Supplemental Methods RNA sequencing experiment
|
|
|
- Thomasine Newman
- 5 years ago
- Views:
Transcription
1 Supplemental Methods RNA sequencing experiment Mice were euthanized as described in the Methods and the right lung was removed, placed in a sterile eppendorf tube, and snap frozen in liquid nitrogen. RNA extraction was performed on each sample by TRIZOL extraction (Life Technologies) following by Qiagen RNEasy Mini kit purification with on-column DNAse digestion. The Vanderbilt Technologies for Advanced Genomics Core Facility (VANTAGE, Nashville, TN) used the Illumina TruSeq Stranded mrna Sample Preparation Kit to convert the mrna in 1 ng of total RNA into a library of template molecules suitable for subsequent cluster generation and sequencing on the Illumina HiSeq 5. The first step was to do a quality check of the input total RNA by running an aliquot on the Agilent Bioanalyzer to confirm integrity. The Qubit RNA fluorometry assay was used to measure concentration. The input to library prep was 5 µl of 1 ng of total RNA ( ng/µl). The total RNA underwent enrichment of the poly- A containing mrna molecules using poly- T oligo- attached magnetic beads. Following enrichment, the eluted poly(a) RNA was cleaved into small fragments of 1-1 bp using divalent cations under elevated temperature. The cleaved RNA fragments were copied into first strand cdna using SuperScript II reverse transcriptase and random hexamer primers. This was followed by second strand cdna synthesis using DNA Polymerase I and RNase H. The cdna fragments then went through an end repair process, the addition of a single A base, and then ligation of the Illumina multiplexing adapters. The products were then purified and enriched with PCR to create the final cdna sequencing library. The cdna library underwent quality control on the Agilent Bioanalyzer HS DNA assay to confirm the final library size and on the Agilent Mx35P qpcr machine using the KAPA Illumina library quantification kit to determine concentration. A nm stock was created and samples pooled by molarity for multiplexing. From the pool 1 pm was loaded into each well for the flow cell on the Illumina cbot for cluster generation. The flow cell was then loaded onto the Illumina HiSeq 5 for Paired-end 5 bp sequencing utilizing v3 chemistry and HTA 1.8. The raw sequencing reads in BCL format are processed through CASAVA-1.8. for FASTQ conversion and demultiplexing. The RTA chastity filter was used and only the PF (pass filter) reads are retained for further analysis. Quality control (QC) assessment was performed on raw data using Fastx Toolkit (1) and FastQC. RNA data was aligned to mouse genome build 1 (mm1) by using TopHat..7 () followed by Cufflinks (3,). Gene-level expression measurements were normalized to reads per kilobase per million reads (RPKM) by Scripture and in fragments per kilobase per million reads (FPKM) by Cufflinks. Cuffdiff from Cufflinks package was used to detect differentially expressed genes for all group comparisons, which was based on the test statistics T = E[log(y)]/Var[log(y)], where y is the RPKM/FPKM value, and the ratio approximately follows a normal distribution. Differential expression was considered significant if both of the following criteria were met: (1) false discovery rate (FDR) corrected P <.1 (5), and () log (fold change) > Fastx Toolkit. Available from: Trapnell C, Pachter L, Salzberg SL. 9. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 5: Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MG, Salzberg SL, Wold BJ, Pachter L. 1. Transcript assembly and quantification by
2 RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 8: Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R, 1 Genome Project Data Processing Subgroup. 9. The Sequence Alignment/Map format and SAMtools. Bioinformatics 5: Benjamini Y, and Hochberg Y Controlling the false discovery rate: A practical and powerful approach to multiple testing. Journal of the Royal Statistical Society. Series B (Methodological) 57: 89 3.
3 A Total Airway Cells B Airway Lymphocytes 1 cells/ml cells/ml day 1 day day 1 day C Airway Macrophages D Blood Neutrophils 1 cells/ml Peripheral Blood Neutrophils per µl 3 1 day 1 day day Figure S1. Total airway cells (A), airway lymphocytes (B), and airway macrophages (C) on day 1 and day post-kp infection in mice that underwent the sensitization, challenge, and KP infection protocol described in Figure 1. (D) Absolute peripheral blood neutrophil count from and mice on day post-infection. n=1 mice per group combined from two representative experiments. p<.1, p<.1, p<.5.
4 log 1 cfu/ml Lung KP Burden Right Lung Left Lung 1 hour post-infection Figure S. Pre-existing allergic airway inflammation does not alter lung KP burden 1 hour post-infection. Left and right lungs from KP-infected mice were harvested at 1 hour post-infection and quantitative bacterial culture was performed. Lower limit of detection was 1 cfu/ml (log 1 cfu/ml = ). For KP-infected groups, values below the limit of detection were assigned half the lower limit of detection (5 cfu/ml; log 1 cfu/ml = 1.67).
5 H&E 1X H&E X Lung bacteria (-) Sirius Red 1X 3 Lung neutrophils (-) 3 Lung granulocytes (-) Lung eosinophils (-) D α-neutrophil X A B C E 1 1
6 Figure S3. Allergic airway inflammation decreases lung neutrophils and bacteria following acute KP infection. Day post-infection lung sections were obtained. (A) H&E X. (B) H&E 1X, arrow indicates bacteria. (C) Sirius Red 1X demonstrating eosinophils (arrows) and neutrophils (arrowheads). (D) α-neutrophil elastase. (E) Lung granulocyte, bacteria, eosinophil, and neutrophil scores. p<.1, p<.1, p<.5., n=5;, n=6;, n=7;, n=1. Combined from two representative experiments.
7 1 Lung KP Burden 8 log 1 cfu/ml 6 No aerosol PBS aerosol OVA aerosol day Figure S. Aerosol exposure to PBS or OVA does not decrease lung KP burden. WT mice were exposed to PBS aerosol or 1% OVA (in PBS) aerosol for consecutive days for minutes per day or were not aerosol exposed. Twenty-four hours following the final aerosol exposure, mice were infected with KP and harvested on day post-infection. Left lung was removed and quantitative bacterial culture was performed. n=9-1 mice per group, combined results from two representative experiments.
8 1 cells/ml A 3 1 Airway Eosinophils pg/ml B 5 3 day 1 day IL-5 1 pg/ml C 8 6 day 1 day IL-13 day 1 day Figure S5. Acute KP lung infection alters post-infection kinetics of allergic airway inflammation. (A) Total airway eosinophils. Lung IL-5 expression (B) and lung IL-13 expression (C) were quantified by cytokine bead array. p<.1, p<.5. n=1 mice per group combined from two representative experiments.
9 A WT- WT- B Mucus Score (-3) 3 1 Airway Mucus WT- WT- WT- WT- WT- WT- day IL-13KO- IL-13KO- IL-13KO- IL-13KO- Figure S6. Decreased airway mucus in the absence of IL-13. (A) Representative PAS stained lung sections from WT and IL-13KO mice that underwent OVA sensitization and challenge or mock sensitization followed by PBS or KP infection. Lungs were harvested on day post-infection. (B) Airway mucus score based on PAS positivity. n=-3 mice per group. p<.1, p<.5.
10 A hour 6 hour WT- ALUM WT- OVA WT- ALUM WT- OVA WT- ALUM WT- OVA STAT6KO- ALUM STAT6KO- OVA WT- ALUM WT- OVA STAT6KO- ALUM STAT6KO- OVA
11 B WT hour STAT6KO WT 6 hour STAT6KO Transcripts Increased in OVA vs. ALUM Transcripts Decreased in OVA vs. ALUM Figure S7. (A) Heatmap showing normalized log-transformed intensities for transcripts with significant different expression levels between ALUM and OVA experimental groups. Significant different expression was defined as q.1 and a log-fold expression difference 1.5. The locus ID is shown when a gene name has not been assigned. n= mice per group for hour timepoint and n=-5 mice per group for the 6 hour timepoint. (B) Venn diagrams demonstrating the number of transcripts for which expression was significantly different in OVA mice compared to ALUM mice. Areas of overlap denote the number of transcripts for which OVA mice had significantly different expression compared to ALUM mice for both WT and STAT6KO genotypes.
12 hours (pre- infection) Transcripts Increased in OVA vs. ALUM lungs in both WT and STAT6KO mice Transcript Name RPKM WT ALUM RPKM WT OVA Log fold change q value RPKM STAT6KO- ALUM RPKM STAT6KO- OVA Log fold change q value Slc6a Ccl Mmp Cxcl E- 7 Cxcl E E- 1 Saa Ccl E- 5 Cxcl E Reg3g E- 13 Bpifb E- 9 Birc E Rgs E Ctsk E- 5 Pigr E- 8 Kifc E Cxcl Ubec Gm E Mki E- 5 Lcn Slfn E Kif E Cenpe Tpx Iqgap Ckap E Ckapl E Adamts Mcm E E- 6 Nusap E
13 hours (pre- infection) Transcripts Decreased in OVA vs. ALUM in both WT and STAT6KO lungs Zbtb E- 5 6 hours Transcripts Increased in OVA vs. ALUM in both WT and STAT6KO lungs Transcript Name RPKM WT ALUM RPKM WT OVA Log fold change q value RPKM STAT6KO- ALUM RPKM STAT6KO- OVA Log fold change q value Ccl Retnla Agr Tff E- 6 Kntc Reg3g Shcbp E Birc E Ckapl E Ndc E Dtl Tpx Rrm E E- 5 Kif18b Ccno Cxcl E- 5 Ubec E Gp AC E31N8Rik hours Transcripts Decreased in OVA vs. ALUM in both WT and STAT6KO lungs None Table S1: Lung transcripts with significantly different expression between OVA and ALUM mice for both WT and STAT6KO genotypes. Transcript name, reads per kilobase per million reads (RPKM), logfold expression change between OVA and ALUM groups, and q value are shown.
Supplementary Online Content
 Supplementary Online Content Fumagalli D, Venet D, Ignatiadis M, et al. RNA Sequencing to predict response to neoadjuvant anti-her2 therapy: a secondary analysis of the NeoALTTO randomized clinical trial.
Supplementary Online Content Fumagalli D, Venet D, Ignatiadis M, et al. RNA Sequencing to predict response to neoadjuvant anti-her2 therapy: a secondary analysis of the NeoALTTO randomized clinical trial.
Selective depletion of abundant RNAs to enable transcriptome analysis of lowinput and highly-degraded RNA from FFPE breast cancer samples
 DNA CLONING DNA AMPLIFICATION & PCR EPIGENETICS RNA ANALYSIS Selective depletion of abundant RNAs to enable transcriptome analysis of lowinput and highly-degraded RNA from FFPE breast cancer samples LIBRARY
DNA CLONING DNA AMPLIFICATION & PCR EPIGENETICS RNA ANALYSIS Selective depletion of abundant RNAs to enable transcriptome analysis of lowinput and highly-degraded RNA from FFPE breast cancer samples LIBRARY
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
genomics for systems biology / ISB2020 RNA sequencing (RNA-seq)
 RNA sequencing (RNA-seq) Module Outline MO 13-Mar-2017 RNA sequencing: Introduction 1 WE 15-Mar-2017 RNA sequencing: Introduction 2 MO 20-Mar-2017 Paper: PMID 25954002: Human genomics. The human transcriptome
RNA sequencing (RNA-seq) Module Outline MO 13-Mar-2017 RNA sequencing: Introduction 1 WE 15-Mar-2017 RNA sequencing: Introduction 2 MO 20-Mar-2017 Paper: PMID 25954002: Human genomics. The human transcriptome
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Material for IPred - Integrating Ab Initio and Evidence Based Gene Predictions to Improve Prediction Accuracy
 1 SYSTEM REQUIREMENTS 1 Supplementary Material for IPred - Integrating Ab Initio and Evidence Based Gene Predictions to Improve Prediction Accuracy Franziska Zickmann and Bernhard Y. Renard Research Group
1 SYSTEM REQUIREMENTS 1 Supplementary Material for IPred - Integrating Ab Initio and Evidence Based Gene Predictions to Improve Prediction Accuracy Franziska Zickmann and Bernhard Y. Renard Research Group
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
Supplemental Information. Regulatory T Cells Exhibit Distinct. Features in Human Breast Cancer
 Immunity, Volume 45 Supplemental Information Regulatory T Cells Exhibit Distinct Features in Human Breast Cancer George Plitas, Catherine Konopacki, Kenmin Wu, Paula D. Bos, Monica Morrow, Ekaterina V.
Immunity, Volume 45 Supplemental Information Regulatory T Cells Exhibit Distinct Features in Human Breast Cancer George Plitas, Catherine Konopacki, Kenmin Wu, Paula D. Bos, Monica Morrow, Ekaterina V.
Transcriptome Analysis
 Transcriptome Analysis Data Preprocessing Sample Preparation Illumina Sequencing Demultiplexing Raw FastQ Reference Genome (fasta) Reference Annotation (GTF) Reference Genome Analysis Tophat Accepted hits
Transcriptome Analysis Data Preprocessing Sample Preparation Illumina Sequencing Demultiplexing Raw FastQ Reference Genome (fasta) Reference Annotation (GTF) Reference Genome Analysis Tophat Accepted hits
RNA SEQUENCING AND DATA ANALYSIS
 RNA SEQUENCING AND DATA ANALYSIS Length of mrna transcripts in the human genome 5,000 5,000 4,000 3,000 2,000 4,000 1,000 0 0 200 400 600 800 3,000 2,000 1,000 0 0 2,000 4,000 6,000 8,000 10,000 Length
RNA SEQUENCING AND DATA ANALYSIS Length of mrna transcripts in the human genome 5,000 5,000 4,000 3,000 2,000 4,000 1,000 0 0 200 400 600 800 3,000 2,000 1,000 0 0 2,000 4,000 6,000 8,000 10,000 Length
RNA-Seq Preparation Comparision Summary: Lexogen, Standard, NEB
 RNA-Seq Preparation Comparision Summary: Lexogen, Standard, NEB CSF-NGS January 22, 214 Contents 1 Introduction 1 2 Experimental Details 1 3 Results And Discussion 1 3.1 ERCC spike ins............................................
RNA-Seq Preparation Comparision Summary: Lexogen, Standard, NEB CSF-NGS January 22, 214 Contents 1 Introduction 1 2 Experimental Details 1 3 Results And Discussion 1 3.1 ERCC spike ins............................................
A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis
 APPLICATION NOTE Cell-Free DNA Isolation Kit A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis Abstract Circulating cell-free DNA (cfdna) has been shown
APPLICATION NOTE Cell-Free DNA Isolation Kit A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis Abstract Circulating cell-free DNA (cfdna) has been shown
National Disease Research Interchange Annual Progress Report: 2010 Formula Grant
 National Disease Research Interchange Annual Progress Report: 2010 Formula Grant Reporting Period July 1, 2012 June 30, 2013 Formula Grant Overview The National Disease Research Interchange received $62,393
National Disease Research Interchange Annual Progress Report: 2010 Formula Grant Reporting Period July 1, 2012 June 30, 2013 Formula Grant Overview The National Disease Research Interchange received $62,393
fl/+ KRas;Atg5 fl/+ KRas;Atg5 fl/fl KRas;Atg5 fl/fl KRas;Atg5 Supplementary Figure 1. Gene set enrichment analyses. (a) (b)
 KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
Ambient temperature regulated flowering time
 Ambient temperature regulated flowering time Applications of RNAseq RNA- seq course: The power of RNA-seq June 7 th, 2013; Richard Immink Overview Introduction: Biological research question/hypothesis
Ambient temperature regulated flowering time Applications of RNAseq RNA- seq course: The power of RNA-seq June 7 th, 2013; Richard Immink Overview Introduction: Biological research question/hypothesis
KAPA Stranded RNA-Seq Library Preparation Kit
 KAPA Stranded RNA-Seq Illumina platforms KR0934 v1.13 This provides product information and a detailed protocol for the KAPA Stranded RNA- Seq (Illumina platforms), product codes KK8400 and KK8401. Contents
KAPA Stranded RNA-Seq Illumina platforms KR0934 v1.13 This provides product information and a detailed protocol for the KAPA Stranded RNA- Seq (Illumina platforms), product codes KK8400 and KK8401. Contents
Iso-Seq Method Updates and Target Enrichment Without Amplification for SMRT Sequencing
 Iso-Seq Method Updates and Target Enrichment Without Amplification for SMRT Sequencing PacBio Americas User Group Meeting Sample Prep Workshop June.27.2017 Tyson Clark, Ph.D. For Research Use Only. Not
Iso-Seq Method Updates and Target Enrichment Without Amplification for SMRT Sequencing PacBio Americas User Group Meeting Sample Prep Workshop June.27.2017 Tyson Clark, Ph.D. For Research Use Only. Not
GeneOverlap: An R package to test and visualize
 GeneOverlap: An R package to test and visualize gene overlaps Li Shen Contact: li.shen@mssm.edu or shenli.sam@gmail.com Icahn School of Medicine at Mount Sinai New York, New York http://shenlab-sinai.github.io/shenlab-sinai/
GeneOverlap: An R package to test and visualize gene overlaps Li Shen Contact: li.shen@mssm.edu or shenli.sam@gmail.com Icahn School of Medicine at Mount Sinai New York, New York http://shenlab-sinai.github.io/shenlab-sinai/
Inference of Isoforms from Short Sequence Reads
 Inference of Isoforms from Short Sequence Reads Tao Jiang Department of Computer Science and Engineering University of California, Riverside Tsinghua University Joint work with Jianxing Feng and Wei Li
Inference of Isoforms from Short Sequence Reads Tao Jiang Department of Computer Science and Engineering University of California, Riverside Tsinghua University Joint work with Jianxing Feng and Wei Li
Genomics Core Facility of the University of Leicester Technical note March 2012
 Genomics Core Facility of the University of Leicester Technical note March 2012 Consequences of various contaminants on RNA quantification and quality controls Nicolas Sylvius and Mariaelena Repici Department
Genomics Core Facility of the University of Leicester Technical note March 2012 Consequences of various contaminants on RNA quantification and quality controls Nicolas Sylvius and Mariaelena Repici Department
High Throughput TruSeq Stranded mrna Library Construction on the Biomek FX P
 High Throughput TruSeq Stranded mrna Library Construction on the Biomek FX P Zach Smith and Scott D. Michaels The Center for Genomics and Bioinformatics Indiana University Bloomington, IN USA Mary Blair
High Throughput TruSeq Stranded mrna Library Construction on the Biomek FX P Zach Smith and Scott D. Michaels The Center for Genomics and Bioinformatics Indiana University Bloomington, IN USA Mary Blair
BIMM 143. RNA sequencing overview. Genome Informatics II. Barry Grant. Lecture In vivo. In vitro.
 RNA sequencing overview BIMM 143 Genome Informatics II Lecture 14 Barry Grant http://thegrantlab.org/bimm143 In vivo In vitro In silico ( control) Goal: RNA quantification, transcript discovery, variant
RNA sequencing overview BIMM 143 Genome Informatics II Lecture 14 Barry Grant http://thegrantlab.org/bimm143 In vivo In vitro In silico ( control) Goal: RNA quantification, transcript discovery, variant
Supplemental Data. Integrating omics and alternative splicing i reveals insights i into grape response to high temperature
 Supplemental Data Integrating omics and alternative splicing i reveals insights i into grape response to high temperature Jianfu Jiang 1, Xinna Liu 1, Guotian Liu, Chonghuih Liu*, Shaohuah Li*, and Lijun
Supplemental Data Integrating omics and alternative splicing i reveals insights i into grape response to high temperature Jianfu Jiang 1, Xinna Liu 1, Guotian Liu, Chonghuih Liu*, Shaohuah Li*, and Lijun
RNA SEQUENCING AND DATA ANALYSIS
 RNA SEQUENCING AND DATA ANALYSIS Download slides and package http://odin.mdacc.tmc.edu/~rverhaak/package.zip http://odin.mdacc.tmc.edu/~rverhaak/rna-seqlecture.zip Overview Introduction into the topic
RNA SEQUENCING AND DATA ANALYSIS Download slides and package http://odin.mdacc.tmc.edu/~rverhaak/package.zip http://odin.mdacc.tmc.edu/~rverhaak/rna-seqlecture.zip Overview Introduction into the topic
Simple, rapid, and reliable RNA sequencing
 Simple, rapid, and reliable RNA sequencing RNA sequencing applications RNA sequencing provides fundamental insights into how genomes are organized and regulated, giving us valuable information about the
Simple, rapid, and reliable RNA sequencing RNA sequencing applications RNA sequencing provides fundamental insights into how genomes are organized and regulated, giving us valuable information about the
ESCs were lysed with Trizol reagent (Life technologies) and RNA was extracted according to
 Supplemental Methods RNA-seq ESCs were lysed with Trizol reagent (Life technologies) and RNA was extracted according to the manufacturer's instructions. RNAse free DNaseI (Sigma) was used to eliminate
Supplemental Methods RNA-seq ESCs were lysed with Trizol reagent (Life technologies) and RNA was extracted according to the manufacturer's instructions. RNAse free DNaseI (Sigma) was used to eliminate
Introduction to Systems Biology of Cancer Lecture 2
 Introduction to Systems Biology of Cancer Lecture 2 Gustavo Stolovitzky IBM Research Icahn School of Medicine at Mt Sinai DREAM Challenges High throughput measurements: The age of omics Systems Biology
Introduction to Systems Biology of Cancer Lecture 2 Gustavo Stolovitzky IBM Research Icahn School of Medicine at Mt Sinai DREAM Challenges High throughput measurements: The age of omics Systems Biology
Molecular Profiling of Tumor Microenvironment Alex Chenchik, Ph.D. Cellecta, Inc.
 Molecular Profiling of Tumor Microenvironment Alex Chenchik, Ph.D. Cellecta, Inc. Cellecta, Inc. Founded: April 2006 Headquarters: Mountain View, CA 12 SBIR Grants Custom Service Provider for Functional
Molecular Profiling of Tumor Microenvironment Alex Chenchik, Ph.D. Cellecta, Inc. Cellecta, Inc. Founded: April 2006 Headquarters: Mountain View, CA 12 SBIR Grants Custom Service Provider for Functional
For Research Use Only Ver
 INSTRUCTION MANUAL Quick-cfDNA/cfRNA Serum & Plasma Kit Catalog No. R1072 Highlights High-quality cell-free DNA and RNA are easily and robustly purified from up to 3 ml of plasma, serum, or other biological
INSTRUCTION MANUAL Quick-cfDNA/cfRNA Serum & Plasma Kit Catalog No. R1072 Highlights High-quality cell-free DNA and RNA are easily and robustly purified from up to 3 ml of plasma, serum, or other biological
QIAsymphony DSP Circulating DNA Kit
 QIAsymphony DSP Circulating DNA Kit February 2017 Performance Characteristics 937556 Sample to Insight Contents Performance Characteristics... 4 Basic performance... 4 Run precision... 6 Equivalent performance
QIAsymphony DSP Circulating DNA Kit February 2017 Performance Characteristics 937556 Sample to Insight Contents Performance Characteristics... 4 Basic performance... 4 Run precision... 6 Equivalent performance
The$mitochondrial$and$autosomal$mutation$landscapes$of$ prostate$cancer$ $supplementary$material$
 The$mitochondrial$and$autosomal$mutation$landscapes$of$ prostate$cancer$ $supplementary$material$ Johan Lindberg 1, Ian G. Mills 2-4, Daniel Klevebring 1, Wennuan Liu 5, Mårten Neiman 1, Jianfeng Xu 5,
The$mitochondrial$and$autosomal$mutation$landscapes$of$ prostate$cancer$ $supplementary$material$ Johan Lindberg 1, Ian G. Mills 2-4, Daniel Klevebring 1, Wennuan Liu 5, Mårten Neiman 1, Jianfeng Xu 5,
Supplementary Materials and Methods
 Supplementary Materials and Methods Whole Mount X-Gal Staining Whole tissues were collected, rinsed with PBS and fixed with 4% PFA. Tissues were then rinsed in rinse buffer (100 mm Sodium Phosphate ph
Supplementary Materials and Methods Whole Mount X-Gal Staining Whole tissues were collected, rinsed with PBS and fixed with 4% PFA. Tissues were then rinsed in rinse buffer (100 mm Sodium Phosphate ph
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Arabidopsis thaliana small RNA Sequencing. Report
 Arabidopsis thaliana small RNA Sequencing Report September 2015 Project Information Client Name Client Company / Institution Macrogen Order Number Order ID Species Arabidopsis thaliana Reference UCSC hg19
Arabidopsis thaliana small RNA Sequencing Report September 2015 Project Information Client Name Client Company / Institution Macrogen Order Number Order ID Species Arabidopsis thaliana Reference UCSC hg19
For the 5 GATC-overhang two-oligo adaptors set up the following reactions in 96-well plate format:
 Supplementary Protocol 1. Adaptor preparation: For the 5 GATC-overhang two-oligo adaptors set up the following reactions in 96-well plate format: Per reaction X96 10X NEBuffer 2 10 µl 10 µl x 96 5 -GATC
Supplementary Protocol 1. Adaptor preparation: For the 5 GATC-overhang two-oligo adaptors set up the following reactions in 96-well plate format: Per reaction X96 10X NEBuffer 2 10 µl 10 µl x 96 5 -GATC
well for 2 h at rt. Each dot represents an individual mouse and bar is the mean ±
 Supplementary data: Control DC Blimp-1 ko DC 8 6 4 2-2 IL-1β p=.5 medium 8 6 4 2 IL-2 Medium p=.16 8 6 4 2 IL-6 medium p=.3 5 4 3 2 1-1 medium IL-1 n.s. 25 2 15 1 5 IL-12(p7) p=.15 5 IFNγ p=.65 4 3 2 1
Supplementary data: Control DC Blimp-1 ko DC 8 6 4 2-2 IL-1β p=.5 medium 8 6 4 2 IL-2 Medium p=.16 8 6 4 2 IL-6 medium p=.3 5 4 3 2 1-1 medium IL-1 n.s. 25 2 15 1 5 IL-12(p7) p=.15 5 IFNγ p=.65 4 3 2 1
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits
 AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
PG-Seq NGS Kit for Preimplantation Genetic Screening
 Application Note: PG-Seq Validation Study PG-Seq NGS Kit for Preimplantation Genetic Screening Validation using Multi (5-10) Cells and Single Cells from euploid and aneuploid cell lines Introduction Advances
Application Note: PG-Seq Validation Study PG-Seq NGS Kit for Preimplantation Genetic Screening Validation using Multi (5-10) Cells and Single Cells from euploid and aneuploid cell lines Introduction Advances
CONTRACTING ORGANIZATION: Joan & Sanford I. Weill Medical College of Cornell University New York, NY 10065
 AWARD NUMBER: W81XWH-14-1-0263 TITLE: Early Detection of NSCLC Using Stromal Markers in Peripheral Blood PRINCIPAL INVESTIGATOR: Dingcheng Gao CONTRACTING ORGANIZATION: Joan & Sanford I. Weill Medical
AWARD NUMBER: W81XWH-14-1-0263 TITLE: Early Detection of NSCLC Using Stromal Markers in Peripheral Blood PRINCIPAL INVESTIGATOR: Dingcheng Gao CONTRACTING ORGANIZATION: Joan & Sanford I. Weill Medical
CONTRACTING ORGANIZATION: Johns Hopkins University, Baltimore, MD
 AD Award Number: W81XWH-12-1-0480 TITLE: Molecular Characterization of Indolent Prostate Cancer PRINCIPAL INVESTIGATOR: Jun Luo, Ph.D. CONTRACTING ORGANIZATION: Johns Hopkins University, Baltimore, MD
AD Award Number: W81XWH-12-1-0480 TITLE: Molecular Characterization of Indolent Prostate Cancer PRINCIPAL INVESTIGATOR: Jun Luo, Ph.D. CONTRACTING ORGANIZATION: Johns Hopkins University, Baltimore, MD
For Research Use Only Ver
 INSTRUCTION MANUAL Quick-RNA Miniprep Kit Catalog Nos. R1054 & R1055 Highlights High-quality total RNA (including small RNAs) from a wide range of samples. You can opt to isolate small and large RNAs in
INSTRUCTION MANUAL Quick-RNA Miniprep Kit Catalog Nos. R1054 & R1055 Highlights High-quality total RNA (including small RNAs) from a wide range of samples. You can opt to isolate small and large RNAs in
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library
 Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
4. Th1-related gene expression in infected versus mock-infected controls from Fig. 2 with gene annotation.
 List of supplemental information 1. Graph of mouse weight loss during course of infection- Line graphs showing mouse weight data during course of infection days 1 to 10 post-infections (p.i.). 2. Graph
List of supplemental information 1. Graph of mouse weight loss during course of infection- Line graphs showing mouse weight data during course of infection days 1 to 10 post-infections (p.i.). 2. Graph
For Research Use Only Ver
 INSTRUCTION MANUAL Quick-RNA Miniprep Kit Catalog Nos. R1054 & R1055 Highlights High-quality total RNA (including small RNAs) from a wide range of samples. You can opt to isolate small and large RNAs in
INSTRUCTION MANUAL Quick-RNA Miniprep Kit Catalog Nos. R1054 & R1055 Highlights High-quality total RNA (including small RNAs) from a wide range of samples. You can opt to isolate small and large RNAs in
Supplementary Figure 1. SA-β-Gal positive senescent cells in various cancer tissues. Representative frozen sections of breast, thyroid, colon and
 Supplementary Figure 1. SA-β-Gal positive senescent cells in various cancer tissues. Representative frozen sections of breast, thyroid, colon and stomach cancer were stained with SA-β-Gal and nuclear fast
Supplementary Figure 1. SA-β-Gal positive senescent cells in various cancer tissues. Representative frozen sections of breast, thyroid, colon and stomach cancer were stained with SA-β-Gal and nuclear fast
York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
 MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
Antibodies: LB1 buffer For 50 ml For 10ml For 30 ml Final 1 M HEPES, ph 2.5 ml 0.5 ml 1.5 ml 50mM. 5 M NaCl 1.4 ml 280 µl 0.
 Experiment: Date: Tissue: Purpose: ChIP-Seq Antibodies: 11x cross-link buffer: Regent Stock Solution Final Vol for 10 ml of 11xstock concentration 5 M NaCl 0.1M 0.2 ml 0.5 M EDTA 1 mm 20 ul 0.5 M EGTA,
Experiment: Date: Tissue: Purpose: ChIP-Seq Antibodies: 11x cross-link buffer: Regent Stock Solution Final Vol for 10 ml of 11xstock concentration 5 M NaCl 0.1M 0.2 ml 0.5 M EDTA 1 mm 20 ul 0.5 M EGTA,
Alternative splicing detection workflow needs a careful combination of sample prep and bioinformatics analysis
 RESEARCH Open Access Alternative splicing detection workflow needs a careful combination of sample prep and bioinformatics analysis Matteo Carrara 1, Josephine Lum 2, Francesca Cordero 3, Marco Beccuti
RESEARCH Open Access Alternative splicing detection workflow needs a careful combination of sample prep and bioinformatics analysis Matteo Carrara 1, Josephine Lum 2, Francesca Cordero 3, Marco Beccuti
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Accessing and Using ENCODE Data Dr. Peggy J. Farnham
 1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
Effects of UBL5 knockdown on cell cycle distribution and sister chromatid cohesion
 Supplementary Figure S1. Effects of UBL5 knockdown on cell cycle distribution and sister chromatid cohesion A. Representative examples of flow cytometry profiles of HeLa cells transfected with indicated
Supplementary Figure S1. Effects of UBL5 knockdown on cell cycle distribution and sister chromatid cohesion A. Representative examples of flow cytometry profiles of HeLa cells transfected with indicated
Products for cfdna and mirna isolation. Subhead Circulating Cover nucleic acids from plasma
 MACHEREY-NAGEL Products for cfdna and mirna isolation Bioanalysis Subhead Circulating Cover nucleic acids from plasma n Flexible solutions for small and large blood plasma volumes n Highly efficient recovery
MACHEREY-NAGEL Products for cfdna and mirna isolation Bioanalysis Subhead Circulating Cover nucleic acids from plasma n Flexible solutions for small and large blood plasma volumes n Highly efficient recovery
ExpressArt FFPE Clear RNAready kit
 Features and Example Results General problems with FFPE samples Formalin-fixation of tissues results in severe RNA fragmentation, as well as in RNA RNA, RNA-DNA and RNA protein cross-linking, which impairs
Features and Example Results General problems with FFPE samples Formalin-fixation of tissues results in severe RNA fragmentation, as well as in RNA RNA, RNA-DNA and RNA protein cross-linking, which impairs
WHITE PAPER. Increasing Ligation Efficiency and Discovery of mirnas for Small RNA NGS Sequencing Library Prep with Plant Samples
 WHITE PPER Increasing Ligation Efficiency and iscovery of mirns for Small RN NGS Sequencing Library Prep with Plant Samples ioo Scientific White Paper May 2017 bstract MicroRNs (mirns) are 18-22 nucleotide
WHITE PPER Increasing Ligation Efficiency and iscovery of mirns for Small RN NGS Sequencing Library Prep with Plant Samples ioo Scientific White Paper May 2017 bstract MicroRNs (mirns) are 18-22 nucleotide
Supplemental Table S1. Clinical Characteristics of the Study Populations. Values are presented
 Supplemental Table S1. Clinical Characteristics of the Study Populations. Values are presented as means ± standard deviation (STD) in LTRC Yale Cohort (), Pittsburgh Cohort () and Korean san cohort (C),
Supplemental Table S1. Clinical Characteristics of the Study Populations. Values are presented as means ± standard deviation (STD) in LTRC Yale Cohort (), Pittsburgh Cohort () and Korean san cohort (C),
For Research Use Only Ver
 INSTRUCTION MANUAL Quick-cfDNA Serum & Plasma Kit Catalog No. D4076 Highlights High-quality DNA, including cell-free, is easily and robustly purified from up to 10 ml of serum/plasma, up to 1 ml amniotic
INSTRUCTION MANUAL Quick-cfDNA Serum & Plasma Kit Catalog No. D4076 Highlights High-quality DNA, including cell-free, is easily and robustly purified from up to 10 ml of serum/plasma, up to 1 ml amniotic
Supplementary Materials for
 immunology.sciencemag.org/cgi/content/full/2/16/eaan6049/dc1 Supplementary Materials for Enzymatic synthesis of core 2 O-glycans governs the tissue-trafficking potential of memory CD8 + T cells Jossef
immunology.sciencemag.org/cgi/content/full/2/16/eaan6049/dc1 Supplementary Materials for Enzymatic synthesis of core 2 O-glycans governs the tissue-trafficking potential of memory CD8 + T cells Jossef
PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland
 AD Award Number: W81XWH-12-1-0480 TITLE: Molecular Characterization of Indolent Prostate Cancer PRINCIPAL INVESTIGATOR: Jun Luo, Ph.D. CONTRACTING ORGANIZATION: Johns Hopkins University Baltimore, MD 21218-2680
AD Award Number: W81XWH-12-1-0480 TITLE: Molecular Characterization of Indolent Prostate Cancer PRINCIPAL INVESTIGATOR: Jun Luo, Ph.D. CONTRACTING ORGANIZATION: Johns Hopkins University Baltimore, MD 21218-2680
Extracellular Vesicle RNA isolation Kits
 Extracellular Vesicle RNA isolation Kits Summary Section 4 Introduction 42 Exo-TotalRNA and TumorExo-TotalRNA isolation kits 43 Extracellular Vesicle RNA extraction kits Ordering information Products can
Extracellular Vesicle RNA isolation Kits Summary Section 4 Introduction 42 Exo-TotalRNA and TumorExo-TotalRNA isolation kits 43 Extracellular Vesicle RNA extraction kits Ordering information Products can
Transcript reconstruction
 Transcript reconstruction Summary I Data types, file formats and utilities Annotation: Genomic regions Genes Peaks bedtools Alignment: Map reads BAM/SAM Samtools Aggregation: Summary files Wig (UCSC) TDF
Transcript reconstruction Summary I Data types, file formats and utilities Annotation: Genomic regions Genes Peaks bedtools Alignment: Map reads BAM/SAM Samtools Aggregation: Summary files Wig (UCSC) TDF
qpcr-array Analysis Service
 qpcr-array Analysis Service Customer Name Institute Telephone Address E-mail PO Number Service Code Report Date Service Laboratory Department Phalanx Biotech Group, Inc 6 Floor, No.6, Technology Road 5,
qpcr-array Analysis Service Customer Name Institute Telephone Address E-mail PO Number Service Code Report Date Service Laboratory Department Phalanx Biotech Group, Inc 6 Floor, No.6, Technology Road 5,
Obstacles and challenges in the analysis of microrna sequencing data
 Obstacles and challenges in the analysis of microrna sequencing data (mirna-seq) David Humphreys Genomics core Dr Victor Chang AC 1936-1991, Pioneering Cardiothoracic Surgeon and Humanitarian The ABCs
Obstacles and challenges in the analysis of microrna sequencing data (mirna-seq) David Humphreys Genomics core Dr Victor Chang AC 1936-1991, Pioneering Cardiothoracic Surgeon and Humanitarian The ABCs
Supplementary Information
 Supplementary Information TABLE S1. SUBJECT CHARACTERISTICS* Normal Control Subjects Subjects with Asthma p Value Number 23 48 Age (years) 35±10 35±10 0.75 Sex, M:F (% F) 9:12 (57) 17:26 (60) 0.76 FEV1
Supplementary Information TABLE S1. SUBJECT CHARACTERISTICS* Normal Control Subjects Subjects with Asthma p Value Number 23 48 Age (years) 35±10 35±10 0.75 Sex, M:F (% F) 9:12 (57) 17:26 (60) 0.76 FEV1
Exosome DNA Extraction Kits
 Exosome DNA Extraction Kits Summary Section 5 Introduction 40 EXO-DNAc 41 EXO-DNA 43 Introduction Genomic DNA Extractiom Kits Ordering informations Products can be purchased directly in our on-line shop:
Exosome DNA Extraction Kits Summary Section 5 Introduction 40 EXO-DNAc 41 EXO-DNA 43 Introduction Genomic DNA Extractiom Kits Ordering informations Products can be purchased directly in our on-line shop:
P. Tang ( 鄧致剛 ); PJ Huang ( 黄栢榕 ) g( ); g ( ) Bioinformatics Center, Chang Gung University.
 Databases and Tools for High Throughput Sequencing Analysis P. Tang ( 鄧致剛 ); PJ Huang ( 黄栢榕 ) g( ); g ( ) Bioinformatics Center, Chang Gung University. HTseq Platforms Applications on Biomedical Sciences
Databases and Tools for High Throughput Sequencing Analysis P. Tang ( 鄧致剛 ); PJ Huang ( 黄栢榕 ) g( ); g ( ) Bioinformatics Center, Chang Gung University. HTseq Platforms Applications on Biomedical Sciences
MODULE 3: TRANSCRIPTION PART II
 MODULE 3: TRANSCRIPTION PART II Lesson Plan: Title S. CATHERINE SILVER KEY, CHIYEDZA SMALL Transcription Part II: What happens to the initial (premrna) transcript made by RNA pol II? Objectives Explain
MODULE 3: TRANSCRIPTION PART II Lesson Plan: Title S. CATHERINE SILVER KEY, CHIYEDZA SMALL Transcription Part II: What happens to the initial (premrna) transcript made by RNA pol II? Objectives Explain
A Statistical Framework for Classification of Tumor Type from microrna Data
 DEGREE PROJECT IN MATHEMATICS, SECOND CYCLE, 30 CREDITS STOCKHOLM, SWEDEN 2016 A Statistical Framework for Classification of Tumor Type from microrna Data JOSEFINE RÖHSS KTH ROYAL INSTITUTE OF TECHNOLOGY
DEGREE PROJECT IN MATHEMATICS, SECOND CYCLE, 30 CREDITS STOCKHOLM, SWEDEN 2016 A Statistical Framework for Classification of Tumor Type from microrna Data JOSEFINE RÖHSS KTH ROYAL INSTITUTE OF TECHNOLOGY
Lecture 8 Understanding Transcription RNA-seq analysis. Foundations of Computational Systems Biology David K. Gifford
 Lecture 8 Understanding Transcription RNA-seq analysis Foundations of Computational Systems Biology David K. Gifford 1 Lecture 8 RNA-seq Analysis RNA-seq principles How can we characterize mrna isoform
Lecture 8 Understanding Transcription RNA-seq analysis Foundations of Computational Systems Biology David K. Gifford 1 Lecture 8 RNA-seq Analysis RNA-seq principles How can we characterize mrna isoform
Phosphate buffered saline (PBS) for washing the cells TE buffer (nuclease-free) ph 7.5 for use with the PrimePCR Reverse Transcription Control Assay
 Catalog # Description 172-5080 SingleShot Cell Lysis Kit, 100 x 50 µl reactions 172-5081 SingleShot Cell Lysis Kit, 500 x 50 µl reactions For research purposes only. Introduction The SingleShot Cell Lysis
Catalog # Description 172-5080 SingleShot Cell Lysis Kit, 100 x 50 µl reactions 172-5081 SingleShot Cell Lysis Kit, 500 x 50 µl reactions For research purposes only. Introduction The SingleShot Cell Lysis
Small RNA Sequencing. Project Workflow. Service Description. Sequencing Service Specification BGISEQ-500 SERVICE OVERVIEW SAMPLE PREPARATION
 BGISEQ-500 SERVICE OVERVIEW Small RNA Sequencing Service Description Small RNAs are a type of non-coding RNA (ncrna) molecules that are less than 200nt in length. They are often involved in gene silencing
BGISEQ-500 SERVICE OVERVIEW Small RNA Sequencing Service Description Small RNAs are a type of non-coding RNA (ncrna) molecules that are less than 200nt in length. They are often involved in gene silencing
ncounter Assay Automated Process Immobilize and align reporter for image collecting and barcode counting ncounter Prep Station
 ncounter Assay ncounter Prep Station Automated Process Hybridize Reporter to RNA Remove excess reporters Bind reporter to surface Immobilize and align reporter Image surface Count codes Immobilize and
ncounter Assay ncounter Prep Station Automated Process Hybridize Reporter to RNA Remove excess reporters Bind reporter to surface Immobilize and align reporter Image surface Count codes Immobilize and
A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis
 A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis Jian Xu, Ph.D. Children s Research Institute, UTSW Introduction Outline Overview of genomic and next-gen sequencing technologies
A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis Jian Xu, Ph.D. Children s Research Institute, UTSW Introduction Outline Overview of genomic and next-gen sequencing technologies
Norgen s HIV proviral DNA PCR Kit was developed and validated to be used with the following PCR instruments: Qiagen Rotor-Gene Q BioRad icycler
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product # 33840 Product Insert Background Information
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product # 33840 Product Insert Background Information
SUPPLEMENTARY METHODS
 SUPPLEMENTARY METHODS Histological analysis. Colonic tissues were collected from 5 parts of the middle colon on day 7 after the start of DSS treatment, and then were cut into segments, fixed with 4% paraformaldehyde,
SUPPLEMENTARY METHODS Histological analysis. Colonic tissues were collected from 5 parts of the middle colon on day 7 after the start of DSS treatment, and then were cut into segments, fixed with 4% paraformaldehyde,
Norgen s HIV Proviral DNA PCR Kit was developed and validated to be used with the following PCR instruments: Qiagen Rotor-Gene Q BioRad T1000 Cycler
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product# 33840 Product Insert Intended
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product# 33840 Product Insert Intended
Hepatitis B Virus Genemer
 Product Manual Hepatitis B Virus Genemer Primer Pair for amplification of HBV Viral Specific Fragment Catalog No.: 60-2007-10 Store at 20 o C For research use only. Not for use in diagnostic procedures
Product Manual Hepatitis B Virus Genemer Primer Pair for amplification of HBV Viral Specific Fragment Catalog No.: 60-2007-10 Store at 20 o C For research use only. Not for use in diagnostic procedures
High-Throughput Sequencing Course
 High-Throughput Sequencing Course Introduction Biostatistics and Bioinformatics Summer 2017 From Raw Unaligned Reads To Aligned Reads To Counts Differential Expression Differential Expression 3 2 1 0 1
High-Throughput Sequencing Course Introduction Biostatistics and Bioinformatics Summer 2017 From Raw Unaligned Reads To Aligned Reads To Counts Differential Expression Differential Expression 3 2 1 0 1
% of live splenocytes. STAT5 deletion. (open shapes) % ROSA + % floxed
 Supp. Figure 1. a 14 1 1 8 6 spleen cells (x1 6 ) 16 % of live splenocytes 5 4 3 1 % of live splenocytes 8 6 4 b 1 1 c % of CD11c + splenocytes (closed shapes) 8 6 4 8 6 4 % ROSA + (open shapes) % floxed
Supp. Figure 1. a 14 1 1 8 6 spleen cells (x1 6 ) 16 % of live splenocytes 5 4 3 1 % of live splenocytes 8 6 4 b 1 1 c % of CD11c + splenocytes (closed shapes) 8 6 4 8 6 4 % ROSA + (open shapes) % floxed
Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder Labs August 30, 2012
 Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
Nature Immunology: doi: /ni Supplementary Figure 1. Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice.
 Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
Comprehensive nucleosome mapping of the human genome in cancer progression
 /, Vol. 7, No. 12 Comprehensive nucleosome mapping of the human genome in cancer progression Brooke R. Druliner 1,5, Daniel Vera 1,6, Ruth Johnson 2, Xiaoyang Ruan 3, Lynn M. Apone 4, Eileen T. Dimalanta
/, Vol. 7, No. 12 Comprehensive nucleosome mapping of the human genome in cancer progression Brooke R. Druliner 1,5, Daniel Vera 1,6, Ruth Johnson 2, Xiaoyang Ruan 3, Lynn M. Apone 4, Eileen T. Dimalanta
RNA Seq Analysis of Non-Alcoholic Fatty Liver Disease (NAFLD) Induced by Metabolic Syndrome in a Mouse Model
 University of Massachusetts Boston ScholarWorks at UMass Boston Honors College Theses 5-2016 RNA Seq Analysis of Non-Alcoholic Fatty Liver Disease (NAFLD) Induced by Metabolic Syndrome in a Mouse Model
University of Massachusetts Boston ScholarWorks at UMass Boston Honors College Theses 5-2016 RNA Seq Analysis of Non-Alcoholic Fatty Liver Disease (NAFLD) Induced by Metabolic Syndrome in a Mouse Model
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research
 Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
MODULE 4: SPLICING. Removal of introns from messenger RNA by splicing
 Last update: 05/10/2017 MODULE 4: SPLICING Lesson Plan: Title MEG LAAKSO Removal of introns from messenger RNA by splicing Objectives Identify splice donor and acceptor sites that are best supported by
Last update: 05/10/2017 MODULE 4: SPLICING Lesson Plan: Title MEG LAAKSO Removal of introns from messenger RNA by splicing Objectives Identify splice donor and acceptor sites that are best supported by
ncounter Assay Automated Process Capture & Reporter Probes Bind reporter to surface Remove excess reporters Hybridize CodeSet to RNA
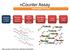 ncounter Assay Automated Process Hybridize CodeSet to RNA Remove excess reporters Bind reporter to surface Immobilize and align reporter Image surface Count codes mrna Capture & Reporter Probes slides
ncounter Assay Automated Process Hybridize CodeSet to RNA Remove excess reporters Bind reporter to surface Immobilize and align reporter Image surface Count codes mrna Capture & Reporter Probes slides
Supplementary Figures. Supplementary Figure 1. Treatment schematic of SIV infection and ARV and PP therapies.
 Supplementary Figures Supplementary Figure 1. Treatment schematic of SIV infection and ARV and PP therapies. Supplementary Figure 2. SIV replication and CD4 + T cell count. (A) Log SIVmac239 copies/ml
Supplementary Figures Supplementary Figure 1. Treatment schematic of SIV infection and ARV and PP therapies. Supplementary Figure 2. SIV replication and CD4 + T cell count. (A) Log SIVmac239 copies/ml
Multiplex target enrichment using DNA indexing for ultra-high throughput variant detection
 Multiplex target enrichment using DNA indexing for ultra-high throughput variant detection Dr Elaine Kenny Neuropsychiatric Genetics Research Group Institute of Molecular Medicine Trinity College Dublin
Multiplex target enrichment using DNA indexing for ultra-high throughput variant detection Dr Elaine Kenny Neuropsychiatric Genetics Research Group Institute of Molecular Medicine Trinity College Dublin
ncounter TM Analysis System
 ncounter TM Analysis System Molecules That Count TM www.nanostring.com Agenda NanoString Technologies History Introduction to the ncounter Analysis System CodeSet Design and Assay Principals System Performance
ncounter TM Analysis System Molecules That Count TM www.nanostring.com Agenda NanoString Technologies History Introduction to the ncounter Analysis System CodeSet Design and Assay Principals System Performance
Interleukin-20 is associated with delayed healing in diabetic wounds
 Interleukin-20 is associated with delayed healing in diabetic wounds Phillip Finley, PhD Integrated and Applied Sciences Program Biology and Statistics/Research Methodology Normal Healing Body s natural
Interleukin-20 is associated with delayed healing in diabetic wounds Phillip Finley, PhD Integrated and Applied Sciences Program Biology and Statistics/Research Methodology Normal Healing Body s natural
RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
 Xu et al. Molecular Cancer (2019) 18:8 https://doi.org/10.1186/s12943-018-0932-8 LETTER TO THE EDITOR RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
Xu et al. Molecular Cancer (2019) 18:8 https://doi.org/10.1186/s12943-018-0932-8 LETTER TO THE EDITOR RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS
 Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS dr sc. Ana Krivokuća Laboratory for molecular genetics Institute for Oncology and
Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS dr sc. Ana Krivokuća Laboratory for molecular genetics Institute for Oncology and
Supplementary Figure 1 a
 Supplementary Figure 1 a b d c Supplementary Figure 1. Poor sensitivity for mutation detection using established single-cell RNA-sequencing methods. (a) Representative bioanalyzer trace showing expected
Supplementary Figure 1 a b d c Supplementary Figure 1. Poor sensitivity for mutation detection using established single-cell RNA-sequencing methods. (a) Representative bioanalyzer trace showing expected
Supplement Figure S1. Real Time PCR analysis of mrna levels of C/EBPα and PU.1 in wild type (WT) and NQO1-null (NQO1-/-) mice.
 competes with 20S proteasome for binding with C/EBP leading to its stabilization and Relative mrna levels Supplement Figure S1. Real Time PCR analysis of mrna levels of C/EBPα and PU.1 in wild type (WT)
competes with 20S proteasome for binding with C/EBP leading to its stabilization and Relative mrna levels Supplement Figure S1. Real Time PCR analysis of mrna levels of C/EBPα and PU.1 in wild type (WT)
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH
 EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
ChIP-seq hands-on. Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs
 ChIP-seq hands-on Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs Main goals Becoming familiar with essential tools and formats Visualizing and contextualizing raw data Understand
ChIP-seq hands-on Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs Main goals Becoming familiar with essential tools and formats Visualizing and contextualizing raw data Understand
Table S1. Total and mapped reads produced for each ChIP-seq sample
 Tale S1. Total and mapped reads produced for each ChIP-seq sample Sample Total Reads Mapped Reads Col- H3K27me3 rep1 125662 1334323 (85.76%) Col- H3K27me3 rep2 9176437 7986731 (87.4%) atmi1a//c H3K27m3
Tale S1. Total and mapped reads produced for each ChIP-seq sample Sample Total Reads Mapped Reads Col- H3K27me3 rep1 125662 1334323 (85.76%) Col- H3K27me3 rep2 9176437 7986731 (87.4%) atmi1a//c H3K27m3
Single Cell Quantitative Polymer Chain Reaction (sc-qpcr)
 Single Cell Quantitative Polymer Chain Reaction (sc-qpcr) Analyzing gene expression profiles from a bulk population of cells provides an average profile which may obscure important biological differences
Single Cell Quantitative Polymer Chain Reaction (sc-qpcr) Analyzing gene expression profiles from a bulk population of cells provides an average profile which may obscure important biological differences
Product Contents. 1 Specifications 1 Product Description. 2 Buffer Preparation... 3 Protocol. 3 Ordering Information 4
 INSTRUCTION MANUAL Quick-RNA Midiprep Kit Catalog No. R1056 Highlights 10 minute method for isolating RNA (up to 1 mg) from a wide range of cell types and tissue samples. Clean-Spin column technology allows
INSTRUCTION MANUAL Quick-RNA Midiprep Kit Catalog No. R1056 Highlights 10 minute method for isolating RNA (up to 1 mg) from a wide range of cell types and tissue samples. Clean-Spin column technology allows
Supplementary Figure 1. General strategy to classify genes and identify TSGs.
 Supplementary Figure 1. General strategy to classify genes and identify TSGs. Supplementary Figure 2. EE patterns of validated EECTPs (7 novel) in our 24 LUAD samples. Red indicates extremely highly expressed
Supplementary Figure 1. General strategy to classify genes and identify TSGs. Supplementary Figure 2. EE patterns of validated EECTPs (7 novel) in our 24 LUAD samples. Red indicates extremely highly expressed
