Jovicic 1, Julien Couthouis 1, Maria Fasolino 3, Alya R.
Supplementary Figure 1 ITGB1 and ITGA11 increase with evidence for heterodimers following HSC activation. (a) Time course of rat HSC activation
 Supplementary Figure 1 ITGB1 and ITGA11 increase with evidence for heterodimers following HSC activation. (a) Time course of rat HSC activation indicated by the detection of -SMA and COL1 (log scale).
Supplementary Figure 1 ITGB1 and ITGA11 increase with evidence for heterodimers following HSC activation. (a) Time course of rat HSC activation indicated by the detection of -SMA and COL1 (log scale).
Supplementary Online Content
 Supplementary Online Content Lyall DM, Celis-Morales C, Ward J, et al. Association of body mass index with cardiometabolic disease in the UK Biobank: a mendelian randomization study. JAMA Cardiol. Published
Supplementary Online Content Lyall DM, Celis-Morales C, Ward J, et al. Association of body mass index with cardiometabolic disease in the UK Biobank: a mendelian randomization study. JAMA Cardiol. Published
Nature Genetics: doi: /ng Supplementary Figure 1. HOX fusions enhance self-renewal capacity.
 Supplementary Figure 1 HOX fusions enhance self-renewal capacity. Mouse bone marrow was transduced with a retrovirus carrying one of three HOX fusion genes or the empty mcherry reporter construct as described
Supplementary Figure 1 HOX fusions enhance self-renewal capacity. Mouse bone marrow was transduced with a retrovirus carrying one of three HOX fusion genes or the empty mcherry reporter construct as described
Supplementary information
 Supplementary information High fat diet-induced changes of mouse hepatic transcription and enhancer activity can be reversed by subsequent weight loss Majken Siersbæk, Lyuba Varticovski, Shutong Yang,
Supplementary information High fat diet-induced changes of mouse hepatic transcription and enhancer activity can be reversed by subsequent weight loss Majken Siersbæk, Lyuba Varticovski, Shutong Yang,
Supplementary Figure 1: TSLP receptor skin expression in dcssc. A: Healthy control (HC) skin with TSLP receptor expression in brown (10x
 Supplementary Figure 1: TSLP receptor skin expression in dcssc. A: Healthy control (HC) skin with TSLP receptor expression in brown (10x magnification). B: Second HC skin stained for TSLP receptor in brown
Supplementary Figure 1: TSLP receptor skin expression in dcssc. A: Healthy control (HC) skin with TSLP receptor expression in brown (10x magnification). B: Second HC skin stained for TSLP receptor in brown
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
Phenotype Report. Num. Positions Not Called (Missing data) Num. Variants Assessed
 Report Date: August 19, 2015 Software Annotation Version: 8 Report Name: NA12144 NW European Genome : NA12144_S1 Sequencing Provider: Illumina Sequencing Type: Exome : Retinitis Pigmentosa Description:
Report Date: August 19, 2015 Software Annotation Version: 8 Report Name: NA12144 NW European Genome : NA12144_S1 Sequencing Provider: Illumina Sequencing Type: Exome : Retinitis Pigmentosa Description:
The pluripotency factor NANOG promotes the formation of squamous cell carcinomas
 SUPPLEMENTAL DATA The pluripotency factor NANOG promotes the formation of squamous cell carcinomas Adelaida R. Palla, Daniela Piazzolla, Noelia Alcazar, Marta Cañamero, Osvaldo Graña, Gonzalo Gómez-López,
SUPPLEMENTAL DATA The pluripotency factor NANOG promotes the formation of squamous cell carcinomas Adelaida R. Palla, Daniela Piazzolla, Noelia Alcazar, Marta Cañamero, Osvaldo Graña, Gonzalo Gómez-López,
Supplementary Note. (Quesada et al., Exome-Sequencing Identifies Recurrent Mutations of the Splicing Factor SF3B1 in Chronic Lymphocytic Leukemia)
 Supplementary Note (Quesada et al., Exome-Sequencing Identifies Recurrent Mutations of the Splicing Factor SF3B1 in Chronic Lymphocytic Leukemia) The clinical and biological characteristics of the 105
Supplementary Note (Quesada et al., Exome-Sequencing Identifies Recurrent Mutations of the Splicing Factor SF3B1 in Chronic Lymphocytic Leukemia) The clinical and biological characteristics of the 105
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
Nature Genetics: doi: /ng Supplementary Figure 1. Clinical timeline for the discovery WES cases.
 Supplementary Figure 1 Clinical timeline for the discovery WES cases. This illustrates the timeline of the disease events during the clinical course of each patient s disease, further indicating the available
Supplementary Figure 1 Clinical timeline for the discovery WES cases. This illustrates the timeline of the disease events during the clinical course of each patient s disease, further indicating the available
Supplemental Tables and Figures
 Supplemental Tables and Figures Post-zygotic de novo changes in glutamate and dopamine pathways may explain discordance of monozygotic twins for schizophrenia Castellani, CA., Melka, MG., Gui, JL., Gallo,
Supplemental Tables and Figures Post-zygotic de novo changes in glutamate and dopamine pathways may explain discordance of monozygotic twins for schizophrenia Castellani, CA., Melka, MG., Gui, JL., Gallo,
Identifying Thyroid Carcinoma Subtypes and Outcomes through Gene Expression Data Kun-Hsing Yu, Wei Wang, Chung-Yu Wang
 Identifying Thyroid Carcinoma Subtypes and Outcomes through Gene Expression Data Kun-Hsing Yu, Wei Wang, Chung-Yu Wang Abstract: Unlike most cancers, thyroid cancer has an everincreasing incidence rate
Identifying Thyroid Carcinoma Subtypes and Outcomes through Gene Expression Data Kun-Hsing Yu, Wei Wang, Chung-Yu Wang Abstract: Unlike most cancers, thyroid cancer has an everincreasing incidence rate
Integrated network analysis reveals distinct regulatory roles of transcription factors and micrornas
 Integrated network analysis reveals distinct regulatory roles of transcription factors and micrornas Yu Guo 1,2,4, Katherine Alexander 1, Andrew G Clark 1, Andrew Grimson 1 and Haiyuan Yu 2,3* 1 Department
Integrated network analysis reveals distinct regulatory roles of transcription factors and micrornas Yu Guo 1,2,4, Katherine Alexander 1, Andrew G Clark 1, Andrew Grimson 1 and Haiyuan Yu 2,3* 1 Department
Supplementary Figure 1
 Supplementary Figure 1 5 microns C7 B6 unclassified H19 C7 signal H19 guide signal H19 B6 signal C7 SNP spots H19 RNA spots B6 SNP spots colocalization H19 RNA classification Supplementary Figure 1. Allele-specific
Supplementary Figure 1 5 microns C7 B6 unclassified H19 C7 signal H19 guide signal H19 B6 signal C7 SNP spots H19 RNA spots B6 SNP spots colocalization H19 RNA classification Supplementary Figure 1. Allele-specific
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplemental Information. Genetic Regulation of Plasma Lipid Species. and Their Association with Metabolic Phenotypes
 Cell Systems, Volume 6 Supplemental Information Genetic Regulation of Plasma Lipid Species and Their Association with Metabolic Phenotypes Pooja Jha, Molly T. McDevitt, Emina Halilbasic, Evan G. Williams,
Cell Systems, Volume 6 Supplemental Information Genetic Regulation of Plasma Lipid Species and Their Association with Metabolic Phenotypes Pooja Jha, Molly T. McDevitt, Emina Halilbasic, Evan G. Williams,
Expert-guided Visual Exploration (EVE) for patient stratification. Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland
 Expert-guided Visual Exploration (EVE) for patient stratification Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland Oncoscape.sttrcancer.org Paul Lisa Ken Jenny Desert Eric The challenge Given - patient clinical
Expert-guided Visual Exploration (EVE) for patient stratification Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland Oncoscape.sttrcancer.org Paul Lisa Ken Jenny Desert Eric The challenge Given - patient clinical
Single-cell RNA-Seq profiling of human pre-implantation embryos and embryonic stem cells
 Single-cell RNA-Seq profiling of human pre-implantation embryos and embryonic stem cells Liying Yan,2,5, Mingyu Yang,5, Hongshan Guo, Lu Yang, Jun Wu, Rong Li,2, Ping Liu, Ying Lian, Xiaoying Zheng, Jie
Single-cell RNA-Seq profiling of human pre-implantation embryos and embryonic stem cells Liying Yan,2,5, Mingyu Yang,5, Hongshan Guo, Lu Yang, Jun Wu, Rong Li,2, Ping Liu, Ying Lian, Xiaoying Zheng, Jie
Myeloma Genetics what do we know and where are we going?
 in partnership with Myeloma Genetics what do we know and where are we going? Dr Brian Walker Thames Valley Cancer Network 14 th September 2015 Making the discoveries that defeat cancer Myeloma Genome:
in partnership with Myeloma Genetics what do we know and where are we going? Dr Brian Walker Thames Valley Cancer Network 14 th September 2015 Making the discoveries that defeat cancer Myeloma Genome:
Nature Genetics: doi: /ng Supplementary Figure 1. Mutational signatures in BCC compared to melanoma.
 Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
fl/+ KRas;Atg5 fl/+ KRas;Atg5 fl/fl KRas;Atg5 fl/fl KRas;Atg5 Supplementary Figure 1. Gene set enrichment analyses. (a) (b)
 KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
SOPten flox/flox (KO) Pten flox/flox (WT) flox allele 6.0 kb. Pten. Actin. ! allele 2.3 kb. Supplementary Figure S1. Yanagi, et al.
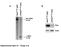 s1 A Pten flox/flox () SOPten flox/flox () flox allele 6. kb B Pten flox/flox () SOPten flox/flox () Pten Actin! allele 2.3 kb Supplementary Figure S1. Yanagi, et al. A B BrdU BrdU positive cells ( ) 3
s1 A Pten flox/flox () SOPten flox/flox () flox allele 6. kb B Pten flox/flox () SOPten flox/flox () Pten Actin! allele 2.3 kb Supplementary Figure S1. Yanagi, et al. A B BrdU BrdU positive cells ( ) 3
SUPPLEMENTARY INFORMATION. Intron retention is a widespread mechanism of tumor suppressor inactivation.
 SUPPLEMENTARY INFORMATION Intron retention is a widespread mechanism of tumor suppressor inactivation. Hyunchul Jung 1,2,3, Donghoon Lee 1,4, Jongkeun Lee 1,5, Donghyun Park 2,6, Yeon Jeong Kim 2,6, Woong-Yang
SUPPLEMENTARY INFORMATION Intron retention is a widespread mechanism of tumor suppressor inactivation. Hyunchul Jung 1,2,3, Donghoon Lee 1,4, Jongkeun Lee 1,5, Donghyun Park 2,6, Yeon Jeong Kim 2,6, Woong-Yang
The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis
 The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis Tieliu Shi tlshi@bio.ecnu.edu.cn The Center for bioinformatics
The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis Tieliu Shi tlshi@bio.ecnu.edu.cn The Center for bioinformatics
Integrative modeling of transcriptional regulatory networks in head and neck cancer
 Integrative modeling of transcriptional regulatory networks in head and neck cancer Bin Yan 1*, Huai Li, Zhong Chen 3, Jiaofang Shao 1, Ming Zhan 4* 1 Department of Biology, Hong Kong Baptist University,
Integrative modeling of transcriptional regulatory networks in head and neck cancer Bin Yan 1*, Huai Li, Zhong Chen 3, Jiaofang Shao 1, Ming Zhan 4* 1 Department of Biology, Hong Kong Baptist University,
Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes.
 Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Figure 1 Gene expression profiles define 3 molecular sub-groups of CNS-PNET
 Supplemental Figure Legends Figure 1 Gene expression profiles define 3 molecular sub-groups of CNS-PNET Multiple unsupervised analyses were performed on human HT-1v4 expression array (Illumina) data from
Supplemental Figure Legends Figure 1 Gene expression profiles define 3 molecular sub-groups of CNS-PNET Multiple unsupervised analyses were performed on human HT-1v4 expression array (Illumina) data from
Chapter 1. Introduction
 Chapter 1 Introduction 1.1 Motivation and Goals The increasing availability and decreasing cost of high-throughput (HT) technologies coupled with the availability of computational tools and data form a
Chapter 1 Introduction 1.1 Motivation and Goals The increasing availability and decreasing cost of high-throughput (HT) technologies coupled with the availability of computational tools and data form a
Dominic J Smiraglia, PhD Department of Cancer Genetics. DNA methylation in prostate cancer
 Dominic J Smiraglia, PhD Department of Cancer Genetics DNA methylation in prostate cancer Overarching theme Epigenetic regulation allows the genome to be responsive to the environment Sets the tone for
Dominic J Smiraglia, PhD Department of Cancer Genetics DNA methylation in prostate cancer Overarching theme Epigenetic regulation allows the genome to be responsive to the environment Sets the tone for
Variant association and prioritization
 Variant association and prioritization Edinburgh Genomics Marta Bleda Latorre Edinburgh, UK mb2033@cam.ac.uk 23rd October 2015 Research Assistant at the Department of Medicine University of Cambridge Cambridge,
Variant association and prioritization Edinburgh Genomics Marta Bleda Latorre Edinburgh, UK mb2033@cam.ac.uk 23rd October 2015 Research Assistant at the Department of Medicine University of Cambridge Cambridge,
Supplementary appendix
 Supplementary appendix This appendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Ellingford JM, Sergouniotis PI, Lennon R, et
Supplementary appendix This appendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Ellingford JM, Sergouniotis PI, Lennon R, et
IPA Advanced Training Course
 IPA Advanced Training Course October 2013 Academia sinica Gene (Kuan Wen Chen) IPA Certified Analyst Agenda I. Data Upload and How to Run a Core Analysis II. Functional Interpretation in IPA Hands-on Exercises
IPA Advanced Training Course October 2013 Academia sinica Gene (Kuan Wen Chen) IPA Certified Analyst Agenda I. Data Upload and How to Run a Core Analysis II. Functional Interpretation in IPA Hands-on Exercises
HALLA KABAT * Outreach Program, mircore, 2929 Plymouth Rd. Ann Arbor, MI 48105, USA LEO TUNKLE *
 CERNA SEARCH METHOD IDENTIFIED A MET-ACTIVATED SUBGROUP AMONG EGFR DNA AMPLIFIED LUNG ADENOCARCINOMA PATIENTS HALLA KABAT * Outreach Program, mircore, 2929 Plymouth Rd. Ann Arbor, MI 48105, USA Email:
CERNA SEARCH METHOD IDENTIFIED A MET-ACTIVATED SUBGROUP AMONG EGFR DNA AMPLIFIED LUNG ADENOCARCINOMA PATIENTS HALLA KABAT * Outreach Program, mircore, 2929 Plymouth Rd. Ann Arbor, MI 48105, USA Email:
underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
 Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Supplementary Information. Preferential associations between co-regulated genes reveal a. transcriptional interactome in erythroid cells
 Supplementary Information Preferential associations between co-regulated genes reveal a transcriptional interactome in erythroid cells Stefan Schoenfelder, * Tom Sexton, * Lyubomira Chakalova, * Nathan
Supplementary Information Preferential associations between co-regulated genes reveal a transcriptional interactome in erythroid cells Stefan Schoenfelder, * Tom Sexton, * Lyubomira Chakalova, * Nathan
Predicting Kidney Cancer Survival from Genomic Data
 Predicting Kidney Cancer Survival from Genomic Data Christopher Sauer, Rishi Bedi, Duc Nguyen, Benedikt Bünz Abstract Cancers are on par with heart disease as the leading cause for mortality in the United
Predicting Kidney Cancer Survival from Genomic Data Christopher Sauer, Rishi Bedi, Duc Nguyen, Benedikt Bünz Abstract Cancers are on par with heart disease as the leading cause for mortality in the United
Supplemental Figure S1. RANK expression on human lung cancer cells.
 Supplemental Figure S1. RANK expression on human lung cancer cells. (A) Incidence and H-Scores of RANK expression determined from IHC in the indicated primary lung cancer subgroups. The overall expression
Supplemental Figure S1. RANK expression on human lung cancer cells. (A) Incidence and H-Scores of RANK expression determined from IHC in the indicated primary lung cancer subgroups. The overall expression
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Metabolic ER stress and inflammation in white adipose tissue (WAT) of mice with dietary obesity.
 Supplementary Figure 1 Metabolic ER stress and inflammation in white adipose tissue (WAT) of mice with dietary obesity. Male C57BL/6J mice were fed a normal chow (NC, 10% fat) or a high-fat diet (HFD,
Supplementary Figure 1 Metabolic ER stress and inflammation in white adipose tissue (WAT) of mice with dietary obesity. Male C57BL/6J mice were fed a normal chow (NC, 10% fat) or a high-fat diet (HFD,
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Multiple samples from five patients (P4, P8, P14, P15 and P17) with Barrett s esophagus and adjacent EAC show that the poor overlap is not a result of sampling bias. Bar graphs showing
Supplementary Figure 1 Multiple samples from five patients (P4, P8, P14, P15 and P17) with Barrett s esophagus and adjacent EAC show that the poor overlap is not a result of sampling bias. Bar graphs showing
