Supplementary Materials for
|
|
|
- Anis Erica Charles
- 5 years ago
- Views:
Transcription
1 Supplementary Materials for Magnetic resonance image features identify glioblastoma phenotypic subtypes with distinct molecular pathway activities Haruka Itakura, Achal S. Achrol, Lex A. Mitchell, Joshua J. Loya, Tiffany Liu, Erick M. Westbroek, Abdullah H. Feroze, Scott Rodriguez, Sebastian Echegaray, Tej D. Azad, Kristen W. Yeom, Sandy Napel, Daniel L. Rubin, Steven D. Chang, Griffith R. Harsh IV, Olivier Gevaert* The PDF file includes: *Corresponding author. Published 2 September 2015, Sci. Transl. Med. 7, 303ra139 (2015) DOI: /scitranslmed.aaa7582 Methods Fig. S1. Relative contributions of the top 24 imaging features in characterizing each of the clusters. Table S1. Clinical characteristics of the Stanford cohort before and after selection of the development cohort. Table S2. Clinical characteristics of the TCGA cohort before and after selection of the validation cohort. Table S3. All 2D and multislice 2D quantitative MR image features used for analysis. Table S4. Two-dimensional and multislice 2D quantitative MR image features significantly associated with each cluster. Table S5. Cluster assignment by subjects in the development cohort with and without midline-crossing lesions. Table S6. Regulatory signaling pathways significantly associated with each cluster. References (45 47)
2 SUPPLEMENTARY METHODS Two-dimensional feature representations To represent 2D features, we computed quantitative image features from the largest slice of the tumor volume. To represent multislice 2D features, we aggregated 2D features from all slices within the three-dimensional tumor volume. The quantitative imaging feature extraction pipeline thus generated separate 2D and multislice 2D features for each patient. For each subject in each cohort, this generated a feature vector of 193 2D, and 195 multislice 2D quantitative image features, resulting in a total of 388 features (table S3). Validation of clusters To validate the reproducibility of the clusters derived from consensus clustering in the development cohort, we used the IGP analysis to demonstrate the existence of these clusters in the validation cohort. IGP, which measures the proportion of samples in the group whose nearest neighbor is also classified to the same group, validated the reproducibility and robustness of the generated clusters by quantifying how well centroids from the development cohort predicted test set co-memberships (24). Statistical significance in IGP denotes low probabilities that the clusters originated from a null distribution, in which centroids are placed randomly in the data and, consequently, result in low quality clusters. The p-value of a cluster is the fraction of the null distribution IGPs that approximate the value of 1 (perfect IGP) more than the actual IGP of the cluster. The null hypothesis that a cluster is not a high-quality cluster is rejected if the actual IGP of the group is high enough (i.e. approximates 1). A significant p-value indicates that a high-quality cluster corresponding to the original cluster was found in an independent
3 dataset (validation cohort). Hence, the development cohort cluster is deemed reproducible and validated. Next we trained a PAM model on the development cohort, fitting a nearest shrunken centroid classifier on the centroids generated from the development cohort clusters. By fitting these centroids in the validation cohort, we assigned individual patients to imaging subtypes defined by the development cohort. Mapping canonical pathways to imaging subtypes We used the matched molecular data in the TCGA validation cohort to assign canonical signaling pathways to the discovered imaging subtypes. TCGA provided CNV and gene expression data for 560 and 495 subjects, respectively. These data were integrated with curated pathways from the NCI Pathway Interaction Database using the PARADIGM algorithm to infer signaling pathway activities (up-regulation or down-regulation) to generate a molecular signaling pathway consisting of 131 features. Next we used the SAM algorithm to identify specific regulatory pathways that were significantly associated with each of the identified imaging subtypes for each of the 144 subjects. We placed the FDR threshold at <5%. Evaluating imaging subtypes for traditional risk factors We examined the subtypes for statistically significant differences in historically associated risk factors. Such factors included both clinical (age, KPS) (4, 45, 46) and molecular factors, including previously described gene expression cluster molecular subgroups (8), as well as CIMP status (10), IDH1 mutation (6, 7), MGMT hypermethylation (47), and EGFR amplification (9). Tumor volume data were available only for the development cohort. We
4 compared across-cluster values for all available data, using the Kruskal-Wallis one-way analysis of variance by ranks test for continuous variables and Fisher s exact test for categorical variables.
5 SUPPLEMENTARY FIGURES 2D-MS-LAII2R3.iqr 2D-MS-LAII2R3.madmedian 2D-MS-LAII2R3.madmean 2D-MS-LAII2R3.var 2D-MS-LAII2R3.min 2D-MS-LAII2R5.iqr 2D-MS-RDS.robustmin 2D-MS-RDS.trimmedmean25perc 2D-MS-RDS.mean 2D-MS-RDS.min 2D-MS-EdgeSharpnessWindow.robustmin 2D-MS-EdgeSharpnessWindow.mode 2D-MS-EdgeSharpnessWindow.min 2D-MS-compactness2dmean 2D-LAII2R5.range 2D-LAII2R5.madmean 2D-LAII2R8.range 2D-RDS.robustmin 2D-EdgeSharpnessScale.range 2D-EdgeSharpnessScale.iqr 2D-EdgeSharpnessScale.madmedian 2D-EdgeSharpnessScale.madmean 2D-EdgeSharpnessWindow.min 2D-Compactness2dmean Cluster 1 Cluster 2 Cluster Figure S1. Relative contributions of the top 24 imaging features in characterizing each of the clusters. The graphic displays the multivariate combination of the top 24 differentially expressed 2D and multislice 2D (2D-MS) imaging features across the three clusters (table S4). Displacement of colored bars to the right of the central vertical line demonstrates over-expression, whereas displacement to the left indicates under-expression of the given imaging feature. The length of the bar signifies magnitude.
6 SUPPLEMENTARY TABLES Table S1. Clinical characteristics of the Stanford cohort before and after selection of the development cohort. Stanford Cohort Stanford Development Cohort Characteristics (n = 364) (n = 121) Age, mean (SD) 62.2 (14.1) 64.6 (14.1) Sex, n male (%) 209 (57) 69 (57) KPS, n (%) >70% 192 (53) 65 (54) 50-70% 141 (39) 44 (37) <50% 28 (8) 11 (9) Mean ± SD 71.7 ± ± 17
7 Table S2. Clinical characteristics of the TCGA cohort before and after selection of the validation cohort. Of the validation cohort, 114 patients had clinical data. TCGA Cohort TCGA Validation Cohort Characteristic (n = 575) (n = 144) Age, mean ± SD 57.9 ± ± 15 Sex, n male (%) 352 (61) 73 (64) KPS, n (%) >70% 317 (55) 78 (68) 50-70% 89 (15) 19 (17) <50% 18 (3) 1 (1) Missing Mean ± SD 77.2 ± ± 12.3
8 Table S3. All 2D and multislice 2D quantitative MR image features used for analysis. Quantitative image features were extracted using a pipeline previously applied in non small cell lung cancer (19) and glioblastoma (15) to characterize intensity, shape, edge sharpness, and texture of lesions from images. Number Image feature 1 2D-spherecity 2 2D-volume 3 2D-surfaceArea 4 2D-roughness2dmean 5 2D-compactness2dmean 6 2D-LowIntensityProportion 7 2D-HistogramRaw-min 8 2D-HistogramRaw-mean 9 2D-HistogramRaw-median 10 2D-HistogramRaw-max 11 2D-HistogramRaw-var 12 2D-HistogramRaw-skewness 13 2D-HistogramRaw-kurtosis 14 2D-HistogramRaw-madmean 15 2D-HistogramRaw-madmedian 16 2D-HistogramRaw-iqr 17 2D-HistogramRaw-mode 18 2D-HistogramRaw-range 19 2D-HistogramRaw-trimmedmean25perc 20 2D-HistogramRaw-harmmean 21 2D-HistogramRaw-robustmin 22 2D-HistogramRaw-robustmax 23 2D-HistogramRaw-robustskewness 24 2D-HistogramRaw-robustkurtosis 25 2D-HistogramRaw-entropy 26 2D-EdgeSharpnessWindow-min 27 2D-EdgeSharpnessWindow-mean 28 2D-EdgeSharpnessWindow-median 29 2D-EdgeSharpnessWindow-max 30 2D-EdgeSharpnessWindow-var 31 2D-EdgeSharpnessWindow-skewness 32 2D-EdgeSharpnessWindow-kurtosis 33 2D-EdgeSharpnessWindow-madmean 34 2D-EdgeSharpnessWindow-madmedian 35 2D-EdgeSharpnessWindow-iqr 36 2D-EdgeSharpnessWindow-mode 37 2D-EdgeSharpnessWindow-range 38 2D-EdgeSharpnessWindow-trimmedmean25perc 39 2D-EdgeSharpnessWindow-harmmean
9 40 2D-EdgeSharpnessWindow-robustmin 41 2D-EdgeSharpnessWindow-robustmax 42 2D-EdgeSharpnessWindow-robustskewness 43 2D-EdgeSharpnessWindow-robustkurtosis 44 2D-EdgeSharpnessWindow-entropy 45 2D-EdgeSharpnessScale-min 46 2D-EdgeSharpnessScale-mean 47 2D-EdgeSharpnessScale-median 48 2D-EdgeSharpnessScale-max 49 2D-EdgeSharpnessScale-var 50 2D-EdgeSharpnessScale-skewness 51 2D-EdgeSharpnessScale-kurtosis 52 2D-EdgeSharpnessScale-madmean 53 2D-EdgeSharpnessScale-madmedian 54 2D-EdgeSharpnessScale-iqr 55 2D-EdgeSharpnessScale-mode 56 2D-EdgeSharpnessScale-range 57 2D-EdgeSharpnessScale-trimmedmean25perc 58 2D-EdgeSharpnessScale-harmmean 59 2D-EdgeSharpnessScale-robustmin 60 2D-EdgeSharpnessScale-robustmax 61 2D-EdgeSharpnessScale-robustskewness 62 2D-EdgeSharpnessScale-robustkurtosis 63 2D-EdgeSharpnessScale-entropy 64 2D-EdgeSharpnessBlurryness-min 65 2D-EdgeSharpnessBlurryness-mean 66 2D-EdgeSharpnessBlurryness-median 67 2D-EdgeSharpnessBlurryness-max 68 2D-EdgeSharpnessBlurryness-var 69 2D-EdgeSharpnessBlurryness-skewness 70 2D-EdgeSharpnessBlurryness-kurtosis 71 2D-EdgeSharpnessBlurryness-madmean 72 2D-EdgeSharpnessBlurryness-madmedian 73 2D-EdgeSharpnessBlurryness-iqr 74 2D-EdgeSharpnessBlurryness-mode 75 2D-EdgeSharpnessBlurryness-range 76 2D-EdgeSharpnessBlurryness-trimmedmean25perc 77 2D-EdgeSharpnessBlurryness-harmmean 78 2D-EdgeSharpnessBlurryness-robustmin 79 2D-EdgeSharpnessBlurryness-robustmax 80 2D-EdgeSharpnessBlurryness-robustskewness 81 2D-EdgeSharpnessBlurryness-robustkurtosis 82 2D-EdgeSharpnessBlurryness-entropy 83 2D-RDS-min
10 84 2D-RDS-mean 85 2D-RDS-median 86 2D-RDS-var 87 2D-RDS-skewness 88 2D-RDS-kurtosis 89 2D-RDS-madmean 90 2D-RDS-madmedian 91 2D-RDS-iqr 92 2D-RDS-range 93 2D-RDS-trimmedmean25perc 94 2D-RDS-harmmean 95 2D-RDS-robustmin 96 2D-RDS-robustskewness 97 2D-RDS-robustkurtosis 98 2D-RDS-entropy 99 2D-LAII2R10-min 100 2D-LAII2R10-mean 101 2D-LAII2R10-median 102 2D-LAII2R10-max 103 2D-LAII2R10-var 104 2D-LAII2R10-skewness 105 2D-LAII2R10-kurtosis 106 2D-LAII2R10-madmean 107 2D-LAII2R10-madmedian 108 2D-LAII2R10-iqr 109 2D-LAII2R10-mode 110 2D-LAII2R10-range 111 2D-LAII2R10-trimmedmean25perc 112 2D-LAII2R10-harmmean 113 2D-LAII2R10-robustmin 114 2D-LAII2R10-robustmax 115 2D-LAII2R10-robustskewness 116 2D-LAII2R10-robustkurtosis 117 2D-LAII2R10-entropy 118 2D-LAII2R8-min 119 2D-LAII2R8-mean 120 2D-LAII2R8-median 121 2D-LAII2R8-max 122 2D-LAII2R8-var 123 2D-LAII2R8-skewness 124 2D-LAII2R8-kurtosis 125 2D-LAII2R8-madmean 126 2D-LAII2R8-madmedian 127 2D-LAII2R8-iqr
11 128 2D-LAII2R8-mode 129 2D-LAII2R8-range 130 2D-LAII2R8-trimmedmean25perc 131 2D-LAII2R8-harmmean 132 2D-LAII2R8-robustmin 133 2D-LAII2R8-robustmax 134 2D-LAII2R8-robustskewness 135 2D-LAII2R8-robustkurtosis 136 2D-LAII2R8-entropy 137 2D-LAII2R5-min 138 2D-LAII2R5-mean 139 2D-LAII2R5-median 140 2D-LAII2R5-max 141 2D-LAII2R5-var 142 2D-LAII2R5-skewness 143 2D-LAII2R5-kurtosis 144 2D-LAII2R5-madmean 145 2D-LAII2R5-madmedian 146 2D-LAII2R5-iqr 147 2D-LAII2R5-mode 148 2D-LAII2R5-range 149 2D-LAII2R5-trimmedmean25perc 150 2D-LAII2R5-harmmean 151 2D-LAII2R5-robustmin 152 2D-LAII2R5-robustmax 153 2D-LAII2R5-robustskewness 154 2D-LAII2R5-robustkurtosis 155 2D-LAII2R5-entropy 156 2D-LAII2R3-min 157 2D-LAII2R3-mean 158 2D-LAII2R3-median 159 2D-LAII2R3-max 160 2D-LAII2R3-var 161 2D-LAII2R3-skewness 162 2D-LAII2R3-kurtosis 163 2D-LAII2R3-madmean 164 2D-LAII2R3-madmedian 165 2D-LAII2R3-iqr 166 2D-LAII2R3-mode 167 2D-LAII2R3-range 168 2D-LAII2R3-trimmedmean25perc 169 2D-LAII2R3-harmmean 170 2D-LAII2R3-robustmin 171 2D-LAII2R3-robustmax
12 172 2D-LAII2R3-robustskewness 173 2D-LAII2R3-robustkurtosis 174 2D-LAII2R3-entropy 175 2D-LAII2R2-min 176 2D-LAII2R2-mean 177 2D-LAII2R2-median 178 2D-LAII2R2-max 179 2D-LAII2R2-var 180 2D-LAII2R2-skewness 181 2D-LAII2R2-kurtosis 182 2D-LAII2R2-madmean 183 2D-LAII2R2-madmedian 184 2D-LAII2R2-iqr 185 2D-LAII2R2-mode 186 2D-LAII2R2-range 187 2D-LAII2R2-trimmedmean25perc 188 2D-LAII2R2-harmmean 189 2D-LAII2R2-robustmin 190 2D-LAII2R2-robustmax 191 2D-LAII2R2-robustskewness 192 2D-LAII2R2-robustkurtosis 193 2D-LAII2R2-entropy 194 2D-MultiSlice-spherecity 195 2D-MultiSlice-volume 196 2D-MultiSlice-surfaceArea 197 2D-MultiSlice-roughness2dmean 198 2D-MultiSlice-roughness2dvar 199 2D-MultiSlice-compactness2dmean 200 2D-MultiSlice-compactness2dvar 201 2D-MultiSlice-LowIntensityProportion 202 2D-MultiSlice-HistogramRaw-min 203 2D-MultiSlice-HistogramRaw-mean 204 2D-MultiSlice-HistogramRaw-median 205 2D-MultiSlice-HistogramRaw-max 206 2D-MultiSlice-HistogramRaw-var 207 2D-MultiSlice-HistogramRaw-skewness 208 2D-MultiSlice-HistogramRaw-kurtosis 209 2D-MultiSlice-HistogramRaw-madmean 210 2D-MultiSlice-HistogramRaw-madmedian 211 2D-MultiSlice-HistogramRaw-iqr 212 2D-MultiSlice-HistogramRaw-mode 213 2D-MultiSlice-HistogramRaw-range 214 2D-MultiSlice-HistogramRaw-trimmedmean25perc 215 2D-MultiSlice-HistogramRaw-harmmean
13 216 2D-MultiSlice-HistogramRaw-robustmin 217 2D-MultiSlice-HistogramRaw-robustmax 218 2D-MultiSlice-HistogramRaw-robustskewness 219 2D-MultiSlice-HistogramRaw-robustkurtosis 220 2D-MultiSlice-HistogramRaw-entropy 221 2D-MultiSlice-EdgeSharpnessWindow-min 222 2D-MultiSlice-EdgeSharpnessWindow-mean 223 2D-MultiSlice-EdgeSharpnessWindow-median 224 2D-MultiSlice-EdgeSharpnessWindow-max 225 2D-MultiSlice-EdgeSharpnessWindow-var 226 2D-MultiSlice-EdgeSharpnessWindow-skewness 227 2D-MultiSlice-EdgeSharpnessWindow-kurtosis 228 2D-MultiSlice-EdgeSharpnessWindow-madmean 229 2D-MultiSlice-EdgeSharpnessWindow-madmedian 230 2D-MultiSlice-EdgeSharpnessWindow-iqr 231 2D-MultiSlice-EdgeSharpnessWindow-mode 232 2D-MultiSlice-EdgeSharpnessWindow-range 233 2D-MultiSlice-EdgeSharpnessWindow-trimmedmean25perc 234 2D-MultiSlice-EdgeSharpnessWindow-harmmean 235 2D-MultiSlice-EdgeSharpnessWindow-robustmin 236 2D-MultiSlice-EdgeSharpnessWindow-robustmax 237 2D-MultiSlice-EdgeSharpnessWindow-robustskewness 238 2D-MultiSlice-EdgeSharpnessWindow-robustkurtosis 239 2D-MultiSlice-EdgeSharpnessWindow-entropy 240 2D-MultiSlice-EdgeSharpnessScale-min 241 2D-MultiSlice-EdgeSharpnessScale-mean 242 2D-MultiSlice-EdgeSharpnessScale-median 243 2D-MultiSlice-EdgeSharpnessScale-max 244 2D-MultiSlice-EdgeSharpnessScale-var 245 2D-MultiSlice-EdgeSharpnessScale-skewness 246 2D-MultiSlice-EdgeSharpnessScale-kurtosis 247 2D-MultiSlice-EdgeSharpnessScale-madmean 248 2D-MultiSlice-EdgeSharpnessScale-madmedian 249 2D-MultiSlice-EdgeSharpnessScale-iqr 250 2D-MultiSlice-EdgeSharpnessScale-mode 251 2D-MultiSlice-EdgeSharpnessScale-range 252 2D-MultiSlice-EdgeSharpnessScale-trimmedmean25perc 253 2D-MultiSlice-EdgeSharpnessScale-harmmean 254 2D-MultiSlice-EdgeSharpnessScale-robustmin 255 2D-MultiSlice-EdgeSharpnessScale-robustmax 256 2D-MultiSlice-EdgeSharpnessScale-robustskewness 257 2D-MultiSlice-EdgeSharpnessScale-robustkurtosis 258 2D-MultiSlice-EdgeSharpnessScale-entropy 259 2D-MultiSlice-EdgeSharpnessBlurryness-min
14 260 2D-MultiSlice-EdgeSharpnessBlurryness-mean 261 2D-MultiSlice-EdgeSharpnessBlurryness-median 262 2D-MultiSlice-EdgeSharpnessBlurryness-max 263 2D-MultiSlice-EdgeSharpnessBlurryness-var 264 2D-MultiSlice-EdgeSharpnessBlurryness-skewness 265 2D-MultiSlice-EdgeSharpnessBlurryness-kurtosis 266 2D-MultiSlice-EdgeSharpnessBlurryness-madmean 267 2D-MultiSlice-EdgeSharpnessBlurryness-madmedian 268 2D-MultiSlice-EdgeSharpnessBlurryness-iqr 269 2D-MultiSlice-EdgeSharpnessBlurryness-mode 270 2D-MultiSlice-EdgeSharpnessBlurryness-range 271 2D-MultiSlice-EdgeSharpnessBlurrynesstrimmedmean25perc 272 2D-MultiSlice-EdgeSharpnessBlurryness-harmmean 273 2D-MultiSlice-EdgeSharpnessBlurryness-robustmin 274 2D-MultiSlice-EdgeSharpnessBlurryness-robustmax 275 2D-MultiSlice-EdgeSharpnessBlurryness-robustskewness 276 2D-MultiSlice-EdgeSharpnessBlurryness-robustkurtosis 277 2D-MultiSlice-EdgeSharpnessBlurryness-entropy 278 2D-MultiSlice-RDS-min 279 2D-MultiSlice-RDS-mean 280 2D-MultiSlice-RDS-median 281 2D-MultiSlice-RDS-var 282 2D-MultiSlice-RDS-skewness 283 2D-MultiSlice-RDS-kurtosis 284 2D-MultiSlice-RDS-madmean 285 2D-MultiSlice-RDS-madmedian 286 2D-MultiSlice-RDS-iqr 287 2D-MultiSlice-RDS-range 288 2D-MultiSlice-RDS-trimmedmean25perc 289 2D-MultiSlice-RDS-harmmean 290 2D-MultiSlice-RDS-robustmin 291 2D-MultiSlice-RDS-robustskewness 292 2D-MultiSlice-RDS-robustkurtosis 293 2D-MultiSlice-RDS-entropy 294 2D-MultiSlice-LAII2R10-min 295 2D-MultiSlice-LAII2R10-mean 296 2D-MultiSlice-LAII2R10-median 297 2D-MultiSlice-LAII2R10-max 298 2D-MultiSlice-LAII2R10-var 299 2D-MultiSlice-LAII2R10-skewness 300 2D-MultiSlice-LAII2R10-kurtosis 301 2D-MultiSlice-LAII2R10-madmean 302 2D-MultiSlice-LAII2R10-madmedian
15 303 2D-MultiSlice-LAII2R10-iqr 304 2D-MultiSlice-LAII2R10-mode 305 2D-MultiSlice-LAII2R10-range 306 2D-MultiSlice-LAII2R10-trimmedmean25perc 307 2D-MultiSlice-LAII2R10-harmmean 308 2D-MultiSlice-LAII2R10-robustmin 309 2D-MultiSlice-LAII2R10-robustmax 310 2D-MultiSlice-LAII2R10-robustskewness 311 2D-MultiSlice-LAII2R10-robustkurtosis 312 2D-MultiSlice-LAII2R10-entropy 313 2D-MultiSlice-LAII2R8-min 314 2D-MultiSlice-LAII2R8-mean 315 2D-MultiSlice-LAII2R8-median 316 2D-MultiSlice-LAII2R8-max 317 2D-MultiSlice-LAII2R8-var 318 2D-MultiSlice-LAII2R8-skewness 319 2D-MultiSlice-LAII2R8-kurtosis 320 2D-MultiSlice-LAII2R8-madmean 321 2D-MultiSlice-LAII2R8-madmedian 322 2D-MultiSlice-LAII2R8-iqr 323 2D-MultiSlice-LAII2R8-mode 324 2D-MultiSlice-LAII2R8-range 325 2D-MultiSlice-LAII2R8-trimmedmean25perc 326 2D-MultiSlice-LAII2R8-harmmean 327 2D-MultiSlice-LAII2R8-robustmin 328 2D-MultiSlice-LAII2R8-robustmax 329 2D-MultiSlice-LAII2R8-robustskewness 330 2D-MultiSlice-LAII2R8-robustkurtosis 331 2D-MultiSlice-LAII2R8-entropy 332 2D-MultiSlice-LAII2R5-min 333 2D-MultiSlice-LAII2R5-mean 334 2D-MultiSlice-LAII2R5-median 335 2D-MultiSlice-LAII2R5-max 336 2D-MultiSlice-LAII2R5-var 337 2D-MultiSlice-LAII2R5-skewness 338 2D-MultiSlice-LAII2R5-kurtosis 339 2D-MultiSlice-LAII2R5-madmean 340 2D-MultiSlice-LAII2R5-madmedian 341 2D-MultiSlice-LAII2R5-iqr 342 2D-MultiSlice-LAII2R5-mode 343 2D-MultiSlice-LAII2R5-range 344 2D-MultiSlice-LAII2R5-trimmedmean25perc 345 2D-MultiSlice-LAII2R5-harmmean 346 2D-MultiSlice-LAII2R5-robustmin
16 347 2D-MultiSlice-LAII2R5-robustmax 348 2D-MultiSlice-LAII2R5-robustskewness 349 2D-MultiSlice-LAII2R5-robustkurtosis 350 2D-MultiSlice-LAII2R5-entropy 351 2D-MultiSlice-LAII2R3-min 352 2D-MultiSlice-LAII2R3-mean 353 2D-MultiSlice-LAII2R3-median 354 2D-MultiSlice-LAII2R3-max 355 2D-MultiSlice-LAII2R3-var 356 2D-MultiSlice-LAII2R3-skewness 357 2D-MultiSlice-LAII2R3-kurtosis 358 2D-MultiSlice-LAII2R3-madmean 359 2D-MultiSlice-LAII2R3-madmedian 360 2D-MultiSlice-LAII2R3-iqr 361 2D-MultiSlice-LAII2R3-mode 362 2D-MultiSlice-LAII2R3-range 363 2D-MultiSlice-LAII2R3-trimmedmean25perc 364 2D-MultiSlice-LAII2R3-harmmean 365 2D-MultiSlice-LAII2R3-robustmin 366 2D-MultiSlice-LAII2R3-robustmax 367 2D-MultiSlice-LAII2R3-robustskewness 368 2D-MultiSlice-LAII2R3-robustkurtosis 369 2D-MultiSlice-LAII2R3-entropy 370 2D-MultiSlice-LAII2R2-min 371 2D-MultiSlice-LAII2R2-mean 372 2D-MultiSlice-LAII2R2-median 373 2D-MultiSlice-LAII2R2-max 374 2D-MultiSlice-LAII2R2-var 375 2D-MultiSlice-LAII2R2-skewness 376 2D-MultiSlice-LAII2R2-kurtosis 377 2D-MultiSlice-LAII2R2-madmean 378 2D-MultiSlice-LAII2R2-madmedian 379 2D-MultiSlice-LAII2R2-iqr 380 2D-MultiSlice-LAII2R2-mode 381 2D-MultiSlice-LAII2R2-range 382 2D-MultiSlice-LAII2R2-trimmedmean25perc 383 2D-MultiSlice-LAII2R2-harmmean 384 2D-MultiSlice-LAII2R2-robustmin 385 2D-MultiSlice-LAII2R2-robustmax 386 2D-MultiSlice-LAII2R2-robustskewness 387 2D-MultiSlice-LAII2R2-robustkurtosis 388 2D-MultiSlice-LAII2R2-entropy
17 Table S4. Two-dimensional and multislice 2D quantitative MR image features significantly associated with each cluster. For each feature the fold change is reported and defined as the number of times the feature is expressed in samples in the cluster versus in samples not in the cluster. For example, a 3.1-fold increase was observed for the top feature in Cluster 1 compared with patients not in Cluster 1. The features are ranked by descending order of fold changes. Cluster 1 Image feature Fold change 2D-MultiSlice-LAII2R3-iqr D-MultiSlice-LAII2R3-madmedian D-LAII2R5-madmean D-MultiSlice-LAII2R5-iqr D-MultiSlice-LAII2R5-madmedian D-LAII2R3-madmean D-LAII2R5-iqr D-LAII2R3-iqr D-MultiSlice-LAII2R8-robustmax D-MultiSlice-LAII2R3-robustmax D-LAII2R3-madmedian D-LAII2R2-var D-MultiSlice-LAII2R5-madmean D-LAII2R5-madmedian D-MultiSlice-LAII2R5-robustmax D-MultiSlice-LAII2R8-iqr D-LAII2R8-madmean D-MultiSlice-LAII2R3-madmean D-LAII2R2-madmean D-RDS-range D-MultiSlice-LAII2R2-madmedian D-RDS-var D-MultiSlice-LAII2R8-madmedian D-MultiSlice-LAII2R8-madmean D-MultiSlice-LAII2R2-iqr D-MultiSlice-LAII2R10-madmedian D-MultiSlice-LAII2R5-var D-LAII2R3-range D-MultiSlice-RDS-madmean D-LAII2R3-var D-RDS-madmean D-LAII2R5-var D-MultiSlice-RDS-iqr D-LAII2R10-madmean D-LAII2R2-madmedian 2.67
18 2D-LAII2R8-var D-MultiSlice-RDS-var D-LAII2R3-robustmax D-LAII2R5-robustmax D-MultiSlice-RDS-madmedian D-LAII2R8-robustmax D-MultiSlice-LAII2R3-var D-LAII2R2-iqr D-MultiSlice-LAII2R10-robustmax D-MultiSlice-LAII2R10-madmean D-LAII2R10-robustmax D-LAII2R2-range D-LAII2R5-range D-MultiSlice-LAII2R10-iqr D-RDS-iqr D-MultiSlice-LAII2R2-madmean D-LAII2R10-var D-MultiSlice-LAII2R8-var D-LAII2R8-range D-LAII2R8-madmedian D-LAII2R8-iqr D-RDS-madmedian D-LAII2R10-range D-MultiSlice-LAII2R10-var D-roughness2dmean D-LAII2R3-max D-MultiSlice-roughness2dmean D-MultiSlice-LAII2R2-var D-LAII2R10-iqr D-LAII2R5-max D-MultiSlice-LAII2R10-entropy D-MultiSlice-LAII2R2-entropy D-LAII2R8-max D-MultiSlice-LAII2R2-robustmax D-LAII2R10-max D-LAII2R10-madmedian D-MultiSlice-LAII2R8-entropy D-MultiSlice-LAII2R5-entropy D-MultiSlice-LAII2R3-entropy D-MultiSlice-roughness2dvar D-MultiSlice-RDS-entropy D-MultiSlice-compactness2dvar D-MultiSlice-LAII2R5-range D-MultiSlice-LAII2R3-robustskewness 1.84
19 2D-MultiSlice-LAII2R3-range D-MultiSlice-HistogramRaw-robustkurtosis D-LAII2R2-robustmax D-MultiSlice-RDS-skewness D-MultiSlice-LAII2R2-robustskewness D-MultiSlice-LAII2R8-range D-MultiSlice-LAII2R10-range D-MultiSlice-RDS-range D-MultiSlice-RDS-robustskewness D-MultiSlice-LAII2R5-max D-LAII2R2-entropy D-LAII2R2-max D-MultiSlice-HistogramRaw-kurtosis D-EdgeSharpnessWindow-robustskewness D-MultiSlice-LAII2R3-max D-MultiSlice-EdgeSharpnessWindow-robustskewness D-LAII2R3-robustskewness D-MultiSlice-EdgeSharpnessBlurrynessrobustskewness D-compactness2dmean D-MultiSlice-RDS-mean D-MultiSlice-RDS-trimmedmean25perc D-RDS-robustmin D-RDS-min D-RDS-harmmean D-MultiSlice-RDS-harmmean D-MultiSlice-RDS-median D-RDS-mean D-MultiSlice-LAII2R2-median D-MultiSlice-compactness2dmean D-MultiSlice-LAII2R2-trimmedmean25perc D-LAII2R2-median D-LAII2R2-robustmin D-RDS-trimmedmean25perc D-LAII2R2-min D-LAII2R2-harmmean D-LAII2R5-robustmin D-LAII2R3-robustmin D-LAII2R2-trimmedmean25perc D-MultiSlice-LAII2R2-mean D-LAII2R5-harmmean D-LAII2R5-min D-RDS-median D-LAII2R3-median 0.40
20 2D-LAII2R3-min D-LAII2R3-harmmean D-LAII2R8-robustmin D-MultiSlice-LAII2R3-median D-MultiSlice-LAII2R3-trimmedmean25perc D-LAII2R8-min D-MultiSlice-LAII2R5-robustmin D-LAII2R2-mean D-LAII2R10-robustmin D-MultiSlice-LAII2R3-harmmean D-LAII2R10-min D-spherecity D-MultiSlice-LAII2R2-harmmean D-LAII2R3-trimmedmean25perc D-LAII2R8-harmmean D-MultiSlice-LAII2R8-robustmin D-MultiSlice-LAII2R3-mean D-MultiSlice-LAII2R3-robustmin D-MultiSlice-spherecity D-MultiSlice-LAII2R10-robustmin D-MultiSlice-RDS-robustmin D-MultiSlice-LAII2R2-robustkurtosis D-MultiSlice-LAII2R3-robustkurtosis D-LAII2R5-median D-MultiSlice-LAII2R5-trimmedmean25perc D-LAII2R10-harmmean D-MultiSlice-LAII2R5-median D-MultiSlice-LAII2R2-robustmin D-MultiSlice-RDS-robustkurtosis D-MultiSlice-LAII2R3-min D-MultiSlice-LAII2R2-kurtosis D-MultiSlice-LAII2R5-min D-MultiSlice-LAII2R2-mode D-MultiSlice-RDS-kurtosis D-LAII2R3-mode D-LAII2R2-mode D-MultiSlice-LAII2R5-harmmean D-MultiSlice-LAII2R2-min D-LAII2R3-mean D-MultiSlice-LAII2R3-kurtosis D-MultiSlice-LAII2R5-kurtosis D-LAII2R5-trimmedmean25perc D-HistogramRaw-robustskewness D-MultiSlice-HistogramRaw-robustskewness 0.61
21 2D-MultiSlice-LAII2R5-mean D-MultiSlice-LAII2R3-mode D-MultiSlice-RDS-min D-MultiSlice-HistogramRaw-skewness D-HistogramRaw-skewness D-MultiSlice-LAII2R8-min 0.67 Cluster 2 Image feature Fold change 2D-MultiSlice-compactness2dmean D-MultiSlice-RDS-min D-MultiSlice-RDS-robustmin D-MultiSlice-LAII2R3-min D-MultiSlice-RDS-harmmean D-compactness2dmean D-MultiSlice-LAII2R2-robustmin D-MultiSlice-LAII2R3-robustmin D-MultiSlice-LAII2R2-min D-MultiSlice-RDS-mean D-RDS-min D-RDS-robustmin D-LAII2R5-min D-LAII2R8-min D-LAII2R3-min D-LAII2R5-robustmin D-LAII2R8-robustmin D-LAII2R3-robustmin D-MultiSlice-RDS-trimmedmean25perc D-LAII2R2-min D-LAII2R2-robustmin D-MultiSlice-LAII2R5-robustmin D-RDS-harmmean D-RDS-mean D-RDS-trimmedmean25perc D-MultiSlice-RDS-median D-RDS-median D-LAII2R10-robustmin D-MultiSlice-LAII2R2-harmmean D-LAII2R10-min D-MultiSlice-LAII2R2-trimmedmean25perc D-MultiSlice-LAII2R2-median D-MultiSlice-LAII2R2-mean D-MultiSlice-LAII2R5-min D-MultiSlice-LAII2R3-harmmean 1.77
22 2D-MultiSlice-LAII2R8-robustmin D-LAII2R2-harmmean D-MultiSlice-EdgeSharpnessBlurryness-entropy D-LAII2R2-trimmedmean25perc D-LAII2R3-harmmean D-LAII2R2-median D-LAII2R3-median D-MultiSlice-EdgeSharpnessWindow-min D-MultiSlice-EdgeSharpnessWindow-mode D-MultiSlice-EdgeSharpnessWindow-robustmin D-LAII2R2-mean D-MultiSlice-EdgeSharpnessScale-robustmin D-LAII2R5-harmmean D-MultiSlice-EdgeSharpnessScale-min D-MultiSlice-EdgeSharpnessScale-mode D-MultiSlice-LAII2R3-trimmedmean25perc D-MultiSlice-LAII2R3-median D-MultiSlice-EdgeSharpnessBlurryness-robustmin D-MultiSlice-spherecity D-MultiSlice-LAII2R10-robustmin D-EdgeSharpnessWindow-min D-EdgeSharpnessWindow-mode D-EdgeSharpnessWindow-robustmin D-LAII2R3-trimmedmean25perc D-MultiSlice-LAII2R3-mean D-LAII2R8-range D-LAII2R5-range D-MultiSlice-LAII2R3-var D-MultiSlice-LAII2R3-madmean D-MultiSlice-LAII2R3-range D-LAII2R10-range D-LAII2R8-max D-MultiSlice-LAII2R5-range D-MultiSlice-LAII2R8-robustmax D-MultiSlice-RDS-range D-LAII2R3-range D-MultiSlice-RDS-var D-MultiSlice-LAII2R5-madmean D-LAII2R10-max D-MultiSlice-LAII2R5-var D-MultiSlice-RDS-madmean D-MultiSlice-roughness2dmean D-LAII2R5-max D-MultiSlice-LAII2R2-madmean 0.47
23 2D-MultiSlice-LAII2R2-range D-LAII2R5-madmean D-MultiSlice-LAII2R5-robustmax D-MultiSlice-LAII2R2-var D-MultiSlice-LAII2R3-robustmax D-RDS-range D-MultiSlice-LAII2R8-max D-LAII2R2-range D-MultiSlice-LAII2R3-iqr D-MultiSlice-LAII2R3-madmedian D-MultiSlice-compactness2dvar D-LAII2R3-madmean D-MultiSlice-LAII2R5-max D-MultiSlice-LAII2R5-iqr D-MultiSlice-LAII2R2-madmedian D-MultiSlice-LAII2R5-madmedian D-MultiSlice-LAII2R8-range D-LAII2R2-var D-LAII2R8-madmean D-MultiSlice-RDS-iqr D-LAII2R5-var D-MultiSlice-LAII2R10-robustmax D-LAII2R3-max D-MultiSlice-LAII2R2-iqr D-LAII2R2-madmean D-MultiSlice-LAII2R10-max D-MultiSlice-RDS-madmedian D-LAII2R8-var D-RDS-madmean D-RDS-var D-surfaceArea D-LAII2R3-var D-MultiSlice-LAII2R10-range D-MultiSlice-LAII2R8-iqr D-MultiSlice-LAII2R8-madmean D-MultiSlice-LAII2R2-robustmax D-MultiSlice-LAII2R3-max D-LAII2R3-madmedian D-LAII2R10-kurtosis D-LAII2R3-iqr D-LAII2R8-kurtosis D-MultiSlice-LAII2R8-madmedian D-LAII2R10-madmean D-LAII2R10-var 0.56
24 2D-MultiSlice-LAII2R8-var D-MultiSlice-roughness2dvar D-LAII2R3-robustmax D-LAII2R5-madmedian D-roughness2dmean D-LAII2R10-robustmax D-LAII2R5-iqr D-RDS-iqr D-MultiSlice-surfaceArea D-RDS-madmedian D-LAII2R8-robustmax D-LAII2R2-iqr D-LAII2R5-robustmax D-LAII2R2-madmedian D-EdgeSharpnessScale-madmedian D-LAII2R10-robustkurtosis D-MultiSlice-LAII2R10-madmedian D-LAII2R8-robustkurtosis D-volume D-MultiSlice-EdgeSharpnessScale-range D-EdgeSharpnessScale-range D-MultiSlice-LAII2R10-madmean D-EdgeSharpnessScale-iqr D-EdgeSharpnessScale-madmean D-MultiSlice-EdgeSharpnessScale-madmean D-LAII2R5-kurtosis D-MultiSlice-volume D-MultiSlice-EdgeSharpnessScale-iqr D-MultiSlice-EdgeSharpnessScale-madmedian D-MultiSlice-EdgeSharpnessScale-var D-EdgeSharpnessScale-var D-LAII2R8-iqr D-MultiSlice-LAII2R10-iqr D-MultiSlice-LAII2R10-var D-LAII2R8-madmedian D-MultiSlice-EdgeSharpnessBlurryness-madmean D-MultiSlice-HistogramRaw-range D-LAII2R10-iqr D-LAII2R5-robustkurtosis D-LAII2R10-madmedian D-HistogramRaw-range D-MultiSlice-EdgeSharpnessBlurryness-range D-MultiSlice-EdgeSharpnessBlurryness-robustkurtosis D-MultiSlice-LAII2R2-max 0.70
25 2D-LAII2R2-max D-LAII2R2-robustmax D-MultiSlice-EdgeSharpnessBlurryness-iqr D-LAII2R3-kurtosis D-MultiSlice-EdgeSharpnessWindow-range D-MultiSlice-HistogramRaw-max D-MultiSlice-EdgeSharpnessScale-robustmax D-MultiSlice-EdgeSharpnessBlurryness-max D-EdgeSharpnessWindow-range D-MultiSlice-EdgeSharpnessBlurryness-kurtosis D-HistogramRaw-max D-MultiSlice-EdgeSharpnessBlurryness-madmedian D-LAII2R10-mean D-MultiSlice-EdgeSharpnessScale-max D-EdgeSharpnessScale-max 0.74 Cluster 3 Image feature Fold change 2D-EdgeSharpnessScale-madmedian D-EdgeSharpnessScale-iqr D-EdgeSharpnessScale-range D-EdgeSharpnessScale-madmean D-EdgeSharpnessScale-var D-MultiSlice-EdgeSharpnessScale-madmean D-MultiSlice-HistogramRaw-robustskewness D-MultiSlice-EdgeSharpnessScale-madmedian D-MultiSlice-EdgeSharpnessScale-range D-MultiSlice-EdgeSharpnessScale-var D-MultiSlice-EdgeSharpnessScale-iqr D-spherecity D-HistogramRaw-robustskewness D-MultiSlice-HistogramRaw-skewness D-HistogramRaw-skewness D-volume D-MultiSlice-volume D-MultiSlice-EdgeSharpnessBlurryness-madmean D-LAII2R8-kurtosis D-MultiSlice-HistogramRaw-madmean D-LAII2R5-kurtosis D-HistogramRaw-range D-MultiSlice-LowIntensityProportion D-MultiSlice-EdgeSharpnessBlurryness-robustkurtosis D-MultiSlice-HistogramRaw-iqr D-MultiSlice-HistogramRaw-range 1.86
26 2D-LowIntensityProportion D-HistogramRaw-madmean D-MultiSlice-RDS-robustkurtosis D-MultiSlice-LAII2R2-robustkurtosis D-MultiSlice-EdgeSharpnessBlurryness-range D-LAII2R10-kurtosis D-MultiSlice-LAII2R2-range D-MultiSlice-surfaceArea D-MultiSlice-HistogramRaw-var D-MultiSlice-LAII2R3-robustkurtosis D-MultiSlice-LAII2R10-trimmedmean25perc D-HistogramRaw-var D-MultiSlice-LAII2R10-median D-MultiSlice-LAII2R10-mean D-MultiSlice-LAII2R3-kurtosis D-MultiSlice-LAII2R8-trimmedmean25perc D-HistogramRaw-max D-MultiSlice-HistogramRaw-max D-MultiSlice-LAII2R2-kurtosis D-MultiSlice-EdgeSharpnessWindow-range D-MultiSlice-LAII2R8-mean D-MultiSlice-LAII2R8-median D-HistogramRaw-iqr D-MultiSlice-RDS-kurtosis D-MultiSlice-EdgeSharpnessBlurryness-max D-MultiSlice-HistogramRaw-madmedian D-MultiSlice-LAII2R5-kurtosis D-MultiSlice-HistogramRaw-robustmax D-MultiSlice-EdgeSharpnessWindow-robustkurtosis D-LAII2R3-kurtosis D-MultiSlice-LAII2R2-max D-MultiSlice-EdgeSharpnessBlurryness-madmedian D-MultiSlice-EdgeSharpnessBlurryness-iqr D-surfaceArea D-MultiSlice-LAII2R5-median D-MultiSlice-LAII2R5-trimmedmean25perc D-MultiSlice-EdgeSharpnessBlurryness-var D-HistogramRaw-robustmax D-LAII2R10-harmmean D-MultiSlice-EdgeSharpnessScale-robustmax D-MultiSlice-LAII2R8-max D-MultiSlice-EdgeSharpnessScale-harmmean D-EdgeSharpnessScale-robustmax D-MultiSlice-EdgeSharpnessBlurryness-kurtosis 1.57
27 2D-MultiSlice-RDS-range D-MultiSlice-LAII2R5-mean D-LAII2R8-harmmean D-EdgeSharpnessBlurryness-robustkurtosis D-EdgeSharpnessBlurryness-kurtosis D-LAII2R8-robustkurtosis D-EdgeSharpnessWindow-robustkurtosis D-EdgeSharpnessScale-max D-MultiSlice-EdgeSharpnessBlurryness-robustmax D-MultiSlice-EdgeSharpnessScale-max D-LAII2R3-robustkurtosis D-LAII2R5-median D-MultiSlice-LAII2R10-max D-LAII2R2-kurtosis D-LAII2R10-median D-LAII2R2-median D-LAII2R10-mean D-LAII2R5-robustkurtosis D-MultiSlice-LAII2R3-range D-MultiSlice-LAII2R5-max D-MultiSlice-EdgeSharpnessWindow-min D-MultiSlice-EdgeSharpnessWindow-mode D-MultiSlice-EdgeSharpnessWindow-robustmin D-EdgeSharpnessWindow-min D-EdgeSharpnessWindow-mode D-EdgeSharpnessWindow-robustmin D-MultiSlice-EdgeSharpnessScale-robustmin D-MultiSlice-EdgeSharpnessScale-min D-MultiSlice-EdgeSharpnessScale-mode D-EdgeSharpnessScale-robustmin D-MultiSlice-EdgeSharpnessBlurryness-robustmin D-MultiSlice-EdgeSharpnessWindow-robustskewness D-EdgeSharpnessScale-min D-EdgeSharpnessScale-mode D-MultiSlice-LAII2R10-entropy D-LAII2R8-entropy D-EdgeSharpnessBlurryness-entropy D-MultiSlice-LAII2R8-entropy D-LAII2R10-entropy D-MultiSlice-EdgeSharpnessBlurryness-entropy D-MultiSlice-LAII2R2-entropy D-MultiSlice-RDS-entropy D-LAII2R5-entropy D-MultiSlice-LAII2R3-entropy 0.55
28 2D-MultiSlice-LAII2R5-entropy D-MultiSlice-EdgeSharpnessBlurryness-min D-MultiSlice-EdgeSharpnessBlurryness-mode D-MultiSlice-RDS-skewness D-MultiSlice-RDS-robustskewness D-MultiSlice-EdgeSharpnessBlurrynessrobustskewness D-EdgeSharpnessWindow-entropy D-MultiSlice-EdgeSharpnessWindow-skewness D-MultiSlice-LAII2R2-robustskewness D-LAII2R3-entropy D-MultiSlice-RDS-min D-LAII2R5-iqr 0.64 Table S5. Cluster assignment by subjects in the development cohort with and without midline-crossing lesions. With midline-crossing, n = 140; without midline crossing, n = 121. With midlinecrossing lesions Cluster Without midlinecrossing lesions Cluster Patient_033 1 Patient_033 1 Patient_047 3 Patient_047 1 Patient_054 2 Patient_054 2 Patient_056 2 Patient_056 2 Patient_066 2 Patient_066 2 Patient_080 1 Patient_080 1 Patient_083 3 Patient_083 3 Patient_084 2 Patient_084 2 Patient_085 3 Patient_085 3 Patient_086 2 Patient_086 2 Patient_088 2 Patient_088 2 Patient_091 2 Patient_091 2 Patient_092 2 Patient_092 2 Patient_094 2 Patient_094 2 Patient_097 3 Patient_097 3 Patient_098 3 Patient_098 3 Patient_099 3 Patient_099 3 Patient_104 2 Patient_104 2 Patient_105 3 Patient_105 2 Patient_106 1 Patient_106 1 Patient_109 1 Patient_109 1 Patient_110 1 Patient_110 1 Patient_114 1 Patient_114 1 Patient_116 3 Patient_116 1
29 Patient_117 2 Patient_117 2 Patient_121 1 Patient_121 1 Patient_124 1 Patient_124 1 Patient_125 1 Patient_125 1 Patient_129 3 Patient_129 3 Patient_133 2 Patient_133 2 Patient_135 1 Patient_135 1 Patient_136 2 Patient_136 2 Patient_137 3 Patient_137 3 Patient_142 1 Patient_142 1 Patient_144 1 Patient_144 1 Patient_145 3 Patient_145 3 Patient_146 2 Patient_146 2 Patient_147 2 Patient_147 2 Patient_149 3 Patient_149 3 Patient_150 2 Patient_150 2 Patient_151 2 Patient_151 2 Patient_154 1 Patient_154 1 Patient_155 1 Patient_155 1 Patient_156 3 Patient_156 3 Patient_157 3 Patient_157 3 Patient_158 3 Patient_158 3 Patient_159 2 Patient_159 2 Patient_160 2 Patient_160 2 Patient_161 2 Patient_161 2 Patient_164 3 Patient_164 3 Patient_166 3 Patient_166 3 Patient_167 2 Patient_167 2 Patient_168 3 Patient_168 3 Patient_169 1 Patient_169 1 Patient_170 1 Patient_170 1 Patient_175 2 Patient_175 2 Patient_178 3 Patient_178 3 Patient_182 2 Patient_182 1 Patient_189 1 Patient_189 1 Patient_190 3 Patient_190 3 Patient_201 2 Patient_201 2 Patient_203 1 Patient_203 1 Patient_204 2 Patient_204 2 Patient_206 2 Patient_206 2 Patient_218 2 Patient_218 2 Patient_220 2 Patient_220 2 Patient_222 3 Patient_222 3 Patient_223 1 Patient_223 1 Patient_224 3 Patient_224 3 Patient_225 3 Patient_225 3 Patient_235 2 Patient_235 2 Patient_236 1 Patient_236 1
30 Patient_238 2 Patient_238 2 Patient_240 2 Patient_240 2 Patient_242 2 Patient_242 2 Patient_243 2 Patient_243 2 Patient_248 1 Patient_248 1 Patient_251 3 Patient_251 3 Patient_252 1 Patient_252 1 Patient_267 1 Patient_267 1 Patient_268 3 Patient_268 3 Patient_272 1 Patient_272 1 Patient_273 1 Patient_273 1 Patient_274 2 Patient_274 2 Patient_275 3 Patient_275 3 Patient_277 2 Patient_277 2 Patient_279 2 Patient_279 2 Patient_283 3 Patient_283 3 Patient_287 2 Patient_287 2 Patient_289 3 Patient_289 3 Patient_290 3 Patient_290 3 Patient_297 2 Patient_297 2 Patient_300 3 Patient_300 3 Patient_305 3 Patient_305 3 Patient_307 1 Patient_307 1 Patient_309 2 Patient_309 2 Patient_315 2 Patient_315 2 Patient_317 2 Patient_317 2 Patient_323 2 Patient_323 2 Patient_324 1 Patient_324 1 Patient_327 1 Patient_327 1 Patient_333 2 Patient_333 2 Patient_335 1 Patient_335 1 Patient_338 3 Patient_338 3 Patient_345 2 Patient_345 2 Patient_356 2 Patient_356 2 Patient_357 3 Patient_357 3 Patient_358 2 Patient_358 2 Patient_362 1 Patient_362 1 Patient_364 1 Patient_364 1 Patient_367 3 Patient_367 3 Patient_375 1 Patient_375 1 Patient_385 2 Patient_385 2 Patient_386 2 Patient_386 2 Patient_387 3 Patient_387 3 Patient_388 2 Patient_388 2 Patient_395 2 Patient_395 2 Patient_400 1 Patient_400 1 Patient_403 1 Patient_403 1 Patient_409 3 Patient_409 1
31 Patient_413 1 Patient_413 1 Patient_128 2 Patient_130 3 Patient_152 3 Patient_165 3 Patient_231 1 Patient_234 3 Patient_241 3 Patient_257 2 Patient_270 2 Patient_271 2 Patient_285 3 Patient_291 1 Patient_293 3 Patient_302 2 Patient_311 1 Patient_312 3 Patient_342 2 Patient_343 2 Patient_379 2
32 Table S6. Regulatory signaling pathways significantly associated with each cluster. Each cluster is associated with up-regulated ( 1.0 fold change) or down-regulated (< 1.0 fold change) signaling pathways as determined by individual gene expression and copy number variation data (CNV) integrated using the Pathway Recognition Algorithm Using Data Integration on Genomic Models (PARADIGM). Only signaling pathways with a false discovery rate (FDR or q-value) <5% are included. Signaling pathways are ranked in descending order of fold changes. Signaling pathway Fold change Cluster 1 Signaling events mediated by stem cell factor receptor (c-kit) Cluster 2 Insulin-mediated glucose transport q-value (%) Signaling events mediated by prolactin (PRL) Platelet-derived growth factor receptor-alpha (PDGFR-α) signaling pathway Reelin signaling pathway Vascular endothelial growth factor receptor 1 (VEGFR1) specific signals Polo-like kinase 1 (PLK1) signaling events Regulation of nuclear mothers against decapentaplegic homolog /3 (SMAD2/3) signaling Signaling events mediated by protein-tyrosine phosphatase 1B (PTP1B) Osteopontin-mediated events Signaling events activated by hepatocyte growth factor receptor (c-met) Alpha-M beta-2 (αmβ2) Integrin signaling Syndecan-1 mediated signaling events Erythropoietin (EPO) signaling pathway Endothelins Forkhead box protein A2 (FOXA2) and FOXA3 transcription factor networks Signaling events mediated by vascular endothelial growth factor receptor 1 (VEGFR1) and VEGFR2 Interleukin 6 (IL6) mediated signaling events Angiopoietin receptor Tie2-mediated signaling IL23-mediated signaling events Signaling events mediated by c-kit Forkhead box protein M1 (FOXM1) transcription factor network Cluster 3 FOXA2 and FOXA3 transcription factor networks PDGFR-β signaling pathway Endothelins
33 Thromboxane A2 receptor signaling Syndecan-4 mediated signaling events Syndecan-1 mediated signaling events IL6 mediated signaling events Osteopontin-mediated events Fc-ε receptor I signaling in mast cells αmβ2 Integrin signaling Angiopoietin receptor Tie2-mediated signaling PTP1B B cell receptor (BCR) signaling pathway Retinoic acid receptor and retinoid X receptor (RXR and RAR) heterodimerization with other nuclear receptor Atypical nuclear factor-κ B pathway PLK1 signaling events Ceramide signaling pathway Noncanonical Wnt signaling pathway SMAD2/3 signaling T cell receptor (TCR) signaling in naïve CD8 + T cells IL1 mediated signaling events Calcium signaling in the CD4 + TCR pathway Ras signaling in the CD4 + TCR pathway Nongenotropic androgen signaling Regulation of p38-α and p38-β VEGFR1 specific signals E-cadherin signaling in the nascent adherens junction PRL PDGFR-α signaling pathway IL27 mediated signaling events Canonical Wnt signaling pathway
Additional file 2 List of pathway from PID
 Additional file 2 List of pathway from PID Pathway ID Pathway name # components # enriched GO terms a4b1_paxdep_pathway Paxillin-dependent events mediated by a4b1 20 179 a4b1_paxindep_pathway Paxillin-independent
Additional file 2 List of pathway from PID Pathway ID Pathway name # components # enriched GO terms a4b1_paxdep_pathway Paxillin-dependent events mediated by a4b1 20 179 a4b1_paxindep_pathway Paxillin-independent
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Nature Medicine: doi: /nm.3967
 Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
Supplementary Figure 1: Func8onal Network Analysis of Kinases Significantly Modulated by MERS CoV Infec8on and Conserved Across All Time Points
 A. B. 8 4 Supplementary Figure : Func8onal Network Analysis of Kinases Significantly Modulated by MERS CoV Infec8on and Conserved Across All Time Points Examined. A) Venn diagram analysis of kinases significantly
A. B. 8 4 Supplementary Figure : Func8onal Network Analysis of Kinases Significantly Modulated by MERS CoV Infec8on and Conserved Across All Time Points Examined. A) Venn diagram analysis of kinases significantly
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
SUPPLEMENTARY FIGURES: Supplementary Figure 1
 SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
8/1/2017. Imaging and Molecular Biomarkers of Lung Cancer Prognosis. Disclosures. The Era of Precision Oncology
 Imaging and Molecular Biomarkers of Lung Cancer Prognosis Ruijiang Li, PhD Assistant Professor of Radiation Oncology 08/01/2017 Stanford University Department of Radiation Oncology School of Medicine Disclosures
Imaging and Molecular Biomarkers of Lung Cancer Prognosis Ruijiang Li, PhD Assistant Professor of Radiation Oncology 08/01/2017 Stanford University Department of Radiation Oncology School of Medicine Disclosures
What you should know before you collect data. BAE 815 (Fall 2017) Dr. Zifei Liu
 What you should know before you collect data BAE 815 (Fall 2017) Dr. Zifei Liu Zifeiliu@ksu.edu Types and levels of study Descriptive statistics Inferential statistics How to choose a statistical test
What you should know before you collect data BAE 815 (Fall 2017) Dr. Zifei Liu Zifeiliu@ksu.edu Types and levels of study Descriptive statistics Inferential statistics How to choose a statistical test
Expert-guided Visual Exploration (EVE) for patient stratification. Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland
 Expert-guided Visual Exploration (EVE) for patient stratification Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland Oncoscape.sttrcancer.org Paul Lisa Ken Jenny Desert Eric The challenge Given - patient clinical
Expert-guided Visual Exploration (EVE) for patient stratification Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland Oncoscape.sttrcancer.org Paul Lisa Ken Jenny Desert Eric The challenge Given - patient clinical
CoINcIDE: A framework for discovery of patient subtypes across multiple datasets
 Planey and Gevaert Genome Medicine (2016) 8:27 DOI 10.1186/s13073-016-0281-4 METHOD CoINcIDE: A framework for discovery of patient subtypes across multiple datasets Catherine R. Planey and Olivier Gevaert
Planey and Gevaert Genome Medicine (2016) 8:27 DOI 10.1186/s13073-016-0281-4 METHOD CoINcIDE: A framework for discovery of patient subtypes across multiple datasets Catherine R. Planey and Olivier Gevaert
Digitizing the Proteomes From Big Tissue Biobanks
 Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Journal: Nature Methods
 Journal: Nature Methods Article Title: Network-based stratification of tumor mutations Corresponding Author: Trey Ideker Supplementary Item Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure
Journal: Nature Methods Article Title: Network-based stratification of tumor mutations Corresponding Author: Trey Ideker Supplementary Item Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure
Gene Ontology and Functional Enrichment. Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein
 Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Supplementary Materials for
 www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Identification of Tissue Independent Cancer Driver Genes
 Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
DNA methylation signatures for 2016 WHO classification subtypes of diffuse gliomas
 Paul et al. Clinical Epigenetics (2017) 9:32 DOI 10.1186/s13148-017-0331-9 RESEARCH Open Access DNA methylation signatures for 2016 WHO classification subtypes of diffuse gliomas Yashna Paul, Baisakhi
Paul et al. Clinical Epigenetics (2017) 9:32 DOI 10.1186/s13148-017-0331-9 RESEARCH Open Access DNA methylation signatures for 2016 WHO classification subtypes of diffuse gliomas Yashna Paul, Baisakhi
Broad GDAC. Lung Adenocarcinoma AWG Run 2013_02_07. Dan DiCara Hailei Zhang Michael Noble
 Broad GDAC Lung Adenocarcinoma AWG Run 2013_02_07 Dan DiCara Hailei Zhang Michael Noble Copyright 2013 Broad Institute. All rights reserved. gdac@broadinstitute.org http://gdac.broadinstitute.org GDAC
Broad GDAC Lung Adenocarcinoma AWG Run 2013_02_07 Dan DiCara Hailei Zhang Michael Noble Copyright 2013 Broad Institute. All rights reserved. gdac@broadinstitute.org http://gdac.broadinstitute.org GDAC
Inference of patient-specific pathway activities from multi-dimensional cancer genomics data using PARADIGM. Bioinformatics, 2010
 Inference of patient-specific pathway activities from multi-dimensional cancer genomics data using PARADIGM. Bioinformatics, 2010 C.J.Vaske et al. May 22, 2013 Presented by: Rami Eitan Complex Genomic
Inference of patient-specific pathway activities from multi-dimensional cancer genomics data using PARADIGM. Bioinformatics, 2010 C.J.Vaske et al. May 22, 2013 Presented by: Rami Eitan Complex Genomic
TITLE: A Data-Driven Approach to Patient Risk Stratification for Acute Respiratory Distress Syndrome (ARDS)
 TITLE: A Data-Driven Approach to Patient Risk Stratification for Acute Respiratory Distress Syndrome (ARDS) AUTHORS: Tejas Prahlad INTRODUCTION Acute Respiratory Distress Syndrome (ARDS) is a condition
TITLE: A Data-Driven Approach to Patient Risk Stratification for Acute Respiratory Distress Syndrome (ARDS) AUTHORS: Tejas Prahlad INTRODUCTION Acute Respiratory Distress Syndrome (ARDS) is a condition
Expanded View Figures
 Solip Park & Ben Lehner Epistasis is cancer type specific Molecular Systems Biology Expanded View Figures A B G C D E F H Figure EV1. Epistatic interactions detected in a pan-cancer analysis and saturation
Solip Park & Ben Lehner Epistasis is cancer type specific Molecular Systems Biology Expanded View Figures A B G C D E F H Figure EV1. Epistatic interactions detected in a pan-cancer analysis and saturation
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Lecture Outline. Biost 517 Applied Biostatistics I. Purpose of Descriptive Statistics. Purpose of Descriptive Statistics
 Biost 517 Applied Biostatistics I Scott S. Emerson, M.D., Ph.D. Professor of Biostatistics University of Washington Lecture 3: Overview of Descriptive Statistics October 3, 2005 Lecture Outline Purpose
Biost 517 Applied Biostatistics I Scott S. Emerson, M.D., Ph.D. Professor of Biostatistics University of Washington Lecture 3: Overview of Descriptive Statistics October 3, 2005 Lecture Outline Purpose
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH
 EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Results and Discussion of Receptor Tyrosine Kinase. Activation
 Results and Discussion of Receptor Tyrosine Kinase Activation To demonstrate the contribution which RCytoscape s molecular maps can make to biological understanding via exploratory data analysis, we here
Results and Discussion of Receptor Tyrosine Kinase Activation To demonstrate the contribution which RCytoscape s molecular maps can make to biological understanding via exploratory data analysis, we here
Colon cancer subtypes from gene expression data
 Colon cancer subtypes from gene expression data Nathan Cunningham Giuseppe Di Benedetto Sherman Ip Leon Law Module 6: Applied Statistics 26th February 2016 Aim Replicate findings of Felipe De Sousa et
Colon cancer subtypes from gene expression data Nathan Cunningham Giuseppe Di Benedetto Sherman Ip Leon Law Module 6: Applied Statistics 26th February 2016 Aim Replicate findings of Felipe De Sousa et
Supplementary Appendix
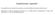 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Bredel M, Scholtens DM, Yadav AK, et al. NFKBIA deletion in
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Bredel M, Scholtens DM, Yadav AK, et al. NFKBIA deletion in
Six Sigma Glossary Lean 6 Society
 Six Sigma Glossary Lean 6 Society ABSCISSA ACCEPTANCE REGION ALPHA RISK ALTERNATIVE HYPOTHESIS ASSIGNABLE CAUSE ASSIGNABLE VARIATIONS The horizontal axis of a graph The region of values for which the null
Six Sigma Glossary Lean 6 Society ABSCISSA ACCEPTANCE REGION ALPHA RISK ALTERNATIVE HYPOTHESIS ASSIGNABLE CAUSE ASSIGNABLE VARIATIONS The horizontal axis of a graph The region of values for which the null
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE
 Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
List of Figures. List of Tables. Preface to the Second Edition. Preface to the First Edition
 List of Figures List of Tables Preface to the Second Edition Preface to the First Edition xv xxv xxix xxxi 1 What Is R? 1 1.1 Introduction to R................................ 1 1.2 Downloading and Installing
List of Figures List of Tables Preface to the Second Edition Preface to the First Edition xv xxv xxix xxxi 1 What Is R? 1 1.1 Introduction to R................................ 1 1.2 Downloading and Installing
Biomarker adaptive designs in clinical trials
 Review Article Biomarker adaptive designs in clinical trials James J. Chen 1, Tzu-Pin Lu 1,2, Dung-Tsa Chen 3, Sue-Jane Wang 4 1 Division of Bioinformatics and Biostatistics, National Center for Toxicological
Review Article Biomarker adaptive designs in clinical trials James J. Chen 1, Tzu-Pin Lu 1,2, Dung-Tsa Chen 3, Sue-Jane Wang 4 1 Division of Bioinformatics and Biostatistics, National Center for Toxicological
Interpreting Diverse Genomic Data Using Gene Sets
 Interpreting Diverse Genomic Data Using Gene Sets giovanni parmigiani@dfci.harvard.edu FHCRC February 2012 Gene Sets Leary 2008 PNAS Alterations in the combined FGF, EGFR, ERBB2 and PIK3 pathways. Red:
Interpreting Diverse Genomic Data Using Gene Sets giovanni parmigiani@dfci.harvard.edu FHCRC February 2012 Gene Sets Leary 2008 PNAS Alterations in the combined FGF, EGFR, ERBB2 and PIK3 pathways. Red:
A quick review. The clustering problem: Hierarchical clustering algorithm: Many possible distance metrics K-mean clustering algorithm:
 The clustering problem: partition genes into distinct sets with high homogeneity and high separation Hierarchical clustering algorithm: 1. Assign each object to a separate cluster. 2. Regroup the pair
The clustering problem: partition genes into distinct sets with high homogeneity and high separation Hierarchical clustering algorithm: 1. Assign each object to a separate cluster. 2. Regroup the pair
Unit 1 Exploring and Understanding Data
 Unit 1 Exploring and Understanding Data Area Principle Bar Chart Boxplot Conditional Distribution Dotplot Empirical Rule Five Number Summary Frequency Distribution Frequency Polygon Histogram Interquartile
Unit 1 Exploring and Understanding Data Area Principle Bar Chart Boxplot Conditional Distribution Dotplot Empirical Rule Five Number Summary Frequency Distribution Frequency Polygon Histogram Interquartile
The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis
 The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis Tieliu Shi tlshi@bio.ecnu.edu.cn The Center for bioinformatics
The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis Tieliu Shi tlshi@bio.ecnu.edu.cn The Center for bioinformatics
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies
 OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
Nature Neuroscience: doi: /nn Supplementary Figure 1. Behavioral training.
 Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
19(7), , 2016 doi: /neuonc/now270 Advance Access date 22 December 2016
 Neuro-Oncology 19(7), 997 1007, 2016 doi:10.1093/neuonc/now270 Advance Access date 22 December 2016 997 Magnetic resonance perfusion image features uncover an angiogenic subgroup of glioblastoma patients
Neuro-Oncology 19(7), 997 1007, 2016 doi:10.1093/neuonc/now270 Advance Access date 22 December 2016 997 Magnetic resonance perfusion image features uncover an angiogenic subgroup of glioblastoma patients
Supplementary Figure 1. Estimation of tumour content
 Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
T. R. Golub, D. K. Slonim & Others 1999
 T. R. Golub, D. K. Slonim & Others 1999 Big Picture in 1999 The Need for Cancer Classification Cancer classification very important for advances in cancer treatment. Cancers of Identical grade can have
T. R. Golub, D. K. Slonim & Others 1999 Big Picture in 1999 The Need for Cancer Classification Cancer classification very important for advances in cancer treatment. Cancers of Identical grade can have
RNA preparation from extracted paraffin cores:
 Supplementary methods, Nielsen et al., A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor positive breast cancer.
Supplementary methods, Nielsen et al., A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor positive breast cancer.
Supplementary Materials
 Supplementary Materials Supplementary figure 1. Taxonomic representation summarized at genus level. Fecal microbiota from a separate set of Jackson and Harlan mice prior to irradiation. A taxon was included
Supplementary Materials Supplementary figure 1. Taxonomic representation summarized at genus level. Fecal microbiota from a separate set of Jackson and Harlan mice prior to irradiation. A taxon was included
for the TCGA Breast Phenotype Research Group
 Decoding Breast Cancer with Quantitative Radiomics & Radiogenomics: Imaging Phenotypes in Breast Cancer Risk Assessment, Diagnosis, Prognosis, and Response to Therapy Maryellen Giger & Yuan Ji The University
Decoding Breast Cancer with Quantitative Radiomics & Radiogenomics: Imaging Phenotypes in Breast Cancer Risk Assessment, Diagnosis, Prognosis, and Response to Therapy Maryellen Giger & Yuan Ji The University
Nature Genetics: doi: /ng.2995
 Supplementary Figure 1 Kaplan-Meier survival curves of patients with brainstem tumors. (a) Comparison of patients with PPM1D mutation versus wild-type PPM1D. (b) Comparison of patients with PPM1D mutation
Supplementary Figure 1 Kaplan-Meier survival curves of patients with brainstem tumors. (a) Comparison of patients with PPM1D mutation versus wild-type PPM1D. (b) Comparison of patients with PPM1D mutation
Cancer Informatics Lecture
 Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
S1 Appendix: Figs A G and Table A. b Normal Generalized Fraction 0.075
 Aiello & Alter (216) PLoS One vol. 11 no. 1 e164546 S1 Appendix A-1 S1 Appendix: Figs A G and Table A a Tumor Generalized Fraction b Normal Generalized Fraction.25.5.75.25.5.75 1 53 4 59 2 58 8 57 3 48
Aiello & Alter (216) PLoS One vol. 11 no. 1 e164546 S1 Appendix A-1 S1 Appendix: Figs A G and Table A a Tumor Generalized Fraction b Normal Generalized Fraction.25.5.75.25.5.75 1 53 4 59 2 58 8 57 3 48
Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities
 Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities Gábor Boross,2, Katalin Orosz,2 and Illés J. Farkas 2, Department of Biological Physics,
Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities Gábor Boross,2, Katalin Orosz,2 and Illés J. Farkas 2, Department of Biological Physics,
PCxN: The pathway co-activity map: a resource for the unification of functional biology
 PCxN: The pathway co-activity map: a resource for the unification of functional biology Sheffield Institute for Translational Neurosciences Center for Integrative Genome Translation GENOME INFORMATICS
PCxN: The pathway co-activity map: a resource for the unification of functional biology Sheffield Institute for Translational Neurosciences Center for Integrative Genome Translation GENOME INFORMATICS
Interpreting Diverse Genomic Data Using Gene Sets
 Interpreting Diverse Genomic Data Using Gene Sets giovanni parmigiani@dfci.harvard.edu FHCRC February 2012 Gene Sets Leary 2008 PNAS Alterations in the combined FGF, EGFR, ERBB2 and PIK3 pathways. Red:
Interpreting Diverse Genomic Data Using Gene Sets giovanni parmigiani@dfci.harvard.edu FHCRC February 2012 Gene Sets Leary 2008 PNAS Alterations in the combined FGF, EGFR, ERBB2 and PIK3 pathways. Red:
Supplementary Figure 1
 Supplementary Figure 1 An example of the gene-term-disease network automatically generated by Phenolyzer web server for 'autism'. The largest word represents the user s input term, Autism. The pink round
Supplementary Figure 1 An example of the gene-term-disease network automatically generated by Phenolyzer web server for 'autism'. The largest word represents the user s input term, Autism. The pink round
Protocol Development: The Guiding Light of Any Clinical Study
 Protocol Development: The Guiding Light of Any Clinical Study Susan G. Fisher, Ph.D. Chair, Department of Clinical Sciences 1 Introduction Importance/ relevance/ gaps in knowledge Specific purpose of the
Protocol Development: The Guiding Light of Any Clinical Study Susan G. Fisher, Ph.D. Chair, Department of Clinical Sciences 1 Introduction Importance/ relevance/ gaps in knowledge Specific purpose of the
Learning Objectives 9/9/2013. Hypothesis Testing. Conflicts of Interest. Descriptive statistics: Numerical methods Measures of Central Tendency
 Conflicts of Interest I have no conflict of interest to disclose Biostatistics Kevin M. Sowinski, Pharm.D., FCCP Last-Chance Ambulatory Care Webinar Thursday, September 5, 2013 Learning Objectives For
Conflicts of Interest I have no conflict of interest to disclose Biostatistics Kevin M. Sowinski, Pharm.D., FCCP Last-Chance Ambulatory Care Webinar Thursday, September 5, 2013 Learning Objectives For
9/4/2013. Decision Errors. Hypothesis Testing. Conflicts of Interest. Descriptive statistics: Numerical methods Measures of Central Tendency
 Conflicts of Interest I have no conflict of interest to disclose Biostatistics Kevin M. Sowinski, Pharm.D., FCCP Pharmacotherapy Webinar Review Course Tuesday, September 3, 2013 Descriptive statistics:
Conflicts of Interest I have no conflict of interest to disclose Biostatistics Kevin M. Sowinski, Pharm.D., FCCP Pharmacotherapy Webinar Review Course Tuesday, September 3, 2013 Descriptive statistics:
Numerous hypothesis tests were performed in this study. To reduce the false positive due to
 Two alternative data-splitting Numerous hypothesis tests were performed in this study. To reduce the false positive due to multiple testing, we are not only seeking the results with extremely small p values
Two alternative data-splitting Numerous hypothesis tests were performed in this study. To reduce the false positive due to multiple testing, we are not only seeking the results with extremely small p values
Genetic alterations of histone lysine methyltransferases and their significance in breast cancer
 Genetic alterations of histone lysine methyltransferases and their significance in breast cancer Supplementary Materials and Methods Phylogenetic tree of the HMT superfamily The phylogeny outlined in the
Genetic alterations of histone lysine methyltransferases and their significance in breast cancer Supplementary Materials and Methods Phylogenetic tree of the HMT superfamily The phylogeny outlined in the
Generation of post-germinal centre myeloma plasma B cell.
 Generation of post-germinal centre myeloma. DNA DAMAGE CXCR4 Homing to Lytic lesion activation CD38 CD138 CD56 Phenotypic markers Naive Secondary lymphoid organ Multiple myeloma is a malignancy of s caused
Generation of post-germinal centre myeloma. DNA DAMAGE CXCR4 Homing to Lytic lesion activation CD38 CD138 CD56 Phenotypic markers Naive Secondary lymphoid organ Multiple myeloma is a malignancy of s caused
Brain Tumour Detection of MR Image Using Naïve Beyer classifier and Support Vector Machine
 International Journal of Scientific Research in Computer Science, Engineering and Information Technology 2018 IJSRCSEIT Volume 3 Issue 3 ISSN : 2456-3307 Brain Tumour Detection of MR Image Using Naïve
International Journal of Scientific Research in Computer Science, Engineering and Information Technology 2018 IJSRCSEIT Volume 3 Issue 3 ISSN : 2456-3307 Brain Tumour Detection of MR Image Using Naïve
Supplementary Methods
 Supplementary Methods Analysis of time course gene expression data. The time course data of the expression level of a representative gene is shown in the below figure. The trajectory of longitudinal expression
Supplementary Methods Analysis of time course gene expression data. The time course data of the expression level of a representative gene is shown in the below figure. The trajectory of longitudinal expression
Introduction to Gene Sets Analysis
 Introduction to Svitlana Tyekucheva Dana-Farber Cancer Institute May 15, 2012 Introduction Various measurements: gene expression, copy number variation, methylation status, mutation profile, etc. Main
Introduction to Svitlana Tyekucheva Dana-Farber Cancer Institute May 15, 2012 Introduction Various measurements: gene expression, copy number variation, methylation status, mutation profile, etc. Main
Predicting Non-Small Cell Lung Cancer Diagnosis and Prognosis by Fully Automated Microscopic Pathology Image Features
 Predicting Non-Small Cell Lung Cancer Diagnosis and Prognosis by Fully Automated Microscopic Pathology Image Features Kun-Hsing Yu, MD, PhD Department of Biomedical Informatics, Harvard Medical School
Predicting Non-Small Cell Lung Cancer Diagnosis and Prognosis by Fully Automated Microscopic Pathology Image Features Kun-Hsing Yu, MD, PhD Department of Biomedical Informatics, Harvard Medical School
Introduction to statistics Dr Alvin Vista, ACER Bangkok, 14-18, Sept. 2015
 Analysing and Understanding Learning Assessment for Evidence-based Policy Making Introduction to statistics Dr Alvin Vista, ACER Bangkok, 14-18, Sept. 2015 Australian Council for Educational Research Structure
Analysing and Understanding Learning Assessment for Evidence-based Policy Making Introduction to statistics Dr Alvin Vista, ACER Bangkok, 14-18, Sept. 2015 Australian Council for Educational Research Structure
ASMS 2015 ThP 459 Glioblastoma Multiforme Subtype Classification: Integrated Analysis of Protein and Gene Expression Data
 ASMS 2015 ThP 459 Glioblastoma Multiforme Subtype Classification: Integrated Analysis of Protein and Gene Expression Data Durairaj Renu 1, Vadiraja Bhat 2, Mona Al-Gizawiy 3, Carolina B. Livi 2, Stephen
ASMS 2015 ThP 459 Glioblastoma Multiforme Subtype Classification: Integrated Analysis of Protein and Gene Expression Data Durairaj Renu 1, Vadiraja Bhat 2, Mona Al-Gizawiy 3, Carolina B. Livi 2, Stephen
Supplemental Information. Molecular, Pathological, Radiological, and Immune. Profiling of Non-brainstem Pediatric High-Grade
 Cancer Cell, Volume 33 Supplemental Information Molecular, Pathological, Radiological, and Immune Profiling of Non-brainstem Pediatric High-Grade Glioma from the HERBY Phase II Randomized Trial Alan Mackay,
Cancer Cell, Volume 33 Supplemental Information Molecular, Pathological, Radiological, and Immune Profiling of Non-brainstem Pediatric High-Grade Glioma from the HERBY Phase II Randomized Trial Alan Mackay,
Cell Signaling (part 1)
 15 Cell Signaling (part 1) Introduction Bacteria and unicellular eukaryotes respond to environmental signals and to signaling molecules secreted by other cells for mating and other communication. In multicellular
15 Cell Signaling (part 1) Introduction Bacteria and unicellular eukaryotes respond to environmental signals and to signaling molecules secreted by other cells for mating and other communication. In multicellular
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded
 Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
CHAPTER 9 SUMMARY AND CONCLUSION
 CHAPTER 9 SUMMARY AND CONCLUSION 9.1 SUMMARY In this thesis, the CAD system for early detection and classification of ischemic stroke in CT image, hemorrhage and hematoma in brain CT image and brain tumor
CHAPTER 9 SUMMARY AND CONCLUSION 9.1 SUMMARY In this thesis, the CAD system for early detection and classification of ischemic stroke in CT image, hemorrhage and hematoma in brain CT image and brain tumor
Gene expression profiling predicts clinical outcome of prostate cancer. Gennadi V. Glinsky, Anna B. Glinskii, Andrew J. Stephenson, Robert M.
 SUPPLEMENTARY DATA Gene expression profiling predicts clinical outcome of prostate cancer Gennadi V. Glinsky, Anna B. Glinskii, Andrew J. Stephenson, Robert M. Hoffman, William L. Gerald Table of Contents
SUPPLEMENTARY DATA Gene expression profiling predicts clinical outcome of prostate cancer Gennadi V. Glinsky, Anna B. Glinskii, Andrew J. Stephenson, Robert M. Hoffman, William L. Gerald Table of Contents
chapter 1 - fig. 2 Mechanism of transcriptional control by ppar agonists.
 chapter 1 - fig. 1 The -omics subdisciplines. chapter 1 - fig. 2 Mechanism of transcriptional control by ppar agonists. 201 figures chapter 1 chapter 2 - fig. 1 Schematic overview of the different steps
chapter 1 - fig. 1 The -omics subdisciplines. chapter 1 - fig. 2 Mechanism of transcriptional control by ppar agonists. 201 figures chapter 1 chapter 2 - fig. 1 Schematic overview of the different steps
Oncogenes. Dr. S Hosseini-Asl
 Oncogenes Dr. S Hosseini-Asl An oncogene is a mutated form of a normal cellular gene called a proto-oncogene that contributes to the development of a cancer. Proto-oncogenes typically regulate cell growth
Oncogenes Dr. S Hosseini-Asl An oncogene is a mutated form of a normal cellular gene called a proto-oncogene that contributes to the development of a cancer. Proto-oncogenes typically regulate cell growth
Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the
 Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the deep dermis. (b) The melanocytes demonstrate abundant
Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the deep dermis. (b) The melanocytes demonstrate abundant
DeSigN: connecting gene expression with therapeutics for drug repurposing and development. Bernard lee GIW 2016, Shanghai 8 October 2016
 DeSigN: connecting gene expression with therapeutics for drug repurposing and development Bernard lee GIW 2016, Shanghai 8 October 2016 1 Motivation Average cost: USD 1.8 to 2.6 billion ~2% Attrition rate
DeSigN: connecting gene expression with therapeutics for drug repurposing and development Bernard lee GIW 2016, Shanghai 8 October 2016 1 Motivation Average cost: USD 1.8 to 2.6 billion ~2% Attrition rate
Supplementary Information
 Supplementary Information Guided Visual Exploration of Genomic Stratifications in Cancer Marc Streit 1,6, Alexander Lex 2,6, Samuel Gratzl¹, Christian Partl³, Dieter Schmalstieg³, Hanspeter Pfister², Peter
Supplementary Information Guided Visual Exploration of Genomic Stratifications in Cancer Marc Streit 1,6, Alexander Lex 2,6, Samuel Gratzl¹, Christian Partl³, Dieter Schmalstieg³, Hanspeter Pfister², Peter
# of transcr ipts FDR q- value transcripts PGF,CTNNB1,AKT2,FARP2,VEGFB,PDGFRA,DRD 2,CSF1,CDH1,ITGB1,SRC,KRAS,RASGRP3,MAP2. Pathway
 Supplementary Table 9. Pathways and biological processes associated with transcripts specific to SEB in both the pre- and post- RASP leukocytes (transcripts included in the Reactome Function Interaction
Supplementary Table 9. Pathways and biological processes associated with transcripts specific to SEB in both the pre- and post- RASP leukocytes (transcripts included in the Reactome Function Interaction
Clustered mutations of oncogenes and tumor suppressors.
 Supplementary Figure 1 Clustered mutations of oncogenes and tumor suppressors. For each oncogene (red dots) and tumor suppressor (blue dots), the number of mutations found in an intramolecular cluster
Supplementary Figure 1 Clustered mutations of oncogenes and tumor suppressors. For each oncogene (red dots) and tumor suppressor (blue dots), the number of mutations found in an intramolecular cluster
Sudin Bhattacharya Institute for Integrative Toxicology
 Beyond the AHRE: the Role of Epigenomics in Gene Regulation by the AHR (or, Varied Applications of Computational Modeling in Toxicology and Ingredient Safety) Sudin Bhattacharya Institute for Integrative
Beyond the AHRE: the Role of Epigenomics in Gene Regulation by the AHR (or, Varied Applications of Computational Modeling in Toxicology and Ingredient Safety) Sudin Bhattacharya Institute for Integrative
Principles of Genetics and Molecular Biology
 Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
Physicochemical Properties Can Be Key Determinants of Mesoporous Silica Nanoparticle Potency In Vitro
 1 Physicochemical Properties Can Be Key Determinants of Mesoporous Silica Nanoparticle Potency In Vitro Dalibor Breznan 1*, Dharani D. Das 1, Christine MacKinnon-Roy 1, Stéphane Bernatchez 2, Abdelhamid
1 Physicochemical Properties Can Be Key Determinants of Mesoporous Silica Nanoparticle Potency In Vitro Dalibor Breznan 1*, Dharani D. Das 1, Christine MacKinnon-Roy 1, Stéphane Bernatchez 2, Abdelhamid
POST GRADUATE DIPLOMA IN BIOETHICS (PGDBE) Term-End Examination June, 2016 MHS-014 : RESEARCH METHODOLOGY
 No. of Printed Pages : 12 MHS-014 POST GRADUATE DIPLOMA IN BIOETHICS (PGDBE) Term-End Examination June, 2016 MHS-014 : RESEARCH METHODOLOGY Time : 2 hours Maximum Marks : 70 PART A Attempt all questions.
No. of Printed Pages : 12 MHS-014 POST GRADUATE DIPLOMA IN BIOETHICS (PGDBE) Term-End Examination June, 2016 MHS-014 : RESEARCH METHODOLOGY Time : 2 hours Maximum Marks : 70 PART A Attempt all questions.
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
Basic Biostatistics. Chapter 1. Content
 Chapter 1 Basic Biostatistics Jamalludin Ab Rahman MD MPH Department of Community Medicine Kulliyyah of Medicine Content 2 Basic premises variables, level of measurements, probability distribution Descriptive
Chapter 1 Basic Biostatistics Jamalludin Ab Rahman MD MPH Department of Community Medicine Kulliyyah of Medicine Content 2 Basic premises variables, level of measurements, probability distribution Descriptive
Plasma-Seq conducted with blood from male individuals without cancer.
 Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
White Paper Estimating Complex Phenotype Prevalence Using Predictive Models
 White Paper 23-12 Estimating Complex Phenotype Prevalence Using Predictive Models Authors: Nicholas A. Furlotte Aaron Kleinman Robin Smith David Hinds Created: September 25 th, 2015 September 25th, 2015
White Paper 23-12 Estimating Complex Phenotype Prevalence Using Predictive Models Authors: Nicholas A. Furlotte Aaron Kleinman Robin Smith David Hinds Created: September 25 th, 2015 September 25th, 2015
Quantitative Data and Measurement. POLI 205 Doing Research in Politics. Fall 2015
 Quantitative Fall 2015 Theory and We need to test our theories with empirical data Inference : Systematic observation and representation of concepts Quantitative: measures are numeric Qualitative: measures
Quantitative Fall 2015 Theory and We need to test our theories with empirical data Inference : Systematic observation and representation of concepts Quantitative: measures are numeric Qualitative: measures
ChIP-seq data analysis
 ChIP-seq data analysis Harri Lähdesmäki Department of Computer Science Aalto University November 24, 2017 Contents Background ChIP-seq protocol ChIP-seq data analysis Transcriptional regulation Transcriptional
ChIP-seq data analysis Harri Lähdesmäki Department of Computer Science Aalto University November 24, 2017 Contents Background ChIP-seq protocol ChIP-seq data analysis Transcriptional regulation Transcriptional
Quantitative Radiomics System Decoding the Tumor Phenotype. John Quackenbush and Hugo Aerts
 Quantitative Radiomics System Decoding the Tumor Phenotype John Quackenbush and Hugo Aerts The Radiomics Hypothesis The tumor s structural phenotype reflects its molecular and clinical properties. This
Quantitative Radiomics System Decoding the Tumor Phenotype John Quackenbush and Hugo Aerts The Radiomics Hypothesis The tumor s structural phenotype reflects its molecular and clinical properties. This
SUPPLEMENTARY APPENDIX
 SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
Measuring Focused Attention Using Fixation Inner-Density
 Measuring Focused Attention Using Fixation Inner-Density Wen Liu, Mina Shojaeizadeh, Soussan Djamasbi, Andrew C. Trapp User Experience & Decision Making Research Laboratory, Worcester Polytechnic Institute
Measuring Focused Attention Using Fixation Inner-Density Wen Liu, Mina Shojaeizadeh, Soussan Djamasbi, Andrew C. Trapp User Experience & Decision Making Research Laboratory, Worcester Polytechnic Institute
EMOTION CLASSIFICATION: HOW DOES AN AUTOMATED SYSTEM COMPARE TO NAÏVE HUMAN CODERS?
 EMOTION CLASSIFICATION: HOW DOES AN AUTOMATED SYSTEM COMPARE TO NAÏVE HUMAN CODERS? Sefik Emre Eskimez, Kenneth Imade, Na Yang, Melissa Sturge- Apple, Zhiyao Duan, Wendi Heinzelman University of Rochester,
EMOTION CLASSIFICATION: HOW DOES AN AUTOMATED SYSTEM COMPARE TO NAÏVE HUMAN CODERS? Sefik Emre Eskimez, Kenneth Imade, Na Yang, Melissa Sturge- Apple, Zhiyao Duan, Wendi Heinzelman University of Rochester,
Deep Learning Analytics for Predicting Prognosis of Acute Myeloid Leukemia with Cytogenetics, Age, and Mutations
 Deep Learning Analytics for Predicting Prognosis of Acute Myeloid Leukemia with Cytogenetics, Age, and Mutations Andy Nguyen, M.D., M.S. Medical Director, Hematopathology, Hematology and Coagulation Laboratory,
Deep Learning Analytics for Predicting Prognosis of Acute Myeloid Leukemia with Cytogenetics, Age, and Mutations Andy Nguyen, M.D., M.S. Medical Director, Hematopathology, Hematology and Coagulation Laboratory,
MethylMix An R package for identifying DNA methylation driven genes
 MethylMix An R package for identifying DNA methylation driven genes Olivier Gevaert May 3, 2016 Stanford Center for Biomedical Informatics Department of Medicine 1265 Welch Road Stanford CA, 94305-5479
MethylMix An R package for identifying DNA methylation driven genes Olivier Gevaert May 3, 2016 Stanford Center for Biomedical Informatics Department of Medicine 1265 Welch Road Stanford CA, 94305-5479
Supplementary figures
 Supplementary figures Supplementary figure 1: Pathway enrichment for each comparison. The first column shows enrichment with differentially expressed genes between the diet groups at t = 0, the second
Supplementary figures Supplementary figure 1: Pathway enrichment for each comparison. The first column shows enrichment with differentially expressed genes between the diet groups at t = 0, the second
T-cell activation T cells migrate to secondary lymphoid tissues where they interact with antigen, antigen-presenting cells, and other lymphocytes:
 Interactions between innate immunity & adaptive immunity What happens to T cells after they leave the thymus? Naïve T cells exit the thymus and enter the bloodstream. If they remain in the bloodstream,
Interactions between innate immunity & adaptive immunity What happens to T cells after they leave the thymus? Naïve T cells exit the thymus and enter the bloodstream. If they remain in the bloodstream,
T-cell activation T cells migrate to secondary lymphoid tissues where they interact with antigen, antigen-presenting cells, and other lymphocytes:
 Interactions between innate immunity & adaptive immunity What happens to T cells after they leave the thymus? Naïve T cells exit the thymus and enter the bloodstream. If they remain in the bloodstream,
Interactions between innate immunity & adaptive immunity What happens to T cells after they leave the thymus? Naïve T cells exit the thymus and enter the bloodstream. If they remain in the bloodstream,
Analysis of EWS Test Data for Reliability Improvement through Outlier Detection and Automated Ink Map Generation
 Analysis of EWS Test Data for Reliability Improvement through Outlier Detection and Automated Ink Map Generation Michael Scott Test Advantage Inc. June 8, 2005 Toward ZERO DEFECTS There is an established
Analysis of EWS Test Data for Reliability Improvement through Outlier Detection and Automated Ink Map Generation Michael Scott Test Advantage Inc. June 8, 2005 Toward ZERO DEFECTS There is an established
microrna PCR System (Exiqon), following the manufacturer s instructions. In brief, 10ng of
 SUPPLEMENTAL MATERIALS AND METHODS Quantitative RT-PCR Quantitative RT-PCR analysis was performed using the Universal mircury LNA TM microrna PCR System (Exiqon), following the manufacturer s instructions.
SUPPLEMENTAL MATERIALS AND METHODS Quantitative RT-PCR Quantitative RT-PCR analysis was performed using the Universal mircury LNA TM microrna PCR System (Exiqon), following the manufacturer s instructions.
Classification of Benign and Malignant Vertebral Compression Fractures in Magnetic Resonance Images
 Classification of Benign and Malignant Vertebral Compression Fractures in Magnetic Resonance Images Lucas Frighetto-Pereira, Guilherme Augusto Metzner, Paulo Mazzoncini de Azevedo-Marques, Rangaraj Mandayam
Classification of Benign and Malignant Vertebral Compression Fractures in Magnetic Resonance Images Lucas Frighetto-Pereira, Guilherme Augusto Metzner, Paulo Mazzoncini de Azevedo-Marques, Rangaraj Mandayam
Using Bayesian Networks to Analyze Expression Data. Xu Siwei, s Muhammad Ali Faisal, s Tejal Joshi, s
 Using Bayesian Networks to Analyze Expression Data Xu Siwei, s0789023 Muhammad Ali Faisal, s0677834 Tejal Joshi, s0677858 Outline Introduction Bayesian Networks Equivalence Classes Applying to Expression
Using Bayesian Networks to Analyze Expression Data Xu Siwei, s0789023 Muhammad Ali Faisal, s0677834 Tejal Joshi, s0677858 Outline Introduction Bayesian Networks Equivalence Classes Applying to Expression
mir-509-5p and mir-1243 increase the sensitivity to gemcitabine by inhibiting
 mir-509-5p and mir-1243 increase the sensitivity to gemcitabine by inhibiting epithelial-mesenchymal transition in pancreatic cancer Hidekazu Hiramoto, M.D. 1,3, Tomoki Muramatsu, Ph.D. 1, Daisuke Ichikawa,
mir-509-5p and mir-1243 increase the sensitivity to gemcitabine by inhibiting epithelial-mesenchymal transition in pancreatic cancer Hidekazu Hiramoto, M.D. 1,3, Tomoki Muramatsu, Ph.D. 1, Daisuke Ichikawa,
