Automated image analysis of nuclear shape: What can we learn from a prematurely aged cell?
|
|
|
- Marvin Fitzgerald
- 6 years ago
- Views:
Transcription
1 AGING, February 2012, Vol. 4, No 2 Research Paper Automated image analysis of nuclear shape: What can we learn from a prematurely aged cell? Meghan K. Driscoll 1, Jason L. Albanese 2, Zheng Mei Xiong 2, Mitch Mailman 1, Wolfgang Losert 1, and Kan Cao 2 1 Department of Physics, University of Maryland, College Park, MD 20742, USA 2 Department of Cell Biology and Molecular Genetics, University of Maryland, College Park, MD 20742, USA Key words: progeria, aging, rapamycin, mtor, nucleus Received: 1/12/12; Accepted: 2/15/12; Published: 2/16/12 Correspondence to: Kan Cao, PhD; Wolfgang Losert, PhD; E mail: kcao@umd.edu; wlosert@umd.edu Copyright: Driscoll et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited Abstract: The premature aging disorder, Hutchinson Gilford progeria syndrome (HGPS), is caused by mutant lamin A, which affects the nuclear scaffolding. The phenotypic hallmark of HGPS is nuclear blebbing. Interestingly, similar nuclear blebbing has also been observed in aged cells from healthy individuals. Recent work has shown that treatment with rapamycin, an inhibitor of the mtor pathway, reduced nuclear blebbing in HGPS fibroblasts. However, the extent of blebbing varies considerably within each cell population, which makes manual blind counting challenging and subjective. Here, we show a novel, automated and high throughput nuclear shape analysis that quantitatively measures curvature, area, perimeter, eccentricity and additional metrics of nuclear morphology for large populations of cells. We examined HGPS fibroblast cells treated with rapamycin and RAD001 (an analog to rapamycin). Our analysis shows that treatment with RAD001 and rapamycin reduces nuclear blebbing, consistent with blind counting controls. In addition, we find that rapamycin treatment reduces the area of the nucleus, but leaves the eccentricity unchanged. Our nuclear shape analysis provides an unbiased, multidimensional fingerprint for a population of cells, which can be used to quantify treatment efficacy and analyze cellular aging. INTRODUCTION Hutchinson-Gilford progeria syndrome (HGPS) is a rare genetic disease that occurs in approximately 1 out of 4 million live births [1]. Visible symptoms of patients with HGPS include a pronounced forehead, short stature, receding mandible, conspicuous veins in the scalp, alopecia and diminished subcutaneous fat [1-4]. Internally, such patients undergo accelerated organ degeneration. The average life expectancy of HGPS patients is just 14 years, with death typically resulting from heart attacks or stroke [1-4]. The genetic mutation that leads to HGPS occurs in exon 11 of the human LMNA gene, which plays a role in nuclear scaffolding [5, 6]. This HGPS mutation is a de novo single nucleotide substitution (1824 C => T), which does not change the amino acid coding sequence [GGC (glycine) => GGT (glycine)]. However, this mutation partially activates a cryptic splice donor site, which causes a 150-nucleotide sequence to be spliced out of exon 11 and leads to the production of the mutant protein progerin, also known as LAΔ50 [7]. Because of this internal deletion, progerin does not contain the cleavage site required for the removal of the farnesyl group by protease Zempste 24, so the farnesyl group remains attached to progerin [1, 8]. The farnesyl chain is hydrophobic and has a strong affinity for the inner nuclear membrane. As a result, progerin abnormally inserts into the nuclear membrane, resulting in bulging of the nuclear envelope. This abnormal nuclear shape, commonly referred to as nuclear blebbing, has been the hallmark cellular phenotype for HGPS cells [1, 8], yet the molecular and physical mechanisms of nuclear blebbing are not well understood. In addition, the presence of progerin results in alterations in histone methylation, a thickened nuclear lamina, genome instability, clustering of nuclear pores, and loss of heterochromatin [9]. As progerin continues to build up inside prematurely aged cells, the nuclear blebbing AGING, February 2012, Vol.4 No.2
2 phenotype and other damaging effects become more severe [9]. Cellular division is also affected in HGPS cells: during mitosis, when the nuclear envelope disassembles, the progerin forms aggregates with membranes, interferes with nuclear membrane disassembly, and mislocalizes to the cytoplasm after mitosis, leading to chromosome mis-segregation and binucleation [10, 11]. Much work has also been done in an effort to develop a cure for HGPS. Children with HGPS are currently participating in the first clinical trial, testing a drug therapy that uses farnesyl transferase inhibitors (FTIs), which block the addition of the farnesyl group to progerin (Progeria Research Foundation 2011)[8, 12-14]. More recently, we showed that the macrolide antibiotic rapamycin can reverse the nuclear blebbing and other phenotypes in HGPS cells through downregulating progerin, which suggests its potential as a treatment for HGPS [15-17]. In both FTI and rapamycin studies, the percentages of nuclear blebbing, as scored by blind observers, were used as the first indication of the effectiveness of the drugs. However, it is not possible to define whether a cell is blebbed unambiguously because many cells in both healthy and diseased populations contain minor abnormalities in nuclear shape. Hence, the fraction of cells counted as blebbed can vary considerably among different observers, making blebbing quantification an inherently statistical problem. A number of studies have suggested a strong connection between HGPS and the normal aging processes. In 2006, Misteli s group reported the detection of progerin mrna and protein in cells obtained from healthy individuals, indicating that the cryptic splice site in exon 11 is also used in the presence of the normal sequence of exon 11 [18]. Similar to the results describe above, we detected low levels of progerin in normal cells, and a significant percentage of these cells had mitotic defects similar to those found in HGPS cells [10]. Our recent study further revealed a causative connection between dysfunctional telomeres and the cryptic splicing of lamin A [19]. Moreover, studies using tissues taken from normal human subjects revealed that at any age, the cryptic splicing event takes place in skin, and as we grow older, progerin-positive fibroblasts become more abundant [20, 21]. Thus, one may expect a broad distribution in the severity of blebbing in a normal cell population and an increase in blebbing with aging. Here, we report an automated, quantitative method that we used suitable to study distributions of blebbing in a large cell population. In this method, the nuclear morphology, as visualized by immunofluorescence staining of lamin A/C, is quantified using image analysis software that extracts the nuclear boundary from microscopy images and then calculates measures such as area, perimeter, and curvature for each nucleus. For each set of treated cells, the curvature of all of the nuclei can further be visualized in a single plot. Since curvature is a mathematically complete description of shape, these plots allow for the quick assessment of the severity of blebbing in a population of nuclei. We applied our method to examine HGPS fibroblasts treated with either mock, rapamycin, or RAD001, a derivative of rapamycin with better tolerance in patients. We found that treatment with RAD001 or rapamycin decreases both blebbing and nuclear area in a dosage dependent manner, but leaves nuclear eccentricity unchanged. Our study presents a novel, unbiased, quantitative method for analyzing HGPS and aging cells. This method could be useful for future drug screenings for HGPS or other age related diseases, patient diagnostics, and quantitative modeling of nuclear shape. RESULTS The nuclear shape of HGPS and normal fibroblast cells can be automatically extracted, visualized, and analyzed In order to test our automatic analysis of nuclear shape, we first cultured fibroblasts from two HGPS fibroblast cell lines (HGADFN155-p15 and HGADFN167-p12, HGPS1 and HGPS2 respectively) and from one normal control (HGFDFN168-p14, normal). The cells were fed with fresh MEM medium containing 15% FBS and grown at 37 C. To visualize the nuclei, we performed immunofluorescence staining of the nuclear membrane with a mouse monoclonal antibody raised against lamin A/C (MAB3211). This antibody has been well characterized in HGPS cells and has also been used in studies on other laminopathies. Fluorescence images of about 100 randomly selected nuclei per cell line were taken with a Zeiss fluorescence microscope at 400X magnification (examples are shown in Figure 1a). A custom-written MATLAB program (details as described in Material and Methods) was used to extract nuclear shapes and properties of the shape, such as boundary curvature. In Figures 1a and b, the nuclear boundaries, which are colored by curvature, are shown overlaid on the microscopy images. Convex curvatures were kept positive, while concave curvatures were made negative. As shown by the color bar in Figure 1c, blue represents regions of large positive curvature, and red represents regions of large negative curvature. A blebbed cell, such as the cell shown in the top of Figure 1b, will have AGING, February 2012, Vol.4 No.2
3 boundary regions that are dark red and dark blue, whereas a non-blebbed cell, such as the cell shown in the bottom of Figure 1b, will have boundary regions that are mostly blue and green with almost no red. Figure 1. The boundary curvature of HGPS and normal nuclei. (a) The curvature of nuclei is automatically extracted from fluorescence images of anti lamina/c immunostaining. Here, the curvature of HGPS and normal nuclei is shown as a colored outline, where blue represents regions of large positive curvature, and red regions of large negative curvature (scale bar: 10 μm). Blebbed nuclei have more regions of negative curvature, and so have more red signals. (b) High magnification examples of the extracted boundary curvature of a blebbed, HGPS nucleus, and a more oval, normal nucleus (scale bar: 10 μm). (c) The boundary curvature of hundreds of nuclei can be represented in a single heat map. In these heat maps, which here correspond to two HGPS cell lines (HGPS1 and HGPS2, respectively) and one control cell line (Normal), each vertical line is the stretched, colored outline of a single nucleus. Regions of large negative curvature are colored blue while regions of large negative curvature are colored red. (d) The nuclei are ordered from left to right by increasing mean negative curvature (MNC), which is shown in the line plots. (e) The MNC of populations from both HGPS cell lines is statistically different from the population from the normal control, as illustrated in this histogram AGING, February 2012, Vol.4 No.2
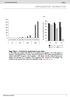
4 We next assembled plots that, for each cell line, display the boundary curvatures of all of the measured nuclei, as shown in Figure 1c. In these heat maps, each vertical line represents the boundary curvature of one nucleus. To construct such a plot, imagine cutting each colored boundary at the location farthest from the nucleus center (that location is labeled 0 on the heat map), pulling the boundary straight, and then lining it up next to the colored boundaries of the other nuclei. The heat maps of blebbed populations, such as the HGPS cell lines, have many red speckles, whereas the heat maps of unblebbed populations, such as the control cell line, have few red speckles. Within each plot, the nuclei are ordered from left to right by increasing mean negative curvature (MNC), a measure of nuclear blebbing. We defined the MNC of each nucleus by averaging all negative curvatures, excluding the positive curvatures entirely, and taking the absolute value. As shown in Figures 1d and 1e, the HGPS1 and HGPS2 cell lines have larger MNCs, and hence are more blebbed, than the control cell line. HGPS1 also has a larger MNC than HGPS2, perhaps because HGPS1 is at a later cellular passage, and thus more senesced. We found that both HGPS MNC distributions are statistically different from the MNC distribution of the control (see Figure 1e for p-values). To validate the automated nuclear shape analysis, we also assessed nuclear morphology using the standard technique: manual blind counting. Nuclei with protrusions or invaginations were counted as blebed, while other nuclei were counted as normal. We found that 73% of HGPS1 nuclei, 63% of HGPS2 nuclei, and 24% of normal nuclei were abnormal. These counts are in quantitative agreement with the MNC distributions of the respective cell lines (Figure 1e). In order to better evaluate how the results of manual counting correlate with quantitative shape metrics, we had experienced human counters label individual nuclei as either normal or blebbed, and analyzed the MNC of the two populations. The analysis shows a positive correlation between MNC and the percentage of nuclei that were labeled as blebbed (Figure S1). Since the automated analysis extracts the boundary of each nucleus, we can assess nuclear morphology using various shape metrics besides boundary curvature. For each nucleus, we also calculated area, perimeter, number of invaginations, eccentricity and other metrics. In analogy to how microarray data is analyzed to find relationships between genes, we used correlation as a measure of interrelationship between the 15 different measures of nuclear shape determined in this study. We hierarchically clustered the 15 measures of nuclear shape and lamina/c fluorescence intensity (Figure S2). We found several families of nuclear measures that roughly correspond to size, extent of blebbing, eccentricity, lamina/c fluorescence intensity, and the standard deviation of fluorescence intensity. The clustering reassures us that the intensity, which is affected by immunostaining and imaging details, does not notably affect the measured MNC. The clustering also indicates that the standard deviation in MNC and the tortuosity are measures related to MNC. Also related to mean MNC is the solidity, which is the ratio of the measured area and the area of 'convex hull,' or the minimal convex shape that bounds the measured shape of the nucleus. As a control experiment, we tested whether the cell density would influence the MNC. We seeded cells from the same HGPS cell line at densities of 3000, 9000, and cells per well in 4-well chamber slides (Figure S3). The three densities did not appear to have different MNC distributions, nor were the measured MNC distributions statistically distinct. RAD001 and rapamycin similarly reduce blebbing in HGPS fibroblast cells Recent work has revealed that rapamycin, an mtor (mammalian target of rapamycin) inhibitor, significantly reduces the phenotypic hallmarks of progeria in HGPS cells by down-regulating progerin [15]. Everolimus (RAD001), which is the 40-O-(2- hydroxyethyl) derivative of rapamycin, works similarly to sirolimus as an mtor inhibitor but is better tolerated by patients. In order to compare the efficacy of RAD001 to rapamycin, we treated HGPS fibroblasts with rapamycin, RAD001, or mock, and then analyzed the nuclear morphology of each treatment group. Cultured fibroblasts from an HGPS patient (HGADFN155-p15) and a normal individual (HGFDFN168-p15) were used in this experiment. The cells were fed every other day with fresh MEM medium containing 0.68 µm rapamycin, 0.1 µm RAD001, 0.5 µm RAD001, or the same volume of vehicle (DMSO, 0.025% v/v) for a duration of seven weeks. To examine the effects on nuclear morphology, we labeled cells with an antibody for lamin A/C and an antibody specific for progerin (Figure 2a). To judge the impact of rapamycin and RAD001, we first scored the percentage of nuclei with abnormal morphology in the usual way by manual blind counting. At least 200 randomly selected cells were scored by fluorescence microscopy for each cell line under each condition. In comparison with the passage-matched, mock-treated HGPS cells, the rapamycin or RAD001 treated HGPS cells exhibited AGING, February 2012, Vol.4 No.2
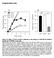
5 a clear reduction in nuclear blebbing (Figure 2b). Since increased genome instability was reported in HGPS cells [15, 22, 23], we also examined whether RAD001 treatment can improve this phenotype. Using immunofluorescence staining, we observed a reduction in 53BP1 foci in rapamycin or RAD001 treated cells, indicating that inhibition of mtor prevents DNA damage induced in prematurely senescent cells by progerin (Figure S4). Quantification of progerin protein by western blotting analysis also revealed an over 50% reduction in progerin levels in rapamycin and RAD001 treated HGPS cells (Figure 2c). We also detected a weaker progerin-staining signal in almost all of the rapamycin or RAD001 treated HGPS cells, and their nuclear morphology appeared substantially improved compared to untreated cells (Figure 2a). As reported in Cao et al. [15], we again noticed a reduction in overall cell proliferation at the beginning of the treatments (less than two weeks) with either rapamycin or RAD001 compared to the mock treated samples. We used a tetrazolium salt-based cell proliferation assay (WST1 assay) to analyze this apparent cell growth inhibition for the concentrations of RAD001 used in the experiment: 0, 20, 60, 100, and 500 nm. Figure S5 shows that all treatments for both control and HGPS cell lines had a similar reduction in cell proliferation compared to the mock treatments (less than 40%), suggesting that any effective dose of RAD001 may have similar anti-hypertrophic effects [24]. Figure 2. The nuclear morphology and progerin levels of rapamycin and RAD001 treated HGPS cells. (a) The phenotype of nuclear blebbing was improved in RAD001 and rapamycin treated HGPS fibroblast cells. Cells were stained with DAPI (blue), lamina/c antibody (red), and progerin antibody (green) to show nuclear location and morphology. The treatment duration is for seven weeks. Mock: vehicle (DMSO, 0.025% v/v); Rap: 0.68 µm rapamycin, Rad: 0.1 µm RAD001. (Scale bar: 10 µm) (b) Quantification of the percentage of blebbing in all treatments. At least 200 nuclei were counted blindly. M: vehicle (DMSO, 0.025% v/v); Rap: 0.68 µm rapamycin; RAD100: 0.1 µm RAD001 treatment; RAD500: 0.5 µm RAD001 treatment. (c) Progerin was decreased in rapamycin or RAD001 treated HGPS fibroblasts. The relative amount of progerin was quantified using quantitative western blotting analysis and compared to the mock treated HGPS cells AGING, February 2012, Vol.4 No.2
6 Figure 3. Imaging analysis of RAD001 and rapamycin treated HGPS cells. (a) Heat maps that represent the boundary curvature of the HGPS cells treated with the vehicle (mock), 100 nm and 500 nm RAD001 (Rad), and rapamycin (Rap). (b) MNC of populations from the mock treated HGPS cell lines is statistically different from the populations of RAD001 or rapamycin treated cells (p < 0.001) (c e) Compared to the mock treated nuclei, the RAD001 and rapamycin treated nuclei have smaller area (c) and fewer invaginations (d), but similar eccentricity (e). In parallel to the blind counting, we took immunofluorescence images of about 100 randomly selected nuclei per treatment group and automatically analyzed their nuclear morphology. Heat maps, which display the boundary curvature of the treated HGPS cells, are shown in Figure 3a. From the heat maps we see that the mock treated cells are much more blebbed than the rapamycin or RAD001 treated cells, which is consistent with our blinded counting. Indeed, we found that the MNC distributions of the rapamycin and RAD001 treated cells were statistically different from that of the control group (Figure 3b). Similarly, our analysis showed a reduction in the number of invaginations in treated HGPS cells (Figure 3d). Interestingly, we also found that the RAD001 and rapamycin treated nuclei had a smaller area than the mock treated nuclei (Figure 3c). Moreover, we noticed that the eccentricity, which is a measure of how elongated the nuclei are, did not change as a result of the rapamycin or RAD001 treatments (Figure 3e). Our analysis suggested that rapamycin or RAD001 treatments appear to locally improve abnormal morphology, without affecting the overall shape of the nuclei, though still altering nuclear size (see discussion). In summary, our data suggest that, similar to rapamycin, RAD001 can reverse the nuclear phenotypes in HGPS cells through promoting progerin clearance AGING, February 2012, Vol.4 No.2
7 RAD001 induces a gradual change in nuclear morphology in a dosage dependant manner Based on the above analysis, we suggested RAD001 could be used at 100 nm concentration to achieve similar beneficial effects in HGPS cell cultures as rapamycin at 0.68 µm as described in Cao et al. [15]. Next, we explored the sensitivity of the curvature analysis program, since quantitative image analysis is most useful if it can reveal small changes that are difficult to observe. Thus, we lowered the dosage of RAD001 to 20 or 60 nm, and shortened the duration of treatment to two weeks. An HGPS fibroblast cell line (HGADFN167-p15) and a control fibroblast cell line (HGFDFN168-p15) were fed with fresh MEM medium containing 20 nm RAD001, 60nM RAD001 or the same volume of vehicle (0.3% DMSO) every other day. Nuclear curvature outline and heat map analyses of MNC were carried out at the end of the two-week treatment (Figure 4a). Box plot analysis indicated a significant reduction of MNC in the HGPS cell line, even in the cells receiving 20 nm RAD001 (Figure 4b), while those minor morphological improvements were not visible with the traditional blinded counting method, suggesting that the automated analysis is more sensitive. Importantly, we noticed a dosage dependent reduction in area of the nuclei of both treated HGPS and treated control cells (Figure 4c): the area of mock treated nuclei was greater than both doses of RAD001 treated nuclei, but the nuclei that received the smaller dose of RAD001 (20 nm) had greater area than the nuclei that received the larger dose (60 nm). This result suggests the improvement in nuclear shape is a gradual process, the area reduction is primarily due to non-specific effects of the drug treatment, and incrementtal improvement during treatment can be captured and quantified by this curvature outline imaging analysis. Figure 4. RAD001 induces a gradual change in nuclear morphology in a dosage dependant manner. (a) Heat maps that represent the boundary curvature of HGPS cells treated with the vehicle (mock) or 20 or 60 nm RAD001. (b) Both doses significantly reduce the mean negative curvature as shown by the histogram. (c) The area of mock treated nuclei is greater than both doses of RAD001 treated nuclei, but the nuclei that received the smaller dose of RAD001 have greater area than the nuclei that received the larger dose. The same trend in area change is apparent to the same extent in the treated control normal fibroblasts AGING, February 2012, Vol.4 No.2
8 DISCUSSION One of the hallmarks of HGPS is the abnormal nuclear shape known as blebbing. This has been the main morphological feature identifying an HGPS cell line and has been used to determine the effectiveness of treatments for HGPS. The traditional method of measuring blebbing is a manual, blind count of the percentage of blebbed nuclei. However, this method has no standard criteria and is extremely time consuming. Sorting the nuclei into two categories, normal and blebbed, also obscures the fact that blebbing is not an either/or phenomenon, but varies continuously. The subjectivity and variability of the threshold for blebbed nuclei makes it impossible to compare values obtained by different counters. The need for an unbiased, quantitative method of measuring the degree of blebbing in a cell sample is clear. In an effort towards solving this issue, we present an automated image analysis method using curvature as the primary measure of blebbing. We used a custom-written program to extract the boundaries of immuno-stained nuclei and calculate a curvature contour for each nucleus among other measures of shape. We found that several measures of the shape differentiate between HGPS and normal control cell lines. We focused on the most intuitive measure, the mean negative curvature (MNC), which is the average of all the concave curvatures on the boundary of a nucleus. MNC provides a continuous measure of blebbing that can be used in quantitative and statistical methods. We analyzed different seeding densities (Figure S3) and exposure times (data not shown) to demonstrate that MNC is also a consistent measure that does not vary significantly between experiments. The cluster analysis also shows that intensity does not affect the measured MNC (Figure S2). Thus MNC values can be compared between samples and experiments, unlike values obtained from the traditional blebbing count method. One caveat is that MNC is affected by pixel size and smoothing, thus care should be taken when comparing results from different laboratories. Of the other measures that strongly correlate with MNC, according to our clustering analysis (Figure S2), solidity should not be significantly affected by pixel size or smoothing and thus may be a viable alternative. To better demonstrate the usefulness of this novel analysis, we treated HGPS and control cell lines with rapamycin, an mtor pathway inhibitor that has been shown to improve nuclear morphology of HGPS cells, and with one of its analogues, RAD001, which is better tolerated by treated patients. The cells were treated for 7 weeks, stained with an anti-lamin A/C antibody, and analyzed using the program. Results of the treatment are presented through heat maps and box plots of MNC in Figure 3. Blinded blebbing counts were also performed (Figure 2), demonstrating that MNC agrees with the established method: RAD001 and rapamycin treated HGPS cells had significantly improved nuclear morphology to the same extent. Consistent with Cao et al. [15], we found that RAD001 promoted progerin degradation (Figure 2). In addition, we reported that RAD001 and rapamycin treatments decreased the DNAdamage induced 53BP1 foci formation in HGPS cells (Figure S4), likely through down-regulation of progerin. Consistent with our observation, previous studies have shown that rapamycin can inhibit the DNA-damage independent pseudo DNA-damage response, which might be caused by general over-activation in senescent cells [25, 26]. To demonstrate the sensitivity of this method, we used this program to distinguish between treatment doses that cannot be differentiated by the traditional bleb counting method (Figure 4). We treated HGPS and control cell lines with lower doses of RAD001 and used Student s t- test to show a statistically significant increase in MNC with lower doses. A blinded bleb count was unable to demonstrate any difference between the treatments (data not shown). In this treatment, we again noticed a dosedependent change in nuclear area (Figures 3c and 4c). However, the same area change was observed in the treated normal control cell line, suggesting that this area change is primarily due to the action of mtor inhibition and not an improvement of nuclear morphology in HGPS cells. We also showed the anti-hypertrophic effects of RAD001 in the early stages of treatment within the first week at the indicated concentrations (Figure S5). This reduced cellular growth in the initial period of treatment and the area decrease of nuclei may be explained by the inhibition of the mtor pathway. After the initial slowdown in growth during the first two weeks of treatment, rapamycin and RAD001 treated cells showed a greatly improved proliferation rate, better than their mock treated counterparts [15], which is consistent with the previously established role of rapamycin in preventing the loss of proliferative potential in cultured cells [27]. Notably, our multidimensional analysis of cell shapes provides unexpected hints into the mechanical aspects of mtor inhibition: while RAD001 or rapamycin treatment decreases blebbing and nuclear size, they do not alter the eccentricity of the nuclear shape (Figure 3e). We expect that our high throughput, multidimensional measures will provide a solid foundation for developing mechanical models of the nucleus. HGPS is a devastating and well-studied premature aging disease that currently has no effective treatment AGING, February 2012, Vol.4 No.2
9 HGPS also has strong connections with the general aging process. In both scenarios, the broad distribution of nuclear shape abnormality in a single population of cells hampers manual analysis. Our automated nuclear shape analysis software provides a high-throughput and easy-to-use method of quantifying nuclear morphology. Heat maps of curvature (Figures 1, 3, 4) allow us to directly visualize the broad distribution of nuclear blebbing in a large cell population. Comparing measures between samples allows us to assess treatment efficacy for HGPS and other age-related diseases. We use this method to demonstrate the potential of RAD001 as a treatment alternative for HGPS, being similarly effective to rapamycin. Our nuclear shape analysis provides an unbiased multidimensional fingerprint for a population of cells, which can be used to quantify treatment efficacy and analyze cellular aging. MATERIALS AND METHODS Cells, Cell Culture, and Treatments. Primary human dermal fibroblasts used in this study were obtained from the Progeria Research Foundation (PRF): two HGPS fibroblasts, HGADFN155 and HGADFN167, and a control normal fibroblast line, HGFDFN168. Fibroblasts were cultured in MEM medium (Invitrogen) supplemented with 15% FBS and 2 mm L-glutamine under 5% CO2 at 37 C. Normal and HGPS fibroblasts were replenished with fresh MEM medium containing 0.68 µm rapamycin/dmso, or indicated concentration of RAD001/DMSO, every other day for up to 50 days. Control cells were treated with vehicle (DMSO) in MEM medium. Rapamycin (Sirolimus) was purchased from Sigma, and RAD001 (Everolimus) was obtained from Selleck. Immunofluorescence Staining. For immunofluorescence, cells were seeded in 4-well chamber slides. After fixation in 4% paraformaldehyde/pbs at room temperature for 15 min, cells were permeabilized with 0.5% Triton X-100/PBS at room temperature for 5 min, followed by an overnight incubation in the blocking solution (4% BSA/TBS) at 4 C. Cells were then stained with primary antibodies for 3 hours at room temperature on the following day. The primary antibodies used in this study were: a rabbit polyclonal antibody against progerin (custom peptide antibody, Yenzym); a goat anti-lamin A/C antibody (N-18, Santa Cruz); and a mouse anti-lamin A/C antibody (MAB3211, Chemicon). After primary antibody incubation, primary antibodies were detected with Alexa Fluor-labeled secondary antibodies (Invitrogen). Slides were mounted with Vectashield mounting medium containing DAPI and observed with a Zeiss fluorescence microscope. Images were taken using a 40x objective (about 100 nuclei per condition). Exposure times and acquisition settings were established at the beginning of each set of experiments and kept constant for all treatments. Extraction of Nuclei Boundaries. A custom-written MATLAB program was used to extract nuclei boundaries. In order to reduce image histogram variability both between and within images, we first used contrast-limited adaptive histogram equalization. An 8 x 8 grid of tiles, a clip limit of 0.02, and a Rayleigh distribution were employed. Next, we binarized the images using MATLAB s built-in thresholding function, which uses Otsu s method. Holes within bright regions were then filled, and regions that either overlapped the image boundary or were smaller than 800 square pixels were removed. In order to smooth and enlarge the regions, which correspond to nuclei, the images were morphologically eroded with a disk of radius 3 pixels and then morphologically dilated with a disk of radius 6 pixels. The convex hulls of these smoothed and enlarged regions were next calculated. These outer convex hulls were later used as an outer limit on the nuclei boundary positions. Next, the images were morphologically eroded with a disk of radius 2 pixels. The convex hulls of these smoothed and slightly enlarged regions were used to initialize an active contour based boundary extraction algorithm. We next processed the original images for use by the active contour based boundary extraction algorithm. First, the brightness and contrast of the image was adjusted so that 1% of the pixels was saturated at the lowest intensity and 1% was saturated at the highest intensity. For each nucleus, any pixel outside of its outer convex hull, which was found as described above, was set to zero, and the brightness and contrast were again adjusted as before. We next calculated the binarization threshold using MATLAB s built-in thresholding function, but did not binarize the image. Instead, we nearly binarized the image by setting any pixel whose value was less than 70% of the threshold value to the lowest intensity and any pixel whose value was greater than 130% of the threshold value to the highest intensity. The remaining, non-saturated pixel intensities were then stretched to fill the entire intensity range. The holes in this gray-scale, nucleus image were next filled. An active contour, or snake algorithm, was used to extract nuclei boundaries with sub-pixel resolution, as described in Xu and Prince [28]. The gradient vector flow (GVF) field of the processed, nucleus image was calculated. The inner convex hull of the nucleus, found as described above, was next interpolated and used as the initial position of the active contour. The contour, AGING, February 2012, Vol.4 No.2
10 which is a polygon, was then repeatedly deformed 75 times and interpolated until the change in area from one set of deformations to the next was no more than 10 square pixels. Contours were not allowed to deform more than 50,025 times. The GVF, snake interpolation and snake deformation functions are from Xu and Prince [28]. (The following parameters were supplied to these three functions: number of GVF iterations, 80; d min, 0.5; d max, 2; tension or alpha, 0.02; rigidity or beta, 0.05; step size or gamma, 1; external force or kappa, 0.6.) The contour was interpolated a final time, resulting in an outputted polygon with sides of constant length (d min ). Some contours do not correspond to single nuclei, but rather are multiple, overlapped nuclei or are autofluorescent regions of cells. The user is next given an opportunity to remove such unwanted contours. Each analyzed image is sequentially displayed and polygons clicked on by the user are removed from further analysis. Analysis of Boundary Shape. We calculated the boundary curvature at each boundary point by fitting a circle to that boundary point and the two points 25 boundary points away from it. The curvature was then calculated as the reciprocal of the radius of that circle. Convex curvatures were kept positive, while concave curvatures were made negative. For each nucleus, the boundary point farthest from the centroid was labeled boundary point 0. When visualized with color, curvature values were cut-off such that magnitudes above a cut-off value were set to that cut-off value. For each nucleus, the number of invaginations was calculated by simply counting the number of boundary regions, of any length, where negative curvature was uninterrupted by positive curvature, and eccentricity was defined as the eccentricity of an ellipse with the same second-moments as the nuclear shape. The eccentricity of an ellipse describes how elongated the ellipse is; a circle would have an eccentricity of 0, and a line segment has an eccentricity of 1. We have previously similarly analyzed the shape of migrating amoebae [29] WST-1 Cell Proliferation Assay. A WST-1 cell proliferation assay (Roche) was used to analyze the effects of RAD001 on cellular growth. HGPS cell line HGADFN167 p12 and control cell line HGADFN168 p14 were seeded in separate standard 24-well plates at 10,000 cells in 500 µl fibroblast medium per well. Wells were treated with 0, 20, 60, 100 and 500 nm RAD001/DMSO in triplicate and the solvent controlled at 0.1% for all wells. The cells were then incubated with treatment for 72 hours. The medium was removed from each well and 500 µl of 10% WST-1 reagent in fibroblast medium was applied to each well following the incubation. Three blanks, consisting of 500 µl of 10% WST-1 reagent in fibroblast medium, were also created. The absorbance (450nm) of each well was read after 3 more hours of incubation using a SpectraMax M5 e plate reader (Molecular Devices), and the average absorbance of the blanks was subtracted from each measurement. Cell numbers were calculated from the absorbance values using a standard curve established by repeating the experiment without treatment and seeding at 1000, 2000, 4000, 8000 and cells per well in duplicate. The percent survival was calculated for each sample by the equation: percent survival = 100 * (cell #)/(average cell # of 0 nm treatment), then averaged by treatment. The error was calculated using the standard deviation of the percent survivals of the 3 samples for each treatment. Statistics. Statistical analysis of MNC and other values was performed using the ttest2 function in MATLAB v that performs a Student s t-test. The parameters were set to a 1-tailed t-test, assuming unequal variances, and a p-value of 0.05 or less was considered significant. ACKNOWLEDGMENTS We thank members in Cao s lab and Dr. Haibei Luo for technical assistance, and Dr. Norma Andrews for sharing her equipment and helpful discussion. We also thank the HGPS group in Dr. Francis Collins laboratory for suggestions. This work was supported by a grant to the University of Maryland from the Howard Hughes Medical Institute Undergraduate Science Education Program (JA), a grant from the National Institute on Aging R00AG (KC), a grant from the Progeria Research Foundation (KC), and by a NIGMS grant R01GM (WL). CONFLICT OF INTERESTS STATEMENT The authors of this manuscript have no conflict of interest to declare. REFERENCES 1. Capell BC and Collins FS. Human laminopathies: nuclei gone genetically awry. Nat Rev Genet. 2006; 7: Gordon LB, Harling Berg CJ and Rothman FG. Highlights of the 2007 Progeria Research Foundation scientific workshop: progress in translational science. J Gerontol A Biol Sci Med Sci. 2008; 63: AGING, February 2012, Vol.4 No.2
11 3. Gordon LB, McCarten KM, Giobbie Hurder A, Machan JT, Campbell SE, Berns SD and Kieran MW. Disease progression in Hutchinson Gilford progeria syndrome: impact on growth and development. Pediatrics. 2007; 120: Merideth MA, Gordon LB, Clauss S, Sachdev V, Smith AC, Perry MB, Brewer CC, Zalewski C, Kim HJ, Solomon B, Brooks BP, Gerber LH, Turner ML, et al. Phenotype and course of Hutchinson Gilford progeria syndrome. N Engl J Med. 2008; 358: Dahl KN, Scaffidi P, Islam MF, Yodh AG, Wilson KL and Misteli T. Distinct structural and mechanical properties of the nuclear lamina in Hutchinson Gilford progeria syndrome. Proc Natl Acad Sci U S A. 2006; 103: Dechat T, Adam SA, Taimen P, Shimi T and Goldman RD. Nuclear lamins. Cold Spring Harb Perspect Biol. 2010; 2:a Eriksson M, Brown WT, Gordon LB, Glynn MW, Singer J, Scott L, Erdos MR, Robbins CM, Moses TY, Berglund P, Dutra A, Pak E, Durkin S, et al. Recurrent de novo point mutations in lamin A cause Hutchinson Gilford progeria syndrome. Nature. 2003; 423: Capell BC, Erdos MR, Madigan JP, Fiordalisi JJ, Varga R, Conneely KN, Gordon LB, Der CJ, Cox AD and Collins FS. Inhibiting farnesylation of progerin prevents the characteristic nuclear blebbing of Hutchinson Gilford progeria syndrome. Proc Natl Acad Sci U S A. 2005; 102: Goldman RD, Shumaker DK, Erdos MR, Eriksson M, Goldman AE, Gordon LB, Gruenbaum Y, Khuon S, Mendez M, Varga R and Collins FS. Accumulation of mutant lamin A causes progressive changes in nuclear architecture in Hutchinson Gilford progeria syndrome. Proc Natl Acad Sci U S A. 2004; 101: Cao K, Capell BC, Erdos MR, Djabali K and Collins FS. A lamin A protein isoform overexpressed in Hutchinson Gilford progeria syndrome interferes with mitosis in progeria and normal cells. Proc Natl Acad Sci U S A. 2007; 104: Dechat T, Shimi T, Adam SA, Rusinol AE, Andres DA, Spielmann HP, Sinensky MS and Goldman RD. Alterations in mitosis and cell cycle progression caused by a mutant lamin A known to accelerate human aging. Proc Natl Acad Sci U S A. 2007; 104: Mallampalli MP, Huyer G, Bendale P, Gelb MH and Michaelis S. Inhibiting farnesylation reverses the nuclear morphology defect in a HeLa cell model for Hutchinson Gilford progeria syndrome. Proc Natl Acad Sci U S A. 2005; 102: Young SG, Fong LG and Michaelis S. Prelamin A, Zmpste24, misshapen cell nuclei, and progeria new evidence suggesting that protein farnesylation could be important for disease pathogenesis. J Lipid Res. 2005; 46: Capell BC, Olive M, Erdos MR, Cao K, Faddah DA, Tavarez UL, Conneely KN, Qu X, San H, Ganesh SK, Chen X, Avallone H, Kolodgie FD, et al. A farnesyltransferase inhibitor prevents both the onset and late progression of cardiovascular disease in a progeria mouse model. Proc Natl Acad Sci U S A. 2008; 105: Cao K, Graziotto JJ, Blair CD, Mazzulli JR, Erdos MR, Krainc D and Collins FS. Rapamycin reverses cellular phenotypes and enhances mutant protein clearance in Hutchinson Gilford progeria syndrome cells. Sci Transl Med. 2011; 3:89ra Graziotto JJ, Cao K, Collins FS and Krainc D. Rapamycin activates autophagy in Hutchinson Gilford progeria syndrome: Implications for normal aging and age dependent neurodegenerative disorders. Autophagy. 2012; Blagosklonny MV. Progeria, rapamycin and normal aging: recent breakthrough. Aging (Albany NY). 2011; 3: Scaffidi P and Misteli T. Lamin A dependent nuclear defects in human aging. Science. 2006; 312: Cao K, Blair CD, Faddah DA, Kieckhaefer JE, Olive M, Erdos MR, Nabel EG and Collins FS. Progerin and telomere dysfunction collaborate to trigger cellular senescence in normal human fibroblasts. J Clin Invest. 2011; 121: McClintock D, Ratner D, Lokuge M, Owens DM, Gordon LB, Collins FS and Djabali K. The mutant form of lamin A that causes Hutchinson Gilford progeria is a biomarker of cellular aging in human skin. PLoS One. 2007; 2:e Olive M, Harten I, Mitchell R, Beers JK, Djabali K, Cao K, Erdos MR, Blair C, Funke B, Smoot L, Gerhard Herman M, Machan JT, Kutys R, et al. Cardiovascular pathology in Hutchinson Gilford progeria: correlation with the vascular pathology of aging. Arterioscler Thromb Vasc Biol. 2010; 30: Richards SA, Muter J, Ritchie P, Lattanzi G and Hutchison CJ. The accumulation of un repairable DNA damage in laminopathy progeria fibroblasts is caused by ROS generation and is prevented by treatment with N acetyl cysteine. Hum Mol Genet. 2011; 20: Hutchison CJ. The role of DNA damage in laminopathy progeroid syndromes. Biochem Soc Trans. 2011; 39: Demidenko ZN and Blagosklonny MV. Quantifying pharmacologic suppression of cellular senescence: prevention of cellular hypertrophy versus preservation of proliferative potential. Aging Us. 2009; 1: Pospelova TV, Demidenko ZN, Bukreeva EI, Pospelov VA, Gudkov AV and Blagosklonny MV. Pseudo DNA damage response in senescent cells. Cell Cycle. 2009; 8: Pankotai T, Hoffbeck AS, Boumendil C and Soutoglou E. DNA damage response in the absence of DNA lesions continued. Cell Cycle. 2009; 8: Demidenko ZN, Zubova SG, Bukreeva EI, Pospelov VA, Pospelova TV and Blagosklonny MV. Rapamycin decelerates cellular senescence. Cell Cycle. 2009; 8: Xu C and Prince JL. Snakes, shapes, and gradient vector flow. IEEE Trans Image Process. 1998; 7: Driscoll MK, Fourkas JT and Losert W. Local and global measures of shape dynamics. Physical Biology. 2011; 8: AGING, February 2012, Vol.4 No.2
12 SUPPLEMENTARY FIGURES Figure S1. Mean negative curvature (MNC) correlates with bleb counting, Images of 847 nuclei from HGPS and normal control cell lines were separately displayed to 3 members of the Cao lab. Each member selected the nuclei he or she considered to be blebbed using the standard criteria established in the Materials and Methods. The selections were saved and analyzed through a custom written MATLAB program. The percentage of all nuclei with an MNC above a certain threshold that were labeled blebbed by the majority of counters (2 or more) is displayed for various MNC thresholds. xxx Figure S2. Covariance matrix (right) and hierarchical clustering plot (left) of 15 measures of nuclear shape and lamin A/C fluorescence intensity. Each box in the covariance matrix indicates the amount of correlation between two measures. High covariance is indicated in red (see color bar) AGING, February 2012, Vol.4 No.2
13 Figure S3. The plating density of the cells does not affect the curvature analysis. (a) HGPS fibroblast cells (HGADFN155 p15) were seeded at various cell densities (3000, 9000, and per well) in 4 well chamber slides and immuno stained with anti lamin A/C (N18, Santa Cruz) and DAPI. Images of the nuclear staining with DAPI and brightfield images are presented to help visualize the various cell densities. (b) Heat maps of curvature contours of 100 randomly selected nuclei for each density and sorted by MNC show a similar degree of blebbing at all cell densities. (c) Box plots of MNC created using MATLAB s boxplot function of nuclei imaged in the heat maps AGING, February 2012, Vol.4 No.2
14 Figure S4. Genome stability is improved in rapamycin or RAD001 treated cells. (a) Fibroblast cells were immuno stained with DAPI (blue) and TP53BP1 antibody (green); scale bar: 20 µm. (b) Percentage of TP53BP1 foci positive cells in Rad100, Rad500, rapamycin and mock treated HGPS cells. Mock: mock treatment; Rap: 0.68 µm rapamycin treatment; Rad100: 0.1 µm RAD001 treatment; Rad500: 0.5 µm RAD001 treatment. The percentage of cells with no TP53BP1 staining is shown in blue, the percentage of cells with one TP53BP1 is in red, and percentage of cells with more than one TP53BP1 foci is shown in yellow. Figure S5. A cell proliferation assay shows that all treatments of RAD001/DMSO at indicated dosages for both normal (blue) and HGPS (red) fibroblast cell lines had similarly reduced growth compared to the mock treatments. All treatments were controlled for DMSO at 0.1% and percent survival calculated relative to the cell numbers of the mock treatments AGING, February 2012, Vol.4 No.2
Supplementary Figure S1: Defective heterochromatin repair in HGPS progeroid cells
 Supplementary Figure S1: Defective heterochromatin repair in HGPS progeroid cells Immunofluorescence staining of H3K9me3 and 53BP1 in PH and HGADFN003 (HG003) cells at 24 h after γ-irradiation. Scale bar,
Supplementary Figure S1: Defective heterochromatin repair in HGPS progeroid cells Immunofluorescence staining of H3K9me3 and 53BP1 in PH and HGADFN003 (HG003) cells at 24 h after γ-irradiation. Scale bar,
Importance of molecular cell biology investigations in human medicine in the story of the Hutchinson-Gilford progeria syndrome
 Interdisc Toxicol. 2010; Vol. 3(3): 89 93. doi: 10.2478/v10102-010-0018-y Published online in: www.intertox.sav.sk & www.versita.com/science/medicine/it/ This is an Open Access article distributed under
Interdisc Toxicol. 2010; Vol. 3(3): 89 93. doi: 10.2478/v10102-010-0018-y Published online in: www.intertox.sav.sk & www.versita.com/science/medicine/it/ This is an Open Access article distributed under
CORRECTIONS. PNAS August 4, 2009 vol. 106 no
 MEDICAL SCIENCES Correction for A farnesyltransferase inhibitor prevents both the onset and late progression of cardiovascular disease in a progeria mouse model, by Brian C. Capell, Michelle Olive, Michael
MEDICAL SCIENCES Correction for A farnesyltransferase inhibitor prevents both the onset and late progression of cardiovascular disease in a progeria mouse model, by Brian C. Capell, Michelle Olive, Michael
Intermittent treatment with farnesyltransferase inhibitor and sulforaphane improves cellular homeostasis in Hutchinson- Gilford progeria fibroblasts
 /, 2017, Vol. 8, (No. 39), pp: 64809-64826 Intermittent treatment with farnesyltransferase inhibitor and sulforaphane improves cellular homeostasis in Hutchinson- Gilford progeria fibroblasts Diana Gabriel
/, 2017, Vol. 8, (No. 39), pp: 64809-64826 Intermittent treatment with farnesyltransferase inhibitor and sulforaphane improves cellular homeostasis in Hutchinson- Gilford progeria fibroblasts Diana Gabriel
The LMNA gene encodes the A-type lamins, including lamin A
 A lamin A protein isoform overexpressed in Hutchinson Gilford progeria syndrome interferes with mitosis in progeria and normal cells Kan Cao*, Brian C. Capell*, Michael R. Erdos*, Karima Djabali, and Francis
A lamin A protein isoform overexpressed in Hutchinson Gilford progeria syndrome interferes with mitosis in progeria and normal cells Kan Cao*, Brian C. Capell*, Michael R. Erdos*, Karima Djabali, and Francis
Hutchinson-Gilford Progeria Syndrome and its Relevance to Cardiovascular Diseases and Normal Aging
 382 Biomed Environ Sci, 2013; 26(5): 382-389 Review Hutchinson-Gilford Progeria Syndrome and its Relevance to Cardiovascular Diseases and Normal Aging QI Ying Chun and XIE Xiao Hua # First Department of
382 Biomed Environ Sci, 2013; 26(5): 382-389 Review Hutchinson-Gilford Progeria Syndrome and its Relevance to Cardiovascular Diseases and Normal Aging QI Ying Chun and XIE Xiao Hua # First Department of
(a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable
 Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
RefCell: multi-dimensional analysis of image-based high-throughput screens based on typical cells
 Shen et al. BMC Bioinformatics (2018) 19:427 https://doi.org/10.1186/s12859-018-2454-1 METHODOLOGY ARTICLE Open Access RefCell: multi-dimensional analysis of image-based high-throughput screens based on
Shen et al. BMC Bioinformatics (2018) 19:427 https://doi.org/10.1186/s12859-018-2454-1 METHODOLOGY ARTICLE Open Access RefCell: multi-dimensional analysis of image-based high-throughput screens based on
 amycin Reverses Cellular Phenotypes and Enhances Mutant Protein Clearance in Hutchinson-Gilford Progeria Syndrome Cells Kan Cao, et al. Sci Transl Med 3, 89ra58 (211); DOI: 1.1126/scitranslmed.32346 Editor's
amycin Reverses Cellular Phenotypes and Enhances Mutant Protein Clearance in Hutchinson-Gilford Progeria Syndrome Cells Kan Cao, et al. Sci Transl Med 3, 89ra58 (211); DOI: 1.1126/scitranslmed.32346 Editor's
LDL Uptake Cell-Based Assay Kit
 LDL Uptake Cell-Based Assay Kit Item No. 10011125 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
LDL Uptake Cell-Based Assay Kit Item No. 10011125 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
Cell Migration and Invasion Assays INCUCYTE LIVE-CELL ANALYSIS SYSTEM. Real-time automated measurements of cell motility inside your incubator
 Cell Migration and Invasion Assays INCUCYTE LIVE-CELL ANALYSIS SYSTEM Real-time automated measurements of cell motility inside your incubator See the whole story Real-time cell motility visualization and
Cell Migration and Invasion Assays INCUCYTE LIVE-CELL ANALYSIS SYSTEM Real-time automated measurements of cell motility inside your incubator See the whole story Real-time cell motility visualization and
HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation
 SUPPLEMENTARY INFORMATION Materials and Methods Human cell lines and culture conditions HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation in exon 20 of BRCA1
SUPPLEMENTARY INFORMATION Materials and Methods Human cell lines and culture conditions HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation in exon 20 of BRCA1
LDL Uptake Cell-Based Assay Kit
 LDL Uptake Cell-Based Assay Kit Catalog Number KA1327 100 assays Version: 07 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
LDL Uptake Cell-Based Assay Kit Catalog Number KA1327 100 assays Version: 07 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
Cesarini et al., http ://www.jcb.org /cgi /content /full /jcb /DC1
 Supplemental material JCB Cesarini et al., http ://www.jcb.org /cgi /content /full /jcb.201504035 /DC1 THE JOU RNAL OF CELL BIO LOGY Figure S1. Lamin A/C depletion generates two distinct phenotypes in
Supplemental material JCB Cesarini et al., http ://www.jcb.org /cgi /content /full /jcb.201504035 /DC1 THE JOU RNAL OF CELL BIO LOGY Figure S1. Lamin A/C depletion generates two distinct phenotypes in
(a) Significant biological processes (upper panel) and disease biomarkers (lower panel)
 Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Progerin expression disrupts critical adult stem cell functions involved in tissue repair
 www.impactaging.com AGING, December 2014, Vol 6, N 12 Research Paper Progerin expression disrupts critical adult stem cell functions involved in tissue repair Laurin Marie Pacheco 1,2, Lourdes Adriana
www.impactaging.com AGING, December 2014, Vol 6, N 12 Research Paper Progerin expression disrupts critical adult stem cell functions involved in tissue repair Laurin Marie Pacheco 1,2, Lourdes Adriana
Prelamin A Acts to Accelerate Smooth Muscle Cell Senescence and Is a Novel Biomarker of Human Vascular Aging
 Prelamin A Acts to Accelerate Smooth Muscle Cell Senescence and Is a Novel Biomarker of Human Vascular Aging Cassandra D. Ragnauth, PhD*; Derek T. Warren, PhD*; Yiwen Liu, PhD; Rosamund McNair; Tamara
Prelamin A Acts to Accelerate Smooth Muscle Cell Senescence and Is a Novel Biomarker of Human Vascular Aging Cassandra D. Ragnauth, PhD*; Derek T. Warren, PhD*; Yiwen Liu, PhD; Rosamund McNair; Tamara
Progeria, the nucleolus and farnesyltransferase inhibitors
 Nuclear Envelope Disease and Chromatin Organization 2009 287 Progeria, the nucleolus and farnesyltransferase inhibitors Ishita S. Mehta*, Joanna M. Bridger* and Ian R. Kill 1 *Laboratory of Nuclear and
Nuclear Envelope Disease and Chromatin Organization 2009 287 Progeria, the nucleolus and farnesyltransferase inhibitors Ishita S. Mehta*, Joanna M. Bridger* and Ian R. Kill 1 *Laboratory of Nuclear and
Reversal of the cellular phenotype in the premature aging disease Hutchinson-Gilford progeria syndrome
 Reversal of the cellular phenotype in the premature aging disease Hutchinson-Gilford progeria syndrome Paola Scaffidi & Tom Misteli Hutchinson-Gilford progeria syndrome (HGPS) is a childhood premature
Reversal of the cellular phenotype in the premature aging disease Hutchinson-Gilford progeria syndrome Paola Scaffidi & Tom Misteli Hutchinson-Gilford progeria syndrome (HGPS) is a childhood premature
Proteomic profiling of small-molecule inhibitors reveals dispensability of MTH1 for cancer cell survival
 Supplementary Information for Proteomic profiling of small-molecule inhibitors reveals dispensability of MTH1 for cancer cell survival Tatsuro Kawamura 1, Makoto Kawatani 1, Makoto Muroi, Yasumitsu Kondoh,
Supplementary Information for Proteomic profiling of small-molecule inhibitors reveals dispensability of MTH1 for cancer cell survival Tatsuro Kawamura 1, Makoto Kawatani 1, Makoto Muroi, Yasumitsu Kondoh,
Nature Medicine: doi: /nm.4322
 1 2 3 4 5 6 7 8 9 10 11 Supplementary Figure 1. Predicted RNA structure of 3 UTR and sequence alignment of deleted nucleotides. (a) Predicted RNA secondary structure of ZIKV 3 UTR. The stem-loop structure
1 2 3 4 5 6 7 8 9 10 11 Supplementary Figure 1. Predicted RNA structure of 3 UTR and sequence alignment of deleted nucleotides. (a) Predicted RNA secondary structure of ZIKV 3 UTR. The stem-loop structure
EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric)
 EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric)
EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric)
Supplementary Figure 1 Expression of Crb3 in mouse sciatic nerve: biochemical analysis (a) Schematic of Crb3 isoforms, ERLI and CLPI, indicating the
 Supplementary Figure 1 Expression of Crb3 in mouse sciatic nerve: biochemical analysis (a) Schematic of Crb3 isoforms, ERLI and CLPI, indicating the location of the transmembrane (TM), FRM binding (FB)
Supplementary Figure 1 Expression of Crb3 in mouse sciatic nerve: biochemical analysis (a) Schematic of Crb3 isoforms, ERLI and CLPI, indicating the location of the transmembrane (TM), FRM binding (FB)
Focus Application. Compound-Induced Cytotoxicity
 xcelligence System Real-Time Cell Analyzer Focus Application Compound-Induced Cytotoxicity For life science research only. Not for use in diagnostic procedures. Featured Study: Using the Time Resolving
xcelligence System Real-Time Cell Analyzer Focus Application Compound-Induced Cytotoxicity For life science research only. Not for use in diagnostic procedures. Featured Study: Using the Time Resolving
Total Histone H3 Acetylation Detection Fast Kit (Colorimetric)
 Total Histone H3 Acetylation Detection Fast Kit (Colorimetric) Catalog Number KA1538 48 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use...
Total Histone H3 Acetylation Detection Fast Kit (Colorimetric) Catalog Number KA1538 48 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use...
Supplementary Figure 1
 Supplementary Figure 1 The average sigmoid parametric curves of capillary dilation time courses and average time to 50% peak capillary diameter dilation computed from individual capillary responses averaged
Supplementary Figure 1 The average sigmoid parametric curves of capillary dilation time courses and average time to 50% peak capillary diameter dilation computed from individual capillary responses averaged
Focus Application. Compound-Induced Cytotoxicity
 xcelligence System Real-Time Cell Analyzer Focus Application Compound-Induced Cytotoxicity Featured Study: Using the Time Resolving Function of the xcelligence System to Optimize Endpoint Viability and
xcelligence System Real-Time Cell Analyzer Focus Application Compound-Induced Cytotoxicity Featured Study: Using the Time Resolving Function of the xcelligence System to Optimize Endpoint Viability and
EpiQuik Total Histone H3 Acetylation Detection Fast Kit (Colorimetric)
 EpiQuik Total Histone H3 Acetylation Detection Fast Kit (Colorimetric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Total Histone H3 Acetylation Detection Fast Kit (Colorimetric)
EpiQuik Total Histone H3 Acetylation Detection Fast Kit (Colorimetric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Total Histone H3 Acetylation Detection Fast Kit (Colorimetric)
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Identification of Sickle Cells using Digital Image Processing. Academic Year Annexure I
 Academic Year 2014-15 Annexure I 1. Project Title: Identification of Sickle Cells using Digital Image Processing TABLE OF CONTENTS 1.1 Abstract 1-1 1.2 Motivation 1-1 1.3 Objective 2-2 2.1 Block Diagram
Academic Year 2014-15 Annexure I 1. Project Title: Identification of Sickle Cells using Digital Image Processing TABLE OF CONTENTS 1.1 Abstract 1-1 1.2 Motivation 1-1 1.3 Objective 2-2 2.1 Block Diagram
INTRODUCTION. This material is quoted from below PDF and modified.
 INTRODUCTION While drug response profiling of cancer cells in two-dimensional culture has been a mainstay of predictive biomarker discovery and anti-cancer drug development, there are aspects of tumor
INTRODUCTION While drug response profiling of cancer cells in two-dimensional culture has been a mainstay of predictive biomarker discovery and anti-cancer drug development, there are aspects of tumor
Nature Biotechnology: doi: /nbt.3828
 Supplementary Figure 1 Development of a FRET-based MCS. (a) Linker and MA2 modification are indicated by single letter amino acid code. indicates deletion of amino acids and N or C indicate the terminus
Supplementary Figure 1 Development of a FRET-based MCS. (a) Linker and MA2 modification are indicated by single letter amino acid code. indicates deletion of amino acids and N or C indicate the terminus
Progeria. Premature human aging: t he progerias. Progerias as models for aging. Hutchinson-Gilford syndrome. Definition:
 Premature human aging: t he progerias Reading: Genetic alterations in accelerated ageing syndromes Do they play a role in natural ageing? Monika Puzianowska- Kuznicka. Jacek Kuznicki. 2005. IJBCB, 37;
Premature human aging: t he progerias Reading: Genetic alterations in accelerated ageing syndromes Do they play a role in natural ageing? Monika Puzianowska- Kuznicka. Jacek Kuznicki. 2005. IJBCB, 37;
Supplementary Information. Preferential associations between co-regulated genes reveal a. transcriptional interactome in erythroid cells
 Supplementary Information Preferential associations between co-regulated genes reveal a transcriptional interactome in erythroid cells Stefan Schoenfelder, * Tom Sexton, * Lyubomira Chakalova, * Nathan
Supplementary Information Preferential associations between co-regulated genes reveal a transcriptional interactome in erythroid cells Stefan Schoenfelder, * Tom Sexton, * Lyubomira Chakalova, * Nathan
EPIGENTEK. EpiQuik Global Acetyl Histone H3K27 Quantification Kit (Colorimetric) Base Catalog # P-4059 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Global Acetyl Histone H3K27 Quantification Kit (Colorimetric) Base Catalog # P-4059 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Global Acetyl Histone H3K27 Quantification Kit (Colorimetric)
EpiQuik Global Acetyl Histone H3K27 Quantification Kit (Colorimetric) Base Catalog # P-4059 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Global Acetyl Histone H3K27 Quantification Kit (Colorimetric)
Longitudinal tracking of single live cancer cells to understand cell cycle effects of the
 Longitudinal tracking of single live cancer cells to understand cell cycle effects of the nuclear export inhibitor, selinexor Joshua M. Marcus 1, Russell T. Burke 1, John A. DeSisto 1, Yosef Landesman
Longitudinal tracking of single live cancer cells to understand cell cycle effects of the nuclear export inhibitor, selinexor Joshua M. Marcus 1, Russell T. Burke 1, John A. DeSisto 1, Yosef Landesman
Eiger BioPharmaceuticals Reports on 2018 R&D Day
 Eiger BioPharmaceuticals Reports on 2018 R&D Day Late Stage Rare and Ultra-Rare Disease Pipeline Advancing >$100M in Cash Available to Achieve Key Milestones PALO ALTO, Calif., December 11, 2018 Eiger
Eiger BioPharmaceuticals Reports on 2018 R&D Day Late Stage Rare and Ultra-Rare Disease Pipeline Advancing >$100M in Cash Available to Achieve Key Milestones PALO ALTO, Calif., December 11, 2018 Eiger
In Vitro Studies of the Aurora-Kinase Inhibitor MLN8237 in Prostate Cancer Cell Lines
 In Vitro Studies of the Aurora-Kinase Inhibitor MLN837 in Prostate Cancer Cell Lines Nicole V. Julian Mentor: Ana Aparicio, M.D. Special Thanks to Brittany Kleb Department of Genitourinary Medical Oncology
In Vitro Studies of the Aurora-Kinase Inhibitor MLN837 in Prostate Cancer Cell Lines Nicole V. Julian Mentor: Ana Aparicio, M.D. Special Thanks to Brittany Kleb Department of Genitourinary Medical Oncology
TFEB-mediated increase in peripheral lysosomes regulates. Store Operated Calcium Entry
 TFEB-mediated increase in peripheral lysosomes regulates Store Operated Calcium Entry Luigi Sbano, Massimo Bonora, Saverio Marchi, Federica Baldassari, Diego L. Medina, Andrea Ballabio, Carlotta Giorgi
TFEB-mediated increase in peripheral lysosomes regulates Store Operated Calcium Entry Luigi Sbano, Massimo Bonora, Saverio Marchi, Federica Baldassari, Diego L. Medina, Andrea Ballabio, Carlotta Giorgi
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
IN the spring of 2007, a 2-year clinical drug trial began at
 Journal of Gerontology: BIOLOGICAL SCIENCES 2008, Vol. 63A, No. 8, 777 787 Copyright 2008 by The Gerontological Society of America Meeting Report Highlights of the 2007 Progeria Research Foundation Scientific
Journal of Gerontology: BIOLOGICAL SCIENCES 2008, Vol. 63A, No. 8, 777 787 Copyright 2008 by The Gerontological Society of America Meeting Report Highlights of the 2007 Progeria Research Foundation Scientific
STUDIES ON MUSTARD-STIMULATED PROTEASES AND INHIBITORS IN HUMAN EPIDERMAL KERATINOCYTES (HEK): DEVELOPMENT OF ANTIVESICANT DRUGS
 STUDIES ON MUSTARD-STIMULATED PROTEASES AND INHIBITORS IN HUMAN EPIDERMAL KERATINOCYTES (HEK): DEVELOPMENT OF ANTIVESICANT DRUGS Xiannu Jin 1, Radharaman Ray 2, Guang Xu 1 and Prabhati Ray 1 1 Department
STUDIES ON MUSTARD-STIMULATED PROTEASES AND INHIBITORS IN HUMAN EPIDERMAL KERATINOCYTES (HEK): DEVELOPMENT OF ANTIVESICANT DRUGS Xiannu Jin 1, Radharaman Ray 2, Guang Xu 1 and Prabhati Ray 1 1 Department
NF-κB p65 (Phospho-Thr254)
 Assay Biotechnology Company www.assaybiotech.com Tel: 1-877-883-7988 Fax: 1-877-610-9758 NF-κB p65 (Phospho-Thr254) Colorimetric Cell-Based ELISA Kit Catalog #: OKAG02015 Please read the provided manual
Assay Biotechnology Company www.assaybiotech.com Tel: 1-877-883-7988 Fax: 1-877-610-9758 NF-κB p65 (Phospho-Thr254) Colorimetric Cell-Based ELISA Kit Catalog #: OKAG02015 Please read the provided manual
Clinical Cell Biology Organelles in Health and Disease
 Department of Ophthalmology University of Kiel, University Medical Center Director: Prof. Dr. Johann Roider Clinical Cell Biology Organelles in Health and Disease Prof. Dr. Alexa Klettner Clinical cell
Department of Ophthalmology University of Kiel, University Medical Center Director: Prof. Dr. Johann Roider Clinical Cell Biology Organelles in Health and Disease Prof. Dr. Alexa Klettner Clinical cell
Figure S1. Standard curves for amino acid bioassays. (A) The standard curve for leucine concentration versus OD600 values for L. casei.
 Figure S1. Standard curves for amino acid bioassays. (A) The standard curve for leucine concentration versus OD600 values for L. casei. (B) The standard curve for lysine concentrations versus OD600 values
Figure S1. Standard curves for amino acid bioassays. (A) The standard curve for leucine concentration versus OD600 values for L. casei. (B) The standard curve for lysine concentrations versus OD600 values
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Lu et al., http://www.jcb.org/cgi/content/full/jcb.201012063/dc1 Figure S1. Kinetics of nuclear envelope assembly, recruitment of Nup133
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Lu et al., http://www.jcb.org/cgi/content/full/jcb.201012063/dc1 Figure S1. Kinetics of nuclear envelope assembly, recruitment of Nup133
Mechanisms of genome instability in Hutchinson-Gilford progeria
 Front. Biol. 2017, 12(1): 49 62 DOI 10.1007/s11515-016-1435-x REVIEW Mechanisms of genome instability in Hutchinson-Gilford progeria Haoyue Zhang, Kan Cao ( ) Department of Cell Biology and Molecular Genetics,
Front. Biol. 2017, 12(1): 49 62 DOI 10.1007/s11515-016-1435-x REVIEW Mechanisms of genome instability in Hutchinson-Gilford progeria Haoyue Zhang, Kan Cao ( ) Department of Cell Biology and Molecular Genetics,
Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation.
 Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation. (a) Embryonic fibroblasts isolated from wildtype (WT), BRAF -/-, or CRAF -/- mice were irradiated (6 Gy) and DNA damage
Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation. (a) Embryonic fibroblasts isolated from wildtype (WT), BRAF -/-, or CRAF -/- mice were irradiated (6 Gy) and DNA damage
Use of online cell counting for micronucleus and neurite outgrowth assays on IN Cell Analyzer 2000
 GE Healthcare Application note 28-9673-96 AA IN Cell Analyzer 2000 Use of online cell counting for micronucleus and neurite outgrowth assays on IN Cell Analyzer 2000 Key words: IN Cell Analyzer online
GE Healthcare Application note 28-9673-96 AA IN Cell Analyzer 2000 Use of online cell counting for micronucleus and neurite outgrowth assays on IN Cell Analyzer 2000 Key words: IN Cell Analyzer online
SUPPLEMENTARY INFORMATION
 DOI: 1.138/ncb222 / b. WB anti- WB anti- ulin Mitotic index (%) 14 1 6 2 T (h) 32 48-1 1 2 3 4 6-1 4 16 22 28 3 33 e. 6 4 2 Time (min) 1-6- 11-1 > 1 % cells Figure S1 depletion leads to mitotic defects
DOI: 1.138/ncb222 / b. WB anti- WB anti- ulin Mitotic index (%) 14 1 6 2 T (h) 32 48-1 1 2 3 4 6-1 4 16 22 28 3 33 e. 6 4 2 Time (min) 1-6- 11-1 > 1 % cells Figure S1 depletion leads to mitotic defects
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2
 Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Supplementary Figure 1. Mother centrioles can reduplicate while in the close association
 C1-GFP distance (nm) C1-GFP distance (nm) a arrested HeLa cell expressing C1-GFP and Plk1TD-RFP -3 s 1 2 3 4 5 6 7 8 9 11 12 13 14 16 17 18 19 2 21 22 23 24 26 27 28 29 3 b 9 8 7 6 5 4 3 2 arrested HeLa
C1-GFP distance (nm) C1-GFP distance (nm) a arrested HeLa cell expressing C1-GFP and Plk1TD-RFP -3 s 1 2 3 4 5 6 7 8 9 11 12 13 14 16 17 18 19 2 21 22 23 24 26 27 28 29 3 b 9 8 7 6 5 4 3 2 arrested HeLa
PHYSIOEX 3.0 EXERCISE 33B: CARDIOVASCULAR DYNAMICS
 PHYSIOEX 3.0 EXERCISE 33B: CARDIOVASCULAR DYNAMICS Objectives 1. To define the following: blood flow; viscosity; peripheral resistance; systole; diastole; end diastolic volume; end systolic volume; stroke
PHYSIOEX 3.0 EXERCISE 33B: CARDIOVASCULAR DYNAMICS Objectives 1. To define the following: blood flow; viscosity; peripheral resistance; systole; diastole; end diastolic volume; end systolic volume; stroke
Superior Fluorescent Labeling Dyes Spanning the Full Visible Spectrum...1. Trademarks: HiLyte Fluor (AnaSpec, Inc.)
 Table of Contents Fluor TM Labeling Dyes Superior Fluorescent Labeling Dyes Spanning the Full Visible Spectrum....1 Fluor TM 405 Dye, an Excellent Alternative to Alexa Fluor 405 & DyLight 405....2 Fluor
Table of Contents Fluor TM Labeling Dyes Superior Fluorescent Labeling Dyes Spanning the Full Visible Spectrum....1 Fluor TM 405 Dye, an Excellent Alternative to Alexa Fluor 405 & DyLight 405....2 Fluor
Fig. S1. Schematic representation of the two RBC membrane:cytoskeleton anchorage
 Supplemental data Fig. S1. Schematic representation of the two RBC membrane:cytoskeleton anchorage complexes. Left, 4.1R complex; right, ankyrin-based complex. Adapted from (1) where abbreviations are
Supplemental data Fig. S1. Schematic representation of the two RBC membrane:cytoskeleton anchorage complexes. Left, 4.1R complex; right, ankyrin-based complex. Adapted from (1) where abbreviations are
1 Introduction. Abstract: Accurate optic disc (OD) segmentation and fovea. Keywords: optic disc segmentation, fovea detection.
 Current Directions in Biomedical Engineering 2017; 3(2): 533 537 Caterina Rust*, Stephanie Häger, Nadine Traulsen and Jan Modersitzki A robust algorithm for optic disc segmentation and fovea detection
Current Directions in Biomedical Engineering 2017; 3(2): 533 537 Caterina Rust*, Stephanie Häger, Nadine Traulsen and Jan Modersitzki A robust algorithm for optic disc segmentation and fovea detection
Supplemental Information. Otic Mesenchyme Cells Regulate. Spiral Ganglion Axon Fasciculation. through a Pou3f4/EphA4 Signaling Pathway
 Neuron, Volume 73 Supplemental Information Otic Mesenchyme Cells Regulate Spiral Ganglion Axon Fasciculation through a Pou3f4/EphA4 Signaling Pathway Thomas M. Coate, Steven Raft, Xiumei Zhao, Aimee K.
Neuron, Volume 73 Supplemental Information Otic Mesenchyme Cells Regulate Spiral Ganglion Axon Fasciculation through a Pou3f4/EphA4 Signaling Pathway Thomas M. Coate, Steven Raft, Xiumei Zhao, Aimee K.
THE ROLE OF ALTERED CALCIUM AND mtor SIGNALING IN THE PATHOGENESIS OF CYSTINOSIS
 Research Foundation, 18 month progress report THE ROLE OF ALTERED CALCIUM AND mtor SIGNALING IN THE PATHOGENESIS OF CYSTINOSIS Ekaterina Ivanova, doctoral student Elena Levtchenko, MD, PhD, PI Antonella
Research Foundation, 18 month progress report THE ROLE OF ALTERED CALCIUM AND mtor SIGNALING IN THE PATHOGENESIS OF CYSTINOSIS Ekaterina Ivanova, doctoral student Elena Levtchenko, MD, PhD, PI Antonella
Rescue of heterochromatin organization in Hutchinson- Gilford progeria by drug treatment
 Cell. Mol. Life Sci. 62 (2005) 2669 2678 1420-682X/05/222669-10 DOI 10.1007/s00018-005-5318-6 Birkhäuser Verlag, Basel, 2005 Cellular and Molecular Life Sciences Research Article Rescue of heterochromatin
Cell. Mol. Life Sci. 62 (2005) 2669 2678 1420-682X/05/222669-10 DOI 10.1007/s00018-005-5318-6 Birkhäuser Verlag, Basel, 2005 Cellular and Molecular Life Sciences Research Article Rescue of heterochromatin
Supplemental Data Figure S1 Effect of TS2/4 and R6.5 antibodies on the kinetics of CD16.NK-92-mediated specific lysis of SKBR-3 target cells.
 Supplemental Data Figure S1. Effect of TS2/4 and R6.5 antibodies on the kinetics of CD16.NK-92-mediated specific lysis of SKBR-3 target cells. (A) Specific lysis of IFN-γ-treated SKBR-3 cells in the absence
Supplemental Data Figure S1. Effect of TS2/4 and R6.5 antibodies on the kinetics of CD16.NK-92-mediated specific lysis of SKBR-3 target cells. (A) Specific lysis of IFN-γ-treated SKBR-3 cells in the absence
APPLICATION SPECIFIC PROTOCOL ANGIOGENESIS... 1 TABLE OF CONTENTS... 1 MONOLAYER FORMATION... 2 OPTION 1: APPLICATION OF ANGIOGENIC STIMULI...
 APPLICATION SPECIFIC PROTOCOL ANGIOGENESIS AIM 3D Cell Culture Chips offer a new perspective in studying angiogenesis by allowing the growth of new vascular sprouts in a 3D matrix from a pre-existing endothelial
APPLICATION SPECIFIC PROTOCOL ANGIOGENESIS AIM 3D Cell Culture Chips offer a new perspective in studying angiogenesis by allowing the growth of new vascular sprouts in a 3D matrix from a pre-existing endothelial
Supplementary Figure 1. Generation of knockin mice expressing L-selectinN138G. (a) Schematics of the Sellg allele (top), the targeting vector, the
 Supplementary Figure 1. Generation of knockin mice expressing L-selectinN138G. (a) Schematics of the Sellg allele (top), the targeting vector, the targeted allele in ES cells, and the mutant allele in
Supplementary Figure 1. Generation of knockin mice expressing L-selectinN138G. (a) Schematics of the Sellg allele (top), the targeting vector, the targeted allele in ES cells, and the mutant allele in
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
SUPPLEMENTARY INFORMATION
 b 350 300 250 200 150 100 50 0 E0 E10 E50 E0 E10 E50 E0 E10 E50 E0 E10 E50 Number of organoids per well 350 300 250 200 150 100 50 0 R0 R50 R100 R500 1st 2nd 3rd Noggin 100 ng/ml Noggin 10 ng/ml Noggin
b 350 300 250 200 150 100 50 0 E0 E10 E50 E0 E10 E50 E0 E10 E50 E0 E10 E50 Number of organoids per well 350 300 250 200 150 100 50 0 R0 R50 R100 R500 1st 2nd 3rd Noggin 100 ng/ml Noggin 10 ng/ml Noggin
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH
 EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
Structural Variation and Medical Genomics
 Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Supplementary Materials for
 www.sciencetranslationalmedicine.org/cgi/content/full/4/117/117ra8/dc1 Supplementary Materials for Notch4 Normalization Reduces Blood Vessel Size in Arteriovenous Malformations Patrick A. Murphy, Tyson
www.sciencetranslationalmedicine.org/cgi/content/full/4/117/117ra8/dc1 Supplementary Materials for Notch4 Normalization Reduces Blood Vessel Size in Arteriovenous Malformations Patrick A. Murphy, Tyson
International Journal of Pharma and Bio Sciences
 International Journal of Pharma and Bio Sciences HUTCHINSON-GILFORD SYNDROME (PROGERIA): A REVIEW SACHIN KUMAR 1 *, AMIT KUMAR 2, MOHIT SINGLA 1 AND ABHISHEK SINGH 3 1. K.N.G.D Modi Institute of Pharmaceutical
International Journal of Pharma and Bio Sciences HUTCHINSON-GILFORD SYNDROME (PROGERIA): A REVIEW SACHIN KUMAR 1 *, AMIT KUMAR 2, MOHIT SINGLA 1 AND ABHISHEK SINGH 3 1. K.N.G.D Modi Institute of Pharmaceutical
Figure S1. PMVs from THP-1 cells expose phosphatidylserine and carry actin. A) Flow
 SUPPLEMENTARY DATA Supplementary Figure Legends Figure S1. PMVs from THP-1 cells expose phosphatidylserine and carry actin. A) Flow cytometry analysis of PMVs labelled with annexin-v-pe (Guava technologies)
SUPPLEMENTARY DATA Supplementary Figure Legends Figure S1. PMVs from THP-1 cells expose phosphatidylserine and carry actin. A) Flow cytometry analysis of PMVs labelled with annexin-v-pe (Guava technologies)
SUPPLEMENT. Materials and methods
 SUPPLEMENT Materials and methods Cell culture and reagents Cell media and reagents were from Invitrogen unless otherwise indicated. Antibiotics and Tet-certified serum were from Clontech. In experiments
SUPPLEMENT Materials and methods Cell culture and reagents Cell media and reagents were from Invitrogen unless otherwise indicated. Antibiotics and Tet-certified serum were from Clontech. In experiments
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
A Review on Hutchinson-gilford Syndrome PROGERIA. Kinja K. and Gupta N. NIMS Institute of Pharmacy, Shobha Nagar, Jaipur , Rajasthan, India
 I J P International Journal of Pharmagenesis 2(1), January-June 2011, pp. 115-120 A Review on Hutchinson-gilford Syndrome PROGERIA Kinja K. and Gupta N. NIMS Institute of Pharmacy, Shobha Nagar, Jaipur-303121,
I J P International Journal of Pharmagenesis 2(1), January-June 2011, pp. 115-120 A Review on Hutchinson-gilford Syndrome PROGERIA Kinja K. and Gupta N. NIMS Institute of Pharmacy, Shobha Nagar, Jaipur-303121,
SUPPLEMENTARY INFORMATION
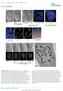 DOI: 0.038/ncb33 a b c 0 min 6 min 7 min (fixed) DIC -GFP, CenpF 3 µm Nocodazole Single optical plane -GFP, CenpF Max. intensity projection d µm -GFP, CenpF, -GFP CenpF 3-D rendering e f 0 min 4 min 0
DOI: 0.038/ncb33 a b c 0 min 6 min 7 min (fixed) DIC -GFP, CenpF 3 µm Nocodazole Single optical plane -GFP, CenpF Max. intensity projection d µm -GFP, CenpF, -GFP CenpF 3-D rendering e f 0 min 4 min 0
Moore s law in information technology. exponential growth!
 Moore s law in information technology exponential growth! ... and how it compares to developments in bio sciences Biology is driven by an avalanche of new information... DNA sequencing methods and throughput
Moore s law in information technology exponential growth! ... and how it compares to developments in bio sciences Biology is driven by an avalanche of new information... DNA sequencing methods and throughput
Expression of acid base transporters in the kidney collecting duct in Slc2a7 -/-
 Supplemental Material Results. Expression of acid base transporters in the kidney collecting duct in Slc2a7 -/- and Slc2a7 -/- mice. The expression of AE1 in the kidney was examined in Slc26a7 KO mice.
Supplemental Material Results. Expression of acid base transporters in the kidney collecting duct in Slc2a7 -/- and Slc2a7 -/- mice. The expression of AE1 in the kidney was examined in Slc26a7 KO mice.
The Cell Cycle. Materials 2 pipe cleaners of one color and 2 of another color Drawing paper
 The Cell Cycle Introduction When the cell has reached its growth potential it will begin to divide. Additionally, if a cell has become damaged or worn out it can be replaced by surrounding cells through
The Cell Cycle Introduction When the cell has reached its growth potential it will begin to divide. Additionally, if a cell has become damaged or worn out it can be replaced by surrounding cells through
of an untreated HS-stained BAEC monolayer viewed using a laser confocal microscope; Bar = 10 µm.
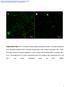 Supplemental Figure 1: EC monolayer heparan sulfate fluorescence intensity. (A) Enface perspective of an untreated HS-stained BAEC monolayer viewed using a laser confocal microscope; Bar = 10 µm. (B) Enface
Supplemental Figure 1: EC monolayer heparan sulfate fluorescence intensity. (A) Enface perspective of an untreated HS-stained BAEC monolayer viewed using a laser confocal microscope; Bar = 10 µm. (B) Enface
Cerebral Small Vessel Disease and HAND in ARV-treated Subjects
 Cerebral Small Vessel Disease and HAND in ARV-treated Subjects Cristian L. Achim, MD, PhD Ronald J. Ellis, MD, PhD Virawudh Soontornniyomkij, MD ARROW, Bucharest, Romania October 5-6, 2015 Rationale and
Cerebral Small Vessel Disease and HAND in ARV-treated Subjects Cristian L. Achim, MD, PhD Ronald J. Ellis, MD, PhD Virawudh Soontornniyomkij, MD ARROW, Bucharest, Romania October 5-6, 2015 Rationale and
Requirements for Efficient Proteolytic Cleavage of Prelamin A by ZMPSTE24
 Requirements for Efficient Proteolytic Cleavage of Prelamin A by ZMPSTE24 Jemima Barrowman, Corinne Hamblet, Megan S. Kane, Susan Michaelis* Department of Cell Biology, The Johns Hopkins School of Medicine,
Requirements for Efficient Proteolytic Cleavage of Prelamin A by ZMPSTE24 Jemima Barrowman, Corinne Hamblet, Megan S. Kane, Susan Michaelis* Department of Cell Biology, The Johns Hopkins School of Medicine,
Influenza virus exploits tunneling nanotubes for cell-to-cell spread
 Supplementary Information Influenza virus exploits tunneling nanotubes for cell-to-cell spread Amrita Kumar 1, Jin Hyang Kim 1, Priya Ranjan 1, Maureen G. Metcalfe 2, Weiping Cao 1, Margarita Mishina 1,
Supplementary Information Influenza virus exploits tunneling nanotubes for cell-to-cell spread Amrita Kumar 1, Jin Hyang Kim 1, Priya Ranjan 1, Maureen G. Metcalfe 2, Weiping Cao 1, Margarita Mishina 1,
Relative SOD1 activity. Relative SOD2 activity. Relative SOD activity (Infected:Mock) + CP + DDC
 Supplementary Figure 1. SOD1 activity is significantly increased relative to SOD1 levels. SOD1 and SOD2 activities in the infected mork13 cells are shown normalised to their corresponding levels and relative
Supplementary Figure 1. SOD1 activity is significantly increased relative to SOD1 levels. SOD1 and SOD2 activities in the infected mork13 cells are shown normalised to their corresponding levels and relative
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
QUANTIFICATION OF PROGRESSION OF RETINAL NERVE FIBER LAYER ATROPHY IN FUNDUS PHOTOGRAPH
 QUANTIFICATION OF PROGRESSION OF RETINAL NERVE FIBER LAYER ATROPHY IN FUNDUS PHOTOGRAPH Hyoun-Joong Kong *, Jong-Mo Seo **, Seung-Yeop Lee *, Hum Chung **, Dong Myung Kim **, Jeong Min Hwang **, Kwang
QUANTIFICATION OF PROGRESSION OF RETINAL NERVE FIBER LAYER ATROPHY IN FUNDUS PHOTOGRAPH Hyoun-Joong Kong *, Jong-Mo Seo **, Seung-Yeop Lee *, Hum Chung **, Dong Myung Kim **, Jeong Min Hwang **, Kwang
Automated Assessment of Diabetic Retinal Image Quality Based on Blood Vessel Detection
 Y.-H. Wen, A. Bainbridge-Smith, A. B. Morris, Automated Assessment of Diabetic Retinal Image Quality Based on Blood Vessel Detection, Proceedings of Image and Vision Computing New Zealand 2007, pp. 132
Y.-H. Wen, A. Bainbridge-Smith, A. B. Morris, Automated Assessment of Diabetic Retinal Image Quality Based on Blood Vessel Detection, Proceedings of Image and Vision Computing New Zealand 2007, pp. 132
HYBRID GENE THERAPY FOR AD-EDMD
 HYBRID GENE THERAPY FOR AD-EDMD Gene Therapy Prof. Isabella Saggio 2017/2018 Bertani Camilla Dezi Clara Difeo Giorgia di Palma Carmen AUTOSOMAL DOMINANT EMERY-DREIFUSS MUSCULAR DYSTROPHY Fig.1 Adapted
HYBRID GENE THERAPY FOR AD-EDMD Gene Therapy Prof. Isabella Saggio 2017/2018 Bertani Camilla Dezi Clara Difeo Giorgia di Palma Carmen AUTOSOMAL DOMINANT EMERY-DREIFUSS MUSCULAR DYSTROPHY Fig.1 Adapted
Supplementary Figure 1: GFAP positive nerves in patients with adenocarcinoma of
 SUPPLEMENTARY FIGURES AND MOVIE LEGENDS Supplementary Figure 1: GFAP positive nerves in patients with adenocarcinoma of the pancreas. (A) Images of nerves stained for GFAP (green), S100 (red) and DAPI
SUPPLEMENTARY FIGURES AND MOVIE LEGENDS Supplementary Figure 1: GFAP positive nerves in patients with adenocarcinoma of the pancreas. (A) Images of nerves stained for GFAP (green), S100 (red) and DAPI
Supplementary Figure 1 Induction of cellular senescence and isolation of exosome. a to c, Pre-senescent primary normal human diploid fibroblasts
 Supplementary Figure 1 Induction of cellular senescence and isolation of exosome. a to c, Pre-senescent primary normal human diploid fibroblasts (TIG-3 cells) were rendered senescent by either serial passage
Supplementary Figure 1 Induction of cellular senescence and isolation of exosome. a to c, Pre-senescent primary normal human diploid fibroblasts (TIG-3 cells) were rendered senescent by either serial passage
Supplemental Information. Myocardial Polyploidization Creates a Barrier. to Heart Regeneration in Zebrafish
 Developmental Cell, Volume 44 Supplemental Information Myocardial Polyploidization Creates a Barrier to Heart Regeneration in Zebrafish Juan Manuel González-Rosa, Michka Sharpe, Dorothy Field, Mark H.
Developmental Cell, Volume 44 Supplemental Information Myocardial Polyploidization Creates a Barrier to Heart Regeneration in Zebrafish Juan Manuel González-Rosa, Michka Sharpe, Dorothy Field, Mark H.
Flow Quantification from 2D Phase Contrast MRI in Renal Arteries using Clustering
 Flow Quantification from 2D Phase Contrast MRI in Renal Arteries using Clustering Frank G. Zöllner 1,2, Jan Ankar Monnsen 1, Arvid Lundervold 2, Jarle Rørvik 1 1 Department for Radiology, University of
Flow Quantification from 2D Phase Contrast MRI in Renal Arteries using Clustering Frank G. Zöllner 1,2, Jan Ankar Monnsen 1, Arvid Lundervold 2, Jarle Rørvik 1 1 Department for Radiology, University of
Image Processing and Data Mining Techniques in the Detection of Diabetic Retinopathy in Fundus Images
 I J C T A, 10(8), 2017, pp. 755-762 International Science Press ISSN: 0974-5572 Image Processing and Data Mining Techniques in the Detection of Diabetic Retinopathy in Fundus Images Payal M. Bante* and
I J C T A, 10(8), 2017, pp. 755-762 International Science Press ISSN: 0974-5572 Image Processing and Data Mining Techniques in the Detection of Diabetic Retinopathy in Fundus Images Payal M. Bante* and
Supplementary materials
 Supplementary materials Chemical library from ChemBridge 50,240 structurally diverse small molecule compounds dissolved in DMSO Hits Controls: No virus added μ Primary screening at 20 g/ml of compounds
Supplementary materials Chemical library from ChemBridge 50,240 structurally diverse small molecule compounds dissolved in DMSO Hits Controls: No virus added μ Primary screening at 20 g/ml of compounds
A novel and automatic pectoral muscle identification algorithm for mediolateral oblique (MLO) view mammograms using ImageJ
 A novel and automatic pectoral muscle identification algorithm for mediolateral oblique (MLO) view mammograms using ImageJ Chao Wang Wolfson Institute of Preventive Medicine Queen Mary University of London
A novel and automatic pectoral muscle identification algorithm for mediolateral oblique (MLO) view mammograms using ImageJ Chao Wang Wolfson Institute of Preventive Medicine Queen Mary University of London
Image Analysis and Cytometry in Three-Dimensional Digital Reconstruction of Porcine Native Aortic Valve Leaflets
 Image Analysis and Cytometry in Three-Dimensional Digital Reconstruction of Porcine Native Aortic Valve Leaflets Introduction Chi Zheng, M1 - University of Pittsburgh; BSE - University of Michigan In association
Image Analysis and Cytometry in Three-Dimensional Digital Reconstruction of Porcine Native Aortic Valve Leaflets Introduction Chi Zheng, M1 - University of Pittsburgh; BSE - University of Michigan In association
Microtubule Teardrop Patterns
 Supporting Information Microtubule Teardrop Patterns Kosuke Okeyoshi 1, Ryuzo Kawamura 1, Ryo Yoshida 2, and Yoshihito Osada 1 * 1 RIKEN Advanced Science Institute, 2-1 Hirosawa, Wako-shi, Saitama 351-0198,
Supporting Information Microtubule Teardrop Patterns Kosuke Okeyoshi 1, Ryuzo Kawamura 1, Ryo Yoshida 2, and Yoshihito Osada 1 * 1 RIKEN Advanced Science Institute, 2-1 Hirosawa, Wako-shi, Saitama 351-0198,
Assay Name: Transwell migration using DAPI
 Assay Name: Transwell migration using DAPI Assay ID: Celigo_04_0001 Table of Contents Experiment: Identification of cells which have migrated through the membrane of a transwell insert....2 Celigo Setup...2
Assay Name: Transwell migration using DAPI Assay ID: Celigo_04_0001 Table of Contents Experiment: Identification of cells which have migrated through the membrane of a transwell insert....2 Celigo Setup...2
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Inhibiting farnesylation of progerin prevents the characteristic nuclear blebbing of Hutchinson Gilford progeria syndrome
 Inhibiting farnesylation of progerin prevents the characteristic nuclear blebbing of Hutchinson Gilford progeria syndrome Brian C. Capell*, Michael R. Erdos*, James P. Madigan, James J. Fiordalisi, Renee
Inhibiting farnesylation of progerin prevents the characteristic nuclear blebbing of Hutchinson Gilford progeria syndrome Brian C. Capell*, Michael R. Erdos*, James P. Madigan, James J. Fiordalisi, Renee

 Supplementary Figure 1 (Mu) SBP (mmhg) 2 18 16 p
Supplementary Figure 1 (Mu) SBP (mmhg) 2 18 16 p