Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos 3, Hui Yang 1, and Guillermo Garcia-Manero 1
|
|
|
- Bridget Parsons
- 5 years ago
- Views:
Transcription
1 Genome-wide CHIP-Seq Analysis of Histone Methylation Reveals Modulators of NF- B Signaling And the Histone Demethylase JMJD3 Implicated in Myelodysplastic Syndrome Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos 3, Hui Yang 1, and Guillermo Garcia-Manero 1 1. Department of Leukemia, MD Anderson Cancer Center, Houston, TX 2. Human Genome Center, Baylor College of Medicine, Houston, TX 3. Department of Hematopathology, MD Anderson Cancer Center, Houston, TX
2 CHIP-Seq in MDS: Hypothesis Histone H3 Khairul 2009 Analysis of activating histone modification (H3K4me3) can provide a stable view of gene activation in MDS This will allow identification of altered signaling modules and/ or related epigenetic regulators as new therapeutic and/or prognostic targets in this disease
3 Chromatin Immunoprecipitation (CHIP) CHIP-Seq: Methodology Direct Sequencing (Seq) Bioinformatics
4 CHIP-Seq Strategy PATIENT CHARACTERISTICS Dx IPSS CYTO MDS BM (N=5) Control BM (N=4) RAEB INT-2-5/5q- & -7/7q- CMML INT-2 DIP RAEB INT-1 DIP RAEB INT-1 MIS RAEB-T HR DIP CD34+ CD34- CD34+ CD34- Ch-IP with H3K4me3 Antibody Solexa Sequencing of H3K4me3 Rich Chromatin Fragments Bio-info analysis Gene promoters specifically bearing high levels of H3K4me3 in MDS (Potential aberrantly active genes in MDS) Validations (in a large cohort of patient samples)
5 H3K4me3 CHIP-Seq in MDS criteria for peak selection K4me3 signals: within 2kb to transcription starting site (TSS) of a known gene K4me3 up-regulation: MDS vs control > 3 fold p value < 10-6 chr19
6 Validation of Gene Expression in Primary MDS CD34+ Cells 350 Relative RNA level in CD34+ (MDS vs control) P< C5AR1 FPR1 FPR2 AQP9 PRX FYB C19ORF133 FCAR IL8RA N=57 (previously reported CD34+ samples) Heinrichs et al. Leukemia 2008
7 H3K4me3 CHIP-Seq identify NF- B Effectors in MDS CD34+ Cells NF- B Genes involved in NF- B activation (11/36) C5AR1, FPR1, FPR2, FYB, FCGR2A, MEFV, IL8-RB, TYROBP, PTAFR, BCL2A1, NCF2 phospho-nf B-p65 *
8 Summary of Part I Defect of Epigenetic Regulation? NF- B activators promoter H3K4me3 MDS CD34+ NF- B activators expression NF- B signaling
9 Histone Demethylase JMJD3 is aberrantly up-regulated in MDS CD34+ Cells Relative Expression in CD34+ (MDS v.s. ctrl) JARID1A in MDS CD34+ cells --- Medium correlation with CHIP-Seq identified genes --- Strong correlation with PTAFR and TYROBP (NF- B effectors) JARID1B JARID1C JARID1D JHDM1A JMJD2A JMJD2B JMJD2C UTX UTY P<0.01 AJmjc domain H3K27 demethylase A positive regulator of H3K4me3 A direct transcriptional target of NF- B JMJD3 JMJD1A JMJD1B H3K4me H3K36me H3K9/36me H3K27me H3K9me1/2
10 Hypothesis: Positive Signaling Circuitry Between JMJD3 And NF- B JMJD3 (positive regulator of H3K4me3) (NF- B target) NF- B activators promoter H3K4me3 MDS CD34+ NF- B activators expression NF- B signaling
11 JMJD3 positively regulates NF- B JMJD3 knock-down: --- NF- B effectors suppressed --- NF- B signal inhibited phospho-nf B-p65 C5AR1 FPR1 FPR2 FYB FCGR2A MEFV IL8RB TYROBP PTAFR BCL2A1 NCF2 PRX AQP9 JMJD3 JMJD3? NF- B activators promoter H3K4me3 JMJD3 overexpression: --- NF- B effectors up-reuglated --- promoter H3k4me3 increase MDS CD34+ NF- B activators expression NF- B signaling
12 NF- B effectors positively regulate JMJD3 Knock-down 4 NF- B effectors: --- JMJD3 expression suppressed --- Dissociation of NF- B from JMJD3 control sirna sirna targeting NF-kB effectors sirna targeting NF-kB effectors control sirna C5AR1 PTAFR FPR2 TYROBP JMJD3 JMJD3 NF- B activators promoter H3K4me3 MDS CD34+? NF- B activators expression NF- B signaling
13 Conclusion 1. CHIP-Seq is applied to a human disease, identifying unique H3K4me3 code in MDS 2. This analysis reveals effectors of NF- B and JMJD3 as important signal modulators in MDS CD34+ cells 3. Future plans systematic analysis of clinical implications of JMJD3 and NF- B effectors in large cohort/ subtypes of MDS Confirm the direct association between JMJD3 and CHIP-Seq detected gene promoters in MDS validation/ analysis of CHIP-Seq data in MDS CD34- cells
14 Acknowledgement University of Texas MD Anderson Cancer Center Baylor College of Medicine Human Genome Center Dr. R Chen Group Leukemia Department Dr. G Garcia-Manero Group Hematophathology Department Dr. C Bueso-Ramos Group Bioinformatics Department Dr. H Yao Dr. J Wang Lixia Diao Dr. M Zhang This project is supported by NIH and LLSA
15 JMJD3 is H3K27 histone demethylase Involved in Up-regulation of H3K4me3 JMJD3 is a Jmjc domain H3K27 demethylase JMJD3 physically integrates in MLL2 complex and positively regulates H3K4me3 JMJD3 is crucial in lineage determination and gene activation during inflammatory response JMJD3 is a direct transcriptional target of NF- B Lan et al Nature 2007 De Santa et al Cell 2007
16 CHIP-Seq in MDS: Introduction MDS consists of a group of hematopoietic stem cell disorders characterized by: Inefficient hematopoiesis Increase rate of transformation to AML At cellular and molecular level, MDS is characterized by: Frequent cytogenetic alterations Aberrant DNA methylation Low prevalence of genetic lesions Research in MDS at molecular level is limited by: Lack of MDS cell lines Paucity of animal models
17 H3K4me3 CHIP-Seq in Hela Cells Overall Signal Distribution Chip-Seq Overlapping Chip-Chip Chip-Seq Only Chip-Chip Only ,000-20,000-10, ,000 20,000 30,000 Base pairs from TSS Alignment with CHIP-chip 21q22.11 IgG control ChIP-Seq: H3K4me3 ChIP-chip: H3K4me3 Known Genes
18 Molecular Research of MDS Search for genetic/ epigenetic alterations has been less informative, although recent SNP arrays and analysis of aberrant CpG methylation in MDS may help to identify genetic/ epigenetic lesions. -- Diverse molecular backgrounds; -- Lack of cell lines and representative animal models;
Innate Immunity Dysregulation in Myelodysplastic Syndromes. CONTRACTING ORGANIZATION: University of Texas MD Anderson Cancer Center Houston, TX 77030
 AWARD NUMBER: W81XWH-12-1-0221 TITLE: Innate Immunity Dysregulation in Myelodysplastic Syndromes PRINCIPAL INVESTIGATOR: Yue Wei CONTRACTING ORGANIZATION: University of Texas MD Anderson Cancer Center
AWARD NUMBER: W81XWH-12-1-0221 TITLE: Innate Immunity Dysregulation in Myelodysplastic Syndromes PRINCIPAL INVESTIGATOR: Yue Wei CONTRACTING ORGANIZATION: University of Texas MD Anderson Cancer Center
Therapeutic options after first line treatment failure
 Therapeutic options after first line treatment failure Orlando ASH 215 Guillermo Garcia-Manero MD Professor of Medicine Chief Section of MDS Deputy Chair, Translational Research Department of Leukemia
Therapeutic options after first line treatment failure Orlando ASH 215 Guillermo Garcia-Manero MD Professor of Medicine Chief Section of MDS Deputy Chair, Translational Research Department of Leukemia
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes
 Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data
 Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier ChIP-seq experiment To
Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier ChIP-seq experiment To
Therapeutic and Prognostic Role of Epigenetic Abnormalities in MDS. Stephen D. Nimer, MD Sylvester Comprehensive Cancer Center December 5, 2014
 Therapeutic and Prognostic Role of Epigenetic Abnormalities in MDS Stephen D. Nimer, MD Sylvester Comprehensive Cancer Center December 5, 2014 DISCLOSURE I have no relevant financial relationships to disclose.
Therapeutic and Prognostic Role of Epigenetic Abnormalities in MDS Stephen D. Nimer, MD Sylvester Comprehensive Cancer Center December 5, 2014 DISCLOSURE I have no relevant financial relationships to disclose.
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf. Rachel Schwope EPIGEN May 24-27, 2016
 Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf Rachel Schwope EPIGEN May 24-27, 2016 What does H3K4 methylation do? Plant of interest: Vitis vinifera Culturally important
Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf Rachel Schwope EPIGEN May 24-27, 2016 What does H3K4 methylation do? Plant of interest: Vitis vinifera Culturally important
An epigenetic approach to understanding (and predicting?) environmental effects on gene expression
 www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
Myelodysplastic Syndromes: Hematopathology. Analysis of SHIP1 as a potential biomarker of Disease Progression
 Myelodysplastic Syndromes: Hematopathology. Analysis of SHIP1 as a potential biomarker of Disease Progression Carlos E. Bueso-Ramos, M.D., Ph.D Department of Hematopathology The University of Texas M.
Myelodysplastic Syndromes: Hematopathology. Analysis of SHIP1 as a potential biomarker of Disease Progression Carlos E. Bueso-Ramos, M.D., Ph.D Department of Hematopathology The University of Texas M.
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Accessing and Using ENCODE Data Dr. Peggy J. Farnham
 1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
Results. Abstract. Introduc4on. Conclusions. Methods. Funding
 . expression that plays a role in many cellular processes affecting a variety of traits. In this study DNA methylation was assessed in neuronal tissue from three pigs (frontal lobe) and one great tit (whole
. expression that plays a role in many cellular processes affecting a variety of traits. In this study DNA methylation was assessed in neuronal tissue from three pigs (frontal lobe) and one great tit (whole
Lecture 8. Eukaryotic gene regulation: post translational modifications of histones
 Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
Epigenetics and Cancer
 Epigenetics and Cancer Ya-Hui Chi March 24, 2016 1 Institute of Biotechnology and Pharmaceutical Research, NHRI, Zhunan, Taiwan Question 1: What causes cancer? Douglas Hanahan and Robert A. Weinberg 2
Epigenetics and Cancer Ya-Hui Chi March 24, 2016 1 Institute of Biotechnology and Pharmaceutical Research, NHRI, Zhunan, Taiwan Question 1: What causes cancer? Douglas Hanahan and Robert A. Weinberg 2
Histone modifications in kidney disease
 Histone modifications in kidney disease Masaomi Nangaku Division of Nephrology and Endocrinology The University of Tokyo Graduate School of Medicine Japan Mimura, Tanaka, & Nangaku. Semin Nephrol 2013
Histone modifications in kidney disease Masaomi Nangaku Division of Nephrology and Endocrinology The University of Tokyo Graduate School of Medicine Japan Mimura, Tanaka, & Nangaku. Semin Nephrol 2013
The Epigenome Tools 2: ChIP-Seq and Data Analysis
 The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
Histones modifications and variants
 Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Genomic Methods in Cancer Epigenetic Dysregulation
 Genomic Methods in Cancer Epigenetic Dysregulation Clara, Lyon 2018 Jacek Majewski, Associate Professor Department of Human Genetics, McGill University Montreal, Canada A few words about my lab Genomics
Genomic Methods in Cancer Epigenetic Dysregulation Clara, Lyon 2018 Jacek Majewski, Associate Professor Department of Human Genetics, McGill University Montreal, Canada A few words about my lab Genomics
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
ChIP-seq analysis. J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand
 ChIP-seq analysis J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand Tuesday : quick introduction to ChIP-seq and peak-calling (Presentation + Practical session)
ChIP-seq analysis J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand Tuesday : quick introduction to ChIP-seq and peak-calling (Presentation + Practical session)
Sirt1 Hmg20b Gm (0.17) 24 (17.3) 877 (857)
 3 (0.17) 24 (17.3) Sirt1 Hmg20 Gm4763 877 (857) c d Suppl. Figure 1. Screen validation for top candidate antagonists of Dot1L (a) Numer of genes with one (gray), two (cyan) or three (red) shrna scored
3 (0.17) 24 (17.3) Sirt1 Hmg20 Gm4763 877 (857) c d Suppl. Figure 1. Screen validation for top candidate antagonists of Dot1L (a) Numer of genes with one (gray), two (cyan) or three (red) shrna scored
CTCF-Mediated Functional Chromatin Interactome in Pluripotent Cells
 SUPPLEMENTARY INFORMATION CTCF-Mediated Functional Chromatin Interactome in Pluripotent Cells Lusy Handoko 1,*, Han Xu 1,*, Guoliang Li 1,*, Chew Yee Ngan 1, Elaine Chew 1, Marie Schnapp 1, Charlie Wah
SUPPLEMENTARY INFORMATION CTCF-Mediated Functional Chromatin Interactome in Pluripotent Cells Lusy Handoko 1,*, Han Xu 1,*, Guoliang Li 1,*, Chew Yee Ngan 1, Elaine Chew 1, Marie Schnapp 1, Charlie Wah
September 20, Submitted electronically to: Cc: To Whom It May Concern:
 History Study (NOT-HL-12-147), p. 1 September 20, 2012 Re: Request for Information (RFI): Building a National Resource to Study Myelodysplastic Syndromes (MDS) The MDS Cohort Natural History Study (NOT-HL-12-147).
History Study (NOT-HL-12-147), p. 1 September 20, 2012 Re: Request for Information (RFI): Building a National Resource to Study Myelodysplastic Syndromes (MDS) The MDS Cohort Natural History Study (NOT-HL-12-147).
STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells
 CAMDA 2009 October 5, 2009 STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells Guohua Wang 1, Yadong Wang 1, Denan Zhang 1, Mingxiang Teng 1,2, Lang Li 2, and Yunlong Liu 2 Harbin
CAMDA 2009 October 5, 2009 STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells Guohua Wang 1, Yadong Wang 1, Denan Zhang 1, Mingxiang Teng 1,2, Lang Li 2, and Yunlong Liu 2 Harbin
The epigenetic landscape of T cell subsets in SLE identifies known and potential novel drivers of the autoimmune response
 Abstract # 319030 Poster # F.9 The epigenetic landscape of T cell subsets in SLE identifies known and potential novel drivers of the autoimmune response Jozsef Karman, Brian Johnston, Sofija Miljovska,
Abstract # 319030 Poster # F.9 The epigenetic landscape of T cell subsets in SLE identifies known and potential novel drivers of the autoimmune response Jozsef Karman, Brian Johnston, Sofija Miljovska,
EPIGENOMICS PROFILING SERVICES
 EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
Large conserved domains of low DNA methylation maintained by Dnmt3a
 Supplementary information Large conserved domains of low DNA methylation maintained by Dnmt3a Mira Jeong# 1, Deqiang Sun # 2, Min Luo# 1, Yun Huang 3, Grant A. Challen %1, Benjamin Rodriguez 2, Xiaotian
Supplementary information Large conserved domains of low DNA methylation maintained by Dnmt3a Mira Jeong# 1, Deqiang Sun # 2, Min Luo# 1, Yun Huang 3, Grant A. Challen %1, Benjamin Rodriguez 2, Xiaotian
Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD,
 Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD, Crawford, GE, Ren, B (2007) Distinct and predictive chromatin
Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD, Crawford, GE, Ren, B (2007) Distinct and predictive chromatin
Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder Labs August 30, 2012
 Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
Disclosure Slide. Research Support: Onconova Therapeutics, Celgene
 Oral Rigosertib Combined with Azacitidine in Patients with Acute Myeloid Leukemia (AML) and Myelodysplastic Syndromes (MDS): Effects in Treatment Naïve and Relapsed- Refractory Patients Shyamala C. Navada,
Oral Rigosertib Combined with Azacitidine in Patients with Acute Myeloid Leukemia (AML) and Myelodysplastic Syndromes (MDS): Effects in Treatment Naïve and Relapsed- Refractory Patients Shyamala C. Navada,
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region
 Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
Breast cancer. Risk factors you cannot change include: Treatment Plan Selection. Inferring Transcriptional Module from Breast Cancer Profile Data
 Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
EHA 2017 Abstracts: 4 Abstracts ( 1 Oral Presentation, 2 EPosters, 1 Publication)
 EHA 2017 Abstracts: 4 Abstracts ( 1 Oral Presentation, 2 EPosters, 1 Publication) ORAL RIGOSERTIB COMBINED WITH AZACITIDINE IN PATIENTS WITH ACUTE MYELOID LEUKEMIA (AML) AND MYELODYSPLASTIC SYNDROMES (MDS):
EHA 2017 Abstracts: 4 Abstracts ( 1 Oral Presentation, 2 EPosters, 1 Publication) ORAL RIGOSERTIB COMBINED WITH AZACITIDINE IN PATIENTS WITH ACUTE MYELOID LEUKEMIA (AML) AND MYELODYSPLASTIC SYNDROMES (MDS):
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development
 Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
FOLLICULAR LYMPHOMA- ILLUMINA METHYLATION. Jude Fitzgibbon
 FOLLICULAR LYMPHOMA- ILLUMINA METHYLATION Jude Fitzgibbon j.fitzgibbon@qmul.ac.uk Molecular Predictors of Clinical outcome Responders Alive Non-responders Dead TUMOUR GENETICS Targeted therapy, Prognostic
FOLLICULAR LYMPHOMA- ILLUMINA METHYLATION Jude Fitzgibbon j.fitzgibbon@qmul.ac.uk Molecular Predictors of Clinical outcome Responders Alive Non-responders Dead TUMOUR GENETICS Targeted therapy, Prognostic
Research Article Identifying Liver Cancer-Related Enhancer SNPs by Integrating GWAS and Histone Modification ChIP-seq Data
 BioMed Volume 2016, Article ID 2395341, 6 pages http://dx.doi.org/10.1155/2016/2395341 Research Article Identifying Liver Cancer-Related Enhancer SNPs by Integrating GWAS and Histone Modification ChIP-seq
BioMed Volume 2016, Article ID 2395341, 6 pages http://dx.doi.org/10.1155/2016/2395341 Research Article Identifying Liver Cancer-Related Enhancer SNPs by Integrating GWAS and Histone Modification ChIP-seq
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10860 Supplementary Discussion It remains unclear why H3K9 demethylation appeared to be more sensitive to suppression than at least some other histone methylation marks as a result of
doi:10.1038/nature10860 Supplementary Discussion It remains unclear why H3K9 demethylation appeared to be more sensitive to suppression than at least some other histone methylation marks as a result of
Peak-calling for ChIP-seq and ATAC-seq
 Peak-calling for ChIP-seq and ATAC-seq Shamith Samarajiwa CRUK Autumn School in Bioinformatics 2017 University of Cambridge Overview Peak-calling: identify enriched (signal) regions in ChIP-seq or ATAC-seq
Peak-calling for ChIP-seq and ATAC-seq Shamith Samarajiwa CRUK Autumn School in Bioinformatics 2017 University of Cambridge Overview Peak-calling: identify enriched (signal) regions in ChIP-seq or ATAC-seq
Bivalent-chromatin and epigenetics alterations in Glioma
 Bivalent-chromatin and epigenetics alterations in Glioma Philippe ARNAUD GReD CNRS 6293 - Clermont Universités- INSERM U1103 Clermont-Ferrand, FRANCE Forum de la Recherche en Cancérologie Auvergne-Rhône
Bivalent-chromatin and epigenetics alterations in Glioma Philippe ARNAUD GReD CNRS 6293 - Clermont Universités- INSERM U1103 Clermont-Ferrand, FRANCE Forum de la Recherche en Cancérologie Auvergne-Rhône
Supplemental Figure 1. Genes showing ectopic H3K9 dimethylation in this study are DNA hypermethylated in Lister et al. study.
 mc mc mc mc SUP mc mc Supplemental Figure. Genes showing ectopic HK9 dimethylation in this study are DNA hypermethylated in Lister et al. study. Representative views of genes that gain HK9m marks in their
mc mc mc mc SUP mc mc Supplemental Figure. Genes showing ectopic HK9 dimethylation in this study are DNA hypermethylated in Lister et al. study. Representative views of genes that gain HK9m marks in their
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
ACK1 Tyrosine Kinases: A Critical Regulator of Prostate Cancer
 ACK1 Tyrosine Kinases: A Critical Regulator of Prostate Cancer Nupam Mahajan Moffitt Cancer Center Learners Objectives How Androgen Receptor (AR) signaling is accomplished in absence of androgen What are
ACK1 Tyrosine Kinases: A Critical Regulator of Prostate Cancer Nupam Mahajan Moffitt Cancer Center Learners Objectives How Androgen Receptor (AR) signaling is accomplished in absence of androgen What are
TMEFF2 is an epigenetic modulator that promotes androgen independent growth in castrationresistant. prostate cancer cells. Joshua Corbin.
 TMEFF2 is an epigenetic modulator that promotes androgen independent growth in castrationresistant prostate cancer cells by Joshua Corbin August, 2014 Director of Thesis: Maria Ruiz-Echevarria, Ph.D. Major
TMEFF2 is an epigenetic modulator that promotes androgen independent growth in castrationresistant prostate cancer cells by Joshua Corbin August, 2014 Director of Thesis: Maria Ruiz-Echevarria, Ph.D. Major
Transduction of lentivirus to human primary CD4+ T cells
 Transduction of lentivirus to human primary CD4 + T cells Human primary CD4 T cells were stimulated with anti-cd3/cd28 antibodies (10 µl/2 5 10^6 cells of Dynabeads CD3/CD28 T cell expander, Invitrogen)
Transduction of lentivirus to human primary CD4 + T cells Human primary CD4 T cells were stimulated with anti-cd3/cd28 antibodies (10 µl/2 5 10^6 cells of Dynabeads CD3/CD28 T cell expander, Invitrogen)
of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed.
 Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
RUNX1 and FPD/AML Translational Research. The Leukemia and Lymphoma Society / Babich Family Foundation Partnership. September 2016
 www.lls.org www.runx1.com RUNX1 and FPD/AML Translational Research The Leukemia and Lymphoma Society / Babich Family Foundation Partnership September 2016 Prepared by L. Greenberger, PhD Chief Scientific
www.lls.org www.runx1.com RUNX1 and FPD/AML Translational Research The Leukemia and Lymphoma Society / Babich Family Foundation Partnership September 2016 Prepared by L. Greenberger, PhD Chief Scientific
Expert Intelligence for Better Decisions Epigenetics:
 Expert Intelligence for Better Decisions Epigenetics: Emerging Targets, Available Technologies, Expert Interviews, and an Epigenetic Community Perspective Using This Document Insight Pharma Reports are
Expert Intelligence for Better Decisions Epigenetics: Emerging Targets, Available Technologies, Expert Interviews, and an Epigenetic Community Perspective Using This Document Insight Pharma Reports are
Chromatin-Based Regulation of Gene Expression
 Chromatin-Based Regulation of Gene Expression.George J. Quellhorst, Jr., PhD.Associate Director, R&D.Biological Content Development Topics to be Discussed Importance of Chromatin-Based Regulation Mechanism
Chromatin-Based Regulation of Gene Expression.George J. Quellhorst, Jr., PhD.Associate Director, R&D.Biological Content Development Topics to be Discussed Importance of Chromatin-Based Regulation Mechanism
Transcriptional and Epigenetic Mechanisms of Addiction
 Transcriptional and Epigenetic Mechanisms of Addiction Eric J. Nestler Mount Sinai School of Medicine New York, NY Dr. Ray Fuller There is every reason to be optimistic that in the future we will find
Transcriptional and Epigenetic Mechanisms of Addiction Eric J. Nestler Mount Sinai School of Medicine New York, NY Dr. Ray Fuller There is every reason to be optimistic that in the future we will find
Dominic J Smiraglia, PhD Department of Cancer Genetics. DNA methylation in prostate cancer
 Dominic J Smiraglia, PhD Department of Cancer Genetics DNA methylation in prostate cancer Overarching theme Epigenetic regulation allows the genome to be responsive to the environment Sets the tone for
Dominic J Smiraglia, PhD Department of Cancer Genetics DNA methylation in prostate cancer Overarching theme Epigenetic regulation allows the genome to be responsive to the environment Sets the tone for
MDS 101. What is bone marrow? Myelodysplastic Syndrome: Let s build a definition. Dysplastic? Syndrome? 5/22/2014. What does bone marrow do?
 101 May 17, 2014 Myelodysplastic Syndrome: Let s build a definition Myelo bone marrow Gail J. Roboz, M.D. Director, Leukemia Program Associate Professor of Medicine What is bone marrow? What does bone
101 May 17, 2014 Myelodysplastic Syndrome: Let s build a definition Myelo bone marrow Gail J. Roboz, M.D. Director, Leukemia Program Associate Professor of Medicine What is bone marrow? What does bone
Computational aspects of ChIP-seq. John Marioni Research Group Leader European Bioinformatics Institute European Molecular Biology Laboratory
 Computational aspects of ChIP-seq John Marioni Research Group Leader European Bioinformatics Institute European Molecular Biology Laboratory ChIP-seq Using highthroughput sequencing to investigate DNA
Computational aspects of ChIP-seq John Marioni Research Group Leader European Bioinformatics Institute European Molecular Biology Laboratory ChIP-seq Using highthroughput sequencing to investigate DNA
Molecular Functions of MLL PHD3 Binding to Its Ligands Cyp33 and H3K4Me3
 Loyola University Chicago Loyola ecommons Dissertations Theses and Dissertations 2013 Molecular Functions of MLL PHD3 Binding to Its Ligands Cyp33 and H3K4Me3 Gayathree Raman Loyola University Chicago
Loyola University Chicago Loyola ecommons Dissertations Theses and Dissertations 2013 Molecular Functions of MLL PHD3 Binding to Its Ligands Cyp33 and H3K4Me3 Gayathree Raman Loyola University Chicago
IPSS Modified 7/27/2011. WHO-Based Prognostic Scoring System (WPSS)
 Advances in MDS Treatment: What s on the Horizon? New Prognostic Models and Therapies Jason Gotlib, MD, MS Assistant Professor of Medicine (Hematology) Stanford Cancer Center AA&MDSIF July 3, 011 WHO-Based
Advances in MDS Treatment: What s on the Horizon? New Prognostic Models and Therapies Jason Gotlib, MD, MS Assistant Professor of Medicine (Hematology) Stanford Cancer Center AA&MDSIF July 3, 011 WHO-Based
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
CBL and EZH2 as new molecular markers in MPN
 CBL and EZH2 as new molecular markers in MPN Andy Chase University of Southampton and Wessex Regional Genetics Laboratory Salisbury, UK Munich 2011 * Myeloproliferative neoplasms MDS/MPN Myelodysplastic
CBL and EZH2 as new molecular markers in MPN Andy Chase University of Southampton and Wessex Regional Genetics Laboratory Salisbury, UK Munich 2011 * Myeloproliferative neoplasms MDS/MPN Myelodysplastic
Nature Structural & Molecular Biology: doi: /nsmb.2419
 Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Combination of Oral Rigosertib and Injectable Azacitidine in Patients with Myelodysplastic Syndromes (MDS)
 Combination of Oral Rigosertib and Injectable Azacitidine in Patients with Myelodysplastic Syndromes (MDS) Shyamala C. Navada, MD 1, Guillermo Garcia-Manero, MD 2, Katherine Hearn, RN 2, Rosalie Odchimar-Reissig,
Combination of Oral Rigosertib and Injectable Azacitidine in Patients with Myelodysplastic Syndromes (MDS) Shyamala C. Navada, MD 1, Guillermo Garcia-Manero, MD 2, Katherine Hearn, RN 2, Rosalie Odchimar-Reissig,
Genome-Wide Localization of Protein-DNA Binding and Histone Modification by a Bayesian Change-Point Method with ChIP-seq Data
 Genome-Wide Localization of Protein-DNA Binding and Histone Modification by a Bayesian Change-Point Method with ChIP-seq Data Haipeng Xing, Yifan Mo, Will Liao, Michael Q. Zhang Clayton Davis and Geoffrey
Genome-Wide Localization of Protein-DNA Binding and Histone Modification by a Bayesian Change-Point Method with ChIP-seq Data Haipeng Xing, Yifan Mo, Will Liao, Michael Q. Zhang Clayton Davis and Geoffrey
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression. Chromatin Array of nuc
 Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Stem Cell Epigenetics
 Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
MIR retrotransposon sequences provide insulators to the human genome
 Supplementary Information: MIR retrotransposon sequences provide insulators to the human genome Jianrong Wang, Cristina Vicente-García, Davide Seruggia, Eduardo Moltó, Ana Fernandez- Miñán, Ana Neto, Elbert
Supplementary Information: MIR retrotransposon sequences provide insulators to the human genome Jianrong Wang, Cristina Vicente-García, Davide Seruggia, Eduardo Moltó, Ana Fernandez- Miñán, Ana Neto, Elbert
Table S1. Total and mapped reads produced for each ChIP-seq sample
 Tale S1. Total and mapped reads produced for each ChIP-seq sample Sample Total Reads Mapped Reads Col- H3K27me3 rep1 125662 1334323 (85.76%) Col- H3K27me3 rep2 9176437 7986731 (87.4%) atmi1a//c H3K27m3
Tale S1. Total and mapped reads produced for each ChIP-seq sample Sample Total Reads Mapped Reads Col- H3K27me3 rep1 125662 1334323 (85.76%) Col- H3K27me3 rep2 9176437 7986731 (87.4%) atmi1a//c H3K27m3
Statistical Genetics. Matthew Stephens. Statistics Retreat, October 26th 2012
 Statistical Genetics Statistics Retreat, October 26th 2012 Two stories The two most influential statistical ideas in analysis of genetic association studies. 1 Sequence, sequence, everywhere. 1 With apologies
Statistical Genetics Statistics Retreat, October 26th 2012 Two stories The two most influential statistical ideas in analysis of genetic association studies. 1 Sequence, sequence, everywhere. 1 With apologies
Epigenetic programming in chronic lymphocytic leukemia
 Epigenetic programming in chronic lymphocytic leukemia Christopher Oakes 10 th Canadian CLL Research Meeting September 18-19 th, 2014 Epigenetics and DNA methylation programming in normal and tumor cells:
Epigenetic programming in chronic lymphocytic leukemia Christopher Oakes 10 th Canadian CLL Research Meeting September 18-19 th, 2014 Epigenetics and DNA methylation programming in normal and tumor cells:
Predictive Blood DNA Markers for Breast Cancer Xiang Zhang, Ph.D.
 Predictive Blood DNA Markers for Breast Cancer Xiang Zhang, Ph.D. Department of Environmental Health University of Cincinnati Background Breast cancer (BCa) The second most common cancer among women in
Predictive Blood DNA Markers for Breast Cancer Xiang Zhang, Ph.D. Department of Environmental Health University of Cincinnati Background Breast cancer (BCa) The second most common cancer among women in
Molecular Hematopathology Leukemias I. January 14, 2005
 Molecular Hematopathology Leukemias I January 14, 2005 Chronic Myelogenous Leukemia Diagnosis requires presence of Philadelphia chromosome t(9;22)(q34;q11) translocation BCR-ABL is the result BCR on chr
Molecular Hematopathology Leukemias I January 14, 2005 Chronic Myelogenous Leukemia Diagnosis requires presence of Philadelphia chromosome t(9;22)(q34;q11) translocation BCR-ABL is the result BCR on chr
Session 6: Integration of epigenetic data. Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016
 Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
ChipSeq. Technique and science. The genome wide dynamics of the binding of ldb1 complexes during erythroid differentiation
 Center for Biomics ChipSeq Technique and science The genome wide dynamics of the binding of ldb1 complexes during erythroid differentiation Wilfred van IJcken Sequencing Seminar Illumina September 8 Scheme
Center for Biomics ChipSeq Technique and science The genome wide dynamics of the binding of ldb1 complexes during erythroid differentiation Wilfred van IJcken Sequencing Seminar Illumina September 8 Scheme
BIO360 Fall 2013 Quiz 1
 BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
Epigenetic Analysis of KSHV Latent and Lytic Genomes
 Epigenetic Analysis of KSHV Latent and Lytic Genomes Zsolt Toth 1, Dennis T. Maglinte 2, Sun Hwa Lee 1, Hye-Ra Lee 1, Lai-Yee Wong 1, Kevin F. Brulois 1, Stacy Lee 1, Jonathan D. Buckley 2,3, Peter W.
Epigenetic Analysis of KSHV Latent and Lytic Genomes Zsolt Toth 1, Dennis T. Maglinte 2, Sun Hwa Lee 1, Hye-Ra Lee 1, Lai-Yee Wong 1, Kevin F. Brulois 1, Stacy Lee 1, Jonathan D. Buckley 2,3, Peter W.
Integrated analysis of sequencing data
 Integrated analysis of sequencing data How to combine *-seq data M. Defrance, M. Thomas-Chollier, C. Herrmann, D. Puthier, J. van Helden *ChIP-seq, RNA-seq, MeDIP-seq, Transcription factor binding ChIP-seq
Integrated analysis of sequencing data How to combine *-seq data M. Defrance, M. Thomas-Chollier, C. Herrmann, D. Puthier, J. van Helden *ChIP-seq, RNA-seq, MeDIP-seq, Transcription factor binding ChIP-seq
Distinct Epigenetic Signatures Delineate Transcriptional Programs during Virus-Specific CD8 + T Cell Differentiation
 Resource Distinct Epigenetic Signatures Delineate Transcriptional Programs during Virus-Specific CD8 + T Cell Differentiation Brendan E. Russ, 1 Moshe Olshanksy, 2 Heather S. Smallwood, 3 Jasmine Li, 1
Resource Distinct Epigenetic Signatures Delineate Transcriptional Programs during Virus-Specific CD8 + T Cell Differentiation Brendan E. Russ, 1 Moshe Olshanksy, 2 Heather S. Smallwood, 3 Jasmine Li, 1
TITLE: Insight Into Skin Tumorigenesis Highlighting the Function of Epigenetic Regulators in SCC Formation
 AD Award Number: W81XWH-12-1-0463 TITLE: Insight Into Skin Tumorigenesis Highlighting the Function of Epigenetic Regulators in SCC Formation PRINCIPAL INVESTIGATOR: Jisheng Zhang CONTRACTING ORGANIZATION:
AD Award Number: W81XWH-12-1-0463 TITLE: Insight Into Skin Tumorigenesis Highlighting the Function of Epigenetic Regulators in SCC Formation PRINCIPAL INVESTIGATOR: Jisheng Zhang CONTRACTING ORGANIZATION:
The corrected Figure S1J is shown below. The text changes are as follows, with additions in bold and deletions in bracketed italics:
 Correction H3K4me3 readth Is Linked to Cell Identity and Transcriptional Consistency érénice A. enayoun, Elizabeth A. Pollina, Duygu Ucar, Salah Mahmoudi, Kalpana Karra, Edith D. Wong, Keerthana Devarajan,
Correction H3K4me3 readth Is Linked to Cell Identity and Transcriptional Consistency érénice A. enayoun, Elizabeth A. Pollina, Duygu Ucar, Salah Mahmoudi, Kalpana Karra, Edith D. Wong, Keerthana Devarajan,
CONTRACTING ORGANIZATION: University of Texas MD Anderson Cancer Center Houston TX 77030
 AD Award Number: W1XWH-1-1-01 TITLE: Innate Immunity Dysregulation in Myelodysplastic Syndromes PRINCIPAL INVESTIGATOR: Yue Wei CONTRACTING ORGANIZATION: University of Texas MD Anderson Cancer Center Houston
AD Award Number: W1XWH-1-1-01 TITLE: Innate Immunity Dysregulation in Myelodysplastic Syndromes PRINCIPAL INVESTIGATOR: Yue Wei CONTRACTING ORGANIZATION: University of Texas MD Anderson Cancer Center Houston
Supplementary Figure 1. Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Nature Immunology: doi: /ni.
 Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
ChIPSeq. Technique and science. The genome wide dynamics of the binding of ldb1 complexes during erythroid differentiation
 Center for Biomics ChIPSeq Technique and science The genome wide dynamics of the binding of ldb1 complexes during erythroid differentiation Wilfred van IJcken EU Sequencing Seminar Illumina June 16 Genomics
Center for Biomics ChIPSeq Technique and science The genome wide dynamics of the binding of ldb1 complexes during erythroid differentiation Wilfred van IJcken EU Sequencing Seminar Illumina June 16 Genomics
3/31/2014. New Directions in Aplastic Anemia Treatment: What s on the Horizon? Objectives
 New Directions in Aplastic Anemia Treatment: What s on the Horizon? AA & MDS International Foundation Living with, MDS, or PNH Patient and Family Conferences in 2014 April 5, 2014 Objectives To provide
New Directions in Aplastic Anemia Treatment: What s on the Horizon? AA & MDS International Foundation Living with, MDS, or PNH Patient and Family Conferences in 2014 April 5, 2014 Objectives To provide
An annotated list of bivalent chromatin regions in human ES cells: a new tool for cancer epigenetic research
 An annotated list of bivalent chromatin regions in human ES cells: a new tool for cancer epigenetic research Franck Court, Philippe Arnaud To cite this version: Franck Court, Philippe Arnaud. An annotated
An annotated list of bivalent chromatin regions in human ES cells: a new tool for cancer epigenetic research Franck Court, Philippe Arnaud To cite this version: Franck Court, Philippe Arnaud. An annotated
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
ChromHMM Tutorial. Jason Ernst Assistant Professor University of California, Los Angeles
 ChromHMM Tutorial Jason Ernst Assistant Professor University of California, Los Angeles Talk Outline Chromatin states analysis and ChromHMM Accessing chromatin state annotations for ENCODE2 and Roadmap
ChromHMM Tutorial Jason Ernst Assistant Professor University of California, Los Angeles Talk Outline Chromatin states analysis and ChromHMM Accessing chromatin state annotations for ENCODE2 and Roadmap
New drugs in Acute Leukemia. Cristina Papayannidis, MD, PhD University of Bologna
 New drugs in Acute Leukemia Cristina Papayannidis, MD, PhD University of Bologna Challenges to targeted therapy in AML Multiple subtypes based upon mutations/cytogenetic aberrations No known uniform genomic
New drugs in Acute Leukemia Cristina Papayannidis, MD, PhD University of Bologna Challenges to targeted therapy in AML Multiple subtypes based upon mutations/cytogenetic aberrations No known uniform genomic
Genome-wide Association Studies (GWAS) Pasieka, Science Photo Library
 Lecture 5 Genome-wide Association Studies (GWAS) Pasieka, Science Photo Library Chi-squared test to evaluate whether the odds ratio is different from 1. Corrected for multiple testing Source: wikipedia.org
Lecture 5 Genome-wide Association Studies (GWAS) Pasieka, Science Photo Library Chi-squared test to evaluate whether the odds ratio is different from 1. Corrected for multiple testing Source: wikipedia.org
Figure S1, Beyer et al.
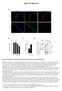 Figure S1, eyer et al. Pax7 Myogenin si sitrl Hoechst T = 72h 14 1.8.6.4.2 12 1 8 6 4 2 24h 48h 96h diff. sitrl siset1 212 72h diff. b1 td r t Se km MyH Vinculin Myogenin β-ctin Vinculin MW b1 ka td r
Figure S1, eyer et al. Pax7 Myogenin si sitrl Hoechst T = 72h 14 1.8.6.4.2 12 1 8 6 4 2 24h 48h 96h diff. sitrl siset1 212 72h diff. b1 td r t Se km MyH Vinculin Myogenin β-ctin Vinculin MW b1 ka td r
Dr Kavita Raj Consultant Haematologist Guys and St Thomas Hospital
 Dr Kavita Raj Consultant Haematologist Guys and St Thomas Hospital IPSS scoring system Blood counts Bone marrow blast percentage Cytogenetics Age as a modulator of median survival IPSS Group Median Survival
Dr Kavita Raj Consultant Haematologist Guys and St Thomas Hospital IPSS scoring system Blood counts Bone marrow blast percentage Cytogenetics Age as a modulator of median survival IPSS Group Median Survival
Molecular Markers. Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC
 Molecular Markers Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC Overview Testing methods Rationale for molecular testing
Molecular Markers Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC Overview Testing methods Rationale for molecular testing
TET2 mutations were predictive of inferior prognosis in the presence of ASXL1 mutations in patients with chronic myelomonocytic leukemia
 Short Report TET2 mutations were predictive of inferior prognosis in the presence of ASXL1 mutations in patients with chronic myelomonocytic leukemia Yajuan Cui 1,2 *, Hongyan Tong 3 *, Xin Du 4 *, Bing
Short Report TET2 mutations were predictive of inferior prognosis in the presence of ASXL1 mutations in patients with chronic myelomonocytic leukemia Yajuan Cui 1,2 *, Hongyan Tong 3 *, Xin Du 4 *, Bing
Department of Leukemia, The University of Texas M.D. Anderson Cancer Center, Houston, Texas; 2 Sunesis Pharmaceuticals, Inc, South San Francisco
 Phase I/II Study of Vosaroxin and Decitabine in Newly Diagnosed Older Patients with Acute Myeloid Leukemia (AML) and High Risk Myelodysplastic Syndrome (MDS) Naval Daver 1, Hagop Kantarjian 1, Guillermo
Phase I/II Study of Vosaroxin and Decitabine in Newly Diagnosed Older Patients with Acute Myeloid Leukemia (AML) and High Risk Myelodysplastic Syndrome (MDS) Naval Daver 1, Hagop Kantarjian 1, Guillermo
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
Measuring DNA Methylation with the MinION. Winston Timp Department of Biomedical Engineering Johns Hopkins University 12/1/16
 Measuring DNA Methylation with the MinION Winston Timp Department of Biomedical Engineering Johns Hopkins University 12/1/16 Epigenetics: Modern Modern Definition of epigenetics involves heritable changes
Measuring DNA Methylation with the MinION Winston Timp Department of Biomedical Engineering Johns Hopkins University 12/1/16 Epigenetics: Modern Modern Definition of epigenetics involves heritable changes
Epigenetic modifications play important roles in
 LIVER BIOLOGY/PATHOBIOLOGY Neonatal Activation of the Nuclear Receptor CAR Results in Epigenetic Memory and Permanent Change of Drug Metabolism in Mouse Liver Wei-Dong Chen, 1,2 * Xianghui Fu, 1 * Bingning
LIVER BIOLOGY/PATHOBIOLOGY Neonatal Activation of the Nuclear Receptor CAR Results in Epigenetic Memory and Permanent Change of Drug Metabolism in Mouse Liver Wei-Dong Chen, 1,2 * Xianghui Fu, 1 * Bingning
Role of Hypoxia-Inducible Factors in Epigenetic Regulation via Histone Demethylases
 HYPOXIA AND CONSEQUENCES Role of Hypoxia-Inducible Factors in Epigenetic Regulation via Histone Demethylases Jun Yang, a Ioanna Ledaki, a Helen Turley, b Kevin C. Gatter, b Juan-Carlos Martinez Montero,
HYPOXIA AND CONSEQUENCES Role of Hypoxia-Inducible Factors in Epigenetic Regulation via Histone Demethylases Jun Yang, a Ioanna Ledaki, a Helen Turley, b Kevin C. Gatter, b Juan-Carlos Martinez Montero,
Transcript-indexed ATAC-seq for immune profiling
 Transcript-indexed ATAC-seq for immune profiling Technical Journal Club 22 nd of May 2018 Christina Müller Nature Methods, Vol.10 No.12, 2013 Nature Biotechnology, Vol.32 No.7, 2014 Nature Medicine, Vol.24,
Transcript-indexed ATAC-seq for immune profiling Technical Journal Club 22 nd of May 2018 Christina Müller Nature Methods, Vol.10 No.12, 2013 Nature Biotechnology, Vol.32 No.7, 2014 Nature Medicine, Vol.24,
Pharmacologic inhibition of histone demethylation as a therapy for pediatric brainstem glioma
 Supplementary information for: Pharmacologic inhibition of histone demethylation as a therapy for pediatric brainstem glioma Rintaro Hashizume 1, Noemi Andor 2, Yuichiro Ihara 2, Robin Lerner 2, Haiyun
Supplementary information for: Pharmacologic inhibition of histone demethylation as a therapy for pediatric brainstem glioma Rintaro Hashizume 1, Noemi Andor 2, Yuichiro Ihara 2, Robin Lerner 2, Haiyun
