Membrane-Mediated Induction and Sorting of K-Ras Microdomain Signaling Platforms
|
|
|
- Abel Golden
- 6 years ago
- Views:
Transcription
1 pubs.acs.org/jacs Membrane-Mediated Induction and Sorting of K-Ras Microdomain Signaling Platforms Katrin Weise, Shobhna Kapoor, Christian Denter, J org Nikolaus, Norbert Opitz, Sebastian Koch,^,# Gemma Triola,^,# Andreas Herrmann, Herbert Waldmann,^,# and Roland Winter*, Physical Chemistry I, Biophysical Chemistry, Faculty of Chemistry, TU Dortmund University, Otto-Hahn-Strasse 6, D Dortmund, Germany Institute of Biology/Biophysics, Humboldt University Berlin, Invalidenstrasse 42, D Berlin, Germany ^Chemical Max Planck Institute of Molecular Physiology, Otto-Hahn-Strasse 11, D Dortmund, Germany Biology, Max Planck Institute of Molecular Physiology, Otto-Hahn-Strasse 11, D Dortmund # Faculty of Chemistry, TU Dortmund University, Otto-Hahn-Strasse 6, D Dortmund, Germany bs Supporting Information ABSTRACT: The K-Ras4B GTPase is a major oncoprotein whose signaling activity depends on its correct localization to negatively charged subcellular membranes and nanoclustering in membrane microdomains. Selective localization and clustering are mediated by the polybasic farnesylated C-terminus of K-Ras4B, but the mechanisms and molecular determinants involved are largely unknown. In a combined chemical biological and biophysical approach we investigated the partitioning of semisynthetic fully functional lipidated K-Ras4B proteins into heterogeneous anionic model membranes and membranes composed of viral lipid extracts. Independent of GDP/GTP-loading, K-Ras4B is preferentially localized in liquid-disordered (l d ) lipid domains and forms new protein-containing fluid domains that are recruiting multivalent acidic lipids by an effective, electrostatic lipid sorting mechanism. In addition, GDP-GTP exchange and, thereby, Ras activation results in a higher concentration of activated K-Ras4B in the nanoscale signaling platforms. Conversely, palmitoylated and farnesylated N-Ras proteins partition into the l d phase and concentrate at the l d /l o phase boundary of heterogeneous membranes. Next to the lipid anchor system, the results reveal an involvement of the G-domain in the membrane interaction process by determining minor but yet significant structural reorientations of the GDP/GTP-K-Ras4B proteins at lipid interfaces. A molecular mechanism for isoform-specific Ras signaling from separate membrane microdomains is postulated from the results of this study. INTRODUCTION Ras proteins are small GTPases that act as molecular switches between a GDP-bound inactive and a GTP-bound active state. 1-3 Constitutively activated K-Ras4B contributes to the initiation and maintenance of many tumor cell types, including those of the pancreas, colon, and lung. 4 A prerequisite for the regulation of signal transmission from cell surface receptors to intracellular signaling cascades including the oncogenic potential of Ras proteins is the anchorage of Ras in the cytosolic leaflet of cellular membranes. 5-7 The cytosolic face of the plasma membrane, endosomes, and lysosomes of mammalian cells is enriched in acidic phospholipids, such as phosphatidylserine (PS) as well as phosphatidylinositol (PI) and polyphosphoinositides (PPI), due to an asymmetric distribution of neutral and anionic phospholipids in the two bilayer leaflets of membranes. 8,9 The highly conserved N-terminal G-domain of Ras encompasses all interaction sites for activators and effectors. Conversely, the plasma membrane localization of Ras has been determined to be directed by the Ras isoform specific, posttranslationally modified C-terminal sequence 6,7,10 that constitutes together with the linker domain the hypervariable region (HVR). For the 4B splice variant of K-Ras (K-Ras4B), this membrane interacting and targeting signal consists of a C-terminal polybasic stretch of six contiguous lysines in a total of eight that immediately precedes the terminal farnesylated and carboxymethylated cysteine residue (Figure 1). 2,7,11 Two models have been proposed for the selective targeting of K-Ras4B to the cytosolic face of the plasma membrane: One involves a nonspecific electrostatic interaction of the lipidated polybasic domain with anionic phospholipids in the plasma membrane, 7,9,12,13 the other involves binding of K-Ras4B to a specific membrane-bound protein receptor. 14 Furthermore, a possible involvement of microtubules in the directing process has Received: August 20, 2010 Published: December 9, 2010 r 2010 American Chemical Society 880 dx.doi.org/ /ja107532q J. Am. Chem. Soc. 2011, 133,
2 Figure 1. Schematic representation of the semisynthetic K-Ras4B protein. The synthesized, lipidated K-Ras4B peptide was ligated to a truncated K-Ras4B protein expressed in E.coli to yield the S-farnesylated K-Ras4B protein bearing an additional cysteine between Gly 174 and Lys 175. The structure of the G-domain was adopted from the pdb (3GFT) to highlight the switch I region (residues 32-38; blue), switch II region (59-67; yellow), and R5-helix ( ; red). For the fluorescence experiments, the protein core is labeled with BODIPY. been suggested. 15 It has also been shown recently that K-Ras4B can be redirected from one target membrane to another depending on the surface charge density of the membrane and the net charge of the protein. 8 Accordingly, an increase in cytosolic calcium will result in a decrease of the most-negative plasma membrane s surface charge, rendering it comparable to the intermediate negative charge density of endomembranes, which now compete effectively for binding of K-Ras4B. 8 On the other hand, a reduction of the positive net charge of the polybasic domain by phosphorylation of serine 181 within the HVR of K-Ras4B leads to an association not only with the plasma membrane, but also to a substantial degree with endocytic membranes. 9,16,17 Hence, the intracellular transport of K-Ras4B between subcellular compartments may be driven by electrostatic interactions, with the polybasic domain acting as a probe for the electronegative surface potential of the membrane. 18,19 Recent experimental work indicates that signaling of K-Ras4B is restricted to particular plasma membrane microdomains in detecting a clustering of GTP- and GDP-loaded K-Ras4B into cholesterol-independent and actin-dependent but spatially distinct nanodomains This K-Ras4B nanocluster assembly is thought to lead to the formation of a local environment highly enriched in acidic phospholipids, thus enabling a localized highaffinity binding domain for effector proteins. However, the mechanism underlying K-Ras4B nanoclustering is currently unknown. To yield a pictorial view of the localization process of K-Ras4B in heterogeneous membranes up to a single molecule level, high-resolution imaging using confocal laser scanning and tapping-mode atomic force microscopy has been used in the present study. Semisynthetic fully functional lipidated, GDPloaded and GTP-loaded (GppNHp-bound state as a nonhydrolyzable GTP analog) K-Ras4B proteins were synthesized, 23 and their partitioning into a newly designed, heterogeneous anionic model membrane system as well as a membrane system that mimics the lipid composition of biological membranes was investigated. The results lead to the postulation of a molecular mechanism for isoform-specific Ras signaling from separate membrane microdomains. Furthermore, time-dependent FTIR experiments reveal changes in G-domain orientation and strength of membrane interaction upon GDP-GTP exchange and thereby Ras activation. RESULTS Establishment of a Heterogeneous Anionic Model Membrane System. Up to now, ternary lipid mixtures such as DOPC/DPPC/Chol have been widely used in studies of the interaction of lipidated proteins, such as N-Ras, with zwitterionic model raft membranes. 24 Since it is known that the positively charged lysines in the K-Ras4B membrane anchor region can strongly interact with negatively charged lipids, a heterogeneous anionic model membrane system needed to be established. Previous reports revealed no specific binding of the polybasic K-Ras4B domain to any of the common mono- and polyanionic phospholipids of the inner plasma membrane. 19 Here we used phosphatidylglycerol (PG) as an established model for anionic phospholipids. Calorimetry, FTIR spectroscopy, confocal fluorescence microscopy, and AFM data revealed that incorporation of 10 to 20 mol % PG (corresponding to a DOPC/DOPG/ DPPC/DPPG/Chol lipid bilayer with a molar ratio of 20:5: 45:5:25 and 15:10:40:10:25, respectively) still leads to segregation of the lipid mixture into liquid-ordered (l o ) and liquiddisordered (l d ) domains at room temperature (data not shown). The corresponding AFM analysis shows one homogeneous lipid bilayer on mica with isolated islands of fluid (l d ) domains in a coherent pool of raft-like (l o ) protruding phase (Supporting Information, Figure S1). 23 Interaction of K-Ras4B with GUVs Containing Charged Domains. In a first set of experiments, the partitioning of GDPand GTP-loaded, farnesylated and BODIPY-labeled K-Ras4B into GUVs composed of DOPC/DOPG/DPPC/DPPG/Chol 15:10:40:10:25 was analyzed by confocal fluorescence microscopy at room temperature. In confocal fluorescence microscopy, phases can be identified as ordered or fluidlike disordered on the basis of the known partitioning behavior of appropriate fluorescence probes, such as N-Rh-DHPE. Figure 2 shows the GUV images taken after the binding process of K-Ras4B to the lipid vesicles was completed (t 2 h). Panels a and c exhibit the round-shaped domains typical for coexisting liquid-like phases. 25,26 The bright red areas with embedded, homogeneously distributed N-Rh-DHPE correspond to l d domains, whereas the dark domains consist mainly of l o phase. By simultaneous inspection of the rhodamine (red) and BODIPY (green) channel, where the latter identifies the localization of the BODIPY-labeled protein within the lipid assembly (panels b and d), one can directly follow the incorporation of the protein into the lipid microdomains. Superposition of both channels clearly shows that both GDPand GTP-loaded K-Ras4B are essentially localized in the l d phase of the lipid bilayer system (the respective images coincide in both cases). Quantification of the protein partitioning can be estimated by determining the background-corrected fluorescence intensity ratio of l o versus l d phases (cf. Materials and Methods). An average fluorescence intensity ratio of 0.18 ( 0.05 and 0.11 ( 0.03 has been determined for the inactive and active K-Ras4B, respectively, thus confirming the preferred l d phase partitioning 881 dx.doi.org/ /ja107532q J. Am. Chem. Soc. 2011, 133,
3 Journal of the American Chemical Society Figure 2. Confocal laser scanning microscopy images of the partitioning of K-Ras4B GDP (a,b) and K-Ras4B GTP (c,d) into GUVs. GUVs were composed of DOPC/DOPG/DPPC/DPPG/Chol 15:10:40:10:25, that segregate into lo and ld domains at room temperature. N-Rh-DHPE was used as membrane marker, labeling preferentially the ld domains (red channel, a and c); the K-Ras4B proteins were BODIPY-labeled (green channel, b and d). The overlay of the red and green channel shows that both K-Ras4B GDP and K-Ras4B GTP insert preferentially into the ld lipid phase, which is confirmed by the determined fluorescence intensity ratio of 0.20 for image b and 0.14 for image d, respectively. The scale bar represents 10 μm. of both K-Ras4B proteins in showing values (82 to 89%) similar to those of ld phase fluorescent markers. AFM Study of K-Ras4B Interaction with Phase-Separated Anionic Membranes. To yield complementary structural data on a smaller, nanometer length scale, the interaction of the GDPand GTP-loaded, farnesylated K-Ras4B with lipid domains of the negatively charged heterogeneous membrane was followed by AFM. A sample consisting of one lipid bilayer with coexisting lo and ld domains on mica was examined at the beginning of each time-lapse tapping-mode AFM experiment (Figure 3, a and d). Using the technique of direct injection of a solution containing K-Ras4B protein into the AFM fluid cell, the same region of the lipid bilayer can be observed before and after incorporation of the protein. The incubation of the lipid bilayer with both active and inactive K-Ras4B did not influence the lateral membrane organization initially, i.e., coexisting ld and lo domains of similar shape and size can still be distinguished, and the height difference between both phases remains 1.0 nm (Figure 4). The data also show that the inactive and active K-Ras4B are readily incorporated (on a minute time scale) and exclusively located in the bulk ld phase upon binding to the lipid bilayer (Figure 3 panels b and e, respectively). A quantitative AFM analysis reveals a mean height of 2.0 ( 0.7 nm (mean value ( s.d., n = 304) for K-Ras4B GDP, and a similar value of 2.5 ( 0.4 nm (n = 296) was determined for the GTP-loaded K-Ras4B (Figure 4). These dimensions correspond roughly to the diameter of a Ras molecule.27 Whereas N-Ras proteins bearing at least one farnesyl anchor showed a Figure 3. AFM images of the time-dependent partitioning of GDP- and GTP-loaded K-Ras4B into lipid bilayers consisting of DOPC/DOPG/ DPPC/DPPG/Chol 20:5:45:5:25. The AFM images are shown before (a and d) and at selected time points (t 6 and 5 h, b and e, respectively; t 24 h, c; and t 20.5 h, f) after injection of 200 μl of K-Ras4B in 20 mm Tris, 7 mm MgCl2, ph 7.4 (ck-ras4b = 2 μm) into the AFM fluid cell. The overall height of the vertical color scale from dark brown to white corresponds to 6 nm for all images. The scale bar represents 2 μm. time dependent diffusion into and subsequent clustering in the lo/ld phase boundary region,24,28 formation of new domains with accumulated protein inside a fluidlike environment was observed for both K-Ras4B proteins after membrane insertion (Figures 3 and 4). These new domains with enriched GDP- and GTPloaded K-Ras4B exhibit an average lateral dimension of 1.5 μm or less and show a thickness which is 0.9 ( 0.1 nm (n = 119) and 1.8 ( 0.2 nm (n = 89) larger than that of the purely fluid lipid phase for the GDP and GTP bound state, respectively. Both the lateral dimensions of the protein clusters and the domain thicknesses critically depend on the protein concentration and preexisting lateral organization of the membrane. Whereas the lateral dimensions of the protein domains enriched in GDPand GTP-loaded K-Ras4B are similar, the protein clusters of active and inactive K-Ras4B can be clearly distinguished by the different domain thicknesses in all cases. This indicates that the clustering effect is even more pronounced for the active K-Ras4B since the thicker domains argue for a stronger enrichment in protein (Figure 4). The increased domain thickness can be 882 dx.doi.org/ /ja107532q J. Am. Chem. Soc. 2011, 133,
4 Figure 4. Analysis of the AFM images of GDP- and GTP-loaded K-Ras4B. Zoom of the lower and middle left area of the AFM images at t 6 and 5 h (panels b and e, respectively, in Figure 3) and analysis of the corresponding AFM images for K-Ras4B GDP (left) and K-Ras4B GTP (right) (c K-Ras4B =2μM). In the upper part, the section profiles of the AFM images are shown with a vertical color scale from dark brown to white corresponding to an overall height of 5 nm. The horizontal black line is the localization of the section analysis given at the bottom, indicating the vertical distances between pairs of arrows (black, l o /l d phase difference; green, thickness difference of the new domains enriched in K-Ras4B compared to l d domains; and red, size of the K-Ras4B proteins). explained by a tighter packing of the Ras molecules via a different orientation of the protein with respect to the membrane interface. Interestingly, these newly formed domains are stable over the whole time range covered, whereas the lipids in the bulk l d phase still exhibit a high lateral mobility (Figure 3, panels c and f). Structural Aspects of the K-Ras4B Membrane Interaction. To reveal possible changes in the structure of K-Ras4B during the membrane interaction process, time-dependent FTIR spectroscopic measurements have been carried out. Structural changes of K-Ras4B in the presence of large unilamellar vesicles (LUVs) composed of DOPC/DOPG/DPPC/DPPG/Chol 20:5:45:5:25 were followed as a function of time at 25 C, with spectra collected every 10 min. The individual secondary structural elements contributing to the amide-i 0 band have been determined by using the local minima of the concomitant second derivative spectra and the local maxima of its Fourier self-deconvolution (FSD; Figure 5 and Supporting Information, Figure S2). Figure 5 shows the FSD of the time evolution of the normalized FTIR spectra of the amide-i 0 band (a and c) and the concomitant second derivative spectra (b and d) for the GDP- and GTP-loaded K-Ras4B, respectively. At the beginning of the membrane interaction process (t = 1.5 min), subbands at about 1673, 1661, 1648, and 1637 cm -1 and a shoulder at 1626 cm -1 can be detected for the inactive K-Ras4B GDP. With time, a shift to lower wavenumbers was observed for the overall spectra that is accompanied by a small change in spectral shape (panel a), suggesting some minor but significant reorientational changes upon membrane binding. The analysis of the second derivative spectra (panel b) revealed a decrease in the band intensity at 1661 cm -1 with time, which is accompanied by a concomitant increase in the band intensity at 1654/1656 cm -1 that is indicative for R-helical structures. The comparison with an equivalent measurement of K-Ras4B GDP in bulk solution (20 mm Tris, 7 mm MgCl 2,pD 7.4; Supporting Information, Figure S3), that is, in the absence of membrane, that exhibits a corresponding subband for R-helical structures at about 1658 cm -1, indicates a marked interaction of the respective R-helical region of the protein with the lipid bilayer interface (shift to lower wavenumbers). Furthermore, a gradual shift to 1671 cm -1 was detected with time for the subband at 1673 cm -1 that is attributed to turns and loops, which again would argue for a stronger interaction of these structure elements with the membrane. 29 The analysis of the amide-i 0 band of K-Ras4B GDP in bulk solution yields a secondary structure content of 24% R-helix, 39% β-sheet, 15% turns and loops, and 21% random coil, in reasonable agreement with literature data (Supporting Information, Figure S3). 30 Minor differences may be due to different transition dipole moments of the various conformers and the absence of the HVR in the NMR and X-ray data. The corresponding fits of the time-dependent FTIR spectra of Ras residing in the membrane revealed, as expected, no significant changes in secondary structure of K-Ras4B GDP within the experimental error over the whole time range covered (Supporting Information, Figure S2c); that is, the structure of K-Ras4B GDP is not drastically affected by the membrane interaction, only minor reorientational changes seem to occur upon membrane binding. The same holds true for the active, GTP-loaded K-Ras4B. Similar to the K-Ras4B GDP behavior, a shift of the overall spectra to lower wavenumbers was observed with time, which is accompanied by small changes in spectral shape (Figure 5c), pointing toward minor but significant structural reorientations of the protein upon membrane insertion. Subbands at about 1672, 1662, 1651, and 1637 cm -1 and a shoulder at 1626 cm -1 can be detected for K-Ras4B GTP at the beginning of the membrane interaction process (t = 1.5 min), that stay constant over the whole time range covered (panel d). However, the intensity of the band at 1672 cm -1 decreases with time, which is accompanied by a corresponding increase at 1662 cm -1. Similar to the results for the inactive K-Ras4B, no marked changes in secondary structure could be detected within the experimental error by the fits of the time-dependent FTIR spectra for K-Ras4B GTP (Supporting Information, Figure S2f), again suggesting minor orientational rearrangements, only. The absence of the shift of the R-helical subband at 1662 cm -1 upon membrane binding might imply a higher reorientational flexibility of the helices of K-Ras4B GTP as compared to K-Ras4B GDP, and hence reveals minor intermolecular interactions of the K-Ras4B in its membrane-bound form upon nucleotide exchange. Orientational changes of the protein can also be revealed by following the insertion of K-Ras4B into lipid monolayers composed of DOPC/DOPG/DPPC/DPPG/Chol with infrared reflection absorption (IRRA) spectroscopy. Corresponding IRRA spectra of the amide-i 0 region of K-Ras4B GDP reveal a band maximum within cm -1 over the whole time course of the membrane interaction process, indicating a rather stable and relatively fixed orientation of the protein at the lipid interface during protein adsorption (Supporting Information, Figure S4a). In contrast, a gradual shift in wavenumber from 1638 to 1647 cm -1 could be observed for K-Ras4B GTP with time (Figure S4c), suggesting a reorientation of the active protein at the lipid interface during membrane insertion. When compared with simulations of the IRRA spectra performed for the Ras G-domain, 31 it can be concluded that K-Ras4B GTP achieves on 883 dx.doi.org/ /ja107532q J. Am. Chem. Soc. 2011, 133,
5 Figure 5. FTIR spectroscopic analysis of the GDP- and GTP-loaded K-Ras4B membrane interaction. Panels a and c show the normalized FSD FTIR spectra for the time evolution of the amide-i 0 band of K-Ras4B GDP (a) and K-Ras4B GTP (c) in the presence of DOPC/DOPG/DPPC/DPPG/Chol 20:5:45:5:25 at 25 C. The corresponding second derivative spectra are given in panels b and d. Arrows indicate the spectral shifts (in b) as well as the increasing (b, at 1654/1656 cm -1 ; d, at 1662 cm -1 ) and decreasing (b, at 1661 cm -1 ; d, at 1672 cm -1 ) intensities as a function of time. average an orientation that is compatible with an essentially random orientation with respect to the lipid interface, suggesting a higher reorientational flexibility for the active K-Ras protein, which would be in agreement with the transmission FTIR experiments. In IRRAS, generally the combined effects of a change in orientation along with a change in secondary structure (including changes in the strength of interaction) are detected. Since there are no secondary structural changes for the protein in the presence of the membrane (as evident from the transmission FTIR experiments), the shift observed for K-Ras4B GTP in the IRRA spectra can be assigned to a reorientation of the protein at the membrane interface. Moreover, the amide-i 0 band for the inactive K-Ras4B appears at lower wavenumbers compared with K-Ras4B GTP at saturation pressures (Figure S4), suggesting a different orientation and stronger interaction with the membrane interface. Interaction of K-Ras4B with Membranes Composed of Viral Lipid Extracts. To evaluate the biological relevance of the membrane partitioning process observed in the heterogeneous anionic model membrane system, corresponding studies with lipid bilayers made from the envelope membrane of influenza viruses (strain A/Japan/305/57) have been carried out. As influenza virus is assembled at and is budding from the plasma membrane of host cells, the envelope mimics essentially the lipid composition of the plasma membranes with some preference for raft lipids. 32 Confocal fluorescence microscopy experiments of GUVs made of this biological mixture indicated the coexistence of raft-like l o and fluid l d domains, with R18 used as a fluorescent marker for the l d phase. Figure 6 shows that both active and inactive K-Ras4B are mainly localized in the fluid l d domains of the viral membrane. Quantification of the protein partitioning yielded average fluorescence intensity ratios of 0.39 ( 0.12 and 0.34 ( 0.13 for the inactive and active K-Ras4B, respectively. Since R18 showed a mean fluorescence intensity ratio of 0.18 ( 0.09 (i.e., 82% l d ), phase partitioning of both K-Ras4B proteins can be essentially assigned to the fluid l d phase (g61%). Whereas for N-Ras a significant incorporation of the protein in the l o /l d phase boundaries of the viral membrane was revealed by fluorescence intensity profiles, 28 no such effect could be detected for K-Ras4B even after incubation times up to 48 h (cf. Figure 7). DISCUSSION AND CONCLUSIONS The Ras oncoproteins most frequently found in human tumors are activated K-Ras and N-Ras. Hence, the Ras isoform specificdifferences in signaling and lateral segregation to specific nanoscale compartments on the plasma membrane are issues under high scrutiny in the fields of biochemistry and biophysics. In a recent comparable approach, we were already able to demonstrate that palmitoylated and farnesylated N-Ras undergoes a time-dependent diffusion and subsequent clustering in the l o /l d phase boundary region of the heterogeneous model and viral membranes, independent of GDP/GTP-loading (cf. Figure7). 24,28,33 A monofarnesylated N-Ras that resembles the depalmitoylated 884 dx.doi.org/ /ja107532q J. Am. Chem. Soc. 2011, 133,
6 Figure 6. Confocal laser scanning microscopy images (equatorial plane, t 12 h) of the partitioning of K-Ras4B GDP (a, b) and K-Ras4B GTP (c, d) into GUVs from viral membranes. GUVs were prepared from lipid extracts of influenza virus (strain A/Japan/305/57) and incubated with BODIPY-labeled K-Ras4B protein for up to 48 h at 20 C. Both BODIPY-labeled K-Ras4B proteins (green channel, b and d) are colocalized with R18 (red channel, a and c) that is enriched in fluid l d phase, indicating a preferential partitioning of K-Ras4B into l d domains of influenza virus lipid-derived membranes. The determined fluorescence intensity ratios are 0.13 (R18), 0.30 (K-Ras4B GDP), 0.09 (R18), and 0.28 (K-Ras4B GTP) for images a, b, c, and d, respectively. The scale bar represents 5 μm. form of the natural N-Ras in the course of the acylation/deacylation cycle 10,34-36 also partitions into the l d phase and concentrates in the l o /l d domain interface. 24 Similar mechanistic studies on the lateral segregation of signaling proteins bearing a farnesyl anchor and a polybasic amino acid stretch, such as K- Ras4B, were missing until now. In contrast to N-Ras, spontaneous formation of new domains with accumulated protein residing within the bulk fluid phase of an anionic model raft membrane has been observed for GDP- and GTP-loaded K-Ras4B in the present study. Thereby, the bulky and branched farnesyl anchor prevents partitioning of K-Ras4B into highly ordered raft-like domains and accommodates more easily into disordered lipid domains. Such sorting effect has also been observed for other bulky and branched lipid anchors such as the tocopherol moiety. 37 A similar membrane partitioning behavior of K-Ras4B as in anionic model membranes was also observed in influenza virus lipid-derived membranes mimicking the composition of mammalian plasma membranes. To further address the question whether the sorting of K-Ras4B is solely goverend by electrostatic interactions with the anionic lipids in the membrane, the partitioning of K-Ras4B was also studied in zwitterionic heterogeneous membranes (DOPC/DPPC/Chol 25:50:25). Rather unexpectedly, a similar membrane partitioning behavior was detected in neutral and negatively charged membranes for both, active and inactive K-Ras4B, with even similar dimensions for the protein enriched domains (Supporting Information, Figures S5 and S6). Hence, electrostatic interactions between the polybasic domain of K-Ras4B Figure 7. Schematic model for N- and K-Ras4B localization in vesicular heterogeneous model biomembranes with liquid-disordered (l d ) and liquid-ordered (l o ) domains (left). Please note that the schematic view is not to scale. Whereas N-Ras partitions into the fluid-like phase of the membrane and subsequently diffuses to the l d /l o phase boundaries of the subcompartments as visualized by fluorescence microscopy and AFM (top right, with the clustered proteins depicted in blue-white in the 3D AFM images), K-Ras4B forms new protein-enriched fluid domains within the bulk fluid phase upon membrane binding and is thought to recruit multivalent anionic lipids by an effective, electrostatic lipid sorting mechanism (bottom). The GDP to GTP nucleotide exchange leads to changes in G-domain orientation and a stronger enrichment in protein in the signaling platform. and anionic lipids do not seem to be essential for the formation of new protein enriched domains in the fluid phase of the membrane. However, this sorting effect of K-Ras4B is nevertheless controlled by electrostatic interactions between the positively charged lysines and the negatively charged phosphate of the lipid head groups since the monofarnesylated and noncharged N-Ras revealed a clustering at the lipid domain boundaries under similar conditions. 24 It has in fact been observed that the polybasic stretch of K-Ras4B is able to induce a local positive electrostatic potential that attracts multivalent acidic lipids. 38 Such scenario would also be in agreement with a model proposed by McLaughlin and co-workers that suggested sequestration of polyvalent (PIP 2 and PIP 3 ) but not monovalent acidic lipids via nonspecific interactions for membrane-bound basic peptides. 38,39 Recent reports also demonstrate the requirement of highly charged polyphosphoinositides for the membrane interaction of K-Ras4B in vivo. 40 Overall, these results provide a molecular mechanism for isoform-specific Ras signaling from separate membrane microdomains (Figure 7). For N-Ras, the expulsion of the protein to the interfacial region combined with a minimization of the line 885 dx.doi.org/ /ja107532q J. Am. Chem. Soc. 2011, 133,
7 energy between neighboring domains is likely to be one of the key parameters controlling the size and dynamic properties of its signaling platform (Figure 7, top). Conversely, for K-Ras4B, electrostatic interactions seem to control the membrane interaction process. As a consequence, polyvalent acidic lipids can be recruited by an effective lipid sorting mechanism and new fluid domains with higher anionic charge density and incorporated K-Ras4B protein are formed (Figure 7, bottom). In fact, the free energy of binding increases by about 1 kcal/mol per Lys under physiological conditions. 41 These results are in accord with in vivo studies that proposed the existence of K-Ras4B nanoclusters with higher contents of acidic phospholipids For both, N-Ras and K-Ras4B no drastic influence of GTP/ GDP-loading on the membrane interaction process could be observed. Both membrane arrested states lead to nanoscale signaling platforms. However, the tendency toward clustering seems to be more pronounced for the active (GTP-bound) states of the proteins (Figure 7). Moreover, as visible in the AFM images, domains seem to largely sort by size. Interestingly, recent experimental and theoretical works have shown that partial budded membrane domains repel due to membrane mediated interactions which can be explained by an interplay between line tension and bending rigidity effects. 42 The spontaneous Ras isoform-specific sorting and the collective lateral organization of induced membrane domains could potentially operate as an effective, high fidelity signaling platform with distinct signal outputs for the Ras isoforms. In fact, nanoclustering has been proposed in in vivo experiments to be critical for signal transmission, as it allows the cell to respond to low signal inputs with a fixed output. 43 Next to the lipid anchor system, the G-domain provides additional determinants for K-Ras4B membrane interactions. Upon membrane binding of the inactive K-Ras4B, a shift of the R- helical subband to lower wavenumbers was observed in the FTIR spectra that is indicative for stronger membrane interactions. Since the R5-helix directly precedes the HVR (Figure 1), this effect can be attributed to an exposure of the R5-helix in the G-domain to the lipid interface. GDP-GTP exchange and thereby K-Ras4B activation leads to structural reorientations mainly in the switch I and switch II regions of the G-domain, 44 which is likely to reduce the R5-helix-membrane interactions observed in the FTIR data. The weaker interaction and higher reorientational flexibility of active K-Ras4B at the membrane interface were further confirmed by corresponding IRRAS experiments. In fact, it appears from recent molecular dynamics (MD) simulations that the HVR is the stabilizing motif for K-Ras4B membrane interaction and that the process involves mainly the C-terminal part of the HVR for K-Ras4B GTP, but the whole HVR for K-Ras4B GDP. 45 The R5-helix is closer to the membrane interface in the inactive state, which enables stronger membrane interaction. Hence, the FTIR and IRRA spectroscopy results support recent insights from MD simulations in showing minor but yet significant structural reorientations of the GDP/ GTP-K-Ras4B proteins at lipid interfaces. However, the effect of the R5-helix-membrane interaction needs to be studied in more detail, for example, by specific protein mutations, in future work. MATERIALS AND METHODS Materials and Sample Preparation. The phospholipids 1,2- dioleoyl-sn-glycero-3-phosphocholine (DOPC), 1,2-dioleoyl-sn-glycero- 3-phospho-(1 0 -rac-glycerol) sodium salt (DOPG), 1,2-dipalmitoyl-snglycero-3-phospho-(1 0 -rac-glycerol) sodium salt (DPPG), and 1,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC) were purchased from Avanti Polar Lipids (Alabaster, AL). Cholesterol (Chol) was from Sigma-Aldrich and the fluorescent lipids N-(lissamine rhodamine B sulfonyl)-1,2-dihexadecanoyl-sn-glycero-3-phosphoethanolamine triethylammonium salt (N-Rh-DHPE), 4,4-difluoro-5,7-dimethyl-4-bora-3a,4adiaza-s-indacene-3-propionic acid, succinimidyl ester (BODIPY FL, SE), and octadecylrhodamine B chloride (R18) from Molecular Probes (Invitrogen). Details on the preparation of lipid films are given in the Supporting Information. Protein Synthesis. The synthesis of the K-Ras4B proteins is described in detail in ref 23. Briefly, the S-farnesylated K-Ras4B protein was synthesized by a combination of expressed protein ligation (EPL) and lipopeptide synthesis. Details on the nucleotide exchange and BODIPY-labeling of K-Ras4B proteins are available in the Supporting Information. Atomic Force Microscopy (AFM). The preparation of the supported lipid bilayers and the AFM setup is described in detail in ref 24 and the Supporting Information. For the protein-membrane interaction studies, 200 μl of K-Ras4B protein in 20 mm Tris, 7 mm MgCl 2, ph 7.4 (c protein =2μM) were injected into the AFM fluid cell at room temperature and allowed to incubate for 1 h. Measurements were performed on a MultiMode scanning probe microscope with a Nano- Scope IIIa controller (Digital Instruments, Santa Barbara, CA) and usage of a J-Scanner (scan size 125 μm). Images were obtained by applying the tapping mode in liquid with oxide-sharpened silicon nitride (DNP-S) or sharp nitride lever (SNL) probes mounted in a fluid cell (MTFML, Veeco, Mannheim, Germany). Confocal Laser Scanning Microscopy of Model Membranes. GUVs were prepared by electroformation 32,46,47 in a temperature controlled homemade chamber on optically transparent and electrically conductive indium tin oxide (ITO) coated glass slides (SPI Supplies, West Chester, USA). The BODIPY-labeled K-Ras4B proteins were added via an injection-channel (c K-Ras4B = μm), yielding a final BODIPY concentration in the cell-chamber of 10 nm and a final molar ratio of lipid/n-rh-dhpe of 500:1. Images were recorded with a confocal laser scanning microscope (Biorad MRC 1024, extended for multiphoton excitation, now Zeiss, Germany) coupled via a side-port to an inverted microscope (Nikon, Eclipse TE-300 DV, infinity corrected optics) enabling fluorescence excitation in the focal plane of an objective lens (Nikon Plan Fluor 40, NA = 0.6, and Nikon Plan Appochromat VC 60 A WI, NA = 1.2 collar rim corr.). Details of the experimental setup are provided in the Supporting Information. Quantification of the protein partitioning was carried out by determining the background (BG) corrected fluorescence intensity ratios (FIR) of l o versus l d phases within the different phase regions of interest, that is, FIR = [I(l o ) - I(BG)]/[I(l d ) - I(BG)] and calculating the average value for at least eight GUVs. The analysis was performed similar to that described in ref 48. Confocal Laser Scanning Microscopy of Viral Membranes. GUVs were prepared by electroformation 46 on titan plates from viral lipids, isolated from influenza virus A/Japan/305/57 according to Bligh and Dyer. 49 A final BODIPY concentration of 1.7 and 2.3 μm R18 was used in a μ-slide VI chamber (ibidi, Martinsried, Germany). Confocal images of the equatorial plane of the GUVs in 250 mm glucose, 5.8 mm NaH 2 PO 4, 5.8 mm Na 2 HPO 4, ph 7.2 were taken using an inverted confocal laser scanning microscope (FV1000, Olympus, Hamburg, Germany) using a 60 (NA 1.20) water-immersion objective with a custom built temperature chamber. R18 and BODIPY were excited with a 559 nm diode laser and the 488 nm line of an Ar-ion laser (Olympus, Hamburg, Germany), respectively. The emissions of R18 and BODIPY were recorded between 570 and 670 nm and between 500 and 530 nm, respectively. FTIR Spectroscopy. For the infrared spectroscopic experiments, details on the preparation of large unilamellar vesicles in 20 mm Tris, 886 dx.doi.org/ /ja107532q J. Am. Chem. Soc. 2011, 133,
8 7 mm MgCl 2, pd 7.4, with included K-Ras4B (c K-Ras = 233 μm) are given in the Supporting Information. Measurements were performed at 25 C with a Nicolet 5700 FTIR spectrometer equipped with a liquid nitrogen cooled MCT (HgCdTe) detector and a cell with CaF 2 windows separated by 50 μm mylar spacers. Typically, FTIR-spectra of 128 scans were taken with a resolution of 2 cm -1 and correspondent processing was performed using GRAMS software (Thermo Electron). Details of the spectra correction and analysis are available in the Supporting Information. ASSOCIATED CONTENT b S Supporting Information. Complete refs 16 and 36. Details on materials and sample preparation. K-Ras4B protein synthesis: general aspects; cloning, expression, and purification of K-Ras4BΔ15-MESNA thioester, BODIPY labeling of the protein core thioester; nucleotide exchange on Ras proteins; ligation of K-Ras4B peptide with K-Ras4BΔ15-MESNA thioester. Details on the conduction of AFM, confocal laser scanning microscopy, FTIR, and IRRA spectroscopy experiments. Figures S1-S6 as described in the text. This material is available free of charge via the Internet at AUTHOR INFORMATION Corresponding Author roland.winter@tu-dortmund.de ACKNOWLEDGMENT This research was supported by the DFG (SFB 642 and SFB 740), the Max Planck Society (IMPRS), and the Leibniz- Graduate School Molecular Biophysics. We are grateful to Christine Nowak for technical assistance. REFERENCES (1) Wittinghofer, A.; Waldmann, H. Angew. Chem., Int. Ed. 2000, 39, (2) Hancock, J. F. Nat. Rev. Mol. Cell Biol. 2003, 4, (3) Wittinghofer, A.; Pai, E. F. Trends Biochem. Sci. 1991, 16, (4) Bos, J. L. Cancer Res. 1989, 49, (5) Marshall, C. J. Curr. Opin. Cell Biol. 1996, 8, (6) Willumsen, B. M.; Christensen, A.; Hubbert, N. L.; Papageorge, A. G.; Lowy, D. R. Nature 1984, 310, (7) Hancock, J. F.; Paterson, H.; Marshall, C. J. Cell 1990, 63, (8) Yeung, T.; Gilbert, G. E.; Shi, J.; Silvius, J.; Kapus, A.; Grinstein, S. Science 2008, 319, (9) Okeley, N. M.; Gelb, M. H. J. Biol. Chem. 2004, 279, (10) Meder, D.; Simons, K. Science 2005, 307, (11) Jackson, J. H.; Li, J. W.; Buss, J. E.; Der, C. J.; Cochrane, C. G. Proc. Natl. Acad. Sci. U.S.A. 1994, 91, (12) Ghomashchi, F.; Zhang, X.; Liu, L.; Gelb, M. H. Biochemistry 1995, 34, (13) Silvius, J. R. J. Membr. Biol. 2002, 190, (14) Jansen, B.; Schlagbauer-Wadl, H.; Kahr, H.; Heere-Ress, E.; Mayer, B. X.; Eichler, H.; Pehamberger, H.; Gana-Weisz, M.; Ben-David, E.; Kloog, Y.; Wolff, K. Proc. Natl. Acad. Sci. U.S.A. 1999, 96, (15) Chen, Z.; Otto, J. C.; Bergo, M. O.; Young, S. G.; Casey, P. J. J. Biol. Chem. 2000, 275, (16) Bivona, T. G.;; et al. Mol. Cell 2006, 21, (17) Roy, M. O.; Leventis, R.; Silvius, J. R. Biochemistry 2000, 39, (18) Gomez, G. A.; Daniotti, J. L. FEBS J. 2007, 274, (19) Leventis, R.; Silvius, J. R. Biochemistry 1998, 37, (20) Plowman, S. J.; Muncke, C.; Parton, R. G.; Hancock, J. F. Proc. Natl. Acad. Sci. U.S.A. 2005, 102, (21) Prior, I. A.; Muncke, C.; Parton, R. G.; Hancock, J. F. J. Cell Biol. 2003, 160, (22) Plowman, S. J.; Ariotti, N.; Goodall, A.; Parton, R. G.; Hancock, J. F. Mol. Cell. Biol. 2008, 28, (23) Chen, Y.-X.; Koch, S.; Uhlenbrock, K.; Weise, K.; Das, D.; Gremer, L.; Brunsveld, L.; Wittinghofer, A.; Winter, R.; Triola, G.; Waldmann, H. Angew. Chem., Int. Ed. 2010, 49, (24) Weise, K.; Triola, G.; Brunsveld, L.; Waldmann, H.; Winter, R. J. Am. Chem. Soc. 2009, 131, (25) Bagatolli, L. A.; Gratton, E. J. Fluoresc. 2001, 11, (26) Weise, K.; Triola, G.; Janosch, S.; Waldmann, H.; Winter, R. Biochim. Biophys. Acta 2010, 1798, (27) Fujisawa, T.; Uruga, T.; Yamaizumi, Z.; Inoko, Y.; Nishimura, S.; Ueki, T. J. Biochem. 1994, 115, (28) Vogel, A.; Reuther, G.; Weise, K.; Triola, G.; Nikolaus, J.; Tan, K.-T.; Nowak, C.; Herrmann, A.; Waldmann, H.; Winter, R.; Huster, D. Angew. Chem., Int. Ed. 2009, 48, (29) Tamm, L. K.; Tatulian, S. A. Q. Rev. Biophys. 1997, 304, (30) Milburn, M. V.; Tong, L.; devos, A. M.; Br unger, A.; Yamaizumi, Z.; Nishimura, S.; Kim, S. H. Science 1990, 247, (31) Meister, A.; Nicolini, C.; Waldmann, H.; Kuhlmann, J.; Kerth, A.; Winter, R.; Blume, A. Biophys. J. 2006, 91, (32) Scheiffele, P.; Rietveld, A.; Wilk, T.; Simons, K. J. Biol. Chem. 1999, 274, (33) Nicolini, C.; Baranski, J.; Schlummer, S.; Palomo, J.; Burgues, M. L.; Kahms, M.; Kuhlmann, J.; Sanchez, S.; Gratton, E.; Waldmann, H.; Winter, R. J. Am. Chem. Soc. 2006, 128, (34) Rocks, O.; Peyker, A.; Kahms, M.; Verveer, P. J.; Koerner, C.; Lumbierres, M.; Kuhlmann, J.; Waldmann, H.; Wittinghofer, A.; Bastiaens, P. I. Science 2005, 307, (35) Rocks, O.; Gerauer, M.; Vartak, N.; Koch, S.; Huang, Z. P.; Pechlivanis, M.; Kuhlmann, J.; Brunsveld, L.; Chandra, A.; Ellinger, B.; Waldmann, H.; Bastiaens, P. I. Cell 2010, 141, (36) Dekker, F. J.;; et al. Nat. Chem. Biol. 2010, 6, (37) Bunge, A.; Kurz, A.; Windeck, A.-K.; Korte, T.; Flasche, W.; Liebscher, J.; Herrmann, A.; Huster, D. Langmuir 2007, 23, (38) McLaughlin, S.; Murray, D. Nature 2005, 438, (39) Golebiewska, U.; Gambhir, A.; Hangyas-Mihalyne, G.; Zaitseva, I.; R adler, J.; McLaughlin, S. Biophys. J. 2006, 91, (40) Heo, W. D.; Inoue, T.; Park, W. S.; Kim, M. L.; Park, B. O.; Wandless, T. J.; Meyer, T. Science 2006, 314, (41) Ben-Tal, N.; Honig, B.; Peitzsch, R. M.; Denisov, G.; McLaughlin, S. Biophys. J. 1996, 71, (42) Idema, T.; Semrau, S.; Storm, C.; Schmidt, T. Phys. Rev. Lett. 2010, 104, (43) Tian, T.; Harding, A.; Inder, K.; Plowman, S.; Parton, R. G.; Hancock, J. F. Nat. Cell Biol. 2007, 9, (44) Lowy, D. R.; Willumsen, B. M. Annu. Rev. Biochem. 1993, 62, (45) Abankwa, D.; Gorfe, A. A.; Inder, K.; Hancock, J. F. Proc. Natl. Acad. Sci. U.S.A. 2010, 107, (46) Angelova, M. I.; Soleau, S.; Meleard, P.; Faucon, F.; Bothorel, P. Prog. Colloid Polym. Sci. 1992, 89, (47) Janosch, S.; Nicolini, C.; Ludolph, B.; Peters, C.; V olkert, M.; Hazlet, T. L.; Gratton, E.; Waldmann, H.; Winter, R. J. Am. Chem. Soc. 2004, 126, (48) Johnson, S. A.; Stinson, B. M.; Go, M. S.; Carmona, L. M.; Reminick, J. I.; Fang, X.; Baumgart, T. Biochim. Biophys. Acta 2010, 1798, (49) Bligh, E. G.; Dyer, W. J. Can. J. Biochem. Physiol. 1959, 37, dx.doi.org/ /ja107532q J. Am. Chem. Soc. 2011, 133,
Pressure Modulation of the Enzymatic Activity of. Phospholipase A2, a Putative Membraneassociated
 SUPPORTING INFORMATION Pressure Modulation of the Enzymatic Activity of Phospholipase A2, a Putative Membraneassociated Pressure Sensor Saba Suladze, Suleyman Cinar, Benjamin Sperlich, and Roland Winter*
SUPPORTING INFORMATION Pressure Modulation of the Enzymatic Activity of Phospholipase A2, a Putative Membraneassociated Pressure Sensor Saba Suladze, Suleyman Cinar, Benjamin Sperlich, and Roland Winter*
STRUCTURE AND MEMBRANE MICRO- DOMAIN LOCALIZATION OF RAS PROTEINS
 Katrin Weise STRUCTURE AND MEMBRANE MICRO- DOMAIN LOCALIZATION OF RAS PROTEINS 1 INTRODUCTION Small GTPases of the rat sarcoma (Ras) superfamily act as binary molecular switches between a GDP-bound inactive
Katrin Weise STRUCTURE AND MEMBRANE MICRO- DOMAIN LOCALIZATION OF RAS PROTEINS 1 INTRODUCTION Small GTPases of the rat sarcoma (Ras) superfamily act as binary molecular switches between a GDP-bound inactive
Electronic Supporting Information
 Modulation of raft domains in a lipid bilayer by boundary-active curcumin Manami Tsukamoto a, Kenichi Kuroda* b, Ayyalusamy Ramamoorthy* c, Kazuma Yasuhara* a Electronic Supporting Information Contents
Modulation of raft domains in a lipid bilayer by boundary-active curcumin Manami Tsukamoto a, Kenichi Kuroda* b, Ayyalusamy Ramamoorthy* c, Kazuma Yasuhara* a Electronic Supporting Information Contents
Supplementary information. Membrane curvature enables N-Ras lipid anchor sorting to. liquid-ordered membrane phases
 Supplementary information Membrane curvature enables N-Ras lipid anchor sorting to liquid-ordered membrane phases Jannik Bruun Larsen 1,2,3, Martin Borch Jensen 1,2,3,6, Vikram K. Bhatia 1,2,3,7, Søren
Supplementary information Membrane curvature enables N-Ras lipid anchor sorting to liquid-ordered membrane phases Jannik Bruun Larsen 1,2,3, Martin Borch Jensen 1,2,3,6, Vikram K. Bhatia 1,2,3,7, Søren
AFM In Liquid: A High Sensitivity Study On Biological Membranes
 University of Wollongong Research Online Faculty of Science - Papers (Archive) Faculty of Science, Medicine and Health 2006 AFM In Liquid: A High Sensitivity Study On Biological Membranes Michael J. Higgins
University of Wollongong Research Online Faculty of Science - Papers (Archive) Faculty of Science, Medicine and Health 2006 AFM In Liquid: A High Sensitivity Study On Biological Membranes Michael J. Higgins
Plasma membrane structure and dynamics explored via a combined AFM/FCS approach
 Plasma membrane structure and dynamics explored via a combined AFM/FCS approach Salvatore Chiantia Molekulare Biophysik, Dept. Of Biology Humboldt-Universität zu Berlin Dresden nanoseminar, May 2013 Outline
Plasma membrane structure and dynamics explored via a combined AFM/FCS approach Salvatore Chiantia Molekulare Biophysik, Dept. Of Biology Humboldt-Universität zu Berlin Dresden nanoseminar, May 2013 Outline
Polyoxometalate Macroion Induced Phase and Morphology
 Polyoxometalate Macroion Induced Phase and Morphology Instability of Lipid Membrane Benxin Jing a, Marie Hutin c, Erin Connor a, Leroy Cronin c,* and Yingxi Zhu a,b,* a Department of Chemical and Biomolecular
Polyoxometalate Macroion Induced Phase and Morphology Instability of Lipid Membrane Benxin Jing a, Marie Hutin c, Erin Connor a, Leroy Cronin c,* and Yingxi Zhu a,b,* a Department of Chemical and Biomolecular
Chemical Surface Transformation 1
 Chemical Surface Transformation 1 Chemical reactions at Si H surfaces (inorganic and organic) can generate very thin films (sub nm thickness up to µm): inorganic layer formation by: thermal conversion:
Chemical Surface Transformation 1 Chemical reactions at Si H surfaces (inorganic and organic) can generate very thin films (sub nm thickness up to µm): inorganic layer formation by: thermal conversion:
The Transport and Organization of Cholesterol in Planar Solid-Supported Lipid Bilayer Depend on the Phospholipid Flip-Flop Rate
 Supporting Information The Transport and Organization of Cholesterol in Planar Solid-Supported Lipid Bilayer Depend on the Phospholipid Flip-Flop Rate Ting Yu, 1,2 Guangnan Zhou, 1 Xia Hu, 1,2 Shuji Ye
Supporting Information The Transport and Organization of Cholesterol in Planar Solid-Supported Lipid Bilayer Depend on the Phospholipid Flip-Flop Rate Ting Yu, 1,2 Guangnan Zhou, 1 Xia Hu, 1,2 Shuji Ye
Supplementary Figure 1. Overview of steps in the construction of photosynthetic protocellular systems
 Supplementary Figure 1 Overview of steps in the construction of photosynthetic protocellular systems (a) The small unilamellar vesicles were made with phospholipids. (b) Three types of small proteoliposomes
Supplementary Figure 1 Overview of steps in the construction of photosynthetic protocellular systems (a) The small unilamellar vesicles were made with phospholipids. (b) Three types of small proteoliposomes
Physical Cell Biology Lecture 10: membranes elasticity and geometry. Hydrophobicity as an entropic effect
 Physical Cell Biology Lecture 10: membranes elasticity and geometry Phillips: Chapter 5, Chapter 11 and Pollard Chapter 13 Hydrophobicity as an entropic effect 1 Self-Assembly of Lipid Structures Lipid
Physical Cell Biology Lecture 10: membranes elasticity and geometry Phillips: Chapter 5, Chapter 11 and Pollard Chapter 13 Hydrophobicity as an entropic effect 1 Self-Assembly of Lipid Structures Lipid
Cellular Biochemistry
 Cellular Biochemistry Fall Semester 2013 Sept. 23 Benoit Kornmann Institute of Biochemistry Introduction to biological membranes General functions and properties Membrane lipids Physical properties Distribution/asymmetry
Cellular Biochemistry Fall Semester 2013 Sept. 23 Benoit Kornmann Institute of Biochemistry Introduction to biological membranes General functions and properties Membrane lipids Physical properties Distribution/asymmetry
H-ras Protein in a Bilayer: Interaction and Structure Perturbation
 Published on Web 09/19/2007 H-ras Protein in a Bilayer: Interaction and Structure Perturbation Alemayehu A. Gorfe,*, Arneh Babakhani, and J. Andrew McCammon,, Contribution from the Department of Chemistry
Published on Web 09/19/2007 H-ras Protein in a Bilayer: Interaction and Structure Perturbation Alemayehu A. Gorfe,*, Arneh Babakhani, and J. Andrew McCammon,, Contribution from the Department of Chemistry
Methods and Materials
 a division of Halcyonics GmbH Anna-Vandenhoeck-Ring 5 37081 Göttingen, Germany Application Note Micostructured lipid bilayers ANDREAS JANSHOFF 1), MAJA GEDIG, AND SIMON FAISS Fig.1: Thickness map of microstructured
a division of Halcyonics GmbH Anna-Vandenhoeck-Ring 5 37081 Göttingen, Germany Application Note Micostructured lipid bilayers ANDREAS JANSHOFF 1), MAJA GEDIG, AND SIMON FAISS Fig.1: Thickness map of microstructured
CHAPTER 4. Tryptophan fluorescence quenching by brominated lipids
 CHAPTER 4 Tryptophan fluorescence quenching by brominated lipids 102 4.1 INTRODUCTION The structure and dynamics of biological macromolecules have been widely studied with fluorescence quenching. The accessibility
CHAPTER 4 Tryptophan fluorescence quenching by brominated lipids 102 4.1 INTRODUCTION The structure and dynamics of biological macromolecules have been widely studied with fluorescence quenching. The accessibility
780 A. Vogel et al.: Ras interaction with raft models
 DOI 10.1515/hsz-2013-0294 Biol. Chem. 2014; 395(7-8): 779 789 Alexander Vogel, Jörg Nikolaus a, Katrin Weise, Gemma Triola, Herbert Waldmann, Roland Winter, Andreas Herrmann and Daniel Huster* Interaction
DOI 10.1515/hsz-2013-0294 Biol. Chem. 2014; 395(7-8): 779 789 Alexander Vogel, Jörg Nikolaus a, Katrin Weise, Gemma Triola, Herbert Waldmann, Roland Winter, Andreas Herrmann and Daniel Huster* Interaction
Protein directed assembly of lipids
 Protein directed assembly of lipids D. Nordin, O. Yarkoni, L. Donlon, N. Savinykh, and D.J. Frankel SUPPLEMENTARY MATERIAL Materials and Methods Supported bilayer preparation 1,2-dioleoyl-sn-glycero-3-phosphocholine
Protein directed assembly of lipids D. Nordin, O. Yarkoni, L. Donlon, N. Savinykh, and D.J. Frankel SUPPLEMENTARY MATERIAL Materials and Methods Supported bilayer preparation 1,2-dioleoyl-sn-glycero-3-phosphocholine
Supplementary Materials. High affinity binding of phosphatidylinositol-4-phosphate. by Legionella pneumophila DrrA
 Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Giant vesicles formed by gentle hydration and electroformation: A comparison by fluorescence microscopy
 Colloids and Surfaces B: Biointerfaces 42 (2005) 125 130 Giant vesicles formed by gentle hydration and electroformation: A comparison by fluorescence microscopy Nicolas Rodriguez a,frédéric Pincet b, Sophie
Colloids and Surfaces B: Biointerfaces 42 (2005) 125 130 Giant vesicles formed by gentle hydration and electroformation: A comparison by fluorescence microscopy Nicolas Rodriguez a,frédéric Pincet b, Sophie
Phase Transition Behaviours of the Supported DPPC Bilayer. Investigated by Sum Frequency Generation (SFG) and Atomic Force
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2015 Supporting Information for Phase Transition Behaviours of the Supported DPPC Bilayer
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2015 Supporting Information for Phase Transition Behaviours of the Supported DPPC Bilayer
Cell Membranes. Dr. Diala Abu-Hassan School of Medicine Cell and Molecular Biology
 Cell Membranes Dr. Diala Abu-Hassan School of Medicine Dr.abuhassand@gmail.com Cell and Molecular Biology Organelles 2Dr. Diala Abu-Hassan Membrane proteins Major components of cells Nucleic acids DNA
Cell Membranes Dr. Diala Abu-Hassan School of Medicine Dr.abuhassand@gmail.com Cell and Molecular Biology Organelles 2Dr. Diala Abu-Hassan Membrane proteins Major components of cells Nucleic acids DNA
Signal Transduction Cascades
 Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
A ph-dependent Charge Reversal Peptide for Cancer Targeting
 Supporting Information A ph-dependent Charge Reversal Peptide for Cancer Targeting Naoko Wakabayashi 1, Yoshiaki Yano 1, Kenichi Kawano 1, and Katsumi Matsuzaki 1 1 Graduate School of Pharmaceutical Sciences,
Supporting Information A ph-dependent Charge Reversal Peptide for Cancer Targeting Naoko Wakabayashi 1, Yoshiaki Yano 1, Kenichi Kawano 1, and Katsumi Matsuzaki 1 1 Graduate School of Pharmaceutical Sciences,
bio-mof-1 DMASM Wavenumber (cm -1 ) Supplementary Figure S1 FTIR spectra of bio-mof-1, DMASMI, and bio-mof-1 DMASM.
 bio-mof-1 Transmittance bio-mof-1 DMASM DMASMI 2000 1500 1000 500 Wavenumber (cm -1 ) Supplementary Figure S1 FTIR spectra of bio-mof-1, DMASMI, and bio-mof-1 DMASM. Intensity (a.u.) bio-mof-1 DMASM as
bio-mof-1 Transmittance bio-mof-1 DMASM DMASMI 2000 1500 1000 500 Wavenumber (cm -1 ) Supplementary Figure S1 FTIR spectra of bio-mof-1, DMASMI, and bio-mof-1 DMASM. Intensity (a.u.) bio-mof-1 DMASM as
Calcium-dependent Hydrolysis of Supported Planar Lipids Were Triggered by honey bee venom Phospholipase A 2 with Right Orientation at Interface
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2017 Calcium-dependent Hydrolysis of Supported Planar Lipids Were Triggered by honey
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2017 Calcium-dependent Hydrolysis of Supported Planar Lipids Were Triggered by honey
Interactions of Polyethylenimines with Zwitterionic and. Anionic Lipid Membranes
 Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Supplementary Information: Liquid-liquid phase coexistence in lipid membranes observed by natural abundance 1 H 13 C solid-state NMR
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the wner Societies 28 Supplementary Information: Liquid-liquid phase coexistence in lipid membranes observed
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the wner Societies 28 Supplementary Information: Liquid-liquid phase coexistence in lipid membranes observed
Phosphatidylcholines are a class of glycerophospholipids which along with other phospholipids
 Phosphatidylcholine Phosphatidylcholines are a class of glycerophospholipids which along with other phospholipids account for more than half of the lipids in most membranes. Phosphatidylcholines can further
Phosphatidylcholine Phosphatidylcholines are a class of glycerophospholipids which along with other phospholipids account for more than half of the lipids in most membranes. Phosphatidylcholines can further
Supplementary Figure S1 Black silicon and dragonfly wing nanotopography.
 Supplementary Figure S1 Black silicon and dragonfly wing nanotopography. Representative low-magnification scanning electron micrographs of a) Black silicon (bsi) and b) Diplacodes bipunctata dragonfly
Supplementary Figure S1 Black silicon and dragonfly wing nanotopography. Representative low-magnification scanning electron micrographs of a) Black silicon (bsi) and b) Diplacodes bipunctata dragonfly
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/6/e1700338/dc1 Supplementary Materials for HIV virions sense plasma membrane heterogeneity for cell entry Sung-Tae Yang, Alex J. B. Kreutzberger, Volker Kiessling,
advances.sciencemag.org/cgi/content/full/3/6/e1700338/dc1 Supplementary Materials for HIV virions sense plasma membrane heterogeneity for cell entry Sung-Tae Yang, Alex J. B. Kreutzberger, Volker Kiessling,
Chapter 1 Membrane Structure and Function
 Chapter 1 Membrane Structure and Function Architecture of Membranes Subcellular fractionation techniques can partially separate and purify several important biological membranes, including the plasma and
Chapter 1 Membrane Structure and Function Architecture of Membranes Subcellular fractionation techniques can partially separate and purify several important biological membranes, including the plasma and
Raman spectroscopy methods for detecting and imaging supported lipid bilayers
 Spectroscopy 24 (2010) 113 117 113 DOI 10.3233/SPE-2010-0426 IOS Press Raman spectroscopy methods for detecting and imaging supported lipid bilayers Claire S. Sweetenham and Ioan Notingher Nanoscience
Spectroscopy 24 (2010) 113 117 113 DOI 10.3233/SPE-2010-0426 IOS Press Raman spectroscopy methods for detecting and imaging supported lipid bilayers Claire S. Sweetenham and Ioan Notingher Nanoscience
Nucleophosmin and Nucleolin regulate K-Ras Signaling. Key words: K-Ras, plasma membrane nanoclusters, ERK activation, NPM, nucleolin
 Nucleophosmin and Nucleolin regulate K-Ras Signaling Kerry L Inder 1, Michelle M Hill 2 and John F Hancock 3 1 Department of Pathology, University of Cambridge, Cambridge, United Kingdom. 2 Diamantina
Nucleophosmin and Nucleolin regulate K-Ras Signaling Kerry L Inder 1, Michelle M Hill 2 and John F Hancock 3 1 Department of Pathology, University of Cambridge, Cambridge, United Kingdom. 2 Diamantina
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Lecture 15. Membrane Proteins I
 Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Nature Biotechnology: doi: /nbt.3828
 Supplementary Figure 1 Development of a FRET-based MCS. (a) Linker and MA2 modification are indicated by single letter amino acid code. indicates deletion of amino acids and N or C indicate the terminus
Supplementary Figure 1 Development of a FRET-based MCS. (a) Linker and MA2 modification are indicated by single letter amino acid code. indicates deletion of amino acids and N or C indicate the terminus
Rama Abbady. Odai Bani-Monia. Diala Abu-Hassan
 5 Rama Abbady Odai Bani-Monia Diala Abu-Hassan Lipid Rafts Lipid rafts are aggregates (accumulations) of sphingolipids. They re semisolid clusters (10-200 nm) of cholesterol and sphingolipids (sphingomyelin
5 Rama Abbady Odai Bani-Monia Diala Abu-Hassan Lipid Rafts Lipid rafts are aggregates (accumulations) of sphingolipids. They re semisolid clusters (10-200 nm) of cholesterol and sphingolipids (sphingomyelin
MOLECULAR DYNAMICS SIMULATION OF MIXED LIPID BILAYER WITH DPPC AND MPPC: EFFECT OF CONFIGURATIONS IN GEL-PHASE
 MOLECULAR DYNAMICS SIMULATION OF MIXED LIPID BILAYER WITH DPPC AND MPPC: EFFECT OF CONFIGURATIONS IN GEL-PHASE A Thesis Presented to The Academic Faculty by Young Kyoung Kim In Partial Fulfillment of the
MOLECULAR DYNAMICS SIMULATION OF MIXED LIPID BILAYER WITH DPPC AND MPPC: EFFECT OF CONFIGURATIONS IN GEL-PHASE A Thesis Presented to The Academic Faculty by Young Kyoung Kim In Partial Fulfillment of the
Tumor Targeting of Functionalized Quantum Dot- Liposome Hybrids by Intravenous Administration
 Tumor Targeting of Functionalized Quantum Dot- Liposome Hybrids by Intravenous Administration Wafa T. Al-Jamal 1, Khuloud T. Al-Jamal 1, Bowen Tian 1, Andrew Cakebread 2, John M. Halket 2 and Kostas Kostarelos
Tumor Targeting of Functionalized Quantum Dot- Liposome Hybrids by Intravenous Administration Wafa T. Al-Jamal 1, Khuloud T. Al-Jamal 1, Bowen Tian 1, Andrew Cakebread 2, John M. Halket 2 and Kostas Kostarelos
Electronic Supplementary Information
 Electronic Supplementary Information Orientation of Lipid Domains in Giant Vesicles Using an Electric Field Frank J. Zendejas, Robert J. Meagher, Jeanne C. Stachowiak, Carl C. Hayden, Bruce C. Bunker,
Electronic Supplementary Information Orientation of Lipid Domains in Giant Vesicles Using an Electric Field Frank J. Zendejas, Robert J. Meagher, Jeanne C. Stachowiak, Carl C. Hayden, Bruce C. Bunker,
Food protein powders classification and discrimination by FTIR spectroscopy and principal component analysis
 APPLICATION NOTE AN53037 Food protein powders classification and discrimination by FTIR spectroscopy and principal component analysis Author Ron Rubinovitz, Ph.D. Thermo Fisher Scientific Key Words FTIR,
APPLICATION NOTE AN53037 Food protein powders classification and discrimination by FTIR spectroscopy and principal component analysis Author Ron Rubinovitz, Ph.D. Thermo Fisher Scientific Key Words FTIR,
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/12/e1601838/dc1 Supplementary Materials for General and programmable synthesis of hybrid liposome/metal nanoparticles Jin-Ho Lee, Yonghee Shin, Wooju Lee, Keumrai
advances.sciencemag.org/cgi/content/full/2/12/e1601838/dc1 Supplementary Materials for General and programmable synthesis of hybrid liposome/metal nanoparticles Jin-Ho Lee, Yonghee Shin, Wooju Lee, Keumrai
Surface-Attached Polyhistidine-Tag Proteins Characterized by FTIR Difference Spectroscopy
 DOI: 10.1002/cphc.201200358 Surface-Attached Polyhistidine-Tag Proteins Characterized by FTIR Difference Spectroscopy Philipp Pinkerneil, Jçrn Güldenhaupt, Klaus Gerwert,* and Carsten Kçtting* [a] FTIR
DOI: 10.1002/cphc.201200358 Surface-Attached Polyhistidine-Tag Proteins Characterized by FTIR Difference Spectroscopy Philipp Pinkerneil, Jçrn Güldenhaupt, Klaus Gerwert,* and Carsten Kçtting* [a] FTIR
Name: Multiple choice questions. Pick the BEST answer (2 pts ea)
 Exam 1 202 Oct. 5, 1999 Multiple choice questions. Pick the BEST answer (2 pts ea) 1. The lipids of a red blood cell membrane are all a. phospholipids b. amphipathic c. glycolipids d. unsaturated 2. The
Exam 1 202 Oct. 5, 1999 Multiple choice questions. Pick the BEST answer (2 pts ea) 1. The lipids of a red blood cell membrane are all a. phospholipids b. amphipathic c. glycolipids d. unsaturated 2. The
Shape transformation of giant phospholipid vesicles at high concentrations of C 12 E 8
 Bioelectrochemistry 63 (2004) 183 187 www.elsevier.com/locate/bioelechem Shape transformation of giant phospholipid vesicles at high concentrations of C 12 E 8 B. Mavčič, B. Babnik, A. Iglič*, M. Kandušer,
Bioelectrochemistry 63 (2004) 183 187 www.elsevier.com/locate/bioelechem Shape transformation of giant phospholipid vesicles at high concentrations of C 12 E 8 B. Mavčič, B. Babnik, A. Iglič*, M. Kandušer,
Supporting material. Membrane permeation induced by aggregates of human islet amyloid polypeptides
 Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
Palmitoylation and localisation of RAS isoforms are modulated by the hypervariable linker domain
 Research Article 421 Palmitoylation and localisation of RAS isoforms are modulated by the hypervariable linker domain Alex J. Laude and Ian A. Prior* Physiological Laboratory, University of Liverpool,
Research Article 421 Palmitoylation and localisation of RAS isoforms are modulated by the hypervariable linker domain Alex J. Laude and Ian A. Prior* Physiological Laboratory, University of Liverpool,
SUPPLEMENTARY INFORMATION
 High-speed atomic force microscopy shows that annexin V stabilizes membranes on the second timescale Atsushi Miyagi, Chris Chipot, Martina Rangl & Simon Scheuring Supplementary Movies: Supplementary Movie
High-speed atomic force microscopy shows that annexin V stabilizes membranes on the second timescale Atsushi Miyagi, Chris Chipot, Martina Rangl & Simon Scheuring Supplementary Movies: Supplementary Movie
Quantifying Lipid Contents in Enveloped Virus Particles with Plasmonic Nanoparticles
 Quantifying Lipid Contents in Enveloped Virus Particles with Plasmonic Nanoparticles Amin Feizpour Reinhard Lab Department of Chemistry and the Photonics Center, Boston University, Boston, MA May 2014
Quantifying Lipid Contents in Enveloped Virus Particles with Plasmonic Nanoparticles Amin Feizpour Reinhard Lab Department of Chemistry and the Photonics Center, Boston University, Boston, MA May 2014
Chapter 14 - Electron Transport and Oxidative Phosphorylation
 Chapter 14 - Electron Transport and Oxidative Phosphorylation The cheetah, whose capacity for aerobic metabolism makes it one of the fastest animals Prentice Hall c2002 Chapter 14 1 14.4 Oxidative Phosphorylation
Chapter 14 - Electron Transport and Oxidative Phosphorylation The cheetah, whose capacity for aerobic metabolism makes it one of the fastest animals Prentice Hall c2002 Chapter 14 1 14.4 Oxidative Phosphorylation
Nucleic acids. Nucleic acids are information-rich polymers of nucleotides
 Nucleic acids Nucleic acids are information-rich polymers of nucleotides DNA and RNA Serve as the blueprints for proteins and thus control the life of a cell RNA and DNA are made up of very similar nucleotides.
Nucleic acids Nucleic acids are information-rich polymers of nucleotides DNA and RNA Serve as the blueprints for proteins and thus control the life of a cell RNA and DNA are made up of very similar nucleotides.
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Lipids and Membranes
 Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Biological membranes are composed of lipid bilayers
Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Biological membranes are composed of lipid bilayers
Polarization and Circular Dichroism (Notes 17)
 Polarization and Circular Dichroism - 2014 (Notes 17) Since is vector, if fix molec. orient., E-field interact (absorb) with molecule differently when change E-orientation (polarization) Transitions can
Polarization and Circular Dichroism - 2014 (Notes 17) Since is vector, if fix molec. orient., E-field interact (absorb) with molecule differently when change E-orientation (polarization) Transitions can
Domain Growth Kinetics in a Cell-sized Liposome
 Domain Growth Kinetics in a Cell-sized Liposome Daisuke SAEKI, Tsutomu HAMADA and Kenichi YOSHIKAWA * Department of Physics, Graduate School of Science, Kyoto University, Kyoto 606-8502 Abstract We investigated
Domain Growth Kinetics in a Cell-sized Liposome Daisuke SAEKI, Tsutomu HAMADA and Kenichi YOSHIKAWA * Department of Physics, Graduate School of Science, Kyoto University, Kyoto 606-8502 Abstract We investigated
Biochemistry 1 Recitation1 Cell & Water
 Biochemistry 1 Recitation1 Cell & Water There are several important themes that transcends the chemistry and bring the importance of understanding the cell biological differences between eukaryotes and
Biochemistry 1 Recitation1 Cell & Water There are several important themes that transcends the chemistry and bring the importance of understanding the cell biological differences between eukaryotes and
Gel-assisted formation of giant unilamellar vesicles
 Gel-assisted formation of giant unilamellar vesicles Andreas Weinberger 1, Feng-Ching Tsai 2, Gijsje H. Koenderink 2, Thais F. Schmidt 3, Rosângela Itri 4, Wolfgang Meier 5, Tatiana Schmatko 1, André Schröder
Gel-assisted formation of giant unilamellar vesicles Andreas Weinberger 1, Feng-Ching Tsai 2, Gijsje H. Koenderink 2, Thais F. Schmidt 3, Rosângela Itri 4, Wolfgang Meier 5, Tatiana Schmatko 1, André Schröder
Supporting Information
 Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany Stepwise Directing Nanocrystals to Self-Assemble at Water/Oil Interfaces Jing Wang, Dayang Wang, Nelli S. Sobal, Michael Giersig, Ming Jiang,
Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany Stepwise Directing Nanocrystals to Self-Assemble at Water/Oil Interfaces Jing Wang, Dayang Wang, Nelli S. Sobal, Michael Giersig, Ming Jiang,
Models of the plasma membrane - from the fluid mosaic to the picket fence model. Mario Schelhaas Institute of Cellular Virology
 Models of the plasma membrane - from the fluid mosaic to the picket fence model Mario Schelhaas Institute of Cellular Virology Today s lecture Central Question: How does the plasma membrane fulfil its
Models of the plasma membrane - from the fluid mosaic to the picket fence model Mario Schelhaas Institute of Cellular Virology Today s lecture Central Question: How does the plasma membrane fulfil its
Structure-Transport Relationship in Organized Soft Matter Systems by Diffusion NMR. Sergey Vasenkov
 Structure-Transport Relationship in Organized Soft Matter Systems by Diffusion NMR Sergey Vasenkov Outline Combining advantages of high field and high gradients in diffusion NMR Relationship between diffusivities
Structure-Transport Relationship in Organized Soft Matter Systems by Diffusion NMR Sergey Vasenkov Outline Combining advantages of high field and high gradients in diffusion NMR Relationship between diffusivities
Microtubule Teardrop Patterns
 Supporting Information Microtubule Teardrop Patterns Kosuke Okeyoshi 1, Ryuzo Kawamura 1, Ryo Yoshida 2, and Yoshihito Osada 1 * 1 RIKEN Advanced Science Institute, 2-1 Hirosawa, Wako-shi, Saitama 351-0198,
Supporting Information Microtubule Teardrop Patterns Kosuke Okeyoshi 1, Ryuzo Kawamura 1, Ryo Yoshida 2, and Yoshihito Osada 1 * 1 RIKEN Advanced Science Institute, 2-1 Hirosawa, Wako-shi, Saitama 351-0198,
Noninvasive Blood Glucose Analysis using Near Infrared Absorption Spectroscopy. Abstract
 Progress Report No. 2-3, March 31, 1999 The Home Automation and Healthcare Consortium Noninvasive Blood Glucose Analysis using Near Infrared Absorption Spectroscopy Prof. Kamal Youcef-Toumi Principal Investigator
Progress Report No. 2-3, March 31, 1999 The Home Automation and Healthcare Consortium Noninvasive Blood Glucose Analysis using Near Infrared Absorption Spectroscopy Prof. Kamal Youcef-Toumi Principal Investigator
H 2 O. Liquid, solid, and vapor coexist in the same environment
 Water H 2 O Liquid, solid, and vapor coexist in the same environment WATER MOLECULES FORM HYDROGEN BONDS Water is a fundamental requirement for life, so it is important to understand the structural and
Water H 2 O Liquid, solid, and vapor coexist in the same environment WATER MOLECULES FORM HYDROGEN BONDS Water is a fundamental requirement for life, so it is important to understand the structural and
Lecture 36: Review of membrane function
 Chem*3560 Lecture 36: Review of membrane function Membrane: Lipid bilayer with embedded or associated proteins. Bilayers: 40-70% neutral phospholipid 10-20% negative phospholipid 10-30% cholesterol 10-30%
Chem*3560 Lecture 36: Review of membrane function Membrane: Lipid bilayer with embedded or associated proteins. Bilayers: 40-70% neutral phospholipid 10-20% negative phospholipid 10-30% cholesterol 10-30%
SUPPORTING INFORMATION. Lysine Carbonylation is a Previously Unrecognized Contributor. to Peroxidase Activation of Cytochrome c by Chloramine-T
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Fluid Mozaic Model of Membranes
 Replacement for the 1935 Davson Danielli model Provided explanation for Gortner-Grendel lack of lipid and permitted the unit membrane model. Trans membrane protein by labelling Fry & Edidin showed that
Replacement for the 1935 Davson Danielli model Provided explanation for Gortner-Grendel lack of lipid and permitted the unit membrane model. Trans membrane protein by labelling Fry & Edidin showed that
Supporting information for
 Supporting information for Nitrogen and Sulfur Co-doped Carbon Dots-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry Imaging for Profiling Bisphenol S Distribution in Mouse Tissues
Supporting information for Nitrogen and Sulfur Co-doped Carbon Dots-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry Imaging for Profiling Bisphenol S Distribution in Mouse Tissues
Biology 4410 First Examination Version B
 Biology 4410 Spring 2006 Name First Examination Version B This examination consists of two parts, a multiple-choice section and an essay section. Be sure to put your name on both the mark-sense sheet and
Biology 4410 Spring 2006 Name First Examination Version B This examination consists of two parts, a multiple-choice section and an essay section. Be sure to put your name on both the mark-sense sheet and
ORGANIZATION OF ANTIBIOTIC AMPHOTERICIN B IN MODEL LIPID MEMBRANES. A MINI REVIEW
 CELLULAR & MOLECULAR BIOLOGY LETTERS Volume 8, (2003) pp 161 170 http://www.cmbl.org.pl Received 30 September 2002 Accepted 13 February 2003 ORGANIZATION OF ANTIBIOTIC AMPHOTERICIN B IN MODEL LIPID MEMBRANES.
CELLULAR & MOLECULAR BIOLOGY LETTERS Volume 8, (2003) pp 161 170 http://www.cmbl.org.pl Received 30 September 2002 Accepted 13 February 2003 ORGANIZATION OF ANTIBIOTIC AMPHOTERICIN B IN MODEL LIPID MEMBRANES.
Cellular Neurophysiology I Membranes and Ion Channels
 Cellular Neurophysiology I Membranes and Ion Channels Reading: BCP Chapter 3 www.bioelectriclab All living cells maintain an electrical potential (voltage) across their membranes (V m ). Resting Potential
Cellular Neurophysiology I Membranes and Ion Channels Reading: BCP Chapter 3 www.bioelectriclab All living cells maintain an electrical potential (voltage) across their membranes (V m ). Resting Potential
Supplementary Information
 Supplementary Information Supplementary Figure 1. Fabrication processes of bud-mimicking PDMS patterns based on a combination of colloidal, soft, and photo-lithography a-d, The polystyrene (PS) particles
Supplementary Information Supplementary Figure 1. Fabrication processes of bud-mimicking PDMS patterns based on a combination of colloidal, soft, and photo-lithography a-d, The polystyrene (PS) particles
Chapter 9 - Biological Membranes. Membranes form a semi-permeable boundary between a cell and its environment.
 Chapter 9 - Biological Membranes www.gsbs.utmb.edu/ microbook/ch037.htmmycoplasma Membranes form a semi-permeable boundary between a cell and its environment. Membranes also permit subcellular organization
Chapter 9 - Biological Membranes www.gsbs.utmb.edu/ microbook/ch037.htmmycoplasma Membranes form a semi-permeable boundary between a cell and its environment. Membranes also permit subcellular organization
Fluorescent Carbon Dots as Off-On Nanosensor for Ascorbic Acid
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Fluorescent Carbon Dots as Off-On Nanosensor for Ascorbic Acid Jun Gong, Xin Lu, Xueqin An*
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Fluorescent Carbon Dots as Off-On Nanosensor for Ascorbic Acid Jun Gong, Xin Lu, Xueqin An*
Biomembranes structure and function. B. Balen
 Biomembranes structure and function B. Balen All cells are surrounded by membranes Selective barrier But also important for: 1. Compartmentalization 2. Biochemical activities 3. Transport of dissolved
Biomembranes structure and function B. Balen All cells are surrounded by membranes Selective barrier But also important for: 1. Compartmentalization 2. Biochemical activities 3. Transport of dissolved
Photochemical Applications to the Study of Complexity Phospholipid Bilayer Environments
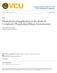 Virginia Commonwealth University VCU Scholars Compass Theses and Dissertations Graduate School 2006 Photochemical Applications to the Study of Complexity Phospholipid Bilayer Environments Christopher John
Virginia Commonwealth University VCU Scholars Compass Theses and Dissertations Graduate School 2006 Photochemical Applications to the Study of Complexity Phospholipid Bilayer Environments Christopher John
The further from the nucleus, the higher the electron s energy Valence shell electrons participate in biological reactions
 Chemistry of Life Revision: The further from the nucleus, the higher the electron s energy Valence shell electrons participate in biological reactions Atoms exchange electrons with other elements to form
Chemistry of Life Revision: The further from the nucleus, the higher the electron s energy Valence shell electrons participate in biological reactions Atoms exchange electrons with other elements to form
Effects of Second Messengers
 Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Protein Trafficking in the Secretory and Endocytic Pathways
 Protein Trafficking in the Secretory and Endocytic Pathways The compartmentalization of eukaryotic cells has considerable functional advantages for the cell, but requires elaborate mechanisms to ensure
Protein Trafficking in the Secretory and Endocytic Pathways The compartmentalization of eukaryotic cells has considerable functional advantages for the cell, but requires elaborate mechanisms to ensure
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
BIOLOGY 103 Spring 2001 MIDTERM LAB SECTION
 BIOLOGY 103 Spring 2001 MIDTERM NAME KEY LAB SECTION ID# (last four digits of SS#) STUDENT PLEASE READ. Do not put yourself at a disadvantage by revealing the content of this exam to your classmates. Your
BIOLOGY 103 Spring 2001 MIDTERM NAME KEY LAB SECTION ID# (last four digits of SS#) STUDENT PLEASE READ. Do not put yourself at a disadvantage by revealing the content of this exam to your classmates. Your
Enrichment of Phospholipids from Biological Matrices with Zirconium Oxide-Modified Silica Sorbents
 Enrichment of Phospholipids from Biological Matrices with Zirconium Oxide-Modified Silica Sorbents Xiaoning Lu, Jennifer E. Claus, and David S. Bell Supelco, Div. of Sigma-Aldrich Bellefonte, PA 16823
Enrichment of Phospholipids from Biological Matrices with Zirconium Oxide-Modified Silica Sorbents Xiaoning Lu, Jennifer E. Claus, and David S. Bell Supelco, Div. of Sigma-Aldrich Bellefonte, PA 16823
Using Infrared and Raman Microspectroscopies to Compare Ex Vivo Involved Psoriatic Skin and Normal Human Skin
 Using Infrared and Raman Microspectroscopies to Compare Ex Vivo Involved Psoriatic Skin and Normal Human Skin Leroy, Marie (Laboratoire d'ingénierie de Surface (LIS)) Lefèvre, Thierry (Groupe de recherche
Using Infrared and Raman Microspectroscopies to Compare Ex Vivo Involved Psoriatic Skin and Normal Human Skin Leroy, Marie (Laboratoire d'ingénierie de Surface (LIS)) Lefèvre, Thierry (Groupe de recherche
The effect of hydrostatic pressure on membrane-bound proteins
 Brazilian Journal of Medical and Biological Research (2005) 38: 1203-1208 High pressure studies on membrane-associated proteins ISSN 0100-879X Review 1203 The effect of hydrostatic pressure on membrane-bound
Brazilian Journal of Medical and Biological Research (2005) 38: 1203-1208 High pressure studies on membrane-associated proteins ISSN 0100-879X Review 1203 The effect of hydrostatic pressure on membrane-bound
Biochemistry - I. Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I
 Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
Measures of Membrane Fluidity: Melting Temperature
 Measures of Membrane Fluidity: Melting Temperature T m (melting temperature) is a phase transition, a change from a more rigid solid-like state to a fluid-like state The fluidity - ease with which lipids
Measures of Membrane Fluidity: Melting Temperature T m (melting temperature) is a phase transition, a change from a more rigid solid-like state to a fluid-like state The fluidity - ease with which lipids
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Membrane Structure. Membrane Structure. Membrane Structure. Membranes
 Membrane Structure Membranes Chapter 5 The fluid mosaic model of membrane structure contends that membranes consist of: -phospholipids arranged in a bilayer -globular proteins inserted in the lipid bilayer
Membrane Structure Membranes Chapter 5 The fluid mosaic model of membrane structure contends that membranes consist of: -phospholipids arranged in a bilayer -globular proteins inserted in the lipid bilayer
Lecture 15. Signal Transduction Pathways - Introduction
 Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
PHSI3009 Frontiers in Cellular Physiology 2017
 Overview of PHSI3009 L2 Cell membrane and Principles of cell communication L3 Signalling via G protein-coupled receptor L4 Calcium Signalling L5 Signalling via Growth Factors L6 Signalling via small G-protein
Overview of PHSI3009 L2 Cell membrane and Principles of cell communication L3 Signalling via G protein-coupled receptor L4 Calcium Signalling L5 Signalling via Growth Factors L6 Signalling via small G-protein
Membranes & Membrane Proteins
 School on Biomolecular Simulations Membranes & Membrane Proteins Vani Vemparala The Institute of Mathematical Sciences Chennai November 13 2007 JNCASR, Bangalore Cellular Environment Plasma membrane extracellular
School on Biomolecular Simulations Membranes & Membrane Proteins Vani Vemparala The Institute of Mathematical Sciences Chennai November 13 2007 JNCASR, Bangalore Cellular Environment Plasma membrane extracellular
1.2 introduction to the cell. me239 mechanics of the cell. 1.2 introduction to the cell. 1.2 introduction to the cell.
 2. introduction to mechanics prokaryotic cells Figure 1.1 Prokaryotic cell. Cell without a nucleus. the inner life of a cell, viel & lue, harvard [2006] me239 mechanics of the cell 1 eukaryotic cells 1.2
2. introduction to mechanics prokaryotic cells Figure 1.1 Prokaryotic cell. Cell without a nucleus. the inner life of a cell, viel & lue, harvard [2006] me239 mechanics of the cell 1 eukaryotic cells 1.2
Early scientists who observed cells made detailed sketches of what they saw.
 Early scientists who observed cells made detailed sketches of what they saw. Early scientists who observed cells made detailed sketches of what they saw. CORK Early scientists who observed cells made detailed
Early scientists who observed cells made detailed sketches of what they saw. Early scientists who observed cells made detailed sketches of what they saw. CORK Early scientists who observed cells made detailed
The electrical properties of ZnO MSM Photodetector with Pt Contact Electrodes on PPC Plastic
 Journal of Electron Devices, Vol. 7, 21, pp. 225-229 JED [ISSN: 1682-3427 ] Journal of Electron Devices www.jeldev.org The electrical properties of ZnO MSM Photodetector with Pt Contact Electrodes on PPC
Journal of Electron Devices, Vol. 7, 21, pp. 225-229 JED [ISSN: 1682-3427 ] Journal of Electron Devices www.jeldev.org The electrical properties of ZnO MSM Photodetector with Pt Contact Electrodes on PPC
Molecular Graphics Perspective of Protein Structure and Function
 Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Biological Membranes. Lipid Membranes. Bilayer Permeability. Common Features of Biological Membranes. A highly selective permeability barrier
 Biological Membranes Structure Function Composition Physicochemical properties Self-assembly Molecular models Lipid Membranes Receptors, detecting the signals from outside: Light Odorant Taste Chemicals
Biological Membranes Structure Function Composition Physicochemical properties Self-assembly Molecular models Lipid Membranes Receptors, detecting the signals from outside: Light Odorant Taste Chemicals
Cholesterol modulates amyloid beta peptide 1-42 channel formation in planar lipid membranes
 Cholesterol modulates amyloid beta peptide 1-42 channel formation in planar lipid membranes Meleleo D., Notarachille G., Gallucci E. and Micelli S. Dept. Farmaco-Biologico, Università degli Studi di Bari,
Cholesterol modulates amyloid beta peptide 1-42 channel formation in planar lipid membranes Meleleo D., Notarachille G., Gallucci E. and Micelli S. Dept. Farmaco-Biologico, Università degli Studi di Bari,
Supplementary Figure 1. Sample preparation schematic. First (Stage I), square islands of MoO 3 are prepared by either photolithography followed by
 Supplementary Figure 1. Sample preparation schematic. First (Stage I), square islands of MoO 3 are prepared by either photolithography followed by thermal evaporation and liftoff or by a process where
Supplementary Figure 1. Sample preparation schematic. First (Stage I), square islands of MoO 3 are prepared by either photolithography followed by thermal evaporation and liftoff or by a process where
Chapter 2 Transport Systems
 Chapter 2 Transport Systems The plasma membrane is a selectively permeable barrier between the cell and the extracellular environment. It permeability properties ensure that essential molecules such as
Chapter 2 Transport Systems The plasma membrane is a selectively permeable barrier between the cell and the extracellular environment. It permeability properties ensure that essential molecules such as
