Supplemental Information for: Localization of anionic phospholipids in Escherichia coli cells
|
|
|
- Russell Morgan
- 5 years ago
- Views:
Transcription
1 Supplemental Information for: Localization of anionic phospholipids in Escherichia coli cells Piercen M. Oliver, a John A. Crooks, a Mathias Leidl, b Earl J. Yoon, a Alan Saghatelian, b and Douglas B. Weibel a,c,d # Department of Biochemistry, University of Wisconsin-Madison, Madison, WI a ; Department of Chemistry and Chemical Biology, Harvard University, Cambridge, MA b ; Department of Chemistry, University of Wisconsin-Madison, Madison, WI c ; Department of Biomedical Engineering, University of Wisconsin-Madison, Madison, WI d #Author to whom correspondence should be addressed Douglas B. Weibel Departments of Biochemistry, Chemistry, and Biomedical Engineering 440 Henry Mall Madison, WI United States Phone: (608) weibel@biochem.wisc.edu 1
2 Figure S1. Fluorescence emission spectra of liposomes mixed with increasing concentrations of NAO. (A, D, G, J, M) Fluorescence versus wavelength spectra of NAO at seven concentrations of CL, PG, PA, PS, and PE, respectively. (B, E, H, K, N) Expanded view of the spectra immediately to the left. (C, F, I, L, O) Spectra (from the left) normalized to a maximum intensity of 1. NAO concentration is 5 µm in 10 mm Tris buffer, ph 8. Excitation is at a wavelength of 456 nm. 2
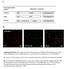
3 Figure S2. Comparison of green and red fluorescence emission of E. coli cells labeled with NAO. (A) Representative images of wild-type (MG1655), clsabc (MG1655 BKT12), pgsa parent (UE53), and pgsa (UE54) E. coli cells labeled with 2 µm NAO and cultured at 30 C to OD = 0.55 ( = 600 nm, see Materials and Methods for details). Green and red channels are shown for each mutant, along with a merged image. The brightness of each channel in the merged images was altered until some yellow color was visible. Scale bars represent 5 µm. (B-D) Averaged intensity profiles of cells labeled with NAO versus the normalized cell length at concentrations of (B) 2 µm, (C) 1 µm, and (D) 0.5 µm NAO. The intensities along the long-axis of each cell were normalized to an area of 1 and averaged (N=166 to 733). The averages in the red channel are shown in red and green in green. The solid black line in (B) shows the average line profile from staining 198 wild-type cells with FM4-64. The shaded space surrounding the NAO fluorescence intensity profiles designates the standard error in the intensity at each point. Septating cells were intentionally left out of the analysis. 3
4 Figure S3. Comparison of green and red fluorescence emission of E. coli cells labeled with NAO from the Keio collection.1 (A) Representative images of wild-type (BW25113), ΔclsA (JW1241), ΔclsB (JW0772), and ΔclsC (JW5150) E. coli cells labeled with 2 µm NAO and cultured at 30 C to OD = 0.55 (λ = 600 nm, see Materials and Methods for details). Green and red channels are shown for each mutant, along with a merged image. The brightness of each channel in the merged images was altered until some yellow color was visible. Scale bars represent 5 µm. (B-D) Averaged intensity profiles of cells labeled with NAO versus the normalized cell length at a concentration of 2 µm NAO. The intensities along the long-axis of each cell were normalized to an area of 1 and averaged (N=246 to 492). The averages in the red channel are shown in red and green in green. The solid black line in (B) shows the average line profile from staining 198 wild-type (MG1655) cells with FM4-64. The shaded space surrounding the NAO fluorescence intensity profiles designates the standard error in the intensity at each point. Septating cells were intentionally left out of the analysis. 1 Baba T, Ara T, Hasegawa M, Takai Y, Okumura Y, Baba M, Datsenko KA, Tomita M, Wanner BL, Mori H Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol 2:
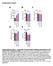
5 Figure S4. Relative abundances of PE, PG, CL, and PA determined by LC-MS (A-C) and TLC/phosphorus analysis (D & E) of minicell producing mutants of (A & D) wild-type (WM1032), (B & E) clsabc (POM10), and (C) pgsa (UEM543) E. coli. To assess differences between the lipid abundances in whole cells and poles in the various E. coli strains, we performed Chisquared tests. All tests were two-sided and statistical significance was considered when a p-value < 0.05 was observed (ns, nonsignificant; *, p < 0.05; **, p < 0.01; ***, p < 0.001). Error bars indicate the standard deviation of the mean (n = 4+ replicates). 5
6 Figure S5. Fractional abundances of lipid species by LC-MS in whole cells and poles of the wild-type (WM1032), clsabc (POM10), and pgsa (UEM543) mutants of E. coli. Lipid species that were analyzed with respect to a corresponding internal standard are presented in this figure: (A) PE, (B) PG, (C) CL, and (D) PA. Each lipid species is presented as the total number of carbons in all alkyl chains on a PL, the total number of unsaturated bonds (after the colon), and whether or not a cyclopropyl group is present (i.e., cp ). Error bars indicate the standard deviation of the mean (n = 4+ replicates). The fractional abundance of each species was calculated by dividing the average standardized intensity of one species by the average standardized summed total of all lipid species for all headgroups (i.e., PEs + PGs + CLs + PAs) for either whole cells or poles. 6
7 Figure S6. Semi-standardized intensities of lipid species by LC-MS in whole cells and poles of the wild-type (WM1032), clsabc (POM10), and pgsa (UEM543) mutants of E. coli. Lipid species that were not analyzed with respect to a corresponding internal standard are presented in this figure: (A) PS, (B) N-acyl-PE, and (C) CDP-DAG. Each lipid species is presented as the total number of carbons in all alkyl chains on a PL and the total number of unsaturated bonds (after the colon). Error bars indicate the standard deviation of the mean (n = 4+ replicates). The intensity of each species was calculated by dividing the average intensity of one species by the average summed total of all standardized lipid species (e.g., PEs + PGs + CLs + PAs) for either whole cells or poles. Since standards for PS, N-acyl-PE, and CDP-DAG were not added, relative comparisons can be made only within one family of lipids. 7
8 Figure S7. Normalized intensities of PE species by LC-MS in whole cells and poles of the A) wild-type (WM1032), B) clsabc (POM10), and C) pgsa (UEM543) mutants of E. coli. Intensities were normalized so that all the intensities of PE species for a particular mutant (whole cells or poles) sum to 1. This comparison is independent of standards and other lipids. To assess differences between the lipid species abundances in whole cells and poles in the various E. coli strains, we performed Chisquared tests. All tests were two-sided and statistical significance was considered when a p-value < 0.05 was observed (ns, nonsignificant; *, p < 0.05; **, p < 0.01; ***, p < 0.001). Error bars indicate the standard deviation of the mean (n = 4+ replicates). 8
9 Figure S8. Normalized intensities of PG species by LC-MS in whole cells and poles of the A) wild-type (WM1032) and B) clsabc (POM10) mutants of E. coli. The pgsa (UEM543) mutant is not shown due to its low measured abundance of PGs. Intensities were normalized so that all the intensities of PG species for a particular mutant (whole cells or poles) sum to 1. This comparison is independent of standards and other lipids. To assess differences between the lipid species abundances in whole cells and poles in the various E. coli strains, we performed Chi-squared tests. All tests were two-sided and statistical significance was considered when a p-value < 0.05 was observed (ns, non-significant; *, p < 0.05; **, p < 0.01; ***, p < 0.001). Error bars indicate the standard deviation of the mean (n = 4+ replicates). 9
10 Figure S9. Normalized intensities of CL species by LC-MS in whole cells and poles of the wild-type (WM1032) E. coli. The clsabc (POM10) and pgsa (UEM543) mutants are not shown due to their low measured abundance of CLs. Intensities were normalized so that all the intensities of CL species for a particular mutant (whole cells or poles) sum to 1. This comparison is independent of standards and other lipids. To assess differences between the lipid species abundances in the whole cells and poles in the E. coli strain, we performed Chi-squared tests. All tests were two-sided and statistical significance was considered when a p-value < 0.05 was observed (ns, non-significant; *, p < 0.05; **, p < 0.01; ***, p < 0.001). Error bars indicate the standard deviation of the mean (n = 4+ replicates). 10
11 Figure S10. Normalized intensities of PA and PS species by LC-MS in whole cells and poles of the A & D) wild-type (WM1032), B & E) clsabc (POM10), and C & F) pgsa (UEM543) mutants of E. coli. Intensities were normalized so that all the intensities of PA or PS species for a particular mutant (whole cells or poles) sum to 1. This comparison is independent of standards and other lipids. To assess differences between the lipid species abundances in whole cells and poles in the various E. coli strains, we performed Chi-squared tests. All tests were two-sided and statistical significance was considered when a p-value < 0.05 was observed (ns, non-significant; *, p < 0.05; **, p < 0.01; ***, p < 0.001). Error bars indicate the standard deviation of the mean (n = 4+ replicates). 11
12 Figure S11. Normalized intensities of N-acyl-PE species by LC-MS in whole cells and poles of the pgsa (UEM543) mutant of E. coli. The wild-type (WM1032) and clsabc (POM10) mutants are not shown due to their low measured abundance of N- acyl-pes. Intensities were normalized so that all the intensities of N-acyl-PE species for a particular mutant (whole cells or poles) sum to 1. This comparison is independent of standards and other lipids. To assess differences between the lipid species abundances in whole cells and poles in the E. coli strain, we performed Chi-squared tests. All tests were two-sided and statistical significance was considered when a p-value < 0.05 was observed (ns, non-significant; *, p < 0.05; **, p < 0.01; ***, p < 0.001). Error bars indicate the standard deviation of the mean (n = 4 replicates). 12
13 Figure S12. Normalized intensities of CDP-DAG species by LC-MS in whole cells and poles of the pgsa (UEM543) mutant of E. coli. The wild-type (WM1032) and clsabc (POM10) mutants are not shown due to their low measured abundance of CDP-DAGs. Intensities were normalized so that all the intensities of CDP-DAG species for a particular mutant (whole cells or poles) sum to 1. This comparison is independent of standards and other lipids. To assess differences between the lipid species abundances in whole cells and poles in the E. coli strain, we performed Chi-squared tests. All tests were two-sided and statistical significance was considered when a p-value < 0.05 was observed (ns, non-significant; *, p < 0.05; **, p < 0.01; ***, p < 0.001). Error bars indicate the standard deviation of the mean (n = 4 replicates). 13
14 Table S1. Semi-standardized total intensities (arbitrary units) of lipid species PS, N-acyl-PE, and CDP-DAG by LC-MS in whole cells and poles of the wild-type (WM1032), clsabc (POM10), and pgsa (UEM543) mutants of E. coli. Each of these total lipid species was normalized to the average summed total of all standardized lipid species (e.g., PEs + PGs + CLs + PAs) for either whole cells or poles. Since standards for PS, N-acyl-PE, and CDP-DAG were not added, comparisons can only be made within one family of lipids. To assess differences between the lipid abundances in whole cells and poles in the various E. coli strains, we performed Chi-squared tests. All tests were two-sided and statistical significance was considered when a p-value < 0.05 was observed (ns, non-significant; *, p < 0.05; **, p < 0.01; ***, p < 0.001). PS N-acyl-PE CDP-DAG Genotype Whole cells p Poles Whole cells p Poles Whole cells p Poles Wild-type 0.12 (±0.09) * 0.2 (±0.1) 1 (±1) 10-3 * 60 (±50) 10-6 n.d. n.d. ΔclsABC 0.04 (±0.02) * 0.10 (±0.03) 0.5 (±0.1) 10-3 n.d. n.d. n.d. ΔpgsA (±0.007) *** (±0.009) 1.42 (±0.06) 10-3 **** 0.65 (±0.06) (±6) 10-3 ** 20 (±12) 10-3 n.d. = none detected 14
15 Table S2. Molecular mass of PE and PG species detected in E. coli. Each lipid species is presented as the total number of carbons in all alkyl chains on a PL, the total number of unsaturated bonds (after the colon), and whether or not a cyclopropyl group is present (i.e., cp ). PE acyl chain PE mass PG acyl chain PG mass compositions Exact Measured compositions Exact Measured Carbons:unsaturations [M-H] - Carbons:unsaturations [M-H] - C28: C30: C29cp* C31: C29: C31: C30: C32: C30: C32: C30: C32: C31: C33: C31: C33: C32: C33: C32: C34: C32: C34: C32: C34: C33: C35: C34: C35: C34: C36: C35: C36: C35cp* C36: C36: C36: C36: C37: * cp designates that one of the fatty acids is cyclopropanated 15
16 Table S3. Molecular mass of CL and PA species detected in E. coli. Each lipid species is presented as the total number of carbons in all alkyl chains on a PL, the total number of unsaturated bonds (after the colon), and whether or not a cyclopropyl group is present (i.e., cp ) CL acyl chain compositions CL mass PA acyl chain PA mass Exact Measured compositions Exact Measured Carbons:unsaturations Most likely structure 2 [M-H] - Carbons:unsaturations [M-H] - C62:1 C16:1/C16:0 C16:0/C14: C30: C62:0 C16:0/C16:0 C16:0/C14: C30: C63:1 C17:1/C16:0 C16:0/C14: C32: C64:2 C16:1/C16:0 C16:1/C16: C32: C64:1 C16:1/C16:0 C16:0/C16: C32: C64:0 C16:0/C16:0 C16:0/C16: C34: C65:2 C17:1/C16:0 C16:1/C16: C34: C65:1 C17:1/C16:0 C16:0/C16: C35cp* C66:3 C18:1/C16:1 C16:1/C16: C36: C66:2 C18:1/C16:0 C16:1/C16: C67:2 C18:1/C16:0 C17:1/C16: C67:1 C18:0/C16:0 C17:1/C16: C68:2 C18:1/C16:0 C18:1/C16: C68:0 C18:0/C16:0 C18:0/C16: C69:2 C18:0/C16:1 C18:0/C17: C70:3 C18:1/C16:0 C18:1/C18: C70:2 C18:1/C16:1 C18:0/C18: * cp designates that one of the fatty acids is cyclopropanated 2 Hsu, F. F., & Turk, J. (2006). Characterization of cardiolipin from Escherichia coli by electrospray ionization with multiple stage quadrupole ion-trap mass spectrometric analysis of [M 2H+Na] ions. Journal of the American Society for Mass Spectrometry, 17(3), doi: /j.jasms
17 Table S4. Molecular mass of PS and N-acyl-PE species detected in E. coli. Each lipid species is presented as the total number of carbons in all alkyl chains on a PL and the total number of unsaturated bonds (after the colon). PS acyl chain PS mass N-acyl-PE acyl chain N-acyl-PE mass compositions Exact Measured compositions 3 Exact Measured Carbons:unsaturations [M-H] - Carbons:unsaturations [M-H] - C34: C46: C34: C48: C36: C48: C36: C49: C36: C49: C38: C50: C50: C50: C52: C52: C54: n.d. 3 Mileykovskaya, E., Ryan, A., Mo, X., Lin, C., Khalaf, K., Dowhan, W., & Garrett, T. (2009). Phosphatidic acid and N- acylphosphatidylethanolamine form membrane domains in Escherichia coli mutant lacking cardiolipin and phosphatidylglycerol. The Journal of Biological Chemistry, 284(5),
18 Table S5. Molecular mass of CDP-DAG species detected in E. coli. Each lipid species is presented as the total number of carbons in all alkyl chains on a PL and the total number of unsaturated bonds (after the colon). CDP-DAG acyl chain CDP-DAG mass compositions Exact Measured Carbons:unsaturations [M-H] - C28: C30: n.d. C30: C30: C31: C31: n.d. C32: C32: C32: C33: C33: n.d. C33: C34: n.d. C34: C34: C35: C35: n.d. C36: C36: C36: C36: C36: C36: C38: C38: n.d. C38: C38: n.d. C38: n.d. 18
Table S1. Strains, Plasmids and Phage
 Table S1. Strains, Plasmids and Phage Strain, Plasmid, Phage Genotype or Description Reference S. enterica 14028 wild type ATCC strain (1) HK132 yfin; 14028 with deletion of yfin This study QW262 yhjh
Table S1. Strains, Plasmids and Phage Strain, Plasmid, Phage Genotype or Description Reference S. enterica 14028 wild type ATCC strain (1) HK132 yfin; 14028 with deletion of yfin This study QW262 yhjh
Mass Spectrometry based metabolomics
 Mass Spectrometry based metabolomics Metabolomics- A realm of small molecules (
Mass Spectrometry based metabolomics Metabolomics- A realm of small molecules (
Protein-Lipid Interactions: Structural and Functional Effects Anthony Lee (Southampton)
 Saulieu ctober 2004 Protein-Lipid Interactions: Structural and Functional Effects Anthony Lee (Southampton) The membrane as a system Co-evolution of lipids and membrane proteins R P - R Phosphatidylcholine
Saulieu ctober 2004 Protein-Lipid Interactions: Structural and Functional Effects Anthony Lee (Southampton) The membrane as a system Co-evolution of lipids and membrane proteins R P - R Phosphatidylcholine
The Mycobacterium tuberculosis MmpL11 cell wall lipid transporter is important for
 Supplemental materials The Mycobacterium tuberculosis MmpL11 cell wall lipid transporter is important for biofilm formation, intracellular growth and non-replicating persistence Catherine C. Wright, 1
Supplemental materials The Mycobacterium tuberculosis MmpL11 cell wall lipid transporter is important for biofilm formation, intracellular growth and non-replicating persistence Catherine C. Wright, 1
Loss of protein association causes cardiolipin degradation in Barth syndrome
 SUPPLEMENTARY INFORMATION Loss of protein association causes cardiolipin degradation in Barth syndrome Yang Xu 1, Colin K.L. Phoon 2, Bob Berno 5, Kenneth D Souza 6, Esthelle Hoedt 4, Guoan Zhang 4, Thomas
SUPPLEMENTARY INFORMATION Loss of protein association causes cardiolipin degradation in Barth syndrome Yang Xu 1, Colin K.L. Phoon 2, Bob Berno 5, Kenneth D Souza 6, Esthelle Hoedt 4, Guoan Zhang 4, Thomas
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
(03) WMP/Jun13/BIOL1
 3 Do not write outside the box Answer all questions in the spaces provided. 1 The diagram shows the structure of the cell-surface membrane of a cell. C B A 1 (a) Name A and B. A... B... (2 marks) 1 (b)
3 Do not write outside the box Answer all questions in the spaces provided. 1 The diagram shows the structure of the cell-surface membrane of a cell. C B A 1 (a) Name A and B. A... B... (2 marks) 1 (b)
SUPPLEMENTARY DATA. Materials and Methods
 SUPPLEMENTARY DATA Materials and Methods HPLC-UV of phospholipid classes and HETE isomer determination. Fractionation of platelet lipid classes was undertaken on a Spherisorb S5W 150 x 4.6 mm column (Waters
SUPPLEMENTARY DATA Materials and Methods HPLC-UV of phospholipid classes and HETE isomer determination. Fractionation of platelet lipid classes was undertaken on a Spherisorb S5W 150 x 4.6 mm column (Waters
A highly selective AIE fluorogen for lipid droplet imaging in live cells and green algae
 Electronic Supporting Information highly selective IE fluorogen for lipid droplet imaging in live cells and green algae Erjing Wang, ab Engui Zhao, ab Yuning Hong, ab Jacky W. Y. Lam, ab and en Zhong Tang*
Electronic Supporting Information highly selective IE fluorogen for lipid droplet imaging in live cells and green algae Erjing Wang, ab Engui Zhao, ab Yuning Hong, ab Jacky W. Y. Lam, ab and en Zhong Tang*
ALLOSTERIC REGULATION OF GPCR ACTIVITY BY PHOSPHOLIPIDS
 Supplementary Information ALLOSTERIC REGULATION OF GPCR ACTIVITY BY PHOSPHOLIPIDS Rosie Dawaliby 1, Cataldo Trubbia 1, Cédric Delporte 3,4, Matthieu Masureel 2, Pierre Van Antwerpen 3,4, Brian K. Kobilka
Supplementary Information ALLOSTERIC REGULATION OF GPCR ACTIVITY BY PHOSPHOLIPIDS Rosie Dawaliby 1, Cataldo Trubbia 1, Cédric Delporte 3,4, Matthieu Masureel 2, Pierre Van Antwerpen 3,4, Brian K. Kobilka
Impact of Chromatography on Lipid Profiling of Liver Tissue Extracts
 Impact of Chromatography on Lipid Profiling of Liver Tissue Extracts Application Note Clinical Research Authors Mark Sartain and Theodore Sana Agilent Technologies, Inc. Santa Clara, California, USA Introduction
Impact of Chromatography on Lipid Profiling of Liver Tissue Extracts Application Note Clinical Research Authors Mark Sartain and Theodore Sana Agilent Technologies, Inc. Santa Clara, California, USA Introduction
Supporting Information
 Supporting Information Mass Spectrometry Imaging Shows Cocaine and Methylphenidate have Opposite Effects on Major Lipids in Drosophila Brain Mai H. Philipsen *, Nhu T. N. Phan *, John S. Fletcher *, Per
Supporting Information Mass Spectrometry Imaging Shows Cocaine and Methylphenidate have Opposite Effects on Major Lipids in Drosophila Brain Mai H. Philipsen *, Nhu T. N. Phan *, John S. Fletcher *, Per
Physical effects underlying the transition from primitive to modern cell membranes
 Physical effects underlying the transition from primitive to modern cell membranes Itay Budin and Jack W. Szostak* *To whom correspondence should be addressed. Email: szostak@molbio.mgh.harvard.edu This
Physical effects underlying the transition from primitive to modern cell membranes Itay Budin and Jack W. Szostak* *To whom correspondence should be addressed. Email: szostak@molbio.mgh.harvard.edu This
7/11/17. Cell Function & Chemistry. Molecular and Cellular Biology. 2. Bio-Chemical Foundations & Key Molecules of a Cell
 Molecular and Cellular Biology Cell Function & Chemistry 2. Bio-Chemical Foundations & Key Molecules of a Cell Prof. Dr. Klaus Heese Interaction Molecular Bonds Define Cellular Functions Water H 2 O Interactions
Molecular and Cellular Biology Cell Function & Chemistry 2. Bio-Chemical Foundations & Key Molecules of a Cell Prof. Dr. Klaus Heese Interaction Molecular Bonds Define Cellular Functions Water H 2 O Interactions
Development of a near-infrared fluorescent probe for monitoring hydrazine in serum and living cells
 Supporting Information for Development of a near-infrared fluorescent probe for monitoring hydrazine in serum and living cells Sasa Zhu, Weiying Lin,* Lin Yuan State Key Laboratory of Chemo/Biosensing
Supporting Information for Development of a near-infrared fluorescent probe for monitoring hydrazine in serum and living cells Sasa Zhu, Weiying Lin,* Lin Yuan State Key Laboratory of Chemo/Biosensing
The use of mass spectrometry in lipidomics. Outlines
 The use of mass spectrometry in lipidomics Jeevan Prasain jprasain@uab.edu 6-2612 utlines Brief introduction to lipidomics Analytical methodology: MS/MS structure elucidation of phospholipids Phospholipid
The use of mass spectrometry in lipidomics Jeevan Prasain jprasain@uab.edu 6-2612 utlines Brief introduction to lipidomics Analytical methodology: MS/MS structure elucidation of phospholipids Phospholipid
Supporting Information. Kinetic and Conformational Insights into Islet Amyloid Polypeptide Self-Assembly using a Biarsenical Fluorogenic Probe
 Kinetic and Conformational Insights into Islet Amyloid Polypeptide Self-Assembly using a Biarsenical Fluorogenic Probe Noé Quittot, Mathew Sebastiao, Soultan Al-Halifa and Steve Bourgault* Department of
Kinetic and Conformational Insights into Islet Amyloid Polypeptide Self-Assembly using a Biarsenical Fluorogenic Probe Noé Quittot, Mathew Sebastiao, Soultan Al-Halifa and Steve Bourgault* Department of
Interactions of Polyethylenimines with Zwitterionic and. Anionic Lipid Membranes
 Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
UNIVERSITY OF CALGARY. Interactions of Inorganic Mercury and Inorganic Cadmium with Biomimetic and Complex
 UNIVERSITY OF CALGARY Interactions of Inorganic Mercury and Inorganic Cadmium with Biomimetic and Complex Biological Membranes and their Influence on Membrane Packing and Size by Evan Kerek A THESIS SUBMITTED
UNIVERSITY OF CALGARY Interactions of Inorganic Mercury and Inorganic Cadmium with Biomimetic and Complex Biological Membranes and their Influence on Membrane Packing and Size by Evan Kerek A THESIS SUBMITTED
Achieve Broad Lipid Quantitation using a High-Throughput Targeted Lipidomics Method
 Achieve Broad Lipid Quantitation using a High-Throughput Targeted Lipidomics Method LC-Based Approach for Lipid Class Separation and Quantitation on QTRAP 6500+ System Mackenzie Pearson, Santosh Kumar
Achieve Broad Lipid Quantitation using a High-Throughput Targeted Lipidomics Method LC-Based Approach for Lipid Class Separation and Quantitation on QTRAP 6500+ System Mackenzie Pearson, Santosh Kumar
MASS SPECTROMETRY STRATEGIES FOR COMPREHENSIVE LIPIDOME ANALYSIS OF COLORECTAL CANCER CELLS AND THEIR SECRETED EXOSOMES. Cassie J.
 MASS SPECTROMETRY STRATEGIES FOR COMPREHENSIVE LIPIDOME ANALYSIS OF COLORECTAL CANCER CELLS AND THEIR SECRETED EXOSOMES By Cassie J. Fhaner A DISSERTATION Submitted to Michigan State University in partial
MASS SPECTROMETRY STRATEGIES FOR COMPREHENSIVE LIPIDOME ANALYSIS OF COLORECTAL CANCER CELLS AND THEIR SECRETED EXOSOMES By Cassie J. Fhaner A DISSERTATION Submitted to Michigan State University in partial
3.1.3 Lipids. Source: AQA Spec
 alevelbiology.co.uk SPECIFICATION Triglycerides and phospholipids are two groups of lipid. Triglycerides are formed by the condensation of one molecule of glycerol and three molecules of fatty acid. A
alevelbiology.co.uk SPECIFICATION Triglycerides and phospholipids are two groups of lipid. Triglycerides are formed by the condensation of one molecule of glycerol and three molecules of fatty acid. A
Glycerolipid Analysis. LC/MS/MS Analytical Services
 Glycerolipid Analysis LC/MS/MS Analytical Services Molecular Characterization and Quantitation of Glycerophospholipids in Commercial Lecithins by High Performance Liquid Chromatography with Mass Spectrometric
Glycerolipid Analysis LC/MS/MS Analytical Services Molecular Characterization and Quantitation of Glycerophospholipids in Commercial Lecithins by High Performance Liquid Chromatography with Mass Spectrometric
SUPPLEMENTAL DATA AGING, April 2013, Vol.5 No.4
 SUPPLEMENTAL DATA Figure S1. Under CR conditions, the atg32δ mutation elevates the extent of oxidative damage to proteins. WT and atg32δ strains were cultured in the nutrient rich YP medium initially containing
SUPPLEMENTAL DATA Figure S1. Under CR conditions, the atg32δ mutation elevates the extent of oxidative damage to proteins. WT and atg32δ strains were cultured in the nutrient rich YP medium initially containing
Received: 21 September 2009 / Revised: 2 November 2009 / Accepted: 3 November 2009 / Published online: 25 November 2009 # Springer-Verlag 2009
 Anal Bioanal Chem (2010) 396:1273 1280 DOI 10.1007/s00216-009-3292-9 ORIGINAL PAPER Quantitative analysis of urinary phospholipids found in patients with breast cancer by nanoflow liquid chromatography
Anal Bioanal Chem (2010) 396:1273 1280 DOI 10.1007/s00216-009-3292-9 ORIGINAL PAPER Quantitative analysis of urinary phospholipids found in patients with breast cancer by nanoflow liquid chromatography
Supporting Information
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2018 Copyright WILEY-VCH Verlag GmbH & Co. KGaA, 69469 Weinheim, Germany, 2016. Supporting Information
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2018 Copyright WILEY-VCH Verlag GmbH & Co. KGaA, 69469 Weinheim, Germany, 2016. Supporting Information
Lipid metabolism of phenol tolerant Rhodococcus opacus strains for lignin bioconversion
 https://doi.org/10.1186/s13068-018-1337-z Biotechnology for Biofuels RESEARCH Open Access Lipid metabolism of phenol tolerant Rhodococcus opacus strains for lignin bioconversion William R. Henson 1, Fong
https://doi.org/10.1186/s13068-018-1337-z Biotechnology for Biofuels RESEARCH Open Access Lipid metabolism of phenol tolerant Rhodococcus opacus strains for lignin bioconversion William R. Henson 1, Fong
ABSTRACT INTRODUCTION
 /, 2017, Vol. 8, (No. 19), pp: 30672-30691 Specific changes in mitochondrial lipidome alter mitochondrial proteome and increase the geroprotective efficiency of lithocholic acid in chronologically aging
/, 2017, Vol. 8, (No. 19), pp: 30672-30691 Specific changes in mitochondrial lipidome alter mitochondrial proteome and increase the geroprotective efficiency of lithocholic acid in chronologically aging
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry. Supporting Information
 Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Engineering a polarity-sensitive biosensor for time-lapse imaging of apoptotic processes and degeneration
 nature methods Engineering a polarity-sensitive biosensor for time-lapse imaging of apoptotic processes and degeneration Yujin E Kim, Jeannie Chen, Jonah R Chan & Ralf Langen Supplementary figures and
nature methods Engineering a polarity-sensitive biosensor for time-lapse imaging of apoptotic processes and degeneration Yujin E Kim, Jeannie Chen, Jonah R Chan & Ralf Langen Supplementary figures and
1. Methodology of lipidomics: Tandem mass spectrometry of ether phospholipids
 Transworld Research Network 37/661 (2), Fort P.O. Trivandrum-695 023 Kerala, India Lipidomics: Sea Food, Marine Based Dietary Supplement, Fruit and Seed, 2012: 1-19 ISBN: 978-81-7895-573-5 Editor: 1. Methodology
Transworld Research Network 37/661 (2), Fort P.O. Trivandrum-695 023 Kerala, India Lipidomics: Sea Food, Marine Based Dietary Supplement, Fruit and Seed, 2012: 1-19 ISBN: 978-81-7895-573-5 Editor: 1. Methodology
Ozonolysis of phospholipid double bonds during electrospray. ionization: a new tool for structure determination
 Ozonolysis of phospholipid double bonds during electrospray ionization: a new tool for structure determination Michael C. Thomas, Todd W. Mitchell, Stephen J. Blanksby Departments of Chemistry and Biomedical
Ozonolysis of phospholipid double bonds during electrospray ionization: a new tool for structure determination Michael C. Thomas, Todd W. Mitchell, Stephen J. Blanksby Departments of Chemistry and Biomedical
Graphene Quantum Dots-Band-Aids Used for Wound Disinfection
 Supporting information Graphene Quantum Dots-Band-Aids Used for Wound Disinfection Hanjun Sun, Nan Gao, Kai Dong, Jinsong Ren, and Xiaogang Qu* Laboratory of Chemical Biology, Division of Biological Inorganic
Supporting information Graphene Quantum Dots-Band-Aids Used for Wound Disinfection Hanjun Sun, Nan Gao, Kai Dong, Jinsong Ren, and Xiaogang Qu* Laboratory of Chemical Biology, Division of Biological Inorganic
Importance of Direct Interactions with Lipids for the Function of the Mechanosensitive Channel MscL
 Biochemistry 2008, 47, 12175 12184 12175 Importance of Direct Interactions with Lipids for the Function of the Mechanosensitive Channel MscL Andrew M. Powl, J. Malcolm East, and Anthony G. Lee* School
Biochemistry 2008, 47, 12175 12184 12175 Importance of Direct Interactions with Lipids for the Function of the Mechanosensitive Channel MscL Andrew M. Powl, J. Malcolm East, and Anthony G. Lee* School
ANSC (NUTR) 618 LIPIDS & LIPID METABOLISM Membrane Lipids and Sphingolipidsd
 ANSC (NUTR) 618 LIPIDS & LIPID METABOLISM Membrane Lipids and Sphingolipidsd I. Classes of membrane lipids A. Glycerolipids (quantitatively the most important of the three membrane lipids) B. Shingolipids
ANSC (NUTR) 618 LIPIDS & LIPID METABOLISM Membrane Lipids and Sphingolipidsd I. Classes of membrane lipids A. Glycerolipids (quantitatively the most important of the three membrane lipids) B. Shingolipids
Supporting Information
 Supporting Information Toward High-Efficient Red Emissive Carbon Dots: Facile Preparation, Unique Properties, and Applications as Multifunctional Theranostic Agents Shan Sun,, Ling Zhang, Kai Jiang, Aiguo
Supporting Information Toward High-Efficient Red Emissive Carbon Dots: Facile Preparation, Unique Properties, and Applications as Multifunctional Theranostic Agents Shan Sun,, Ling Zhang, Kai Jiang, Aiguo
Poly-unsaturated phospholipids WE (FA18:1) A B C D OAHFA 18:1/24:0 OAHFA 18:1/30:2 PS 38:4 C18:1C23:0 OAHFA 18:1/25:0 OAHFA 18:1/30:1
 Figure S1, related to Figure 1. Correlation scatter plots illustrating individual lipid species from various classes that exhibited statistically significant correlations to age. (A-B) Various species
Figure S1, related to Figure 1. Correlation scatter plots illustrating individual lipid species from various classes that exhibited statistically significant correlations to age. (A-B) Various species
A mutant in Arabidopsis Lacking a Chloroplast Specific Lipid. Lewis Kurschner and Karen Thulasi Masters in Botany
 A mutant in Arabidopsis Lacking a Chloroplast Specific Lipid Lewis Kurschner and Karen Thulasi Masters in Botany Fatty acid nomenclature Fatty acyl composition Chain length Degree of unsaturation and position
A mutant in Arabidopsis Lacking a Chloroplast Specific Lipid Lewis Kurschner and Karen Thulasi Masters in Botany Fatty acid nomenclature Fatty acyl composition Chain length Degree of unsaturation and position
ASSESSMENT OF CELLULAR OXYGEN GRADIENTS WITH A PANEL OF PHOSPHORESCENT OXYGEN-SENSITIVE PROBES
 ASSESSMENT OF CELLULAR OXYGEN GRADIENTS WITH A PANEL OF PHOSPHORESCENT OXYGEN-SENSITIVE PROBES Ruslan I. Dmitriev, Alexander V. Zhdanov, Greg Jasionek, Dmitri B. Papkovsky Biochemistry Department, University
ASSESSMENT OF CELLULAR OXYGEN GRADIENTS WITH A PANEL OF PHOSPHORESCENT OXYGEN-SENSITIVE PROBES Ruslan I. Dmitriev, Alexander V. Zhdanov, Greg Jasionek, Dmitri B. Papkovsky Biochemistry Department, University
Zn 2+ Triggered Amide Tautomerization Produces a Highly Zn 2+ Selective, Cell Permeable and Ratiometric Fluorescent Sensor
 Supporting Information For Zn 2+ Triggered Amide Tautomerization Produces a Highly Zn 2+ Selective, Cell Permeable and Ratiometric Fluorescent Sensor Zhaochao Xu,*,, Kyung-Hwa Baek, Ha Na Kim, Jingnan
Supporting Information For Zn 2+ Triggered Amide Tautomerization Produces a Highly Zn 2+ Selective, Cell Permeable and Ratiometric Fluorescent Sensor Zhaochao Xu,*,, Kyung-Hwa Baek, Ha Na Kim, Jingnan
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB
 Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Desorption Electrospray Ionization Coupled with Ultraviolet Photodissociation for Characterization of Phospholipid Isomers in Tissue Sections
 Desorption Electrospray Ionization Coupled with Ultraviolet Photodissociation for Characterization of Phospholipid Isomers in Tissue Sections Dustin R. Klein, Clara L. Feider, Kyana Y. Garza, John Q. Lin,
Desorption Electrospray Ionization Coupled with Ultraviolet Photodissociation for Characterization of Phospholipid Isomers in Tissue Sections Dustin R. Klein, Clara L. Feider, Kyana Y. Garza, John Q. Lin,
Phospholipids Metabolism
 Chapter VI: Phospholipids Metabolism Dr. Sameh Sarray Hlaoui Phospholipids Features: Amphipatic: - Hydrophobic head: fatty acids - Hydropholic head: P group+ alcohol Composed of alcohol attached by a phosphodiester
Chapter VI: Phospholipids Metabolism Dr. Sameh Sarray Hlaoui Phospholipids Features: Amphipatic: - Hydrophobic head: fatty acids - Hydropholic head: P group+ alcohol Composed of alcohol attached by a phosphodiester
Supplementary Figure 1. Overview of steps in the construction of photosynthetic protocellular systems
 Supplementary Figure 1 Overview of steps in the construction of photosynthetic protocellular systems (a) The small unilamellar vesicles were made with phospholipids. (b) Three types of small proteoliposomes
Supplementary Figure 1 Overview of steps in the construction of photosynthetic protocellular systems (a) The small unilamellar vesicles were made with phospholipids. (b) Three types of small proteoliposomes
Thermo Scientific LipidSearch Software for Lipidomics Workflows. Automated Identification and Relative. Quantitation of Lipids by LC/MS
 Thermo Scientific LipidSearch Software for Lipidomics Workflows Automated Identification and Relative of Lipids by LC/MS The promise of lipidomics Lipidomics is a new field of study crucial for understanding
Thermo Scientific LipidSearch Software for Lipidomics Workflows Automated Identification and Relative of Lipids by LC/MS The promise of lipidomics Lipidomics is a new field of study crucial for understanding
Nature Biotechnology: doi: /nbt.3828
 Supplementary Figure 1 Development of a FRET-based MCS. (a) Linker and MA2 modification are indicated by single letter amino acid code. indicates deletion of amino acids and N or C indicate the terminus
Supplementary Figure 1 Development of a FRET-based MCS. (a) Linker and MA2 modification are indicated by single letter amino acid code. indicates deletion of amino acids and N or C indicate the terminus
Supplementary Information: A Critical. Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers
 Supplementary Information: A Critical Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers Kristyna Pluhackova,, Sonja A. Kirsch, Jing Han, Liping Sun,
Supplementary Information: A Critical Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers Kristyna Pluhackova,, Sonja A. Kirsch, Jing Han, Liping Sun,
Supplementary Figure 1: UV-Vis absorption (a) and excitation emission matrix (EEM) (b) spectra of the SN medium (Cyanosite ) used to grow
 Supplementary Figure 1: UV-Vis absorption (a) and excitation emission matrix (EEM) (b) spectra of the SN medium (Cyanosite ) used to grow Synechococcus cultures. Note: The same scale used in Figure 1 was
Supplementary Figure 1: UV-Vis absorption (a) and excitation emission matrix (EEM) (b) spectra of the SN medium (Cyanosite ) used to grow Synechococcus cultures. Note: The same scale used in Figure 1 was
High-Resolution Analysis of Intact Triglycerides by Reversed Phase HPLC Using the Agilent 1290 Infinity LC UHPLC System
 High-Resolution Analysis of Intact Triglycerides by Reversed Phase HPLC Using the Agilent 1290 Infinity LC UHPLC System Application Note Food, Hydrocarbon Processing Authors Michael Woodman Agilent Technologies,
High-Resolution Analysis of Intact Triglycerides by Reversed Phase HPLC Using the Agilent 1290 Infinity LC UHPLC System Application Note Food, Hydrocarbon Processing Authors Michael Woodman Agilent Technologies,
Comparison of PCB analysis by GC Electron Capture Detection and GC-MS Selective Ion Monitoring Analytical Methods
 Comparison of PCB analysis by GC Electron Capture Detection and GC-MS Selective Ion Monitoring Analytical Methods Robert K. Johnston, Ronald Gauthier, William Wild, Jr. Space and Naval Warfare Systems
Comparison of PCB analysis by GC Electron Capture Detection and GC-MS Selective Ion Monitoring Analytical Methods Robert K. Johnston, Ronald Gauthier, William Wild, Jr. Space and Naval Warfare Systems
Imaging of lipid species by MALDI mass spectrometry
 Imaging of lipid species by MALDI mass spectrometry Robert C. Murphy, 1 Joseph A. Hankin, and Robert M. Barkley Department of Pharmacology, MS 8303 University of Colorado Denver, Aurora, CO 80045 Abstract
Imaging of lipid species by MALDI mass spectrometry Robert C. Murphy, 1 Joseph A. Hankin, and Robert M. Barkley Department of Pharmacology, MS 8303 University of Colorado Denver, Aurora, CO 80045 Abstract
Supporting Information
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2015 Supporting Information A new ICT and CHEF based visible light excitable fluorescent probe easily
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2015 Supporting Information A new ICT and CHEF based visible light excitable fluorescent probe easily
Lipid Mass Spectrometry: Core Technologic Research and Development I.) INTRODUCTION TO MASS SPECTROMETRY OF COMPLEX LIPIDS 4
 Lipid Mass Spectrometry: Core Technologic Research and Development PAGE I.) INTRODUCTION TO MASS SPECTROMETRY OF COMPLEX LIPIDS 4 II.) MASS SPECTROMETRY OF GLYCEROLIPIDS.. 8 A.) Neutral Acylglycerols and
Lipid Mass Spectrometry: Core Technologic Research and Development PAGE I.) INTRODUCTION TO MASS SPECTROMETRY OF COMPLEX LIPIDS 4 II.) MASS SPECTROMETRY OF GLYCEROLIPIDS.. 8 A.) Neutral Acylglycerols and
Core E Analysis of Neutral Lipids from Human Plasma June 4, 2010 Thomas J. Leiker and Robert M. Barkley
 Core E Analysis of Neutral Lipids from Human Plasma June 4, 2010 Thomas J. Leiker and Robert M. Barkley This protocol describes the extraction and direct measurement of cholesterol esters (CEs) and triacylglycerols
Core E Analysis of Neutral Lipids from Human Plasma June 4, 2010 Thomas J. Leiker and Robert M. Barkley This protocol describes the extraction and direct measurement of cholesterol esters (CEs) and triacylglycerols
The detergent-solubilized and gel filtration purified rhodopsin was partitioned against
 Supplement Jastrzebska et al. Materials and Methods The detergent-solubilized and gel filtration purified rhodopsin was partitioned against H 2 O/MeOH/CHCl 3, and the bottom layer was removed, dried down,
Supplement Jastrzebska et al. Materials and Methods The detergent-solubilized and gel filtration purified rhodopsin was partitioned against H 2 O/MeOH/CHCl 3, and the bottom layer was removed, dried down,
MEMBRANE LIPIDS I and II: GLYCEROPHOSPHOLIPIDS AND SPHINGOLIPIDS
 December 6, 2011 Lecturer: Eileen M. Lafer MEMBRANE LIPIDS I and II: GLYCEROPHOSPHOLIPIDS AND SPHINGOLIPIDS Reading: Stryer Edition 6: Chapter 26 Images: All images in these notes were taken from Lehninger,
December 6, 2011 Lecturer: Eileen M. Lafer MEMBRANE LIPIDS I and II: GLYCEROPHOSPHOLIPIDS AND SPHINGOLIPIDS Reading: Stryer Edition 6: Chapter 26 Images: All images in these notes were taken from Lehninger,
Synthesis and elongation of fatty acids
 Synthesis and elongation of fatty acids A molecular caliper mechanism for determining very long-chain fatty acid length Vladimir Denic and Jonathan S. Weissman (2007) Cell 130, 663-677 February 28, 2008
Synthesis and elongation of fatty acids A molecular caliper mechanism for determining very long-chain fatty acid length Vladimir Denic and Jonathan S. Weissman (2007) Cell 130, 663-677 February 28, 2008
Molecular and Cellular Biology. 2. Bio-Chemical Foundations & Key Molecules of a Cell
 Molecular and Cellular Biology 2. Bio-Chemical Foundations & Key Molecules of a Cell Prof. Dr. Klaus Heese Cell Function & Chemistry Interaction 1 Molecular Bonds Define Cellular Functions Interactions
Molecular and Cellular Biology 2. Bio-Chemical Foundations & Key Molecules of a Cell Prof. Dr. Klaus Heese Cell Function & Chemistry Interaction 1 Molecular Bonds Define Cellular Functions Interactions
Babu Antharavally, Ryan Bomgarden, and John Rogers Thermo Fisher Scientific, Rockford, IL
 A Versatile High-Recovery Method for Removing Detergents from Low-Concentration Protein or Peptide Samples for Mass Spectrometry Sample Preparation and Analysis Babu Antharavally, Ryan Bomgarden, and John
A Versatile High-Recovery Method for Removing Detergents from Low-Concentration Protein or Peptide Samples for Mass Spectrometry Sample Preparation and Analysis Babu Antharavally, Ryan Bomgarden, and John
Quantifying Lipid Contents in Enveloped Virus Particles with Plasmonic Nanoparticles
 Quantifying Lipid Contents in Enveloped Virus Particles with Plasmonic Nanoparticles Amin Feizpour Reinhard Lab Department of Chemistry and the Photonics Center, Boston University, Boston, MA May 2014
Quantifying Lipid Contents in Enveloped Virus Particles with Plasmonic Nanoparticles Amin Feizpour Reinhard Lab Department of Chemistry and the Photonics Center, Boston University, Boston, MA May 2014
LC/MS/MS SOLUTIONS FOR LIPIDOMICS. Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS
 LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
Principles of Shotgun Lipidomics
 Principles of Shotgun Lipidomics Xianlin Han Diabetes and Obesity Research Center Sanford-Burnham Medical Research Institute Lake Nona Orlando, FL 32827 What is shotgun lipidomics? Original definition
Principles of Shotgun Lipidomics Xianlin Han Diabetes and Obesity Research Center Sanford-Burnham Medical Research Institute Lake Nona Orlando, FL 32827 What is shotgun lipidomics? Original definition
Supporting Information
 Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany Tissue Imaging at Atmospheric Pressure using Desorption Electrospray Ionization (DESI) Mass Spectrometry Justin M. Wiseman, Demian R. Ifa,
Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany Tissue Imaging at Atmospheric Pressure using Desorption Electrospray Ionization (DESI) Mass Spectrometry Justin M. Wiseman, Demian R. Ifa,
Supporting information
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2014 Supporting information Seeing the Diabetes: Visual Detection of Glucose Based on the Intrinsic
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2014 Supporting information Seeing the Diabetes: Visual Detection of Glucose Based on the Intrinsic
SUPPLEMENTARY INFORMATION
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2919 Direct observation of the influence of cardiolipin and antibiotics on lipid II binding to MurJ Jani
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2919 Direct observation of the influence of cardiolipin and antibiotics on lipid II binding to MurJ Jani
Suppl. Table 1: CV of pooled lipoprotein fractions analysed by ESI-MS/MS
 Supplement VLDL LDL HDL PC 3.3 1.77 1.3 LPC 4.82 2.5.35 SM 3.1 4.6 1.92 CER 2.17 6.3 4.15 PE 3.18 1.93 2.79 PE-pl 13.18 1.9 2.32 CE 2.9.65.4 FC.36 3.5 2.54 Suppl. Table 1: CV of pooled lipoprotein fractions
Supplement VLDL LDL HDL PC 3.3 1.77 1.3 LPC 4.82 2.5.35 SM 3.1 4.6 1.92 CER 2.17 6.3 4.15 PE 3.18 1.93 2.79 PE-pl 13.18 1.9 2.32 CE 2.9.65.4 FC.36 3.5 2.54 Suppl. Table 1: CV of pooled lipoprotein fractions
Photo-reduction and Stabilization Capability of Molecular Weight. Fractionated Natural Organic Matter in Transformation of Silver Ion to
 Supporting information Photo-reduction and Stabilization Capability of Molecular Weight Fractionated Natural Organic Matter in Transformation of Silver Ion to Metallic Nanoparticle 7 8 9 Yongguang Yin
Supporting information Photo-reduction and Stabilization Capability of Molecular Weight Fractionated Natural Organic Matter in Transformation of Silver Ion to Metallic Nanoparticle 7 8 9 Yongguang Yin
Supplementary Figure 1. Structure models of the c4 variant proteins based on the structures of maltose binding protein (MBP) in red and TEM-1
 Supplementary Figure 1. Structure models of the c4 variant proteins based on the structures of maltose binding protein (MBP) in red and TEM-1 β-lactamase (BLA) in blue. (a) c4 model and the close-up view
Supplementary Figure 1. Structure models of the c4 variant proteins based on the structures of maltose binding protein (MBP) in red and TEM-1 β-lactamase (BLA) in blue. (a) c4 model and the close-up view
Supporting Information For
 Supporting Information For MicroRNA-Catalyzed Cancer Therapeutics Based on DNA-Programmed Nanoparticle Complex Xucheng Luo, 1 Zhi Li, 1 Ganglin Wang, 1 Xuewen He, 2,3 Xiaoqin Shen, 1 Quanhong Sun, 1 Li
Supporting Information For MicroRNA-Catalyzed Cancer Therapeutics Based on DNA-Programmed Nanoparticle Complex Xucheng Luo, 1 Zhi Li, 1 Ganglin Wang, 1 Xuewen He, 2,3 Xiaoqin Shen, 1 Quanhong Sun, 1 Li
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors
 Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
A Definitive Lipidomics Workflow for Human Plasma Utilizing Off-line Enrichment and Class Specific Separation of Phospholipids
 A Definitive Lipidomics Workflow for Human Plasma Utilizing Off-line Enrichment and Class Specific Separation of Phospholipids Jeremy Netto, 1 Stephen Wong, 1 Federico Torta, 2 Pradeep Narayanaswamy, 2
A Definitive Lipidomics Workflow for Human Plasma Utilizing Off-line Enrichment and Class Specific Separation of Phospholipids Jeremy Netto, 1 Stephen Wong, 1 Federico Torta, 2 Pradeep Narayanaswamy, 2
bio-mof-1 DMASM Wavenumber (cm -1 ) Supplementary Figure S1 FTIR spectra of bio-mof-1, DMASMI, and bio-mof-1 DMASM.
 bio-mof-1 Transmittance bio-mof-1 DMASM DMASMI 2000 1500 1000 500 Wavenumber (cm -1 ) Supplementary Figure S1 FTIR spectra of bio-mof-1, DMASMI, and bio-mof-1 DMASM. Intensity (a.u.) bio-mof-1 DMASM as
bio-mof-1 Transmittance bio-mof-1 DMASM DMASMI 2000 1500 1000 500 Wavenumber (cm -1 ) Supplementary Figure S1 FTIR spectra of bio-mof-1, DMASMI, and bio-mof-1 DMASM. Intensity (a.u.) bio-mof-1 DMASM as
Aminoglycoside activity observed on single pre-translocation ribosome complexes
 correction notice Nat. Chem. Biol. 6, 54 62 (2010) Aminoglycoside activity observed on single pre-translocation ribosome complexes Michael B Feldman, Daniel S Terry, Roger B Altman & Scott C Blanchard
correction notice Nat. Chem. Biol. 6, 54 62 (2010) Aminoglycoside activity observed on single pre-translocation ribosome complexes Michael B Feldman, Daniel S Terry, Roger B Altman & Scott C Blanchard
Phospholipids of Lactobacillus spp.
 JOURNAL OF BACTERIOLOGY, Nov. 1995, p. 6304 6308 Vol. 177, No. 21 0021-9193/95/$04.00 0 Copyright 1995, American Society for Microbiology Phospholipids of Lactobacillus spp. D. B. DRUCKER, 1,2 * G. MEGSON,
JOURNAL OF BACTERIOLOGY, Nov. 1995, p. 6304 6308 Vol. 177, No. 21 0021-9193/95/$04.00 0 Copyright 1995, American Society for Microbiology Phospholipids of Lactobacillus spp. D. B. DRUCKER, 1,2 * G. MEGSON,
Comparative Lipid Profiling of the Cnidarian Aiptasia pallida and Its Dinoflagellate Symbiont
 Comparative Lipid Profiling of the Cnidarian Aiptasia pallida and Its Dinoflagellate Symbiont Teresa A. Garrett 1 *, John L. Schmeitzel 1, Joshua A. Klein 3, Janice J. Hwang 1, Jodi A. Schwarz 2 1 Department
Comparative Lipid Profiling of the Cnidarian Aiptasia pallida and Its Dinoflagellate Symbiont Teresa A. Garrett 1 *, John L. Schmeitzel 1, Joshua A. Klein 3, Janice J. Hwang 1, Jodi A. Schwarz 2 1 Department
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Selective phosphatidylcholine double bond. fragmentation and localization using. Paternó-Büchi reactions and ultraviolet.
 Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2017 Selective phosphatidylcholine double bond fragmentation and localization using Paternó-Büchi reactions
Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2017 Selective phosphatidylcholine double bond fragmentation and localization using Paternó-Büchi reactions
Targeting RNA-Protein interactions within. the HIV-1 lifecycle.
 SUPPORTING INFORMATION for Targeting RNA-Protein interactions within the HIV-1 lifecycle. Neil M. Bell, 1,2 Anne L Hernault, 2 Pierre Murat, 1 James E. Richards, 2 Andrew M.L. Lever 2 & Shankar Balasubramanian
SUPPORTING INFORMATION for Targeting RNA-Protein interactions within the HIV-1 lifecycle. Neil M. Bell, 1,2 Anne L Hernault, 2 Pierre Murat, 1 James E. Richards, 2 Andrew M.L. Lever 2 & Shankar Balasubramanian
Supplementary Figure 1. Ent inhibits LPO activity in a dose- and time-dependent fashion.
 Supplementary Information Supplementary Figure 1. Ent inhibits LPO activity in a dose- and time-dependent fashion. Various concentrations of Ent, DHBA or ABAH were pre-incubated for 10 min with LPO (50
Supplementary Information Supplementary Figure 1. Ent inhibits LPO activity in a dose- and time-dependent fashion. Various concentrations of Ent, DHBA or ABAH were pre-incubated for 10 min with LPO (50
MALDI Activity 4 MALDI-TOF Mass Spectrometry: Data Analysis
 MALDI Activity 4 MALDI-TOF Mass Spectrometry: Data Analysis Model 1: Introduction to Triacylglycerides (TAGs) In MALDI Activity 3 you learned how to open your raw mass spectrum data and use the MMass program
MALDI Activity 4 MALDI-TOF Mass Spectrometry: Data Analysis Model 1: Introduction to Triacylglycerides (TAGs) In MALDI Activity 3 you learned how to open your raw mass spectrum data and use the MMass program
CHAPTER 4. Tryptophan fluorescence quenching by brominated lipids
 CHAPTER 4 Tryptophan fluorescence quenching by brominated lipids 102 4.1 INTRODUCTION The structure and dynamics of biological macromolecules have been widely studied with fluorescence quenching. The accessibility
CHAPTER 4 Tryptophan fluorescence quenching by brominated lipids 102 4.1 INTRODUCTION The structure and dynamics of biological macromolecules have been widely studied with fluorescence quenching. The accessibility
Expression constructs
 Gene expressed in bebe3 ZmBEa Expression constructs 35S ZmBEa Pnos:Hygromycin r 35S Pnos:Hygromycin r 35S ctp YFP Pnos:Hygromycin r B -1 Chl YFP- Merge Supplemental Figure S1: Constructs Used for the Expression
Gene expressed in bebe3 ZmBEa Expression constructs 35S ZmBEa Pnos:Hygromycin r 35S Pnos:Hygromycin r 35S ctp YFP Pnos:Hygromycin r B -1 Chl YFP- Merge Supplemental Figure S1: Constructs Used for the Expression
9/11/18. Cell Function & Chemistry. Molecular and Cellular Biology. 2. Bio-Chemical Foundations & Key Molecules of a Cell
 Molecular and Cellular Biology Cell Function & Chemistry 2. Bio-Chemical Foundations & Key Molecules of a Cell Prof. Dr. Klaus Heese Interaction Molecular Bonds Define Cellular Functions Water H 2 O Interactions
Molecular and Cellular Biology Cell Function & Chemistry 2. Bio-Chemical Foundations & Key Molecules of a Cell Prof. Dr. Klaus Heese Interaction Molecular Bonds Define Cellular Functions Water H 2 O Interactions
Supplementary Information. Conformational states of Lck regulate clustering in early T cell signaling
 Supplementary Information Conformational states of Lck regulate clustering in early T cell signaling Jérémie Rossy, Dylan M. Owen, David J. Williamson, Zhengmin Yang and Katharina Gaus Centre for Vascular
Supplementary Information Conformational states of Lck regulate clustering in early T cell signaling Jérémie Rossy, Dylan M. Owen, David J. Williamson, Zhengmin Yang and Katharina Gaus Centre for Vascular
Supporting Information: Millimeter-area, free. standing, phospholipid bilayers
 Electronic Supplementary Material (ESI) for Soft Matter. This journal is The Royal Society of Chemistry 2016 Supporting Information: Millimeter-area, free standing, phospholipid bilayers Peter J. Beltramo,
Electronic Supplementary Material (ESI) for Soft Matter. This journal is The Royal Society of Chemistry 2016 Supporting Information: Millimeter-area, free standing, phospholipid bilayers Peter J. Beltramo,
Putting bee honey on a cut kills bacteria. Honey contains a high concentration of sugar
 Q1.(a) Give two ways in which pathogens can cause disease. 1... 2... (b) Putting bee honey on a cut kills bacteria. Honey contains a high concentration of sugar. Use your knowledge of water potential to
Q1.(a) Give two ways in which pathogens can cause disease. 1... 2... (b) Putting bee honey on a cut kills bacteria. Honey contains a high concentration of sugar. Use your knowledge of water potential to
PHOSPHOLIPIDS METABOLISM. BY Dr. Walid Said Zaki Dr. Marwa Ali LECTURER OF BIOCHEMISTRY AND MOLECULAR BIOLOGY
 PHOSPHOLIPIDS METABOLISM BY Dr. Walid Said Zaki Dr. Marwa Ali LECTURER OF BIOCHEMISTRY AND MOLECULAR BIOLOGY 1. State the definition and classification of Phospholipids. 2. Describe the general structure
PHOSPHOLIPIDS METABOLISM BY Dr. Walid Said Zaki Dr. Marwa Ali LECTURER OF BIOCHEMISTRY AND MOLECULAR BIOLOGY 1. State the definition and classification of Phospholipids. 2. Describe the general structure
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction
 Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Table S1: Kinetic parameters of drug and substrate binding to wild type and HIV-1 protease variants. Data adapted from Ref. 6 in main text.
 Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Supplementary Information
 Supplementary Information Molecular imaging of brain localization of liposomes in mice using MALDI mass spectrometry Annabelle Fülöp 1,2, Denis A. Sammour 1,2, Katrin Erich 1,2, Johanna von Gerichten 4,
Supplementary Information Molecular imaging of brain localization of liposomes in mice using MALDI mass spectrometry Annabelle Fülöp 1,2, Denis A. Sammour 1,2, Katrin Erich 1,2, Johanna von Gerichten 4,
Phospholipid characterization by a TQ-MS data based identification scheme
 P-CN1716E Phospholipid characterization by a TQ-MS data based identification scheme ASMS 2017 MP-406 Tsuyoshi Nakanishi 1, Masaki Yamada 1, Ningombam Sanjib Meitei 2, 3 1 Shimadzu Corporation, Kyoto, Japan,
P-CN1716E Phospholipid characterization by a TQ-MS data based identification scheme ASMS 2017 MP-406 Tsuyoshi Nakanishi 1, Masaki Yamada 1, Ningombam Sanjib Meitei 2, 3 1 Shimadzu Corporation, Kyoto, Japan,
Tandem mass spectrometry analysis of prostaglandins and isoprostanes
 Tandem mass spectrometry analysis of prostaglandins and isoprostanes Jeevan Prasain jprasain@uab.edu 6-2612 verview Introduction to PGs and their synthesis Mass spectrometry characterization of PGs and
Tandem mass spectrometry analysis of prostaglandins and isoprostanes Jeevan Prasain jprasain@uab.edu 6-2612 verview Introduction to PGs and their synthesis Mass spectrometry characterization of PGs and
Phase Transition Behaviours of the Supported DPPC Bilayer. Investigated by Sum Frequency Generation (SFG) and Atomic Force
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2015 Supporting Information for Phase Transition Behaviours of the Supported DPPC Bilayer
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2015 Supporting Information for Phase Transition Behaviours of the Supported DPPC Bilayer
Lipids Analysis. Lipids
 Lipids Analysis Stephen Barnes 3 5 15 Lipids Lipids are mostly very hydrophobic Most are conjugates of fatty acids of a variety of chain lengths, which have different degrees of unsaturation, cis trans
Lipids Analysis Stephen Barnes 3 5 15 Lipids Lipids are mostly very hydrophobic Most are conjugates of fatty acids of a variety of chain lengths, which have different degrees of unsaturation, cis trans
Inorganic compounds: Usually do not contain carbon H 2 O Ca 3 (PO 4 ) 2 NaCl Carbon containing molecules not considered organic: CO 2
 Organic Chemistry The study of carbon-containing compounds and their properties. Biochemistry: Made by living things All contain the elements carbon and hydrogen Inorganic: Inorganic compounds: All other
Organic Chemistry The study of carbon-containing compounds and their properties. Biochemistry: Made by living things All contain the elements carbon and hydrogen Inorganic: Inorganic compounds: All other
Fluorescent Carbon Dots as Off-On Nanosensor for Ascorbic Acid
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Fluorescent Carbon Dots as Off-On Nanosensor for Ascorbic Acid Jun Gong, Xin Lu, Xueqin An*
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Fluorescent Carbon Dots as Off-On Nanosensor for Ascorbic Acid Jun Gong, Xin Lu, Xueqin An*
Essential Lipidomics Experiments using the LTQ Orbitrap Hybrid Mass Spectrometer
 Application Note: 367 Essential Lipidomics Experiments using the LTQ rbitrap Hybrid Mass Spectrometer Thomas Moehring 1, Michaela Scigelova 2, Christer S. Ejsing 3, Dominik Schwudke 3, Andrej Shevchenko
Application Note: 367 Essential Lipidomics Experiments using the LTQ rbitrap Hybrid Mass Spectrometer Thomas Moehring 1, Michaela Scigelova 2, Christer S. Ejsing 3, Dominik Schwudke 3, Andrej Shevchenko
Supplementary Figures
 Supplementary Figures a b c d PDI activity in % ERp72 activity in % 4 3 2 1 1 1 ERp activity in % e ΔRFU min -1 1 1 ERp7 activity in % 1 1 Supplementary Figure 1. Selectivity of the bepristat-mediated
Supplementary Figures a b c d PDI activity in % ERp72 activity in % 4 3 2 1 1 1 ERp activity in % e ΔRFU min -1 1 1 ERp7 activity in % 1 1 Supplementary Figure 1. Selectivity of the bepristat-mediated
Supplementary Information. Effects of Perfluorooctanoic Acid on Metabolic Profiles in Brain and Liver of Mouse by a
 Supplementary Information Effects of Perfluorooctanoic Acid on Metabolic Profiles in Brain and Liver of Mouse by a High-throughput Targeted Metabolomics Approach Nanyang Yu, Si Wei, *, Meiying Li, Jingping
Supplementary Information Effects of Perfluorooctanoic Acid on Metabolic Profiles in Brain and Liver of Mouse by a High-throughput Targeted Metabolomics Approach Nanyang Yu, Si Wei, *, Meiying Li, Jingping
Figure S1. Standard curves for amino acid bioassays. (A) The standard curve for leucine concentration versus OD600 values for L. casei.
 Figure S1. Standard curves for amino acid bioassays. (A) The standard curve for leucine concentration versus OD600 values for L. casei. (B) The standard curve for lysine concentrations versus OD600 values
Figure S1. Standard curves for amino acid bioassays. (A) The standard curve for leucine concentration versus OD600 values for L. casei. (B) The standard curve for lysine concentrations versus OD600 values
