ISOXAZOLE INHIBITORS OF BROMODOMAINS
|
|
|
- Nora Carson
- 6 years ago
- Views:
Transcription
1 ISOXAZOLE INHIBITORS OF BROMODOMAINS Paul Brennan SGC Oxford Nuffield Dept. of Clinical Medicine University of Oxford RSC Advances in Synthesis and Medicinal Chemistry 1 May 2012
2 INTRODUCING THE SGC A model for open access public-private partnership The SGC is a public-private partnership with a mandate to place protein structures of relevance to human health into the public domain, free from restrictions on use. Focus on proteins from human and human parasites. To promote drug discovery by substantially increasing the number of medically relevant protein structures, as well as related reagents and protocols, available in the public domain Human proteins (main effort) Proteins from pathogens (e.g. Plasmodium) Chemical probes Biological probes Open Source science All structures/results are made freely available promptly Funding partners receive no prior access or rights to data or progress information No IP
3 PARTNERS Pharma GSK Novartis Pfizer Eli Lilly Abbott Takeda SGC Academia Oxford University Wellcome Trust
4 LEADER IN STRUCTURAL GENOMICS Kinase Structures Epigenetic Structures Family Lysine demethylase (KDM) Number of targets Purified in SGC Structures deposited [SGC/Total] /8 Bromodomain (BRD) /17 R O Y A L Tudor domain /19 Chromo domain /16 MBT domain 9 8 4/7 PHD /23 Histone acetyltransferase (HAT) Histone methyltransferase (HMT) / /18 TOTAL /116
5 EPIGENETICS Wikipedia: In biology, and specifically genetics, epigenetics is the study of heritable changes in gene expression or cellular phenotype caused by mechanisms other than changes in the underlying DNA sequence. Underlying Mechanism of: Cell differentiation Disease progression Molecular Mechanism: Reversible DNA modification Reversible histone modification
6 CHROMATIN The largest human cells are 0.1 μm ( m) wide. There are 2 m of DNA in every cell.
7 HISTONE CODE: WRITERS AND ERASERS
8 HISTONE CODE: READER
9 EPIGENETIC CODE Histones can be modified and recognized Histone Modification Write Read Erase Acetyl HAT Bromo HDAC Methyl HMT Chromo, PHD, Tudor, MBT HDM Different modifications affect gene transcription Modification H3K4 H3K9 H3K27 KMe Activation Activation Activation KMe 3 Activation Repression Repression KAc Activation Activation Activation HAT: histone acetyl transferase, Bromo: bromodomain; HDAC: histone deacetylase; HMT: histone methyl transferase Chromodomain, PHD-domain, Tudor-domain, MBT-domain; HDM: histone demethylase
10 SGC PROGRESS FOR EPIGENETICS TARGETS Pure at SGC Assay at SGC SGC structure Methylation PHD Non-SGC structure PKMT/PRMT Acetylation BRD KDM CHROMO TUDOR PolyADP Ribosylation HAT PARP MACRO 17 Targets 55 Targets 60 Targets 30 Targets 9 Targets MBT 34 Targets 36 Targets 97 Targets 17 Targets 15 Targets
11 EPIGENETIC TARGETS IN DISEASE Early position in signalling cascades: - Epigenetic modifications regulate multiple genes implicated in chronic diseases Potential for many drugs, in many therapeutic areas Francesca De Santa, Vipin Narang Zhei Hwee Yap, Betsabeh Khoramian Tusi, Thomas Burgold, Liv Austenaa, Gabriele Bucci, Marieta Caganova, Samuele Notarbartolo, Stefano Casola, Giuseppe Testa, Wing-Kin Sung, Chia-Lin Wei,* and Gioacchino Natoli,*
12 EPIGENETIC TARGETS IN DISEASE Neurology & Psychiatry Inflammatory Cancer No disease link Neurological disease Inflammation Cancer
13 SGC CHEMICAL PROBE DISCOVERY Probe Criteria In vitro activity: IC 50 or Kd 100nM In vitro selectivity: 30-fold vs. other branches of phylogenetic tree Cellular activity: IC 50 1μM Purified Protein & Structure Hit ID Hit Optimization Probe Characterisation Probe Datasheet Sigma Catalogue Publications Scientific Community Epigenetics Biology Validate Drug Targets Assay development Focused sets VLS Fragments HTS Analogue purchase Synthesis SAR generation SAR analysis Complex structures Selectivity Secondary assays Cellular assays Selectivity
14 CHEMICAL PROBES HAVE BIG IMPACT Number of Papers Nuclear Hormone Receptors * * *Chemical Probe * * * * * * Open access chemical and clinical probes to support drug discovery Nat. Chem. Bio. 2009, 5 (7),
15 EPIGENETIC CHEMICAL PROBES Epigenetics Biology Chemical Probes
16 BROMODOMAIN STRUCTURAL ALIGNMENT TREE AlphaScreens BET subfamily
17 ACETYL LYSINE READERS: BROMODOMAINS 62 domains in Human ~120 residue Kac interaction module ( reader ) Clinical POC targeting Kac regulation (HDACs) Inflammation Cancer Metabolic disease Neurological diseases Cardiovascular diseases
18 BROMODOMAIN CONTAINING PROTEINS
19 BROMODOMAINS AND CANCER Domain organization of bromodomain proteins and translocations in cancer. Bromodomain modules are shown in green (labelled BRD). Other domain types are labelled directly in the figure and breakpoints are indicated by arrows. Wild type domain arrangements are shown in the upper panel. Oncogenic fusions of CBP in acute leukemia. CBP contributes to tumorigenesis of fusion NUP98-HoxA9 and MOZ-TIF2 proteins. CBP mutations in relapsed acute lymphoblastic leukaemia (AML) and are very common in diffuse large B-cell lymphoma and follicular lymphoma. CBP and the related HAT EP300 are also highly expressed in advanced prostate cancer and expression levels have been linked with cancer patient survival.
20 ACETYL LYSINE BINDING SITES
21 ACETYL LYSINE MIMETICS Ligand BRD2 BRD4 BAZ2B CREBBP FALZ Kac 3,740 7,210 2, ,350 DMSO 255, , ,500 17,600 78,200 NMP 8,900 6,000 34, ,400 E >> IC 50 [ M]
22 BROMODOMAIN BINDING ASSAYS Assay Alpha- Screen Potent Output Sensitivity Through -put Ligand needed Low IC50 High High Yes Drawbacks High false positive rate Tm Shift High Δ C Medium High No Indirect Octet BLI Low Kd Medium Medium No Biotinylated protein Data Points 4383 compounds 6 proteins 4615 compounds 21 proteins 219 compounds 4 proteins Micro ITC Low Kd Low Low No Protein consumption Very few
23 BROMODOMAIN BINDING ASSAYS BRD ITC pkd (+)-JQ1 Tm Shift vs ITC CBP AlphaScreen pic y = x CBP Tm Shift 10 um R² = BRD Tm Shift 10 um ITC pkd = 0.14*[Tm Shift C] +6.1 R 2 = 0.92 BRD4~1 AlphaScreen pic y = x BRD4~1 Tm Shift 10 um R² =
24 DISCOVERY OF ISOXAZOLES BRD4* 36 um BRD4* 3.3 um BRD4* 0.7 um Panagis Fillipakopoulos *AlphaScreen IC50
25 3,5-DIMETHYLISOXAZOLES AS BET INHIBITORS
26 AlphaScreen IC50 μm COMMERCIAL ISOXAZOLES FOR BRD4
27 ISOXAZOLES FOR BRD4 BDOIA114 BRD4* 0.91 um CBP* ND BDOIA79 BRD4* 3.5 um BDOIA116 BRD4* 1.4 um CBP* 24 um BDOIA115 BRD4* 2.6 um CBP* 3.8 um 2 step synthesis for mixture 3-4 step synthesis for single regioisomer *AlphaScreen IC50
28 ISOXAZOLES FOR CBP 75 compounds selected from Pfizer collection > 50 mg available Published scaffolds Acid, base, neutral R1, R2, R3
29 ISOXAZOLES FOR CBP
30 NON-SELECTIVE ISOXAZOLES
31 SELECTIVE ISOXAZOLES
32 CBP SELECTIVE ISOXAZOLES
33 BDOIA220 MODELLED IN CBP AND BRD4
34 BDOIA220 MODELLED IN CBP AND BRD4
35 CBP SELECTIVE ISOXAZOLE OPTIMIZATION R1 ID CREBBP IC50 CREBBP Tm Shift BRD4_Tm Shift Bn BDOIA BDOIA ID R1 CREBBP IC50 um CREBBP Tm Shift BRD4 Tm Shift BDOIA220 H BDOIA299 3-MeO BDOIA301 4-Me 0.30 R2 ID CREBBP IC50 um CREBBP Tm Shift BRD4 Tm Shift BDOIA BDOIA BDOIA BDOIA BDOIA302 2-Me 0.38 BDOIA313 2-Cl BDOIA314 3-Cl BDOIA315 4-Cl BDOIA316 2-CN BDOIA317 2-OMe BDOIA323 3-CN BDOIA324 4-CN
36 CBP SELECTIVE ISOXAZOLE OPTIMIZATION ID R2 CREBBP IC50 um CREBBP Tm Shift BRD4 Tm Shift BDOIA000322a Piperazine BDOIA000321a Boc-Piperazine BDOIA000320a N-Me-Piperazine BDOIA000319a Azetidine BDOIA000298a Morpholine BDOIA000297a Pyrrolidine 0.57 BDOIA000296a Piperidine
37 BDOIA298 IS A CBP/EP300 INHIBITOR Tm shift assay AlphaScreen IC50 um Fold Selective CREBBP 0.17 BRD CECR % Inhibition CREBBP %I CECR2 %I BRD9 %I BAZ2A %I FALZ %I PHIP %I ATAD2 %I BRPF3 6.3 Selectivity panel: 43 bromodomains
38 kcal mol -1 of injectant kcal/mole of injectant µcal/sec µcal/sec % Inhibition BDOIA298 IS A Potent CBP/EP300 INHIBITOR AlphaScreen IC nm (n =5) BLI Kd 105 nm (like SPR) 100 BDOIA000298a Bottom Top LogEC50 HillSlope EC e [Cmpd] Log M CBP ITC Kd 323 nm BRD4 ITC Kd 1050 nm 0.10 Time (min) Time (min) Data: Data1_NDH Model: OneSites Chi^2/DoF = 2385 N 1.00 ± Sites K 3.10E6 ±1.06E5 M -1 H ±18.40 cal/mol S 9.72 cal/mol/deg -4-6 Data: BRD41E11469_NDH Model: OneSites Chi^2/DoF = 3625 N 1.10 ± K 1.05E6 ±5.09E ±33.81 H S Molar Ratio Molar Ratio
39 BDOIA298 Displaces CBP from Chromatin transfect FRAP bleach assay Δtime zf-taz KIX PAT1 Bromodomain DU KAT11 (HAT) ZZ zf-taz CREB binding NLS Full length protein nuclear, but no ΔFRAP with mutants or inhibitors NLS+3x CREBBP bromodomain (with flanking sequence) gave nuclear expression x3 Tra nsla tion of pdest pcdna5 e GFP CREBBP 2444 aa No SAHA 1.0 um BDOIA298 SAHA 5 um Pre 0 s 5 s 10 s 20 s 30 s Pre 0 s 5 s 10 s 20 s 30 s Pre 0 s 5 s 10 s 20 s 30 s 40 s 40 s 40 s
40 BDOIA298 Displaces CBP from Chromatin Cytotoxicity in HELA CC 50 ~40 μm (non-toxic) CBP Frap IC nm
41 BDOIA298 Attenuates DNA Damage Induced p53 Reporter Expression p53 reporter assay CBP in stress response: IC 50 : 3 μm CBP binding to p53 at the C-terminal acetylated lysine 382 upon DNA damage Results in p53 acetylation-dependent coactivator and transcriptional activation of the cyclin-dependent kinase inhibitor p21 in G1 cell cycle arrest.
42 BDOIA383 IS MORE SELECTIVE FOR CBP
43 BDOIA220 in CBP
44 BDOIA220 in CBP
45 SUMMARY Bromodomains and histones form a switchable protein-protein interaction important in transcriptional regulation. Ligand binding induces a more druggable pocket. Cyclic ureas and isoxazoles are good acetyl lysine mimetics. Selective chemical probes are possible.
46 ACKNOWLEDGEMENTS SGC Aled Edwards Brian Marsden Chas Bountra Chris Wells Duncan Hay Frank von Delft Ildiko Felletar Nicola Burgess-Brown Panagis Filippakopoulos Sarah Picaud Stefan Knapp Susanne Muller-Knapp Tracy Keates Martin Philpott Chris Wells Oleg Fedorov Tony Tumber ICR Lewis Vidler Swen Hoelder Julian Blagg Changchun Discovery Sciences Yue Zhu Mimi Zhang Zhihui Zhao Xuehua Zheng FUNDING PARTNERS Canadian Institutes for Health Research, Canadian Foundation for Innovation, Genome Canada through the Ontario Genomics Institute, GlaxoSmithKline, Knut and Alice Wallenberg Foundation, Pfizer, Eli Lilly, Abbott, Novartis Research Foundation, Ontario Innovation Trust, Ontario Ministry for Research and Innovation, Swedish Agency for Innovation Systems, Swedish Foundation for Strategic Research, and Wellcome Trust.
A Public Private Partnership facilitating Drug Discovery
 A Public Private Partnership facilitating Drug Discovery SGC Toronto SGC Oxford Outline Challenges in Drug discovery: science and organisational/ process We do not know how to pick the right targets Epigenetic
A Public Private Partnership facilitating Drug Discovery SGC Toronto SGC Oxford Outline Challenges in Drug discovery: science and organisational/ process We do not know how to pick the right targets Epigenetic
Selective Targeting of Protein Interactions Mediated by Epigenetic Effector Domains
 Selective Targeting of Protein Interactions Mediated by Epigenetic Effector Domains Stefan Knapp Structural Genomics Consortium Oxford University, Nuffield Department of Clinical Medicine Oxford, United
Selective Targeting of Protein Interactions Mediated by Epigenetic Effector Domains Stefan Knapp Structural Genomics Consortium Oxford University, Nuffield Department of Clinical Medicine Oxford, United
Developing Best in Class BET Inhibitors for Oncology & AI: from Discovery to the Clinic. Kevin G. McLure, PhD EpiCongress July 2014
 Developing Best in Class BET Inhibitors for Oncology & AI: from Discovery to the Clinic Kevin G. McLure, PhD EpiCongress July 2014 Zenith Epigenetics Overview Formed from Resverlogix as an independent
Developing Best in Class BET Inhibitors for Oncology & AI: from Discovery to the Clinic Kevin G. McLure, PhD EpiCongress July 2014 Zenith Epigenetics Overview Formed from Resverlogix as an independent
Epigenetics q&more 01.11
 Laurie. Knight, istockphoto.com Epigenetics 6 Bookmarks About the reading of genes in the Book of Life Prof. Dr. Manfred Jung, Julia M. Wagner, Institute for Pharmaceutical Sciences, Albert-Ludwig-University
Laurie. Knight, istockphoto.com Epigenetics 6 Bookmarks About the reading of genes in the Book of Life Prof. Dr. Manfred Jung, Julia M. Wagner, Institute for Pharmaceutical Sciences, Albert-Ludwig-University
Lecture 8. Eukaryotic gene regulation: post translational modifications of histones
 Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
4 th Oxford Symposium Epigenetic Mechanisms in Health & Disease New Frontiers in Epigenetics
 4 th Oxford Symposium Epigenetic Mechanisms in Health & Disease New Frontiers in Epigenetics 24-26 June 2015 Saïd Business School Park End Street Oxford PROGRAMME Day 1 24 June 2015 Inflammation 09.00
4 th Oxford Symposium Epigenetic Mechanisms in Health & Disease New Frontiers in Epigenetics 24-26 June 2015 Saïd Business School Park End Street Oxford PROGRAMME Day 1 24 June 2015 Inflammation 09.00
PRC2 crystal clear. Matthieu Schapira
 PRC2 crystal clear Matthieu Schapira Epigenetic mechanisms control the combination of genes that are switched on and off in any given cell. In turn, this combination, called the transcriptional program,
PRC2 crystal clear Matthieu Schapira Epigenetic mechanisms control the combination of genes that are switched on and off in any given cell. In turn, this combination, called the transcriptional program,
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Epigenetic Mechanisms
 RCPA Lecture Epigenetic chanisms Jeff Craig Early Life Epigenetics Group, MCRI Dept. of Paediatrics Overview What is epigenetics? Chromatin The epigenetic code What is epigenetics? the interactions of
RCPA Lecture Epigenetic chanisms Jeff Craig Early Life Epigenetics Group, MCRI Dept. of Paediatrics Overview What is epigenetics? Chromatin The epigenetic code What is epigenetics? the interactions of
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Histones modifications and variants
 Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Identification of Oral Bioavailable, Type2 Inhibitors of Discoidin Domain-containing Receptor 1/2 (DDR1/DDR2) using Back-to-Front X-Ray FBDD
 Identification of ral Bioavailable, Type2 Inhibitors of Discoidin Domain-containing Receptor 1/2 (DDR1/DDR2) using Back-to-Front X-Ray FBDD Emiliano Tamanini 26 th Symposium on Medicinal Chemistry in Eastern
Identification of ral Bioavailable, Type2 Inhibitors of Discoidin Domain-containing Receptor 1/2 (DDR1/DDR2) using Back-to-Front X-Ray FBDD Emiliano Tamanini 26 th Symposium on Medicinal Chemistry in Eastern
Expert Intelligence for Better Decisions Epigenetics:
 Expert Intelligence for Better Decisions Epigenetics: Emerging Targets, Available Technologies, Expert Interviews, and an Epigenetic Community Perspective Using This Document Insight Pharma Reports are
Expert Intelligence for Better Decisions Epigenetics: Emerging Targets, Available Technologies, Expert Interviews, and an Epigenetic Community Perspective Using This Document Insight Pharma Reports are
Gene Expression DNA RNA. Protein. Metabolites, stress, environment
 Gene Expression DNA RNA Protein Metabolites, stress, environment 1 EPIGENETICS The study of alterations in gene function that cannot be explained by changes in DNA sequence. Epigenetic gene regulatory
Gene Expression DNA RNA Protein Metabolites, stress, environment 1 EPIGENETICS The study of alterations in gene function that cannot be explained by changes in DNA sequence. Epigenetic gene regulatory
Transcription and chromatin. General Transcription Factors + Promoter-specific factors + Co-activators
 Transcription and chromatin General Transcription Factors + Promoter-specific factors + Co-activators Cofactor or Coactivator 1. work with DNA specific transcription factors to make them more effective
Transcription and chromatin General Transcription Factors + Promoter-specific factors + Co-activators Cofactor or Coactivator 1. work with DNA specific transcription factors to make them more effective
The Epigenome Tools 2: ChIP-Seq and Data Analysis
 The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS
 R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS EPIGENETICS THE STUDY OF CHANGES IN GENE EXPRESSION THAT ARE POTENTIALLY HERITABLE AND THAT DO NOT ENTAIL A
R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS EPIGENETICS THE STUDY OF CHANGES IN GENE EXPRESSION THAT ARE POTENTIALLY HERITABLE AND THAT DO NOT ENTAIL A
Chromatin Structure & Gene activity part 2
 Chromatin Structure & Gene activity part 2 Figure 13.30 Make sure that you understand it. It is good practice for identifying the purpose for various controls. Chromatin remodeling Acetylation helps to
Chromatin Structure & Gene activity part 2 Figure 13.30 Make sure that you understand it. It is good practice for identifying the purpose for various controls. Chromatin remodeling Acetylation helps to
New horizons for small cell lung cancers. Charles Rudin MD PhD
 New horizons for small cell lung cancers Charles Rudin MD PhD Annual deaths (US) US cancer deaths 140000 120000 100000 80000 60000 40000 20000 0 Cancer type Small cell lung cancer: a disease in need of
New horizons for small cell lung cancers Charles Rudin MD PhD Annual deaths (US) US cancer deaths 140000 120000 100000 80000 60000 40000 20000 0 Cancer type Small cell lung cancer: a disease in need of
Epigenetics. Lyle Armstrong. UJ Taylor & Francis Group. f'ci Garland Science NEW YORK AND LONDON
 ... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
Intrinsically Disordered Proteins. Alex Cioffi June 22 nd 2013
 Intrinsically Disordered Proteins Alex Cioffi June 22 nd 2013 Implications in Human Disease Adenovirus Early Region 1A (E1A) and Cancer α-synuclein and Parkinson s Disease Amyloid β and Alzheimer s Disease
Intrinsically Disordered Proteins Alex Cioffi June 22 nd 2013 Implications in Human Disease Adenovirus Early Region 1A (E1A) and Cancer α-synuclein and Parkinson s Disease Amyloid β and Alzheimer s Disease
PI3K Background. The SignalRx R & D pipeline is shown below followed by a brief description of each program:
 PI3K Background The phosphatidylinositol 3-kinase (PI3K) pathway is a key cell signaling node whose dysregulation commonly results in the transformation of normal cells into cancer cells. The role of PI3K
PI3K Background The phosphatidylinositol 3-kinase (PI3K) pathway is a key cell signaling node whose dysregulation commonly results in the transformation of normal cells into cancer cells. The role of PI3K
ACK1 Tyrosine Kinases: A Critical Regulator of Prostate Cancer
 ACK1 Tyrosine Kinases: A Critical Regulator of Prostate Cancer Nupam Mahajan Moffitt Cancer Center Learners Objectives How Androgen Receptor (AR) signaling is accomplished in absence of androgen What are
ACK1 Tyrosine Kinases: A Critical Regulator of Prostate Cancer Nupam Mahajan Moffitt Cancer Center Learners Objectives How Androgen Receptor (AR) signaling is accomplished in absence of androgen What are
SensoLyte 520 HDAC Activity Assay Kit *Fluorimetric*
 SensoLyte 520 HDAC Activity Assay Kit *Fluorimetric* Catalog # 72084 Kit Size 100 Assays (96-well plate) Optimized Performance: This kit is optimized to detect HDAC activity. Enhanced Value: It provides
SensoLyte 520 HDAC Activity Assay Kit *Fluorimetric* Catalog # 72084 Kit Size 100 Assays (96-well plate) Optimized Performance: This kit is optimized to detect HDAC activity. Enhanced Value: It provides
TITLE: TREATMENT OF ENDOCRINE-RESISTANT BREAST CANCER WITH A SMALL MOLECULE C-MYC INHIBITOR
 AD Award Number: W81XWH-13-1-0159 TITLE: TREATMENT OF ENDOCRINE-RESISTANT BREAST CANCER WITH A SMALL MOLECULE C-MYC INHIBITOR PRINCIPAL INVESTIGATOR: Qin Feng CONTRACTING ORGANIZATION: Baylor College of
AD Award Number: W81XWH-13-1-0159 TITLE: TREATMENT OF ENDOCRINE-RESISTANT BREAST CANCER WITH A SMALL MOLECULE C-MYC INHIBITOR PRINCIPAL INVESTIGATOR: Qin Feng CONTRACTING ORGANIZATION: Baylor College of
Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos 3, Hui Yang 1, and Guillermo Garcia-Manero 1
 Genome-wide CHIP-Seq Analysis of Histone Methylation Reveals Modulators of NF- B Signaling And the Histone Demethylase JMJD3 Implicated in Myelodysplastic Syndrome Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos
Genome-wide CHIP-Seq Analysis of Histone Methylation Reveals Modulators of NF- B Signaling And the Histone Demethylase JMJD3 Implicated in Myelodysplastic Syndrome Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos
ISOGENIC CELL LINES. ATCC No. Designation Mutation Parental Cell Line Disease Cancer Model
 THE ESSENTILS OF LIFE SCIENCE RESERCH GLOLLY DELIVERED ISOGENIC CELL LINES TCC CRISPR/CS9 GENE-EDITED ISOGENIC CELL LINES Clinically relevant cell models are critical for studies of molecular and cellular
THE ESSENTILS OF LIFE SCIENCE RESERCH GLOLLY DELIVERED ISOGENIC CELL LINES TCC CRISPR/CS9 GENE-EDITED ISOGENIC CELL LINES Clinically relevant cell models are critical for studies of molecular and cellular
Epigenetics Armstrong_Prelims.indd 1 04/11/2013 3:28 pm
 Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
Host cell activation
 Dept. of Internal Medicine/Infectious and Respiratory Diseases Stefan Hippenstiel Epigenetics as regulator of inflammation Host cell activation LPS TLR NOD2 MDP TRAF IKK NF-κB IL-x, TNFα,... Chromatin
Dept. of Internal Medicine/Infectious and Respiratory Diseases Stefan Hippenstiel Epigenetics as regulator of inflammation Host cell activation LPS TLR NOD2 MDP TRAF IKK NF-κB IL-x, TNFα,... Chromatin
HDAC Cell-Based Activity Assay Kit
 HDAC Cell-Based Activity Assay Kit Item No. 600150 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
HDAC Cell-Based Activity Assay Kit Item No. 600150 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
Are you the way you are because of the
 EPIGENETICS Are you the way you are because of the It s my fault!! Nurture Genes you inherited from your parents? Nature Experiences during your life? Similar DNA Asthma, Autism, TWINS Bipolar Disorders
EPIGENETICS Are you the way you are because of the It s my fault!! Nurture Genes you inherited from your parents? Nature Experiences during your life? Similar DNA Asthma, Autism, TWINS Bipolar Disorders
Chromatin-Based Regulation of Gene Expression
 Chromatin-Based Regulation of Gene Expression.George J. Quellhorst, Jr., PhD.Associate Director, R&D.Biological Content Development Topics to be Discussed Importance of Chromatin-Based Regulation Mechanism
Chromatin-Based Regulation of Gene Expression.George J. Quellhorst, Jr., PhD.Associate Director, R&D.Biological Content Development Topics to be Discussed Importance of Chromatin-Based Regulation Mechanism
Targeting Epigenetic Mechanisms to Reverse Stem Cell Programs in Cancer. Scott A. Armstrong MD, Ph.D.
 Targeting Epigenetic Mechanisms to Reverse Stem Cell Programs in Cancer Scott A. Armstrong MD, Ph.D. Disclosures Scientific advisory boards (Compensation or equity stake) 1. C4 Therapeutics 2. Syros Pharma
Targeting Epigenetic Mechanisms to Reverse Stem Cell Programs in Cancer Scott A. Armstrong MD, Ph.D. Disclosures Scientific advisory boards (Compensation or equity stake) 1. C4 Therapeutics 2. Syros Pharma
Biochemistry 673 Lecture 2 Jason Kahn, UMCP Introduction to steroid hormone receptor (nuclear receptor) signalling
 Biochemistry 673 Lecture 2 Jason Kahn, UMCP Introduction to steroid hormone receptor (nuclear receptor) signalling Resources: Latchman Lodish chapter 10, 20 Helmreich, chapter 11 http://www.nursa.org,
Biochemistry 673 Lecture 2 Jason Kahn, UMCP Introduction to steroid hormone receptor (nuclear receptor) signalling Resources: Latchman Lodish chapter 10, 20 Helmreich, chapter 11 http://www.nursa.org,
Designer Affinity Reagents. Brian Kay
 Designer Affinity Reagents Brian Kay bkay@uic.edu Types of Affinity Reagents Src SH3 domain Lysozyme Src SH3 domain FN3 monobody Peptide Ligand Antibody Fragment Scaffold M13 Bacteriophage 900 nm x 10
Designer Affinity Reagents Brian Kay bkay@uic.edu Types of Affinity Reagents Src SH3 domain Lysozyme Src SH3 domain FN3 monobody Peptide Ligand Antibody Fragment Scaffold M13 Bacteriophage 900 nm x 10
Epigenomics. Ivana de la Serna Block Health Science
 Epigenomics Ivana de la Serna Block Health Science 388 383-4111 ivana.delaserna@utoledo.edu Outline 1. Epigenetics-definition and overview 2. DNA methylation/hydroxymethylation 3. Histone modifications
Epigenomics Ivana de la Serna Block Health Science 388 383-4111 ivana.delaserna@utoledo.edu Outline 1. Epigenetics-definition and overview 2. DNA methylation/hydroxymethylation 3. Histone modifications
DOWNLOAD OR READ : PROTEIN METHYLTRANSFERASES PDF EBOOK EPUB MOBI
 DOWNLOAD OR READ : PROTEIN METHYLTRANSFERASES PDF EBOOK EPUB MOBI Page 1 Page 2 protein methyltransferases protein methyltransferases pdf protein methyltransferases N-alpha methyltransferases transfer
DOWNLOAD OR READ : PROTEIN METHYLTRANSFERASES PDF EBOOK EPUB MOBI Page 1 Page 2 protein methyltransferases protein methyltransferases pdf protein methyltransferases N-alpha methyltransferases transfer
Epigenetics and Toxicology
 Epigenetics and Toxicology Aline.deconti@fda.hhs.gov Division of Biochemical Toxicology National Center for Toxicology Research U.S.-Food and Drug Administration The views expressed in this presentation
Epigenetics and Toxicology Aline.deconti@fda.hhs.gov Division of Biochemical Toxicology National Center for Toxicology Research U.S.-Food and Drug Administration The views expressed in this presentation
Designer Affinity Reagents. Reagents. Types of Affinity. M13 Bacteriophage. 900 nm x 10 nm. Brian Kay Src SH3 domain.
 Designer Affinity Reagents Brian Kay bkay@uic.edu Types of Affinity Reagents Src SH3 domain Lysozyme Src SH3 domain FN3 monobody Peptide Ligand Antibody Fragment Scaffold M13 Bacteriophage 900 nm 10 nm
Designer Affinity Reagents Brian Kay bkay@uic.edu Types of Affinity Reagents Src SH3 domain Lysozyme Src SH3 domain FN3 monobody Peptide Ligand Antibody Fragment Scaffold M13 Bacteriophage 900 nm 10 nm
Epigenetics: The Future of Psychology & Neuroscience. Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1
 Epigenetics: The Future of Psychology & Neuroscience Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1 Nature versus Nurture Despite the belief that the Nature vs. Nurture
Epigenetics: The Future of Psychology & Neuroscience Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1 Nature versus Nurture Despite the belief that the Nature vs. Nurture
Beads-on-a- The 30nm Fibre Active Chromosome The Metaphase Chromosome. Less active genes During interphase During cell division
 Overview of Epigenetics Figure 2.2. The Increasing Structural Complexity of Genetic Information from the Double-Helical Structure of DNA, through Nucleosome and Chromatin Structures, to the Chromosome
Overview of Epigenetics Figure 2.2. The Increasing Structural Complexity of Genetic Information from the Double-Helical Structure of DNA, through Nucleosome and Chromatin Structures, to the Chromosome
EPIGENOMICS PROFILING SERVICES
 EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
INTERACTION DRUG BODY
 INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
Genomic Methods in Cancer Epigenetic Dysregulation
 Genomic Methods in Cancer Epigenetic Dysregulation Clara, Lyon 2018 Jacek Majewski, Associate Professor Department of Human Genetics, McGill University Montreal, Canada A few words about my lab Genomics
Genomic Methods in Cancer Epigenetic Dysregulation Clara, Lyon 2018 Jacek Majewski, Associate Professor Department of Human Genetics, McGill University Montreal, Canada A few words about my lab Genomics
Gene Regulation Part 2
 Michael Cummings Chapter 9 Gene Regulation Part 2 David Reisman University of South Carolina Other topics in Chp 9 Part 2 Protein folding diseases Most diseases are caused by mutations in the DNA that
Michael Cummings Chapter 9 Gene Regulation Part 2 David Reisman University of South Carolina Other topics in Chp 9 Part 2 Protein folding diseases Most diseases are caused by mutations in the DNA that
Personalized Therapeutics The Power of Epigenetics. Company Overview. June 2014
 Personalized Therapeutics The Power of Epigenetics Company Overview June 2014 2013 Accomplishments Forward Looking Statements This presentation contains forward-looking statements that involve substantial
Personalized Therapeutics The Power of Epigenetics Company Overview June 2014 2013 Accomplishments Forward Looking Statements This presentation contains forward-looking statements that involve substantial
Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc
 Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc P. Matthias, 16 April 2014 DNA methylation basics Acetylation Acetyltransferases/Deacetylases
Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc P. Matthias, 16 April 2014 DNA methylation basics Acetylation Acetyltransferases/Deacetylases
Lecture 10. Eukaryotic gene regulation: chromatin remodelling
 Lecture 10 Eukaryotic gene regulation: chromatin remodelling Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
Lecture 10 Eukaryotic gene regulation: chromatin remodelling Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature23267 Discussion Our findings reveal unique roles for the methylation states of histone H3K9 in RNAi-dependent and - independent heterochromatin formation. Clr4 is the sole S. pombe enzyme
doi:10.1038/nature23267 Discussion Our findings reveal unique roles for the methylation states of histone H3K9 in RNAi-dependent and - independent heterochromatin formation. Clr4 is the sole S. pombe enzyme
This PDF file includes: Supplementary Figures 1 to 6 Supplementary Tables 1 to 2 Supplementary Methods Supplementary References
 Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
TITLE: Inhibitors of Histone Deacetylases for Radiosensitization of Prostate Cancer
 AD Award Number: W81XWH-04-1-0170 TITLE: Inhibitors of Histone Deacetylases for Radiosensitization of Prostate Cancer PRINCIPAL INVESTIGATOR: Mira O. Jung, Ph.D. CONTRACTING ORGANIZATION: Georgetown University
AD Award Number: W81XWH-04-1-0170 TITLE: Inhibitors of Histone Deacetylases for Radiosensitization of Prostate Cancer PRINCIPAL INVESTIGATOR: Mira O. Jung, Ph.D. CONTRACTING ORGANIZATION: Georgetown University
SUPPLEMENTARY INFORMATION
 Supplementary Discussion The cell cycle machinery and the DNA damage response network are highly interconnected and co-regulated in assuring faithful duplication and partition of genetic materials into
Supplementary Discussion The cell cycle machinery and the DNA damage response network are highly interconnected and co-regulated in assuring faithful duplication and partition of genetic materials into
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system
 Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Targeting MAT2A in MTAP-deleted Cancers
 Targeting MAT2A in MTAP-deleted Cancers Presented at the American Association for Cancer Research (AACR) Annual Meeting, April 14-18, 2018, Chicago, IL, USA 1 Acknowledgements Agios 2017 Founders Day Retreat
Targeting MAT2A in MTAP-deleted Cancers Presented at the American Association for Cancer Research (AACR) Annual Meeting, April 14-18, 2018, Chicago, IL, USA 1 Acknowledgements Agios 2017 Founders Day Retreat
Data Sheet. Fluorogenic HDAC 8 Assay Kit Catalog #: 50068
 Data Sheet Fluorogenic HDAC 8 Assay Kit Catalog #: 50068 DESCRIPTION: The Fluorogenic HDAC 8 Assay Kit is a complete assay system designed to measure histone deacetylase (HDAC) 8 activity for screening
Data Sheet Fluorogenic HDAC 8 Assay Kit Catalog #: 50068 DESCRIPTION: The Fluorogenic HDAC 8 Assay Kit is a complete assay system designed to measure histone deacetylase (HDAC) 8 activity for screening
Dominic J Smiraglia, PhD Department of Cancer Genetics. DNA methylation in prostate cancer
 Dominic J Smiraglia, PhD Department of Cancer Genetics DNA methylation in prostate cancer Overarching theme Epigenetic regulation allows the genome to be responsive to the environment Sets the tone for
Dominic J Smiraglia, PhD Department of Cancer Genetics DNA methylation in prostate cancer Overarching theme Epigenetic regulation allows the genome to be responsive to the environment Sets the tone for
Supplementary Figure 1. MAT IIα is Acetylated at Lysine 81.
 IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
HDAC1 Inhibitor Screening Assay Kit
 HDAC1 Inhibitor Screening Assay Kit Item No. 10011564 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
HDAC1 Inhibitor Screening Assay Kit Item No. 10011564 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
Comparative effect of Various HDAC-inhibitors in-vitro on T- Cell Lymphoma cell lines alone and in combination with conventional anti-cancer drugs
 Comparative effect of Various HDAC-inhibitors in-vitro on T- Cell Lymphoma cell lines alone and in combination with conventional anti-cancer drugs Arshad H. Banday Mentor:Dr. Francisco Hernandez-Illizaliturri
Comparative effect of Various HDAC-inhibitors in-vitro on T- Cell Lymphoma cell lines alone and in combination with conventional anti-cancer drugs Arshad H. Banday Mentor:Dr. Francisco Hernandez-Illizaliturri
AlphaScreen TNFα Binding Assay Kit: A Homogeneous, Sensitive and High-Throughput Assay for Screening TNFα Receptors
 AlphaScreen TNFα Binding Assay Kit: A Homogeneous, Sensitive and High-Throughput Assay for Screening TNFα Receptors Bouchard N., Legault M. and Wenham D. PerkinElmer BioSignal, 1744 William, suite 3, Montréal,
AlphaScreen TNFα Binding Assay Kit: A Homogeneous, Sensitive and High-Throughput Assay for Screening TNFα Receptors Bouchard N., Legault M. and Wenham D. PerkinElmer BioSignal, 1744 William, suite 3, Montréal,
SUPPLEMENTAL INFORMATION. Evolving Spindlin1 Small Molecule Inhibitors Using Protein Microarrays
 SUPPLEMETAL IFRMATI Evolving Spindlin1 Small Molecule Inhibitors Using Protein Microarrays arkhyun Bae 1, Monica Viviano 2, Xiaonan Su 3, Jie Lu 4, Donghang Cheng 1, Cari Sagum 1, Sabrina Castellano 2,6,
SUPPLEMETAL IFRMATI Evolving Spindlin1 Small Molecule Inhibitors Using Protein Microarrays arkhyun Bae 1, Monica Viviano 2, Xiaonan Su 3, Jie Lu 4, Donghang Cheng 1, Cari Sagum 1, Sabrina Castellano 2,6,
Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure
 Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure Frederick E. Domann, Ph.D. Associate Professor of Radiation Oncology The University of Iowa Iowa City,
Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure Frederick E. Domann, Ph.D. Associate Professor of Radiation Oncology The University of Iowa Iowa City,
Molecular Cell Biology. Prof. D. Karunagaran. Department of Biotechnology. Indian Institute of Technology Madras
 Molecular Cell Biology Prof. D. Karunagaran Department of Biotechnology Indian Institute of Technology Madras Module-9 Molecular Basis of Cancer, Oncogenes and Tumor Suppressor Genes Lecture 6 Epigenetics
Molecular Cell Biology Prof. D. Karunagaran Department of Biotechnology Indian Institute of Technology Madras Module-9 Molecular Basis of Cancer, Oncogenes and Tumor Suppressor Genes Lecture 6 Epigenetics
Identification of a First-In-Class PRMT5 Inhibitor with Potent In Vitro and In Vivo Activity in Preclinical Models of Mantle Cell Lymphoma
 Identification of a First-In-Class PRMT5 Inhibitor with Potent In Vitro and In Vivo Activity in Preclinical Models of Mantle Cell Lymphoma Elayne Penebre, Kristy G Kuplast, Christina R Majer, P Ann Boriak-Sjodin,
Identification of a First-In-Class PRMT5 Inhibitor with Potent In Vitro and In Vivo Activity in Preclinical Models of Mantle Cell Lymphoma Elayne Penebre, Kristy G Kuplast, Christina R Majer, P Ann Boriak-Sjodin,
HDAC1 Inhibitor Screening Assay Kit
 HDAC1 Inhibitor Screening Assay Kit Catalog Number KA1320 96 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
HDAC1 Inhibitor Screening Assay Kit Catalog Number KA1320 96 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
Inhibitors of Methionine Aminopeptidase-2 2 in the Treatment of Non-Hodgkin
 Inhibitors of Methionine Aminopeptidase-2 2 in the Treatment of Non-Hodgkin Hodgkin s s Lymphoma Targeted Cancer Therapies William Westlin, Ph.D. Vice President, Preclinical Research Discovery of Fumagillin
Inhibitors of Methionine Aminopeptidase-2 2 in the Treatment of Non-Hodgkin Hodgkin s s Lymphoma Targeted Cancer Therapies William Westlin, Ph.D. Vice President, Preclinical Research Discovery of Fumagillin
RAS Genes. The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes.
 ۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
Unique, expert portfolio
 EPIGENETICS Enzymes Kits Cell Lines Screening Services INNOVATIVE PRODUCTS TO FUEL YOUR EXPERIMENTS Unique, expert portfolio www.bpsbioscience.com HISTONE METHYLTRANSFERASES HISTONE DEMETHYLASES ACETYLTRANSFERASES
EPIGENETICS Enzymes Kits Cell Lines Screening Services INNOVATIVE PRODUCTS TO FUEL YOUR EXPERIMENTS Unique, expert portfolio www.bpsbioscience.com HISTONE METHYLTRANSFERASES HISTONE DEMETHYLASES ACETYLTRANSFERASES
EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric)
 EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric)
EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric)
Part-4. Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death
 Part-4 Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death 95 1. Introduction The process of replicating DNA and dividing cells can be described as a series of coordinated
Part-4 Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death 95 1. Introduction The process of replicating DNA and dividing cells can be described as a series of coordinated
This article appeared in a journal published by Elsevier. The attached copy is furnished to the author for internal non-commercial research and
 This article appeared in a journal published by Elsevier. The attached copy is furnished to the author for internal non-commercial research and education use, including for instruction at the authors institution
This article appeared in a journal published by Elsevier. The attached copy is furnished to the author for internal non-commercial research and education use, including for instruction at the authors institution
The Role of Protein Domains in Cell Signaling
 The Role of Protein Domains in Cell Signaling Growth Factors RTK Ras Shc PLCPLC-γ P P P P P P GDP Ras GTP PTB CH P Sos Raf SH2 SH2 PI(3)K SH3 SH3 MEK Grb2 MAPK Phosphotyrosine binding domains: PTB and
The Role of Protein Domains in Cell Signaling Growth Factors RTK Ras Shc PLCPLC-γ P P P P P P GDP Ras GTP PTB CH P Sos Raf SH2 SH2 PI(3)K SH3 SH3 MEK Grb2 MAPK Phosphotyrosine binding domains: PTB and
HDAC Assay Kit. (Fluorescent) (version B1) Catalog No
 HDAC Assay Kit (Fluorescent) (version B1) Catalog No. 56200 Active Motif North America 1914 Palomar Oaks Way, Suite 150 Carlsbad, California 92008, USA Toll free: 877 222 9543 Telephone: 760 431 1263 Fax:
HDAC Assay Kit (Fluorescent) (version B1) Catalog No. 56200 Active Motif North America 1914 Palomar Oaks Way, Suite 150 Carlsbad, California 92008, USA Toll free: 877 222 9543 Telephone: 760 431 1263 Fax:
Minjia Tan, Ph.D Shanghai Institute of Materia Medica, Chinese Academy of Sciences
 Lysine Glutarylation Is a Protein Posttranslational Modification Regulated by SIRT5 Minjia Tan, Ph.D Shanghai Institute of Materia Medica, Chinese Academy of Sciences Lysine: the most frequently modified
Lysine Glutarylation Is a Protein Posttranslational Modification Regulated by SIRT5 Minjia Tan, Ph.D Shanghai Institute of Materia Medica, Chinese Academy of Sciences Lysine: the most frequently modified
Epigene.cs: What is it and how it effects our health? Overview. Dr. Bill Stanford, PhD OFawa Hospital Research Ins.tute University of OFawa
 Epigene.cs: What is it and how it effects our health? Dr. Bill Stanford, PhD OFawa Hospital Research Ins.tute University of OFawa Overview Basic Background Epigene.cs in general Epigene.cs in cancer Epigene.cs
Epigene.cs: What is it and how it effects our health? Dr. Bill Stanford, PhD OFawa Hospital Research Ins.tute University of OFawa Overview Basic Background Epigene.cs in general Epigene.cs in cancer Epigene.cs
Epigenetic Inheritance
 (2) The role of Epigenetic Inheritance Lamarck Revisited Lamarck was incorrect in thinking that the inheritance of acquired characters is the main mechanism of evolution (Natural Selection more common)
(2) The role of Epigenetic Inheritance Lamarck Revisited Lamarck was incorrect in thinking that the inheritance of acquired characters is the main mechanism of evolution (Natural Selection more common)
p.r623c p.p976l p.d2847fs p.t2671 p.d2847fs p.r2922w p.r2370h p.c1201y p.a868v p.s952* RING_C BP PHD Cbp HAT_KAT11
 ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
Total Histone H3 Acetylation Detection Fast Kit (Colorimetric)
 Total Histone H3 Acetylation Detection Fast Kit (Colorimetric) Catalog Number KA1538 48 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use...
Total Histone H3 Acetylation Detection Fast Kit (Colorimetric) Catalog Number KA1538 48 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use...
Supplementary Information. Dual kinase/bromodomain inhibitors for rationally designed polypharmacology
 Supplementary Information Dual kinase/bromodomain inhibitors for rationally designed polypharmacology Pietro Ciceri 4,7, Susanne Müller 1,3,7, Alison O Mahony 5,7, Oleg Fedorov 3, Panagis Filippakopoulos
Supplementary Information Dual kinase/bromodomain inhibitors for rationally designed polypharmacology Pietro Ciceri 4,7, Susanne Müller 1,3,7, Alison O Mahony 5,7, Oleg Fedorov 3, Panagis Filippakopoulos
EZH2 Inhibitors as Novel Cancer Therapeutics!!!! Robert A. Copeland, Ph.D.! Epizyme, Inc.!!! 20 November 2014!
 EZH2 Inhibitors as Novel Cancer Therapeutics Robert A. Copeland, Ph.D. Epizyme, Inc. 20 November 2014 2013 Accomplishments Disclosure Information EORTC-NCI-AACR Meeting, 20 November 2014 Robert A. Copeland,
EZH2 Inhibitors as Novel Cancer Therapeutics Robert A. Copeland, Ph.D. Epizyme, Inc. 20 November 2014 2013 Accomplishments Disclosure Information EORTC-NCI-AACR Meeting, 20 November 2014 Robert A. Copeland,
September 20, Submitted electronically to: Cc: To Whom It May Concern:
 History Study (NOT-HL-12-147), p. 1 September 20, 2012 Re: Request for Information (RFI): Building a National Resource to Study Myelodysplastic Syndromes (MDS) The MDS Cohort Natural History Study (NOT-HL-12-147).
History Study (NOT-HL-12-147), p. 1 September 20, 2012 Re: Request for Information (RFI): Building a National Resource to Study Myelodysplastic Syndromes (MDS) The MDS Cohort Natural History Study (NOT-HL-12-147).
Conference Reports for NATAP. S/GSK Integrase Inhibitor Resistance Profile
 Conference Reports for NATAP EACS - 12th European AIDS Conference November 11-14, 2009 Cologne, Germany Back S/GSK1349572 Integrase Inhibitor Resistance Profile Reported by Jules Levin EACS Oct 31-Nov
Conference Reports for NATAP EACS - 12th European AIDS Conference November 11-14, 2009 Cologne, Germany Back S/GSK1349572 Integrase Inhibitor Resistance Profile Reported by Jules Levin EACS Oct 31-Nov
Harnessing Killer T Cells
 Harnessing Killer T Cells Julie Nielsen, PhD BC Cancer Agency May 2, 2015 Overview The immune response to cancer What T cells are & how they work Immune-based therapies for cancer Using T cells to target
Harnessing Killer T Cells Julie Nielsen, PhD BC Cancer Agency May 2, 2015 Overview The immune response to cancer What T cells are & how they work Immune-based therapies for cancer Using T cells to target
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression. Chromatin Array of nuc
 Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Design and Synthesis of Chemical Probes for the BRPF Bromodomains
 Design and Synthesis of Chemical Probes for the BRPF Bromodomains Niall Michael Igoe September 2016 A thesis submitted for the degree of Doctor of Philosophy of University College London Department of
Design and Synthesis of Chemical Probes for the BRPF Bromodomains Niall Michael Igoe September 2016 A thesis submitted for the degree of Doctor of Philosophy of University College London Department of
Assay Report. Histone Deacetylase (HDAC) Inhibitor Assays Enzymatic Study of Compounds from Client
 Assay Report Histone Deacetylase (HDAC) Inhibitor Assays Enzymatic Study of Compounds from Client Page 1 of 27 Client_HDAC _Year Month Day 1 Client_HDAC_Year Month Year HDAC Inhibitor Assays Study Sponsor:
Assay Report Histone Deacetylase (HDAC) Inhibitor Assays Enzymatic Study of Compounds from Client Page 1 of 27 Client_HDAC _Year Month Day 1 Client_HDAC_Year Month Year HDAC Inhibitor Assays Study Sponsor:
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3
 Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
Post-translational modifications of proteins in gene regulation under hypoxic conditions
 203 Review Article Post-translational modifications of proteins in gene regulation under hypoxic conditions 1, 2) Olga S. Safronova 1) Department of Cellular Physiological Chemistry, Tokyo Medical and
203 Review Article Post-translational modifications of proteins in gene regulation under hypoxic conditions 1, 2) Olga S. Safronova 1) Department of Cellular Physiological Chemistry, Tokyo Medical and
Epigenetic Variation in Human Health and Disease
 Epigenetic Variation in Human Health and Disease Michael S. Kobor Centre for Molecular Medicine and Therapeutics Department of Medical Genetics University of British Columbia www.cmmt.ubc.ca Understanding
Epigenetic Variation in Human Health and Disease Michael S. Kobor Centre for Molecular Medicine and Therapeutics Department of Medical Genetics University of British Columbia www.cmmt.ubc.ca Understanding
DNA methylation: a potential clinical biomarker for the detection of human cancers
 DNA methylation: a potential clinical biomarker for the detection of human cancers Name: Tong Samuel Supervisor: Zigui CHEN Date: 1 st December 2016 Department: Microbiology Source: cited from Jakubowski,
DNA methylation: a potential clinical biomarker for the detection of human cancers Name: Tong Samuel Supervisor: Zigui CHEN Date: 1 st December 2016 Department: Microbiology Source: cited from Jakubowski,
Activation of cellular proto-oncogenes to oncogenes. How was active Ras identified?
 Dominant Acting Oncogenes Eugene E. Marcantonio, M.D. Ph.D. Oncogenes are altered forms of normal cellular genes called proto-oncogenes that are involved in pathways regulating cell growth, differentiation,
Dominant Acting Oncogenes Eugene E. Marcantonio, M.D. Ph.D. Oncogenes are altered forms of normal cellular genes called proto-oncogenes that are involved in pathways regulating cell growth, differentiation,
Brd4 Coactivates Transcriptional Activation of NF- B via Specific Binding to Acetylated RelA
 MOLECULAR AND CELLULAR BIOLOGY, Mar. 2009, p. 1375 1387 Vol. 29, No. 5 0270-7306/09/$08.00 0 doi:10.1128/mcb.01365-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Brd4 Coactivates
MOLECULAR AND CELLULAR BIOLOGY, Mar. 2009, p. 1375 1387 Vol. 29, No. 5 0270-7306/09/$08.00 0 doi:10.1128/mcb.01365-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Brd4 Coactivates
Molecular Hematopathology Leukemias I. January 14, 2005
 Molecular Hematopathology Leukemias I January 14, 2005 Chronic Myelogenous Leukemia Diagnosis requires presence of Philadelphia chromosome t(9;22)(q34;q11) translocation BCR-ABL is the result BCR on chr
Molecular Hematopathology Leukemias I January 14, 2005 Chronic Myelogenous Leukemia Diagnosis requires presence of Philadelphia chromosome t(9;22)(q34;q11) translocation BCR-ABL is the result BCR on chr
MYC Translocations In Multiple Myeloma Involve Recruitment Of Enhancer Elements Resulting In Over- Expression and Decreased Overall Survival
 in partnership with MYC Translocations In Multiple Myeloma Involve Recruitment Of Enhancer Elements Resulting In Over- Expression and Decreased Overall Survival Brian A Walker Centre for Myeloma Research,
in partnership with MYC Translocations In Multiple Myeloma Involve Recruitment Of Enhancer Elements Resulting In Over- Expression and Decreased Overall Survival Brian A Walker Centre for Myeloma Research,
EPIGENTEK. EpiQuik Global Acetyl Histone H3K27 Quantification Kit (Colorimetric) Base Catalog # P-4059 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Global Acetyl Histone H3K27 Quantification Kit (Colorimetric) Base Catalog # P-4059 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Global Acetyl Histone H3K27 Quantification Kit (Colorimetric)
EpiQuik Global Acetyl Histone H3K27 Quantification Kit (Colorimetric) Base Catalog # P-4059 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Global Acetyl Histone H3K27 Quantification Kit (Colorimetric)
On the way of revealing coactivator complexes cross talk during transcriptional activation
 DOI 10.1186/s13578-016-0081-y Cell & Bioscience REVIEW Open Access On the way of revealing coactivator complexes cross talk during transcriptional activation Aleksey N. Krasnov *, Marina Yu. Mazina, Julia
DOI 10.1186/s13578-016-0081-y Cell & Bioscience REVIEW Open Access On the way of revealing coactivator complexes cross talk during transcriptional activation Aleksey N. Krasnov *, Marina Yu. Mazina, Julia
CRC 992 Symposium on Medical Epigenetics 2018
 CRC 992 Symposium on Medical Epigenetics 2018 Monday, March 12 th Wednesday, March 14 th Agenda Lecture Hall Otto-Krayer-Haus Albertstr. 25 79104 Freiburg, Germany Day 1 Monday, March 12 th 2018, 14:30-19:30
CRC 992 Symposium on Medical Epigenetics 2018 Monday, March 12 th Wednesday, March 14 th Agenda Lecture Hall Otto-Krayer-Haus Albertstr. 25 79104 Freiburg, Germany Day 1 Monday, March 12 th 2018, 14:30-19:30
Repression of IP-10 by Interactions between Histone Deacetylation and Hypermethylation in Idiopathic Pulmonary Fibrosis
 MOLECULAR AND CELLULAR BIOLOGY, June 2010, p. 2874 2886 Vol. 30, No. 12 0270-7306/10/$12.00 doi:10.1128/mcb.01527-09 Copyright 2010, American Society for Microbiology. All Rights Reserved. Repression of
MOLECULAR AND CELLULAR BIOLOGY, June 2010, p. 2874 2886 Vol. 30, No. 12 0270-7306/10/$12.00 doi:10.1128/mcb.01527-09 Copyright 2010, American Society for Microbiology. All Rights Reserved. Repression of
LPS LPS P6 - + Supplementary Fig. 1.
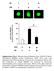 P6 LPS - - - + + + - LPS + + - - P6 + Supplementary Fig. 1. Pharmacological inhibition of the JAK/STAT blocks LPS-induced HMGB1 nuclear translocation. RAW 267.4 cells were stimulated with LPS in the absence
P6 LPS - - - + + + - LPS + + - - P6 + Supplementary Fig. 1. Pharmacological inhibition of the JAK/STAT blocks LPS-induced HMGB1 nuclear translocation. RAW 267.4 cells were stimulated with LPS in the absence
