10802) hypothetical protein
|
|
|
- Paul Whitehead
- 5 years ago
- Views:
Transcription
1 upplementary Table 1. Annotation of the divamide A biosynthetic cluster. CDs of the div A biosynthetic gene cluster (GenBank: KY11568) from the E11-36 metagenome listed alongside their most similar BLAT search results. Conserved protein domains and biochemical characterizations are included. Individual accession numbers are listed for gene products (ND=data not deposited). Gene Protein accession number Length (bp) Closest BLAT hit ID -2 ND 369 WP_ ND 321 XP_ diva ARD WP_ divm ARD WP_ divx ARD WP_ divy ARD WP_ divmt ARD WP_ divt ARD WP_ divu ARD WP_ divn ARD WP_ ND 15 WP_ ND 195 WP_ Annotated function (Organism) hypothetical protein (Kamptonema) uncharacterized protein LOC (Rhinopithecus roxellana) hypothetical protein (Oscillatoria sp. PCC 182) hypothetical protein (Oscillatoria sp. PCC 182) hypothetical protein (Oscillatoria sp. PCC 182) hypothetical protein (Tolypothrix bouteillei) AM-dependent methyltransferase (tanieria cyanosphaera) ABC transporter permease and ATPase (Moorea producens) hypothetical protein (cytonema millei) hypothetical protein (Oscillatoria sp. PCC 182) phosphopeptidebinding protein (Pleurocapsa minor) serine/threonine protein phosphatase (Microcoleus sp. PCC 7113) Conserved protein domains (accession) none none none LanM-like (cd4792) Nif11 (pfam7862) Het-C (pfam7217) Methyltransf_18 (pfam12847) ABC_membrane_2 (pfam6472), ABC_tran (pfam5) FGE-sulfatase (pfam3781) none CHAT (pfam1277), FHA (pfam498) PP2Cc (cd143) Functional characterization not characterized not characterized precursor gene class II lanthionine synthetase putative Asp ß- hydroxylase (16) not characterized methyltransferase not characterized not characterized putative role in Lal formation (14-16) not characterized not characterized Nature Chemical Biology: doi:1.138/nchembio.2537
2 upplementary Table 2. Relationship of divamide B cluster genes to divamide A cluster. CDs of the partial div B biosynthetic gene cluster (GenBank: KY11569) from the E11-37 metagenome compared with their respective div A homologues. Individual accession numbers are listed for gene products (ND=data not deposited). Gene ID Protein accession Coverage with ID to corresponding Length (bp) number corresponding div A gene div A gene -1 ND 286 (partial) 88 1 diva ARD divm ARD (partial) 3 1 divx ARD (partial) 66 1 divy ARD divmt ARD (partial) Nature Chemical Biology: doi:1.138/nchembio.2537
3 upplementary Table 3. ynthetic DNA used in this study. DNA gblocks purchased from Integrated DNA Technologies used to construct pdiv and pdiv-3. DNA equence (5-3 ) 1a 1b 2 3 divadivamide B ctcgtatgttgtgtggaattgtgagcggataacaatttcacacaggaaacaatgccgaccaccctggaaaaaccgagcgtggcctatctggagaaact GTTTCACCAGACCGCCATCGACAGCGAATTTCGTAGCGAACTGCAGAGCCATCCGGAAGCCTTTGGTATTAGCGCCGATCTGGAACTGCCGCAGAGCG TGGAGAAACAGGACGAAAGCTTCGTGGAACTGCTGAACAACGCCCTGGGCGAAATCGATATTGCCGCCGAATGCGCCAGCACCTGTAGCTTTGGCATC GTGACCATCGTGTGCGATGGTACCACCAAATAAGTTGTTTTACTAGCATATTATATTATTAATATGTCTGGGCTAGCACCACATTGTGAAGCTAGCCT CATTAAATTGATTGTTGGGCATGGTTAGAAAGGAACTAAGTACAAGACAAAACTTGGATGAGAAGCGTAGTAAAATGTCCGAAACTATTGGCTCATAA AGTTTCGAGCAATATTTTCCATCTCTCTTGCTCCATATGAGTTTCAGTTGTTTCAGCTG AATCCGGAGACCAAGGAACTGGTGACCCTGGCCGAGAAACCGAAAATCGTGCGCTTCTTCAAAATCGAAAGCGAGCCGAGCAAAATTCTGTGGCACTG CATTGTTAACACCACCAAGCTGAATCGCTGGAGCTTCGGTTTCAACGTGAGCAGCAAGTGGATGGACAAACTGCTGACAGTGTAAGGAGAATAAAATG AAAATTTTCCTGACATGCCTGCTGGCCCTGGCCCTGCTGTTAGGTATGCCGAGCAGTGCCTTTGCCTTCAAAGTGCCGATCCACGAAGAGATCACCCG CGAAGTGTTTGAGGATTTCCAGGTGGTGGTGGAAGGCGAGACCTTTAAGTTCACCGACTATGCCATCGACCAGATCGTGAAGGCCAACAAGGATACCG ATGACCTGCCGAACCAGTTCAATACCGAGATGCACTTCGACGGCGAAGACTTTAGCGGTGGCAGCAATCGTGTGATGTTCCTGAAGGAGCGCACCATT ACCAAAGTGACCGACCCTCAGGATCCGCAGGGCACAAGCGCACGTAATGATCTGGGTACCGCCCTGCATACCGTGCAGGACTTCTATGCCCATAGCAA TTGGGTGGAACTGGGTCACAGCAGCAGCGATATCAACACCAAGATCGGCCGCGAGGTGTTCAGCGGCGCCGATAAAAACACCGCCACCTGCCCGAACG ATCCGGGCATTCTGGGTGGTGCCGGCCTGACCGAACTGACCAGTGGCTACTTCACCTTCATCGGCGTTGTGCCTAGCTGCGATGTTCCTGAGGGTAAA TGCCGCCATGGCGTTCCGATTGTGTGCCCGGATGGCCTGAACAAGGACGATAATAGTCGCCCGGGTTTTCCGACCGCCCGTGCATTAGCCGTTAAAGC CACCGAGGACTTCCTGAATCAGATCTTCAGCGACAGCCGTATGGACGGCAACGTGGATGCCATCAAACTGCTGATGCGCATCCGTAATTAATCCCTTT AACAAGTAAATTGTACCTCGCCTAAACCATACCCGACAGGATTGGTTTAGGCTTATCTGCTGCTTTATAAAAGACAGATAGAGATTAGAATTATCGCA ATGGAAAATAATAATTATCCGTTCGAACTGAAAGCCTATGACTTTAGCTTTAAAATCTTCAAGGAGATCATTTTCGCCCTGAACGCCTTCTTCATTGG CTTTTGGCTGGGTGTGCTGAAACGCGAACATTACCACCTGGTGGATAGCATTTACTACAACCAGACCGAAATGTACCGCGACGAGAACTACAACAAAC GCGGCCTGTGGGACTGGGAAGAGAAAGTTCTGGCCCAGTACTTCCAGCAGTGCCATAATCTGCTGGTGGTTGCAGCAGGCGGTGGTCGTGAGGTTCTG GCCTTATGCAAACGCGGCTATGAAGTGGACGGCTTTGAGTGCAATGCCAACCTGCTGAAATTCGCCAACAACCTGATCAAGCAAGAGGAGTTTGCCAG CCATATCAAACTGGCACCGCGTGATCAGTGCCCGGATAGCCAGAAGGAGTATGATGGCCTGATCGTTGGCTGGGGTGCCTATATGCTGATCCAGGGTA AGGAACGCCGCATTGAATTCCTGCGTCAGCTGCGTACCCAGGCCAAGAAGAACAGTCCGGTGCTGCTGAGCTTCTTCTGCTACAGCGAGACCACCGGC GGCCGTGATTTCAAAGCCATCGCCATGATCGGCAATGCCTTTCGTCGCCTGCTGGGTCGTGAATGCCTGGAAGTTGGCGATAATCTGGCCCCGAACTA CGTGCACTACTTCACCAAGGACGAGATCGCAAGCGAACTGCAGGCAGGCGGCTTTGAACTGAAGATCTACTGCACCAATCAGTATGGCCACGCCGTGG GTATCGCAGTGTAAcacctttaagcctatagtctaggaaaaagaggttgaattagagccaacttggtcatctgaaaccctcatgctgtcgtttccccc tgaacagttacgaaactacgaagatagagaagctttatgtataaaggcctggatcgcgacgaaattcgccagattcaggtgctgatgctgctgtgcct GTGCCTGAGCCCGCAGAGCAAACTGCGCCAGCTGCTGGAAATTGCACTGGCCGCCAGCGAAACCCAGATCATGACCCGCATGACACCGTGCGATGATG TGAATGTGGACGGCCTGTTTACCTGGGTGCAGAGCCTGTTTGCCCAGGGTGGTCTGACCGAAGAAGAGAAACGCCTGCTGAAGTGGCAGAACGAAAGC CGTAACATGCTGCCGGCCATTGACGAACTGAAAACCATTGAGAAGAAGCTGGGCTTCAAGATCCGCATTCAGAAGCTGCAGAGCCACAATTAAttatt ggggtgcggtcaaaaacagttgaattttaagtcgcggcttacccg ATTTTCCATCTCTCTTGCTCCATATGAGTTTCAGTTGTTTCAGCTGCACCGAGCGCAAGGAGAATAACGAACCTTGTTTATTTTTACTGTTAAAGCAA AACACCAATTTCACACTGCGCACCAGCACCATGCTGGACGACTTCAGCCTGTTAAAGTTAGCCAGTCGCGCAAGCAACCTGTGTGAGCAAACCCTGAT TGTGAAAGAGCTGGCAAAGAGCAAAGCACCGATCGCCAGCACAACACAACTGAGTCCTGTTGACAGCTGGAAAATTAAGAAACTGACAGGCAAACTGG CCGTGCAGCCGTTCAAAGAGAGCTACGAGCAGGGCACAATCAGTCAGAGCGTGATTGAGGACCTGCGTAAACTGCTGATCGATTACAAACTGTACGAA CTGAACCTGGCAAACTTAAGCGAGAGCGACCGCCTGGAATTTATTAAGCCTCACAGCCAGTGGTTAAAGGCATACCAGGCCGCCATGGCAACCCTGGA TCTGCCGCGTGAAAAGTTCAGCGGTAGTTGTTGGGGCGAACCGGATATTTACTACGGTAAATTCGCCAAAGTGTGTGAACCGTTTCTGCGCCTGCTGC ACCAAACCCTGCGCGGTACCGGCGACGCAATTAACGCCACCGCCGACAATTACCGCATTAATCCGCAGGTGGCCATCGACATCGAGCTGCATCTGCTG AACCGCTTCGAATTAGCCCTGGCCTGGGCCTTAGAGGCCAACATCAATGTTTATTGTAGTCAGAAAGCAATCGCCAAGAGCGAGGATGATAGCGAAGC CTACATCGCATACCTGGAGGAGACCTTCGACCGCAAGCAGAATTATCATGATTTTTATTGTCGTTTCCCGGTGCTGGCCCGTTGGTTAGCCCAAGTGA CCTATTTCCTGTGCAATTTTGGCGAGGAAACCCTGCAACGCCTGACCAGCGATCGTGAGCAGATTGGCGCCACCTTCTTCGGCAGCAAGCCGATCAGT CAAATCAAAAGTTTTAAACTGGGTAAAAGCGACTACCATGCCGGTGCCAAGAGCGTGGTGATCGTGGAGCTGGAGCTGGCCAACAGCGAACCTGCCAC CCTGGTGTATAAGCCTCGCAGCATTCAGAGCGAAGCCGGCATGCAGGGCCTGCTGGCACAGCTGAACCAAGATAAGGTGGTTCGCTTTGCCCACTATC AGGTGCTGTGTCGCGATGGTTATGGCTATGCAGAGTTCATCCCGAGCGGCAAAAACCAGGTTCAGAATAAAGAAGATCTGAAGAAATTCTACCAACAG CTGGGCGGCTTTTTAAGCATCTTCCATATCCTGGGTGGCGGCGATCTGCACCATGAAAACATCCTGGTTGCAGATGGTAACGCCTTCATCTGTGACTG CGAGACCGTGCTGGAGGTGCTGCCGCAAGGCATGGATAAACTGCCTGGTACAGTTTTAGATAGTGTGTTTAAAACAGCCATGCTGGACTGGCCTCGTG ATAGCGCAAGCCCGGAGAACAGCGAGATGATGAGCATTAGTGGTTACAGCGGTGGCGAAAGCTATGAGGTTGCATTCACCGTGCCGCGCGTTAAGGAG CACCGTATGAGCCTGGATCAGGGTGTTGAGTACAAGACAGGTATCACCGTGGAACTGGAAGGTACAAATCGCATTTACTACAACGGTGAGATTGTGGA TCCGCAGGACTATAAGGATAGCATTGTGGACGGTTTTAACCAGGTTTATACATGGTTTCAGCAGCACCCGACCAAGGCAATTACCCGTATTAAGGAGC TGTTTAGCAGTAGTTTAGTGCGCTTTATTAATTGGGGCACCCAAGCCTACGCCAAGAGCATTGTGGCCGTTCGTCACCCTAAGTGCCTGGCCGACCCT CTGGAGGTGGACCTGATCTTTAATAGCCTGAAAGAGCATAAACGCCAGTGGGACAAAAAGGGCGAACTGGCAGAGTTAGAGCTGGGCAGCCTGTGGCA ACTGGATATCCCGATTTTCACCGCCTTAGCCGCCGAAAGCAAAGACTTAATCTTCAATTATCAGTATAGCGTTAGTGATACCTTAGCCATCAGCCCGT TAGACAATGCCAAACGCCGTCTCGAGCAGCTGAGCACCGAGAACCG TAGACAATGCCAAACGCCGTCTCGAGCAGCTGAGCACCGAGAACCGTATCCGTCAGAATCAATACATCTATACCAGCCTGAGCACCGACGAGATCAAC AGTCCGTACTTTATCGCCGCAGCCGTTAACTATGCCCAGCAGATCGGCTGGCAGCTGTGTGAACAGCTGAGTAGCGATAGTAGCAAAGCACCTTGGCA GACATGGGACTACACCGCAACCGGCAAGCGCTTAGTGGATATCAGCGGCGACCTGTATGATGGCAGCGCCGGCATTTGTCTGTTCCTGGCCTACCTGG ATGCCATCAAACCGCAAGTGGAATTCCGTCAGGCCGCCGAACGCGCCTTAGAATACAGTATCGAGAAACGTAACACAACCCTGATCGGTGCATTCCAG GGCGAAACAGGTCTGATTTATCTGCTGACCCATCTGGCACAGTTATGGGACAAACCGGCATTACTGGACCTGGCAGTTGACCTGAGCGACGAGCTGCT GCCGCGCATCAAACAGGACATCTACTTCGATATCCTGCATGGCGTTGCCGGTATCATTCCGGTTATGCTGGGCCTGGCCGAAGCAACCGGTGGTAAAG GCATTGATTGCGCATTACAGTGTGCCGAGCACCTGCTGGAGCAGGGTATTTACCAGGATAACACCCTGAGCTGGCCTCCGGGTCGTCCTGACCTGGTG CGCGGCAATTTTACAGGCTTCAGCCACGGTGCAAGCGGCATCGGTTGGGCCTTAATTATGCTGGGCTGCCACAGCAATAAGAGCGAGTATATTGAGGC AGGCCGTCAGGGCTTTGCCTACGAAGCCACACAGTTTGATGAGGAACAGCGCGATTGGTACGACTTACGCAAGAGCGTTACCACCGCAGATAGCAACG AACCTCACTTTGCCAACGCATGGTGCAATGGTGCCGCCGGTATTGGCCTGAGTCGCATCATTAGTTGGGCCGCCCTGGGCAAAACAGACGACGACATT CTGCGCGATGCCTACACCGCACTGAATGCAACCTTACGCAATTTCAACAAGCTGGGCAACGATAGCCTGTGTCATGGCAAGAGTGGTAACGCCGAGTT ATTCCTGCGTTTTGCACAGCTGCGCGATACCCCGTATCTGCAAATGGAGGCAAACGTGCAGGCCCAGGCACAATGGCGCAACTTTGAAAAGGCACGCC GTTGGATGTGCGGTAGCACAGGCAACGATGTTTTCCCGGATTTAATGCTGGGCCTGGCCGGCATTGGCATGCACTTCCTGCGCCTGGCCTACCGTGAA CGCGTTCCGAGCCCGTTATTATTAGATCCGCCTCCGCGCGCCATCGACTAAGTTTTGAGTTTTTTAATTCAATATTCAACACCCAAGAGTAATATAGT TTAGTTTGTTTATTAGGATCGAGATGAGCAAAGAAACCGTGATTGAATTTTATGAAGCAATTTTTGAAAGTCCGGAATTCATTCAGGAAATTAAAGCC ATTACCCGCCAAGAGGAACTGATCAAACTGGGTGCCCGCAACGGCTACCATTTTACCATGGAAGACCTGGCCCAGGCCGACGCCAGCTATATCCCGAA GAATAACCAGCCGCTGATCAGCATTGATAGCGACGACCGTGCCCGCGAAGAACTGCCGCGCCCGTATCACTATGAATTCGAGTTCAGTGAGATCCCGG GCTTCGAAGAGATCGATCGTGAACTGAAAAAACTGCAGATCAAACCGAACACCGTGGATCTGGACCTGTACGAGAAGAGTTTCCGCGAAGAAGATTTT AAATTTAACTATATTAGCCCGACCGTGCCGGGCTTCCGCCAGTATTACTACAAGAGCCTGAAAAGCTATCTGGATCTGCCTAGCCCTCAGCCGGAATA TGCCTGGCGTCCGTTTCACCTGATCAATCTGGATTGCCACGTGGAGGATCCGCTGTACGAGGATTATTTCCAGACCAAGGTGCGTCTGCTGAAGCTGC TGGAGAATTGCCTGGAGACAGAGCTGCGCTTTAGTGGCAGCCTGTGGTATCCGCCGAACGCCTATCGCCTGTGGCACACCAATGAAACCCAGCCGGGC TGGCGCATGTACCTGGTGGATTTCGACAATTTTGACGACAACCAGGAAGGCGAGGTGTTCTTCCGCTACATGAATCCGGAGACCAAGGAACTGGTGAC CCTGGCCGAGAAACCGAAAAT GTGGAACTGCTGAACAACGCCCTGGGCGAAATCGATATTGCCGCCGAATGCGCGTCGACCTGTAGCAGCGGCCCGATCACCGCGATCTGCGATGGTAC CACCAAATAAGTTGTTTTACTAGCATATTATATTATTAATATGTCTGGGCTACCACCACATTGTGAAGCTAGCCTCATTAAATTGATTGTTGGGCATG GTTAG Length (bp) Nature Chemical Biology: doi:1.138/nchembio.2537
4 upplementary Table 4. DNA primers used in this study. Primers used in the construction and sequencing of pdiv, pdiv-2, and prfduet-divmt. # Primer name equence (5-3 ) 1 div1a-fwd CTCGTATGTTGTGTGGAATTGTG 2 div2-rev CGGTTCTCGGTGCTCAG 3 div3-fwd TAGACAATGCCAAACGCC 4 div1b-rev CGGGTAAGCCGCGAC 5 divm-23-fwd CGATCGCCAGCACAAC 6 divm-1424-rev CCAGTCCAGCATGGCTG 7 divm-2818-fwd GCAGGCCGTCAGGG 8 divx-65-rev AGCCCGGCTGGGTTTC 9 divx-58-fwd GAGACAGAGCTGCGCTTTAG 1 divmt-79-rev CAATGAAGAAGGCGTTCAG 11 divmt-2-fwd CGTTCGAACTGAAAGCCTATG 12 divm-23-fwd CGATCGCCAGCACAAC 13 divm-1977-f CTTAGCCGCCGAAAGC 14 divx-461-f TGCCTAGCCCTCAGCC 15 divhypo-78-f CGCATCCGTAATTAATCCC 16 intergenic-divmnative-fwd GAGCAATATTTTCCATCTCTCTTGCTCCATATG 17 intergenic-divmnative-rvs ACTCAATC GGGTGTTGAATATTGAATTAAAAAACTCAAA 18 divmp-fwd GTGGAACTGCTGAACAAC 19 divm-non-optimizedpet16b-gibson-fwd cgaaggtcgtcatatgcttgatgatttttctctcc 2 divm-native-fwd-1 CTTTAGCCTGGGCTCTTG 21 divm-native-fwd-2 GGACAGTGCTTGATTCAG 22 divm-native-rvs-4 CTTCATCAAATTGGGTCG 23 divmt-mc1-fwd TTCGAGCTCGATGGAAAATAATAATTATCCGT TCG 24 divmt-mc1-rvs TATGCGGCCGCTTACACTGCGATACCCACG Nature Chemical Biology: doi:1.138/nchembio.2537
5 upplementary Table 5. ummary of quantification of divamide material by NMR. Divamides were quantified using linear equations shown in upplementary Figure 3a. All divamide samples were quantified with (top row) and without (bottom row) inclusion of the methyl region integral because this integral accounts for a large number of protons, which may reduce the accuracy of integration. Compounds 2, 3, and 9 were quantified using integrals for only two proton signals such that numbers obtained with exclusion of methyl integrals represent a single integral. Yields were calculated from the average of the two concentrations. A known concentration of L-Tyr was used to assess the quality of the L-Trp standard curves. A difference of 7.6 was calculated for the NMR-determined concentration relative to the UV-determined concentration. UV NMR ample MW (g/mol) mg/ml mg/ml D mm D yield (µg) L-Tyr E11-36 divamide A (1) E. coli divamide A (1) desmethyl divamide A (9) divamide B (2) divamide C (3) Nature Chemical Biology: doi:1.138/nchembio.2537
6 upplementary Table 6. ummary of quantification of divamide material by LC/M. Divamides were quantified using linear equations shown in upplementary Figure 3b. For mixtures of compounds, the concentrations in mg/ml were calculated based on individual molecular weights before adding together. The concentration of E. coli divamide A determined by LC/M was in agreement with the NMR concentration ( -5.92). The solution of 4 used in flow cytometry based assays was quantified using curve ii (*). ample MW (g/mol) mm D mg/ml D yield (µg) 1 (E. coli) mix mix mix mix biotinylated divamide A mix biotinylated cinnamycin mix * Nature Chemical Biology: doi:1.138/nchembio.2537
7 upplementary Table 7. ummary of flow cytometry experiments. Flow cytometry experiments were performed in triplicate (or duplicate in several cases) for each dose of each sample. A total of four compounds were tested, with eight different doses for dose range 1 and six doses and two infection conditions, uninfected and HIV infected, for dose range 2 (see Figure 4), utilizing four 96-well plates. The total number of events are listed alongside the number of events collected within gate 1, representing live cells, and gate 2, representing living infected cells. Log concentration (viability-dapi) (p24-fitc) ample Plate Replicate ng/ml nm Total events Gate 1 events gate1/total gate2/gate1 Gate 2 events 1 (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) (E11-36) uninfected uninfected uninfected HIV HIV Nature Chemical Biology: doi:1.138/nchembio.2537
8 continued uninfected uninfected uninfected HIV HIV uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected HIV HIV HIV Nature Chemical Biology: doi:1.138/nchembio.2537
9 continued... 9 uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected uninfected HIV HIV HIV Nature Chemical Biology: doi:1.138/nchembio.2537
10 upplementary Table 8. ummary table of assay data. IC 5 (anti-hiv), CC 5 (cytotoxicity), and therapeutic index values for all compounds screened are listed. Values from both cytoprotection (Figure 3) and flow cytometry-based (Figure 4) assays are included for comparison. For some samples, the true value of these parameters could not be determined from curve fitting and was estimated based on neighboring data to lie above or below the indicated number (> or <). ome activities were not detected within the concentration range tested (ND). The therapeutic index was not calculated for data that could not be fitted to a curve (NC). Values from repeated experiments were averaged (*). Assay ample IC5 (µm) CC5 (µm) Therapeutic index ND <27. Cytoprotection 1-12 >.35 <8.42 NC ND < ND ND 4 ND >5.35 NC Flow cytometry * <14.4 NC * 6.25 Nature Chemical Biology: doi:1.138/nchembio.2537
11 upplementary Table 9. Divamide-like masses observed from tunicate extracts. Unidentified ions observed from Didemnum molle extracts by LC/M resembled those of characterized divamides in m/z and isotope distribution. ome masses appear to be derivatives of 1-3, but others may reflect alternative amino acid sequences at various stages of posttranslational modification. everal of these display evidence of Hoffmann elimination in negative ionization mode. Thus, additional, uncharacterized divamides are believed to be encoded by E Observed m/z (z = 2) ource ID 11.9 E E11-37? E11-37? 9.5 E11-37? E11-37? E11-37? E x Me E x Me E11-37? E x Me E E x O E11-37? E11-37? 11.5 E11-37? E11-37? E11-37? E11-37? E11-37? E11-37? E E x O E11-37? E11-37? E11-37? E11-37? E11-37? E11-37? E11-37? E11-37? E11-37? Nature Chemical Biology: doi:1.138/nchembio.2537
12 White3 (n=2) White4 (n=4) White1 White2 (n=4) D. psammatode.6 Large1 (n=4) Large3 (n=3) Large4 Large5 Large2 E11-36 mall1 mall2 2.6 substitution/site Brown1 (n=7) Brown3 Gray1 Gray2 (n=3) Gray4 72 Gray3 E11-37 upplementary Figure 1. Ascidian phylogeny. The phylogenetic relationship between ascidians collected in this study (shown in red) to samples collected by Hirose et al. (shown in black) was estimated by maximum likelihood analysis of Didemnum molle cytochrome oxidase subunit I (COI) gene sequences. 49 The tree was generated using RAxML software with 1 bootstrap replicates, with the resulting bootstrap probabilities shown at each node. According to Hirose et al., each of these named morphotypes (such as Large, Brown, etc.) are likely to be different species, even though they are currently all classified as D. molle. Nature Chemical Biology: doi:1.138/nchembio.2537
13 A DNA Length (kbp) i ii i = 88 Da D = 9 Da ladder P L FT W1 W2 T y y H! N!! ECATCFGIVTIVCDGTTK! m/z m/z i y12 y H H! H N! N N! ECATCFGIVTIVCDGTTK!! ECATCFGIVTIVCDGTTK ECATCFGIVTIVCDGTTK!! y12 y9 y7 y5 y2 y13 y1 y8 y6 y4 y3 1! ECATCFGIVTIVCDGTTK! E E1 E2 E3 E4 y6 y y y y9 y ! ECATCFGIVTIVCDGTTK! 1!!! H +H + N (Me3)NN (Me3)N ECATCFGIVTIVCDGTTK ECATCFGIVTIVCDGTTK m/z H! N!! ECATCFGIVTIVCDGTTK! m/z ii kda 27. C m/z DivMT purification Molecular Weight (kda) = 9 Da iii ii iii! ECATCFGIVTIVCDGTTK! i NiCl2, NaBH4 55 C, 3 min = 88 Da!! ECATCFGIVTIVCDGTTK! ECATCFGIVTIVCDGTTK! ECATCFGIVTIVCDGTTK y12 y9 y7 y5 y2!!! y13 y1 y8 y6 y4 y3!! ECATCFGIVTIVCDGTTK! ECATCFGIVTIVCDGTTK NiCl2, NaBH4, y12 y9 y7 y5 y2 y4 NiCl2, NaBH4 55 C, 3 min 1 ECATCFGIVTIVCDGTTK y13 y1 y8 y6 y4 y3 ECATCFGIVTIVCDGTTK C, 3 min! kda ladder P L FT W1 yw2 T E1 E2 E3 E4 ladder 6 y y7 ECATCFGIVTIVCDGTTK!! ECATCFGIVTIVCDGTTK y1 ECATCFGIVTIVCDGTTK y ECATCFGIVTIVCDGTTK! y y9 y1 y13 H N 42.7 H 34.6 ECATCFGIVTIVCDGTTK 86.1 N H kda NH! 27. H! ECATCFGIVTIVCDGTTK N!! N!! ECATCFGIVTIVCDGTTK 2. ECATCFGIVTIVCDGTTK! ECATCFGIVTIVCDGTTK! ii bp ! ECATCFGIVTIVCDGTTK!!!! 1 B m/z H H N N ECATCGPITAICDGTTK ECATCGPITAICDGTTK H +H + N (Me3)NN (Me 3)N ECATCGPITAICDGTTK ECATCGPITAICDGTTK m/z m/z upplementary Figure 2. Installation and detection of in vitro chemical transformations of pdiv-2 and -3 E. coli products. a, The pdiv expression vector for 1 produced no detectable compounds. The native divm gene was amplified from E11-36 metagenomic DNA by PCR resulting in a faint band (i) that was used as a template for additional PCR amplification (ii). This gene replaced the codon-optimized divm gene of pdiv to yield pdiv-2 (Figure 2a), which produced MeLan-containing products when expressed in E. coli (Figure 2b). b, Desulfurization of 5 (i) results in an 88 Da mass shift (ii) due to the reduction of three MeLan residues and one Dha residue. After base-catalyzed Lal formation, desulfurization yields a 9 Da mass shift (iii), indicating the presence of Lal. c, M/M analysis is hindered by the presence of cyclic residues like MeLan (i), while the linearized desulfurization product is fragmented into y ions (ii). After base treatment, the desulfurization product is resistant to M/M fragmentation due to the Nature Chemical Biology: doi:1.138/nchembio.2537
14 presence of Lal (iii). d, DivMT was purified on Ni-NTA resin from the cell lysate of prfduet- DivMT-expressing E. coli BL21(DE3) and column fractions analyzed by D-PAGE. P = pellet, L = filtered cell lysate (supernatant), FT = Ni-NTA column flowthrough, W = column wash with lysis buffer, T = column wash with transition buffer, E = elutions of increasing imidazole concentration in transition buffer (1 = 5 mm, 2 = 1 mm, 3 = 2 mm, 4 = 5 mm imidazole). A band around 32.6 kda, corresponding to His 6 -DivMT, was observed in E3. e, Methylation was successful for both HPLC-purified 9 (i) and an E. coli C18 fraction containing 17 (ii). Top and bottom spectra show the reaction in absence or presence of DivMT, respectively. A small amount of final product (m/z 965.7) was observed in the C18 fraction prior to DivMT methylation. Nature Chemical Biology: doi:1.138/nchembio.2537
15 A i L-Trp standard curve (! 7.62 ppm) Absolute integral y =.531x +.22 R² = Concentration (mm) ii L-Trp standard curve (! 7.17 ppm) Absolute integral y =.5358x -.13 R² = Concentration (mm) iii L-Trp standard curve (! 3.19 ppm) Absolute integral y =.529x R² = Concentration (mm) upplementary Figure 3. Quantification of divamide material. a, tandard curves for quantification of low abundance materials by NMR were generated using a dilution series of L- Trp in H 2 O, the concentration of which was determined by UV-absorbance (e 28, Trp = nm - 1 ). The absolute integral was plotted as a function of concentration using three different aromatic proton signals: H E3 (i), H Z3 (ii), and methylene H B2 (iii). The linear equations generated were used to determine yields of divamide analytes of unknown concentration in upplementary Table 5. b, tandard curves for quantification by LC/M were generated using a dilution series of synthetic 9, previously quantified by NMR. The resulting linear equations were used to determine concentrations of divamide analytes and estimate relative ratios of compound mixtures, such as those obtained from expression in E. coli, as shown in upplementary Table 6. The majority of analytes were quantified with one curve (i), while a second curve (ii) was used to quantify additional 4. Nature Chemical Biology: doi:1.138/nchembio.2537
16 B i continued Desmethyl divamide A standard curve ii Area Under Curve (AUC) Concentration (µm) Desmethyl divamide A standard curve y = x R² =.9878 Area Under Curve (AUC) y = 25.95x R² = Concentration (µm) Nature Chemical Biology: doi:1.138/nchembio.2537
17 A relative survival B R 2 =.9998 IC 5 =.292 µm CC 5 = 4.43 µm 9 R 2 =.9976 IC 5 =.327 µm CC 5 = 3.38 µm 1 (E11-36) live infected cells (p24-fitc) (E11-36) R 2 =.9987 IC 5 = 1.3 µm CC 5 = >5.35 µm Dose range 1 HIV infected 1-12 log concentration (ng/ml) R 2 =.9935 IC 5 =.874 µm CC 5 = <14.4 µm live cells (viability-dapi) Controls Uninfected AZT Infected Controls Uninfected p24 Infected p24 Uninfected viability Infected viability upplementary Figure 4. Additional cytoprotection and flow-cytometry assay doseresponse curves. a, Cytoprotection assay dose-response curves for divamide species not shown in Figure 3. Experimental controls, shown as dashed lines, include uninfected (blue), infected (red), and AZT (at either.1 or.5 mg/ml) responses. Points represent an average of three assay responses per dose (error bars = ± standard deviation). R 2 values are included for compounds for which curves could be fit with equation 7 (see online methods) or a single fourparameter IC 5 equation. Curves could not be fit for E11-36-derived 1 due to abnormal curve shape. b, Flow-cytometry dose-response curves for divamide species not shown in Figure 4. Cytotoxicity and anti-hiv activities of divamides and 4 were monitored simultaneously via flow cytometry using a fluorescein-labeled antibody to HIV-1 capsid protein p24 (green; live infected cells) and viability stain (purple; live cells) with HIV-infected CEM TART T-cells. The biological marker and corresponding fluorescence channel are shown in parentheses along the vertical axes. Points represent an average of three assay responses per dose (error bars = ± standard deviation). Average uninfected and infected cell control responses are shown as horizontal lines for each biological marker. All plots shown employed dose range 1 with HIVinfected cells, yielding full anti-hiv dose-response curves but only partial cytotoxicity curves. Nature Chemical Biology: doi:1.138/nchembio.2537
18 A 1 [M+H] 2+ = pos i 1 [M-2H] 2- = = 63 Da neg ii 1 1 pos m/z [M-2H] 2- = = neg [M+H] 2+ = = 63 Da iii m/z pos [M-2H] 2-2- = neg [M+H] = = 63 Da m/z upplementary Figure 5. Chemical diversity within the divamide family. a, Hoffman elimination of trimethylamine from a quaternary N-trimethylamine results in a net loss of 6 Da and the elimination of a positive charge. This characteristic was observed for N- trimethylglutamate-containing lanthipeptides 1 (i), 2 (ii), and 3 (iii) by LC/M in negative ionization mode. b, All divamide-like lanthipeptide gene clusters that could be identified by BLAT search of div pathway genes are shown. They occur mostly in cyanobacteria (i) and actinobacteria (ii), with one example from chloroflexi (iii). Cyanobacteria may contain multiple lanthipeptide clusters within a single genome or multiple lana precursor proteins in a single cluster, suggesting multiple products are produced from a single strain. Each pathway maintains the genes responsible for introducing conserved modifications, including Lan/MeLan and Hya. Most also contain divn homologs, but this gene does not appear to be 1 conserved across all pathways. ome cyanobacterial pathways have incorporated unique modifying enzymes. A methyltransferase bearing no resemblance to DivMT can be found in the first pathway from Cylindrospermum stagnale PCC7417 ( orf1 ). The recently reported oscillamycin is the product of the second cluster from Oscillatoria sp. PCC 182 and contains hydroxyproline, presumably introduced by the putative proline hydroxylase located in the first cluster ( orf ) (Yang, J. J., 216, Thesis, University of Helsinki, Finland). Many of the actinobacterial cluster strongly resemble that of cinnamycin, containing cinnamycin regulatory gene homologs, while cinnamycin B, recently reported from Actinomadura atramentaria NRBC (the cluster described here was identified by BLAT search from Actinomadura atramentaria DM 43919), includes only the essential divamide-like biosynthetic genes (Kodani,. et al., 216, J. Ind. Microbiol. Biotechnol. 43, ). c, Alignment of LanA precursor proteins from the divamide extended family, generated using ClustalW2 and then manually adjusted. The Nature Chemical Biology: doi:1.138/nchembio.2537
19 occurrence of multiple cassettes, as is seen for putative proteins 4-6, is a rare feature among RiPPs but common within the diversity-generating cyanobactins. 4 The alignment reveals conservation of Lan/MeLan, Lal, Gly, and Hya positions within the core, as well as hypervariable positions (*). Nature Chemical Biology: doi:1.138/nchembio.2537
20 B i ii iii Cylindrospermumstagnale PCC 7417 Genbank: CP Moorea producens 3L Genbank: GL Oscillatoria sp. PCC 182 Genbank: KB KB cytonemamillei VB Genbank: JTJC Actinomadura atramentaria DM Genbank: KB Actinomadura oligospora ATCC Genbank: JADG111.1 JADG13.1 Actinomadura macra NBRC 1412 Genbank: BCQT13.1 Actinomadura rubrobrunea NBRC Genbank: BCQU11.1 Frankia sp. EUN1f Genbank: ADGX Nocardiopsistrehalosi NBRC 1421 Genbank: BCRK12.1 Nocardiopsis potens DM Genbank: ANBB18.1 Marinaactinospora thermotolerans DM Genbank: NZ_FUW11.1 treptomyces roseoverticillatus strain NRRL B-35 Genbank: JOFL116.1 treptomyces africanus NRRL B Genbank: NZ_MUKA141.1 Anaerolineaceae bacterium 4572_78 Genbank: NBMH continued lana: WP_ lana: WP_ orf1 A orf2 M T orf3 A orf2 M T N X M A-1 A-2 T lana-1: EGJ lana: WP_ oscillamycin lana: WP_ T M orf A X orf M A N T X M A lana: WP_ orf X M A N cinnamycin B lana: WP_ lana: WP_ lana: WP_ cint/h X M A-1 N A-2 cint X M A N lana: WP_ cint X M A N lana: WP_ Y X M A N cinh cint lana: WP_ cinh cint X M A N cinr1 cinorf11 lana: WP_ orf cinh cint X M A N cinr1 cinorf11 lana: WP_ cinh cint X M A N cinr1 cinorf11 lana: WP_ Y cinh cint X M A N cinr cink cinorf1 11 cinr1 lana: WP_ cinh cint X M A N cinr cink cinorf1 11 cinr1 cinorf12 13 lana: WP_ orf M A lana: OQY kb Nature Chemical Biology: doi:1.138/nchembio.2537
21 C continued Prochloron didemni E11-36 (divamidea) 2. Prochloron didemni E11-37 (divamideb) 3. Anaerolineaceae bacterium 4572_78 4. Cylindrospermum stagnale PCC Cylindrospermum stagnale PCC 7417 (2) 6. Oscillatoria sp. PCC cytonema millei VB Moorea producens 3L 9. Moorea producens 3L (2) 1. Oscillatoria sp. PCC 182 (2; oscillamycin) 11. Frankia sp. EUN1f 12. treptomyces roseoverticillatus NRRL B Actinomadura atramentaria DM (cinnamycin B) 14. Actinomadura oligospora ATCC Actinomadura oligospora ATCC (2) 16. Actinomadura macra NRBC Actinomadura rubrobrunea NRBC treptomyces africanus NRRL B Nocardiopsis trehalosi NRBC Marinactinospora thermotolerans DM Nocardiopsis potens DM treptomyces cinnamoneous DM 45 (cinnamycin) 23. treptomyces cinnamoneous ATCC (duramycin) 24. treptomyces cinnamoneous ATCC (2) 25. treptoverticillium sp. R275 (duramycin B) 26. treptoverticillium sp. R217 (duramycin C) 27. treptomyces sp. No. A647P-2 (ancovenin) * * * * * * * * * * * Nature Chemical Biology: doi:1.138/nchembio.2537
Supplementary Results
 upplementary Results upplementary Table 1. Primers used in this study. upplementary Figure 1. PCP3-TE assay in the absence of ul. upplementary Figure 2. Multiple alignment of the sulfazecin TE, highlighting
upplementary Results upplementary Table 1. Primers used in this study. upplementary Figure 1. PCP3-TE assay in the absence of ul. upplementary Figure 2. Multiple alignment of the sulfazecin TE, highlighting
Supplementary Information
 Supplementary Information The conifer biomarkers dehydroabietic and abietic acids are widespread in Cyanobacteria Maria Sofia Costa, Adriana Rego, Vitor Ramos, Tiago Afonso, Sara Freitas, Marco Preto,
Supplementary Information The conifer biomarkers dehydroabietic and abietic acids are widespread in Cyanobacteria Maria Sofia Costa, Adriana Rego, Vitor Ramos, Tiago Afonso, Sara Freitas, Marco Preto,
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Cytochalasins from an Australian marine sediment-derived Phomopsis sp. (CMB-M0042F): Acid-mediated intra-molecular cycloadditions enhance
 SUPPORTING INFORMATION Cytochalasins from an Australian marine sediment-derived Phomopsis sp. (CMB-M42F): Acid-mediated intra-molecular cycloadditions enhance chemical diversity Zhuo Shang, Ritesh Raju,
SUPPORTING INFORMATION Cytochalasins from an Australian marine sediment-derived Phomopsis sp. (CMB-M42F): Acid-mediated intra-molecular cycloadditions enhance chemical diversity Zhuo Shang, Ritesh Raju,
Supporting Information. Lysine Propionylation to Boost Proteome Sequence. Coverage and Enable a Silent SILAC Strategy for
 Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Figure S1. PMVs from THP-1 cells expose phosphatidylserine and carry actin. A) Flow
 SUPPLEMENTARY DATA Supplementary Figure Legends Figure S1. PMVs from THP-1 cells expose phosphatidylserine and carry actin. A) Flow cytometry analysis of PMVs labelled with annexin-v-pe (Guava technologies)
SUPPLEMENTARY DATA Supplementary Figure Legends Figure S1. PMVs from THP-1 cells expose phosphatidylserine and carry actin. A) Flow cytometry analysis of PMVs labelled with annexin-v-pe (Guava technologies)
Supplementary Information
 Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Automated Purification and Analytical Reinjection of a Small Molecule Drug, Probenecid, on a Gilson LC/MS Dual Function System
 Automated Purification and Analytical Reinjection of a Small Molecule Drug, Probenecid, on a Gilson LC/MS Dual Function System Keywords Introduction Application Note PHA0413 High Pressure Liquid Chromatography
Automated Purification and Analytical Reinjection of a Small Molecule Drug, Probenecid, on a Gilson LC/MS Dual Function System Keywords Introduction Application Note PHA0413 High Pressure Liquid Chromatography
Supporting information (protein purification, kinetic characterization, product isolation, and characterization by NMR and mass spectrometry):
 Supporting Information Mechanistic studies of a novel C-S lyase in ergothioneine biosynthesis: the involvement of a sulfenic acid intermediate Heng Song, 1 Wen Hu, 1,2 Nathchar Naowarojna, 1 Ampon Sae
Supporting Information Mechanistic studies of a novel C-S lyase in ergothioneine biosynthesis: the involvement of a sulfenic acid intermediate Heng Song, 1 Wen Hu, 1,2 Nathchar Naowarojna, 1 Ampon Sae
Proteins? Protein function. Protein folding. Protein folding diseases. Protein interactions. Macromolecular assemblies. The end product of Genes
 Proteins? Protein function Protein folding Protein folding diseases Protein interactions Macromolecular assemblies The end product of Genes Protein Unfolding DOD Acid Catalysis DOD HDOD + N H N D C N C
Proteins? Protein function Protein folding Protein folding diseases Protein interactions Macromolecular assemblies The end product of Genes Protein Unfolding DOD Acid Catalysis DOD HDOD + N H N D C N C
Biology Open (2014) 000, 1 10 doi: /bio
 (2014) 000, 1 10 doi:10.1242/bio.201410041 Supplementary Material Michael Brauchle et al. doi: 10.1242/bio.201410041 Fig. S1. Alignment of GFP, sfgfp, egfp, eyfp, mcherry and mruby2. Sequence-based alignment
(2014) 000, 1 10 doi:10.1242/bio.201410041 Supplementary Material Michael Brauchle et al. doi: 10.1242/bio.201410041 Fig. S1. Alignment of GFP, sfgfp, egfp, eyfp, mcherry and mruby2. Sequence-based alignment
Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid
 Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid Joanne ( 乔安妮 ) Mather Senior Scientist Waters Corporation Data courtesy of Erin Chambers and Mary
Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid Joanne ( 乔安妮 ) Mather Senior Scientist Waters Corporation Data courtesy of Erin Chambers and Mary
Manja Henze, Dorothee Merker and Lothar Elling. 1. Characteristics of the Recombinant β-glycosidase from Pyrococcus
 S1 of S17 Supplementary Materials: Microwave-Assisted Synthesis of Glycoconjugates by Transgalactosylation with Recombinant Thermostable β-glycosidase from Pyrococcus Manja Henze, Dorothee Merker and Lothar
S1 of S17 Supplementary Materials: Microwave-Assisted Synthesis of Glycoconjugates by Transgalactosylation with Recombinant Thermostable β-glycosidase from Pyrococcus Manja Henze, Dorothee Merker and Lothar
Supporting Information for:
 Supporting Information for: Methylerythritol Cyclodiphosphate (MEcPP) in Deoxyxylulose Phosphate Pathway: Synthesis from an Epoxide and Mechanisms Youli Xiao, a Rodney L. Nyland II, b Caren L. Freel Meyers
Supporting Information for: Methylerythritol Cyclodiphosphate (MEcPP) in Deoxyxylulose Phosphate Pathway: Synthesis from an Epoxide and Mechanisms Youli Xiao, a Rodney L. Nyland II, b Caren L. Freel Meyers
Gladstone Institutes, University of California (UCSF), San Francisco, USA
 Fluorescence-linked Antigen Quantification (FLAQ) Assay for Fast Quantification of HIV-1 p24 Gag Marianne Gesner, Mekhala Maiti, Robert Grant and Marielle Cavrois * Gladstone Institutes, University of
Fluorescence-linked Antigen Quantification (FLAQ) Assay for Fast Quantification of HIV-1 p24 Gag Marianne Gesner, Mekhala Maiti, Robert Grant and Marielle Cavrois * Gladstone Institutes, University of
Dr. Erin E. Chambers Waters Corporation. Presented by Dr. Diego Rodriguez Cabaleiro Waters Europe Waters Corporation 1
 Development of an SPE-LC/MS/MS Assay for the Simultaneous Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid in Support of Alzheimer s Research Dr. Erin E. Chambers Waters Corporation Presented
Development of an SPE-LC/MS/MS Assay for the Simultaneous Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid in Support of Alzheimer s Research Dr. Erin E. Chambers Waters Corporation Presented
Supplementary Information
 Supplementary Information Structural basis of improved second generation 3-nitro-tyrosine trna synthetases Richard B. Cooley, Jessica L. Feldman, Camden M. Driggers, Taylor Bundy, Audrey L. Stokes, P.
Supplementary Information Structural basis of improved second generation 3-nitro-tyrosine trna synthetases Richard B. Cooley, Jessica L. Feldman, Camden M. Driggers, Taylor Bundy, Audrey L. Stokes, P.
2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry
 Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring
 Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Proteins are sometimes only produced in one cell type or cell compartment (brain has 15,000 expressed proteins, gut has 2,000).
 Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Isolation of Cytochrome C from Beef Heart BCHM 3100K-02
 Isolation of Cytochrome C from Beef Heart BCHM 3100K-02 John T. Johnson April 15, 2013 Dates Performed: Partner: Instructor: 01-Mar-2013 08-Mar-2013 22-Mar-2013 29-Mar-2013 05-Apr-2013 Anthony Ferrante
Isolation of Cytochrome C from Beef Heart BCHM 3100K-02 John T. Johnson April 15, 2013 Dates Performed: Partner: Instructor: 01-Mar-2013 08-Mar-2013 22-Mar-2013 29-Mar-2013 05-Apr-2013 Anthony Ferrante
<Supplemental information>
 The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
Product # Kit Components
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Pneumocystis jirovecii PCR Kit Product # 42820 Product Insert Background Information
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Pneumocystis jirovecii PCR Kit Product # 42820 Product Insert Background Information
Biological Mass Spectrometry. April 30, 2014
 Biological Mass Spectrometry April 30, 2014 Mass Spectrometry Has become the method of choice for precise protein and nucleic acid mass determination in a very wide mass range peptide and nucleotide sequencing
Biological Mass Spectrometry April 30, 2014 Mass Spectrometry Has become the method of choice for precise protein and nucleic acid mass determination in a very wide mass range peptide and nucleotide sequencing
MS/MS as an LC Detector for the Screening of Drugs and Their Metabolites in Race Horse Urine
 Application Note: 346 MS/MS as an LC Detector for the Screening of Drugs and Their Metabolites in Race Horse Urine Gargi Choudhary and Diane Cho, Thermo Fisher Scientific, San Jose, CA Wayne Skinner and
Application Note: 346 MS/MS as an LC Detector for the Screening of Drugs and Their Metabolites in Race Horse Urine Gargi Choudhary and Diane Cho, Thermo Fisher Scientific, San Jose, CA Wayne Skinner and
Figure S1: Abundance of Fe related proteins in oceanic Synechococcus sp. strain WH8102 in. Ferredoxin. Ferredoxin. Fe ABC transporter
 0.004 SW nm 0.04 0.03 nm nm 0.16 0.1 nm nm 0.40 0.3 nm nm 0.80 0.5 nm 1.6 1 nm 16 10 nm 160 100 nm 0 nm 0.04 nm 0.16 nm 0.40 nm 0.80 nm 1.6 nm 16 nm 160 nm Spectral counts 0 nm 0.04 nm 0.16 nm 0.40 nm
0.004 SW nm 0.04 0.03 nm nm 0.16 0.1 nm nm 0.40 0.3 nm nm 0.80 0.5 nm 1.6 1 nm 16 10 nm 160 100 nm 0 nm 0.04 nm 0.16 nm 0.40 nm 0.80 nm 1.6 nm 16 nm 160 nm Spectral counts 0 nm 0.04 nm 0.16 nm 0.40 nm
Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR
 Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR and LRRK2 WD40 GST fusion proteins (5 µg) were loaded
Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR and LRRK2 WD40 GST fusion proteins (5 µg) were loaded
Use of Capto ViralQ for the removal of genomic DNA from influenza virus produced in MDCK cells
 GE Healthcare Application note 28-9769-69 AA Vaccines Use of Capto ViralQ for the removal of genomic DNA from influenza virus produced in MDCK cells We have developed a chromatographic method to remove
GE Healthcare Application note 28-9769-69 AA Vaccines Use of Capto ViralQ for the removal of genomic DNA from influenza virus produced in MDCK cells We have developed a chromatographic method to remove
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood
 Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB
 Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Rose et al. Supplementary Material. 1. Supplementary Materials and Methods
 Rose et al. Supplementary Material 1. Supplementary Materials and Methods 1.1. Synthesis of the biotinylated CrA modules. 1.1.1. Instrumentation and reagents: All solvents were purchased from Carl Roth
Rose et al. Supplementary Material 1. Supplementary Materials and Methods 1.1. Synthesis of the biotinylated CrA modules. 1.1.1. Instrumentation and reagents: All solvents were purchased from Carl Roth
Tivadar Orban, Beata Jastrzebska, Sayan Gupta, Benlian Wang, Masaru Miyagi, Mark R. Chance, and Krzysztof Palczewski
 Structure, Volume Supplemental Information Conformational Dynamics of Activation for the Pentameric Complex of Dimeric G Protein-Coupled Receptor and Heterotrimeric G Protein Tivadar Orban, Beata Jastrzebska,
Structure, Volume Supplemental Information Conformational Dynamics of Activation for the Pentameric Complex of Dimeric G Protein-Coupled Receptor and Heterotrimeric G Protein Tivadar Orban, Beata Jastrzebska,
Supplementary Figure 1. Chemical structures of activity-based probes (ABPs) and of click reagents used in this study.
 Supplementary Figure 1. Chemical structures of activity-based probes (ABPs) and of click reagents used in this study. In this study, one fluorophosphonate (FP, 1), three nitrophenol phosphonate probes
Supplementary Figure 1. Chemical structures of activity-based probes (ABPs) and of click reagents used in this study. In this study, one fluorophosphonate (FP, 1), three nitrophenol phosphonate probes
BabyBio IMAC columns DATA SHEET DS
 BabyBio IMAC columns DATA SHEET DS 45 655 010 BabyBio columns for Immobilized Metal Ion Affinity Chromatography (IMAC) are ready-to-use for quick and easy purification of polyhistidine-tagged (His-tagged)
BabyBio IMAC columns DATA SHEET DS 45 655 010 BabyBio columns for Immobilized Metal Ion Affinity Chromatography (IMAC) are ready-to-use for quick and easy purification of polyhistidine-tagged (His-tagged)
[ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS
![[ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS [ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS](/thumbs/79/80328542.jpg) Yun Wang Alelyunas, Henry Shion, Mark Wrona Waters Corporation, Milford, MA, USA APPLICATION BENEFITS mab LC-MS method which enables users to achieve highly sensitive bioanalysis of intact trastuzumab
Yun Wang Alelyunas, Henry Shion, Mark Wrona Waters Corporation, Milford, MA, USA APPLICATION BENEFITS mab LC-MS method which enables users to achieve highly sensitive bioanalysis of intact trastuzumab
Graphene Quantum Dots-Band-Aids Used for Wound Disinfection
 Supporting information Graphene Quantum Dots-Band-Aids Used for Wound Disinfection Hanjun Sun, Nan Gao, Kai Dong, Jinsong Ren, and Xiaogang Qu* Laboratory of Chemical Biology, Division of Biological Inorganic
Supporting information Graphene Quantum Dots-Band-Aids Used for Wound Disinfection Hanjun Sun, Nan Gao, Kai Dong, Jinsong Ren, and Xiaogang Qu* Laboratory of Chemical Biology, Division of Biological Inorganic
Supplementary Figure 1. The proposed biosynthetic pathways for the A-ring transformation of the
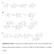 Supplementary Figure 1. The proposed biosynthetic pathways for the A-ring transformation of the kinamycin and lomaiviticin antibiotics. A, by Steven J. Gould; B, by Emily P. Balskus; C, by Bradley S. Moore.
Supplementary Figure 1. The proposed biosynthetic pathways for the A-ring transformation of the kinamycin and lomaiviticin antibiotics. A, by Steven J. Gould; B, by Emily P. Balskus; C, by Bradley S. Moore.
f(x) = x R² = RPKM (M8.MXB) f(x) = x E-014 R² = 1 RPKM (M31.
 14 12 f(x) = 1.633186874x - 21.46732234 R² =.995616541 RPKM (M8.MXA) 1 8 6 4 2 2 4 6 8 1 12 14 RPKM (M8.MXB) 14 12 f(x) =.821767782x - 4.192595677497E-14 R² = 1 RPKM (M31.XA) 1 8 6 4 2 2 4 6 8 1 12 14
14 12 f(x) = 1.633186874x - 21.46732234 R² =.995616541 RPKM (M8.MXA) 1 8 6 4 2 2 4 6 8 1 12 14 RPKM (M8.MXB) 14 12 f(x) =.821767782x - 4.192595677497E-14 R² = 1 RPKM (M31.XA) 1 8 6 4 2 2 4 6 8 1 12 14
Minutes Figure S1. HPLC separation of nucleosides from LC/ESI-MS analysis of a total enzymatic Trp
 100 A % Relative Abundance m m Ø acp 5 m Øm 5 m m 7 m s * 6 ia m * 0 5 10 15 0 5 0 5 40 Minutes Figure S1. HPL separation of nucleosides from L/ESI-MS analysis of a total enzymatic digest of mt trna. V
100 A % Relative Abundance m m Ø acp 5 m Øm 5 m m 7 m s * 6 ia m * 0 5 10 15 0 5 0 5 40 Minutes Figure S1. HPL separation of nucleosides from L/ESI-MS analysis of a total enzymatic digest of mt trna. V
Supporting Information
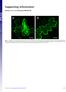 Supporting Information Harries et al. 1.173/pnas.9923916 A Fig. S1. Disruption of microfilaments within epidermal cells after treatment with 5 M Lat. Images of N. benthamiana cells are from plants expressing
Supporting Information Harries et al. 1.173/pnas.9923916 A Fig. S1. Disruption of microfilaments within epidermal cells after treatment with 5 M Lat. Images of N. benthamiana cells are from plants expressing
Gene expression scaled by distance to the genome replication site. Ying et al. Supporting Information. Contents. Supplementary note I p.
 Gene expression scaled by distance to the genome replication site Ying et al Supporting Information Contents Supplementary note I p. 2 Supplementary note II p. 3-5 Supplementary note III p. 5-7 Supplementary
Gene expression scaled by distance to the genome replication site Ying et al Supporting Information Contents Supplementary note I p. 2 Supplementary note II p. 3-5 Supplementary note III p. 5-7 Supplementary
Comparison of mass spectrometers performances
 Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
Supporting Information
 Supporting Information For Heterologous production of thiostrepton A and biosynthetic engineering of thiostrepton analogs Chaoxuan Li, Feifei Zhang, and Wendy L. Kelly* Table of Contents Figure S1. PCR
Supporting Information For Heterologous production of thiostrepton A and biosynthetic engineering of thiostrepton analogs Chaoxuan Li, Feifei Zhang, and Wendy L. Kelly* Table of Contents Figure S1. PCR
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
Transient Ribosomal Attenuation Coordinates Protein Synthesis and Co-translational Folding
 SUPPLEMENTARY INFORMATION: Transient Ribosomal Attenuation Coordinates Protein Synthesis and Co-translational Folding Gong Zhang 1,2, Magdalena Hubalewska 1 & Zoya Ignatova 1,2 1 Department of Cellular
SUPPLEMENTARY INFORMATION: Transient Ribosomal Attenuation Coordinates Protein Synthesis and Co-translational Folding Gong Zhang 1,2, Magdalena Hubalewska 1 & Zoya Ignatova 1,2 1 Department of Cellular
LC-MS/MS for the quantification of Peptide biomarker and mixture of closely related Protein in formulation
 EUROPEAN BIOANALYSIS FORUM Barcelona, November 14-16, 2012 LC-MS/MS for the quantification of Peptide biomarker and mixture of closely related Protein in formulation Luc-Alain SAVOY CONTENT Part I: SGS
EUROPEAN BIOANALYSIS FORUM Barcelona, November 14-16, 2012 LC-MS/MS for the quantification of Peptide biomarker and mixture of closely related Protein in formulation Luc-Alain SAVOY CONTENT Part I: SGS
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
Ion Exchange Chromatography COSMOGEL IEX Series Columns
 Ion Echange Chromatography COSMOGEL IEX Series Columns Available in 3 different ion-echange modes Anion-echange type, Cation-echange type, Amphoteric ion-echange type Available for 3 different application
Ion Echange Chromatography COSMOGEL IEX Series Columns Available in 3 different ion-echange modes Anion-echange type, Cation-echange type, Amphoteric ion-echange type Available for 3 different application
To test the possible source of the HBV infection outside the study family, we searched the Genbank
 Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Phosphate buffered saline (PBS) for washing the cells TE buffer (nuclease-free) ph 7.5 for use with the PrimePCR Reverse Transcription Control Assay
 Catalog # Description 172-5080 SingleShot Cell Lysis Kit, 100 x 50 µl reactions 172-5081 SingleShot Cell Lysis Kit, 500 x 50 µl reactions For research purposes only. Introduction The SingleShot Cell Lysis
Catalog # Description 172-5080 SingleShot Cell Lysis Kit, 100 x 50 µl reactions 172-5081 SingleShot Cell Lysis Kit, 500 x 50 µl reactions For research purposes only. Introduction The SingleShot Cell Lysis
Supplementary Figure 1
 Supplementary Figure 1 Isolation of mt-trnas and RNA-MS analysis of mt-trna Asn from M. nudus (a)m. nudus mt-trnas were isolated by RCC and resolved by 10% denaturing PAGE. The gel was stained with SYBR
Supplementary Figure 1 Isolation of mt-trnas and RNA-MS analysis of mt-trna Asn from M. nudus (a)m. nudus mt-trnas were isolated by RCC and resolved by 10% denaturing PAGE. The gel was stained with SYBR
Key Words: Brassica oleraceae, glucosinolate, liquid chromatography mass spectrometry, FNH-I-003
 IDENTIFICATION OF MAJOR GLUCOSINOLATES IN BROCCOLI (Brassica oleracea var. italica) BY LIQUID CHROMATOGRAPHY MASS SPECTROMETRY (LC-MS) AND DETERMINATION OF ANTICANCER PROPERTIES OF BROCCOLI EXTRACTS Carlos
IDENTIFICATION OF MAJOR GLUCOSINOLATES IN BROCCOLI (Brassica oleracea var. italica) BY LIQUID CHROMATOGRAPHY MASS SPECTROMETRY (LC-MS) AND DETERMINATION OF ANTICANCER PROPERTIES OF BROCCOLI EXTRACTS Carlos
LC/QTOF Identification of Microcystin Variants in Cell- Bound Algae from the San Francisco Estuary
 LC/QTOF Identification of Microcystin Variants in Cell- Bound Algae from the San Francisco Estuary Ralph Hindle Vogon Laboratory Services Ltd. Cochrane, Alberta, Canada Max Mizel Dept. of Civil and Environmental
LC/QTOF Identification of Microcystin Variants in Cell- Bound Algae from the San Francisco Estuary Ralph Hindle Vogon Laboratory Services Ltd. Cochrane, Alberta, Canada Max Mizel Dept. of Civil and Environmental
Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene
 Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene YUELIN ZHANG, WEIHUA FAN, MARK KINKEMA, XIN LI, AND
Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene YUELIN ZHANG, WEIHUA FAN, MARK KINKEMA, XIN LI, AND
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
complemented with SipA ( SipA/pSipA) or SL1344 WT for 48 hours, after which the
 P-gp expression (% of control) 12 1 8 6 4 2 * * Untreated SipA SipA/pSipA WT Supplementary Figure 1. SipA modulates the expression of P-gp in healthy murine intestinal epithelium in vivo. Salmonella Typhimurium
P-gp expression (% of control) 12 1 8 6 4 2 * * Untreated SipA SipA/pSipA WT Supplementary Figure 1. SipA modulates the expression of P-gp in healthy murine intestinal epithelium in vivo. Salmonella Typhimurium
CHAPTER 3 Amino Acids, Peptides, Proteins
 CHAPTER 3 Amino Acids, Peptides, Proteins Learning goals: Structure and naming of amino acids Structure and properties of peptides Ionization behavior of amino acids and peptides Methods to characterize
CHAPTER 3 Amino Acids, Peptides, Proteins Learning goals: Structure and naming of amino acids Structure and properties of peptides Ionization behavior of amino acids and peptides Methods to characterize
ESI and MALDI Mass Spectrometry. S. Sankararaman Department of Chemistry Indian Institute of Technology Madras Chennai
 ESI and MALDI Mass Spectrometry S. Sankararaman Department of Chemistry Indian Institute of Technology Madras Chennai 600036 sanka@iitm.ac.in MODULE 22 ESI-Q MASS SPECTROMETRY - examples Mol. Wt = 1140
ESI and MALDI Mass Spectrometry S. Sankararaman Department of Chemistry Indian Institute of Technology Madras Chennai 600036 sanka@iitm.ac.in MODULE 22 ESI-Q MASS SPECTROMETRY - examples Mol. Wt = 1140
Supplementary Information
 Supplementary Information Precursors of trnas are stabilized by methylguanosine cap structures Takayuki Ohira and Tsutomu Suzuki Department of Chemistry and Biotechnology, Graduate School of Engineering,
Supplementary Information Precursors of trnas are stabilized by methylguanosine cap structures Takayuki Ohira and Tsutomu Suzuki Department of Chemistry and Biotechnology, Graduate School of Engineering,
Total Histone H3 Acetylation Detection Fast Kit (Colorimetric)
 Total Histone H3 Acetylation Detection Fast Kit (Colorimetric) Catalog Number KA1538 48 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use...
Total Histone H3 Acetylation Detection Fast Kit (Colorimetric) Catalog Number KA1538 48 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use...
Learning Objectives. Overview of topics to be discussed 10/25/2013 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS
 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan
 PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
LANCE Eu-W1024 ITC Chelate & Europium Standard AD0013 Development grade
 AD0017P-4 (en) 1 LANCE Eu-W1024 ITC Chelate & Europium Standard AD0013 Development grade INTRODUCTION Fluorescent isothiocyanato-activated (ITC-activated) Eu-W1024 chelate is optimized for labelling proteins
AD0017P-4 (en) 1 LANCE Eu-W1024 ITC Chelate & Europium Standard AD0013 Development grade INTRODUCTION Fluorescent isothiocyanato-activated (ITC-activated) Eu-W1024 chelate is optimized for labelling proteins
Supporting Information
 Supporting Information A single design strategy for dual sensitive ph probe with a suitable range to map ph in living cells Kang-Kang Yu, Ji-Ting Hou, Kun Li, * Qian Yao, Jin Yang, Ming-Yu Wu, Yong-Mei
Supporting Information A single design strategy for dual sensitive ph probe with a suitable range to map ph in living cells Kang-Kang Yu, Ji-Ting Hou, Kun Li, * Qian Yao, Jin Yang, Ming-Yu Wu, Yong-Mei
ab Calcineurin Phosphatase Activity Assay Kit (Colorimetric)
 ab139461 Calcineurin Phosphatase Activity Assay Kit (Colorimetric) Instructions for Use For measuring Calcineurin phosphatase activity. This product is for research use only and is not intended for diagnostic
ab139461 Calcineurin Phosphatase Activity Assay Kit (Colorimetric) Instructions for Use For measuring Calcineurin phosphatase activity. This product is for research use only and is not intended for diagnostic
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Total genomic DNA was extracted from either 6 DBS punches (3mm), or 0.1ml of peripheral or
 Material and methods Measurement of telomere length (TL) Total genomic DNA was extracted from either 6 DBS punches (3mm), or 0.1ml of peripheral or cord blood using QIAamp DNA Mini Kit and a Qiacube (Qiagen).
Material and methods Measurement of telomere length (TL) Total genomic DNA was extracted from either 6 DBS punches (3mm), or 0.1ml of peripheral or cord blood using QIAamp DNA Mini Kit and a Qiacube (Qiagen).
Supporting Information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Supporting Information Enzyme-activatable Probe with a Self-immolative Linker for Rapid and Sensitive
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Supporting Information Enzyme-activatable Probe with a Self-immolative Linker for Rapid and Sensitive
Supplementary Material (ESI) for Chemical Communications This journal is (c) The Royal Society of Chemistry 2008
 Experimental Details Unless otherwise noted, all chemicals were purchased from Sigma-Aldrich Chemical Company and were used as received. 2-DOS and neamine were kindly provided by Dr. F. Huang. Paromamine
Experimental Details Unless otherwise noted, all chemicals were purchased from Sigma-Aldrich Chemical Company and were used as received. 2-DOS and neamine were kindly provided by Dr. F. Huang. Paromamine
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Unsupervised Identification of Isotope-Labeled Peptides
 Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Electronic Supplementary Information. Table of Contents
 Electronic Supplementary Information Examination of native chemical ligation using peptidyl prolyl thioester Takahiro Nakamura, Akira Shigenaga, Kohei Sato, Yusuke Tsuda, Ken Sakamoto, and Akira Otaka*
Electronic Supplementary Information Examination of native chemical ligation using peptidyl prolyl thioester Takahiro Nakamura, Akira Shigenaga, Kohei Sato, Yusuke Tsuda, Ken Sakamoto, and Akira Otaka*
Measuring Lipid Composition LC-MS/MS
 Project: Measuring Lipid Composition LC-MS/MS Verification of expected lipid composition in nanomedical controlled release systems by liquid chromatography tandem mass spectrometry AUTHORED BY: DATE: Sven
Project: Measuring Lipid Composition LC-MS/MS Verification of expected lipid composition in nanomedical controlled release systems by liquid chromatography tandem mass spectrometry AUTHORED BY: DATE: Sven
Supplementary Information
 Supplementary Information Assembly and Clustering of Natural Antibiotics Guides Target Identification Chad W. Johnston 1,2, Michael A. Skinnider 1,2, Chris A. Dejong 1,2, Philip N. Rees 1,2, Gregory M.
Supplementary Information Assembly and Clustering of Natural Antibiotics Guides Target Identification Chad W. Johnston 1,2, Michael A. Skinnider 1,2, Chris A. Dejong 1,2, Philip N. Rees 1,2, Gregory M.
Use of a camp BRET Sensor to Characterize a Novel Regulation of camp by the Sphingosine-1-phosphate/G 13 Pathway
 Use of a camp BRET Sensor to Characterize a Novel Regulation of camp by the Sphingosine-1-phosphate/G 13 Pathway SUPPLEMENTAL DATA Characterization of the CAMYEL sensor and calculation of intracellular
Use of a camp BRET Sensor to Characterize a Novel Regulation of camp by the Sphingosine-1-phosphate/G 13 Pathway SUPPLEMENTAL DATA Characterization of the CAMYEL sensor and calculation of intracellular
Determination of red blood cell fatty acid profiles in clinical research
 Application Note Clinical Research Determination of red blood cell fatty acid profiles in clinical research Chemical ionization gas chromatography tandem mass spectrometry Authors Yvonne Schober 1, Hans
Application Note Clinical Research Determination of red blood cell fatty acid profiles in clinical research Chemical ionization gas chromatography tandem mass spectrometry Authors Yvonne Schober 1, Hans
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
Caspase-3 Assay Cat. No. 8228, 100 tests. Introduction
 Introduction Caspase-3 Assay Cat. No. 8228, 100 tests Caspase-3 is a member of caspases that plays a key role in mediating apoptosis, or programmed cell death. Upon activation, it cleaves a variety of
Introduction Caspase-3 Assay Cat. No. 8228, 100 tests Caspase-3 is a member of caspases that plays a key role in mediating apoptosis, or programmed cell death. Upon activation, it cleaves a variety of
RP-HPLC Analysis of Temozolomide in Pharmaceutical Dosage Forms
 Asian Journal of Chemistry Vol. 22, No. 7 (2010), 5067-5071 RP-HPLC Analysis of Temozolomide in Pharmaceutical Dosage Forms A. LAKSHMANA RAO*, G. TARAKA RAMESH and J.V.L.N.S. RAO Department of Pharmaceutical
Asian Journal of Chemistry Vol. 22, No. 7 (2010), 5067-5071 RP-HPLC Analysis of Temozolomide in Pharmaceutical Dosage Forms A. LAKSHMANA RAO*, G. TARAKA RAMESH and J.V.L.N.S. RAO Department of Pharmaceutical
- 1 - Cell types Monocytes THP-1 cells Macrophages. LPS Treatment time (Hour) IL-6 level (pg/ml)
 Supplementary Table ST1: The dynamic effect of LPS on IL-6 production in monocytes and THP-1 cells after GdA treatment. Monocytes, THP-1 cells and macrophages (5x10 5 ) were incubated with 10 μg/ml of
Supplementary Table ST1: The dynamic effect of LPS on IL-6 production in monocytes and THP-1 cells after GdA treatment. Monocytes, THP-1 cells and macrophages (5x10 5 ) were incubated with 10 μg/ml of
Application Note. Agilent Application Solution Analysis of ascorbic acid, citric acid and benzoic acid in orange juice. Author. Abstract.
 Agilent Application Solution Analysis of ascorbic acid, citric acid and benzoic acid in orange juice Application Note Author Food Syed Salman Lateef Agilent Technologies, Inc. Bangalore, India 8 6 4 2
Agilent Application Solution Analysis of ascorbic acid, citric acid and benzoic acid in orange juice Application Note Author Food Syed Salman Lateef Agilent Technologies, Inc. Bangalore, India 8 6 4 2
Synthetic chemistry-led creation of a difluorinated biaryl ether non-nucleoside reverse transcriptase inhibitor
 upplementary Material for rganic & Biomolecular Chemistry ynthetic chemistry-led creation of a difluorinated biaryl ether non-nucleoside reverse transcriptase inhibitor Lyn. Jones* Amy Randall, scar Barba
upplementary Material for rganic & Biomolecular Chemistry ynthetic chemistry-led creation of a difluorinated biaryl ether non-nucleoside reverse transcriptase inhibitor Lyn. Jones* Amy Randall, scar Barba
Development and Validation of a Polysorbate 20 Assay in a Therapeutic Antibody Formulation by RP-HPLC and Charged Aerosol Detector (CAD)
 LIFE SCIENCES AR ENOUGH? HOW DO YOU KNOW RAW MATERIALS ARE PURE? HOW DO YOU EVALUATE PRODUCT PACKAGING? HOW DO YOU DO YOU KNOW WHAT ANALYTICAL TECHNIQUE TO USE? HOW DO YOU SIMULTANEOUSLY TEST FOR TWO BYPRODUCTS?
LIFE SCIENCES AR ENOUGH? HOW DO YOU KNOW RAW MATERIALS ARE PURE? HOW DO YOU EVALUATE PRODUCT PACKAGING? HOW DO YOU DO YOU KNOW WHAT ANALYTICAL TECHNIQUE TO USE? HOW DO YOU SIMULTANEOUSLY TEST FOR TWO BYPRODUCTS?
SUPPLEMENTARY INFORMATION. An orthogonal ribosome-trnas pair by the engineering of
 SUPPLEMENTARY INFORMATION An orthogonal ribosome-trnas pair by the engineering of peptidyl transferase center Naohiro Terasaka 1 *, Gosuke Hayashi 1 *, Takayuki Katoh 1, and Hiroaki Suga 1,2 1 Department
SUPPLEMENTARY INFORMATION An orthogonal ribosome-trnas pair by the engineering of peptidyl transferase center Naohiro Terasaka 1 *, Gosuke Hayashi 1 *, Takayuki Katoh 1, and Hiroaki Suga 1,2 1 Department
AlphaScreen : A Straightforward and Powerful Alternative to ELISA. Martina Bielefeld-Sévigny Ph.D., R&D Director
 AlphaScreen : A Straightforward and Powerful Alternative to ELISA Martina Bielefeld-Sévigny Ph.D., R&D Director Overview AlphaScreen - an alternative to ELISA Why an alternative to ELISA? Assay principle
AlphaScreen : A Straightforward and Powerful Alternative to ELISA Martina Bielefeld-Sévigny Ph.D., R&D Director Overview AlphaScreen - an alternative to ELISA Why an alternative to ELISA? Assay principle
Supplementary materials
 Supplementary materials Chemical library from ChemBridge 50,240 structurally diverse small molecule compounds dissolved in DMSO Hits Controls: No virus added μ Primary screening at 20 g/ml of compounds
Supplementary materials Chemical library from ChemBridge 50,240 structurally diverse small molecule compounds dissolved in DMSO Hits Controls: No virus added μ Primary screening at 20 g/ml of compounds
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus
 Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
Double charge of 33kD peak A1 A2 B1 B2 M2+ M/z. ABRF Proteomics Research Group - Qualitative Proteomics Study Identifier Number 14146
 Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Supplementary Figure 1. mrna targets were found in exosomes and absent in free-floating supernatant. Serum exosomes and exosome-free supernatant were
 Supplementary Figure 1. mrna targets were found in exosomes and absent in free-floating supernatant. Serum exosomes and exosome-free supernatant were separated via ultracentrifugation and lysed to analyze
Supplementary Figure 1. mrna targets were found in exosomes and absent in free-floating supernatant. Serum exosomes and exosome-free supernatant were separated via ultracentrifugation and lysed to analyze
Application Note # FTMS-46 solarix XR: Analysis of Complex Mixtures
 Application Note # FTMS-46 solarix XR: Analysis of Complex Mixtures Introduction Natural organic matter (NOM) as well as crude oil and crude oil fractions are very complex mixtures of organic compounds
Application Note # FTMS-46 solarix XR: Analysis of Complex Mixtures Introduction Natural organic matter (NOM) as well as crude oil and crude oil fractions are very complex mixtures of organic compounds
Supporting Information Table of Contents
 Supporting Information Table of Contents Supporting Information Figure 1 Page 2 Supporting Information Figure 2 Page 4 Supporting Information Figure 3 Page 5 Supporting Information Figure 4 Page 6 Supporting
Supporting Information Table of Contents Supporting Information Figure 1 Page 2 Supporting Information Figure 2 Page 4 Supporting Information Figure 3 Page 5 Supporting Information Figure 4 Page 6 Supporting
Lecture 33 Membrane Proteins
 Lecture 33 Membrane Proteins Reading for today: Chapter 4, section D Required reading for next Wednesday: Chapter 14, sections A and 14.19 to the end Kuriyan, J., and Eisenberg, D. (2007) The origin of
Lecture 33 Membrane Proteins Reading for today: Chapter 4, section D Required reading for next Wednesday: Chapter 14, sections A and 14.19 to the end Kuriyan, J., and Eisenberg, D. (2007) The origin of
Chapter 8. Interaction between the phosphatidylinositol 3- kinase SH3 domain and a photocleavable cyclic peptide
 Interaction between the phosphatidylinositol 3- kinase SH3 domain and a photocleavable cyclic peptide 129 Abstract The interaction of the PI3K SH3 domain with a cyclic photocleavable peptide and the linear
Interaction between the phosphatidylinositol 3- kinase SH3 domain and a photocleavable cyclic peptide 129 Abstract The interaction of the PI3K SH3 domain with a cyclic photocleavable peptide and the linear
Supporting Information Nitric oxide releasing photoresponsive nanohybrids as excellent therapeutic agent for cervical cancer cell lines
 upporting Information itric oxide releasing photoresponsive nanohybrids as excellent therapeutic agent for cervical cancer cell lines Priya udhesh a, Kaviyarasan Tamilarasan a, Palaniappan Arumugam a and
upporting Information itric oxide releasing photoresponsive nanohybrids as excellent therapeutic agent for cervical cancer cell lines Priya udhesh a, Kaviyarasan Tamilarasan a, Palaniappan Arumugam a and
Determination of Tetracyclines in Chicken by Solid-Phase Extraction and High-Performance Liquid Chromatography
 Determination of Tetracyclines in Chicken by Solid-Phase Extraction and High-Performance Liquid Chromatography Application ote Food Safety Authors Chen-Hao Zhai and Yun Zou Agilent Technologies Co. Ltd.
Determination of Tetracyclines in Chicken by Solid-Phase Extraction and High-Performance Liquid Chromatography Application ote Food Safety Authors Chen-Hao Zhai and Yun Zou Agilent Technologies Co. Ltd.
Supporting information
 Supporting information Figure legends Supplementary Table 1. Specific product ions obtained from fragmentation of lithium adducts in the positive ion mode comparing the different positional isomers of
Supporting information Figure legends Supplementary Table 1. Specific product ions obtained from fragmentation of lithium adducts in the positive ion mode comparing the different positional isomers of
Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved
 1 Supplemental Figure Legends Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved PCSK9 concentrations. 4 Plasma mature and furin-cleaved PCSK9s were measured by a sandwich
1 Supplemental Figure Legends Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved PCSK9 concentrations. 4 Plasma mature and furin-cleaved PCSK9s were measured by a sandwich
