SUPPLEMENTAL INFORMATION. Evolving Spindlin1 Small Molecule Inhibitors Using Protein Microarrays
|
|
|
- Jerome Bailey
- 6 years ago
- Views:
Transcription
1 SUPPLEMETAL IFRMATI Evolving Spindlin1 Small Molecule Inhibitors Using Protein Microarrays arkhyun Bae 1, Monica Viviano 2, Xiaonan Su 3, Jie Lu 4, Donghang Cheng 1, Cari Sagum 1, Sabrina Castellano 2,6, Xue Bai 3,5, Claire Johnson 1, Mahmoud Ibrahim Khalil 1,7, Kaifu Chen 4, aitao Li 3,5*, Gianluca Sbardella 2* and Mark T. Bedford 1* 1 Department of Epigenetics and Molecular Carcinogenesis, The University of Texas MD Anderson Cancer Center, Smithville, TX 78957, USA. 2 Dipartimento di Farmacia, Epigenetic Med Chem Lab, Università degli Studi di Salerno, Via Giovanni Paolo II 132, I Fisciano (SA), Italy. 3 ME Key Laboratory of Protein Sciences, Beijing Advanced Innovation Center for Structural Biology, Department of Basic Medical Sciences, School of Medicine, Tsinghua University, Beijing , China. 4 Center for regenerative Medicine, Department of Cardiovascular Sciences, ouston Methodist Research Institute, ouston, TX 77030, USA 5 Tsinghua-Peking Center for Life Sciences, Beijing, , P.R. China 6 Dipartimento di Medicina e Chirurgia, Università degli Studi di Salerno, Via Salvador Allende, I Baronissi (SA), Italy. 7 Molecular Biology Unit, Department of Zoology, Faculty of Science, Alexandria University, Egypt. These authors contributed equally to this work. * Correspondence should be addressed to - M.T.B. (mtbedford@mdanderson.org) G.S..L. (gsbardella@unisa.it) (lht@tsinghua.edu.cn) ature Chemical Biology: doi: /nchembio.2377
2 SUPPLEMETAL RESULTS Supplemental Figures Supplementary Figure 1 The layout and composition of the protein domain microarray used in Fig. 1b and Supplementary Fig. 3. ature Chemical Biology: doi: /nchembio.2377
3 Supplementary Figure 2 (a) Structure of UC1215, a previously reported antagonist of L3MBTL3 Kme-binding activity. The compound contains a central moiety symmetrically decorated with two nitrogen-containing groups that are mimetic of substituted lysine residues. (b) With the aim to find molecular probes for the various proteins containing Kme- and Rme-reader domains, we designed a series of compounds featuring a central aromatic core substituted, symmetrically and asymmetrically, with basic nitrogen-containing privileged scaffolds, potentially able to give additional binding interactions. ature Chemical Biology: doi: /nchembio.2377
4 Supplementary Figure 3 UC1215 analogs bind different subsets of royal domain family members. A collection of 98 GST fusion proteins was arrayed in duplicate on a nitrocellulose slide. This library is composed of almost complete sets of human Tudor and Chromo domains that are cloned as GST fusions. About 50 tagged compounds were labeled with Cy3 and used to probe these arrays. An example of 17 of the EML series is shown, as well as UC1215. Some of these interactions displayed novel shifts in binding specificity, two of which (EML405 & EML417) are highlighted in Fig. 1b. ature Chemical Biology: doi: /nchembio.2377
5 a b Supplementary Figure 4 (a) Calorimetric studies of EML analogs and UC1215 binding to 53BP1, PF20, L3MBTL1 and L3MBTL3, respectively. (b) Calorimetric studies showing that the SPI1/TCF4 is blocked by EML405 and EML631. ITC titrations were confirmed by three independent experiments. ature Chemical Biology: doi: /nchembio.2377
6 -PEG(10)-Biotin -PEG(10)-Biotin 10 (biotin-eml631) 11 (biotin-eml632) -PEG(10)-Biotin -PEG(10)-Biotin 12 (biotin-eml633) 13 (biotin-eml405*) Supplementary Figure 5 Alternative tagging for EML405 and EML Because the introduction of the pyrrolidinecontaining additional arm in EML the position of the biotinylated PEG linker needed to be altered for these three analogs. A new variant of EML405 with this altered attachment site was also synthesized to confirm that it does not interfere with the SPI1 Tudor interaction see Fig. 4b. ature Chemical Biology: doi: /nchembio.2377
7 Supplementary Figure 6 (a) To reaffirm that EML632 can interact with SPI1 we performed a pulldown assay with tagged and bead immobilized EML405, and efficiently competed off GST-SPI1 with increasing concentrations of untagged EML632. (b) Chemiprecipitation of GFP- SPI family members expressed in EK 293T cells, using biotinylated EML631. ature Chemical Biology: doi: /nchembio.2377
8 ature Chemical Biology: doi: /nchembio.2377
9 Supplementary Figure 7 RA-Seq identification of SPI1 regulated transcripts and inhibition of this co-activator activity by EML631. (a) Representative RA-Seq gene expression profiles for SPI1, IL1B and BST2 gene loci obtained using RA isolated from control (sira control), SPI1 knockdown (sira SPI1) and EML631 (10 µm) treatment T778 cells. (b) The expression levels of the indicated transcripts, determined by RT-qPCR analysis of RA purified from T778 cells, after treatment with or without EML 405 (20 µm) and EML631 (10 µm) for 4 days. (c) The effect of EML405 (10 µm) and EML631 (10 µm) was monitored analyzing the gene expression changes in up-regulated SPI1 target genes. Gene expression was normalized to GAPD. The mean value for the control groups was arbitrarily set as 1. All data represent the average of three independent experiments (biological replicates), each subjected to three independent RT-qPCR (total of n=9). S.D. is denoted by error bar. *P<0.05, **P<0.01, ***P<0.001 (Student s t-test). ature Chemical Biology: doi: /nchembio.2377
10 Supplementary Figure 8 (a) The time dependent effect of EML631 (10 µm) was analyzed by RT-qPCR (b) The dose dependent effect of EML631 (0 ~ 20 µm) was analyzed by RT-qPCR. Gene expression was normalized to GAPD. The mean value for the control groups was arbitrarily set as 1. All data represent the average of three independent experiments (biological replicates), each subjected to three independent RT-qPCR (total of n=9). S.D. is denoted by error bar. *P<0.05, **P<0.01, ***P<0.001 (Student s t-test). ature Chemical Biology: doi: /nchembio.2377
11 Supplementary Figure 9 riginal uncropped blot images from the data presented in Figure 4a. ature Chemical Biology: doi: /nchembio.2377
12 Supplementary Figure 10 riginal uncropped blot images from the data presented in Figure 4b. ature Chemical Biology: doi: /nchembio.2377
13 Supplementary Figure 11 riginal uncropped blot images from the data presented in Figure 4c. ature Chemical Biology: doi: /nchembio.2377
14 Supplementary Figure 12 riginal uncropped blot images from the data presented in Figure 5b. ature Chemical Biology: doi: /nchembio.2377
15 Supplementary Tables Supplementary Table 1. Summary of compounds screened against the array and their binding data. Binding interactions with the protein domains on the array are depicted as heatmap, on the basis of densitometry. For the sake of clarity, proteins that gave no interaction with any of the compounds tested are not shown on the table. Binding affinity (densitometry) compounds # Structure 53BP1(1-2)* 53BP1(1-2) PF20 Tudor PF20-2 Tudor PF20L1 Tudor Pombe1 Tudor* SPF30sp Spindlin 1 Tudor* Engaged proteins SPI1 Tudor MRG15 Chromo L3MBTL 1(1-3) MBT L3MBTL 1(1-3) MBT* CD1 Chromo CD2 Chromo L3MBTL 1(1-2)* MBT L3MBTL 3 MBT UC EML EML EML EML ature Chemical Biology: doi: /nchembio.2377
16 EML S EML S EML EML EML EML EML ature Chemical Biology: doi: /nchembio.2377
17 EML EML EML EML EML ature Chemical Biology: doi: /nchembio.2377
18 EML EML EML EML EML EML EML EML ature Chemical Biology: doi: /nchembio.2377
19 EML EML EML EML EML EML EML ature Chemical Biology: doi: /nchembio.2377
20 EML EML EML EML EML EML EML EML631 -PEG(10)-Biotin ature Chemical Biology: doi: /nchembio.2377
21 EML632 -PEG(10)-Biotin -PEG(10)-Biotin EML EML EML EML EML664 (EML405* alternative linker) -PEG(10)-Biotin ature Chemical Biology: doi: /nchembio.2377
22 Supplementa Table 2. Data collection and refinement statistics of SPI1-EML405 complex structure Spindlin1-EML405 Data collection Space group P21 Cell dimensions a, b, c (Å) 43.1, 123.9, 49.8 α, β, γ ( ) 90, 92.3, 90 Resolution (Å) ( )* Rsym or Rmerge 13.5 (83.4) I / σi 13.5 (2.2) Completeness (%) 99.7 (99.7) Redundancy 3.5 (3.5) Refinement Resolution (Å) o. reflections (1841) Rwork / Rfree 18.6/24.5 o. atoms Protein 3141 Ligand/ion 86 Water 113 B-factors (Ask for input) Protein 40.7 Ligand/ion 40.8 Water 37.8 R.m.s. deviations Bond lengths (Å) Bond angles ( ) * umber of xtals for each structure should be noted in footnote. *ighest-resolution shell is shown in parentheses. ature Chemical Biology: doi: /nchembio.2377
23 Supplementa Table 3. Data collection and refinement statistics of SPI1-EML631 complex structure Spindlin1-EML631 Data collection Space group P21 Cell dimensions a, b, c (Å) 42.8, 123.8, 49.7 α, β, γ ( ) 90, 92.2, 90 Resolution (Å) ( )* Rsym or Rmerge 9.2 (31.8) I / σi 22.1 (3.6) Completeness (%) 98.9 (89.9) Redundancy 5.4 (5.3) Refinement Resolution (Å) o. reflections (1057) Rwork / Rfree 20.9/25.0 o. atoms Protein 3166 Ligand/ion 100 Water 84 B-factors Protein 57.5 Ligand/ion 48.6 Water 51.7 R.m.s. deviations Bond lengths (Å) Bond angles ( ) * umber of xtals for each structure should be noted in footnote. *ighest-resolution shell is shown in parentheses. ature Chemical Biology: doi: /nchembio.2377
24 Supplementa Table 4. Binding affinity for the association of EML405, EML631, EML632, EML633 and UC1215 to five methyl-lysine-binding proteins. ITC titrations were confirmed by three independent experiments. ature Chemical Biology: doi: /nchembio.2377
The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism
 Supplementary Information The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism Address correspondence to Yong Li (yongli@xmu.edu.cn, Tel: 86-592-218151) GW464 CDCA Supplementary
Supplementary Information The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism Address correspondence to Yong Li (yongli@xmu.edu.cn, Tel: 86-592-218151) GW464 CDCA Supplementary
KDM2A. Reactions. containing. Reactions
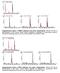 Supplementary Figure 1 KDM2A catalyses only lysine demethylation.. MALDI-TOF MS of demethylation of the shown variant histone peptides as catalysed by recombinant KDM2A. Reactions containing enzyme are
Supplementary Figure 1 KDM2A catalyses only lysine demethylation.. MALDI-TOF MS of demethylation of the shown variant histone peptides as catalysed by recombinant KDM2A. Reactions containing enzyme are
This PDF file includes: Supplementary Figures 1 to 6 Supplementary Tables 1 to 2 Supplementary Methods Supplementary References
 Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1. Differential expression of mirnas from the pri-mir-17-92a locus.
 Supplementary Figure 1 Differential expression of mirnas from the pri-mir-17-92a locus. (a) The mir-17-92a expression unit in the third intron of the host mir-17hg transcript. (b,c) Impact of knockdown
Supplementary Figure 1 Differential expression of mirnas from the pri-mir-17-92a locus. (a) The mir-17-92a expression unit in the third intron of the host mir-17hg transcript. (b,c) Impact of knockdown
Supporting Information
 Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
Figure S1. HP1α localizes to centromeres in mitosis and interacts with INCENP. (A&B) HeLa
 SUPPLEMENTARY FIGURES Figure S1. HP1α localizes to centromeres in mitosis and interacts with INCENP. (A&B) HeLa tet-on cells that stably express HP1α-CFP, HP1β-CFP, or HP1γ-CFP were monitored with livecell
SUPPLEMENTARY FIGURES Figure S1. HP1α localizes to centromeres in mitosis and interacts with INCENP. (A&B) HeLa tet-on cells that stably express HP1α-CFP, HP1β-CFP, or HP1γ-CFP were monitored with livecell
SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES
 SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors:
SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors:
Supplementary Information Janssen et al.
 Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
Alpha thalassemia mental retardation X-linked. Acquired alpha-thalassemia myelodysplastic syndrome
 Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
Biochemistry - I. Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I
 Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/9/e1600292/dc1 Supplementary Materials for Native phasing of x-ray free-electron laser data for a G protein coupled receptor Alexander Batyuk, Lorenzo Galli,
advances.sciencemag.org/cgi/content/full/2/9/e1600292/dc1 Supplementary Materials for Native phasing of x-ray free-electron laser data for a G protein coupled receptor Alexander Batyuk, Lorenzo Galli,
SUPPLEMENTAL MATERIAL. UNC119 is required for G protein trafficking in sensory neurons
 1 SUPPLEMENTAL MATERIAL UNC119 is required for G protein trafficking in sensory neurons Houbin Zhang, Ryan N. Constantine, Sergey Vorobiev, Yang Chen, Jayaraman Seetharaman, Yuanpeng Janet Huang, Rong
1 SUPPLEMENTAL MATERIAL UNC119 is required for G protein trafficking in sensory neurons Houbin Zhang, Ryan N. Constantine, Sergey Vorobiev, Yang Chen, Jayaraman Seetharaman, Yuanpeng Janet Huang, Rong
FCC2 5CY7 FCC1 5CY8. Actinonin 5CVQ
 Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
Supplementary Figure 1. Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
 Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
T Cell Receptor Optimized Peptide Skewing of the T-Cell Repertoire Can Enhance Antigen Targeting
 T Cell Receptor Optimized Peptide Skewing of the T-Cell Repertoire Can Enhance Antigen Targeting Julia Ekeruche-Makinde 1*, Mathew Clement 1*, David K Cole 1*, Emily S J Edwards 1, Kristin Ladell 1, John
T Cell Receptor Optimized Peptide Skewing of the T-Cell Repertoire Can Enhance Antigen Targeting Julia Ekeruche-Makinde 1*, Mathew Clement 1*, David K Cole 1*, Emily S J Edwards 1, Kristin Ladell 1, John
Supporting Information
 Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Nature Genetics: doi: /ng Supplementary Figure 1. Immunofluorescence (IF) confirms absence of H3K9me in met-2 set-25 worms.
 Supplementary Figure 1 Immunofluorescence (IF) confirms absence of H3K9me in met-2 set-25 worms. IF images of wild-type (wt) and met-2 set-25 worms showing the loss of H3K9me2/me3 at the indicated developmental
Supplementary Figure 1 Immunofluorescence (IF) confirms absence of H3K9me in met-2 set-25 worms. IF images of wild-type (wt) and met-2 set-25 worms showing the loss of H3K9me2/me3 at the indicated developmental
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes
 Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose.
 Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose. (a) Relative levels of both new acetylation and all acetylation
Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose. (a) Relative levels of both new acetylation and all acetylation
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information.
 Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein
 Supplementary Methods Thermal shift assays Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein unfolding was examined by monitoring the fluorescence of ANS (1-anilinonaphthalene-8-
Supplementary Methods Thermal shift assays Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein unfolding was examined by monitoring the fluorescence of ANS (1-anilinonaphthalene-8-
m 6 A mrna methylation regulates AKT activity to promote the proliferation and tumorigenicity of endometrial cancer
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
Supporting Information. Structure-based Lead Optimization and Biological Evaluation of BAX Direct Activators as Novel Potential Anticancer Agents
 Supporting Information Structure-based Lead Optimization and Biological Evaluation of BAX Direct Activators as Novel Potential Anticancer Agents Mariano Stornaiuolo, 1 * Giuseppe La Regina, 2 Sara Passacantilli,
Supporting Information Structure-based Lead Optimization and Biological Evaluation of BAX Direct Activators as Novel Potential Anticancer Agents Mariano Stornaiuolo, 1 * Giuseppe La Regina, 2 Sara Passacantilli,
Table S1. Primer sequences used for qrt-pcr. CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT ACTB AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT LCOR
 Table S1. Primer sequences used for qrt-pcr. ACTB LCOR KLF6 CTBP1 CDKN1A CDH1 ATF3 PLAU MMP9 TFPI2 CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT CGGCTGCAGGAAAGTTTACA
Table S1. Primer sequences used for qrt-pcr. ACTB LCOR KLF6 CTBP1 CDKN1A CDH1 ATF3 PLAU MMP9 TFPI2 CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT CGGCTGCAGGAAAGTTTACA
Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS
 Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS nucleotide sequences (a, b) or amino acid sequences (c) from
Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS nucleotide sequences (a, b) or amino acid sequences (c) from
SUPPLEMENTARY INFORMATION FOR. (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation. by COX-2
 SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
supplementary information
 DOI: 1.138/ncb1 Control Atg7 / NAC 1 1 1 1 (mm) Control Atg7 / NAC 1 1 1 1 (mm) Lamin B Gstm1 Figure S1 Neither the translocation of into the nucleus nor the induction of antioxidant proteins in autophagydeficient
DOI: 1.138/ncb1 Control Atg7 / NAC 1 1 1 1 (mm) Control Atg7 / NAC 1 1 1 1 (mm) Lamin B Gstm1 Figure S1 Neither the translocation of into the nucleus nor the induction of antioxidant proteins in autophagydeficient
172R 172K TAM-2/172R TAM-2/172K. AZT concentration [nm] AZT concentration [nm] MgCl 2 2.5K 2.5K 5K 2.5K 5K 2.5K K 5K 2.5K 5K 2.5K 50 2.
![172R 172K TAM-2/172R TAM-2/172K. AZT concentration [nm] AZT concentration [nm] MgCl 2 2.5K 2.5K 5K 2.5K 5K 2.5K K 5K 2.5K 5K 2.5K 50 2. 172R 172K TAM-2/172R TAM-2/172K. AZT concentration [nm] AZT concentration [nm] MgCl 2 2.5K 2.5K 5K 2.5K 5K 2.5K K 5K 2.5K 5K 2.5K 50 2.](/thumbs/82/85138523.jpg) 5 5 5 5 A MgCl 2 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] B 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] ATP + ATP - Supplemental Figure 1. Primer extension of HIV-1 RT polymorphisms
5 5 5 5 A MgCl 2 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] B 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] ATP + ATP - Supplemental Figure 1. Primer extension of HIV-1 RT polymorphisms
effects on organ development. a-f, Eye and wing discs with clones of ε j2b10 show no
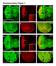 Supplementary Figure 1. Loss of function clones of 14-3-3 or 14-3-3 show no significant effects on organ development. a-f, Eye and wing discs with clones of 14-3-3ε j2b10 show no obvious defects in Elav
Supplementary Figure 1. Loss of function clones of 14-3-3 or 14-3-3 show no significant effects on organ development. a-f, Eye and wing discs with clones of 14-3-3ε j2b10 show no obvious defects in Elav
Supplementary Figure 1. MAT IIα is Acetylated at Lysine 81.
 IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis
 Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
Supplementary Information
 Supplementary Information Using the pimeloyl-coa synthetase adenylation fold to synthesise fatty acid thioesters Menglu Wang 1a ; Lucile Moynié 2a, Peter J. Harrison 1, Van Kelly 1, Andrew Piper 1, James
Supplementary Information Using the pimeloyl-coa synthetase adenylation fold to synthesise fatty acid thioesters Menglu Wang 1a ; Lucile Moynié 2a, Peter J. Harrison 1, Van Kelly 1, Andrew Piper 1, James
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12215 Supplementary Figure 1. The effects of full and dissociated GR agonists in supporting BFU-E self-renewal divisions. BFU-Es were cultured in self-renewal medium with indicated GR
doi:10.1038/nature12215 Supplementary Figure 1. The effects of full and dissociated GR agonists in supporting BFU-E self-renewal divisions. BFU-Es were cultured in self-renewal medium with indicated GR
Breast cancer. Risk factors you cannot change include: Treatment Plan Selection. Inferring Transcriptional Module from Breast Cancer Profile Data
 Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
Guangdong Medical University, Zhanjiang, China; 5 Guangxi Medical University, Nanning, China; 6 Department of Pathology, University of Michigan
 Overexpression of FAM83H-AS1 indicates poor patient survival and knockdown impairs cell proliferation and invasion via MET/EGFR signaling in lung cancer Jie Zhang 1,2, Shumei Feng 3, Wenmei Su 4, Shengbin
Overexpression of FAM83H-AS1 indicates poor patient survival and knockdown impairs cell proliferation and invasion via MET/EGFR signaling in lung cancer Jie Zhang 1,2, Shumei Feng 3, Wenmei Su 4, Shengbin
Table S1. X-ray data collection and refinement statistics
 Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables. File Name: Peer Review File Description:
 File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables File Name: Peer Review File Description: Primer Name Sequence (5'-3') AT ( C) RT-PCR USP21 F 5'-TTCCCATGGCTCCTTCCACATGAT-3'
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables File Name: Peer Review File Description: Primer Name Sequence (5'-3') AT ( C) RT-PCR USP21 F 5'-TTCCCATGGCTCCTTCCACATGAT-3'
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/1/8/e1500296/dc1 Supplementary Materials for Transcriptional regulation of APOBEC3 antiviral immunity through the CBF- /RUNX axis This PDF file includes: Brett
advances.sciencemag.org/cgi/content/full/1/8/e1500296/dc1 Supplementary Materials for Transcriptional regulation of APOBEC3 antiviral immunity through the CBF- /RUNX axis This PDF file includes: Brett
SUPPLEMENTARY INFORMATION
 Supplementary Discussion The HGGG sequence motif forms an oxyanion hole in HLSs. In the case of GID1, the third Gly residue of the sequence motif is replaced with Ser (Ser123 in OsGID1), i.e., the sequence
Supplementary Discussion The HGGG sequence motif forms an oxyanion hole in HLSs. In the case of GID1, the third Gly residue of the sequence motif is replaced with Ser (Ser123 in OsGID1), i.e., the sequence
Infect MCF-7 cells carrying dcas9-vp64 + psm2-p65-hsf1 with SAM library or vector. Introduce AKT reporter
 Infect MCF-7 cells carrying dcas9-vp64 + psm2-p65-hsf1 with SM library or vector Introduce reporter Grow cells in presence of puromycin for 5 days Vector control SM library fewer surviving cells More surviving
Infect MCF-7 cells carrying dcas9-vp64 + psm2-p65-hsf1 with SM library or vector Introduce reporter Grow cells in presence of puromycin for 5 days Vector control SM library fewer surviving cells More surviving
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Electron micrograph of phosphotungstanic acid-stained exosomes derived from murine
 1 SUPPLEMENTARY INFORMATION SUPPLEMENTARY FIGURES Supplementary Figure 1. Physical properties of murine DC-derived exosomes. a, Electron micrograph of phosphotungstanic acid-stained exosomes derived from
1 SUPPLEMENTARY INFORMATION SUPPLEMENTARY FIGURES Supplementary Figure 1. Physical properties of murine DC-derived exosomes. a, Electron micrograph of phosphotungstanic acid-stained exosomes derived from
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/9/439/ra78/dc1 Supplementary Materials for Small heterodimer partner mediates liver X receptor (LXR) dependent suppression of inflammatory signaling by promoting
www.sciencesignaling.org/cgi/content/full/9/439/ra78/dc1 Supplementary Materials for Small heterodimer partner mediates liver X receptor (LXR) dependent suppression of inflammatory signaling by promoting
Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene
 Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene YUELIN ZHANG, WEIHUA FAN, MARK KINKEMA, XIN LI, AND
Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene YUELIN ZHANG, WEIHUA FAN, MARK KINKEMA, XIN LI, AND
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature9814 a A SHARPIN FL B SHARPIN ΔNZF C SHARPIN T38L, F39V b His-SHARPIN FL -1xUb -2xUb -4xUb α-his c Linear 4xUb -SHARPIN FL -SHARPIN TF_LV -SHARPINΔNZF -SHARPIN
SUPPLEMENTARY INFORMATION doi:1.138/nature9814 a A SHARPIN FL B SHARPIN ΔNZF C SHARPIN T38L, F39V b His-SHARPIN FL -1xUb -2xUb -4xUb α-his c Linear 4xUb -SHARPIN FL -SHARPIN TF_LV -SHARPINΔNZF -SHARPIN
ABT-199, a potent and selective BCL-2 inhibitor, achieves antitumor activity while sparing platelets
 Supplementary Information ABT-199, a potent and selective BCL-2 inhibitor, achieves antitumor activity while sparing platelets Andrew J Souers 1, Joel D Leverson 1, Erwin R Boghaert 1, Scott L Ackler 1,
Supplementary Information ABT-199, a potent and selective BCL-2 inhibitor, achieves antitumor activity while sparing platelets Andrew J Souers 1, Joel D Leverson 1, Erwin R Boghaert 1, Scott L Ackler 1,
ncounter TM Analysis System
 ncounter TM Analysis System Molecules That Count TM www.nanostring.com Agenda NanoString Technologies History Introduction to the ncounter Analysis System CodeSet Design and Assay Principals System Performance
ncounter TM Analysis System Molecules That Count TM www.nanostring.com Agenda NanoString Technologies History Introduction to the ncounter Analysis System CodeSet Design and Assay Principals System Performance
Biology Open (2014) 000, 1 10 doi: /bio
 (2014) 000, 1 10 doi:10.1242/bio.201410041 Supplementary Material Michael Brauchle et al. doi: 10.1242/bio.201410041 Fig. S1. Alignment of GFP, sfgfp, egfp, eyfp, mcherry and mruby2. Sequence-based alignment
(2014) 000, 1 10 doi:10.1242/bio.201410041 Supplementary Material Michael Brauchle et al. doi: 10.1242/bio.201410041 Fig. S1. Alignment of GFP, sfgfp, egfp, eyfp, mcherry and mruby2. Sequence-based alignment
Single-cell RNA-Seq profiling of human pre-implantation embryos and embryonic stem cells
 Single-cell RNA-Seq profiling of human pre-implantation embryos and embryonic stem cells Liying Yan,2,5, Mingyu Yang,5, Hongshan Guo, Lu Yang, Jun Wu, Rong Li,2, Ping Liu, Ying Lian, Xiaoying Zheng, Jie
Single-cell RNA-Seq profiling of human pre-implantation embryos and embryonic stem cells Liying Yan,2,5, Mingyu Yang,5, Hongshan Guo, Lu Yang, Jun Wu, Rong Li,2, Ping Liu, Ying Lian, Xiaoying Zheng, Jie
ISG15 sirna # Ctrl sirna+ifn+wt. Virus titer (Pfu/ml) hours post infection. d USP18 sirna #2 IFN
 a ISG15 sirna Ctrl #1 #2 ISG15 conjugates IB: ISG15 Virus titer (Pfu/ml) b ISG15 sirna #2 10 7 Ctrl sirna+ifn+wt Ctrl sirna+ifn+67 10 6 ISG15 sirna+ifn+wt ISG15 sirna+ifn+67 10 5 10 4 10 3 Free ISG15 10
a ISG15 sirna Ctrl #1 #2 ISG15 conjugates IB: ISG15 Virus titer (Pfu/ml) b ISG15 sirna #2 10 7 Ctrl sirna+ifn+wt Ctrl sirna+ifn+67 10 6 ISG15 sirna+ifn+wt ISG15 sirna+ifn+67 10 5 10 4 10 3 Free ISG15 10
Dramatic increase in SHP2 binding activity of Helicobacter. pylori Western CagA by EPIYA-C duplication: its
 Supplementary Information Dramatic increase in SHP2 binding activity of Helicobacter pylori Western CagA by EPIYA-C duplication: its implications in gastric carcinogenesis Lisa Nagase, Takeru Hayashi,
Supplementary Information Dramatic increase in SHP2 binding activity of Helicobacter pylori Western CagA by EPIYA-C duplication: its implications in gastric carcinogenesis Lisa Nagase, Takeru Hayashi,
Nature Immunology: doi: /ni.3631
 Supplementary Figure 1 SPT analyses of Zap70 at the T cell plasma membrane. (a) Total internal reflection fluorescent (TIRF) excitation at 64-68 degrees limits single molecule detection to 100-150 nm above
Supplementary Figure 1 SPT analyses of Zap70 at the T cell plasma membrane. (a) Total internal reflection fluorescent (TIRF) excitation at 64-68 degrees limits single molecule detection to 100-150 nm above
Supplementary Material to Oxidative properties of FeO2+: electronic structure and solvation effects.
 Supplementary Material to Oxidative properties of FeO2+: electronic structure and solvation effects. Manuel J. Louwerse and Evert Jan Baerends Theoretical Chemistry, Vrije Universiteit Amsterdam, De Boelelaan
Supplementary Material to Oxidative properties of FeO2+: electronic structure and solvation effects. Manuel J. Louwerse and Evert Jan Baerends Theoretical Chemistry, Vrije Universiteit Amsterdam, De Boelelaan
Supplementary Figure 1. Structure models of the c4 variant proteins based on the structures of maltose binding protein (MBP) in red and TEM-1
 Supplementary Figure 1. Structure models of the c4 variant proteins based on the structures of maltose binding protein (MBP) in red and TEM-1 β-lactamase (BLA) in blue. (a) c4 model and the close-up view
Supplementary Figure 1. Structure models of the c4 variant proteins based on the structures of maltose binding protein (MBP) in red and TEM-1 β-lactamase (BLA) in blue. (a) c4 model and the close-up view
Molecular Testing in Lung Cancer
 Molecular Testing in Lung Cancer Pimpin Incharoen, M.D. Assistant Professor, Thoracic Pathology Department of Pathology, Ramathibodi Hospital Genetic alterations in lung cancer Source: Khono et al, Trans
Molecular Testing in Lung Cancer Pimpin Incharoen, M.D. Assistant Professor, Thoracic Pathology Department of Pathology, Ramathibodi Hospital Genetic alterations in lung cancer Source: Khono et al, Trans
= (3) = (3) = (4) V = (4) Å 3 Z =2. Data collection. Refinement
 organic compounds Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 = 95.181 (3) = 112.830 (3) = 106.243 (4) V = 948.7 (4) Å 3 Z =2 Mo K radiation = 0.07 mm 1 T = 298 (2) K 0.41
organic compounds Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 = 95.181 (3) = 112.830 (3) = 106.243 (4) V = 948.7 (4) Å 3 Z =2 Mo K radiation = 0.07 mm 1 T = 298 (2) K 0.41
EGFR shrna A: CCGGCGCAAGTGTAAGAAGTGCGAACTCGAGTTCGCACTTCTTACACTTGCG TTTTTG. EGFR shrna B: CCGGAGAATGTGGAATACCTAAGGCTCGAGCCTTAGGTATTCCACATTCTCTT TTTG
 Supplementary Methods Sequence of oligonucleotides used for shrna targeting EGFR EGFR shrna were obtained from the Harvard RNAi consortium. The following oligonucleotides (forward primer) were used to
Supplementary Methods Sequence of oligonucleotides used for shrna targeting EGFR EGFR shrna were obtained from the Harvard RNAi consortium. The following oligonucleotides (forward primer) were used to
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk
 Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
MARTINI Coarse-Grained Model of Triton TX-100 in Pure DPPC. Monolayer and Bilayer Interfaces. Supporting Information
 MARTINI Coarse-Grained Model of Triton TX-100 in Pure DPPC Monolayer and Bilayer Interfaces. Antonio Pizzirusso a, Antonio De Nicola* a, Giuseppe Milano a a Dipartimento di Chimica e Biologia, Università
MARTINI Coarse-Grained Model of Triton TX-100 in Pure DPPC Monolayer and Bilayer Interfaces. Antonio Pizzirusso a, Antonio De Nicola* a, Giuseppe Milano a a Dipartimento di Chimica e Biologia, Università
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Supplementary materials
 Supplementary materials Chemical library from ChemBridge 50,240 structurally diverse small molecule compounds dissolved in DMSO Hits Controls: No virus added μ Primary screening at 20 g/ml of compounds
Supplementary materials Chemical library from ChemBridge 50,240 structurally diverse small molecule compounds dissolved in DMSO Hits Controls: No virus added μ Primary screening at 20 g/ml of compounds
STRUCTURE OF A UBIQUITIN-LOADED HECT LIGASE REVEALS THE MOLECULAR BASIS FOR CATALYTIC PRIMING SUPPLEMENTARY INFORMATION
 STRUCTURE OF A UBIQUITINLOADED ECT LIGASE REVEALS TE MOLECULAR BASIS FOR CATALYTIC PRIMING Elena Maspero 1, Eleonora Valentini 1, Sara Mari 1, Valentina Cecatiello 2, Paolo Soffientini 1, Sebastiano Pasqualato
STRUCTURE OF A UBIQUITINLOADED ECT LIGASE REVEALS TE MOLECULAR BASIS FOR CATALYTIC PRIMING Elena Maspero 1, Eleonora Valentini 1, Sara Mari 1, Valentina Cecatiello 2, Paolo Soffientini 1, Sebastiano Pasqualato
Histone peptide microarray screen of chromo and Tudor domains defines new histone lysine methylation interactions
 DOI 10.1186/s13072-017-0117-5 Epigenetics & Chromatin RESEARCH Open Access Histone peptide microarray screen of chromo and Tudor domains defines new histone lysine methylation interactions Erin K. Shanle
DOI 10.1186/s13072-017-0117-5 Epigenetics & Chromatin RESEARCH Open Access Histone peptide microarray screen of chromo and Tudor domains defines new histone lysine methylation interactions Erin K. Shanle
Biochemistry 15 Doctor /7/2012
 Heme The Heme is a chemical structure that diffracts by light to give a red color. This chemical structure is introduced to more than one protein. So, a protein containing this heme will appear red in
Heme The Heme is a chemical structure that diffracts by light to give a red color. This chemical structure is introduced to more than one protein. So, a protein containing this heme will appear red in
b = (17) Å c = (18) Å = (4) = (4) = (4) V = (3) Å 3 Data collection Refinement
 organic compounds Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 (Z)-N-tert-Butyl-2-(4-methoxyanilino)- N 0 -(4-methoxyphenyl)-2-phenylacetimidamide Sue A. Roberts, a * Biswajit
organic compounds Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 (Z)-N-tert-Butyl-2-(4-methoxyanilino)- N 0 -(4-methoxyphenyl)-2-phenylacetimidamide Sue A. Roberts, a * Biswajit
Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in
 Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in Supplementary Fig. 2 Substitution Sequence Position variant Sequence original APNCYGNIPL original APNCYGNIPL
Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in Supplementary Fig. 2 Substitution Sequence Position variant Sequence original APNCYGNIPL original APNCYGNIPL
Dynamic Partitioning of a GPI-Anchored Protein in Glycosphingolipid-Rich Microdomains Imaged by Single-Quantum Dot Tracking
 Additional data for Dynamic Partitioning of a GPI-Anchored Protein in Glycosphingolipid-Rich Microdomains Imaged by Single-Quantum Dot Tracking Fabien Pinaud 1,3, Xavier Michalet 1,3, Gopal Iyer 1, Emmanuel
Additional data for Dynamic Partitioning of a GPI-Anchored Protein in Glycosphingolipid-Rich Microdomains Imaged by Single-Quantum Dot Tracking Fabien Pinaud 1,3, Xavier Michalet 1,3, Gopal Iyer 1, Emmanuel
Supplementary Information
 Coarse-Grained Molecular Dynamics Simulations of Photoswitchable Assembly and Disassembly Xiaoyan Zheng, Dong Wang,* Zhigang Shuai,* MOE Key Laboratory of Organic Optoelectronics and Molecular Engineering,
Coarse-Grained Molecular Dynamics Simulations of Photoswitchable Assembly and Disassembly Xiaoyan Zheng, Dong Wang,* Zhigang Shuai,* MOE Key Laboratory of Organic Optoelectronics and Molecular Engineering,
Supplementary Information. Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase
 Journal: Nature Medicine Supplementary Information Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase 1,2 Peng Wang PhD, 1,2 Juan-Carlos Alvarez-Perez
Journal: Nature Medicine Supplementary Information Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase 1,2 Peng Wang PhD, 1,2 Juan-Carlos Alvarez-Perez
Experimental. Crystal data. C 20 H 23 N 4 O 6 PS M r = Monoclinic, P2 1 =n a = (2) Å b = (4) Å c = (2) Å = 117.
 organic compounds Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 Diethyl {(4-methoxyphenyl)[5-(4-nitro- phenyl)-1,3,4-thiadiazol-2-ylamino]- methyl}phosphonate Li-He Yin, Rong
organic compounds Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 Diethyl {(4-methoxyphenyl)[5-(4-nitro- phenyl)-1,3,4-thiadiazol-2-ylamino]- methyl}phosphonate Li-He Yin, Rong
organic compounds (E)-Ethyl N 0 -(3-hydroxybenzylidene)- hydrazinecarboxylate dihydrate
 organic compounds Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 (E)-Ethyl N 0 -(3-hydroxybenzylidene)- hydrazinecarboxylate dihydrate Xian-Chao Hu, a,b * Jie Zhang, a Da-Yong
organic compounds Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 (E)-Ethyl N 0 -(3-hydroxybenzylidene)- hydrazinecarboxylate dihydrate Xian-Chao Hu, a,b * Jie Zhang, a Da-Yong
% of Cells A B C. Proliferation Index. T cell count (10 6 ) Division Index. % of Max CFSE. %Ki67+ cells. Supplementary Figure 1.
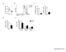 A B C T cell count (1 6 ) 3. 2. 1.. * % of Max CFSE ormoxia ypoxia o Stim. Proliferation Index 2.5 2. 1.5 1. * Division Index 2. 1.5 1..5. D E %Ki67+ cells 1 8 6 4 2 % of Cells 8 ormoxia 6 ypoxia 4 2 *
A B C T cell count (1 6 ) 3. 2. 1.. * % of Max CFSE ormoxia ypoxia o Stim. Proliferation Index 2.5 2. 1.5 1. * Division Index 2. 1.5 1..5. D E %Ki67+ cells 1 8 6 4 2 % of Cells 8 ormoxia 6 ypoxia 4 2 *
How T cells recognize antigen: The T Cell Receptor (TCR) Identifying the TCR: Why was it so hard to do? Monoclonal antibody approach
 How T cells recognize antigen: The T Cell Receptor (TCR) Identifying the TCR: Why was it so hard to do By the early 1980s, much about T cell function was known, but the receptor genes had not been identified
How T cells recognize antigen: The T Cell Receptor (TCR) Identifying the TCR: Why was it so hard to do By the early 1980s, much about T cell function was known, but the receptor genes had not been identified
A second type of TCR TCR: An αβ heterodimer
 How s recognize antigen: The T Cell Receptor (TCR) Identifying the TCR: Why was it so hard to do By the early 1980s, much about function was known, but the receptor genes had not been identified Recall
How s recognize antigen: The T Cell Receptor (TCR) Identifying the TCR: Why was it so hard to do By the early 1980s, much about function was known, but the receptor genes had not been identified Recall
Final Report. Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD. Wendy Fong ( 方美珍 ) CNIC; Beijing, China
 Final Report Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD Wendy Fong ( 方美珍 ) CNIC; Beijing, China Influenza s Viral Life Cycle 1. HA on virus binds to sialic
Final Report Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD Wendy Fong ( 方美珍 ) CNIC; Beijing, China Influenza s Viral Life Cycle 1. HA on virus binds to sialic
Supporting Information
 Supporting Information Bauer et al. 10.1073/pnas.1610421113 SI Materials and Methods Immunofluorescence experiments were performed as described (8). Briefly, NUGC-3 cells were treated with 25 μm PK11007
Supporting Information Bauer et al. 10.1073/pnas.1610421113 SI Materials and Methods Immunofluorescence experiments were performed as described (8). Briefly, NUGC-3 cells were treated with 25 μm PK11007
SUPPLEMENTAL FIGURE LEGENDS
 SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure S1: Endogenous interaction between RNF2 and H2AX: Whole cell extracts from 293T were subjected to immunoprecipitation with anti-rnf2 or anti-γ-h2ax antibodies
SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure S1: Endogenous interaction between RNF2 and H2AX: Whole cell extracts from 293T were subjected to immunoprecipitation with anti-rnf2 or anti-γ-h2ax antibodies
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
a b G75 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server.
 a b G75 2 2 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server. a. Overlay of top 10 models generated by I-TASSER illustrates the potential effect of 7 amino acid insertion
a b G75 2 2 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server. a. Overlay of top 10 models generated by I-TASSER illustrates the potential effect of 7 amino acid insertion
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
EXTENDED DATA FORMATTING GUIDE
 EXTENDED DATA FORMATTING GUIDE Nature paper but are included online within the full-text HTML and at the end of the online PDF. Nature separate panel (for example, see Extended Data Figure 2, panel d)
EXTENDED DATA FORMATTING GUIDE Nature paper but are included online within the full-text HTML and at the end of the online PDF. Nature separate panel (for example, see Extended Data Figure 2, panel d)
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11095 Supplementary Table 1. Summary of the binding between Angptls and various Igdomain containing receptors as determined by flow cytometry analysis. The results were summarized from
doi:10.1038/nature11095 Supplementary Table 1. Summary of the binding between Angptls and various Igdomain containing receptors as determined by flow cytometry analysis. The results were summarized from
sirna count per 50 kb small RNAs matching the direct strand Repeat length (bp) per 50 kb repeats in the chromosome
 Qi et al. 26-3-2564C Qi et al., Figure S1 sirna count per 5 kb small RNAs matching the direct strand sirna count per 5 kb small RNAs matching the complementary strand Repeat length (bp) per 5 kb repeats
Qi et al. 26-3-2564C Qi et al., Figure S1 sirna count per 5 kb small RNAs matching the direct strand sirna count per 5 kb small RNAs matching the complementary strand Repeat length (bp) per 5 kb repeats
Experimental. Crystal data. C 22 H 26 N 2 O 3 M r = Triclinic, P1 a = (3) Å b = (4) Å c = (4) Å = (6) = 100.
 Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 15-Methoxy-14,15-dihydroandranginine Dian-Lei Wang, a Xiang-Hai Cai, b Peng Huang a * and He-Ping Huang a * a School of Pharmacy,
Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 15-Methoxy-14,15-dihydroandranginine Dian-Lei Wang, a Xiang-Hai Cai, b Peng Huang a * and He-Ping Huang a * a School of Pharmacy,
The time has come: SINGLE CELL western blotting. Parrinello Natalia TJC
 The time has come: SINGLE CELL western blotting Parrinello Natalia TJC 08.07.2014 To fully understand diverse and often rare behaviors in complex cell populations, researchers need analytical tools that:
The time has come: SINGLE CELL western blotting Parrinello Natalia TJC 08.07.2014 To fully understand diverse and often rare behaviors in complex cell populations, researchers need analytical tools that:
Research on a Transmit-Receive Method of Ultrasonic Array for Planar Defects
 7 th Asia-Pacific Workshop on Structural Health Monitoring November 12-15, 2018 Hong Kong SAR, P.R. China Research on a Transmit-Receive Method of Ultrasonic Array for Planar Defects Zhenggan Zhou 1,2,3
7 th Asia-Pacific Workshop on Structural Health Monitoring November 12-15, 2018 Hong Kong SAR, P.R. China Research on a Transmit-Receive Method of Ultrasonic Array for Planar Defects Zhenggan Zhou 1,2,3
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/471/eaah5085/dc1 Supplementary Materials for Phosphorylation of the exocyst protein Exo84 by TBK1 promotes insulin-stimulated GLUT4 trafficking Maeran Uhm,
www.sciencesignaling.org/cgi/content/full/10/471/eaah5085/dc1 Supplementary Materials for Phosphorylation of the exocyst protein Exo84 by TBK1 promotes insulin-stimulated GLUT4 trafficking Maeran Uhm,
Ephrin receptor A2 is an epithelial cell receptor for Epstein Barr virus entry
 SUPPLEMENTARY INFORMATION Letters https://doi.org/10.1038/s41564-017-0080-8 In the format provided by the authors and unedited. Ephrin receptor A2 is an epithelial cell receptor for Epstein Barr virus
SUPPLEMENTARY INFORMATION Letters https://doi.org/10.1038/s41564-017-0080-8 In the format provided by the authors and unedited. Ephrin receptor A2 is an epithelial cell receptor for Epstein Barr virus
SUPPLEMENTARY INFORMATION
 Supplementary Table 1. Cell sphingolipids and S1P bound to endogenous TRAF2. Sphingolipid Cell pmol/mg TRAF2 immunoprecipitate pmol/mg Sphingomyelin 4200 ± 250 Not detected Monohexosylceramide 311 ± 18
Supplementary Table 1. Cell sphingolipids and S1P bound to endogenous TRAF2. Sphingolipid Cell pmol/mg TRAF2 immunoprecipitate pmol/mg Sphingomyelin 4200 ± 250 Not detected Monohexosylceramide 311 ± 18
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Supplementary Figure S1: Fibroblast-induced elongation of cancer cells requires direct contact with living fibroblasts. A. Representative images of HT29-GFP cultured in the presence
SUPPLEMENTARY FIGURES Supplementary Figure S1: Fibroblast-induced elongation of cancer cells requires direct contact with living fibroblasts. A. Representative images of HT29-GFP cultured in the presence
SUPPLEMENTARY INFORMATION
 sirna pool: Control Tetherin -HA-GFP HA-Tetherin -Tubulin Supplementary Figure S1. Knockdown of HA-tagged tetherin expression by tetherin specific sirnas. HeLa cells were cotransfected with plasmids expressing
sirna pool: Control Tetherin -HA-GFP HA-Tetherin -Tubulin Supplementary Figure S1. Knockdown of HA-tagged tetherin expression by tetherin specific sirnas. HeLa cells were cotransfected with plasmids expressing
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
