Nature Biotechnology: doi: /nbt Supplementary Figure 1. gene target strain characteristics and respiration culture optimization.
|
|
|
- Erin Jones
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1 gene target strain characteristics and respiration culture optimization. (a) Proteins encoded by the individual genes knocked out of the 174 yeast strains investigated in this study, shown in the context of biological pathways. APS, adenosine-5 -phosphosulfate; CII CV, oxidative phosphorylation complexes II V; ER, endoplasmic reticulum; EMC, ER membrane complex; ERMES, ER-mitochondria encounter structure; ETF, electron transfer flavoprotein complex; MAM, mitochondria-associated membrane; MECA, mitochondria-er-cortex anchor; MICOS, mitochondrial contact site and cristae organizing system; MIM, mitochondrial inner membrane; MOM, mitochondrial outer membrane; mtdna, mitochondrial DNA; mtribosome, mitochondrial ribosome; NAD, nicotinamide adenine dinucleotide; PDH, pyruvate dehydrogenase; TCA, tricarboxylic acid cycle; vclamp, vacuole and mitochondria patch. The pie charts show the total number of characterized and uncharacterized genes profiled (top); the total number of profiled genes that have human homologs (upper middle); of these genes with human homologs, the number of profiled genes that are also associated with disease (lower middle); and of the uncharacterized genes profiled, the number of genes that have human homologs (bottom). (b) Density of yeast cultures in the respiratory growth condition (mean, n = 3) plotted in strain rank order (left) or against fermentation culture density (mean, n = 3) (right). (c) Optical density at 600 nm (OD 600 ) of yeast cultures (media with 3% [w/v] glycerol and 0.1% [w/v] glucose) indicating time points at which yeast were harvested during fermentation (F1 F3) or respiration (R4 R8). Time point R6 (25 h) was selected for the respiration culture condition of the larger study. (d) Wholeproteome plot of protein abundances at time points R5 and R8. (e) Pairwise whole proteome plot comparisons (as in d) across all eight time points (lower left) and linear regression analysis of each comparison (r 2, Pearson correlation coefficients) (upper right).
2 Supplementary Figure 2 Mass spectrometry analysis metrics and quality assessment. (a) Proteins, lipids, and metabolites quantified per gene strain (mean s.d., n = 3). (b) MS experiments conducted per day (top) and phenotypes (molecules) quantified per day (bottom) for proteomics, lipidomics, and metabolomics. (c) Overview of the yeast protein extraction method optimized for this study compared to previous work. (d) Violin plots depicting the range of fold changes in molecule abundance (log 2 [ gene/wt]) across all molecule classes and metabolic states. (e) Density plots of the distribution of coefficients of variation (CVs) (%) for each molecule measured in biological triplicate across all mutants and growth conditions. (f) Venn diagrams depicting the average overlap of molecules quantified within individual gene strains across fermentation and respiration growth conditions. (g) Average profile overlap between different gene strains.
3 Supplementary Figure 3 Features of protein-lipid-metabolite perturbation profiles. (a) Heat maps depicting the number of molecules significantly perturbed within each gene strain (P < 0.05; two-sided Student s t-test). (b) Hierarchical clusters of gene strains and significantly perturbed molecules (relative abundances compared to WT quantified by MS; P < 0.05; two-sided Student s t-test). The center column annotates select clusters with significant functional (GO term) enrichments (P < 0.05; Fisher s exact test followed by Benjamini-Hochberg FDR correction for multiple hypothesis testing). Pie charts indicate proteins in clusters encoded by characterized (gray) or uncharacterized (red) genes.
4 Supplementary Figure 4 Expanded view of two protein clusters from the respiration Y3K dataset heat map (respiration profiles). Heat map indicates relative abundance of proteins in gene strains compared to WT as quantified by MS. See Supplementary Fig. 3 for the full heat map.
5 Supplementary Figure 5 Subsets of the gene-specific phenotypes identified in this study. Relative abundances of individual molecules (mean log 2 [ gene/wt], n = 3) (x-axes) versus statistical significance ( log 10 [p-value]; twosided Student s t-test) (y-axes) as quantified by MS. The plots shown represent a subset of molecules identified as gene-specific phenotypes through an unbiased survey of the Y3K dataset (see Fig. 2a). The array here is limited to the most robust outliers (based on both statistical significance and fold-change, see Supplementary Note 2 and Online Methods) the top 20 upregulated proteins, the top 20 downregulated proteins, the top 10 metabolites, and the top 4 or 5 lipids excluding knocked out proteins (e.g. Fmp52p in the fmp52 strain) and excluding a given gene strain after it appeared twice on the rank list. Biological hypotheses surrounding genephenotype relationship were generated for the starred plots (see Supplementary Fig. 6).
6 Supplementary Figure 6 Examples of hypotheses that can be generated from a subset of the gene-specific phenotypes identified in this study. Subset of gene-specific phenotypes identified in the Y3K dataset. Volcano plots indicate relative molecule abundances (mean log 2 [ gene/wt], n = 3) (x-axes) versus statistical significance ( log 10 [p-value]; two-sided Student s t-test) (y-axes) as quantified by MS. Hypotheses were developed to describe each gene-phenotype relationship reported here.
7 Supplementary Figure 7 Hfd1p supports production of 4-HB for CoQ biosynthesis. (a) Relative lipid abundances (mean, n = 3) versus statistical significance ( log 10 [p-value]; two-sided Student s t-test) as quantified by MS. (b) Relative abundances of 4-HBz (mean, n = 3) versus statistical significance ( log 10 [p-value]; two-sided Student s t-test) across all gene strains in the study. (c) Protein domain structures of Hfd1p, highlighting residues involved in catalysis. (d) Serial dilutions of hfd1 yeast transformed with plasmids encoding the indicated Hfd1p variants grown on paba synthetic solid medias with glucose or glycerol. (e) Relative respiratory growth rates of hfd1 yeast transformed with plasmids encoding the indicated Hfd1p variants and grown in paba synthetic liquid media. (f) Growth curves showing the respiratory growth of hfd1 yeast in paba synthetic media with the additives shown. (g) Relative 4-HB abundance in hfd1 yeast cultured in paba media with the additives shown (mean log 2 [additive/unsupplemented] s.d., n = 3). (h) SDS-PAGE analysis (Coomassie stained gel) of protein fractions from an isolation of MBP-Hfd1p(C 25), MBP-ALDH3A1, and MBP-ALDH3A2(C 25) (WT and catalytically dead mutant for each). (i) Phylogenetic tree of human ALDH superfamily members and yeast Hfd1p. (j) Density of yeast (upon harvest) cultured in paba media 4-HB (mean s.d., n = 3). (k) Relative abundances of 4-HB, PPHB, and CoQ compared to WT yeast cultured in paba media (mean log 2 [ gene/wt with no additive] s.d., n = 3) as quantified by MS. (l) Whole proteome correlation map for yeast grown in paba media 4-HB (mean, n = 3). (m) Relative abundances of select proteins as quantified by MS (mean log 2 [ gene/wt], n = 3) analysis of yeast cultured in paba media 4-HB. (n) Serial dilutions of hfd1 yeast transformed with plasmids encoding the proteins shown and cultured on solid paba
8 synthetic media plates. (o) Enzyme activity of MBP-ALDH3A1 or MBP-ALDH3A2(C 25) against 4-HBz (200 M) or hexadecanal (200 M) (mean s.e.m., n = 3). (p) Table of enzyme kinetic parameters for MBP-Hfd1p(C 25), MBP-ALDH3A1, and MBP- ALDH3A2(C 25) (mean s.e.m., n = 3). (q) Representative enzyme kinetic curves for MBP-ALDH3A1 and MBP-ALDH3A2(C 25). *P < 0.05; **P < 0.01; ***P < (two-sided Student s t-test).
9 Supplementary Figure 8 Identification of respiration deficiency response pathways and potential biomarkers. (a) Projection of RC and RD strains onto the planes defined by principal component (PC) axes 1 and 2 for separate proteome, metabolome, and lipidome PC analyses. (b) RD versus RC proteome perturbation volcano plots (as in Fig. 3e) showing select functional groups (GO terms) significantly enriched (Bonferroni corrected p-values shown in figure) in either upregulated or downregulated proteins. (c) Box plots depicting median molecule fold changes for RC and RD strains (log 2 [RD or RC average/wt]) (n = 111 for RC, 41 for RD). Notch indicates 95% c.i. (d) Receiver operating characteristic (ROC) curves for select molecules depicting the false positive rates and true positive rates for prediction of respiration deficiency associated with particular molecule fold changes. AUC, area under the curve.
10 Supplementary Figure 9 Subtraction of shared responses to reveal deeper biochemical insight. (a) RDR-abundance adjustment of a representative molecule (Mls1p) by subtraction of the average fold change in abundance (mean log 2 [ gene/wt], n = 3) across respiration deficient (RD) strains. This adjustment was only performed within RD strains. (b) Plots comparing relative protein abundances between pairs of gene strains. Linear regression analysis of pairs of perturbation profiles before (left) and after (right) RD-abundance adjustment. Green points indicate molecules significantly perturbed in both mutants ( log 2 (FC) > 0.7; P < 0.05; two-sided Student s t-test) prior to RDR-adjustment. (c) Expanded view of highly correlated strains in the respiration proteomes correlation map (see Fig. 3b). (d) Procedure for normalization of the RDR. (e) Re-clustered respiration proteome strain-strain correlation map following RDR-adjustment (also shown in Fig. 3g).
11 Supplementary Figure 10 Molecular perturbations of yeast lacking yjr120w. (a) Relative protein abundances (mean log 2 [ yjr120w/wt], n = 3) versus statistical significance ( log 10 [p-value]; two-sided Student s t- test) as quantified by MS. (b) Relative Atp2p protein abundance (mean log 2 [ gene/wt], n = 3) versus statistical significance ( log 10 [pvalue]; two-sided Student s t-test) across all mutants in the study. (c) Genomic organization of yjr120w and atp2. (d) Serial dilutions of yeast transformed with the indicated plasmids grown on agar plates with glucose (to enable fermentation) or glycerol (to force respiration). (e) Fold changes in mrna abundances (mean gene/wt, n = 3) as quantified by real time polymerase chain reaction (RT- PCR) analysis. Yjr120w mrna was not detected (n.d.) in WT yeast, so imputation of this missing value was used to calculate the fold increase in yjr120w mrna shown for the atp2 strain. *P < 0.05; **P < 0.01; ***P < (two-sided Student s t-test).
12 Supplementary Figure 11 Features of multi-omic molecule covariance networks. (a) Network of all covariant molecules observed in each dataset ( 0.58, Bonferroni-adjusted P < 0.001; two-sided Student s t-test). (b) Regression analysis of pairs of RDR-associated molecules before and after RDR adjustment using Spearman s rank coefficient ( ). Points corresponding to RD and RC gene strains are indicated. (c) Distribution of calculated Spearman coefficients for all pairwise molecule covariance comparisons ( cutoff at ±0.58 used throughout the study is indicated). (d) Distribution of Bonferroni-adjusted p- values from all pairwise molecule comparisons (p-value cutoff at used throughout the study is indicated). (e) Bar chart indicating number of protein protein (P P), protein metabolite (P M), protein lipid (P L), metabolite metabolite (M M), metabolite lipid (M L), and lipid lipid (L L) edges in each dataset. (f) Box plots indicating the number of edges per node in the respiration, fermentation, and RDR-adjusted networks. (g) Network of all covariant RDR-associated molecules ( 0.58, Bonferroni-adjusted P < 0.001; two-sided Student s t-test) generated using the respiration (left) and RDR-adjusted (right) datasets. Nodes are highlighted according to GO category. (h) Box plots indicating the molecule covariance network (MCN) specificity coefficient for all nodes involved in mitochondrial translation in both the respiration and RDR-adjusted respiration RDR-associated molecule networks (shown in panel G). (i) Relative protein abundances (mean log 2 [ yor020w-a/wt], n = 2) versus statistical significance ( log 10 [p-value]; two-sided Student s t-test) as quantified by MS.
13 Supplementary Figure 12 Molecule covariance networks for uncharacterized proteins. Nearest neighbor molecule covariance networks for all uncharacterized proteins observed across the respiration, fermentation, and RDR-adjusted respiration datasets ( 0.58, Bonferroni-adjusted P < 0.001; two-sided Student s t-test). If more than 14 correlated molecules were present in a given covariance network, only the top 14 correlated molecules (nearest neighbors) are displayed.
14 Supplementary Figure 13 Examples of hypotheses that can be generated from a subset of the molecule covariance network analyses in this study. Nearest neighbor molecule covariance networks from uncharacterized proteins containing more than four connected nodes were tested for GO term enrichment using a Fisher s exact test with Benjamini Hochberg FDR adjustment to account for multiple hypothesis testing. Networks containing four or fewer connected nodes were analyzed manually for functionally related molecules. Based on these MCNA results, biological hypotheses about the functions of the uncharacterized proteins shown were developed.
15 Supplementary Figure 14 Hypothesized pathways for Aro9p, Aro10p, and Aim18p. (a) Putative biochemical functions of Aro9p and Aro10p in catabolism of tyrosine and phenylalanine. (b) Predicted functions for Aro9p and Aro10p in the Tyr-to-4-HB-to-CoQ pathway. (c) Protein sequence alignments of Aim18p (S. cerevisiae) and chalcone isomerases (CHI) from Medicago and Arabidopsis highlighting conservation of putative catalytic residues (starred residues). (d) Example of a CHI catalyzed reaction (upper scheme) and the hypothesized pathway of Aim18p action (lower scheme).
16 Supplementary Notes Supplementary Note 1. Development of a stable and reproducible respiration culture condition. To profile diverse yeast strains during respiratory growth, when mitochondrial OxPhos is highly active, we first needed to develop a distinct respiration condition suitable for large-scale investigation. Early log phase fermentation cultures repress mitochondrial respiration, cultures containing solely non-fermentable sugars preclude growth of respiration deficient yeast, and high glucose cultures grown past the diauxic shift are too biologically dynamic to allow reproducible sampling across a large scale study 43, 44. To overcome these problems, we developed a culture system that includes low glucose (1 g/l) and high glycerol (30 g/l), enabling a short fermentation phase followed by a longer respiration phase. This respiration condition affords steady growth and a stable biological state as reflected by a proteome that is constant over multiple hours (Supplementary Fig. 1c e) and, thus, an essential window for reproducible sample harvesting. Supplementary Note 2. ΔGene-specific phenotype detection. To identify Δgene-specific phenotypes, we broadly surveyed our data for characteristic outlier abundance measurements. For each profiled molecule (in both respiration and fermentation growth conditions) we separated potential Δgene-specific measurements into two groups: positive log 2 fold change (log 2 [Δgene/WT]) and negative log 2 fold change. These two sets were then plotted individually with log 2 fold change and log 10 (p-value [two-sided Student s t-test]) along the x- and y- axes, respectively. Data were normalized such that the largest log 2 fold change and largest log 10 (p-value) were set equal to 1. Considering the three largest fold changes where P < 0.05, we calculated the Euclidean distance to all neighboring data points and stored the smallest result. A requirement was imposed that all considered neighbors have a smaller fold change than the data point being considered. It is anticipated that data points corresponding to Δgene-specific phenotypes will be outliers in the described plots and have large associated nearest-neighbor Euclidean distances. The described routine yielded three separate distances, the largest of which was stored for further analysis. The results of this analysis and representative examples are highlighted (Fig. 2, Supplementary Figs. 5 and 6). We observed maximal Euclidean distances across a range of to We set a cutoff for classification as a Δgene-specific phenotype at 0.70 and report 714 molecules (4.6% of considered cases across both culture conditions) which exceed this threshold (Supplementary Table 4). This procedure provided a useful first pass analysis and afforded a truncated set of leads, which were used to develop biological hypotheses. Supplementary Note 3. Lack of effect of Dpl1p disruption on the Tyr-to-4-HB-to-CoQ pathway. To test the idea that the CoQ biosynthesis and sphingolipid catabolism pathways are independent, we examined Δdpl1 yeast, which lack a known dihydrosphingosine phosphate lyase. Δdpl1 yeast show neither a paba respiratory growth phenotype nor CoQ deficiency (Supplementary Fig. 7j,k). These results demonstrate that disruption of the Tyr-to-4- HB pathway in Δhfd1 yeast is not downstream of a defect in sphingolipid metabolism. Furthermore, proteome analyses showed that Δhfd1 cultured without 4-HB and paba are similar to Δcoq8 yeast but not Δdpl1 yeast and adding 4-HB to Δhfd1 cultures returns their proteomes to WT-like profiles (Supplementary Fig. 7l,m). Supplementary Note 4. Quantitative definition of the respiration deficiency response (RDR). To quantitatively define the RDR, we categorized strains as respiration deficient (RD) or competent (RC) and examined differences between these two groups. Principal component analysis of the Y3K respiration dataset revealed marked separation of RD and RC strains (Fig. 3c and Supplementary Fig. 8a). The underlying phenotype changes that distinguish RD and RC strains include proteins, lipids, and metabolites (Fig. 3d and Supplementary Table 5). RDR perturbations include significant decreases in ATP synthase, TCA cycle, and MICOS proteins (Fig. 3e,f and Supplementary Fig. 8b), likely to decrease allocation of useless proteome mass to dysfunctional mitochondria 45. Importantly, the RDR also includes a positive response, and numerous proteins including protein folding, NADH metabolism, and proteasome assembly proteins are significantly upregulated in RD strains (Fig. 3e,f). Numerous individual molecules including lactate, alanine, 2-hydroxyglutarate, tyrosol, 4-HB, Gpx2p, and Ahp1p, among many others are significantly perturbed in RD strains and strongly predictive of respiration deficiency (Supplementary Fig. 8c,d). Our quantitative assessment of the RDR highlights biochemical features of the cellular response to defects in mitochondrial respiration, and suggests that a multi-omic assessment of proteins, lipids, and metabolites could afford a highly specific biomarker panel for diseases affected by OxPhos deficiency.
17 Supplementary Note 5. RDR normalization procedure. Dgene strains were classified as RD (51) or respiration competent (RC) (123) based on observation of a common perturbation profile signature in the respiration culture condition. For each molecule we calculated an RDR score. This metric represents the proportion of RD Dgene strains over which the molecule was consistently perturbed, relative to all RD Dgene strains where the molecule was quantified. Across all RD Dgene strains, 776 molecules were identified as having an RDR score > 0.95 (consistently perturbed across more than 95% of RD Dgene strains where quantified) and classified as RDR-associated. (Supplementary Table 6). The individual measurements of these RDR-associated molecules were then mean normalized ( RDR-adjusted ) using abundance values from RD Dgene strains. This normalization procedure revealed characteristic deviations from the general RDR (Supplementary Fig. 9). Importantly, this procedure enables visualization of Δgene-specific changes. For example, prior to RDR normalization, the expected decrease in Coq8p in Δcoq8 yeast is obscured by RDR-associated proteins with large abundance changes (Supplementary Fig. 9d). RDR normalization not only uncovers the decrease in Coq8p, but a significant decrease in Coq5p, a functionally-related CoQ biosynthesis protein, also becomes readily apparent (Supplementary Fig. 9d). Supplementary Note 6. Molecular defects of Δyjr120w yeast. To examine the molecular basis for the CoQ deficiency of Δyjr120w yeast, we inspected our proteomics dataset, which revealed significant decreases in ATP synthase proteins, especially Atp2p (Supplementary Fig. 10a). Compared to other strains, the large decrease in Atp2p is unique to Δyjr120w and Δatp2 (Supplementary Fig. 10b). A relationship between yjr120w and atp2 is also suggested by their genetic proximity (Supplementary Fig. 10c). Plasmid overexpression of atp2 rescues the Δyjr120w respiratory growth defect (Supplementary Fig. 10d), indicating a functional relationship between atp2 and yjr120w in vivo. A decrease in atp2 mrna in the Δyjr120w strain is a component of the underlying mechanism (Supplementary Fig. 10e). Interestingly, CoQ deficiency was also observed in Δatp2 yeast (Fig. 3h). Supplementary Note 7. Predicted enzymatic functions of Aim18p, Aro9p, and Aro10p. Since 1907, yeast have been known to catabolize amino acids into fusel (German for bad liquor ) alcohols through the Ehrlich pathway 46, 47, but the physiological roles for the enzymes involved such as Aro9p and Aro10p are not fully understood. Aro9p and Aro10p were previously thought to provide a simple catabolic route for extracting nitrogen from aromatic amino acids 48 (Supplementary Fig. 14a), but our MCNA unexpectedly indicated strong correlations between Aro9p, Aro10p, and proteins involved in mitochondrial respiration (Fig. 4d,e), suggesting a more complicated biological function that supports OxPhos. We hypothesized that this function might be in the Tyr-to-4- HB-to-CoQ pathway (Supplementary Fig. 14b), given the putative enzymatic activities of Aro9p and Aro10p in tyrosine and phenylalanine metabolism. Consistently, when cultured in paba media, Δaro9 and Δaro10 yeast are deficient in CoQ and PPHB (Fig. 4f). Aim18p is a protein of undefined molecular function that has been detected in mitochondria 49 and potentially linked to mitochondrial inheritance (Altered Inheritance of Mitochondria, AIM ) by large-scale studies in yeast 50. Protein sequence alignments show that Aim18p contains a chalcone-flavone isomerase (CHI)-like domain (Supplementary Fig. 14c), whose homologs in plants typically function on aromatic small molecules (chalcones) (Supplementary Fig. 14d) Given the potential for this protein domain to catalyze modifications of aromatic small molecules, we hypothesized that Aim18p might function in the Tyr-to-4-HB pathway to produce the CoQ headgroup (Supplementary Fig. 14d). Consistently, when cultured in paba media, we observed deficiency of PPHB in Δaim18 yeast (Fig. 4f).
18 Supplementary Table Captions Supplementary Table 1. Knockout yeast strains. Table of single-gene deletion ( gene) yeast strains investigated in this study and their harvest culture densities. For each gene deleted in a strain studied, the first tab includes systematic yeast gene name, standard gene name, Entrez gene ID, UniProt ID, and human homolog(s). The second tab shows the culture densities upon harvest (growth phenotypes), and it includes the systematic yeast gene name, the standard gene name, the average culture densities at the harvest time point (mean, n = 3), and the corresponding standard deviations, fold changes (KO/WT), and p-values (Student s t-test) for respiration and fermentation cultures. Supplementary Table 2. Profiled biomolecules. Table of all 4505 molecules profiled in the study. Includes molecule type (protein, lipid, or metabolite), molecule name, standard gene name (for proteins) or standard lipid name, systematic gene name (for proteins), and UniProt ID (for proteins). For lipids, the numbers in parentheses indicate the number of carbons in the acyl tail(s) and the number of carbon-carbon double bonds in the chains (carbons:double_bonds). Supplementary Table 3. Quantitative dataset. Table containing quantitative measurements and descriptive statistics used throughout the Y3K study. Average fold changes in molecule abundances (mean log 2 [ gene/wt], n = 3) for all strains and all molecules in the respiration and fermentation datasets are shown on tabs labeled KO vs WT_Resp ( LFQ) and KO vs WT_Ferm ( LFQ). Corresponding standard deviations, and p-values (2-tailed t-test [homostatic]) for all measured fold changes are shown on separate tabs labeled accordingly with (Std. Dev.) and (P-Value). Each tab contains a table with rows corresponding to molecules and columns corresponding to the 174 single gene knockout ( gene) strains profiled in the study. Supplementary Table 4. Δgene-specific phenotypes. Table of unique Δgene-phenotype relationships identified in this study. Includes molecule name (standard gene name followed by systematic gene name for all proteins), yeast deletion strain (standard gene name), calculated Euclidean distance, and associated growth condition (respiration or fermentation). Supplementary Table 5. Respiration deficient strains vs respiration competent strains. Table of average fold change in molecule abundances (mean log 2 [RD strains/rc strains]). Includes molecule identifiers (including UniProt IDs, symbols, and systematic names for proteins), average fold change (mean log 2 [RD strains/rc strains]), log 10 (p-value), and select GO terms corresponding to those highlighted in Fig. 3 and Supplementary Fig. 8. Supplementary Table 6. Respiration deficient strains versus wild type. Table of average fold change in molecule abundances (mean log 2 [RD strains/wt]). Includes molecule identifiers (including UniProt IDs, symbols, and systematic names for proteins), fraction of RD strains showing consistent perturbation of each molecule, RDR score (see Methods), and average fold change (mean log 2 [RD strains/wt]). Supplementary Table 7. Δgene Δgene perturbation profile correlations. Table of Δgene Δgene perturbation profile correlations (Pearson coefficients). Includes the gene knocked out of strain one in the pairwise comparison, the gene knocked out of strain two in the pairwise comparison, the Pearson coefficient, and the Ome (proteome, lipidome, or metabolome) used for the regression analysis. Coefficients are only reported for Δgene Δgene pairs meeting the criteria outlined in the Methods under the heading Regression analysis of phenotype changes. Separate tabs are included for the respiration (resp), fermentation (ferm), and RDRadjusted respiration (resp-rdr) datasets. Supplementary Table 8. Molecule covariance network analysis results. Table of 288,794 pairs of covariant molecules ( ρ 0.58 and Bonferroni-adjusted P < 0.001) identified in the Y3K dataset. Includes molecule names (standard protein name followed by systematic protein name) and types (protein, lipid, or metabolite) for molecule one and molecule two, Spearman s correlation coefficients (ρ), and Bonferroniadjusted P-values. Separate tabs are included for the respiration (resp), fermentation (ferm), and RDR-adjusted respiration (resp-rdr) datasets.
19 References 43. Picotti, P., Bodenmiller, B., Mueller, L.N., Domon, B. & Aebersold, R. Full dynamic range proteome analysis of S. cerevisiae by targeted proteomics. Cell 138, (2009). 44. Casanovas, A. et al. Quantitative analysis of proteome and lipidome dynamics reveals functional regulation of global lipid metabolism. Chem Biol 22, (2015). 45. Basan, M. et al. Overflow metabolism in Escherichia coli results from efficient proteome allocation. Nature 528, (2015). 46. Ehrlich, F. Über die Bedingungen der Fuselölbildung und über ihren Zusammenhang mit dem Eiweißaufbau der Hefe. Ber. Dtsch. Chem. Ges. 40, (1907). 47. Hazelwood, L.A., Daran, J.M., van Maris, A.J., Pronk, J.T. & Dickinson, J.R. The Ehrlich pathway for fusel alcohol production: a century of research on Saccharomyces cerevisiae metabolism. Appl Environ Microbiol 74, (2008). 48. Kneen, M.M. et al. Characterization of a thiamin diphosphate-dependent phenylpyruvate decarboxylase from Saccharomyces cerevisiae. FEBS J 278, (2011). 49. Reinders, J., Zahedi, R.P., Pfanner, N., Meisinger, C. & Sickmann, A. Toward the complete yeast mitochondrial proteome: multidimensional separation techniques for mitochondrial proteomics. J Proteome Res 5, (2006). 50. Hess, D.C. et al. Computationally driven, quantitative experiments discover genes required for mitochondrial biogenesis. PLoS Genet 5, e (2009). 51. Gensheimer, M. & Mushegian, A. Chalcone isomerase family and fold: no longer unique to plants. Protein Sci 13, (2004). 52. Ngaki, M.N. et al. Evolution of the chalcone-isomerase fold from fatty-acid binding to stereospecific catalysis. Nature 485, (2012). 53. Jez, J.M., Bowman, M.E., Dixon, R.A. & Noel, J.P. Structure and mechanism of the evolutionarily unique plant enzyme chalcone isomerase. Nat Struct Biol 7, (2000).
Department of Chemistry, Université de Montréal, C.P. 6128, Succursale centre-ville, Montréal, Québec, H3C 3J7, Canada.
 Phosphoproteome dynamics of Saccharomyces cerevisiae under heat shock and cold stress Evgeny Kanshin 1,5, Peter Kubiniok 1,2,5, Yogitha Thattikota 1,3, Damien D Amours 1,3 and Pierre Thibault 1,2,4 * 1
Phosphoproteome dynamics of Saccharomyces cerevisiae under heat shock and cold stress Evgeny Kanshin 1,5, Peter Kubiniok 1,2,5, Yogitha Thattikota 1,3, Damien D Amours 1,3 and Pierre Thibault 1,2,4 * 1
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Experimental design and workflow utilized to generate the WMG Protein Atlas.
 Supplementary Figure 1 Experimental design and workflow utilized to generate the WMG Protein Atlas. (a) Illustration of the plant organs and nodule infection time points analyzed. (b) Proteomic workflow
Supplementary Figure 1 Experimental design and workflow utilized to generate the WMG Protein Atlas. (a) Illustration of the plant organs and nodule infection time points analyzed. (b) Proteomic workflow
Cellular Respiration. Cellular Respiration. C 6 H 12 O 6 + 6O > 6CO 2 + 6H energy. Heat + ATP. You need to know this!
 Cellular Respiration LISA Biology Cellular Respiration C 6 H 12 O 6 + 6O 2 - - - - - > 6CO 2 + 6H 2 0 + energy You need to know this! Heat + ATP 1 Did that equation look familiar? * The equation for cellular
Cellular Respiration LISA Biology Cellular Respiration C 6 H 12 O 6 + 6O 2 - - - - - > 6CO 2 + 6H 2 0 + energy You need to know this! Heat + ATP 1 Did that equation look familiar? * The equation for cellular
Metabolism Energy Pathways Biosynthesis. Catabolism Anabolism Enzymes
 Topics Microbial Metabolism Metabolism Energy Pathways Biosynthesis 2 Metabolism Catabolism Catabolism Anabolism Enzymes Breakdown of complex organic molecules in order to extract energy and dform simpler
Topics Microbial Metabolism Metabolism Energy Pathways Biosynthesis 2 Metabolism Catabolism Catabolism Anabolism Enzymes Breakdown of complex organic molecules in order to extract energy and dform simpler
ABSTRACT INTRODUCTION
 /, 2017, Vol. 8, (No. 19), pp: 30672-30691 Specific changes in mitochondrial lipidome alter mitochondrial proteome and increase the geroprotective efficiency of lithocholic acid in chronologically aging
/, 2017, Vol. 8, (No. 19), pp: 30672-30691 Specific changes in mitochondrial lipidome alter mitochondrial proteome and increase the geroprotective efficiency of lithocholic acid in chronologically aging
Higher Biology. Unit 2: Metabolism and Survival Topic 2: Respiration. Page 1 of 25
 Higher Biology Unit 2: Metabolism and Survival Topic 2: Respiration Page 1 of 25 Sub Topic: Respiration I can state that: All living cells carry out respiration. ATP is the energy currency of the cell
Higher Biology Unit 2: Metabolism and Survival Topic 2: Respiration Page 1 of 25 Sub Topic: Respiration I can state that: All living cells carry out respiration. ATP is the energy currency of the cell
Chemical Energy. Valencia College
 9 Pathways that Harvest Chemical Energy Valencia College 9 Pathways that Harvest Chemical Energy Chapter objectives: How Does Glucose Oxidation Release Chemical Energy? What Are the Aerobic Pathways of
9 Pathways that Harvest Chemical Energy Valencia College 9 Pathways that Harvest Chemical Energy Chapter objectives: How Does Glucose Oxidation Release Chemical Energy? What Are the Aerobic Pathways of
ADP, ATP and Cellular Respiration
 ADP, ATP and Cellular Respiration What Is ATP? Energy used by all Cells Adenosine Triphosphate Organic molecule containing highenergy Phosphate bonds Chemical Structure of ATP Adenine Base 3 Phosphates
ADP, ATP and Cellular Respiration What Is ATP? Energy used by all Cells Adenosine Triphosphate Organic molecule containing highenergy Phosphate bonds Chemical Structure of ATP Adenine Base 3 Phosphates
Discussion of Prism modules and predicted interactions (Fig. 4)
 SUPPLEMENTARY NOTES Discussion of Prism modules and predicted interactions (Fig. 4) a. Interactions of the TCA-cycle, respiratory chain, and ATP synthetase with the amino acid biosynthesis modules. Given
SUPPLEMENTARY NOTES Discussion of Prism modules and predicted interactions (Fig. 4) a. Interactions of the TCA-cycle, respiratory chain, and ATP synthetase with the amino acid biosynthesis modules. Given
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
A cell has enough ATP to last for about three seconds.
 Energy Transformation: Cellular Respiration Outline 1. Energy and carbon sources in living cells 2. Sources of cellular ATP 3. Turning chemical energy of covalent bonds between C-C into energy for cellular
Energy Transformation: Cellular Respiration Outline 1. Energy and carbon sources in living cells 2. Sources of cellular ATP 3. Turning chemical energy of covalent bonds between C-C into energy for cellular
Ch. 9 Cell Respiration. Title: Oct 15 3:24 PM (1 of 53)
 Ch. 9 Cell Respiration Title: Oct 15 3:24 PM (1 of 53) Essential question: How do cells use stored chemical energy in organic molecules and to generate ATP? Title: Oct 15 3:28 PM (2 of 53) Title: Oct 19
Ch. 9 Cell Respiration Title: Oct 15 3:24 PM (1 of 53) Essential question: How do cells use stored chemical energy in organic molecules and to generate ATP? Title: Oct 15 3:28 PM (2 of 53) Title: Oct 19
In glycolysis, glucose is converted to pyruvate. If the pyruvate is reduced to lactate, the pathway does not require O 2 and is called anaerobic
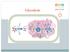 Glycolysis 1 In glycolysis, glucose is converted to pyruvate. If the pyruvate is reduced to lactate, the pathway does not require O 2 and is called anaerobic glycolysis. If this pyruvate is converted instead
Glycolysis 1 In glycolysis, glucose is converted to pyruvate. If the pyruvate is reduced to lactate, the pathway does not require O 2 and is called anaerobic glycolysis. If this pyruvate is converted instead
3.7.1 Define cell respiration [Cell respiration is the controlled release of energy from organic compounds in cells to form ATP]
![3.7.1 Define cell respiration [Cell respiration is the controlled release of energy from organic compounds in cells to form ATP] 3.7.1 Define cell respiration [Cell respiration is the controlled release of energy from organic compounds in cells to form ATP]](/thumbs/72/66350985.jpg) 3.7 Cell respiration ( Chapter 9 in Campbell's book) 3.7.1 Define cell respiration [Cell respiration is the controlled release of energy from organic compounds in cells to form ATP] Organic compounds store
3.7 Cell respiration ( Chapter 9 in Campbell's book) 3.7.1 Define cell respiration [Cell respiration is the controlled release of energy from organic compounds in cells to form ATP] Organic compounds store
Ch 9: Cellular Respiration
 Ch 9: Cellular Respiration Cellular Respiration An overview Exergonic reactions and catabolic pathway Energy stored in bonds of food molecules is transferred to ATP Cellular respiration provides the energy
Ch 9: Cellular Respiration Cellular Respiration An overview Exergonic reactions and catabolic pathway Energy stored in bonds of food molecules is transferred to ATP Cellular respiration provides the energy
Energy Transformation: Cellular Respiration Outline 1. Sources of cellular ATP 2. Turning chemical energy of covalent bonds between C-C into energy
 Energy Transformation: Cellular Respiration Outline 1. Sources of cellular ATP 2. Turning chemical energy of covalent bonds between C-C into energy for cellular work (ATP) 3. Importance of electrons and
Energy Transformation: Cellular Respiration Outline 1. Sources of cellular ATP 2. Turning chemical energy of covalent bonds between C-C into energy for cellular work (ATP) 3. Importance of electrons and
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Chapter 5. Microbial Metabolism
 Chapter 5 Microbial Metabolism Metabolism Collection of controlled biochemical reactions that take place within a microbe Ultimate function of metabolism is to reproduce the organism Metabolic Processes
Chapter 5 Microbial Metabolism Metabolism Collection of controlled biochemical reactions that take place within a microbe Ultimate function of metabolism is to reproduce the organism Metabolic Processes
III. 6. Test. Respiració cel lular
 III. 6. Test. Respiració cel lular Chapter Questions 1) What is the term for metabolic pathways that release stored energy by breaking down complex molecules? A) anabolic pathways B) catabolic pathways
III. 6. Test. Respiració cel lular Chapter Questions 1) What is the term for metabolic pathways that release stored energy by breaking down complex molecules? A) anabolic pathways B) catabolic pathways
Cellular Respiration
 Cellular Respiration 1. To perform cell work, cells require energy. a. A cell does three main kinds of work: i. Mechanical work, such as the beating of cilia, contraction of muscle cells, and movement
Cellular Respiration 1. To perform cell work, cells require energy. a. A cell does three main kinds of work: i. Mechanical work, such as the beating of cilia, contraction of muscle cells, and movement
How Did Energy-Releasing Pathways Evolve? (cont d.)
 How Did Energy-Releasing Pathways Evolve? (cont d.) 7.1 How Do Cells Access the Chemical Energy in Sugars? In order to use the energy stored in sugars, cells must first transfer it to ATP The energy transfer
How Did Energy-Releasing Pathways Evolve? (cont d.) 7.1 How Do Cells Access the Chemical Energy in Sugars? In order to use the energy stored in sugars, cells must first transfer it to ATP The energy transfer
How Cells Harvest Energy. Chapter 7. Respiration
 How Cells Harvest Energy Chapter 7 Respiration Organisms classified on how they obtain energy: autotrophs: produce their own organic molecules through photosynthesis heterotrophs: live on organic compounds
How Cells Harvest Energy Chapter 7 Respiration Organisms classified on how they obtain energy: autotrophs: produce their own organic molecules through photosynthesis heterotrophs: live on organic compounds
MIDDLETOWN HIGH SCHOOL SOUTH BIOLOGY
 MIDDLETOWN HIGH SCHOOL SOUTH BIOLOGY BOOKLET 10 NAME: CLASS: 1 S.Tagore Middletown South High School March 2013 LEARNING OUTCOMES The role and production of ATP (a) Importance, role and structure of ATP
MIDDLETOWN HIGH SCHOOL SOUTH BIOLOGY BOOKLET 10 NAME: CLASS: 1 S.Tagore Middletown South High School March 2013 LEARNING OUTCOMES The role and production of ATP (a) Importance, role and structure of ATP
Metabolism. Chapter 8 Microbial Metabolism. Metabolic balancing act. Catabolism Anabolism Enzymes. Topics. Metabolism Energy Pathways Biosynthesis
 Chapter 8 Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis Catabolism Anabolism Enzymes Metabolism 1 2 Metabolic balancing act Catabolism and anabolism simple model Catabolism Enzymes
Chapter 8 Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis Catabolism Anabolism Enzymes Metabolism 1 2 Metabolic balancing act Catabolism and anabolism simple model Catabolism Enzymes
Cell Respiration. Anaerobic & Aerobic Respiration
 Cell Respiration Anaerobic & Aerobic Respiration Understandings/Objectives 2.8.U1: Cell respiration is the controlled release of energy from organic compounds to produce ATP. Define cell respiration State
Cell Respiration Anaerobic & Aerobic Respiration Understandings/Objectives 2.8.U1: Cell respiration is the controlled release of energy from organic compounds to produce ATP. Define cell respiration State
MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question.
 Respiration Practice Name MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Which of the following statements describes NAD+? A) NAD+ can donate
Respiration Practice Name MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Which of the following statements describes NAD+? A) NAD+ can donate
Cellular Respiration Harvesting Chemical Energy ATP
 Cellular Respiration Harvesting Chemical Energy ATP 2006-2007 What s the point? The point is to make ATP! ATP 2006-2007 Harvesting stored energy Energy is stored in organic molecules carbohydrates, fats,
Cellular Respiration Harvesting Chemical Energy ATP 2006-2007 What s the point? The point is to make ATP! ATP 2006-2007 Harvesting stored energy Energy is stored in organic molecules carbohydrates, fats,
Chapter 8. Metabolism. Topics in lectures 15 and 16. Chemical foundations Catabolism Biosynthesis
 Chapter 8 Topics in lectures 15 and 16 Metabolism Chemical foundations Catabolism Biosynthesis 1 Metabolism Chemical Foundations Enzymes REDOX Catabolism Pathways Anabolism Principles and pathways 2 Enzymes
Chapter 8 Topics in lectures 15 and 16 Metabolism Chemical foundations Catabolism Biosynthesis 1 Metabolism Chemical Foundations Enzymes REDOX Catabolism Pathways Anabolism Principles and pathways 2 Enzymes
BY: RASAQ NURUDEEN OLAJIDE
 BY: RASAQ NURUDEEN OLAJIDE LECTURE CONTENT INTRODUCTION CITRIC ACID CYCLE (T.C.A) PRODUCTION OF ACETYL CoA REACTIONS OF THE CITIRC ACID CYCLE THE AMPHIBOLIC NATURE OF THE T.C.A CYCLE THE GLYOXYLATE CYCLE
BY: RASAQ NURUDEEN OLAJIDE LECTURE CONTENT INTRODUCTION CITRIC ACID CYCLE (T.C.A) PRODUCTION OF ACETYL CoA REACTIONS OF THE CITIRC ACID CYCLE THE AMPHIBOLIC NATURE OF THE T.C.A CYCLE THE GLYOXYLATE CYCLE
4. Which step shows a split of one molecule into two smaller molecules? a. 2. d. 5
 1. Which of the following statements about NAD + is false? a. NAD + is reduced to NADH during both glycolysis and the citric acid cycle. b. NAD + has more chemical energy than NADH. c. NAD + is reduced
1. Which of the following statements about NAD + is false? a. NAD + is reduced to NADH during both glycolysis and the citric acid cycle. b. NAD + has more chemical energy than NADH. c. NAD + is reduced
chapter 1 - fig. 2 Mechanism of transcriptional control by ppar agonists.
 chapter 1 - fig. 1 The -omics subdisciplines. chapter 1 - fig. 2 Mechanism of transcriptional control by ppar agonists. 201 figures chapter 1 chapter 2 - fig. 1 Schematic overview of the different steps
chapter 1 - fig. 1 The -omics subdisciplines. chapter 1 - fig. 2 Mechanism of transcriptional control by ppar agonists. 201 figures chapter 1 chapter 2 - fig. 1 Schematic overview of the different steps
Cellular Pathways That Harvest Chemical Energy. Cellular Pathways That Harvest Chemical Energy. Cellular Pathways In General
 Cellular Pathways That Harvest Chemical Energy A. Obtaining Energy and Electrons from Glucose Lecture Series 12 Cellular Pathways That Harvest Chemical Energy B. An Overview: Releasing Energy from Glucose
Cellular Pathways That Harvest Chemical Energy A. Obtaining Energy and Electrons from Glucose Lecture Series 12 Cellular Pathways That Harvest Chemical Energy B. An Overview: Releasing Energy from Glucose
Biology Kevin Dees. Chapter 9 Harvesting Chemical Energy: Cellular Respiration
 Chapter 9 Harvesting Chemical Energy: Cellular Respiration Life is Work!!! Biology Kevin Dees Catabolic pathways and ATP production Catabolic pathways release energy by breaking down large molecules into
Chapter 9 Harvesting Chemical Energy: Cellular Respiration Life is Work!!! Biology Kevin Dees Catabolic pathways and ATP production Catabolic pathways release energy by breaking down large molecules into
Enzymes what are they?
 Topic 11 (ch8) Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis 1 Catabolism Anabolism Enzymes Metabolism 2 Metabolic balancing act Catabolism Enzymes involved in breakdown of complex
Topic 11 (ch8) Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis 1 Catabolism Anabolism Enzymes Metabolism 2 Metabolic balancing act Catabolism Enzymes involved in breakdown of complex
Chapter 9: Cellular Respiration: Harvesting Chemical Energy
 AP Biology Reading Guide Name: Date: Period Chapter 9: Cellular Respiration: Harvesting Chemical Energy Overview: Before getting involved with the details of cellular respiration and photosynthesis, take
AP Biology Reading Guide Name: Date: Period Chapter 9: Cellular Respiration: Harvesting Chemical Energy Overview: Before getting involved with the details of cellular respiration and photosynthesis, take
Chapter 9: Cellular Respiration
 Chapter 9: Cellular Respiration To perform their many tasks, living cells require energy from outside sources. Energy stored in food utimately comes from the sun. Photosynthesis makes the raw materials
Chapter 9: Cellular Respiration To perform their many tasks, living cells require energy from outside sources. Energy stored in food utimately comes from the sun. Photosynthesis makes the raw materials
Metabolism. Topic 11&12 (ch8) Microbial Metabolism. Metabolic Balancing Act. Topics. Catabolism Anabolism Enzymes
 Topic 11&12 (ch8) Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis 1 Catabolism Anabolism Enzymes Metabolism 2 Metabolic Balancing Act Catabolism Enzymes involved in breakdown of complex
Topic 11&12 (ch8) Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis 1 Catabolism Anabolism Enzymes Metabolism 2 Metabolic Balancing Act Catabolism Enzymes involved in breakdown of complex
Metabolism. Metabolic pathways. BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Metabolism Metabolism is the chemical change of
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Metabolism Metabolism is the chemical change of
Cellular Respiration Let s get energized!
 Copyrighted by Amy Brown Science Stuff Cellular Respiration Let s get energized! Amy Brown Science Food provides living things with the: chemical building blocks they need to grow and reproduce. Food serves
Copyrighted by Amy Brown Science Stuff Cellular Respiration Let s get energized! Amy Brown Science Food provides living things with the: chemical building blocks they need to grow and reproduce. Food serves
Chapter 8. Metabolism. Topics in lectures 15 and 16. Chemical foundations Catabolism Biosynthesis
 Chapter 8 Topics in lectures 15 and 16 Metabolism Chemical foundations Catabolism Biosynthesis 1 Metabolism Chemical Foundations Enzymes REDOX Catabolism Pathways Anabolism Principles and pathways 2 Chemical
Chapter 8 Topics in lectures 15 and 16 Metabolism Chemical foundations Catabolism Biosynthesis 1 Metabolism Chemical Foundations Enzymes REDOX Catabolism Pathways Anabolism Principles and pathways 2 Chemical
Reading Assignments. A. Energy and Energy Conversions. Lecture Series 9 Cellular Pathways That Harvest Chemical Energy. gasoline) or elevated mass.
 Lecture Series 9 Cellular Pathways That Harvest Chemical Energy Reading Assignments Review Chapter 3 Energy, Catalysis, & Biosynthesis Read Chapter 13 How Cells obtain Energy from Food Read Chapter 14
Lecture Series 9 Cellular Pathways That Harvest Chemical Energy Reading Assignments Review Chapter 3 Energy, Catalysis, & Biosynthesis Read Chapter 13 How Cells obtain Energy from Food Read Chapter 14
Cellular Respiration
 Cellular I can describe cellular respiration Cellular respiration is a series of metabolic pathways releasing energy from a foodstuff e.g. glucose. This yields energy in the form of ATP adenosine P i P
Cellular I can describe cellular respiration Cellular respiration is a series of metabolic pathways releasing energy from a foodstuff e.g. glucose. This yields energy in the form of ATP adenosine P i P
Mitochondria and ATP Synthesis
 Mitochondria and ATP Synthesis Mitochondria and ATP Synthesis 1. Mitochondria are sites of ATP synthesis in cells. 2. ATP is used to do work; i.e. ATP is an energy source. 3. ATP hydrolysis releases energy
Mitochondria and ATP Synthesis Mitochondria and ATP Synthesis 1. Mitochondria are sites of ATP synthesis in cells. 2. ATP is used to do work; i.e. ATP is an energy source. 3. ATP hydrolysis releases energy
Cellular Respiration
 Cellular Respiration C 6 H 12 O 6 + 6O 2 -----> 6CO 2 + 6H 2 0 + energy (heat and ATP) 1. Energy Capacity to move or change matter Forms of energy are important to life include Chemical, radiant (heat
Cellular Respiration C 6 H 12 O 6 + 6O 2 -----> 6CO 2 + 6H 2 0 + energy (heat and ATP) 1. Energy Capacity to move or change matter Forms of energy are important to life include Chemical, radiant (heat
INTEGRATION OF GENERAL AMINO ACID CONTROL AND TOR REGULATORY PATHWAYS IN NITROGEN ASSIMILATION IN YEAST
 INTEGRATION OF GENERAL AMINO ACID CONTROL AND TOR REGULATORY PATHWAYS IN NITROGEN ASSIMILATION IN YEAST Kirk A. Staschke 1, Souvik Dey 1, John M. Zaborske 2, Lakshmi Reddy Palam 1, Jeanette N. McClintick
INTEGRATION OF GENERAL AMINO ACID CONTROL AND TOR REGULATORY PATHWAYS IN NITROGEN ASSIMILATION IN YEAST Kirk A. Staschke 1, Souvik Dey 1, John M. Zaborske 2, Lakshmi Reddy Palam 1, Jeanette N. McClintick
MITOCHONDRIA LECTURES OVERVIEW
 1 MITOCHONDRIA LECTURES OVERVIEW A. MITOCHONDRIA LECTURES OVERVIEW Mitochondrial Structure The arrangement of membranes: distinct inner and outer membranes, The location of ATPase, DNA and ribosomes The
1 MITOCHONDRIA LECTURES OVERVIEW A. MITOCHONDRIA LECTURES OVERVIEW Mitochondrial Structure The arrangement of membranes: distinct inner and outer membranes, The location of ATPase, DNA and ribosomes The
1- Which of the following statements is TRUE in regards to eukaryotic and prokaryotic cells?
 Name: NetID: Exam 3 - Version 1 October 23, 2017 Dr. A. Pimentel Each question has a value of 4 points and there are a total of 160 points in the exam. However, the maximum score of this exam will be capped
Name: NetID: Exam 3 - Version 1 October 23, 2017 Dr. A. Pimentel Each question has a value of 4 points and there are a total of 160 points in the exam. However, the maximum score of this exam will be capped
respiration mitochondria mitochondria metabolic pathways reproduction can fuse or split DRP1 interacts with ER tubules chapter DRP1 ER tubule
 mitochondria respiration chapter 3-4 shape highly variable can fuse or split structure outer membrane inner membrane cristae intermembrane space mitochondrial matrix free ribosomes respiratory enzymes
mitochondria respiration chapter 3-4 shape highly variable can fuse or split structure outer membrane inner membrane cristae intermembrane space mitochondrial matrix free ribosomes respiratory enzymes
Bio 111 Study Guide Chapter 7 Cellular Respiration & Fermentation
 Bio 111 Study Guide Chapter 7 Cellular Respiration & Fermentation BEFORE CLASS: Reading: Read the whole chapter from pp. 141-158. In Concept 7.1, pay special attention to oxidation & reduction and the
Bio 111 Study Guide Chapter 7 Cellular Respiration & Fermentation BEFORE CLASS: Reading: Read the whole chapter from pp. 141-158. In Concept 7.1, pay special attention to oxidation & reduction and the
Oxidative Phosphorylation
 Electron Transport Chain (overview) The NADH and FADH 2, formed during glycolysis, β- oxidation and the TCA cycle, give up their electrons to reduce molecular O 2 to H 2 O. Electron transfer occurs through
Electron Transport Chain (overview) The NADH and FADH 2, formed during glycolysis, β- oxidation and the TCA cycle, give up their electrons to reduce molecular O 2 to H 2 O. Electron transfer occurs through
Cellular Respiration Harvesting Chemical Energy ATP
 Cellular Respiration Harvesting Chemical Energy ATP 2006-2007 What s the point? The point is to make ATP! ATP 2006-2007 Harvesting stored energy Energy is stored in organic molecules carbohydrates, fats,
Cellular Respiration Harvesting Chemical Energy ATP 2006-2007 What s the point? The point is to make ATP! ATP 2006-2007 Harvesting stored energy Energy is stored in organic molecules carbohydrates, fats,
Chapter 5: Major Metabolic Pathways
 Chapter 5: Major Metabolic Pathways David Shonnard Department of Chemical Engineering 1 Presentation Outline: Introduction to Metabolism Glucose Metabolism Glycolysis, Kreb s Cycle, Respiration Biosysthesis
Chapter 5: Major Metabolic Pathways David Shonnard Department of Chemical Engineering 1 Presentation Outline: Introduction to Metabolism Glucose Metabolism Glycolysis, Kreb s Cycle, Respiration Biosysthesis
CITRIC ACID CYCLE ERT106 BIOCHEMISTRY SEM /19 BY: MOHAMAD FAHRURRAZI TOMPANG
 CITRIC ACID CYCLE ERT106 BIOCHEMISTRY SEM 1 2018/19 BY: MOHAMAD FAHRURRAZI TOMPANG Chapter Outline (19-1) The central role of the citric acid cycle in metabolism (19-2) The overall pathway of the citric
CITRIC ACID CYCLE ERT106 BIOCHEMISTRY SEM 1 2018/19 BY: MOHAMAD FAHRURRAZI TOMPANG Chapter Outline (19-1) The central role of the citric acid cycle in metabolism (19-2) The overall pathway of the citric
Energy Production In A Cell (Chapter 25 Metabolism)
 Energy Production In A Cell (Chapter 25 Metabolism) Large food molecules contain a lot of potential energy in the form of chemical bonds but it requires a lot of work to liberate the energy. Cells need
Energy Production In A Cell (Chapter 25 Metabolism) Large food molecules contain a lot of potential energy in the form of chemical bonds but it requires a lot of work to liberate the energy. Cells need
SUPPLEMENTARY APPENDIX
 SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
Supplementary Figure 1. Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
 Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Metabolic engineering some basic considerations. Lecture 9
 Metabolic engineering some basic considerations Lecture 9 The 90ties: From fermentation to metabolic engineering Recruiting heterologous activities to perform directed genetic modifications of cell factories
Metabolic engineering some basic considerations Lecture 9 The 90ties: From fermentation to metabolic engineering Recruiting heterologous activities to perform directed genetic modifications of cell factories
MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question.
 Exam 3 BIOL 1406, Fall 2012 HCC Name MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) When biologists wish to study the internal ultrastructure
Exam 3 BIOL 1406, Fall 2012 HCC Name MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) When biologists wish to study the internal ultrastructure
Respiration. Respiration. Respiration. How Cells Harvest Energy. Chapter 7
 How Cells Harvest Energy Chapter 7 Organisms can be classified based on how they obtain energy: autotrophs: are able to produce their own organic molecules through photosynthesis heterotrophs: live on
How Cells Harvest Energy Chapter 7 Organisms can be classified based on how they obtain energy: autotrophs: are able to produce their own organic molecules through photosynthesis heterotrophs: live on
Supplementary. properties of. network types. randomly sampled. subsets (75%
 Supplementary Information Gene co-expression network analysis reveals common system-level prognostic genes across cancer types properties of Supplementary Figure 1 The robustness and overlap of prognostic
Supplementary Information Gene co-expression network analysis reveals common system-level prognostic genes across cancer types properties of Supplementary Figure 1 The robustness and overlap of prognostic
9-1 Chemical Pathways
 2 of 39 Food serves as a source of raw materials for the cells in the body and as a source of energy. Animal Cells Animal Mitochondrion Plant Plant Cells 3 of 39 1 Both plant and animal cells carry out
2 of 39 Food serves as a source of raw materials for the cells in the body and as a source of energy. Animal Cells Animal Mitochondrion Plant Plant Cells 3 of 39 1 Both plant and animal cells carry out
Lesson Overview. Cellular Respiration: An Overview. 9.2 process of cell respiration
 9.2 process of cell respiration Glycolysis During glycolysis, glucose is broken down into 2 molecules of the 3-carbon molecule pyruvic acid. Pyruvic acid is a reactant in the Krebs cycle. ATP and NADH
9.2 process of cell respiration Glycolysis During glycolysis, glucose is broken down into 2 molecules of the 3-carbon molecule pyruvic acid. Pyruvic acid is a reactant in the Krebs cycle. ATP and NADH
7 Cellular Respiration and Fermentation
 CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY REECE 7 Cellular Respiration and Fermentation Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge, Simon Fraser University SECOND EDITION
CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY REECE 7 Cellular Respiration and Fermentation Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge, Simon Fraser University SECOND EDITION
Growth. Principles of Metabolism. Principles of Metabolism 1/18/2011. The role of ATP energy currency. Adenosine triphosphate
 Metabolism: Fueling Cell Growth Principles of Metabolism Cells (including your own) must: Synthesize new components (anabolism/biosynthesis) Harvest energy and convert it to a usable form (catabolism)
Metabolism: Fueling Cell Growth Principles of Metabolism Cells (including your own) must: Synthesize new components (anabolism/biosynthesis) Harvest energy and convert it to a usable form (catabolism)
Introduction. Living is work. To perform their many tasks, cells must bring in energy from outside sources.
 Introduction Living is work. To perform their many tasks, cells must bring in energy from outside sources. In most ecosystems, energy enters as sunlight. Light energy trapped in organic molecules is available
Introduction Living is work. To perform their many tasks, cells must bring in energy from outside sources. In most ecosystems, energy enters as sunlight. Light energy trapped in organic molecules is available
NBCE Mock Board Questions Biochemistry
 1. Fluid mosaic describes. A. Tertiary structure of proteins B. Ribosomal subunits C. DNA structure D. Plasma membrane structure NBCE Mock Board Questions Biochemistry 2. Where in the cell does beta oxidation
1. Fluid mosaic describes. A. Tertiary structure of proteins B. Ribosomal subunits C. DNA structure D. Plasma membrane structure NBCE Mock Board Questions Biochemistry 2. Where in the cell does beta oxidation
Glycolysis. Cellular Respiration
 Glucose is the preferred carbohydrate of cells. In solution, it can change from a linear chain to a ring. Energy is stored in the bonds of the carbohydrates. Breaking these bonds releases that energy.
Glucose is the preferred carbohydrate of cells. In solution, it can change from a linear chain to a ring. Energy is stored in the bonds of the carbohydrates. Breaking these bonds releases that energy.
Biology. Slide 1 of 39. End Show. Copyright Pearson Prentice Hall
 Biology 1 of 39 2 of 39 9-1 Chemical Pathways Food serves as a source of raw materials for the cells in the body and as a source of energy. Animal Cells Animal Mitochondrion Plant Plant Cells 3 of 39 Both
Biology 1 of 39 2 of 39 9-1 Chemical Pathways Food serves as a source of raw materials for the cells in the body and as a source of energy. Animal Cells Animal Mitochondrion Plant Plant Cells 3 of 39 Both
Metabolism. Chapter 5. Catabolism Drives Anabolism 8/29/11. Complete Catabolism of Glucose
 8/29/11 Metabolism Chapter 5 All of the reactions in the body that require energy transfer. Can be divided into: Cell Respiration and Metabolism Anabolism: requires the input of energy to synthesize large
8/29/11 Metabolism Chapter 5 All of the reactions in the body that require energy transfer. Can be divided into: Cell Respiration and Metabolism Anabolism: requires the input of energy to synthesize large
Chapter 8 Mitochondria and Cellular Respiration
 Chapter 8 Mitochondria and Cellular Respiration Cellular respiration is the process of oxidizing food molecules, like glucose, to carbon dioxide and water. The energy released is trapped in the form of
Chapter 8 Mitochondria and Cellular Respiration Cellular respiration is the process of oxidizing food molecules, like glucose, to carbon dioxide and water. The energy released is trapped in the form of
Chapter 7 Cellular Respiration and Fermentation*
 Chapter 7 Cellular Respiration and Fermentation* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. Life Is Work
Chapter 7 Cellular Respiration and Fermentation* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. Life Is Work
Chapter 7: How Cells Harvest Energy AP
 Chapter 7: How Cells Harvest Energy AP Essential Knowledge 1.B.1 distributed among organisms today. (7.1) 1.D.2 Organisms share many conserved core processes and features that evolved and are widely Scientific
Chapter 7: How Cells Harvest Energy AP Essential Knowledge 1.B.1 distributed among organisms today. (7.1) 1.D.2 Organisms share many conserved core processes and features that evolved and are widely Scientific
Chapter 5 Microbial Metabolism: The Chemical Crossroads of Life
 Chapter 5 Microbial Metabolism: The Chemical Crossroads of Life Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. The Metabolism of Microbes metabolism all chemical
Chapter 5 Microbial Metabolism: The Chemical Crossroads of Life Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. The Metabolism of Microbes metabolism all chemical
Name Class Date. 1. Cellular respiration is the process by which the of "food"
 Name Class Date Cell Respiration Introduction Cellular respiration is the process by which the chemical energy of "food" molecules is released and partially captured in the form of ATP. Carbohydrates,
Name Class Date Cell Respiration Introduction Cellular respiration is the process by which the chemical energy of "food" molecules is released and partially captured in the form of ATP. Carbohydrates,
CLASS 11 th. Respiration in Plants
 CLASS 11 th 01. Introduction All living cells require continuous supply of energy to perform various vital activities. This energy is released in controlled manner for cellular use via the process of respiration.
CLASS 11 th 01. Introduction All living cells require continuous supply of energy to perform various vital activities. This energy is released in controlled manner for cellular use via the process of respiration.
RESPIRATION Worksheet
 A.P. Bio L.C. RESPIRATION Worksheet 1. In the conversion of glucose and oxygen to carbon dioxide and water a) which molecule becomes reduced? b) which molecule becomes oxidized? c) what happens to the
A.P. Bio L.C. RESPIRATION Worksheet 1. In the conversion of glucose and oxygen to carbon dioxide and water a) which molecule becomes reduced? b) which molecule becomes oxidized? c) what happens to the
g) Cellular Respiration Higher Human Biology
 g) Cellular Respiration Higher Human Biology What can you remember about respiration? 1. What is respiration? 2. What are the raw materials? 3. What are the products? 4. Where does it occur? 5. Why does
g) Cellular Respiration Higher Human Biology What can you remember about respiration? 1. What is respiration? 2. What are the raw materials? 3. What are the products? 4. Where does it occur? 5. Why does
BIO16 Mapua Institute of Technology
 BIO16 Mapua Institute of Technology The Marathon If somebody challenged you to a run a race, how should you prepare to win? 1. Practice 2. Eat the right foods 3. Drink the right liquids Energy All living
BIO16 Mapua Institute of Technology The Marathon If somebody challenged you to a run a race, how should you prepare to win? 1. Practice 2. Eat the right foods 3. Drink the right liquids Energy All living
Chapter 5 MITOCHONDRIA AND RESPIRATION 5-1
 Chapter 5 MITOCHONDRIA AND RESPIRATION All organisms must transform energy. This energy is required to maintain a dynamic steady state, homeostasis, and to insure continued survival. As will be discussed
Chapter 5 MITOCHONDRIA AND RESPIRATION All organisms must transform energy. This energy is required to maintain a dynamic steady state, homeostasis, and to insure continued survival. As will be discussed
General Biology 1004 Chapter 6 Lecture Handout, Summer 2005 Dr. Frisby
 Slide 1 CHAPTER 6 Cellular Respiration: Harvesting Chemical Energy PowerPoint Lecture Slides for Essential Biology, Second Edition & Essential Biology with Physiology Presentation prepared by Chris C.
Slide 1 CHAPTER 6 Cellular Respiration: Harvesting Chemical Energy PowerPoint Lecture Slides for Essential Biology, Second Edition & Essential Biology with Physiology Presentation prepared by Chris C.
9-1 Cellular Respiration Slide 1 of 39
 9-1 Cellular Respiration 1 of 39 Learning Targets TN State Standards CLE 3210.3.2 Distinguish between aerobic and anaerobic respiration. CLE 3216.3.3 Describe how mitochondria make stored chemical energy
9-1 Cellular Respiration 1 of 39 Learning Targets TN State Standards CLE 3210.3.2 Distinguish between aerobic and anaerobic respiration. CLE 3216.3.3 Describe how mitochondria make stored chemical energy
Cellular Respiration: Harvesting Chemical Energy
 Chapter 9 Cellular Respiration: Harvesting Chemical Energy You should be able to: 1. Explain how redox reactions are involved in energy exchanges. Name and describe the three stages of cellular respiration;
Chapter 9 Cellular Respiration: Harvesting Chemical Energy You should be able to: 1. Explain how redox reactions are involved in energy exchanges. Name and describe the three stages of cellular respiration;
What s the point? The point is to make ATP! ATP
 ATP Chapter 8 What s the point? The point is to make ATP! ATP Flows into an ecosystem as sunlight and leaves as heat Energy is stored in organic compounds Carbohydrates, lipids, proteins Heterotrophs eat
ATP Chapter 8 What s the point? The point is to make ATP! ATP Flows into an ecosystem as sunlight and leaves as heat Energy is stored in organic compounds Carbohydrates, lipids, proteins Heterotrophs eat
Unit 2: Metabolic Processes
 How is energy obtained biologically? Recall: Red Ox Reactions Unit 2: Metabolic Processes Oxidation Is the chief mechanism by which chemical potential energy is released This energy comes from reduced
How is energy obtained biologically? Recall: Red Ox Reactions Unit 2: Metabolic Processes Oxidation Is the chief mechanism by which chemical potential energy is released This energy comes from reduced
Table S1: Kinetic parameters of drug and substrate binding to wild type and HIV-1 protease variants. Data adapted from Ref. 6 in main text.
 Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
How Cells Release Chemical Energy. Chapter 7
 How Cells Release Chemical Energy Chapter 7 7.1 Overview of Carbohydrate Breakdown Pathways All organisms (including photoautotrophs) convert chemical energy of organic compounds to chemical energy of
How Cells Release Chemical Energy Chapter 7 7.1 Overview of Carbohydrate Breakdown Pathways All organisms (including photoautotrophs) convert chemical energy of organic compounds to chemical energy of
Background knowledge
 Background knowledge This is the required background knowledge: State three uses of energy in living things Give an example of an energy conversion in a living organism State that fats and oils contain
Background knowledge This is the required background knowledge: State three uses of energy in living things Give an example of an energy conversion in a living organism State that fats and oils contain
2) The molecule that functions as the reducing agent (electron donor) in a redox or oxidationreduction
 Campbell Biology in Focus (Urry) Chapter 7 Cellular Respiration and Fermentation 7.1 Multiple-Choice Questions 1) What is the term for metabolic pathways that release stored energy by breaking down complex
Campbell Biology in Focus (Urry) Chapter 7 Cellular Respiration and Fermentation 7.1 Multiple-Choice Questions 1) What is the term for metabolic pathways that release stored energy by breaking down complex
7/5/2014. Microbial. Metabolism. Basic Chemical Reactions Underlying. Metabolism. Metabolism: Overview
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University Basic Chemical Reactions Underlying Metabolism Metabolism C H A P T E R 5 Microbial Metabolism Collection
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University Basic Chemical Reactions Underlying Metabolism Metabolism C H A P T E R 5 Microbial Metabolism Collection
Section 1 Proteins and Proteomics
 Section 1 Proteins and Proteomics Learning Objectives At the end of this assignment, you should be able to: 1. Draw the chemical structure of an amino acid and small peptide. 2. Describe the difference
Section 1 Proteins and Proteomics Learning Objectives At the end of this assignment, you should be able to: 1. Draw the chemical structure of an amino acid and small peptide. 2. Describe the difference
How Cells Harvest Chemical Energy
 How Cells Harvest Chemical Energy Global Athlete Outreach Program US CytoThesis Systems Medicine Center www.cytothesis.us US OncoTherapy Systems BioMedicine Group CytoThesis Bioengineering Research Group
How Cells Harvest Chemical Energy Global Athlete Outreach Program US CytoThesis Systems Medicine Center www.cytothesis.us US OncoTherapy Systems BioMedicine Group CytoThesis Bioengineering Research Group
What is Glycolysis? Breaking down glucose: glyco lysis (splitting sugar)
 What is Glycolysis? Breaking down glucose: glyco lysis (splitting sugar) Most ancient form of energy capture. Starting point for all cellular respiration. Inefficient: generates only 2 ATP for every 1
What is Glycolysis? Breaking down glucose: glyco lysis (splitting sugar) Most ancient form of energy capture. Starting point for all cellular respiration. Inefficient: generates only 2 ATP for every 1
Inferring Biological Meaning from Cap Analysis Gene Expression Data
 Inferring Biological Meaning from Cap Analysis Gene Expression Data HRYSOULA PAPADAKIS 1. Introduction This project is inspired by the recent development of the Cap analysis gene expression (CAGE) method,
Inferring Biological Meaning from Cap Analysis Gene Expression Data HRYSOULA PAPADAKIS 1. Introduction This project is inspired by the recent development of the Cap analysis gene expression (CAGE) method,
What s the point? The point is to make ATP! ATP
 2006-2007 What s the point? The point is to make ATP! ATP Glycolysis 2 ATP Kreb s cycle 2 ATP Life takes a lot of energy to run, need to extract more energy than 4 ATP! There s got to be a better way!
2006-2007 What s the point? The point is to make ATP! ATP Glycolysis 2 ATP Kreb s cycle 2 ATP Life takes a lot of energy to run, need to extract more energy than 4 ATP! There s got to be a better way!
Table of Contents. Section 1 Glycolysis and Fermentation. Section 2 Aerobic Respiration
 Table of Contents Section 1 Glycolysis and Fermentation Section 2 Aerobic Respiration Objectives Identify the two major steps of cellular respiration. Describe the major events in glycolysis. Compare lactic
Table of Contents Section 1 Glycolysis and Fermentation Section 2 Aerobic Respiration Objectives Identify the two major steps of cellular respiration. Describe the major events in glycolysis. Compare lactic
chemical compounds
 chemical compounds Adenine 3 Phosphate groups Ribose The three phosphate groups are the key to ATP's ability to store and release energy. Storing Energy ADP has two (di) phosphate groups instead of three.
chemical compounds Adenine 3 Phosphate groups Ribose The three phosphate groups are the key to ATP's ability to store and release energy. Storing Energy ADP has two (di) phosphate groups instead of three.
2. Cellular respiration uses oxygen to convert the chemical energy stored in organic molecules into -?-
 HB Cell Respiration Questions (1/2 point each question or blank to fill in 37 points) 1. Organisms, such as plants that make their own food are called -?- 2. Cellular respiration uses oxygen to convert
HB Cell Respiration Questions (1/2 point each question or blank to fill in 37 points) 1. Organisms, such as plants that make their own food are called -?- 2. Cellular respiration uses oxygen to convert
Cellular Respiration: Harvesting Chemical Energy Chapter 9
 Cellular Respiration: Harvesting Chemical Energy Chapter 9 Assemble polymers, pump substances across membranes, move and reproduce The giant panda Obtains energy for its cells by eating plants which get
Cellular Respiration: Harvesting Chemical Energy Chapter 9 Assemble polymers, pump substances across membranes, move and reproduce The giant panda Obtains energy for its cells by eating plants which get
Digitizing the Proteomes From Big Tissue Biobanks
 Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Biology 30 Structure & Function of Cells (Part 2) Bioenergetics: Energy: Potential energy: Examples: Kinetic energy. Examples:
 Biology 30 Structure & Function of Cells (Part 2) Bioenergetics: Energy: Potential energy: Examples: Kinetic energy Examples: Energy can be transformed: Thermodynamics: First law of Thermodynamics: Second
Biology 30 Structure & Function of Cells (Part 2) Bioenergetics: Energy: Potential energy: Examples: Kinetic energy Examples: Energy can be transformed: Thermodynamics: First law of Thermodynamics: Second
