Protein Name. IFLENVIR,DSVTYTEHAK,TV TALDVVYALK,KTVTALDVV YALK,TVTALDVVYALKR,IF LENVIRDSVTYTEHAK gi Histone H2B
|
|
|
- Buck Miller
- 5 years ago
- Views:
Transcription
1 Table 1. A functional category list of proteins (Lentinula edodes) identified by 1-DGE and nesi-lc-ms/ms. The table lists indicated fraction numbers, matching peptides, scores, accession numbers, protein names, theoretical mass (molecular weight, Da), theoretical pi and analyzed peptides; left to right. Fraction Matching Peptide Score Accession Protein Name Analytical MW Theoretical pi Peptide Chromatin structure and dynamics gi Histone H2B* AMAILNSFVNDIFER gi 3142 Histone H4.2* TVTALDVVYALKR gi Histone H2B AMAILNSFVNDIFER gi 3142 Histone H IFLENVIR,DSVTYTEHAK,TV TALDVVYALK,KTVTALDVV YALK,TVTALDVVYALKR,IF LENVIRDSVTYTEHAK gi Histone H2B LILPGELAK,AMAILNSFVNDI FER Cell division and chromosome partitioning gi α tubulin* LIAQVVSSITASLR,FDGSLNV DLNEFQTNLVPFPR,AFVHW YVGEGMEEGEFSEAREDLAA gi CDC (cell division control protein) 12* LTVIDTPGFGDYVNNR gi Septin* STLINTLFSAHLMDSK Transcription gi Pre-mRNA splicing factor* SATTNLSTADVVNNLKR gi AP-1 like bzip transcriptional activator* ASCYHILEEISSLPK Translation, ribosomal structure and biogenesis gi GNVASDSKNDPAK,EHALLA Τranslation elongation factor FTLGVR,AGMIVTFAPSNVTT 1-α EVK gi Τranslation elongation factor 1 α gi 5059 Ribosomal protein L3* HGSLGFLPR gi Cytoplasmic asparaginyltrna synthetase* FLAWLCAR gi gi gi gi gi gi gi Translation elongation factor 2* Translation elongation factor 2* Translation elongation factor 2* Translation elongation factor 2* Τranslation elongation factor 1-α Τranslation elongation factor 1-α Τranslation elongation factor 1-α EHALLAFTLGVR,YVVTVIDA PGHRDFIK IKPVVIINK,FSVSPVVQVAVE VK IQGPNYIPGK,FSVSPVVQVA VEVK ILADEFGWDVTDAR VLADEFGWDVTDAR ETKAGNAK,GITIDIALWK,Y AWVLDKLK,YVVTVIDAPGH R,EHALLAFTLGVR,ERGITIDI ALWK,YVVTVIDAPGHRDFI KEAAELGK,ERGITIDIALWK, YAWVLDKLK EHALLAFTLGVR,SVEMHHE QLEQGTPGDNVGFNV
2 gi Τranslation elongation factor EHALLAFTLGVR,AGMIVTFA α PSNVTTEVK gi 5059 Ribosomal protein L HGSLGFLPR gi Ribosomal protein L2* RLNLLQLAPGGHLGR LFAIGFTK,APSIFDVR,MFEI S-phase specific ribosomal gi MTR,SIYPLQNVYVR,IIEVSL protein cyc07* GDLNKEEEQSFR TSFFQALGIPTK,GNIGFVFTS gi L10e protein* GDLK APTGSNTVLLR,ILKAGGEVL gi S ribosomal protein L18* TLDQLALR gi Ribosomal protein S2* SMEEIYLFSLPVK gi Cytosolic small ribosomal subunit S4* TDPTYPAGFMDVISIEK gi Ribosomal protein S28* VSGVGLLALWK gi Ribosomal protein L12* ATGGEVGASSALAPK modification, protein gi 2551 Ubiquitin* ESTLHLVLR,IQDKEGIPPDQQ R,TITLEVESSDTIDNVK LYSLDMGALMAGAK,EFVTL gi Heat shock protein 100* FLQVLDDGR,TLATLLFDSPD NALEEYIYDTR,KNALEEYIY gi PSS1* DTR FELSGIPPAPR,DAGTIAGMN VLR,IINEPTAAAIAYGLDK,A gi Heat shock protein 70* TAGDTHLGGEDFDNR,IINEP TAAAIAYGLDKK,KSETFSTY ADNQPGVLIQVFEGER FELSGIPPAPR,LVSDFFNGKE gi Heat shock protein 70* PNK,ATAGDTHLGGEDFDNR, IINEPTAAAIAYGLDKK,TQD LLLLDVAPLSLGIETAGGVM TALIK,ATAGDTHLGGEDFD gi Heat shock protein 70* LVSDFFNGKEPNK,IINEPTAA AIAYGLDKK,KSETFSTFSDN QPGVLIQVFEGER,TLSSAAQ TSIEIDSLYEGIDFYTSITR FELSGIPPAPR,LVDHFIQEFK, gi Heat shock protein 70* IINEPTAAAIAYGLDK,ATAG DTHLGGEDFDNR,NQVAMNP HNTVFDAKR DSPFLEVLKK,HFSVEGQLEF gi Heat shock protein 90* KLGIHEDAQNR,GIVDSEDLPL gi Heat shock protein 90* NISR TIEDELEVTEGMR,VEFEKPLI gi Heat shock protein 60* LLSEK GFISPYFITDVK,VEFEKPLILL gi kda chaperonin* SEK gi 2908 Heat shock protein 70* TFSPQEISSMVLTK gi Heat shock protein 70* DAGTIAGLNILR gi kda cyclophilin* SLLAPLLLNSALAAIR gi S proteaseme regulatory subunit 7* TMLELINQLDGFDPR gi kda cyclophilin SLLAPLLLNSALAAIR gi Thioredoxin reductase Trr1* VVIIGSGPAGHTAAIYLAR gi S proteasome component α 1* LYQVEYAFK gi S proteasome subunit* YNDDLSLEDAIHTALLTLK
3 gi S proteasome component α 5* LFQVEYAIEAIK gi Cyclophilin (peptidyl-prolyl cis-trans isomerase)* IIPNFMIQGGDFTR Energy production and conversion gi Plasma membrane (H+) ATPase* GAADIVLTEPGLSTIVHAIR gi ATP citrate lyase (citrate synthetase)* SIAIIAEGVPER gi ATP citrate lyase (citrate synthetase) SIAIIAEGVPER gi Pyruvate carboxylase* VFEEYQGFLEK gi Pyruvate kinase* QIHLHR,NGVDMIFASFIR,IEN EQGVANFDEILK,GDLGIEIPA SQVFLAQK gi Isocitrate dehydrogenase* GLANPTALLLSSLMMLR gi Inorganic pyrophosphatase* VLGIMALLDEGETDWK gi NADH-ubiquinone oxidoreductase* VVYEPLQLTQAFR Carbohydrate transport gi phosphogluconate dehydrogenase* GILFVGSGVSGGEEGAR gi Glycogen synthase* GVDMFIESLAR gi Trehalose synthase* LADELREK,NNHNILQGVAD PSLR,IGFFSSTPQGGGVALM gi Phosphoglucomutase* FFGNLMDAGR VIPSLNGK,YDSVHGR,EISVF GEK,FKGTVETK,VVDLLVFA AK,SVNGNIIPSSTGAAK,VPT LDVSVVDLVCR,LIVDGKEIS gi Glyceraldehyde-3-phosphate VFGEK,LIAWYDNEWGYSGR dehydrogenase*,fgivealmttvhattatqk, VIHDKFGIVEALMTTVHATT ATQK,DAGAIPWSSVGAEYIV ESTGVFTTIEK,NALLNPEVN VVAVNDPFIALEYMVYMFK gi Glyceraldehyde-3-phosphate dehydrogenase* gi Transaldolase* gi Amino acid transport NAD-specific glutamate dehydrogenase* YDSVHGR,DPAAIPWGSVGA HYIVESTGVFTTIEK LLIEFGK,VSTEVDAR,IASTW EGIQAAR LGPAYLQLK,LPSGEIVLDGT DFR gi Tryptophan synthetase Trp 1* WFLGSLQDMLHAR gi Glutamine synthetase* gi Glutamine synthetase* gi deoxy-7- phosphoheptulonate synthase* KVTDIGQLR,RPASNIDPYR,R HDEHIAVYGEDNDLR,GGDN ILVLAETYNNDGTPNR KVTDIGQLR,RPASNIDPYR,H ETGHIGSFSSGVANR,VQAEY VWIDGDGGLR ELASGVSFPIGFK,VLGYDPL VQPALLR gi Endopeptidase* EPGLAFAFGK
4 gi α-keto-β-hydroxylacyl reductoisomerase* Nucleotide transport gi 2977 ADP/ATP carrier protein* Inorganic ion transport DNGLNVIVGVR TIASDGIAGLYR,QFNGLVDV YRK gi Catalase* YNILDLTK,LFSYPDTHR gi Manganese superoxide dismutase* IALQAALK Cytoskeleton gi Actin* IVAPPERK,AVFPSIVGRPR,I WHHTFYNELR,SYELPDGQVI TIGNER,YPIEHGIVTNWDDM EK,VAPEEHPVLLTEAPLNPK, DLYGNVVLSGGTTMFPGIAD R,TTGIVLDSGDGVTHTVPIY gi Actin* IVAPPERK,VAPEEHPVLLTE APLNPK,FRAPEALFQPGFLG LESAGIHETTYNSIMK Cell envelope biogenesis, outer membrane gi Glutamine:fructose-6- phosphate amidotransferase* TIETLELELAAIMK gi gi gi Intracellular trafficking and secretion Mitochondrial processing peptidase β subunit* Ypt (yeast protein transport) 1 related protein* SAR small monomeric GTPase* gi Ran GTPase Spi1* gi gi Signal transduction CAP (adenyl cyclase associated protein)* Protein phosphatase regulatory subunit Pab1* DVPVAVDIISDILQNSK LLLIGDSGVGK,LQIWDTAGQ ER,FADDTYTESYISTIGVDFK LLFLGLDNAGK,LATLQPTLH PTSEELAIGNVK LVLVGDGGTGK,SNYNFEKP FLWLAR SFAGASVVEIVGLVEK SFFSEIISSISDVK gi Hhp1 protein kinase* TVLLLADQLISR NLLSVAYK,YLAEFATGDK,D gi protein* STLIMQLLR,PETREDSVYLA K,VASSDQELTVEER,QAFDD AIAELDTLSEESYK,DNLTLW TSDMQDSADKPAEKDEAAD APADE,QAFDDAIAELDTLSE ESYKDSTLIMQLLR TIIVWQLTR,DKTIIVWQLTR, DGITMLWDLNEGK,DGITML WDLNEGK,SIVDELKPAYAE Guanine nucleotide binding gi EGGR,SIVDELKPAYAEEGGR protein beta subunit*,fspnvmnpvivssgwdk,hl YSLEAGDIVNALVFSPNR,HL YSLEAGDIVNALVFSPNR,G gi Small GTPase CDC42* NVFDEAIVAALEPPVVK gi Ras * INVDQAFQDLVR
5 Asterisks indicate unique proteins identified from L.edodes fruiting body using 1-DGE and following LC-MS/MS. Commonly identified proteins from 1-DGE and 2-DGE analysis are shown in red letters.
6 Table 2. Proteins (Lentinula edodes) identified by 2-DGE and nesi-lc-ms/ms. The table lists indicated spot numbers, internal amino acid sequences analyzed by tandem MS, matching peptides, scores, accession numbers, protein names, sequence coverage, theoretical mass (molecular weight, kda), theoretical pi, function and N -terminal amino acid sequence analyzed; left to right. Spot Amino Acid Sequence Peptide Score Accession Protein Name Sequence Coverage Analytical MW Theoretical pi Functional Category N -terminal Amino Acid GFFHHESDEAQAYD 1 EAFVEVTK 1 40 gi Hypothetical protein 2% Unknown Q 2 Unanalyzed 3 4 IPTIGIGAGNGTSGQVLV QADMTGNFPPGR TEAILTPLQQAMRQK,QV KESWPSNPVDGYIQSIR, AGPGK 1 26 gi methyl-2-oxobutanoate hydroxymethyltransferase* 9% Coenzyme metabolism DGGME 3 0 gi Hypothetical protein 7% Unknown QGLNL 5 IEKDAVLK 1 39 gi Hypothetical protein 0% Unknown QLERG LYDDVVPK,IIPDFMLQG gi Cyclophilin* 13% modification, protein Blocked GDFTR 7 IDTGSPRYSFLDPR 1 24 gi Hypothetical protein 3% Unknown Unanalyzed 8 MYTTISPMLVVSLATLAF ISLFNIYR 1 25 gi Cytochrome P450 1* 5% Secondary metabolites biosynthesis, transport, and catabolism Unanalyzed 9 DATEAFEDVGHSDEAR 1 52 gi Hypothetical protein 10% Unknown Unanalyzed MPNLYETFVNSIAPSIYG 10 NEDIKK,LSSHFVSIR 2 36 gi Hypothetical protein 4% Unknown Unanalyzed 11 No significant hit found QPNEDR 12 DYGVLIEDEGIALR gi Hypothetical protein 7% Unknown QPXIY 13 KNGHVVIK,NRPCKIVDM Translation iniciation factor Translation, ribosomal 3 82 gi % STSK,VHLVA IDIFTGK 5A* structure and biogenesis Blocked 14 EFGIDQK 1 45 gi Hypothetical protein 1% Unknown Unanalyzed APTIKVGDTVPDGT 15 VEVDLASIPEGK 1 46 gi Hypothetical protein 5% Unknown F SPVKPTAAPITTTPSEPKV SPAK,VSLPR,MSVSQQQ SKQNDR,TEDEHNPMSAI 16 RPTMAPPIR,T AQPQRPAHPM DMAKYASGKI 5 0 gi Hypothetical protein 6% Unknown Unanalyzed PLHAYQHDNNSGG 17 FVNSIVSRLDAK 1 43 gi Hypothetical protein 4% Unknown PL 18 IALQAALK 1 49 gi Manganese superoxide Inorganic ion transport 5% dismutase* QGGPL
7 19 AATPK,IALQAALK 2 56 gi Manganese superoxide Inorganic ion transport 9% dismutase QNTLL 20 Unanalyzed 21 TTKATMAANAK 1 48 gi Hypothetical protein 10% Unknown QFGIE 22 SFGAEEGGSGK 1 44 gi Hypothetical protein 0% Unknown SFGIE 23 Unanalyzed 24 MSGLSKMEK 1 35 gi Ribosome biogenesis protein* 1% Translation, ribosomal structure and biogenesis QFGIE 25 QEQQHLWTR 1 41 gi Hypothetical protein 1% Unknown Unanalyzed 26 LQSTSFDKK,SYLTYLK 2 56 gi Cytoplasm protein* 9% Signal transduction MLLYS 27 QFHIFIVK 1 45 gi Hypothetical protein 7% Unknown Unanalyzed 28 GETVISPR 1 42 gi Hypothetical protein 0% Unknown Unanalyzed 29 FGSPALLLLVAR 1 48 gi Hypothetical protein 1% Unknown Unanalyzed 30 AANRAFASSAATLIK 1 50 gi Hypothetical protein 16% Unknown Unanalyzed 31 LVLVGDGGTGK,RHLTG EFEKK,FGGLR,NVPNWH R,VCENIPIVLCGNK,VDV K,TGAVTFHR,NLQYFEIS gi Small G protein RAN* 31% Intracellular trafficking and secretion 32 GLISEER 1 28 gi Hypothetical protein 0% Unknown Unanalyzed 33 SVYVIDPAKK 1 56 gi Hypothetical protein 3% Unknown Unanalyzed 34 SVYVIDPAKK 1 55 gi Hypothetical protein 3% Unknown QGQDL 35 SVYVIDPAKK 1 45 gi Hypothetical protein 3% Unknown XAPNT 36 NEENILPLPK 1 40 gi Hypothetical protein 1% Unknown Unanalyzed 37 ILCDISNISR 1 33 gi Hypothetical protein 2% Unknown Unanalyzed 38 Unanalyzed EIAPDGHCLFSAFADQLS 39 TLSIPLQQK 1 31 gi Hypothetical protein 4% Unknown Unanalyzed PETR,EDSVYLAK,LAEQ AER,YEEMVENMKR,VA SSDQELTVEER,NLLSVA YK,NVIGAR,IVSSIEQK,E ESK,GNEAQVSMIK,HLIP 40 SAASGESK,MMGDYHR, YLAEFATGDKR,ESADKS LEAYK,AASDVAVTELPP THPIR,DSTLIMQLLR,DN LTLWTSDMQDSADKPAE K,DEAADAPADE gi protein* 68% Signal transduction PETRE Blocked
8 PETR,EDSVYLAK,LAEQ AER,YEEMVENMKR,VA SSDQELTVEER,NLLSVA YK,NVIGAR,IVSSIEQK,E ESK,GNEAQVSMIK,HLIP SAASGESK,MMGDYHR, 41 YLAEFATGDKR,ESADKS gi protein 72% Signal transduction PETRE LAYK,AASDVAVTELPPT HPIR,LGLALNFSVFYYEI LNSPDR,DSTLIMQLLR,D NLTLWTSDMQDSADKP AEK PETR,EDSVYLAK,LAEQ AER,YEEMVENMKR,VA SSDQELTVEER,NLLSVA YK,NVIGARR,IVSSIEQK, EESK,GNEAQVSMIK,HLI PSAASGESK,MMGDYHR, 42 YLAEFATGDKR,ESADKS gi protein 78% Signal transduction QEDIE LEAYK,AASDVAVTELPP THPIR,QAFDDAIAELDTL SEESYK,DSTLIMQLLR,D NLTLWTSDMQDSADKP AEK,DEAADAPADE LAEQAERYEEMVENMK, VASSDQELTVEER,NLLS 43 VAYK,ICEDILDVLDK,YL gi protein 22% Signal transduction QPREE AEFATGDK VASSDQELTVEER,NLLS 44 VAYK,ICEDILDVLDK gi protein 12% Signal transduction ATELI NLLSVAYK,ICEDILDVLD 45 K 2 63 gi protein 7% Signal transduction QYCES 46 EVDLSK 1 41 gi Hypothetical protein 2% Unknown Unanalyzed 47 ELEEELK 1 49 gi Hypothetical protein 0% Unknown Unanalyzed Cell division and Cytoplasmic dynein 48 EEIEEELR 1 44 gi % chromosome QPLPP intermediate chain * partitioning 49 FLGSLTGHK,GWVTAIAT SSENPDMILTASR,DKTII VWQLTR,DDDSYGYPKR, LWDLNTGLTTR,QIVSGS R,LWNTLGECK,YDIK,DD GHSEWVSCVR,FSPNVM NPVIVSSGWDK,VWELSK,TNHYGHTGYINTVSVSP DGSLAASGGK,DGITML WDLNEGK,YWLCAATAS CVK,SIVDELKPAYAEEG GR,VWTVTS gi Guanine nucleotide binding protein beta subunit* 61% Signal transduction QGIEQ
9 50 NSLRIAR 1 53 gi MPENFTYKPK 1 42 gi ATP synthase δ chain, mitochondrial precursor* MAPK activated protein kinase Cmk2* 4% Energy production and conversion QGE 1% Signal transduction Unanalyzed IIAIDVKDPLASK,HLPGL 52 LRATNEWFR,KYADEVIR 3 52 gi Hypothetical protein 12% Unknown QSQYS IIAIDVKDPLASK,HLPGL 53 LR 2 47 gi Hypothetical protein 6% Unknown Unanalyzed 54 HIDTAAVYGTEK,YGKTG Carbohydrate transport 2 55 gi Aldehyde reductase* 8% AQVALAWG ITK QYLET GSVANMSLGGGK,KIVE 55 AGSYK RADGIHLLNIGK,FTPGSF 2 63 gi Hypothetical protein 3% Unknown DEILV 56 TNYITR,SFKEPR,VDHQA 4 60 gi Hypothetical protein 13% Unknown Unanalyzed 57 IR GVFAAGDVQDKR 1 94 gi Hypothetical protein 3% Unknown Unanalyzed 58 DMLASNMLEAVSKPK 1 50 gi Hypothetical protein 0% Unknown QTPLD 59 Unanalyzed 60 QALISNGLK 1 47 gi Palmitoyltransferase* 1% Lipid metabolism Unanalyzed SYWPLDEPSINMLAVSTS KK YDSVHGR,LIVDG K,EISVFGEK,LTGLAFR,V VDLLVFAAK 1 27 gi Hypothetical protein 2% Unknown Unanalyzed 4 43 gi Glyceraldehyde-3-phosphate dehydrogenase* 11% Carbohydrate transport 63 CTS IACFLHAK 1 48 gi Aspartyl proteinase* 6% Amino acid transport Blocked 64 THDGAVAVR 1 36 gi Hypothetical protein 5% Unknown Unanalyzed VTDIGQLR,GGFPGPQGP YYCGAGTGK,DLIEAHYR TTTVSK,KVTDIGQLR,GG FPGPQGPYYCGAGTGK,D LIEAHYR,RHDEHIAVYG EDNDLR,RPASNIDPYR LQDYFK,DNGLNVIVGVR K,EVYSDLYGER GVAEVKLSMFESALK,TS EQTSGALGEDHGPKK 3 69 gi Glutamine synthetase* 9% gi Glutamine synthetase 18% Amino acid transport Amino acid transport 3 98 gi Hypothetical protein 4% Unknown IKTLD QVLRP GILDI QNRVY 2 29 gi Hypothetical protein 3% Unknown QAYKI
10 69 AGFAGDDAPR,AVFPSIV GRPR,HQGVMVGMGQK, DSYVGDEAQSK,RGILTL K,YPIEHGIVTNWDDMEK,IWHHTFYNELR,VAPEE HPVLLTEAPLNPK,LDLA GR,DLTDFLIK,NLMER,G gi β-actin* 56% Cytoskeleton QTMNY 70 YPFTTTAER,LCYVALDF EQELQTAAQSSALEK,SY ELPDGQVITIGNER,CDLD IR,DLYGNVVLSGGTTMF PGIADR,VKIVAPPER,QE YDESGPGIVHRK DLTDFLIK,LCYVALDFE QELQTAAQSSALEK,SYE LPDGQVITIGNER,DLYG gi β-actin 18% Cytoskeleton QPGAK NVVLSGGTTMFPGIADR MQK 71 AAVPSGASTGIHEAVELR,EIDDFLIK,LDGTPNK,YD LDFK,IKTAIEK,KACNGL LLK,SGETEDVTIADLVV GLGVGIIKTGAPCR gi Hypothetical protein 18% Unknown QPNKD AAVPSGASTGIHEAVELR 72,EIDDFLIK 2 84 gi Hypothetical protein 5% Unknown Unanalyzed 73 KDDGTSAIVLENPK 1 52 gi Hypothetical protein 2% Unknown Unanalyzed 74 VLHMDR,DR,DYAVDLIP K,SPLMGLFEK,KVIGDPS YFGAGK 5 82 gi Hypothetical protein 8% Unknown Unanalyzed ILSTLDR,GVLLYGPPGTG K,TMLVK,VVGSEFVQK, YLGEGPR,ENSPAIIFIDEI DAIATK,VIMATNR TGTLSSTGGGGGGGPERP PFAR VWELSK,SIVDELKPAYA EEGGR SPYLYPLYGLGELPQGFT R gi Protease regulatory subunit 6B homolog* 15% modification, protein Blocked 1 28 gi Hypothetical protein 2% Unknown Unanalyzed 2 49 gi Guanine nucleotide binding protein beta subunit 7% Signal transduction Unanalyzed 1 42 gi Hypothetical protein 4% Unknown Unanalyzed 79 ELAALKGK 1 48 gi Hypothetical protein 3% Unknown Unanalyzed DYAVDLIPK,SPLMGLFE gi Hypothetical protein 6% Unknown Unanalyzed K,KVIGDPSYFGAGK 81 GRPI LSIPK 1 41 gi Putative monooxygenase* 1% Secondary metabolites biosynthesis, transport, and catabolism Blocked
11 82 YAWVLDK,YVTVIDAPG HR 2 42 gi Translation elongation factor 1-α 22% Translation, ribosomal structure and biogenesis QVAFE 83 VVDLLAPYAR,IGLFGGA GVGK,VALVFGQMNEPP GAR,FTQAGSEVSALLGR gi Hypothetical protein 14% Unknown Unanalyzed,IPSAVGYQPTLSTDMGG MQER,ITTTK,MLDPR 84 GIAELGIYPAVDPLDSK 1 56 gi Hypothetical protein 3% Unknown Unanalyzed 85 DATDPPDK 1 34 gi Hypothetical protein 1% Unknown Unanalyzed 86 LGRVELLK,GGSEAVISK, RVFNDPPALTANHDPNG MR 3 45 gi Hypothetical protein 1% Unknown Unanalyzed TAVALDTILNQK,ASLAY RQLSLMLR SLFHPETLLTGK,EDAAS NYAR,GHYTIGK,YMACA LLYR,LGICNEPPAHVPG GDLAK,SMCMLSNTTAIS SAWSR,LDHKFDLLYSKR,EDLAALEK NMIYGK,FFGNLMDAGR, LSICGEESFGTGSDHIR 2 78 gi ATPase 1 α subunit* 4% gi α tubulin* 20% gi Phosphoglucomutase* 5% Energy production and conversion Cell division and chromosome partitioning Carbohydrate transport Blocked MREVI QGAIS 90 SGAITGITLKDXK 1 67 gi Integrase* 11% Translation, ribosomal structure and biogenesis MLAGT 91 DAGAIAGLDVLR 1 86 gi Hypothetical protein 1% Unknown Unanalyzed 92 VEIIANDQGNR,TTPSYV AFTEGER,TFSPQEISAMV LTK,KAVVTVPAYFNDSQ gi Hypothetical protein 12% Unknown QGLVA R,LDISDDAR,VDDIVLVG GSTR,DAEQFK 93 IINEPTAAAIAYGLDR 1 77 gi Heat shock protein Ssa2* 2% modification, protein Unanalyzed NQVAMNPHNTVFDAKR, FDDAEVQSDMK,IINEPT AAAIAYGLDKK,ATAGDT HLGGEDFDNR,SINPDEA 94 VAYGAAVQAAILSGDTS EK,NTTVPTK,TKDNNLL GK,FELSGIPPAPR,EGIEW LDSNTTASTDELKDK gi Hypothetical protein 20% Unknown Unanalyzed
12 95 96 NQVAMNPHNTVFDAKR, KFDDAEVQSDMK,IINEP TAAAIAYGLDKK,ATAGD THLGGEDFDNR,NTTVPT KK,TKDNNLLGK,FELSGI PPAPR VEIIANDQGNR,NQVAM NPHNTVFDAKR,IINEPTA AAIAYGLDK,ATAGDTHL GGEDFDNR,DLSSNPR,L VSFFNGK,TKDNNLLGK,I TITNDKGR,LTDEK,LADK gi Hypothetical protein 13% Unknown Unanalyzed gi HSS1 (HSP 70)* 16% modification, protein TLDEQADSAAR,LTIGPR, Carbohydrate transport gi Trehalose phosphorylase* 4% EESNS NNHNILQGVASPDLR 98 TPDILNK 1 45 gi Hypothetical protein 1% Unknown Unanalyzed EIISNASDALDK,YESLSD PSKLDSGK,DTGIGMTK,E TLQQNK,LGIHEDAQNR, AQALR LIPVYK,FPTTTIG SFPQTK,GMLTGPVTILN WSFPR,AIQVDEPAIR,EG LPLR,LDADVISIEASK GVSYLK,AIDINK,DVILK, VAGILTVK,FAEEVK,SQF T ITPGSEQVR,KGEANSIITS YNR,CTTDHISAGGPWLK EHAALEPR IHETNLKK 6 96 gi Heat shock protein 90* 8% gi methyltetrahydropteroyltriglut amate-homocysteine S- methyltransferase* 8% gi Aconitase* 10% modification, protein Amino acid transport Energy production and conversion 102 EVLEILKK 1 41 gi Hypothetical protein 0% Unknown Unanalyzed 103 YK,IDVLSNPK,K NALEEYIYDTR gi Pss1* 3% modification, protein Asterisks indicate unique proteins identified from L. edodes fruiting body using 2-DGE and LC-MS/MS analysis. (In case of overlapping among Table 2, Table 3 and 4, astertisks added to protein name prior Table 2 to Table 3 and 4) Commonly identified proteins from 1-DGE and 2-DGE analysis are shown in red letters. Blue letters indicates consensus amino acid sequences between analyzed protein and matched protein. QGEAT QNAEE QASEV QTAAP Blocked
13 Table 3. Phosphoproteins (Lentinula edodes) identified by 2-DGE, Pro-Q DPGS staining, and nesi-lc-ms/ms. The table lists indicated spot numbers, matching peptide, scores, accession numbers, protein names, analytical mass (molecular weight, kda), theoretical pi, internal amino acid sequence and function; left to right. Spot Matching Peptide Score Accession Protein Name Analytical MW Theoretical pi Peptide Functional Category P1 P gi Hypothetical protein P gi Ring zinc finger transcription negative regulator factor* P4 (13) 3 82 gi Translation initiation factor 5A IGSSPVGAAESSTIR,DT GFDEHARIEIADQSR Unknown LIASGQGGGNFSNIR Transcription KNGHVVIK,NRPCKIV DMSTSK,VHLVA IDIFTGK Translation, ribosomal structure and biogenesis P5 (15) 1 45 gi Hypothetical protein EFGIDQK Unknown P6 P7 (22) 1 44 gi Hypothetical protein SFGAEEGGSGK Unknown P8 P9 P10 P11 (56) 4 60 gi Hypothetical protein RADGIHLLNIGK,FTPG SFTNYITR,SFKEPR,VD HQAIR Unknown P12 P13 (60) 1 47 gi Palmitoyltransferase QALISNGLK Lipid metabolism P14 P15 P16 P17 (75) gi ILSTLDR,GVLLYGPPG 26S protease regulatory TGK,TMLVK,VVGSEF modification, protein subunit 6B homolog VQK,YLGEGPR,ENSPA IIFIDEIDAIATK,VIMAT P18 P19 (89) 3 95 gi Phosphoglucomutase NMIYGK,FFGNLMDAG Carbohydrate transport R,LSICGEESFGTGSDHI R P gi Trehalose phosphorylase TLDEQADSAAR,GIPN VIDSYAR,NNHNILQG Carbohydrate transport VASPDLR,IGFFSSTPQ GGGVALMR,SDLVHIA GSPQEEVWK P gi Trehalose phosphorylase P gi Trehalose phosphorylase P23 (99) 6 96 gi Heat shock protein IGFFSSTPQGGGVALM R,NNHNILQGVASPDL R,SDLVHIAGSPQEEV WK,GIPNVIDSYAR,IAL QLSTR Carbohydrate transport TLDEQADSAARK,LTIG PR,IGFFSSTPQGGGVA LMR,NNHNILQGVASP Carbohydrate transport DLR,VRPELPIIYR,GIP NVIDSYAR,IALQLSTR, EGFEVK EIISNASDALDK,YESL SDPSKLDSGK,DTGIG modification, protein MTK,ETLQQNK,LGIHE DAQNR,AQALR Asterisks indicate unique proteins identified from L. edodes fruiting body using 2-DGE and LC-MS/MS analysis. (In case of overlapping among Table 2, Table 3 and 4, astertisks added to protein name prior Table 2 to Table 3 and 4) in parenthesis of Spot correspond to spot number of Figure 3 and Table 2.
14 Table 4. Glycoproteins (Lentinula edodes) identified by 2-DGE, Pro-Q EGGS, and nesi-lc-ms/ms. The table lists indicated spot numbers, matching peptide, scores, accession numbers, protein names, analytical mass (molecular weight, kda), theoretical pi, internal amino acid sequence and function; left to right. Spot Matching Peptide Score Accession Protein Name Analytical MW Theoretical pi Peptide Functional Category G1 (85) 1 34 gi Hypothetical protein DATDPPDK Unknown G2 (80) 3 63 gi Hypothetical protein DYAVDLIPK,SPLMGLFEK,KVI GDPSYFGAGK Unknown G3 (73) 1 52 gi Hypothetical protein KDDGTSAIV LENPK Unknown G4 G gi Hypothetical protein EQFLDDLEQALYASK,DYFGA HTFR Unknown G6 (41) gi protein PETREDSVYLAK,LAEQAER,Y EEMVENMKR,VASSDQELTVE ER,NLLSVAYK,NVIGARR,IVSS IEQK,EESK,GNEAQVSMIK,HLI PSAASGESK,MMGDYHR,YLAE Signal transduction FATGDKR,ESADKSLEAYK,AA SDVAVTELPPTHPIR,LGLALNF SVFYYEILNSPDR,DSTLIMQLL R,DNLTLWTSDQDSADKPAEK G gi Ras protein* TGEGFLLVYSITSR Signal transduction G8 (22) 1 44 gi Hypothetical protein SFGAEEGGSGK Unknown G9 (19) 2 56 gi Manganese superoxide Inorganic ion transport and AATPK,IALQAALK dismutase metabolism G10 (15) 1 46 gi Hypothetical protein VEVDLASIPEGK Unknown G11 (11) G12 (6)
15 Asterisks indicate unique proteins identified from L. edodes fruiting body using 2-DGE and LC-MS/MS analysis. (In case of overlapping among Table 2, Table 3 and 4, astertisks added to protein name prior Table 2 to Table 3 and 4) in parenthesis of Spot correspond to spot number of Figure 3 and Table 2. Commonly identified proteins from 1-DGE and 2-DGE analysis are shown in red letters.
Biological processes. Mitochondrion Metabolic process Catalytic activity Oxidoreductase
 Full name Glyceraldehyde3 phosphate dehydrogenase Succinatesemialdehyde Glutamate dehydrogenase 1, Alcohol dehydrogenase [NADP+] 2',3'cyclicnucleotide 3' phosphodiesterase Dihydropyrimidinaserelated 2
Full name Glyceraldehyde3 phosphate dehydrogenase Succinatesemialdehyde Glutamate dehydrogenase 1, Alcohol dehydrogenase [NADP+] 2',3'cyclicnucleotide 3' phosphodiesterase Dihydropyrimidinaserelated 2
f(x) = x R² = RPKM (M8.MXB) f(x) = x E-014 R² = 1 RPKM (M31.
 14 12 f(x) = 1.633186874x - 21.46732234 R² =.995616541 RPKM (M8.MXA) 1 8 6 4 2 2 4 6 8 1 12 14 RPKM (M8.MXB) 14 12 f(x) =.821767782x - 4.192595677497E-14 R² = 1 RPKM (M31.XA) 1 8 6 4 2 2 4 6 8 1 12 14
14 12 f(x) = 1.633186874x - 21.46732234 R² =.995616541 RPKM (M8.MXA) 1 8 6 4 2 2 4 6 8 1 12 14 RPKM (M8.MXB) 14 12 f(x) =.821767782x - 4.192595677497E-14 R² = 1 RPKM (M31.XA) 1 8 6 4 2 2 4 6 8 1 12 14
Metabolism. Metabolic pathways. BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Metabolism Metabolism is the chemical change of
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Metabolism Metabolism is the chemical change of
Citric acid cycle and respiratory chain. Pavla Balínová
 Citric acid cycle and respiratory chain Pavla Balínová Mitochondria Structure of mitochondria: Outer membrane Inner membrane (folded) Matrix space (mtdna, ribosomes, enzymes of CAC, β-oxidation of FA,
Citric acid cycle and respiratory chain Pavla Balínová Mitochondria Structure of mitochondria: Outer membrane Inner membrane (folded) Matrix space (mtdna, ribosomes, enzymes of CAC, β-oxidation of FA,
Syllabus for BASIC METABOLIC PRINCIPLES
 Syllabus for BASIC METABOLIC PRINCIPLES The video lecture covers basic principles you will need to know for the lectures covering enzymes and metabolism in Principles of Metabolism and elsewhere in the
Syllabus for BASIC METABOLIC PRINCIPLES The video lecture covers basic principles you will need to know for the lectures covering enzymes and metabolism in Principles of Metabolism and elsewhere in the
TCA CYCLE (Citric Acid Cycle)
 TCA CYCLE (Citric Acid Cycle) TCA CYCLE The Citric Acid Cycle is also known as: Kreb s cycle Sir Hans Krebs Nobel prize, 1953 TCA (tricarboxylic acid) cycle The citric acid cycle requires aerobic conditions!!!!
TCA CYCLE (Citric Acid Cycle) TCA CYCLE The Citric Acid Cycle is also known as: Kreb s cycle Sir Hans Krebs Nobel prize, 1953 TCA (tricarboxylic acid) cycle The citric acid cycle requires aerobic conditions!!!!
Protein carbonylation: a marker of oxidative stress damage
 Protein carbonylation: a marker of oxidative stress damage ITN-TREATMENT Metabolic Dysfunctions associated with Pharmacological Treatment of Schizophrenia TREATMENT Protein carbonyl formation: Metal-Catalyzed
Protein carbonylation: a marker of oxidative stress damage ITN-TREATMENT Metabolic Dysfunctions associated with Pharmacological Treatment of Schizophrenia TREATMENT Protein carbonyl formation: Metal-Catalyzed
SP GPI Found in exosomes
 Accession /Name/Functional group Spectr. counts SP GPI Found in exosomes Description Metabolism and energy production (21) CNAG_06770 fructose-bisphosphate aldolase 0.94 - - + gluconeogenesis and glycolysis
Accession /Name/Functional group Spectr. counts SP GPI Found in exosomes Description Metabolism and energy production (21) CNAG_06770 fructose-bisphosphate aldolase 0.94 - - + gluconeogenesis and glycolysis
Electron Transport Chain and Oxidative phosphorylation
 Electron Transport Chain and Oxidative phosphorylation So far we have discussed the catabolism involving oxidation of 6 carbons of glucose to CO 2 via glycolysis and CAC without any oxygen molecule directly
Electron Transport Chain and Oxidative phosphorylation So far we have discussed the catabolism involving oxidation of 6 carbons of glucose to CO 2 via glycolysis and CAC without any oxygen molecule directly
III. Metabolism The Citric Acid Cycle
 Department of Chemistry and Biochemistry University of Lethbridge III. Metabolism The Citric Acid Cycle Slide 1 The Eight Steps of the Citric Acid Cycle Enzymes: 4 dehydrogenases (2 decarboxylation) 3
Department of Chemistry and Biochemistry University of Lethbridge III. Metabolism The Citric Acid Cycle Slide 1 The Eight Steps of the Citric Acid Cycle Enzymes: 4 dehydrogenases (2 decarboxylation) 3
BIOL 158: BIOLOGICAL CHEMISTRY II
 BIOL 158: BIOLOGICAL CHEMISTRY II Lecture 5: Vitamins and Coenzymes Lecturer: Christopher Larbie, PhD Introduction Cofactors bind to the active site and assist in the reaction mechanism Apoenzyme is an
BIOL 158: BIOLOGICAL CHEMISTRY II Lecture 5: Vitamins and Coenzymes Lecturer: Christopher Larbie, PhD Introduction Cofactors bind to the active site and assist in the reaction mechanism Apoenzyme is an
Amino acids. Side chain. -Carbon atom. Carboxyl group. Amino group
 PROTEINS Amino acids Side chain -Carbon atom Amino group Carboxyl group Amino acids Primary structure Amino acid monomers Peptide bond Peptide bond Amino group Carboxyl group Peptide bond N-terminal (
PROTEINS Amino acids Side chain -Carbon atom Amino group Carboxyl group Amino acids Primary structure Amino acid monomers Peptide bond Peptide bond Amino group Carboxyl group Peptide bond N-terminal (
PCB 3023 Exam 4 - Form A First and Last Name
 PCB 3023 Exam 4 - Form A First and Last Name Student ID # (U Number) A Before beginning this exam, please complete the following instructions: 1) Write your name and U number on the first page of this
PCB 3023 Exam 4 - Form A First and Last Name Student ID # (U Number) A Before beginning this exam, please complete the following instructions: 1) Write your name and U number on the first page of this
Citric Acid Cycle: Central Role in Catabolism. Entry of Pyruvate into the TCA cycle
 Citric Acid Cycle: Central Role in Catabolism Stage II of catabolism involves the conversion of carbohydrates, fats and aminoacids into acetylcoa In aerobic organisms, citric acid cycle makes up the final
Citric Acid Cycle: Central Role in Catabolism Stage II of catabolism involves the conversion of carbohydrates, fats and aminoacids into acetylcoa In aerobic organisms, citric acid cycle makes up the final
Chapter 10. 이화작용 : 에너지방출과보존 (Catabolism: Energy Release and Conservation)
 Chapter 10 이화작용 : 에너지방출과보존 (Catabolism: Energy Release and Conservation) 1 Fueling Processes Respiration 1 Most respiration involves use of an electron transport chain As electrons pass through the electron
Chapter 10 이화작용 : 에너지방출과보존 (Catabolism: Energy Release and Conservation) 1 Fueling Processes Respiration 1 Most respiration involves use of an electron transport chain As electrons pass through the electron
Krebs cycle Energy Petr Tůma Eva Samcová
 Krebs cycle Energy - 215 Petr Tůma Eva Samcová Overview of Citric Acid Cycle Key Concepts The citric acid cycle (Krebs cycle) is a multistep catalytic process that converts acetyl groups derived from carbohydrates,
Krebs cycle Energy - 215 Petr Tůma Eva Samcová Overview of Citric Acid Cycle Key Concepts The citric acid cycle (Krebs cycle) is a multistep catalytic process that converts acetyl groups derived from carbohydrates,
Fatty acid breakdown
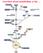 Fatty acids contain a long hydrocarbon chain and a terminal carboxylate group. Most contain between 14 and 24 carbon atoms. The chains may be saturated or contain double bonds. The complete oxidation of
Fatty acids contain a long hydrocarbon chain and a terminal carboxylate group. Most contain between 14 and 24 carbon atoms. The chains may be saturated or contain double bonds. The complete oxidation of
Multiple choice: Circle the best answer on this exam. There are 12 multiple choice questions, each question is worth 3 points.
 CHEM 4420 Exam 4 Spring 2015 Dr. Stone Page 1 of 6 Name Use complete sentences when requested. There are 120 possible points on this exam. Therefore there are 20 bonus points. Multiple choice: Circle the
CHEM 4420 Exam 4 Spring 2015 Dr. Stone Page 1 of 6 Name Use complete sentences when requested. There are 120 possible points on this exam. Therefore there are 20 bonus points. Multiple choice: Circle the
Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function
 Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function 1 BPSL0024 26223 26621 + LrgA family BPSL0025 26690 27412 + hypothetical
Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function 1 BPSL0024 26223 26621 + LrgA family BPSL0025 26690 27412 + hypothetical
7 Pathways That Harvest Chemical Energy
 7 Pathways That Harvest Chemical Energy Pathways That Harvest Chemical Energy How Does Glucose Oxidation Release Chemical Energy? What Are the Aerobic Pathways of Glucose Metabolism? How Is Energy Harvested
7 Pathways That Harvest Chemical Energy Pathways That Harvest Chemical Energy How Does Glucose Oxidation Release Chemical Energy? What Are the Aerobic Pathways of Glucose Metabolism? How Is Energy Harvested
Biology 638 Biochemistry II Exam-2
 Biology 638 Biochemistry II Exam-2 Biol 638, Exam-2 (Code-1) 1. Assume that 16 glucose molecules enter into a liver cell and are attached to a liner glycogen one by one. Later, this glycogen is broken-down
Biology 638 Biochemistry II Exam-2 Biol 638, Exam-2 (Code-1) 1. Assume that 16 glucose molecules enter into a liver cell and are attached to a liner glycogen one by one. Later, this glycogen is broken-down
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/2/e1602038/dc1 Supplementary Materials for Mitochondrial metabolic regulation by GRP78 Manoj Prasad, Kevin J. Pawlak, William E. Burak, Elizabeth E. Perry, Brendan
advances.sciencemag.org/cgi/content/full/3/2/e1602038/dc1 Supplementary Materials for Mitochondrial metabolic regulation by GRP78 Manoj Prasad, Kevin J. Pawlak, William E. Burak, Elizabeth E. Perry, Brendan
BY: RASAQ NURUDEEN OLAJIDE
 BY: RASAQ NURUDEEN OLAJIDE LECTURE CONTENT INTRODUCTION CITRIC ACID CYCLE (T.C.A) PRODUCTION OF ACETYL CoA REACTIONS OF THE CITIRC ACID CYCLE THE AMPHIBOLIC NATURE OF THE T.C.A CYCLE THE GLYOXYLATE CYCLE
BY: RASAQ NURUDEEN OLAJIDE LECTURE CONTENT INTRODUCTION CITRIC ACID CYCLE (T.C.A) PRODUCTION OF ACETYL CoA REACTIONS OF THE CITIRC ACID CYCLE THE AMPHIBOLIC NATURE OF THE T.C.A CYCLE THE GLYOXYLATE CYCLE
ANSC/NUTR 618 Lipids & Lipid Metabolism
 I. Overall concepts A. Definitions ANC/NUTR 618 Lipids & Lipid Metabolism 1. De novo synthesis = synthesis from non-fatty acid precursors a. Carbohydrate precursors (glucose, lactate, and pyruvate) b.
I. Overall concepts A. Definitions ANC/NUTR 618 Lipids & Lipid Metabolism 1. De novo synthesis = synthesis from non-fatty acid precursors a. Carbohydrate precursors (glucose, lactate, and pyruvate) b.
CHY2026: General Biochemistry UNIT 7& 8: CARBOHYDRATE METABOLISM
 CHY2026: General Biochemistry UNIT 7& 8: CARBOHYDRATE METABOLISM Metabolism Bioenergetics is the transfer and utilization of energy in biological systems The direction and extent to which a chemical reaction
CHY2026: General Biochemistry UNIT 7& 8: CARBOHYDRATE METABOLISM Metabolism Bioenergetics is the transfer and utilization of energy in biological systems The direction and extent to which a chemical reaction
Coenzymes, vitamins and trace elements 209. Petr Tůma Eva Samcová
 Coenzymes, vitamins and trace elements 209 Petr Tůma Eva Samcová History and nomenclature of enzymes 1810, Gay-Lussac made an experiment with yeats alter saccharide to ethanol and CO 2 Fermentation From
Coenzymes, vitamins and trace elements 209 Petr Tůma Eva Samcová History and nomenclature of enzymes 1810, Gay-Lussac made an experiment with yeats alter saccharide to ethanol and CO 2 Fermentation From
CITRIC ACID CYCLE ERT106 BIOCHEMISTRY SEM /19 BY: MOHAMAD FAHRURRAZI TOMPANG
 CITRIC ACID CYCLE ERT106 BIOCHEMISTRY SEM 1 2018/19 BY: MOHAMAD FAHRURRAZI TOMPANG Chapter Outline (19-1) The central role of the citric acid cycle in metabolism (19-2) The overall pathway of the citric
CITRIC ACID CYCLE ERT106 BIOCHEMISTRY SEM 1 2018/19 BY: MOHAMAD FAHRURRAZI TOMPANG Chapter Outline (19-1) The central role of the citric acid cycle in metabolism (19-2) The overall pathway of the citric
Pathway overview. Glucose + 2NAD + + 2ADP +2Pi 2NADH + 2pyruvate + 2ATP + 2H 2 O + 4H +
 Glycolysis Glycolysis The conversion of glucose to pyruvate to yield 2ATP molecules 10 enzymatic steps Chemical interconversion steps Mechanisms of enzyme conversion and intermediates Energetics of conversions
Glycolysis Glycolysis The conversion of glucose to pyruvate to yield 2ATP molecules 10 enzymatic steps Chemical interconversion steps Mechanisms of enzyme conversion and intermediates Energetics of conversions
Biological oxidation II. The Cytric acid cycle
 Biological oxidation II The Cytric acid cycle Outline The Cytric acid cycle (TCA tricarboxylic acid) Central role of Acetyl-CoA Regulation of the TCA cycle Anaplerotic reactions The Glyoxylate cycle Localization
Biological oxidation II The Cytric acid cycle Outline The Cytric acid cycle (TCA tricarboxylic acid) Central role of Acetyl-CoA Regulation of the TCA cycle Anaplerotic reactions The Glyoxylate cycle Localization
III. 6. Test. Respiració cel lular
 III. 6. Test. Respiració cel lular Chapter Questions 1) What is the term for metabolic pathways that release stored energy by breaking down complex molecules? A) anabolic pathways B) catabolic pathways
III. 6. Test. Respiració cel lular Chapter Questions 1) What is the term for metabolic pathways that release stored energy by breaking down complex molecules? A) anabolic pathways B) catabolic pathways
Chapter 14. Energy conversion: Energy & Behavior
 Chapter 14 Energy conversion: Energy & Behavior Why do you Eat and Breath? To generate ATP Foods, Oxygen, and Mitochodria Cells Obtain Energy by the Oxidation of Organic Molecules Food making ATP making
Chapter 14 Energy conversion: Energy & Behavior Why do you Eat and Breath? To generate ATP Foods, Oxygen, and Mitochodria Cells Obtain Energy by the Oxidation of Organic Molecules Food making ATP making
Mitochondria and ATP Synthesis
 Mitochondria and ATP Synthesis Mitochondria and ATP Synthesis 1. Mitochondria are sites of ATP synthesis in cells. 2. ATP is used to do work; i.e. ATP is an energy source. 3. ATP hydrolysis releases energy
Mitochondria and ATP Synthesis Mitochondria and ATP Synthesis 1. Mitochondria are sites of ATP synthesis in cells. 2. ATP is used to do work; i.e. ATP is an energy source. 3. ATP hydrolysis releases energy
Vocabulary. Chapter 19: The Citric Acid Cycle
 Vocabulary Amphibolic: able to be a part of both anabolism and catabolism Anaplerotic: referring to a reaction that ensures an adequate supply of an important metabolite Citrate Synthase: the enzyme that
Vocabulary Amphibolic: able to be a part of both anabolism and catabolism Anaplerotic: referring to a reaction that ensures an adequate supply of an important metabolite Citrate Synthase: the enzyme that
Yield of energy from glucose
 Paper : Module : 05 Yield of Energy from Glucose Principal Investigator, Paper Coordinator and Content Writer Prof. Ramesh Kothari, Professor Dept. of Biosciences, Saurashtra University, Rajkot - 360005
Paper : Module : 05 Yield of Energy from Glucose Principal Investigator, Paper Coordinator and Content Writer Prof. Ramesh Kothari, Professor Dept. of Biosciences, Saurashtra University, Rajkot - 360005
2. (12 pts) Given the following metabolic pathway (as it occurs in the cell):
 Answer Sheet 1 (Gold) 1. (1 pt) Write your exam ID (A) in the blank at the upper right of your answer sheet. 2. (12 pts) Given the following metabolic pathway (as it occurs in the cell): a. Would you expect
Answer Sheet 1 (Gold) 1. (1 pt) Write your exam ID (A) in the blank at the upper right of your answer sheet. 2. (12 pts) Given the following metabolic pathway (as it occurs in the cell): a. Would you expect
This is an example outline of 3 lectures in BSC (Thanks to Dr. Ellington for sharing this information.)
 This is an example outline of 3 lectures in BSC 2010. (Thanks to Dr. Ellington for sharing this information.) Topic 10: CELLULAR RESPIRATION (lectures 14-16) OBJECTIVES: 1. Know the basic reactions that
This is an example outline of 3 lectures in BSC 2010. (Thanks to Dr. Ellington for sharing this information.) Topic 10: CELLULAR RESPIRATION (lectures 14-16) OBJECTIVES: 1. Know the basic reactions that
University of Guelph Department of Chemistry and Biochemistry Structure and Function In Biochemistry
 University of Guelph Department of Chemistry and Biochemistry 19-356 Structure and Function In Biochemistry Final Exam, April 21, 1997. Time allowed, 120 min. Answer questions 1-30 on the computer scoring
University of Guelph Department of Chemistry and Biochemistry 19-356 Structure and Function In Biochemistry Final Exam, April 21, 1997. Time allowed, 120 min. Answer questions 1-30 on the computer scoring
CELLULAR METABOLISM. Metabolic pathways can be linear, branched, cyclic or spiral
 CHM333 LECTURE 24 & 25: 3/27 29/13 SPRING 2013 Professor Christine Hrycyna CELLULAR METABOLISM What is metabolism? - How cells acquire, transform, store and use energy - Study reactions in a cell and how
CHM333 LECTURE 24 & 25: 3/27 29/13 SPRING 2013 Professor Christine Hrycyna CELLULAR METABOLISM What is metabolism? - How cells acquire, transform, store and use energy - Study reactions in a cell and how
Bacterial cellulose synthesis mechanism of facultative
 1 2 3 4 Bacterial cellulose synthesis mechanism of facultative anaerobe Kaihua Ji 1 +, Wei Wang 2, 3 +, Bing Zeng 1, Sibin Chen 1, Qianqian Zhao 4, Yueqing Chen 1, Guoqiang Li 1* and Ting Ma 1* 5 6 7 Figure
1 2 3 4 Bacterial cellulose synthesis mechanism of facultative anaerobe Kaihua Ji 1 +, Wei Wang 2, 3 +, Bing Zeng 1, Sibin Chen 1, Qianqian Zhao 4, Yueqing Chen 1, Guoqiang Li 1* and Ting Ma 1* 5 6 7 Figure
Module No. # 01 Lecture No. # 19 TCA Cycle
 Biochemical Engineering Prof. Dr. Rintu Banerjee Department of Agricultural and Food Engineering Asst. Prof. Dr. Saikat Chakraborty Department of Chemical Engineering Indian Institute of Technology, Kharagpur
Biochemical Engineering Prof. Dr. Rintu Banerjee Department of Agricultural and Food Engineering Asst. Prof. Dr. Saikat Chakraborty Department of Chemical Engineering Indian Institute of Technology, Kharagpur
Marah Bitar. Faisal Nimri ... Nafeth Abu Tarboosh
 8 Marah Bitar Faisal Nimri... Nafeth Abu Tarboosh Summary of the 8 steps of citric acid cycle Step 1. Acetyl CoA joins with a four-carbon molecule, oxaloacetate, releasing the CoA group and forming a six-carbon
8 Marah Bitar Faisal Nimri... Nafeth Abu Tarboosh Summary of the 8 steps of citric acid cycle Step 1. Acetyl CoA joins with a four-carbon molecule, oxaloacetate, releasing the CoA group and forming a six-carbon
WT siz1-2 siz1-3. WT siz1-2 siz1-3. -Pi, 0.05 M IAA. -Pi, 2.5 M NPA
 A WT siz1-2 siz1-3 B WT siz1-2 siz1-3 -Pi C +Pi WT siz1-2 siz1-3 D WT siz1-2 siz1-3 E +Pi, 0.05 M IAA -Pi, 0.05 M IAA WT siz1-2 siz1-3 WT siz1-2 siz1-3 F +Pi, 2.5 M NPA -Pi, 2.5 M NPA Supplemental Figure
A WT siz1-2 siz1-3 B WT siz1-2 siz1-3 -Pi C +Pi WT siz1-2 siz1-3 D WT siz1-2 siz1-3 E +Pi, 0.05 M IAA -Pi, 0.05 M IAA WT siz1-2 siz1-3 WT siz1-2 siz1-3 F +Pi, 2.5 M NPA -Pi, 2.5 M NPA Supplemental Figure
7/5/2014. Microbial. Metabolism. Basic Chemical Reactions Underlying. Metabolism. Metabolism: Overview
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University Basic Chemical Reactions Underlying Metabolism Metabolism C H A P T E R 5 Microbial Metabolism Collection
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University Basic Chemical Reactions Underlying Metabolism Metabolism C H A P T E R 5 Microbial Metabolism Collection
METABOLISM Biosynthetic Pathways
 METABOLISM Biosynthetic Pathways Metabolism Metabolism involves : Catabolic reactions that break down large, complex molecules to provide energy and smaller molecules. Anabolic reactions that use ATP energy
METABOLISM Biosynthetic Pathways Metabolism Metabolism involves : Catabolic reactions that break down large, complex molecules to provide energy and smaller molecules. Anabolic reactions that use ATP energy
Lecture 15. Signal Transduction Pathways - Introduction
 Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
INTRODUCTORY BIOCHEMISTRY. BI 28 Second Midterm Examination April 3, 2007
 INTRODUCTORY BIOCHEMISTRY BI 28 Second Midterm Examination April 3, 2007 Name SIS # Make sure that your name or SIS # is on every page. This is the only way we have of matching you with your exam after
INTRODUCTORY BIOCHEMISTRY BI 28 Second Midterm Examination April 3, 2007 Name SIS # Make sure that your name or SIS # is on every page. This is the only way we have of matching you with your exam after
Midterm 2. Low: 14 Mean: 61.3 High: 98. Standard Deviation: 17.7
 Midterm 2 Low: 14 Mean: 61.3 High: 98 Standard Deviation: 17.7 Lecture 17 Amino Acid Metabolism Review of Urea Cycle N and S assimilation Last cofactors: THF and SAM Synthesis of few amino acids Dietary
Midterm 2 Low: 14 Mean: 61.3 High: 98 Standard Deviation: 17.7 Lecture 17 Amino Acid Metabolism Review of Urea Cycle N and S assimilation Last cofactors: THF and SAM Synthesis of few amino acids Dietary
BIO 5099: Molecular Biology for Computer Scientists (et al)
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
PRINT your Name Student (FAMILY, first name) Midterm 7:00 P.M.
 PRINT your Name Student No. (FAMILY, first name) BIOCHEMISTRY 311A VERSION 1 (ONE) Midterm 7:00 P.M. Examiners: Dr. R. E. MacKenzie (69%) Dr. A. Storer (18%) Dr. W. Mushynski (13%) READ THE QUESTIONS CAREFULLY!!
PRINT your Name Student No. (FAMILY, first name) BIOCHEMISTRY 311A VERSION 1 (ONE) Midterm 7:00 P.M. Examiners: Dr. R. E. MacKenzie (69%) Dr. A. Storer (18%) Dr. W. Mushynski (13%) READ THE QUESTIONS CAREFULLY!!
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site.
 Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
Biol 219 Lec 7 Fall 2016
 Cellular Respiration: Harvesting Energy to form ATP Cellular Respiration and Metabolism Glucose ATP Pyruvate Lactate Acetyl CoA NAD + Introducing The Players primary substrate for cellular respiration
Cellular Respiration: Harvesting Energy to form ATP Cellular Respiration and Metabolism Glucose ATP Pyruvate Lactate Acetyl CoA NAD + Introducing The Players primary substrate for cellular respiration
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being A Eukaryote. Eukaryotic Cells. Basic eukaryotes have:
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
Link download full of Test Bank for Fundamentals of Biochemistry 4th Edition by Voet
 Link download full of Test Bank for Fundamentals of Biochemistry 4th Edition by Voet http://testbankair.com/download/test-bank-for-fundamentals-ofbiochemistry-4th-edition-by-voet/ Chapter 16: Glycogen
Link download full of Test Bank for Fundamentals of Biochemistry 4th Edition by Voet http://testbankair.com/download/test-bank-for-fundamentals-ofbiochemistry-4th-edition-by-voet/ Chapter 16: Glycogen
Integration of Metabolism
 Integration of Metabolism Metabolism is a continuous process. Thousands of reactions occur simultaneously in order to maintain homeostasis. It ensures a supply of fuel, to tissues at all times, in fed
Integration of Metabolism Metabolism is a continuous process. Thousands of reactions occur simultaneously in order to maintain homeostasis. It ensures a supply of fuel, to tissues at all times, in fed
Final Review Sessions. 3/16 (FRI) 126 Wellman (4-6 6 pm) 3/19 (MON) 1309 Surge 3 (4-6 6 pm) Office Hours
 Final Review Sessions 3/16 (FRI) 126 Wellman (4-6 6 pm) 3/19 (MON) 1309 Surge 3 (4-6 6 pm) Office ours 3/14 (WED) 9:30 11:30 am (Rebecca) 3/16 (FRI) 9-11 am (Abel) Final ESSENTIALS Posted Lecture 20 ormonal
Final Review Sessions 3/16 (FRI) 126 Wellman (4-6 6 pm) 3/19 (MON) 1309 Surge 3 (4-6 6 pm) Office ours 3/14 (WED) 9:30 11:30 am (Rebecca) 3/16 (FRI) 9-11 am (Abel) Final ESSENTIALS Posted Lecture 20 ormonal
Microbial Metabolism. Chapter 7. Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
 Microbial Metabolism Chapter 7 Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Metabolism and the Role of Enzymes Metabolism: pertains to all chemical reactions
Microbial Metabolism Chapter 7 Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Metabolism and the Role of Enzymes Metabolism: pertains to all chemical reactions
4. Which step shows a split of one molecule into two smaller molecules? a. 2. d. 5
 1. Which of the following statements about NAD + is false? a. NAD + is reduced to NADH during both glycolysis and the citric acid cycle. b. NAD + has more chemical energy than NADH. c. NAD + is reduced
1. Which of the following statements about NAD + is false? a. NAD + is reduced to NADH during both glycolysis and the citric acid cycle. b. NAD + has more chemical energy than NADH. c. NAD + is reduced
Aerobic Respiration. The four stages in the breakdown of glucose
 Aerobic Respiration The four stages in the breakdown of glucose 1 I. Aerobic Respiration Why can t we break down Glucose in one step? (Flaming Gummy Bear) Enzymes gently lower the potential energy until
Aerobic Respiration The four stages in the breakdown of glucose 1 I. Aerobic Respiration Why can t we break down Glucose in one step? (Flaming Gummy Bear) Enzymes gently lower the potential energy until
Lecture #27 Lecturer A. N. Koval
 Lecture #27 Lecturer A. N. Koval Hormones Transduce Signals to Affect Homeostatic Mechanisms Koval A. (C), 2011 2 Lipophilic hormones Classifying hormones into hydrophilic and lipophilic molecules indicates
Lecture #27 Lecturer A. N. Koval Hormones Transduce Signals to Affect Homeostatic Mechanisms Koval A. (C), 2011 2 Lipophilic hormones Classifying hormones into hydrophilic and lipophilic molecules indicates
Glycolysis. Cellular Respiration
 Glucose is the preferred carbohydrate of cells. In solution, it can change from a linear chain to a ring. Energy is stored in the bonds of the carbohydrates. Breaking these bonds releases that energy.
Glucose is the preferred carbohydrate of cells. In solution, it can change from a linear chain to a ring. Energy is stored in the bonds of the carbohydrates. Breaking these bonds releases that energy.
Moving Proteins into Membranes and Organelles
 13 Moving Proteins into Membranes and Organelles Review the Concepts 1. In eukaryotes, protein translocation across the endoplasmic reticulum (ER) membrane is most commonly cotranslational; it can also
13 Moving Proteins into Membranes and Organelles Review the Concepts 1. In eukaryotes, protein translocation across the endoplasmic reticulum (ER) membrane is most commonly cotranslational; it can also
MOLECULAR CELL BIOLOGY
 1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
The citric acid cycle Sitruunahappokierto Citronsyracykeln
 The citric acid cycle Sitruunahappokierto Citronsyracykeln Ove Eriksson BLL/Biokemia ove.eriksson@helsinki.fi Metabolome: The complete set of small-molecule metabolites to be found in a cell or an organism.
The citric acid cycle Sitruunahappokierto Citronsyracykeln Ove Eriksson BLL/Biokemia ove.eriksson@helsinki.fi Metabolome: The complete set of small-molecule metabolites to be found in a cell or an organism.
Comparison of nanoparticle protein corona profiles resulting from changes in nanoparticle environment and engineered properties.
 Comparison of nanoparticle protein corona profiles resulting from changes in nanoparticle environment and engineered properties. Korin Wheeler Dept. of Chemistry & Biochemistry Santa Clara University at
Comparison of nanoparticle protein corona profiles resulting from changes in nanoparticle environment and engineered properties. Korin Wheeler Dept. of Chemistry & Biochemistry Santa Clara University at
CHE 242 Exam 3 Practice Questions
 CHE 242 Exam 3 Practice Questions Glucose metabolism 1. Below is depicted glucose catabolism. Indicate on the pathways the following: A) which reaction(s) of glycolysis are irreversible B) where energy
CHE 242 Exam 3 Practice Questions Glucose metabolism 1. Below is depicted glucose catabolism. Indicate on the pathways the following: A) which reaction(s) of glycolysis are irreversible B) where energy
CELLS. Cells. Basic unit of life (except virus)
 Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled
 Protein Targeting Objectives 1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled As a protein is being synthesized, decisions
Protein Targeting Objectives 1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled As a protein is being synthesized, decisions
Chapter 3 Part 2! Pages (10 th and 11 th eds.)! The Cellular Level of Organization! Cellular Organelles and Protein Synthesis!
 Chapter 3 Part 2! Pages 65 89 (10 th and 11 th eds.)! The Cellular Level of Organization! Cellular Organelles and Protein Synthesis! The Cell Theory! Living organisms are composed of one or more cells.!
Chapter 3 Part 2! Pages 65 89 (10 th and 11 th eds.)! The Cellular Level of Organization! Cellular Organelles and Protein Synthesis! The Cell Theory! Living organisms are composed of one or more cells.!
respiration mitochondria mitochondria metabolic pathways reproduction can fuse or split DRP1 interacts with ER tubules chapter DRP1 ER tubule
 mitochondria respiration chapter 3-4 shape highly variable can fuse or split structure outer membrane inner membrane cristae intermembrane space mitochondrial matrix free ribosomes respiratory enzymes
mitochondria respiration chapter 3-4 shape highly variable can fuse or split structure outer membrane inner membrane cristae intermembrane space mitochondrial matrix free ribosomes respiratory enzymes
Biosynthesis of Fatty Acids
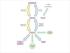 Biosynthesis of Fatty Acids Fatty acid biosynthesis takes place in the cytosol rather than the mitochondria and requires a different activation mechanism and different enzymes and coenzymes than fatty
Biosynthesis of Fatty Acids Fatty acid biosynthesis takes place in the cytosol rather than the mitochondria and requires a different activation mechanism and different enzymes and coenzymes than fatty
LIPID METABOLISM
 LIPID METABOLISM LIPOGENESIS LIPOGENESIS LIPOGENESIS FATTY ACID SYNTHESIS DE NOVO FFA in the blood come from :- (a) Dietary fat (b) Dietary carbohydrate/protein in excess of need FA TAG Site of synthesis:-
LIPID METABOLISM LIPOGENESIS LIPOGENESIS LIPOGENESIS FATTY ACID SYNTHESIS DE NOVO FFA in the blood come from :- (a) Dietary fat (b) Dietary carbohydrate/protein in excess of need FA TAG Site of synthesis:-
AP Bio Photosynthesis & Respiration
 AP Bio Photosynthesis & Respiration Multiple Choice Identify the letter of the choice that best completes the statement or answers the question. 1. What is the term used for the metabolic pathway in which
AP Bio Photosynthesis & Respiration Multiple Choice Identify the letter of the choice that best completes the statement or answers the question. 1. What is the term used for the metabolic pathway in which
Cellular Respiration
 Cellular I can describe cellular respiration Cellular respiration is a series of metabolic pathways releasing energy from a foodstuff e.g. glucose. This yields energy in the form of ATP adenosine P i P
Cellular I can describe cellular respiration Cellular respiration is a series of metabolic pathways releasing energy from a foodstuff e.g. glucose. This yields energy in the form of ATP adenosine P i P
Signal Transduction Cascades
 Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Chapter 14 - Electron Transport and Oxidative Phosphorylation
 Chapter 14 - Electron Transport and Oxidative Phosphorylation The cheetah, whose capacity for aerobic metabolism makes it one of the fastest animals Prentice Hall c2002 Chapter 14 1 14.4 Oxidative Phosphorylation
Chapter 14 - Electron Transport and Oxidative Phosphorylation The cheetah, whose capacity for aerobic metabolism makes it one of the fastest animals Prentice Hall c2002 Chapter 14 1 14.4 Oxidative Phosphorylation
BIOLOGY 103 Spring 2001 MIDTERM LAB SECTION
 BIOLOGY 103 Spring 2001 MIDTERM NAME KEY LAB SECTION ID# (last four digits of SS#) STUDENT PLEASE READ. Do not put yourself at a disadvantage by revealing the content of this exam to your classmates. Your
BIOLOGY 103 Spring 2001 MIDTERM NAME KEY LAB SECTION ID# (last four digits of SS#) STUDENT PLEASE READ. Do not put yourself at a disadvantage by revealing the content of this exam to your classmates. Your
MCB 102 Midterm II October 30, 2003
 MCB 102 Midterm II October 30, 2003 NAME Student ID Signature TA Relax. There is ample time to finish everything. Only exams answered by using non-erasable ink qualify for possible re-grading. I. (15)
MCB 102 Midterm II October 30, 2003 NAME Student ID Signature TA Relax. There is ample time to finish everything. Only exams answered by using non-erasable ink qualify for possible re-grading. I. (15)
number Done by Corrected by Doctor Nayef Karadsheh
 number 11 Done by حسام أبو عوض Corrected by Moayyad Al-Shafei Doctor Nayef Karadsheh 1 P a g e General Regulatory Aspects in Metabolism: We can divide all pathways in metabolism to catabolicand anabolic.
number 11 Done by حسام أبو عوض Corrected by Moayyad Al-Shafei Doctor Nayef Karadsheh 1 P a g e General Regulatory Aspects in Metabolism: We can divide all pathways in metabolism to catabolicand anabolic.
Answer three from questions 5, 6, 7, 8, and 9.
 BCH 4053 May 1, 2003 FINAL EXAM NAME There are 9 pages and 9 questions on the exam. nly five are to be answered, each worth 20 points. Answer two from questions 1, 2, 3, and 4 Answer three from questions
BCH 4053 May 1, 2003 FINAL EXAM NAME There are 9 pages and 9 questions on the exam. nly five are to be answered, each worth 20 points. Answer two from questions 1, 2, 3, and 4 Answer three from questions
Aerobic Fate of Pyruvate. Chapter 16 Homework Assignment. Chapter 16 The Citric Acid Cycle
 Chapter 16 Homework Assignment The following problems will be due once we finish the chapter: 1, 3, 7, 10, 16, 19, 20 Additional Problem: Write out the eight reaction steps of the Citric Acid Cycle, using
Chapter 16 Homework Assignment The following problems will be due once we finish the chapter: 1, 3, 7, 10, 16, 19, 20 Additional Problem: Write out the eight reaction steps of the Citric Acid Cycle, using
CHAPTER 24: Carbohydrate, Lipid, & Protein Metabolism. General, Organic, & Biological Chemistry Janice Gorzynski Smith
 CHAPTER 24: Carbohydrate, Lipid, & Protein Metabolism General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 24: Carbohydrate, Lipid, & Protein Metabolism Learning Objectives: q Role in
CHAPTER 24: Carbohydrate, Lipid, & Protein Metabolism General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 24: Carbohydrate, Lipid, & Protein Metabolism Learning Objectives: q Role in
Metabolic Biochemistry / BIBC 102 Midterm Exam / Spring 2005
 Metabolic Biochemistry / BIBC 102 Midterm Exam / Spring 2005 I. (20 points) Fill in all of the enzyme catalyzed reactions which convert glycogen to lactate. Draw the correct structure for each intermediate
Metabolic Biochemistry / BIBC 102 Midterm Exam / Spring 2005 I. (20 points) Fill in all of the enzyme catalyzed reactions which convert glycogen to lactate. Draw the correct structure for each intermediate
Biochemistry 2 Recita0on Amino Acid Metabolism
 Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
Chapter 7 Cellular Respiration and Fermentation*
 Chapter 7 Cellular Respiration and Fermentation* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. Life Is Work
Chapter 7 Cellular Respiration and Fermentation* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. Life Is Work
Glycolysis Part 2. BCH 340 lecture 4
 Glycolysis Part 2 BCH 340 lecture 4 Regulation of Glycolysis There are three steps in glycolysis that have enzymes which regulate the flux of glycolysis These enzymes catalyzes irreversible reactions of
Glycolysis Part 2 BCH 340 lecture 4 Regulation of Glycolysis There are three steps in glycolysis that have enzymes which regulate the flux of glycolysis These enzymes catalyzes irreversible reactions of
CH395G FINAL (3 rd ) EXAM Kitto/Hackert - Fall 2003
 CH395G FINAL (3 rd ) EXAM Kitto/Hackert - Fall 2003 1. A cell in an active, catabolic state has a. a high (ATP/ADP) and a high (NADH/NAD + ) ratio b. a high (ATP/ADP) and a low (NADH/NAD + ) ratio c. a
CH395G FINAL (3 rd ) EXAM Kitto/Hackert - Fall 2003 1. A cell in an active, catabolic state has a. a high (ATP/ADP) and a high (NADH/NAD + ) ratio b. a high (ATP/ADP) and a low (NADH/NAD + ) ratio c. a
Midterm 2 Results. Standard Deviation:
 Midterm 2 Results High: Low: Mean: Standard Deviation: 97.5% 16% 58% 16.3 Lecture 17 Amino Acid Metabolism Urea Cycle N and S assimilation Last cofactors: THF and SAM Dietary (Exogenous) Proteins Hydrolyzed
Midterm 2 Results High: Low: Mean: Standard Deviation: 97.5% 16% 58% 16.3 Lecture 17 Amino Acid Metabolism Urea Cycle N and S assimilation Last cofactors: THF and SAM Dietary (Exogenous) Proteins Hydrolyzed
Metabolism. Chapter 5. Catabolism Drives Anabolism 8/29/11. Complete Catabolism of Glucose
 8/29/11 Metabolism Chapter 5 All of the reactions in the body that require energy transfer. Can be divided into: Cell Respiration and Metabolism Anabolism: requires the input of energy to synthesize large
8/29/11 Metabolism Chapter 5 All of the reactions in the body that require energy transfer. Can be divided into: Cell Respiration and Metabolism Anabolism: requires the input of energy to synthesize large
Lecture 16. Finish lipid metabolism (Triglycerides, Isoprenoids/Steroids, Glyoxylate cycle) Amino acid metabolism (Urea cycle) Google Man III
 Lecture 16 Finish lipid metabolism (Triglycerides, Isoprenoids/Steroids, Glyoxylate cycle) Amino acid metabolism (Urea cycle) Google Man III The Powertrain of Human Metabolism (verview) CARBHYDRATES PRTEINS
Lecture 16 Finish lipid metabolism (Triglycerides, Isoprenoids/Steroids, Glyoxylate cycle) Amino acid metabolism (Urea cycle) Google Man III The Powertrain of Human Metabolism (verview) CARBHYDRATES PRTEINS
Carbohydrate Metabolism
 OpenStax-CNX module: m46451 1 Carbohydrate Metabolism OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
OpenStax-CNX module: m46451 1 Carbohydrate Metabolism OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
Supplementary Information. Deciphering the Venomic Transcriptome of Killer- Wasp Vespa velutina
 Supplementary Information Deciphering the Venomic Transcriptome of Killer- Wasp Vespa velutina Zhirui Liu, Shuanggang Chen, You Zhou, Cuihong Xie, Bifeng Zhu, Huming Zhu, Shupeng Liu, Wei Wang, Hongzhuan
Supplementary Information Deciphering the Venomic Transcriptome of Killer- Wasp Vespa velutina Zhirui Liu, Shuanggang Chen, You Zhou, Cuihong Xie, Bifeng Zhu, Huming Zhu, Shupeng Liu, Wei Wang, Hongzhuan
Regulation. 1. Short term control 8-1
 Regulation Several aspects of regulation have been alluded to or described in detail as we have progressed through the various sections of the course. These include: (a) compartmentation: This was not
Regulation Several aspects of regulation have been alluded to or described in detail as we have progressed through the various sections of the course. These include: (a) compartmentation: This was not
MITOCHONDRIA LECTURES OVERVIEW
 1 MITOCHONDRIA LECTURES OVERVIEW A. MITOCHONDRIA LECTURES OVERVIEW Mitochondrial Structure The arrangement of membranes: distinct inner and outer membranes, The location of ATPase, DNA and ribosomes The
1 MITOCHONDRIA LECTURES OVERVIEW A. MITOCHONDRIA LECTURES OVERVIEW Mitochondrial Structure The arrangement of membranes: distinct inner and outer membranes, The location of ATPase, DNA and ribosomes The
True or False: 1. Reactions are called endergonic if they occur spontaneously and release free energy.
 True or False: 1. Reactions are called endergonic if they occur spontaneously and release free energy. 2. Enzymes catalyze chemical reactions by lowering the activation energy 3. Biochemical pathways are
True or False: 1. Reactions are called endergonic if they occur spontaneously and release free energy. 2. Enzymes catalyze chemical reactions by lowering the activation energy 3. Biochemical pathways are
Fatty acids synthesis
 Fatty acids synthesis The synthesis start from Acetyl COA the first step requires ATP + reducing power NADPH! even though the oxidation and synthesis are different pathways but from chemical part of view
Fatty acids synthesis The synthesis start from Acetyl COA the first step requires ATP + reducing power NADPH! even though the oxidation and synthesis are different pathways but from chemical part of view
The Citric acid cycle. The Citric Acid Cycle II 11/17/2009. Overview. Pyruvate dehydrogenase
 The itric acid cycle The itric Acid ycle II 11/17/2009 It is called the Krebs cycle or the tricarboxylic and is the hub of the metabolic system. It accounts for the majority of carbohydrate, fatty acid
The itric acid cycle The itric Acid ycle II 11/17/2009 It is called the Krebs cycle or the tricarboxylic and is the hub of the metabolic system. It accounts for the majority of carbohydrate, fatty acid
Biology 638 Biochemistry II Exam-3. (Note that you are not allowed to use any calculator)
 Biology 638 Biochemistry II Exam-3 (Note that you are not allowed to use any calculator) 1. In the non-cyclic pathway, electron pathway is. Select the most accurate one. a. PSII PC Cyt b 6 f PC PSI Fd-NADP
Biology 638 Biochemistry II Exam-3 (Note that you are not allowed to use any calculator) 1. In the non-cyclic pathway, electron pathway is. Select the most accurate one. a. PSII PC Cyt b 6 f PC PSI Fd-NADP
Electron transport chain chapter 6 (page 73) BCH 340 lecture 6
 Electron transport chain chapter 6 (page 73) BCH 340 lecture 6 The Metabolic Pathway of Cellular Respiration All of the reactions involved in cellular respiration can be grouped into three main stages
Electron transport chain chapter 6 (page 73) BCH 340 lecture 6 The Metabolic Pathway of Cellular Respiration All of the reactions involved in cellular respiration can be grouped into three main stages
Coenzyme A is a substrate or a product. (four enzymes) NADH is a substrate or a product. (five enzymes)
 BCH 4053 August 4, 2000 Mini-Exam NAME (25) 1. Following is an alphabetical list of the glycolytic and TCA cycle enzymes plus a few other enzymes we have discussed. Choose enzymes from this list that are
BCH 4053 August 4, 2000 Mini-Exam NAME (25) 1. Following is an alphabetical list of the glycolytic and TCA cycle enzymes plus a few other enzymes we have discussed. Choose enzymes from this list that are
Oxidative Phosphorylation
 Electron Transport Chain (overview) The NADH and FADH 2, formed during glycolysis, β- oxidation and the TCA cycle, give up their electrons to reduce molecular O 2 to H 2 O. Electron transfer occurs through
Electron Transport Chain (overview) The NADH and FADH 2, formed during glycolysis, β- oxidation and the TCA cycle, give up their electrons to reduce molecular O 2 to H 2 O. Electron transfer occurs through
