Supplementary Figures. Supplementary Figure 1. Treatment schematic of SIV infection and ARV and PP therapies.
|
|
|
- Alexis Harrington
- 6 years ago
- Views:
Transcription
1 Supplementary Figures Supplementary Figure 1. Treatment schematic of SIV infection and ARV and PP therapies.
2 Supplementary Figure 2. SIV replication and CD4 + T cell count. (A) Log SIVmac239 copies/ml plasma after SIV infection (day 0). Dotted vertical line at day 100 post-siv represents initiation of probiotics/prebiotics alone in ARV+PP animals (ARV alone not treated). Dotted vertical line at day 155 post-siv represents initiation of antiretroviral therapy in all animals. (B) Absolute CD4 + T cell count/mm 3 blood. Frequency of CD4 + T cells and lymphocyte complete blood cell count used for analysis. Dotted verticle line represents initiation of ARV therapy at 155 days post-siv. (A-B) Solid red lines, animals treated with ARVs and PP with suppressed viremia PTM; dotted red line, slow responder PTM treated with ARVs and PP; Black lines, animals treated with ARVs alone.
3 Supplementary Figure 3. Fecal microbiome analysis. Pyrosequencing of 16s rrna was performed on fecal samples. (a) Ribosomal Database Project (RDP) classification was used for taxonomical assignments. Left, ARV alone PTM, right, ARV + PP PTM, separated left to right in order of: pre-infection, post-infection (D90), after treatment with 5 months ARVs alone or ARVs + PP. Animal names denoted under bars. Supplementary Figure 4. CD4 + T cells in jejunum and without Mane-A1*084+. (A) Longitudinal frequencies of CD4 + T cells in jejunal biopsy samples. Red line, PTM treated with ARVs and probiotics (both suppressed and viremic animals), black line, PTM treated with ARV alone. Verticle dotted line, initiation of ARV at day 155 post-siv infection. Points represent mean, vertical lines represent standard deviation. (B) Frequency of CD4 + T cells from jejunum tissues at necropsy. Circles, ARVs and PP
4 treated PTM with suppressed viremia; square, ARVs and PP treated slow responder PTM; Triangles, PTMs treated with ARVs. Supplementary Figure 5. Microbial translocation and immune activation. (A-B) Microbial translocation measured by the fraction of area stained for lipopolysaccharide (LPS) by immunohistochemistry in (A) axillary LN or (B) mesenteric LN taken at necropsy. (C-D) Peripheral immune activation measured by the fraction of (C) CD4 + T cells or (D) CD8 + T cells that expressed Ki67 in peripheral blood at necropsy. P value from Mann-Whitney T test. Circles, ARVs and PP treated PTM with suppressed viremia; square, ARVs and PP treated slow responder PTM; Triangles, PTMs treated with ARVs.
5 Supplementary Figure 6. The fraction of CD4+ T cells in the colon, with PTM that expressed Mane-A1*084 in red. P values from Mann Whitney T test. Circles, ARVs and PP treated PTM with suppressed viremia; square, ARVs and PP treated slow responder PTM; Triangles, PTMs treated with ARVs.
6 Supplemental Methods Microbiome Analysis DNA from one gram of feces (collected and immediately stored at -80oC) was obtained using the QIAmp protocol for larger volumes of stool. This protocol was modified as follows: We performed initial lysis of bacterial cells for 5 minutes at 95oC (rather than 70oC) as recommended for gram negative bacteria and incubated Proteinase K for 20 minutes at 70oC (rather than 10 minutes). For quantitative analysis of 16S rdna, real time PCR was performed using primers BacF (5 -CGGCAACGAGCGCAACCC-3 ) and BacR (5 -CCATTGTAGCACGTGTGTAGCC-3 ) (2). For sequencing of 16S rdna amplicon libraries were prepared from sample DNA using Accuprime High Fidelity Taq polymerase (Invitrogen) and universal primers flanking variable regions V1 (primer 27F; 5 -AGAGTTTGATCCTGGCTCAG-3 ) and V3 (primer 534R; 5 ATTACCGCGGCTGCTGG-3 ). For each sample, the universal primers were tagged with unique sequences ("barcodes") to allow for multiplexing/demultiplexing (3) PCR products were then purified using the Agencourt Ampure XP Kit (Beckman Counter Genomics) and quantitated using the QuantIT dsdna High-Sensitivity Assay Kit (Invitrogen). Approximately equivalent amounts of each PCR product were then pooled and purified with a Qiagen minelute column (Qiagen) into 30 µl TE buffer prior to sequencing at the NIH Intramural Sequencing Center. Amplicon libraries were sequenced on a 454 FLX instrument using Titanium chemistry. Flowgrams were processed using the 454 Basecalling pipeline (v2.5.3). Sequence pre-processing, alignment and chimera removal: mothur (version ) (4) was used for all 16S rrna gene sequence analysis steps. Prior to analysis, sequences were trimmed of low quality ends and filtered to retain sequences with a minimum length of 200 bp. After alignment to a bacterial reference alignment (SILVA), chimeras were removed using the chimera slayer implementation in the mothur package. Taxonomic classification of reads was done using the RDP Classifier included in mothur.
LIFESTYLE-ASSOCIATED ALTERATIONS OF INTESTINAL
 LIFESTYLE-ASSOCIATED ALTERATIONS OF INTESTINAL MICROBIOTA AND THEIR POTENTIAL ROLE IN ETIOLOGY OF IBD: A METAGENOMIC ANALYSIS Stephan J. Ott, University Hospital (UKSH), Campus Kiel Institute for Clinical
LIFESTYLE-ASSOCIATED ALTERATIONS OF INTESTINAL MICROBIOTA AND THEIR POTENTIAL ROLE IN ETIOLOGY OF IBD: A METAGENOMIC ANALYSIS Stephan J. Ott, University Hospital (UKSH), Campus Kiel Institute for Clinical
The human milk microbiome healthy breasts, mothers, and babies. James A. Foster 22 September 2010
 The human milk microbiome healthy breasts, mothers, and babies James A. Foster 22 September 2010 The human milk microbiome healthy breasts, mothers, and babies James A. Foster 22 September 2010 Who cares
The human milk microbiome healthy breasts, mothers, and babies James A. Foster 22 September 2010 The human milk microbiome healthy breasts, mothers, and babies James A. Foster 22 September 2010 Who cares
Microbiome 101: Sequencing, Nomenclature, and Methodology
 Microbiome 101: Sequencing, Nomenclature, and Methodology Ian Carroll, PhD Assistant Professor Division of Gastroenterology and Hepatology UNC Chapel Hill Background Microbiota: an ecological community
Microbiome 101: Sequencing, Nomenclature, and Methodology Ian Carroll, PhD Assistant Professor Division of Gastroenterology and Hepatology UNC Chapel Hill Background Microbiota: an ecological community
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 NLRP12 is downregulated in biopsy samples from patients with active ulcerative colitis (UC). (a-g) NLRP12 expression in 7 UC mrna profiling studies deposited in NCBI GEO database.
Supplementary Figure 1 NLRP12 is downregulated in biopsy samples from patients with active ulcerative colitis (UC). (a-g) NLRP12 expression in 7 UC mrna profiling studies deposited in NCBI GEO database.
Probiotic/prebiotic supplementation of antiretrovirals improves gastrointestinal immunity in SIV-infected macaques
 Related article, page 544 Brief report Probiotic/prebiotic supplementation of antiretrovirals improves gastrointestinal immunity in SIV-infected macaques Nichole R. Klatt, 1 Lauren A. Canary, 1 Xiaoyong
Related article, page 544 Brief report Probiotic/prebiotic supplementation of antiretrovirals improves gastrointestinal immunity in SIV-infected macaques Nichole R. Klatt, 1 Lauren A. Canary, 1 Xiaoyong
CROI 2016 Review: Immunology and Vaccines
 Frontier AIDS Education and Training Center CROI 2016 Review: Immunology and Vaccines Meena Ramchandani MD MPH Acting Instructor, University of Washington March 2016 This presentation is intended for educational
Frontier AIDS Education and Training Center CROI 2016 Review: Immunology and Vaccines Meena Ramchandani MD MPH Acting Instructor, University of Washington March 2016 This presentation is intended for educational
Are microbes the missing piece of the gut comfort puzzle?
 Priority Research Programme Foods for improving gut function and comfort Are microbes the missing piece of the gut comfort puzzle? Wayne Young Senior Scientist, AgResearch Host institution Foods for gut
Priority Research Programme Foods for improving gut function and comfort Are microbes the missing piece of the gut comfort puzzle? Wayne Young Senior Scientist, AgResearch Host institution Foods for gut
Selective depletion of abundant RNAs to enable transcriptome analysis of lowinput and highly-degraded RNA from FFPE breast cancer samples
 DNA CLONING DNA AMPLIFICATION & PCR EPIGENETICS RNA ANALYSIS Selective depletion of abundant RNAs to enable transcriptome analysis of lowinput and highly-degraded RNA from FFPE breast cancer samples LIBRARY
DNA CLONING DNA AMPLIFICATION & PCR EPIGENETICS RNA ANALYSIS Selective depletion of abundant RNAs to enable transcriptome analysis of lowinput and highly-degraded RNA from FFPE breast cancer samples LIBRARY
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
QIAsymphony DSP Circulating DNA Kit
 QIAsymphony DSP Circulating DNA Kit February 2017 Performance Characteristics 937556 Sample to Insight Contents Performance Characteristics... 4 Basic performance... 4 Run precision... 6 Equivalent performance
QIAsymphony DSP Circulating DNA Kit February 2017 Performance Characteristics 937556 Sample to Insight Contents Performance Characteristics... 4 Basic performance... 4 Run precision... 6 Equivalent performance
in the Gastrointestinal and Reproductive Tracts of Quarter Horse Mares
 Influence of Probiotics on Microflora in the Gastrointestinal and Reproductive Tracts of Quarter Horse Mares Katie Barnhart Research Advisors: Dr. Kimberly Cole and Dr. John Mark Reddish Department of
Influence of Probiotics on Microflora in the Gastrointestinal and Reproductive Tracts of Quarter Horse Mares Katie Barnhart Research Advisors: Dr. Kimberly Cole and Dr. John Mark Reddish Department of
Figure S1. Schematic presentation of genomic replication of idsiv after transfection and infection. After transfection of idsiv plasmid DNA into 293T
 Figure S1. Schematic presentation of genomic replication of idsiv after transfection and infection. After transfection of idsiv plasmid DNA into 293T cells, the RNA genomes with all modifications are generated
Figure S1. Schematic presentation of genomic replication of idsiv after transfection and infection. After transfection of idsiv plasmid DNA into 293T cells, the RNA genomes with all modifications are generated
collected for biochemical and molecular microbiological analyses at baseline, week 4 and
 SUPPLEMENTARY FIGURE LEGENDS Figure S1 Study design. Figure was adapted from Paramsothy et al. 5 Blood and stool samples were collected for biochemical and molecular microbiological analyses at baseline,
SUPPLEMENTARY FIGURE LEGENDS Figure S1 Study design. Figure was adapted from Paramsothy et al. 5 Blood and stool samples were collected for biochemical and molecular microbiological analyses at baseline,
Combined IL-21 and IFNα treatment limits inflammation and delay viral rebound in ART-treated, SIV-infected macaques
 Combined and IFNα treatment limits inflammation and delay viral rebound in ART-treated, SIV-infected macaques Mirko Paiardini Associate Professor, Emory University School of Medicine Yerkes National Primate
Combined and IFNα treatment limits inflammation and delay viral rebound in ART-treated, SIV-infected macaques Mirko Paiardini Associate Professor, Emory University School of Medicine Yerkes National Primate
Nature Medicine: doi: /nm.4411
 Supplementary Figure 1. Plasma viral load decay characteristics in each of 4 monkey cohorts studied after prolonged ART. A. Decay in 5 SIV infected animals treated ART for 20-22 weeks. B, C. Decay in additional
Supplementary Figure 1. Plasma viral load decay characteristics in each of 4 monkey cohorts studied after prolonged ART. A. Decay in 5 SIV infected animals treated ART for 20-22 weeks. B, C. Decay in additional
Supplementary Figure S1. PTPN2 levels are not altered in proliferating CD8+ T cells. Lymph node (LN) CD8+ T cells from C57BL/6 mice were stained with
 Supplementary Figure S1. PTPN2 levels are not altered in proliferating CD8+ T cells. Lymph node (LN) CD8+ T cells from C57BL/6 mice were stained with CFSE and stimulated with plate-bound α-cd3ε (10µg/ml)
Supplementary Figure S1. PTPN2 levels are not altered in proliferating CD8+ T cells. Lymph node (LN) CD8+ T cells from C57BL/6 mice were stained with CFSE and stimulated with plate-bound α-cd3ε (10µg/ml)
The Potential For Microbiome Modification In Critical Illness. Deborah Cook
 The Potential For Microbiome Modification In Critical Illness Deborah Cook To review Objectives The microbiome & concepts about its modification during critical illness Interventions Predisposition to
The Potential For Microbiome Modification In Critical Illness Deborah Cook To review Objectives The microbiome & concepts about its modification during critical illness Interventions Predisposition to
SUPPLEMENTARY INFORMATION. Divergent TLR7/9 signaling and type I interferon production distinguish
 SUPPLEMENTARY INFOATION Divergent TLR7/9 signaling and type I interferon production distinguish pathogenic and non-pathogenic AIDS-virus infections Judith N. Mandl, Ashley P. Barry, Thomas H. Vanderford,
SUPPLEMENTARY INFOATION Divergent TLR7/9 signaling and type I interferon production distinguish pathogenic and non-pathogenic AIDS-virus infections Judith N. Mandl, Ashley P. Barry, Thomas H. Vanderford,
Table S1. Primers and PCR protocols for mutation screening of MN1, NF2, KREMEN1 and ZNRF3.
 Table S1. Primers and PCR protocols for mutation screening of MN1, NF2, KREMEN1 and ZNRF3. MN1 (Accession No. NM_002430) MN1-1514F 5 -GGCTGTCATGCCCTATTGAT Exon 1 MN1-1882R 5 -CTGGTGGGGATGATGACTTC Exon
Table S1. Primers and PCR protocols for mutation screening of MN1, NF2, KREMEN1 and ZNRF3. MN1 (Accession No. NM_002430) MN1-1514F 5 -GGCTGTCATGCCCTATTGAT Exon 1 MN1-1882R 5 -CTGGTGGGGATGATGACTTC Exon
Product # Kit Components
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Pneumocystis jirovecii PCR Kit Product # 42820 Product Insert Background Information
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Pneumocystis jirovecii PCR Kit Product # 42820 Product Insert Background Information
Gut Reaction. Mary ET Boyle, Ph. D. Department of Cognitive Science UCSD
 Gut Reaction Mary ET Boyle, Ph. D. Department of Cognitive Science UCSD Ley, R. et al (2005) PNAS vol. 102 no. 31 Bacterial diversity in the distal gut (ceca) of C57BL6 mice. (A) Phylogenetic tree of
Gut Reaction Mary ET Boyle, Ph. D. Department of Cognitive Science UCSD Ley, R. et al (2005) PNAS vol. 102 no. 31 Bacterial diversity in the distal gut (ceca) of C57BL6 mice. (A) Phylogenetic tree of
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis
 APPLICATION NOTE Cell-Free DNA Isolation Kit A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis Abstract Circulating cell-free DNA (cfdna) has been shown
APPLICATION NOTE Cell-Free DNA Isolation Kit A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis Abstract Circulating cell-free DNA (cfdna) has been shown
Supporting Information Table of Contents
 Supporting Information Table of Contents Supporting Information Figure 1 Page 2 Supporting Information Figure 2 Page 4 Supporting Information Figure 3 Page 5 Supporting Information Figure 4 Page 6 Supporting
Supporting Information Table of Contents Supporting Information Figure 1 Page 2 Supporting Information Figure 2 Page 4 Supporting Information Figure 3 Page 5 Supporting Information Figure 4 Page 6 Supporting
Differences in Vaginal Bacterial Communities of Women in North America: Implications for disease diagnosis and prevention
 Differences in Vaginal Bacterial Communities of Women in North America: Implications for disease diagnosis and prevention Larry J. Forney, Pawel Gajer, Christopher J. Williams, Maria G. Schneider, Stacey
Differences in Vaginal Bacterial Communities of Women in North America: Implications for disease diagnosis and prevention Larry J. Forney, Pawel Gajer, Christopher J. Williams, Maria G. Schneider, Stacey
Supplementary Figure 1. Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
 Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region
 Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
OVERVIEW OF CURRENT IDENTIFICATION SYSTEMS AND DATABASES
 OVERVIEW OF CURRENT IDENTIFICATION SYSTEMS AND DATABASES EVERY STEP OF THE WAY 1 EVERY STEP OF THE WAY MICROBIAL IDENTIFICATION METHODS DNA RNA Genotypic Sequencing of ribosomal RNA regions of bacteria
OVERVIEW OF CURRENT IDENTIFICATION SYSTEMS AND DATABASES EVERY STEP OF THE WAY 1 EVERY STEP OF THE WAY MICROBIAL IDENTIFICATION METHODS DNA RNA Genotypic Sequencing of ribosomal RNA regions of bacteria
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research
 Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Supplementary Information
 Supplementary Information Detection and differential diagnosis of colon cancer by a cumulative analysis of promoter methylation Qiong Yang 1,3*, Ying Dong 2,3, Wei Wu 1, Chunlei Zhu 1, Hui Chong 1, Jiangyang
Supplementary Information Detection and differential diagnosis of colon cancer by a cumulative analysis of promoter methylation Qiong Yang 1,3*, Ying Dong 2,3, Wei Wu 1, Chunlei Zhu 1, Hui Chong 1, Jiangyang
Nature Immunology: doi: /ni Supplementary Figure 1. Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice.
 Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
COPD lungs show an attached stratified mucus layer that separate. bacteria from the epithelial cells resembling the protective colonic
 COPD lungs show an attached stratified mucus layer that separate bacteria from the epithelial cells resembling the protective colonic mucus SUPPLEMENTARY TABLES AND FIGURES Tables S1 S8, page 1 and separate
COPD lungs show an attached stratified mucus layer that separate bacteria from the epithelial cells resembling the protective colonic mucus SUPPLEMENTARY TABLES AND FIGURES Tables S1 S8, page 1 and separate
Supplementary Figure 1. Schematic diagram of o2n-seq. Double-stranded DNA was sheared, end-repaired, and underwent A-tailing by standard protocols.
 Supplementary Figure 1. Schematic diagram of o2n-seq. Double-stranded DNA was sheared, end-repaired, and underwent A-tailing by standard protocols. A-tailed DNA was ligated to T-tailed dutp adapters, circularized
Supplementary Figure 1. Schematic diagram of o2n-seq. Double-stranded DNA was sheared, end-repaired, and underwent A-tailing by standard protocols. A-tailed DNA was ligated to T-tailed dutp adapters, circularized
Diversity and Tropism of HIV-1 Plasma Rebound Virus after Treatment Discontinuation
 Diversity and Tropism of HIV-1 Plasma Rebound Virus after Treatment Discontinuation By: Blake M. Hauser Senior Honors Thesis Department of Biology College of Arts and Sciences The University of North Carolina
Diversity and Tropism of HIV-1 Plasma Rebound Virus after Treatment Discontinuation By: Blake M. Hauser Senior Honors Thesis Department of Biology College of Arts and Sciences The University of North Carolina
JAK3 inhibitor administration in vivo in chronically SIV Infected Rhesus Macaques
 JAK3 inhibitor administration in vivo in chronically SIV Infected Rhesus Macaques preferentially depletes NK Cells Ladawan Kowawisatsut 1 Kovit Pattanapanyasat 1 Aftab A. Ansari 2 1 Department of Immunology
JAK3 inhibitor administration in vivo in chronically SIV Infected Rhesus Macaques preferentially depletes NK Cells Ladawan Kowawisatsut 1 Kovit Pattanapanyasat 1 Aftab A. Ansari 2 1 Department of Immunology
Supplementary Figure 1. Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Nature Immunology: doi: /ni.
 Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Abstract. Optimization strategy of Copy Number Variant calling using Multiplicom solutions APPLICATION NOTE. Introduction
 Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Antibodies: LB1 buffer For 50 ml For 10ml For 30 ml Final 1 M HEPES, ph 2.5 ml 0.5 ml 1.5 ml 50mM. 5 M NaCl 1.4 ml 280 µl 0.
 Experiment: Date: Tissue: Purpose: ChIP-Seq Antibodies: 11x cross-link buffer: Regent Stock Solution Final Vol for 10 ml of 11xstock concentration 5 M NaCl 0.1M 0.2 ml 0.5 M EDTA 1 mm 20 ul 0.5 M EGTA,
Experiment: Date: Tissue: Purpose: ChIP-Seq Antibodies: 11x cross-link buffer: Regent Stock Solution Final Vol for 10 ml of 11xstock concentration 5 M NaCl 0.1M 0.2 ml 0.5 M EDTA 1 mm 20 ul 0.5 M EGTA,
Phosphate buffered saline (PBS) for washing the cells TE buffer (nuclease-free) ph 7.5 for use with the PrimePCR Reverse Transcription Control Assay
 Catalog # Description 172-5080 SingleShot Cell Lysis Kit, 100 x 50 µl reactions 172-5081 SingleShot Cell Lysis Kit, 500 x 50 µl reactions For research purposes only. Introduction The SingleShot Cell Lysis
Catalog # Description 172-5080 SingleShot Cell Lysis Kit, 100 x 50 µl reactions 172-5081 SingleShot Cell Lysis Kit, 500 x 50 µl reactions For research purposes only. Introduction The SingleShot Cell Lysis
KAPA Stranded RNA-Seq Library Preparation Kit
 KAPA Stranded RNA-Seq Illumina platforms KR0934 v1.13 This provides product information and a detailed protocol for the KAPA Stranded RNA- Seq (Illumina platforms), product codes KK8400 and KK8401. Contents
KAPA Stranded RNA-Seq Illumina platforms KR0934 v1.13 This provides product information and a detailed protocol for the KAPA Stranded RNA- Seq (Illumina platforms), product codes KK8400 and KK8401. Contents
Supplementary Table 1. Oligonucleotide primers used in quantitative and qualitative PCRs of this study.
 Supplementary Table 1. Oligonucleotide primers used in quantitative and qualitative PCRs of this study. Primer Sequence Qualitative PCR primers Universal 16S rdna forward 5 - GACGCTGAGGCATGAGAGCA-3 Universal
Supplementary Table 1. Oligonucleotide primers used in quantitative and qualitative PCRs of this study. Primer Sequence Qualitative PCR primers Universal 16S rdna forward 5 - GACGCTGAGGCATGAGAGCA-3 Universal
For the 5 GATC-overhang two-oligo adaptors set up the following reactions in 96-well plate format:
 Supplementary Protocol 1. Adaptor preparation: For the 5 GATC-overhang two-oligo adaptors set up the following reactions in 96-well plate format: Per reaction X96 10X NEBuffer 2 10 µl 10 µl x 96 5 -GATC
Supplementary Protocol 1. Adaptor preparation: For the 5 GATC-overhang two-oligo adaptors set up the following reactions in 96-well plate format: Per reaction X96 10X NEBuffer 2 10 µl 10 µl x 96 5 -GATC
Supplementary Information
 Supplementary Information Methods Growth Characterisation Total Adenosine Triphosphate (ATP) was determined using a Molecular Probes ATP Determination Kit (Life Technologies, USA). 0.5 ml of sample was
Supplementary Information Methods Growth Characterisation Total Adenosine Triphosphate (ATP) was determined using a Molecular Probes ATP Determination Kit (Life Technologies, USA). 0.5 ml of sample was
general meeting 1 20 October 2016
 general meeting 1 20 October 2016 introductions today s topics the human microbiome about the study the human microbiome the human microbiome human microbiota: the microorganisms that live within and on
general meeting 1 20 October 2016 introductions today s topics the human microbiome about the study the human microbiome the human microbiome human microbiota: the microorganisms that live within and on
HIV-1 Viral Load Real Time (RG)
 -1 Viral Load Real Time (RG) Real Time RT-PCR type 1 RNA quantification assay MSP Reg. pending Valdense 3616. 11700. Montevideo. Uruguay. phone (598) 2 336 83 01. Fax (598) 2 336 71 60. Info@atgen.com.uy
-1 Viral Load Real Time (RG) Real Time RT-PCR type 1 RNA quantification assay MSP Reg. pending Valdense 3616. 11700. Montevideo. Uruguay. phone (598) 2 336 83 01. Fax (598) 2 336 71 60. Info@atgen.com.uy
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2
 Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Supplementary Online Content
 Supplementary Online Content Fumagalli D, Venet D, Ignatiadis M, et al. RNA Sequencing to predict response to neoadjuvant anti-her2 therapy: a secondary analysis of the NeoALTTO randomized clinical trial.
Supplementary Online Content Fumagalli D, Venet D, Ignatiadis M, et al. RNA Sequencing to predict response to neoadjuvant anti-her2 therapy: a secondary analysis of the NeoALTTO randomized clinical trial.
Purification of viral nucleic acid from serum, plasma, cell-free biological fluids MACHEREY- NAGEL
 Purification of viral nucleic acid from serum, plasma, cell-free biological fluids Purification of viral nucleic acid from serum, plasma, cell-free biological fluids viral RNA: viral DNA: NucleoSpin RNA
Purification of viral nucleic acid from serum, plasma, cell-free biological fluids Purification of viral nucleic acid from serum, plasma, cell-free biological fluids viral RNA: viral DNA: NucleoSpin RNA
Supplementary Figure 1. Gating strategy and quantification of integrated HIV DNA in sorted CD4 + T-cell subsets.
 Supplementary information HIV reservoir size and persistence are driven by T-cell survival and homeostatic proliferation. Chomont, N., M. El Far, P. Ancuta, L. Trautmann, F. A. Procopio, B. Yassine-Diab,
Supplementary information HIV reservoir size and persistence are driven by T-cell survival and homeostatic proliferation. Chomont, N., M. El Far, P. Ancuta, L. Trautmann, F. A. Procopio, B. Yassine-Diab,
Figure S1. Standard curves for amino acid bioassays. (A) The standard curve for leucine concentration versus OD600 values for L. casei.
 Figure S1. Standard curves for amino acid bioassays. (A) The standard curve for leucine concentration versus OD600 values for L. casei. (B) The standard curve for lysine concentrations versus OD600 values
Figure S1. Standard curves for amino acid bioassays. (A) The standard curve for leucine concentration versus OD600 values for L. casei. (B) The standard curve for lysine concentrations versus OD600 values
Mitochondrial DNA Isolation Kit
 Mitochondrial DNA Isolation Kit Catalog Number KA0895 50 assays Version: 01 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information... 4 Materials
Mitochondrial DNA Isolation Kit Catalog Number KA0895 50 assays Version: 01 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information... 4 Materials
fl/+ KRas;Atg5 fl/+ KRas;Atg5 fl/fl KRas;Atg5 fl/fl KRas;Atg5 Supplementary Figure 1. Gene set enrichment analyses. (a) (b)
 KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
Janina A. Krumbeck 1, Heather E. Rasmussen 2, Robert W. Hutkins 1*, Jennifer Clarke 1, Krista Shawron 3, Ali Keshavarzian 3* and Jens Walter 1,4,5,6*
 Krumbeck et al. Microbiome (2018) 6:121 https://doi.org/10.1186/s40168-018-0494-4 RESEARCH Open Access Probiotic Bifidobacterium strains and galactooligosaccharides improve intestinal barrier function
Krumbeck et al. Microbiome (2018) 6:121 https://doi.org/10.1186/s40168-018-0494-4 RESEARCH Open Access Probiotic Bifidobacterium strains and galactooligosaccharides improve intestinal barrier function
PG-Seq NGS Kit for Preimplantation Genetic Screening
 Application Note: PG-Seq Validation Study PG-Seq NGS Kit for Preimplantation Genetic Screening Validation using Multi (5-10) Cells and Single Cells from euploid and aneuploid cell lines Introduction Advances
Application Note: PG-Seq Validation Study PG-Seq NGS Kit for Preimplantation Genetic Screening Validation using Multi (5-10) Cells and Single Cells from euploid and aneuploid cell lines Introduction Advances
Pyrosequencing Reveals Restricted Patterns of CD8 T Cell Escape-Associated Compensatory Mutations in Simian Immunodeficiency Virus
 JOURNAL OF VIROLOGY, Dec. 2011, p. 13088 13096 Vol. 85, No. 24 0022-538X/11/$12.00 doi:10.1128/jvi.05650-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. Pyrosequencing Reveals
JOURNAL OF VIROLOGY, Dec. 2011, p. 13088 13096 Vol. 85, No. 24 0022-538X/11/$12.00 doi:10.1128/jvi.05650-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. Pyrosequencing Reveals
Cytochalasins from an Australian marine sediment-derived Phomopsis sp. (CMB-M0042F): Acid-mediated intra-molecular cycloadditions enhance
 SUPPORTING INFORMATION Cytochalasins from an Australian marine sediment-derived Phomopsis sp. (CMB-M42F): Acid-mediated intra-molecular cycloadditions enhance chemical diversity Zhuo Shang, Ritesh Raju,
SUPPORTING INFORMATION Cytochalasins from an Australian marine sediment-derived Phomopsis sp. (CMB-M42F): Acid-mediated intra-molecular cycloadditions enhance chemical diversity Zhuo Shang, Ritesh Raju,
by Micah Johnson, Michael Turner, Elena Poiata, Stephanie Vadasz, Nathan Yardley, and Maria Moreno
 MCDB 201L: Molecular Biology Laboratory Professor: Maria Moreno By submitting this essay, I attest that it is my own work, completed in accordance with University regulations. Micah Johnson Abstract Cloning
MCDB 201L: Molecular Biology Laboratory Professor: Maria Moreno By submitting this essay, I attest that it is my own work, completed in accordance with University regulations. Micah Johnson Abstract Cloning
Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice
 Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
Supplementary Figure 1. H-PGDS deficiency does not affect GI tract functions and anaphylactic reaction. (a) Representative pictures of H&E-stained
 1 2 3 4 5 6 7 8 9 10 11 Supplementary Figure 1. H-PGDS deficiency does not affect GI tract functions and anaphylactic reaction. (a) Representative pictures of H&E-stained jejunum sections ( 200 magnification;
1 2 3 4 5 6 7 8 9 10 11 Supplementary Figure 1. H-PGDS deficiency does not affect GI tract functions and anaphylactic reaction. (a) Representative pictures of H&E-stained jejunum sections ( 200 magnification;
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Online Appendix Material and Methods: Pancreatic RNA isolation and quantitative real-time (q)rt-pcr. Mice were fasted overnight and killed 1 hour (h)
 Online Appendix Material and Methods: Pancreatic RNA isolation and quantitative real-time (q)rt-pcr. Mice were fasted overnight and killed 1 hour (h) after feeding. A small slice (~5-1 mm 3 ) was taken
Online Appendix Material and Methods: Pancreatic RNA isolation and quantitative real-time (q)rt-pcr. Mice were fasted overnight and killed 1 hour (h) after feeding. A small slice (~5-1 mm 3 ) was taken
Detection of low-frequent mitochondrial DNA variants using SMRT sequencing
 Detection of low-frequent mitochondrial DNA variants using SMRT sequencing Marjolein J.A. Weerts SMRT Leiden 2018 June 13 Content Mitochondrial DNA & liquid biopsy in oncology Pitfalls when studying human
Detection of low-frequent mitochondrial DNA variants using SMRT sequencing Marjolein J.A. Weerts SMRT Leiden 2018 June 13 Content Mitochondrial DNA & liquid biopsy in oncology Pitfalls when studying human
HIV-1 Genemer Detection Kit Ready to Use Amplification Kit for HIV-1 Specific DNA Fragment Analysis
 Product Manual HIV-1 Genemer Detection Kit Ready to Use Amplification Kit for HIV-1 Specific DNA Fragment Analysis For research use only. Not for use in diagnostic procedures for clinical purposes Catalog
Product Manual HIV-1 Genemer Detection Kit Ready to Use Amplification Kit for HIV-1 Specific DNA Fragment Analysis For research use only. Not for use in diagnostic procedures for clinical purposes Catalog
Technical Bulletin No. 161
 CPAL Central Pennsylvania Alliance Laboratory Technical Bulletin No. 161 cobas 6800 HIV-1 Viral Load Assay - New Platform - June 1, 2017 Contact: Heather Habig, MLS (ASCP) CM, MB CM, 717-851-1422 Operations
CPAL Central Pennsylvania Alliance Laboratory Technical Bulletin No. 161 cobas 6800 HIV-1 Viral Load Assay - New Platform - June 1, 2017 Contact: Heather Habig, MLS (ASCP) CM, MB CM, 717-851-1422 Operations
For purification of viral DNA and RNA from a wide range of sample materials
 QIAamp virus kits For purification of viral DNA and RNA from a wide range of sample materials Automatable on QIAGEN s proven QIAamp Kits set the standard for purification of viral DNA and RNA. QIAamp virus
QIAamp virus kits For purification of viral DNA and RNA from a wide range of sample materials Automatable on QIAGEN s proven QIAamp Kits set the standard for purification of viral DNA and RNA. QIAamp virus
Kit Components Product # EP42720 (24 preps) MDx 2X PCR Master Mix 350 µl Cryptococcus neoformans Primer Mix 70 µl Cryptococcus neoformans Positive
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Cryptococcus neoformans End-Point PCR Kit Product# EP42720 Product
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Cryptococcus neoformans End-Point PCR Kit Product# EP42720 Product
Diversified pattern of the human colorectal cancer microbiome
 Geng et al. Gut Pathogens 2013, 5:2 SHORT REPORT Open Access Diversified pattern of the human colorectal cancer microbiome Jiawei Geng 1,2, Hong Fan 1, Xiaodan Tang 1, Huiqin Zhai 1 and Zhigang Zhang 2*
Geng et al. Gut Pathogens 2013, 5:2 SHORT REPORT Open Access Diversified pattern of the human colorectal cancer microbiome Jiawei Geng 1,2, Hong Fan 1, Xiaodan Tang 1, Huiqin Zhai 1 and Zhigang Zhang 2*
New technologies reaching the clinic
 New technologies reaching the clinic Martin Däumer May 31, 2018 Deep-sequencing Standard Sanger-sequencing...PQIYMDDHTRE... Ultra-deep-sequencing...PQIYMDDHTRE......PQIYMDDHTRE......PQIYVDDHTRE......PQIYMDDHTRE......PQIYMDDHTRE......PQIYMDDHTRE...
New technologies reaching the clinic Martin Däumer May 31, 2018 Deep-sequencing Standard Sanger-sequencing...PQIYMDDHTRE... Ultra-deep-sequencing...PQIYMDDHTRE......PQIYMDDHTRE......PQIYVDDHTRE......PQIYMDDHTRE......PQIYMDDHTRE......PQIYMDDHTRE...
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A.
 Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
Effects of probiotics in the treatment of alcoholic hepatitis: randomized controlled multicenter study
 Effects of probiotics in the treatment of alcoholic hepatitis: randomized controlled multicenter study Lactobacillus subtilis/streptococcus faecium Lactobacillus rhamnosus R0011/acidophilus R0052 Ki Tae
Effects of probiotics in the treatment of alcoholic hepatitis: randomized controlled multicenter study Lactobacillus subtilis/streptococcus faecium Lactobacillus rhamnosus R0011/acidophilus R0052 Ki Tae
High-throughput transcriptome sequencing
 High-throughput transcriptome sequencing Erik Kristiansson (erik.kristiansson@zool.gu.se) Department of Zoology Department of Neuroscience and Physiology University of Gothenburg, Sweden Outline Genome
High-throughput transcriptome sequencing Erik Kristiansson (erik.kristiansson@zool.gu.se) Department of Zoology Department of Neuroscience and Physiology University of Gothenburg, Sweden Outline Genome
Supplementary Materials for
 www.sciencetranslationalmedicine.org/cgi/content/full/8/352/352ra110/dc1 Supplementary Materials for Spatially selective depletion of tumor-associated regulatory T cells with near-infrared photoimmunotherapy
www.sciencetranslationalmedicine.org/cgi/content/full/8/352/352ra110/dc1 Supplementary Materials for Spatially selective depletion of tumor-associated regulatory T cells with near-infrared photoimmunotherapy
Cell counting (EtBr) Before cell-lysis. Cell-lysis by 3% SDS beads beating. After cell-lysis
 Key words Sample Cell counting (EtBr) DNA extraction Cell-lysis by 3% SDS beads beating Before cell-lysis After cell-lysis Cell-lysis efficiency (Maintained70%) PCR PCR amplification of partial fragments
Key words Sample Cell counting (EtBr) DNA extraction Cell-lysis by 3% SDS beads beating Before cell-lysis After cell-lysis Cell-lysis efficiency (Maintained70%) PCR PCR amplification of partial fragments
The enteric microbiota: Implications for IBD. Eugene B. Chang, M.D. University of Chicago
 The enteric microbiota: Implications for IBD Eugene B. Chang, M.D. University of Chicago On a per cell basis, humans are mostly prokaryote 100 90 80 70 60 50 40 30 20 10 0 EuK ProK The microbial flora
The enteric microbiota: Implications for IBD Eugene B. Chang, M.D. University of Chicago On a per cell basis, humans are mostly prokaryote 100 90 80 70 60 50 40 30 20 10 0 EuK ProK The microbial flora
Supplementary Figure 1. Characterization of basophils after reconstitution of SCID mice
 Supplementary figure legends Supplementary Figure 1. Characterization of after reconstitution of SCID mice with CD4 + CD62L + T cells. (A-C) SCID mice (n = 6 / group) were reconstituted with 2 x 1 6 CD4
Supplementary figure legends Supplementary Figure 1. Characterization of after reconstitution of SCID mice with CD4 + CD62L + T cells. (A-C) SCID mice (n = 6 / group) were reconstituted with 2 x 1 6 CD4
Human Immunodeficiency Virus-1 (HIV-1) Genemer. Primer Pair for amplification of HIV-1 Specific DNA Fragment
 Product Manual Human Immunodeficiency Virus-1 (HIV-1) Genemer Primer Pair for amplification of HIV-1 Specific DNA Fragment Catalog No.: 60-2002-10 Store at 20 o C For research use only. Not for use in
Product Manual Human Immunodeficiency Virus-1 (HIV-1) Genemer Primer Pair for amplification of HIV-1 Specific DNA Fragment Catalog No.: 60-2002-10 Store at 20 o C For research use only. Not for use in
iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers).
 Supplementary Materials Supplementary Methods iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers). IDH1 R132H and R132L Forward:
Supplementary Materials Supplementary Methods iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers). IDH1 R132H and R132L Forward:
HIV Reservoirs in Developing Countries: Implication for HIV CURE Strategies
 HIV Reservoirs in Developing Countries: Implication for HIV CURE Strategies 10 th INTEREST CONFERENCE 2016, 3-6 TH May 2016, Yaoundé, Cameroon Cissy Kityo 1 ; Thomas Schacker 2 ; Francis Ssali 1 ; Jeffrey
HIV Reservoirs in Developing Countries: Implication for HIV CURE Strategies 10 th INTEREST CONFERENCE 2016, 3-6 TH May 2016, Yaoundé, Cameroon Cissy Kityo 1 ; Thomas Schacker 2 ; Francis Ssali 1 ; Jeffrey
Impact of antiretroviral therapy on gut immunology and the HIV reservoir in elite controllers
 Impact of antiretroviral therapy on gut immunology and the HIV reservoir in elite controllers Connie J Kim (Abstract #185) Session: Building Better Therapeutics November 19, 2013 10:00am HIV Elite Controllers
Impact of antiretroviral therapy on gut immunology and the HIV reservoir in elite controllers Connie J Kim (Abstract #185) Session: Building Better Therapeutics November 19, 2013 10:00am HIV Elite Controllers
Current Concepts in VAP: Stress Ulcer Prophylaxis & Probiotics. Deborah Cook
 Current Concepts in VAP: Stress Ulcer Prophylaxis & Probiotics Deborah Cook Objectives VAP The Old: Gastropulmonary route of infection The New: Microbiome modification Role of acid suppression Influence
Current Concepts in VAP: Stress Ulcer Prophylaxis & Probiotics Deborah Cook Objectives VAP The Old: Gastropulmonary route of infection The New: Microbiome modification Role of acid suppression Influence
FIG S1 Replication rates of S. suis strain 10, 10Δsly, and 10cpsΔEF on mono- and virus pre-infected porcine PCLS.
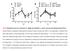 A strain 10 10cps EF B strain 10 H1N1 + strain 10 10cps EF H1N1 + 10cps EF 10 8 10 sly 10 7 H3N2 + strain 10 H3N2 + 10cps EF CFU/ml media 10 7 10 6 10 5 10 4 CFU/ml media 10 6 10 5 10 4 10 3 0 2 4 8 12
A strain 10 10cps EF B strain 10 H1N1 + strain 10 10cps EF H1N1 + 10cps EF 10 8 10 sly 10 7 H3N2 + strain 10 H3N2 + 10cps EF CFU/ml media 10 7 10 6 10 5 10 4 CFU/ml media 10 6 10 5 10 4 10 3 0 2 4 8 12
WHO Prequalification of Diagnostics Programme PUBLIC REPORT. Product: VERSANT HIV-1 RNA 1.0 Assay (kpcr) Number: PQDx
 WHO Prequalification of Diagnostics Programme PUBLIC REPORT Product: VERSANT HIV-1 RNA 1.0 Assay (kpcr) Number: PQDx 0115-041-00 Abstract The VERSANT HIV-1 RNA 1.0 Assay (kpcr) with product codes 10375763,
WHO Prequalification of Diagnostics Programme PUBLIC REPORT Product: VERSANT HIV-1 RNA 1.0 Assay (kpcr) Number: PQDx 0115-041-00 Abstract The VERSANT HIV-1 RNA 1.0 Assay (kpcr) with product codes 10375763,
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Randomized pilot trial of a synbiotic dietary supplement in chronic HIV-1 infection
 Schunter et al. BMC Complementary and Alternative Medicine 212, 12:84 CLINICAL RESEARCH Randomized pilot trial of a synbiotic dietary supplement in chronic HIV-1 infection Open Access Marco Schunter 1,
Schunter et al. BMC Complementary and Alternative Medicine 212, 12:84 CLINICAL RESEARCH Randomized pilot trial of a synbiotic dietary supplement in chronic HIV-1 infection Open Access Marco Schunter 1,
Analysis of the effect of probiotics on shaping human gut microbiota
 Analysis of the effect of probiotics on shaping human gut microbiota Masahira HATTORI Center for Omics and Bioinformatics, The University of Tokyo http://www.cb.k.u-tokyo.ac.jp/hattorilab
Analysis of the effect of probiotics on shaping human gut microbiota Masahira HATTORI Center for Omics and Bioinformatics, The University of Tokyo http://www.cb.k.u-tokyo.ac.jp/hattorilab
Supplementary Figure 1. mrna targets were found in exosomes and absent in free-floating supernatant. Serum exosomes and exosome-free supernatant were
 Supplementary Figure 1. mrna targets were found in exosomes and absent in free-floating supernatant. Serum exosomes and exosome-free supernatant were separated via ultracentrifugation and lysed to analyze
Supplementary Figure 1. mrna targets were found in exosomes and absent in free-floating supernatant. Serum exosomes and exosome-free supernatant were separated via ultracentrifugation and lysed to analyze
For Research Use Only Ver
 INSTRUCTION MANUAL Quick-RNA Miniprep Kit Catalog Nos. R1054 & R1055 Highlights High-quality total RNA (including small RNAs) from a wide range of samples. You can opt to isolate small and large RNAs in
INSTRUCTION MANUAL Quick-RNA Miniprep Kit Catalog Nos. R1054 & R1055 Highlights High-quality total RNA (including small RNAs) from a wide range of samples. You can opt to isolate small and large RNAs in
For Research Use Only Ver
 INSTRUCTION MANUAL Quick-RNA Miniprep Kit Catalog Nos. R1054 & R1055 Highlights High-quality total RNA (including small RNAs) from a wide range of samples. You can opt to isolate small and large RNAs in
INSTRUCTION MANUAL Quick-RNA Miniprep Kit Catalog Nos. R1054 & R1055 Highlights High-quality total RNA (including small RNAs) from a wide range of samples. You can opt to isolate small and large RNAs in
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Norgen s HIV proviral DNA PCR Kit was developed and validated to be used with the following PCR instruments: Qiagen Rotor-Gene Q BioRad icycler
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product # 33840 Product Insert Background Information
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product # 33840 Product Insert Background Information
Benakis et al. Supplementary Figure 1
 Benakis et al. Supplementary Figure a flora Naive C7BL/ d AC weeks d S rrna sequencing Antibiotic in drinking water Stool pellet collection riginal seeder flora flora Naive C7BL/ d AC weeks d S rrna sequencing
Benakis et al. Supplementary Figure a flora Naive C7BL/ d AC weeks d S rrna sequencing Antibiotic in drinking water Stool pellet collection riginal seeder flora flora Naive C7BL/ d AC weeks d S rrna sequencing
Norgen s HIV Proviral DNA PCR Kit was developed and validated to be used with the following PCR instruments: Qiagen Rotor-Gene Q BioRad T1000 Cycler
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product# 33840 Product Insert Intended
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product# 33840 Product Insert Intended
HIV-1 Dual Infection and Neurocognitive Impairment
 HIV-1 Dual Infection and Neurocognitive Impairment Gabriel Wagner, MD Assistant Professor of Medicine Infectious Diseases & Global Public Health UC San Diego HIV-Associated End Organ Damage Antiretroviral
HIV-1 Dual Infection and Neurocognitive Impairment Gabriel Wagner, MD Assistant Professor of Medicine Infectious Diseases & Global Public Health UC San Diego HIV-Associated End Organ Damage Antiretroviral
by nexttec TM 1 -Step
 Protocol DNA Isolation from Bacteria by nexttec TM 1 -Step - nexttec cleancolumns - Cat. No. 20N.010 Cat. No. 20N.050 Cat. No. 20N.250 Version 1.0 For research only Principle nexttec 1 -Step is the easiest
Protocol DNA Isolation from Bacteria by nexttec TM 1 -Step - nexttec cleancolumns - Cat. No. 20N.010 Cat. No. 20N.050 Cat. No. 20N.250 Version 1.0 For research only Principle nexttec 1 -Step is the easiest
Hepatitis B Virus Genemer
 Product Manual Hepatitis B Virus Genemer Primer Pair for amplification of HBV Viral Specific Fragment Catalog No.: 60-2007-10 Store at 20 o C For research use only. Not for use in diagnostic procedures
Product Manual Hepatitis B Virus Genemer Primer Pair for amplification of HBV Viral Specific Fragment Catalog No.: 60-2007-10 Store at 20 o C For research use only. Not for use in diagnostic procedures
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/3/114/ra23/dc1 Supplementary Materials for Regulation of Zap70 Expression During Thymocyte Development Enables Temporal Separation of CD4 and CD8 Repertoire Selection
www.sciencesignaling.org/cgi/content/full/3/114/ra23/dc1 Supplementary Materials for Regulation of Zap70 Expression During Thymocyte Development Enables Temporal Separation of CD4 and CD8 Repertoire Selection
Supplementary Information
 Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
