Supplementary figures
|
|
|
- Ralph Woods
- 6 years ago
- Views:
Transcription
1 Supplementary figures Supplementary figure 1: Pathway enrichment for each comparison. The first column shows enrichment with differentially expressed genes between the diet groups at t = 0, the second and thirds columns show enrichment with differentially expressed genes for t = 0 compared to the other time points in LF diet and HF diet respectively. Only pathways with an enrichment p-value < 0.01 in at least one of the tests are shown. The pathways are sorted by hierarchical clustering over all columns to group pathways with a similar response. Saturation of each box indicates enrichment significance, the more saturated blue, the more significant. In addition, the boxes are starred by significance threshold (* for p < 0.05, ** for p < 0.01). 1
2 Supplementary figure 2: In and out degree (number of interactions) for the most connected pathways per network. Only pathways with a degree >=4 in at least one of the networks are included. 2
3 Supplementary figure 3: Normalized node betweenness centrality per pathway, which measures how often a node occurs on shortest paths between the other nodes in the network. The value drawn in each cell is the normalized by dividing by the maximum betweenness centrality (values multiplied by 10 4 ). 3
4 Supplementary figure 4: Total number of protein interaction paths between pathways for each pathway interaction network. Bars are colored by the length (number of interactions) of each path. 4
5 Supplementary figure 5: Interactions with Sgk3 that are part of identified indirect paths between pathways in the network for the response in LF at t=2 and t=48. 5
6 Supplementary figure 6: The protein interactions that compose several pathway interactions in the subgraph as shown in Figure 3A (see insets, included interactions are marked orange). Protein nodes have a thick border when they are significantly differentially expressed (q < 0.05). A: interactions between the ESC Pluripotency pathway and its neighbors, B: interaction between the Proteasome pathway and ESC Pluripotency pathway, C: interactions between the TNF-alpha NF-kB Signaling pathway and the two versions of the Wnt Signaling pathway. 6
7 Supplementary figure 7: The diagram for the ESC Pluripotency pathways. Highlighted in blue are the proteins Apc, Ctnnb1 and Gsk3b which are identified in the interaction with the Proteasome pathway in the network for the comparison between diets at t=0. Supplementary figure 8: Protein interactions contributing to the pathways between the Primary Bile Acid Biosynthesis and Peroxisome pathways and several other metabolic pathways in the network for comparison between diets at t = 0. The three proteins are all significantly up-regulated in the HF group and their interactions are binding. 7
8 Supplementary Figure 9: Protein interactions playing a role in the interactions between pathways for the networks based on differential expression between t=0 and t=0.6 in the LF diet group (left) and the HF diet group (right). Supplementary figure 10: Protein interactions playing a role in the interactions between pathways in the component composed by the Antigen processing and presentation, the Spliceosome and Lysosome pathways for the network based on differential expression between t=0 and t=0.6 in the HF diet group. 8
9 Supplementary figure 11: Protein interactions playing a role in the interactions between pathways in the component composed by the S1P Receptor, Nuclear Receptors, Cytoplasmic Ribosomal Proteins and Insulin Signaling pathways for the network based on differential expression between t=0 and t=0.6 in HF diet. 9
10 Supplementary figure 12: Protein interactions for the paths between the Endocytosis pathway and its neighbors in the network for differential expression between t=2 and t=0 in the HF diet group. 10
11 Supplementary figure 13: Proteins forming the paths between the Proteasome and TGF-beta Receptor Signaling pathways for the networks based on differential expression between t=48 and t=0 in the LF group (left) and HF group (right). Supplementary figure 14: Paths between the Antigen processing and presentation, T Cell Receptor Signaling, Cell Cycle and TNF alpha NF-kB pathways in the network for differential expression between t=48 and t=0 in the LF diet. 11
12 Supplementary figure 15: Paths between the pathways in the second order neighborhood of the Insulin Signaling pathway in the network for differential expression between t=48 and t=0 in the LF diet. Supplementary figure 16: Interaction between the Insulin and Cytokine-cytokine receptor interaction pathways for the network based on differential expression between t=0 and t=2 in the HF group. 12
13 Supplementary figure 17: Transformation of T statistic to edge weights using the sigmoid function, with parameters µ=3 and α=2. 13
14 Supplementary tables HF vs LF, t = 0 Nr. paths Kras 109 Raf1 96 Nras 85 Ctnnb1 64 Sos1 62 LF, t = 0.6 vs t = 0 Jun 25 Il1b 20 Fos 17 Il1a 11 Atf3 5 LF, t = 2 vs t = 0 Mapk1 21 Prkacb 16 Involved in interactions with pathways Dorso-ventral axis formation (15), FAS pathway and Stress induction of HSP regulation (14), ESC Pluripotency Pathways (13), 47 more. ESC Pluripotency Pathways (13), FAS pathway and Stress induction of HSP regulation (11), Oxidative Damage; Apoptosis (11), 45 more. FAS pathway and Stress induction of HSP regulation (11), Oxidative Damage; Apoptosis (11), Long-term depression (10), 34 more. Ptf1a related regulatory pathway (12), Dorso-ventral axis formation (10), Proteasome; Proteasome Degradation (7), 43 more. ESC Pluripotency Pathways (13), Dorso-ventral axis formation (10), FAS pathway and Stress induction of HSP regulation (7), 40 more. FAS pathway and Stress induction of HSP regulation (11), Selenium metabolism/selenoproteins (11), Oxidative Stress (3), 14 more. FAS pathway and Stress induction of HSP regulation (10), Selenium metabolism/selenoproteins (9), Tolllike receptor signaling pathway; NLR proteins (3), 10 more. Selenium metabolism/selenoproteins (10), FAS pathway and Stress induction of HSP regulation (4), Oxidative Stress (3), 13 more. FAS pathway and Stress induction of HSP regulation (10), Toll-like receptor signaling pathway; NLR proteins (2), 10 more. Selenium metabolism/selenoproteins (3), Hypertrophy Model (2), Myometrial Relaxation and Contraction Pathways (2), 3 more. Signal Transduction of S1P Receptor (6), Cytoplasmic Ribosomal Proteins; Ribosome (5), Vasopressinregulated water reabsorption (5), 18 more. Taste transduction (5), Vasopressin-regulated water reabsorption (5), Hypothetical Network for Drug Addiction (3), 13 more. Rps6ka1 10 Signal Transduction of S1P Receptor (6), Cytoplasmic Ribosomal Proteins; Ribosome (5), 9 more. Spliceosome (5), SNARE interactions in vesicular transport (3), Antigen processing and presentation (2), 5 Hspa1b 8 more. Mapk6 6 Signal Transduction of S1P Receptor (6), 6 more. LF, t = 48 vs t = 0 Prkaca 48 Cep57 44 Nedd1 44 Mapre3 44 Tubg1 44 HF, t = 0.6 vs t = 0 Jun 34 Fos 28 Il1b 23 Il1a 17 Atf3 15 HF, t = 2 vs t = 0 Endochondral Ossification (9), Antigen processing and presentation (9), T Cell Receptor Signaling Pathway (8), 25 more. Antigen processing and presentation (11), T Cell Receptor Signaling Pathway (9), Endochondral Ossification (8), 16 more. Antigen processing and presentation (11), T Cell Receptor Signaling Pathway (9), Endochondral Ossification (8), 16 more. Antigen processing and presentation (11), T Cell Receptor Signaling Pathway (9), Endochondral Ossification (8), 16 more. Antigen processing and presentation (11), T Cell Receptor Signaling Pathway (9), Endochondral Ossification (8), 16 more. Hypertrophy Model (13), Selenium metabolism/selenoproteins (10), FAS pathway and Stress induction of HSP regulation (8), 19 more. Hypertrophy Model (15), Selenium metabolism/selenoproteins (9), T cell receptor signaling pathway (4), 18 more. FAS pathway and Stress induction of HSP regulation (8), Selenium metabolism/selenoproteins (6), Hypertrophy Model (4), 11 more. FAS pathway and Stress induction of HSP regulation (9), Hypertrophy Model (4), Toll Like Receptor signaling (4), 9 more. Hypertrophy Model (13), Selenium metabolism/selenoproteins (3), Myometrial Relaxation and Contraction Pathways (2), 11 more. 14
15 Pik3r1 60 Pik3r2 51 Egfr 45 Akt1 41 Pik3cg 33 HF, t = 48 vs t = 0 Cytokine-cytokine receptor interaction; Cytokines and Inflammatory Response (BioCarta) (14), Dorsoventral axis formation (12), Aldosterone-regulated sodium reabsorption (7), 35 more. Dorso-ventral axis formation (13), Cytokine-cytokine receptor interaction; Cytokines and Inflammatory Response (BioCarta) (12), Aldosterone-regulated sodium reabsorption (7), 32 more. Cytokine-cytokine receptor interaction; Cytokines and Inflammatory Response (BioCarta) (15), Dorsoventral axis formation (15), Endocytosis (7), 33 more. Endochondral Ossification (6), EPO Receptor Signaling (6), Signal Transduction of S1P Receptor (6), 31 more. Dorso-ventral axis formation (12), Cytokine-cytokine receptor interaction; Cytokines and Inflammatory Response (BioCarta) (11), Endocytosis (6), 22 more. Pik3r1 39 Axon guidance (8), Tight junction (8), IL-4 signaling Pathway (5), 33 more. Kras 37 Axon guidance (8), Tight junction (8), IL-4 signaling Pathway (5), 33 more. Ctnnb1 35 Proteasome; Proteasome Degradation (10), Tight junction (8), Ptf1a related regulatory pathway (6), 25 more. Prkca 22 Keap1-Nrf2 (5), Axon guidance (4), Phosphatidylinositol signaling system; Inositol phosphate metabolism (4), 20 more. Mapk14 18 Axon guidance (6), IL-4 signaling Pathway (5), Dorso-ventral axis formation (4), 15 more. Supplementary Table 1: Top 5 proteins involved in most significant pathway interactions for each comparison. The last column displays the pathways in which interactions the protein is most involved, with the number of interactions in parentheses. 15
Supplementary data Table S3. GO terms, pathways and networks enriched among the significantly correlating genes using Tox-Profiler
 Supplementary data Table S3. GO terms, pathways and networks enriched among the significantly correlating genes using Tox-Profiler DR CALUX Boys Girls Database Systemic lupus erythematosus 4.4 0.0021 6.7
Supplementary data Table S3. GO terms, pathways and networks enriched among the significantly correlating genes using Tox-Profiler DR CALUX Boys Girls Database Systemic lupus erythematosus 4.4 0.0021 6.7
Pathways of changes in hypermethylation in F1 offspring exposed to intrauterine hyperglycemia
 Supplementary Table 1. Pathways of changes in hypermethylation in F1 offspring exposed to intrauterine hyperglycemia Pathway Enrichment Score (-log10[p value]) p-value genes Glycosaminoglycan degradation
Supplementary Table 1. Pathways of changes in hypermethylation in F1 offspring exposed to intrauterine hyperglycemia Pathway Enrichment Score (-log10[p value]) p-value genes Glycosaminoglycan degradation
Late regulation of immune genes and micrornas in circulating leukocytes in a pig model of
 1 Supplementary material for: 2 3 4 5 6 Late regulation of immune genes and micrornas in circulating leukocytes in a pig model of influenza A (H1N2) infection Louise Brogaard, Peter M. H. Heegaard, Lars
1 Supplementary material for: 2 3 4 5 6 Late regulation of immune genes and micrornas in circulating leukocytes in a pig model of influenza A (H1N2) infection Louise Brogaard, Peter M. H. Heegaard, Lars
Supplementary Figure 1: Func8onal Network Analysis of Kinases Significantly Modulated by MERS CoV Infec8on and Conserved Across All Time Points
 A. B. 8 4 Supplementary Figure : Func8onal Network Analysis of Kinases Significantly Modulated by MERS CoV Infec8on and Conserved Across All Time Points Examined. A) Venn diagram analysis of kinases significantly
A. B. 8 4 Supplementary Figure : Func8onal Network Analysis of Kinases Significantly Modulated by MERS CoV Infec8on and Conserved Across All Time Points Examined. A) Venn diagram analysis of kinases significantly
Additional file 2 List of pathway from PID
 Additional file 2 List of pathway from PID Pathway ID Pathway name # components # enriched GO terms a4b1_paxdep_pathway Paxillin-dependent events mediated by a4b1 20 179 a4b1_paxindep_pathway Paxillin-independent
Additional file 2 List of pathway from PID Pathway ID Pathway name # components # enriched GO terms a4b1_paxdep_pathway Paxillin-dependent events mediated by a4b1 20 179 a4b1_paxindep_pathway Paxillin-independent
Supplementary Figure 1
 Supplementary Figure 1 Expression of apoptosis-related genes in tumor T reg cells. (a) Identification of FOXP3 T reg cells by FACS. CD45 + cells were gated as enriched lymphoid cell populations with low-granularity.
Supplementary Figure 1 Expression of apoptosis-related genes in tumor T reg cells. (a) Identification of FOXP3 T reg cells by FACS. CD45 + cells were gated as enriched lymphoid cell populations with low-granularity.
The Major Histocompatibility Complex (MHC)
 The Major Histocompatibility Complex (MHC) An introduction to adaptive immune system before we discuss MHC B cells The main cells of adaptive immune system are: -B cells -T cells B cells: Recognize antigens
The Major Histocompatibility Complex (MHC) An introduction to adaptive immune system before we discuss MHC B cells The main cells of adaptive immune system are: -B cells -T cells B cells: Recognize antigens
Supplementary Files. Novel blood-based microrna biomarker panel for early diagnosis of chronic pancreatitis
 Supplementary Files Novel blood-based microrna biomarker panel for early diagnosis of chronic pancreatitis Lei Xin 1 *, M.D., Jun Gao 1 *, Ph.D., Dan Wang 1 *, M.D., Jin-Huan Lin 1, M.D., Zhuan Liao 1,
Supplementary Files Novel blood-based microrna biomarker panel for early diagnosis of chronic pancreatitis Lei Xin 1 *, M.D., Jun Gao 1 *, Ph.D., Dan Wang 1 *, M.D., Jin-Huan Lin 1, M.D., Zhuan Liao 1,
* Kyoto Encyclopedia of Genes and Genomes.
 Supplemental Material Complete gene expression data using Affymetrix 3PRIME IVT ID Chip (54,614 genes) and human immature dendritic cells stimulated with rbmasnrs, IL-8 and control (media) has been deposited
Supplemental Material Complete gene expression data using Affymetrix 3PRIME IVT ID Chip (54,614 genes) and human immature dendritic cells stimulated with rbmasnrs, IL-8 and control (media) has been deposited
Supplementary Material Correlation matrices on FP and FN profiles
 Supplementary Material Correlation matrices on FP and FN profiles The following two tables give the correlation coefficients for the FP profiles and the FN profiles of a single tagging solutions against
Supplementary Material Correlation matrices on FP and FN profiles The following two tables give the correlation coefficients for the FP profiles and the FN profiles of a single tagging solutions against
Signaling Through Immune System Receptors (Ch. 7)
 Signaling Through Immune System Receptors (Ch. 7) 1. General principles of signal transduction and propagation. 2. Antigen receptor signaling and lymphocyte activation. 3. Other receptors and signaling
Signaling Through Immune System Receptors (Ch. 7) 1. General principles of signal transduction and propagation. 2. Antigen receptor signaling and lymphocyte activation. 3. Other receptors and signaling
2.28E E AKT/CAT P-3-4:
 Supplemental Table 1: IPA network analysis of the microarray data presented in Figure 6 showing the most significant molecular and cellular function classifications along with the respective number of
Supplemental Table 1: IPA network analysis of the microarray data presented in Figure 6 showing the most significant molecular and cellular function classifications along with the respective number of
Supplementary Figures
 Supplementary Figures Figure S1. Urodynamic recording of BOO-induced LUTD patients. (A) DO group. BOO patients with increased detrusor pressure and reduced urine flow during pressure flow in combination
Supplementary Figures Figure S1. Urodynamic recording of BOO-induced LUTD patients. (A) DO group. BOO patients with increased detrusor pressure and reduced urine flow during pressure flow in combination
Principles of Genetics and Molecular Biology
 Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
Phosphorylation Site Company Cat #
 Supplemental Table 1. Antibodies used for RPPA analysis. Label Protein Phosphorylation Site Company Cat # Used on MDA_CLSS Used on MDA_Pilot 4EBP1 4EBP1 Cell Signaling 9452 No 4EBP1.pS65 4EBP1 S65 Cell
Supplemental Table 1. Antibodies used for RPPA analysis. Label Protein Phosphorylation Site Company Cat # Used on MDA_CLSS Used on MDA_Pilot 4EBP1 4EBP1 Cell Signaling 9452 No 4EBP1.pS65 4EBP1 S65 Cell
Supplementary Information
 Supplementary Information Supplementary Figure 1. Short-term coreceptor and costimulation blockade induces tolerance to pluripotent human ESCs. (A) Schematic diagram showing experimental approaches and
Supplementary Information Supplementary Figure 1. Short-term coreceptor and costimulation blockade induces tolerance to pluripotent human ESCs. (A) Schematic diagram showing experimental approaches and
Molecular Cell Biology - Problem Drill 19: Cell Signaling Pathways and Gene Expression
 Molecular Cell Biology - Problem Drill 19: Cell Signaling Pathways and Gene Expression Question No. 1 of 10 1. Which statement about cell signaling is correct? Question #1 (A) Cell signaling involves receiving
Molecular Cell Biology - Problem Drill 19: Cell Signaling Pathways and Gene Expression Question No. 1 of 10 1. Which statement about cell signaling is correct? Question #1 (A) Cell signaling involves receiving
Lecture #27 Lecturer A. N. Koval
 Lecture #27 Lecturer A. N. Koval Hormones Transduce Signals to Affect Homeostatic Mechanisms Koval A. (C), 2011 2 Lipophilic hormones Classifying hormones into hydrophilic and lipophilic molecules indicates
Lecture #27 Lecturer A. N. Koval Hormones Transduce Signals to Affect Homeostatic Mechanisms Koval A. (C), 2011 2 Lipophilic hormones Classifying hormones into hydrophilic and lipophilic molecules indicates
Cellular Physiology (PHSI3009) Contents:
 Cellular Physiology (PHSI3009) Contents: Cell membranes and communication 2 nd messenger systems G-coupled protein signalling Calcium signalling Small G-protein signalling o RAS o MAPK o PI3K RHO GTPases
Cellular Physiology (PHSI3009) Contents: Cell membranes and communication 2 nd messenger systems G-coupled protein signalling Calcium signalling Small G-protein signalling o RAS o MAPK o PI3K RHO GTPases
Systems biology approaches and pathway tools for investigating cardiovascular disease
 Systems biology approaches and pathway tools for investigating cardiovascular disease Craig E. Wheelock 1,2*, Åsa M. Wheelock 2,3,4, Shuichi Kawashima 5, Diego Diez 2, Minoru Kanehisa 2,5, Marjan van Erk
Systems biology approaches and pathway tools for investigating cardiovascular disease Craig E. Wheelock 1,2*, Åsa M. Wheelock 2,3,4, Shuichi Kawashima 5, Diego Diez 2, Minoru Kanehisa 2,5, Marjan van Erk
Antigen presenting cells
 Antigen recognition by T and B cells - T and B cells exhibit fundamental differences in antigen recognition - B cells recognize antigen free in solution (native antigen). - T cells recognize antigen after
Antigen recognition by T and B cells - T and B cells exhibit fundamental differences in antigen recognition - B cells recognize antigen free in solution (native antigen). - T cells recognize antigen after
ras Multikinase Inhibitor Multikinase Inhibitor 0.1
 a ras ** * ** * ** ** ** ** un in et m lu Se ib SL G 32 W 7 So 50 ra 74 fe ni b LY W 294 or 0 tm 02 R an ap n a in Ev my er cin ol im BE us Z2 3 En P 5 za I13 st 0 au rin D as SP ati 60 nib C 012 is 5
a ras ** * ** * ** ** ** ** un in et m lu Se ib SL G 32 W 7 So 50 ra 74 fe ni b LY W 294 or 0 tm 02 R an ap n a in Ev my er cin ol im BE us Z2 3 En P 5 za I13 st 0 au rin D as SP ati 60 nib C 012 is 5
Plasma-Seq conducted with blood from male individuals without cancer.
 Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
TCR, MHC and coreceptors
 Cooperation In Immune Responses Antigen processing how peptides get into MHC Antigen processing involves the intracellular proteolytic generation of MHC binding proteins Protein antigens may be processed
Cooperation In Immune Responses Antigen processing how peptides get into MHC Antigen processing involves the intracellular proteolytic generation of MHC binding proteins Protein antigens may be processed
Signal transduction camp signaling pathway ko Signal transduction MAPK signaling pathway ko
 Table S2 - Summary of the KEGG pathway annotation results for the P transcriptome. Pathway Hierarchy1 Pathway Hierarchy2 KEGG Pathway Pathway ID Gene Metabolism Amino acid metabolism Lysine degradation
Table S2 - Summary of the KEGG pathway annotation results for the P transcriptome. Pathway Hierarchy1 Pathway Hierarchy2 KEGG Pathway Pathway ID Gene Metabolism Amino acid metabolism Lysine degradation
Nature Structural & Molecular Biology: doi: /nsmb.3218
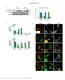 Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh-
 1 a b Supplementary Figure 1. Effects of GSK3b knockdown on poly I:C-induced cytokine production. RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh- GSK3b) were stimulated
1 a b Supplementary Figure 1. Effects of GSK3b knockdown on poly I:C-induced cytokine production. RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh- GSK3b) were stimulated
Pathway Map Reference Guide
 Pathway Map Reference Guide Focus Attention-grabber Your Pathway The most popular cell signaling pathways Sample & Assay Technologies Table of contents AKT Signaling 4 camp Signaling 5 Cellular Apoptosis
Pathway Map Reference Guide Focus Attention-grabber Your Pathway The most popular cell signaling pathways Sample & Assay Technologies Table of contents AKT Signaling 4 camp Signaling 5 Cellular Apoptosis
4. Th1-related gene expression in infected versus mock-infected controls from Fig. 2 with gene annotation.
 List of supplemental information 1. Graph of mouse weight loss during course of infection- Line graphs showing mouse weight data during course of infection days 1 to 10 post-infections (p.i.). 2. Graph
List of supplemental information 1. Graph of mouse weight loss during course of infection- Line graphs showing mouse weight data during course of infection days 1 to 10 post-infections (p.i.). 2. Graph
Enzyme-coupled Receptors. Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors
 Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Table S1: Data-generating Model:
 Table S1: Data-generating Model: Below are the parameters used for the data-generating model (section 1.5). The model was generated using CNORode. The data generating model is included below (figure S1).
Table S1: Data-generating Model: Below are the parameters used for the data-generating model (section 1.5). The model was generated using CNORode. The data generating model is included below (figure S1).
Interpreting Diverse Genomic Data Using Gene Sets
 Interpreting Diverse Genomic Data Using Gene Sets giovanni parmigiani@dfci.harvard.edu FHCRC February 2012 Gene Sets Leary 2008 PNAS Alterations in the combined FGF, EGFR, ERBB2 and PIK3 pathways. Red:
Interpreting Diverse Genomic Data Using Gene Sets giovanni parmigiani@dfci.harvard.edu FHCRC February 2012 Gene Sets Leary 2008 PNAS Alterations in the combined FGF, EGFR, ERBB2 and PIK3 pathways. Red:
Allergy and Immunology Review Corner: Chapter 13 of Immunology IV: Clinical Applications in Health and Disease, by Joseph A. Bellanti, MD.
 Allergy and Immunology Review Corner: Chapter 13 of Immunology IV: Clinical Applications in Health and Disease, by Joseph A. Bellanti, MD. Chapter 13: Mechanisms of Immunity to Viral Disease Prepared by
Allergy and Immunology Review Corner: Chapter 13 of Immunology IV: Clinical Applications in Health and Disease, by Joseph A. Bellanti, MD. Chapter 13: Mechanisms of Immunity to Viral Disease Prepared by
TNFSF13B tumor necrosis factor (ligand) superfamily, member 13b NF-kB pathway cluster, Enrichment Score: 3.57
 Appendix 2. Highly represented clusters of genes in the differential expression of data. Immune Cluster, Enrichment Score: 5.17 GO:0048584 positive regulation of response to stimulus GO:0050778 positive
Appendix 2. Highly represented clusters of genes in the differential expression of data. Immune Cluster, Enrichment Score: 5.17 GO:0048584 positive regulation of response to stimulus GO:0050778 positive
Supplementary Table 1 Clinicopathological characteristics of 35 patients with CRCs
 Supplementary Table Clinicopathological characteristics of 35 patients with CRCs Characteristics Type-A CRC Type-B CRC P value Sex Male / Female 9 / / 8.5 Age (years) Median (range) 6. (9 86) 6.5 (9 76).95
Supplementary Table Clinicopathological characteristics of 35 patients with CRCs Characteristics Type-A CRC Type-B CRC P value Sex Male / Female 9 / / 8.5 Age (years) Median (range) 6. (9 86) 6.5 (9 76).95
Supplementary Material. Part I: Sample Information. Part II: Pathway Information
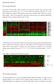 Supplementary Material Part I: Sample Information Three NPC cell lines, CNE1, CNE2, and HK1 were treated with CYC202. Gene expression of 380 selected genes were collected at 0, 2, 4, 6, 12 and 24 hours
Supplementary Material Part I: Sample Information Three NPC cell lines, CNE1, CNE2, and HK1 were treated with CYC202. Gene expression of 380 selected genes were collected at 0, 2, 4, 6, 12 and 24 hours
FOR OPTIMAL GUT HEALTH KEMIN.COM/GUTHEALTH
 FOR OPTIMAL GUT HEALTH KEMIN.COM/GUTHEALTH ALETA A SOURCE OF 1,3-BETA GLUCANS Aleta is highly bioavailable, offering a concentration greater than 5% of 1,3-beta glucans. Aleta provides a consistent response
FOR OPTIMAL GUT HEALTH KEMIN.COM/GUTHEALTH ALETA A SOURCE OF 1,3-BETA GLUCANS Aleta is highly bioavailable, offering a concentration greater than 5% of 1,3-beta glucans. Aleta provides a consistent response
Interpreting Diverse Genomic Data Using Gene Sets
 Interpreting Diverse Genomic Data Using Gene Sets giovanni parmigiani@dfci.harvard.edu FHCRC February 2012 Gene Sets Leary 2008 PNAS Alterations in the combined FGF, EGFR, ERBB2 and PIK3 pathways. Red:
Interpreting Diverse Genomic Data Using Gene Sets giovanni parmigiani@dfci.harvard.edu FHCRC February 2012 Gene Sets Leary 2008 PNAS Alterations in the combined FGF, EGFR, ERBB2 and PIK3 pathways. Red:
Contents. Preface XV Acknowledgments XXI List of Abbreviations XXIII About the Companion Website XXIX
 Contents Preface XV Acknowledgments XXI List of Abbreviations XXIII About the Companion Website XXIX 1 General Aspects of Signal Transduction and Cancer Therapy 1 1.1 General Principles of Signal Transduction
Contents Preface XV Acknowledgments XXI List of Abbreviations XXIII About the Companion Website XXIX 1 General Aspects of Signal Transduction and Cancer Therapy 1 1.1 General Principles of Signal Transduction
CH 03 CELLS: THE LIVING UNITS
 CH 03 CELLS: THE LIVING UNITS This chapter provides a review of critical information regarding cells the basic units of structure and function of all living things. CELL THEORY The cell theory resulted
CH 03 CELLS: THE LIVING UNITS This chapter provides a review of critical information regarding cells the basic units of structure and function of all living things. CELL THEORY The cell theory resulted
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplementary Figure 1. Copy Number Alterations TP53 Mutation Type. C-class TP53 WT. TP53 mut. Nature Genetics: doi: /ng.
 Supplementary Figure a Copy Number Alterations in M-class b TP53 Mutation Type Recurrent Copy Number Alterations 8 6 4 2 TP53 WT TP53 mut TP53-mutated samples (%) 7 6 5 4 3 2 Missense Truncating M-class
Supplementary Figure a Copy Number Alterations in M-class b TP53 Mutation Type Recurrent Copy Number Alterations 8 6 4 2 TP53 WT TP53 mut TP53-mutated samples (%) 7 6 5 4 3 2 Missense Truncating M-class
Screening Protocol and Assay Conditions
 Page 1 of 20 OVERVIEW AND ASSAY THEORY 2 SELECTSCREEN ASSAY CONDITIONS 4 SELECTSCREEN ASSAY CONTROLS 6 SELECTSCREEN DATA ANALYSIS 7 PATHWAY PROFILING CELL LINES AVAILABLE FOR SCREENING 8 CELL LINE-SPECIFIC
Page 1 of 20 OVERVIEW AND ASSAY THEORY 2 SELECTSCREEN ASSAY CONDITIONS 4 SELECTSCREEN ASSAY CONTROLS 6 SELECTSCREEN DATA ANALYSIS 7 PATHWAY PROFILING CELL LINES AVAILABLE FOR SCREENING 8 CELL LINE-SPECIFIC
1. Activated receptor tyrosine kinases (RTKs) phosphorylates themselves
 Enzyme-coupled receptors Transmembrane proteins Ligand-binding domain on the outer surface Cytoplasmic domain acts as an enzyme itself or forms a complex with enzyme 1. Activated receptor tyrosine kinases
Enzyme-coupled receptors Transmembrane proteins Ligand-binding domain on the outer surface Cytoplasmic domain acts as an enzyme itself or forms a complex with enzyme 1. Activated receptor tyrosine kinases
blood mononuclear cells were stained with AAD, annexin, CD3, CD4, CD25, LAP and specific
 Supplementary figure legends Figure 1: Selective expression of regulatory markers on CD4 + LAP + T cells. Human peripheral blood mononuclear cells were stained with AAD, annexin, CD3, CD4, CD25, LAP and
Supplementary figure legends Figure 1: Selective expression of regulatory markers on CD4 + LAP + T cells. Human peripheral blood mononuclear cells were stained with AAD, annexin, CD3, CD4, CD25, LAP and
Cell Biology. A discipline of biology: 1. Cell structure 2. Cellular processes 3. Cell division
 The Cell Cell Biology 1 A discipline of biology: 1. Cell structure 2. Cellular processes 3. Cell division Tight connection with 1. Molecular biology 2. Biochemistry Cell theory 2 1838, 1839 Theodor Schwann
The Cell Cell Biology 1 A discipline of biology: 1. Cell structure 2. Cellular processes 3. Cell division Tight connection with 1. Molecular biology 2. Biochemistry Cell theory 2 1838, 1839 Theodor Schwann
Biology 12 November 2001 Provincial Examination
 Biology 12 November 2001 Provincial Examination ANSWER KEY / SCORING GUIDE CURRICULUM: Organizers 1. Cell Biology 2. Cell Processes and Applications 3. Human Biology Sub-Organizers A, B, C, D E, F, G,
Biology 12 November 2001 Provincial Examination ANSWER KEY / SCORING GUIDE CURRICULUM: Organizers 1. Cell Biology 2. Cell Processes and Applications 3. Human Biology Sub-Organizers A, B, C, D E, F, G,
Characterization of mirna-regulated networks, hubs of signaling, and biomarkers in obstruction-induced bladder dysfunction
 Characterization of mirna-regulated networks, hubs of signaling, and biomarkers in obstruction-induced bladder dysfunction Ali Hashemi Gheinani, 1 Bernhard Kiss, 2 Felix Moltzahn, 2 Irene Keller, 3 Rémy
Characterization of mirna-regulated networks, hubs of signaling, and biomarkers in obstruction-induced bladder dysfunction Ali Hashemi Gheinani, 1 Bernhard Kiss, 2 Felix Moltzahn, 2 Irene Keller, 3 Rémy
T Cell Effector Mechanisms I: B cell Help & DTH
 T Cell Effector Mechanisms I: B cell Help & DTH Ned Braunstein, MD The Major T Cell Subsets p56 lck + T cells γ δ ε ζ ζ p56 lck CD8+ T cells γ δ ε ζ ζ Cα Cβ Vα Vβ CD3 CD8 Cα Cβ Vα Vβ CD3 MHC II peptide
T Cell Effector Mechanisms I: B cell Help & DTH Ned Braunstein, MD The Major T Cell Subsets p56 lck + T cells γ δ ε ζ ζ p56 lck CD8+ T cells γ δ ε ζ ζ Cα Cβ Vα Vβ CD3 CD8 Cα Cβ Vα Vβ CD3 MHC II peptide
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk
 Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Figure 1:
 Supplementary Figure 1: (A) Whole aortic cross-sections stained with Hematoxylin and Eosin (H&E), 7 days after porcine-pancreatic-elastase (PPE)-induced AAA compared to untreated, healthy control aortas
Supplementary Figure 1: (A) Whole aortic cross-sections stained with Hematoxylin and Eosin (H&E), 7 days after porcine-pancreatic-elastase (PPE)-induced AAA compared to untreated, healthy control aortas
PCxN: The pathway co-activity map: a resource for the unification of functional biology
 PCxN: The pathway co-activity map: a resource for the unification of functional biology Sheffield Institute for Translational Neurosciences Center for Integrative Genome Translation GENOME INFORMATICS
PCxN: The pathway co-activity map: a resource for the unification of functional biology Sheffield Institute for Translational Neurosciences Center for Integrative Genome Translation GENOME INFORMATICS
Cell Signaling part 2
 15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
Antigen Presentation and T Lymphocyte Activation. Abul K. Abbas UCSF. FOCiS
 1 Antigen Presentation and T Lymphocyte Activation Abul K. Abbas UCSF FOCiS 2 Lecture outline Dendritic cells and antigen presentation The role of the MHC T cell activation Costimulation, the B7:CD28 family
1 Antigen Presentation and T Lymphocyte Activation Abul K. Abbas UCSF FOCiS 2 Lecture outline Dendritic cells and antigen presentation The role of the MHC T cell activation Costimulation, the B7:CD28 family
The T cell receptor for MHC-associated peptide antigens
 1 The T cell receptor for MHC-associated peptide antigens T lymphocytes have a dual specificity: they recognize polymporphic residues of self MHC molecules, and they also recognize residues of peptide
1 The T cell receptor for MHC-associated peptide antigens T lymphocytes have a dual specificity: they recognize polymporphic residues of self MHC molecules, and they also recognize residues of peptide
Cholera host response gene networks: preliminary studies
 www.bioinformation.net Volume 13(10) Hypothesis Cholera host response gene networks: preliminary studies Paul Shapshak 1 *, Charurut Somboonwit 1, 2, Shu Cao 1, John T. Sinnott 1, 2 1Division of Infectious
www.bioinformation.net Volume 13(10) Hypothesis Cholera host response gene networks: preliminary studies Paul Shapshak 1 *, Charurut Somboonwit 1, 2, Shu Cao 1, John T. Sinnott 1, 2 1Division of Infectious
Basic Immunology. Lecture 5 th and 6 th Recognition by MHC. Antigen presentation and MHC restriction
 Basic Immunology Lecture 5 th and 6 th Recognition by MHC. Antigen presentation and MHC restriction Molecular structure of MHC, subclasses, genetics, functions. Antigen presentation and MHC restriction.
Basic Immunology Lecture 5 th and 6 th Recognition by MHC. Antigen presentation and MHC restriction Molecular structure of MHC, subclasses, genetics, functions. Antigen presentation and MHC restriction.
Supplementary Figure 1.
 Supplementary Figure 1. Increased expression of cell cycle pathway genes in insulin + Glut2 low cells of STZ-induced diabetic islets. A) random blood glucose measuers of STZ and vehicle treated MIP-GFP
Supplementary Figure 1. Increased expression of cell cycle pathway genes in insulin + Glut2 low cells of STZ-induced diabetic islets. A) random blood glucose measuers of STZ and vehicle treated MIP-GFP
T cell maturation. T-cell Maturation. What allows T cell maturation?
 T-cell Maturation What allows T cell maturation? Direct contact with thymic epithelial cells Influence of thymic hormones Growth factors (cytokines, CSF) T cell maturation T cell progenitor DN DP SP 2ry
T-cell Maturation What allows T cell maturation? Direct contact with thymic epithelial cells Influence of thymic hormones Growth factors (cytokines, CSF) T cell maturation T cell progenitor DN DP SP 2ry
Nature Immunology: doi: /ni Supplementary Figure 1. Cytokine pattern in skin in response to urushiol.
 Supplementary Figure 1 Cytokine pattern in skin in response to urushiol. Wild-type (WT) and CD1a-tg mice (n = 3 per group) were sensitized and challenged with urushiol (uru) or vehicle (veh). Quantitative
Supplementary Figure 1 Cytokine pattern in skin in response to urushiol. Wild-type (WT) and CD1a-tg mice (n = 3 per group) were sensitized and challenged with urushiol (uru) or vehicle (veh). Quantitative
HORMONES AND CELL SIGNALLING
 HORMONES AND CELL SIGNALLING TYPES OF CELL JUNCTIONS CHEMICAL SIGNALS AND MODES OF ACTION Endocrine system produces chemical messages = hormones that are transported from endocrine gland to target cell
HORMONES AND CELL SIGNALLING TYPES OF CELL JUNCTIONS CHEMICAL SIGNALS AND MODES OF ACTION Endocrine system produces chemical messages = hormones that are transported from endocrine gland to target cell
G-Protein-Coupled Receptors
 Cellular Signalling Cells must be ready to respond to essential signals in their environment. These are often chemicals in the extracellular fluid (ECF) from distant locations in a multicellular organism
Cellular Signalling Cells must be ready to respond to essential signals in their environment. These are often chemicals in the extracellular fluid (ECF) from distant locations in a multicellular organism
fasting blood glucose [mg/dl]
![fasting blood glucose [mg/dl] fasting blood glucose [mg/dl]](/thumbs/80/82129652.jpg) SUPPLEMENTL MTERIL Supplemental Figure I body weight [g] 5 5 5 fasting blood glucose [mg/dl] 5 5 5 C total cholesterol [mg/dl] 8 6 4 WT Has -/- Supplemental Figure I: Has-deficient mice exhibited no apparent
SUPPLEMENTL MTERIL Supplemental Figure I body weight [g] 5 5 5 fasting blood glucose [mg/dl] 5 5 5 C total cholesterol [mg/dl] 8 6 4 WT Has -/- Supplemental Figure I: Has-deficient mice exhibited no apparent
Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction. Marker Sequence (5 3 ) Accession No.
 Supplementary Tables Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction Marker Sequence (5 3 ) Accession No. Angiopoietin 1, ANGPT1 A CCCTCCGGTGAATATTGGCTGG NM_001146.3
Supplementary Tables Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction Marker Sequence (5 3 ) Accession No. Angiopoietin 1, ANGPT1 A CCCTCCGGTGAATATTGGCTGG NM_001146.3
silent epidemic,. (WHO),
 Tel: 02-740-8686; E-mail: hhbkim@snu.ac.kr silent epidemic,. (WHO),. 5 3, 1. 50 70. 50%, 25%, 20% (12~35%). 2.8% 0.7% 4. ( ). bone remodeling (osteoblast), (osteoclast),.. 3~4.. 70% (osteocyte) (bone lining
Tel: 02-740-8686; E-mail: hhbkim@snu.ac.kr silent epidemic,. (WHO),. 5 3, 1. 50 70. 50%, 25%, 20% (12~35%). 2.8% 0.7% 4. ( ). bone remodeling (osteoblast), (osteoclast),.. 3~4.. 70% (osteocyte) (bone lining
T cell-mediated immunity
 T cell-mediated immunity Overview For microbes within phagosomes in phagocytes.cd4+ T lymphocytes (TH1) Activate phagocyte by cytokines studies on Listeria monocytogenes For microbes infecting and replicating
T cell-mediated immunity Overview For microbes within phagosomes in phagocytes.cd4+ T lymphocytes (TH1) Activate phagocyte by cytokines studies on Listeria monocytogenes For microbes infecting and replicating
Significance of the MHC
 CHAPTER 8 Major Histocompatibility Complex (MHC) What is is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) Significance of the MHC role in immune response role in organ
CHAPTER 8 Major Histocompatibility Complex (MHC) What is is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) Significance of the MHC role in immune response role in organ
A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy se
 A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy serves as a defence mechanism that prevents or retards
A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy serves as a defence mechanism that prevents or retards
Significance of the MHC
 CHAPTER 8 Major Histocompatibility Complex (MHC) What is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) - MHC molecules were initially discovered during studies aimed
CHAPTER 8 Major Histocompatibility Complex (MHC) What is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) - MHC molecules were initially discovered during studies aimed
Mass is the key. Maartje van den Biggelaar. British Society for Proteome Research meeting CIDA September 29 th Sanquin Plasma Proteins
 Mass is the key Maartje van den Biggelaar British Society for Proteome Research meeting 2013 CIDA September 29 th 2016 Maartje van den Biggelaar Sanquin Plasma Proteins Mass spectrometry approaches to
Mass is the key Maartje van den Biggelaar British Society for Proteome Research meeting 2013 CIDA September 29 th 2016 Maartje van den Biggelaar Sanquin Plasma Proteins Mass spectrometry approaches to
The Adaptive Immune Response: T lymphocytes and Their Functional Types *
 OpenStax-CNX module: m46560 1 The Adaptive Immune Response: T lymphocytes and Their Functional Types * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution
OpenStax-CNX module: m46560 1 The Adaptive Immune Response: T lymphocytes and Their Functional Types * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution
Cornstarch
 Electronic Supplementary Material (ESI) for Food & Function. This journal is The Royal Society of Chemistry 2018 Supplementary data : Supplementary Table 1: Diet composition (g/kg) on the basis of the
Electronic Supplementary Material (ESI) for Food & Function. This journal is The Royal Society of Chemistry 2018 Supplementary data : Supplementary Table 1: Diet composition (g/kg) on the basis of the
SUPPLEMENTARY DATA Supplementary Figure 1. Body weight and fat mass of AdicerKO mice.
 SUPPLEMENTARY DATA Supplementary Figure 1. Body weight and fat mass of AdicerKO mice. Twelve week old mice were subjected to ad libitum (AL) or dietary restriction (DR) regimens for three months. (A) Body
SUPPLEMENTARY DATA Supplementary Figure 1. Body weight and fat mass of AdicerKO mice. Twelve week old mice were subjected to ad libitum (AL) or dietary restriction (DR) regimens for three months. (A) Body
Trim29 gene-targeting strategy. (a) Genotyping of wildtype mice (+/+), Trim29 heterozygous mice (+/ ) and homozygous mice ( / ).
 Supplementary Figure 1 Trim29 gene-targeting strategy. (a) Genotyping of wildtype mice (+/+), Trim29 heterozygous mice (+/ ) and homozygous mice ( / ). (b) Immunoblot analysis of TRIM29 in lung primary
Supplementary Figure 1 Trim29 gene-targeting strategy. (a) Genotyping of wildtype mice (+/+), Trim29 heterozygous mice (+/ ) and homozygous mice ( / ). (b) Immunoblot analysis of TRIM29 in lung primary
Supplemental Figure S1. RANK expression on human lung cancer cells.
 Supplemental Figure S1. RANK expression on human lung cancer cells. (A) Incidence and H-Scores of RANK expression determined from IHC in the indicated primary lung cancer subgroups. The overall expression
Supplemental Figure S1. RANK expression on human lung cancer cells. (A) Incidence and H-Scores of RANK expression determined from IHC in the indicated primary lung cancer subgroups. The overall expression
Network-assisted data analysis
 Network-assisted data analysis Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Protein identification in shotgun proteomics Protein digestion LC-MS/MS Protein
Network-assisted data analysis Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Protein identification in shotgun proteomics Protein digestion LC-MS/MS Protein
fl/+ KRas;Atg5 fl/+ KRas;Atg5 fl/fl KRas;Atg5 fl/fl KRas;Atg5 Supplementary Figure 1. Gene set enrichment analyses. (a) (b)
 KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001
 BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001 SS# Name This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. Good luck! 1. (2) The
BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001 SS# Name This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. Good luck! 1. (2) The
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/264/rs4/dc1 Supplementary Materials for A Systems Approach for Decoding Mitochondrial Retrograde Signaling Pathways Sehyun Chae, Byung Yong Ahn, Kyunghee Byun,
www.sciencesignaling.org/cgi/content/full/6/264/rs4/dc1 Supplementary Materials for A Systems Approach for Decoding Mitochondrial Retrograde Signaling Pathways Sehyun Chae, Byung Yong Ahn, Kyunghee Byun,
Identification of Tissue-Specific Protein-Coding and Noncoding. Transcripts across 14 Human Tissues Using RNA-seq
 Identification of Tissue-Specific Protein-Coding and Noncoding Transcripts across 14 Human Tissues Using RNA-seq Jinhang Zhu 1, Geng Chen 1, Sibo Zhu 1,2, Suqing Li 3, Zhuo Wen 3, Bin Li 1, Yuanting Zheng
Identification of Tissue-Specific Protein-Coding and Noncoding Transcripts across 14 Human Tissues Using RNA-seq Jinhang Zhu 1, Geng Chen 1, Sibo Zhu 1,2, Suqing Li 3, Zhuo Wen 3, Bin Li 1, Yuanting Zheng
Basic Immunology. Immunological tolerance. Cellular and molecular mechanisms of the immunological tolerance. Lecture 23 rd
 Basic Immunology Lecture 23 rd Immunological tolerance Cellular and molecular mechanisms of the immunological tolerance Tolerated skin grafts on MHC (H2) identical mice TOLERANCE & AUTOIMMUNITY Upon encountering
Basic Immunology Lecture 23 rd Immunological tolerance Cellular and molecular mechanisms of the immunological tolerance Tolerated skin grafts on MHC (H2) identical mice TOLERANCE & AUTOIMMUNITY Upon encountering
Patient characteristics of training and validation set. Patient selection and inclusion overview can be found in Supp Data 9. Training set (103)
 Roepman P, et al. An immune response enriched 72-gene prognostic profile for early stage Non-Small- Supplementary Data 1. Patient characteristics of training and validation set. Patient selection and inclusion
Roepman P, et al. An immune response enriched 72-gene prognostic profile for early stage Non-Small- Supplementary Data 1. Patient characteristics of training and validation set. Patient selection and inclusion
Molecular Pathways Involved in Prostate Carcinogenesis: Insights from Public Microarray Datasets
 Molecular Pathways Involved in Prostate Carcinogenesis: Insights from Public Microarray Datasets Sarah C. Baetke 1,2, Michiel E. Adriaens 1,3, Renaud Seigneuric 4,5, Chris T. Evelo 1, Lars M. T. Eijssen
Molecular Pathways Involved in Prostate Carcinogenesis: Insights from Public Microarray Datasets Sarah C. Baetke 1,2, Michiel E. Adriaens 1,3, Renaud Seigneuric 4,5, Chris T. Evelo 1, Lars M. T. Eijssen
Supplementary Information
 Supplementary Information Modelling the Yeast Interactome Vuk Janjić, Roded Sharan 2 and Nataša Pržulj, Department of Computing, Imperial College London, London, United Kingdom 2 Blavatnik School of Computer
Supplementary Information Modelling the Yeast Interactome Vuk Janjić, Roded Sharan 2 and Nataša Pržulj, Department of Computing, Imperial College London, London, United Kingdom 2 Blavatnik School of Computer
Molecular Cell Biology - Problem Drill 17: Intracellular Vesicular Traffic
 Molecular Cell Biology - Problem Drill 17: Intracellular Vesicular Traffic Question No. 1 of 10 1. Which of the following statements about clathrin-coated vesicles is correct? Question #1 (A) There are
Molecular Cell Biology - Problem Drill 17: Intracellular Vesicular Traffic Question No. 1 of 10 1. Which of the following statements about clathrin-coated vesicles is correct? Question #1 (A) There are
Nature Genetics: doi: /ng Supplementary Figure 1. Phenotypic characterization of MES- and ADRN-type cells.
 Supplementary Figure 1 Phenotypic characterization of MES- and ADRN-type cells. (a) Bright-field images showing cellular morphology of MES-type (691-MES, 700-MES, 717-MES) and ADRN-type (691-ADRN, 700-
Supplementary Figure 1 Phenotypic characterization of MES- and ADRN-type cells. (a) Bright-field images showing cellular morphology of MES-type (691-MES, 700-MES, 717-MES) and ADRN-type (691-ADRN, 700-
Lecture 6 - Intracellular compartments and transport I
 01.25.10 Lecture 6 - Intracellular compartments and transport I Intracellular transport and compartments 1. Protein sorting: How proteins get to their appropriate destinations within the cell 2. Vesicular
01.25.10 Lecture 6 - Intracellular compartments and transport I Intracellular transport and compartments 1. Protein sorting: How proteins get to their appropriate destinations within the cell 2. Vesicular
Cellular Messengers. Intracellular Communication
 Cellular Messengers Intracellular Communication Most common cellular communication is done through extracellular chemical messengers: Ligands Specific in function 1. Paracrines Local messengers (neighboring
Cellular Messengers Intracellular Communication Most common cellular communication is done through extracellular chemical messengers: Ligands Specific in function 1. Paracrines Local messengers (neighboring
Physiology Unit 1 CELL SIGNALING: CHEMICAL MESSENGERS AND SIGNAL TRANSDUCTION PATHWAYS
 Physiology Unit 1 CELL SIGNALING: CHEMICAL MESSENGERS AND SIGNAL TRANSDUCTION PATHWAYS In Physiology Today Cell Communication Homeostatic mechanisms maintain a normal balance of the body s internal environment
Physiology Unit 1 CELL SIGNALING: CHEMICAL MESSENGERS AND SIGNAL TRANSDUCTION PATHWAYS In Physiology Today Cell Communication Homeostatic mechanisms maintain a normal balance of the body s internal environment
Receptor mediated Signal Transduction
 Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
T-cell activation T cells migrate to secondary lymphoid tissues where they interact with antigen, antigen-presenting cells, and other lymphocytes:
 Interactions between innate immunity & adaptive immunity What happens to T cells after they leave the thymus? Naïve T cells exit the thymus and enter the bloodstream. If they remain in the bloodstream,
Interactions between innate immunity & adaptive immunity What happens to T cells after they leave the thymus? Naïve T cells exit the thymus and enter the bloodstream. If they remain in the bloodstream,
T-cell activation T cells migrate to secondary lymphoid tissues where they interact with antigen, antigen-presenting cells, and other lymphocytes:
 Interactions between innate immunity & adaptive immunity What happens to T cells after they leave the thymus? Naïve T cells exit the thymus and enter the bloodstream. If they remain in the bloodstream,
Interactions between innate immunity & adaptive immunity What happens to T cells after they leave the thymus? Naïve T cells exit the thymus and enter the bloodstream. If they remain in the bloodstream,
q 2017 by The University of Chicago. All rights reserved. DOI: /690105
 q 2017 by The University of Chicago. All rights reserved. DOI: 10.1086/690105 Appendix from R. M. Cox et al., Hormonally Mediated Increases in Sex-Biased Gene Expression Accompany the Breakdown of Between-
q 2017 by The University of Chicago. All rights reserved. DOI: 10.1086/690105 Appendix from R. M. Cox et al., Hormonally Mediated Increases in Sex-Biased Gene Expression Accompany the Breakdown of Between-
BIOL 4374/BCHS 4313 Cell Biology Exam #2 March 22, 2001
 BIOL 4374/BCHS 4313 Cell Biology Exam #2 March 22, 2001 SS# Name This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. Good luck! 1. (2) In the
BIOL 4374/BCHS 4313 Cell Biology Exam #2 March 22, 2001 SS# Name This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. Good luck! 1. (2) In the
Supplementary Materials for
 www.sciencetranslationalmedicine.org/cgi/content/full/7/303/303ra139/dc1 Supplementary Materials for Magnetic resonance image features identify glioblastoma phenotypic subtypes with distinct molecular
www.sciencetranslationalmedicine.org/cgi/content/full/7/303/303ra139/dc1 Supplementary Materials for Magnetic resonance image features identify glioblastoma phenotypic subtypes with distinct molecular
Clinical significance of CD44 expression in children with hepatoblastoma
 Clinical significance of CD44 expression in children with hepatoblastoma H.-Y. Cai 1 *, B. Yu 1 *, Z.-C. Feng 2, X. Qi 1 and X.-J. Wei 1 1 Department of General Surgery, General Hospital of Beijing Military
Clinical significance of CD44 expression in children with hepatoblastoma H.-Y. Cai 1 *, B. Yu 1 *, Z.-C. Feng 2, X. Qi 1 and X.-J. Wei 1 1 Department of General Surgery, General Hospital of Beijing Military
Supplementary figure 1
 Supplementary figure 1 Nature Medicine: doi:1.138/nm.275 CLUSTER BY SELF-ORGANIZING MAPS SELECTED PATHWAY ANALISYS TERMS Cluster : up-regulated genes in acute patients Cell cycle/dna repair Fatty acid
Supplementary figure 1 Nature Medicine: doi:1.138/nm.275 CLUSTER BY SELF-ORGANIZING MAPS SELECTED PATHWAY ANALISYS TERMS Cluster : up-regulated genes in acute patients Cell cycle/dna repair Fatty acid
Table S7B- Biocarta functional annotation of celecoxib-modulated genes unique to COX-2 expressers
 Table S7B- Biocarta functional annotation of celecoxib-modulated genes unique to COX-2 expressers ListHits ListTotal PopulationHits PopulationTotal Terms 3 72 10 1429 Glycolysis Pathway 3 72 11 1429 Regulation
Table S7B- Biocarta functional annotation of celecoxib-modulated genes unique to COX-2 expressers ListHits ListTotal PopulationHits PopulationTotal Terms 3 72 10 1429 Glycolysis Pathway 3 72 11 1429 Regulation
QUESTIONSHEET 1. The diagram shows some of the cell structures involved in the secretion of an extracellular enzyme. C D
 QUESTIONSHEET 1 The diagram shows some of the cell structures involved in the secretion of an extracellular enzyme. C D A (a) Identify A,, C, and D. A:... :... C:... D:... [4] (b) Outline the role of each
QUESTIONSHEET 1 The diagram shows some of the cell structures involved in the secretion of an extracellular enzyme. C D A (a) Identify A,, C, and D. A:... :... C:... D:... [4] (b) Outline the role of each
