RNA DECAY PART I - GENERAL MECHANISMS PART II - SPECIFIC PATHWAYS
|
|
|
- Dinah Page
- 5 years ago
- Views:
Transcription
1 RNA DECAY PART I - GENERAL MECHANISMS PART II - SPECIFIC PATHWAYS
2 REGULATION OF GENE EXPRESSION 1) PolII assembly 2) transcription 3) mrna processing 4) mrna export 5) translation 6) protein stability 7) RNA degradation
3 RNases Endonucleases processing (RNase P, RNase III, RNase E): specific, cleavage results in 3 -OH and 5 -P (monophosphate) degrading (RNase I, RNAse A): unspecific, cleavage results in 5 -OH and 3 -P (cyclic phosphate) Exonucleases hydrolytic: attacking group H 2 O, results in 3 -OH and 5 -P phosphorolytic: attacking group inorganic phosphate, results in 3 -OH and 5 -PP distributive processive Symmons et al, TiBS, 2002
4 Prokaryotic RNases Family RNases Characteristics Exonucleases 3 5 RNR RNase II nonspecific processive, degrades only ssrna, mrna decay RNase R nonspecific processive, degrades ssrna and dsrna, mrna decay DEDD RNase D distributive, small RNA and stabile RNA processing RNase T Oligoribonuclease specific for oligoribonucleotides RBN RNase BN/Z distributive exonuclease 3-5 and endonuclease, trna processing PDX PNPase phosphorolytic processive, degradosome subunit, KH/S1 RNA BD domains, degrades ss/dsrna RNase PH phosphorolytic distributive Exonucleases 5 3 *RNAse J1/J2 present in Bacillus subtilis, specific for 5 monop ssrna, mrna decay Endonucleases RNase III RNase E RNase G RNase I RNase H RNase P RNase Z Rae1/YacP *RNAse J1/J2 MazF/EndoA RNAse M5 dsrna specific, rrna, trna, mrna processing, mrna degradation degradosome subunit, mrna decay; rrna trna and RNaseP RNA processing similar to RNase E nonspecific, mrna degradation specific for RNA:DNA hybrid trna 5 end processing trna 3 end processing ribosome-dependent mrna decay in Bacillus subtilis mrna decay in Bacillus subtilis toxin, mrna degradation in stress conditions, sequence specific
5 Mackie, Nat. Rev. Microbiol, 2012
6 Bacterial exo- and endo-nucleases Huiet al, Annu Rev genet, 2015 Novel bacterial endonuclease Rae1 involved in ribosomedependent mrna decay in Bacillus subtilis Leroy et al, EMBO, 2017
7 Degradation of bacterial mrnas Degradosome - major complex involved in mrna decay in bacteria, functions as dimer RNase E PNPase RhIB Enolase additional 5 -phosphate -dependent endoribonuclease, N-terminal nucleolytic domain, C-terminal protein binding domain, central RNA binding domain (BD) phosphorolytic processive exonuclease 3-5, KH and S1 RNA BD ATP-dependent helicase, DEAD box, stimulate degradation of structured RNA regions glycolytic enzyme DnaK/GroEL chaperons, poliphosphate kinase, poly(a) polymerase, S1 ribosomal protein RNase E NH 2 Catalytic domain RNA BD (RRD) C-terminal COOH PNPaza PDX domain RNA BD (KH) RNA BD (S1) RhIB PNPaza PNPazy trimer Symmons et al, Structure, 2000 Symmons et al, TiBS, 2002
8 Degradation of bacterial mrnas General endo-dependend pathway 3 exo-dependend pathway RNAse E or Y 5 -dependent pathway Huiet al, Annu Rev genet, 2015
9 Degradation of bacterial mrnas 3 end stem-loop structure of transcripts targeted for degradation becomes often polyadenylated by PAP (poly(a) polymerase) and PNPase (polynucleotide phosphatase), with the help of Hfq (hexameric RNA chaperone). RNase E cleavage initiates degradation by 3-5 exonucleases, mainly RNase II, RNase R and PNPase. PNPase Hfq PAP I Mohanty et al, Mol. Microbiol., 2004 Symmons et al, TiBS, 2002
10 Protein Function Characteristics Exonucleases 5 3 Xrn1 Rat1 Rrp17/Nol12 Eukaryotic RNases cytoplasmic, mrna degradation nuclear, pre-rrna, sn/snorna, pre-mrna processing and degradation nuclear, pre-rrna processing Exosome 3 5 multisubunit exo/endo complex subunits organized as in bacterial PNPazy Rrp44/Dis3 catalytic subunit Exo/PIN domains, distributive, hydrolytic Rrp4, Rrp40 pre-rrna, sn/snorna processing, mrna degradation Rrp41-43, participates in NMD, ARE-dependent, non-stop decay Mtr3, Ski4 Mtr4 nuclear helicase cofactor Rrp6 / Rrp47p exonuclease nuclear cofactor RNase D homolog /RNA binding Ski2,3,7,8 cytoplasmic exosome cofactors DEAD box helicase, GTPase Other 3 5 exonucleases Rex exonucleases, rrna, snorna, trna processing RNase D homolog mtexo 3 5 mitochondrial degradosome RNA degradation in yeast Suv3/ Dss1 helicase/ 3-5 exonuclease DExH box/ RNase II homolog Deadenylation Ccr4/NOT major deadenylase complex (Ccr, Caf, Pop, Not proteins) Ccr4- Mg 2+ dependent endonuclease Pop2 deadenylation regulator, deadenylase activity RNase D homolog Pan2p/Pan3 additional deadenylases (polia tail length) RNase D homolog, poly(a) specific nuclease PARN mammalian deadenylase RNase D homolog, poly(a) specific nuclease Endonucleases RNase III -Rnt1 pre-rrna, sn/snorna processing, mrna degradation dsrna specific -Dicer, Drosha sirna/mirna biogenesis, functions in RNAi PAZ, RNA BD, RNase III domains Ago2 Slicer mrna cleavage in RNAi Utp24 early 35S pre-rrna processing PIN domain Nob1 18S pre-rrna processing PIN domain SMG6 mrna cleavage in NMD PIN domain RNase P 5 trna end processing RNP complex RNase MRP pre-rrna processing RNP complex, similar to RNase P RNase L rrna degradation in apoptosis oligo 2-5A dependent (ppp(a2 p) n A) ELAC2/Trz1 3 trna endonuclease PDE motif and Zn 2+ - binding motif
11 Eukaryotic auxiliary decay factors Protein Function / Characteristics 5 3 decay: decapping Dcp1/Dcp2 Dcp2- pyrophosphatase catalytic activity, Nudix domain, Dcp1- protein binding Hedls/Ge-1/Edc4 decapping cofactor, WD40 domain Edc1,2,3 decapping enhancers, stimulate cap binding/catalysis, Edc1-2 (yeast), Edc3 (all eykaryotes) Dhh1 DexD/H ATPase, decapping activator by translation repression Lsm1-7 decapping activator, heptameric complex, binds mrna 3 end-u rich tracts Pat1 decapping activator by translation repression TRAMP complex: nuclear RNA surveillance, polyadenylation-dependent degradation Trf4/Trf5 nuclear alternative poly(a) polymerases Mtr4 DEAD box helicase Air1/Air2 RNA binding proteins, also nuclear exosome cofactor Nrd1-Nab3-Sen1 complex: PolII termination of small RNAs, TRAMP-depdendent degradation Nrd1 Pol II C-terminal domain (CTD) binding, RNA binding Nab3 RNA binding Sen1 RNA helicase
12 RNases Stoecklin and M. hlemann, BBA, 2013
13 EXOSOME: 3 5 decay machinery Kilchert et al, Nat Rev Mol Cell biol, 2016 Dis3 3 5 exo/endo nuclease complex; 10 core components (RNA BP) catalytically active exo hydrolytic Dis3/Rrp44 (RNase II) PIN domain with endo activity nuclear cofactors- RNA BP Rrp47, nuclease Rrp6 (RNase D), RNA helicase Mtr4 cytoplasmic cofactors- Ski2-3-8 complex (RNA helicase Ski2), GTPase Ski7 subtrates- processing and/or degradation of almost all RNAs Dziembowski et al, Mol.Cell, 2008; Nature, 2008
14 EXOSOME: 3 5 decay machinery: FUNCTIONS NUCLEAR: Rrp6 and core components have partly separate functions 3 end processing of 5.8S rrna, sn/snornas, trnas, SRP RNA degradation of pre-mrnas, trnas, sn/snornas degradation of other ncrnas: CUTs, PROMPTS CYTOPLASMIC: generic mrna decay specialised mrna decay pathways: NMD, NSD, NO-GO decay, AREdependent decay
15 XRN family: 5 3 processive exonucleases Kastenmayer and Green, 2000, PNAS Crystal structure of S. pombe Rat1/Rai1 complex NUCLEAR Rat1/XRN2 with Rai1 activator (5 -ppp pyrophosphohydrolase and phoshodiesterase-decapping nuclease) 5 end processing of 5.8S and 25S rrnas, snornas degradation of pre-mrnas, trnas, sn/snornas degradation of some ncrnas: CUTs transcription termination of Pol I and II (torpedo mechanism) Xiang et al, 2009, Nature CYTOPLASMIC XRN1 generic mrna decay specialised mrna decay pathways: NMD, NSD, NO-GO decay, ARE-dependent decay degradation of mirna-dependent mrna cleavage products (in plants) degradation of some ncrnas: CUTs, SUTs, XUTs
16 She et al. Nat.Struct. Mol. Biol, 2004 Dcp2 DCP- DECAPPING ENZYMES Dcp1 Dcp1/Dcp2 complex participates in mrna 5 decay catalyses the reaction m 7 GpppX-mRNA -> m 7 GDP + 5 p-mrna Dcp2 is the catalytic subunit (pyrophosphatase Nudix domain) Dcp1 is required for activity in vivo, interacts with other proteins Dcp1/Dcp2p is regulated by Pab1 and activating factors (yeast Lsm1-7, Dhh1, Pat1, Edc1-3, Upf1-3) Wang et al. PNAS, 2002 Nudt proteins (22), Nudt16, Nudt3 with in vivo decapping activity in mammalian cells DcpS DcpS: HIT pyrophosphatase ( histidine triad on the C-terminus) catalyses the cleavage of m 7 GDP -> m 7 GMP + Pi remaining after decapping during mrna 5 decay cooperates with the exosome during mrna 3 decay (m 7 GpppX-oligoRNA -> m 7 GMP+ pp-oligorna) Gu et al., M.Cell, 2004 functions as an asymmetric dimer
17 LSM PROTEINS Sm motif Involved in pre-mrna splicing associates with U6 snrna Achsel et al, EMBO J, 2001 required for U6 RNA accumulation and U6 snrnp biogenesis nuclear interacts with the U4/U6.U5 tri-snrnp cytoplasmic Functions in mrna decapping and decay activator of decapping interacts with components of the mrna decapping and degradation machinery (XRN, DCP, Pat1)
18 EUKARYOTIC mrna CBP80 NUCLEUS SPLICEOSOME 5 UTR INTRON 3 UTR m 7 GpppG AUG UAA 5 ss 3 ss CBP20 PABP2 AAAAAAAAAAAAA nts CYTOPLASM eif3 EJC PABP1 5 UTR 3 UTR m 7 GpppG eif4e eif4g AUG ORF UAA AAAAAAAAAAAAA nts m 7 Gppp eif4e eif4g AUG A A Pab1p A A A A A UAA mrna t 1/2 = few minutes to 2 hours (yeast) to >90 hours (mammals) UTR- UnTranslated Region
19 mrna DEGRADATION in the CYTOPLASM DEADENYLATION RELEASE OF RIBOSOMES the RELEASE OF TRANSLATION FACTORS t he RECRUITMENT OF DECAY FACTORS (decapping) P-BODY ASSEMBLY RNA DEGRADATION or storage? Coller and Parker, Annu. Rev. Biochem., 2004
20 NORMAL mrna DECAY IN THE CYTOPLASM eif4g eif4e m 7 Gppp Pab1p AAAAA..AAAAA Pat1p
21 NORMAL mrna DECAY IN THE CYTOPLASM dissociation m 7 Gppp Pat1p AAAAA..AAAAA Ccr4/Pop2/Not Pan2/Pan3 deadenylacja deadenylation
22 NORMAL mrna DECAY IN THE CYTOPLASM Recruitment of Lsm1-7 Lsm/Dcp m 7 Gppp AAAA AAA Pat1p
23 NORMAL mrna DECAY IN THE CYTOPLASM Xrn1p 5-3 degradation decapping Lsm Pat1p Ski2p Egzosom C AAAA AAA 3-5 degradation normal mrna decay involves deadenylation LSM/Pat1 binds and protects deadenylated mrna 3 ends against 3-5 degradation and recruite Dcp complex to activate 5-3 decay depending on organism different pathway (5-3 or 3-5 ) dominates
24 TAIL-seq: poly(a) tail analysis newly transcribed mrnas long poly(a) tails > 200nt TAIL- In strong translation mrnas PABP-eIF4G interaction stabilizes PABP binding to poly(a) allowing for poly(a) pruning to a defined length; weak translation mrnas are not protected by translation, poly(a) tails are shortened by deadenylases recruited to PABC domain, which triggers decapping and 5-3 decay
25 mrna DEGRADATION in the CYTOPLASM mrna DEGRADATION in P-BODIES mrna DEGRADATION on POLYSOMES Balagopal and Parker, Cur.Op.Cel.Biol., 2009
26 P- bodies: P bodies- processing bodies (decay bodies, DCP bodies, GW bodies) cytoplasmic dynamic structures of mrna and decay factors storage (LSM, DCP, XRN, GW182) sites of mrna degradation hlsm1-gfp -hdcp1 overlay SPECIES: Yeast Human cells Drosophila C. elegans CONTENT: Dcp1/2 Lsm Edc1/2/3 Dhh1, Pat1 general Xrn1 sirna SMG5-7 GW182 AIN-1, ALG-1 human C. elegans mirisc Cougot et al, JCB, 2004 mrna decay factors co-localize and polya + RNA accumulates in P bodies degradation bodies differ from stress granules
27 Mutations in mrna decay affect size and number of P-bodies Assembly of P bodies Stress increases P-bodies (aging, glucose deprivation, osmotic stress) Dcp2-GFP Dcp2-GFP -GLU Inhibition of translation activates P-bodies Dcp2-GFP prt1ts yeast prt1ts 23C 37C 15 min 1M KCl Sheth and Parker, 2003, Science; Brengues et al, 2005, Science Franks and Lykke-Andersen, 2008, Mol.Cell
28 P-bodies: mrna decay or storage? Purified PB contain mrna regulons: translationally repressed mrnas with their regulatory proteins mrnas with low protein yield are targeted to P-bodies mrnas in PB are translationally repressed but not decayed Hubstenberger et al, Mol Cell, 2017
29 mrna STABILITY Elements in cis: Chen and Shyu, TiBS 2016
30 LINK: mrna DECAY and TRANSCRIPTION
31 Coupling between transcription and mrna decay Haimovich et al, BBA 2013
32 Coupling between transcription and mrna decay Dahan and Choder, BBA 2013 Transcriptional machinery regulates mrna translation and decay in the cytoplasm - PolII and promoters regulate cytoplasmic post-transcriptional stages - Rpb4/7 subunits of PolII regulates trx initiation, elongation and polyadenylation by binding to the emerging transcript and remaining associated throughout its lifecycle: (i) mrna export; (ii) translation initiation via interaction with eif3; (iii) deadenylation and decay by Xrn1 and exosome via interaction with Pat1/Lsm1-7 complex
33 Coupling between transcription and mrna decay Haimovich et al, BBA 2013
34 mrna DECAY and TRANSCRIPTION mrna homeostasis Rpb4/7 Rpb4/7 Rpb4/7 Lsm1-7 Pat1 Perez-Ortin et al, 2013, JMB
35 mrna SYNTHESIS and DECAY Kervestin and Jacobson, Nat Rev Mol Cel Biol,2012
36 RNA DECAY PART II - SPECIFIC PATHWAYS
37 RNA SURVEILLANCE = RNA QUALITY CONTROL MECHANISMS NMD- (nonsense mediated decay) - degradation of mrnas with premature stop codons (PTC) NSD- (non-stop decay) - degradation of mrnas with no stop codons NO-GO decay- degradation of mrnas stalled in translation elongation AMD - ARE mediated decay- rapid degradation of mrnas with specific instability elements (e.g. AU-rich) NRD- Non-functional rrna decay nuclear RNA degradation (mrna, pre-mrna, rrna, trna, ncrnas) - degradation of RNA species that were not properly processed i.e. spliced, end-matured, modified or of unstable species (CUTs)
38 THE IMPORTANCE OF RNA QUALITY CONTROL these mechanisms control the synthesis, integrity and lifespan of all cellular RNA molecules to assure optimal functioning of the cell deficient QC and mutations in QC components lead to severe defects and diseases - several genetic disorders (30%!- e.g. β-thalasemia, ostegenesis imperfecta, Marfan syndrome, Stickler s syndrome, neurologic syndromes) result from inefficient NMD and other QC mechanisms due to frameshift mutations and premature translation termination - mutations in RNA enzymes (exosome, decapping, deadenylases) confer autoimmune diseases in humans and developmental defects in plants, worms and flies
39 NMD - degradation of mrnas containing premature STOP codons (PTC) - prevents expression of truncated, possibly harmful, proteins - 33% of yeast intron-containing mrnas undergo NMD - 30% of alternatively spliced human mrnas generate NMD substrates 40S Pab1 Translation termination problem: - premature termination or - ribosome stalling on PTC mrna degradation Amrani et al. Nat.Rev.Cel.Biol., 2006
40 NMD FACTORS Stalder and Muhlemann, TiCB, 2008 Llorka. Cur. Op. Chem. Biol SMG7 Behm-Ansmant et al., FEBS Lett, 2007
41 metazoan EXON JUNCTION COMPLEX (EJC) core eif-4aiii - DEAD box RNA helicase MLN51 - stimulates eif-4aiii Magoh/Y14 - heterodimer binds to eif-4aiii Associated factors UAP56, SRm160, Acinus, SAP18, Pinin, RNPS1 - splicing REF, TAP, p15 - export Upf2, Upf3 - NMD EJC contains proteins linked to different functions
42 MECHANISM OF NMD 1. Recognition of premature stop codon during translation EJC PTC splicing-related mechanism - EJC deposited as a mark of splicing - Upf3 is bound to mrna via EJC - mrna is exported and Upf2 joins Upf3 translation termination and unified 3 UTR mechanism: ribosome not interacting with 3 UTR factors is arrested on the PTC 2. Assembly of the active NMD complex and repression of translation active NMD complex Upf1-3 + SMG proteins EJC downstream of PTC is not removed by the advancing ribosome SURF complex, Upf1.SMG1 and erf1-2, is recruited by the stalled ribosome Upf1 is phosphorylated by SMG1 erf1-erf2 are released mrna is directed for degradation Singh and Lykke-Andersen, TiBS 2003; Isken and Maquat, Gene Dev. 2007
43 MECHANISMS OF NMD EJC-dependent Alternative long 3 UTR Kervestin Jacobson, Nat Rev Mol Cel Biol,2012
44 MECHANISM OF NMD 3. mrna degradation DCP1/2 XRN1 NMD 5 3 deadenylation-independent decapping exonucleolytic 5 3 degradation (XRN1) NMD endonucleolytic cleavage (Drosophila melanogaster, humans) SMG6 decapping is triggered by SMG7 recruitment or dephosphorylation of Upf1 NMD 3 5 egzosom C recruitment of the Exosome by Upf1 exonucleolytic 3 5 degradation Parker and Song, Nat. Struct. Mol. Biol. 2004; Isken and Maquat, Gene Dev. 2007
45 NGD and NSD NGD (non-go decay) - degradation of mrnas stalled on ribosomes NSD- (non-stop decay) - degradation of mrnas with no stop codons NGD strong RNA structure NSD Dom34/Hbs1 stalled translation endonucleolytic cleavage (?) ribosome stalls on poly(a) tail Ski7 recruits the exosome alternative pathway ExosomeC Xrn1 Exosome C Dom34 has RNA-binding Sm fold Hbs1- GTPase binding activity Xrn1 Garneau et al, Nat. Rev. Mol. Cel. Biol. 2007
46 NGD and NSD NGD (non-go decay) - degradation of mrnas stalled on ribosomes NSD- (non-stop decay) - degradation of mrnas with no stop codons Tsuboi et al, Mol. Cell 2012 Dom34:Hbs1 stimulates degradation of the 5 -NGD intermediate and nonstop mrna by dissociating the ribosome that is stalled at the 3 end of the mrna
47 Inada, TiBS 2016 NMD, NGD and NSD
48 AMD ARE - mediated decay AMD degradation of mrnas containing AU-rich instability elements present in mrna 3 UTR ARE-binding proteins (AUBP) TTP BRF1 HuR AUF1 KSRP mrna instability elements: - U rich - AU rich AUUUA, UUAUUUA(U/A) n - oligo(u) (or CUCU) tails - DST (plants) GGAnnAUAGAUUnnnCAUUnnGUAU Exosome (RRP45, RRP41, RRP43) is recruited by ARE-binding proteins AUBP (AUF1) Exosomal subunits interact directly with ARE sequences PARN and CCR4, deadenylases and XRN1, DCP1/2 interact with AUBP (TTP, KSRP, BRF1) Roretz and Galouzi, J. Cell. Biol. 2008
49 AMD - connections with RNAi? The RNAi connection? TTP binds Ago2 Georg Stoecklin s website Ago1/Ago2 and Dicer are required for the rapid decay of some ARE-mRNAs Cooperation with the RNA-induced silencing complex (RISC) may contribute to controlling the translation and decay rate of ARE-mRNAs
50 RNA REPAIR Simms and Zaher, CellMolLifeSci 2016
51 STAUFEN-mediated DECAY (SMD) Isken and Maquat, Nat Rev Genet 2008 STAU1, a dsrna binding protein, recruits UPF1 to target mrna 3'UTRs to elicit SMD in a translation-dependent fashion. SMD targets contain a STAU1 binding site (SBS) within their 3' UTR. They include newly synthesized CBC-bound mrnas and steady-state eif4e-bound mrnas. Some mrnas are targeted to SMD by lncrnas (Alu elements) which form SBS with SMD substrates. Gong and Maquat, Nature 2011
52 OTHER MECHANISMS Endonuclease mediated decay RNase MPR, RNase III (Rnt1) IRE1, PMR1 Exosome C Xrn1 PMR1 - degradation of translationally active mrnas on polysomes IRE1 - degradation of mrnas in endoplasmic reticulum during Unfoded Protein Response stress MRP (RNP responsible for pre-rrna processing in the nucleolus) - cleaves CLB2 mrna within its 5 UTR in yeast Rnt1 (RNAse III endonuclease, involved in pre-rrna and pre-sn/snorna processing) - cleaves stem-loops structures in some ribosomal protein mrnas Garneau et al, Nat. Rev. Mol. Cel. Biol. 2007
53 Deadenylation-independent decay (RPS28B and EDC1 mrnas) OTHER MECHANISMS Dcp2 RPS28B Edc3 Rps28B Dcp1 Xrn1 Autoregulatory mechanism: Rps28B protein binds to a stem-loop structure within the 3 UTR of its own mrna recruits enhancer of decapping Edc3 stimulates decapping by Dcp1/2 Garneau et al, Nat. Rev. Mol. Cel. Biol. 2007
54 mrna DECAY in the NUCLEUS nuclear retention of intron-containing pre-mrnas Processing Quality Control Export RES complex (REtention and Splicing): Snu17/Bud13/Pml1 Casolari and Silver, TiCB., 2004
55 mrna DECAY in the NUCLEUS pre-mrna with unspliced introns AAAA AAA m 7 Rat1p m 7 Gppp Gppp 7 Gppp Exosome N AAAAAAA AAAAAA Lsm2-8 Mtr4p
56 mrna DECAY in the NUCLEUS NUCLEUS mrna arrested in the nucleus Dcp1/2p Rat1p m 7 G Lsm2-8 mrna export blocked AAAA AAA AAAAAA Rrp6 m 7 G AAAAAA CYTOPLASM
57 TRAMP - EXOSOME COFACTOR (yeast) TRAMP = Trf4/5 + Air1/2 + Mtr4 polyadenylation complex poly(a) polymerases RNA binding proteins RNA DEVH helicase YEAST some RNAs degraded by Rat1/Xrn1 Polyadenylation-mediated nuclear discard pathway for defective RNAs hypomodified trnas, pre-trnas ncrnas: sn/snornas, rrnas, some mrnas CUTs (Cryptic Unstable Transcripts) Interacts with - exosome via Mtr4 - Nrd1/Nab3 complex LaCava et al., Cell, 2005; Vanacova et al., PLoS Biol. 2005; Wyers et al., Cell, 2005; Lubas et al. Mol. Cell, 2011
58 TRAMP + EXOSOME NUCLEAR RNA SURVEILLANCE TRAMP interacts with the Exosome via Mtr4 - role in degradation interacts with Nrd1/Nab3 complex - role in ncrna Pol II termination role in transcription silencing in S. cerevisiae and S. pombe (Cid14) Vanacova and Stefl, Embo Rep, 2007
59 TRAMP/EXOSOME Houseley et al., Nat. Rev. Mol. Cel. Biol., 2006 Tuttuci and Stutz., Nat. Rev. Mol. Cel. Biol., 2011
60 NEXT and PAXT - EXOSOME COFACTORS (humans) Nuclear Exosome Targetting NEXT HUMAN ZCCHC8 Zn-knucle RMB7 RNA binding ZFC3H1 (Zn-knuckle protein) links MTR4 with PABPN1 in PAXT ZFC3H1/PABPN1 and RBM7/ZCCHC8 (NEXT) interact with MTR4 in a mutually exclusive manner PAXT and NEXT direct distinct RNA species for nuclear exosome degradation PAXT targets tend to be longer and more extensively polyadenylated than NEXT targets Lubas et al. Mol. Cell, 2011; Meola et al.,. Mol. Cell, 2016
61 Rat1 - NUCLEAR RNA SURVEILLANCE 5-3 Rat1/Xrn2 decay of transcripts with aberrant cap structure degradation of prematurely terminated nascent transcripts degradation of readthrough transcripts Tutucci and Stutz., Nat. Rev. Mol. Cel. Biol., 2011
62 CAP NUCLEAR RNA SURVEILLANCE 5-3 S. cerevisiae Rai1 Rat1 activator 5 -ppp pyrophosphohydrolase and phoshodiesterase-decapping nuclease (unmethylated cap-specific)
63 CAP NUCLEAR RNA SURVEILLANCE 5-3 S. cerevisiae Dxo1: -phoshodiesterase -decapping nuclease exonuclease - no pyrophosphohydrolase Human DXO - 5 ppp pyrophosphohydrolase - phoshodiesterase -decapping nuclease exonuclease
64 trna SURVEILLANCE RAPID trna DECAY occurs for precursors and mature trnas with mutations which destabilize tertiary structure (modifications) - in the nucleus (polyadenylation via TRAMP and degradation by the exosome or degradation by Rat1) - in the cytoplasm (degradation by Xrn1) Phizicky and Hopper, GeneGev.,2010
65 trna SURVEILLANCE mature trna (hypo-modified) pre-trna (hypo-modified)
66 Interplay between trna synthesis and degradation trna primary transcript synthesized by RNA Pol III, regulated by Maf1 Initial processing in the nucleus: 5 leader and 3 trailer removed pre-trna exported to the cytoplasm CCA on the 3 terminus and some modifications added to pre-trna Intron spliced out on the outer surface of the mitochondrial membrane trna charged by trna synthetase, bound by elongation factor (eef1a) and delivered to ribosomes for translation trna turnover: (i) in the nucleus pre-trnas degraded by the exosome or by Rat1 in Rapid trna Decay (RTD) (ii) in the cytoplasm mature trnas degraded by Xrn1-mediated RTD. Mature trna cleaved into trna halves under stress Wichtowska, Turowski, Boguta, WIREsRNA, 2013
67 pre-trnas are DEGRADED by the EXOSOME Dis3 endo Dis3 exo Important contribution of the endo Dis3 activity to the degradation of structured substrates
68 rrna SURVEILLANCE Nucleus: pre-rrnas Nrd1/Nab3 (via pre-mature termination) Lafontaine, TiBS.,2010
69 NRD- nonfunctional rrna decay Cytoplasm: mature ribosomes rrna SURVEILLANCE mutations in peptidyl transferase centre mutations in decoding site Mms1, Rtt101- subunits of E3 ubiquitin ligase complex Dom34::Hbs1 factors involved in NGD and NSD Lafontaine, TiBS.,2010
70 RNA SURVEILLANCE pre-trnas cytoplasm nucleus nucleus NRD Nonfunctional rrna Decay nucleus cytoplasm nucleus
71 RNA SURVEILLANCE Vohhodina et al., WIREsRNA 16
72 RNA SURVEILLANCE Stoecklin and Mu hlemann, BBA, 2013
73 OLIGO-URIDYLATION PUP Poly(U) Polymerases TUTase Terminal Uridylyl Transferase 1. Histone mrna degradation (metazoans) 3 oligouridylation 3 hexo Eri-1 Lsm1-7 3 hexo Eri-1 3 hexo Eri-1 Mullen and Marzluff, Genes Dev., 2008
74 OLIGO-URIDYLATION PUP Poly(U) Polymerases TUTase Terminal Uridylyl Transferase 1. Histone mrna degradation (metazoans) 3 oligouridylation 3 hexo Eri-1 3 hexo Eri-1 3 hexo Eri-1 Mullen and Marzluff, Genes Dev., 2008
75 1. Histone mrna degradation (metazoans) histone mrna synthesis histone mrna decay Scheer et al, TiG., 2016
76 2. mirna degradation precursors C. elegans mammals OLIGO-URIDYLATION DIS3L2 mature 3. mrna degradation? (plants) Arabidopsis Chlamydomonas Lsm1-7 Krol et al., Nat Rev Genet, 2010; Kim et al., Cell, 2010
77 hdis3l2 - EXOSOME independent DECAY DIS3L2 pre-mirna degradation Krol et al., Nat Rev Genet, cytoplasmic mrna decay 3 degradation of aberrant, structured ncrnas: trna, sn/snorna, rrna, lncrna, Y RNA, vault RNA, surveillance of 3 snrna processing Pirouz et al., Cell Rep, 2016; Ustianenko et al., EMBO, 2016
78 STRESS-INDUCED ENZYMATIC trna and rrna DEGRADATION (S. cerevisiae, D. melanogaster, A. thaliana, A. nidulans, human cell lines) Thompson and Parker, Cell, 2009
79 RNA DECAY TAKE-HOME MESSAGE NORMAL (usually in the cytoplasm) SPECIALIZED RNA SURVEILLANCE targeting aberrant or unstable transcripts for the discard pathway (NMD, NSD, NGD, ARE, NRD) 1. deadenylation decapping exonucleolytic degradation 5-3 by Xrn1/Rat1 or 3-5 by exosome 2. by endo- cleavage (mirna-dependent, RNAse III/Rnt1, MRP, SMG6, Dom34) followed by exo- digestion (Xrn1/Rat1, exosome) NUCLEAR RNA SURVEILLANCE - polyadenylation-mediated by TRAMP (Trf4/5) followed by degradation by the exosome or Rat1 - related to pre-mature termination or aberrant cap structure by Rat1 MEDIATED BY OLIGOURIDYLATION Processing and degradation are often carried out by the same machineries
80 TAKE-HOME MESSAGE RNA DECAY normal (usually in the cytoplasm) specialized, RNA surveillance: targeting aberrant, unstable transcripts for the discard pathway (NMD, NSD, NGD, ARE, NRD) 1. deadenylation decapping exonucleolytic degradation 5-3 by Xrn1/Rat1 or 3-5 by exosome 2. by endo- cleavage (mirna-dependent, RNAse III/Rnt1, MRP, SMG6, Dom34) followed by exo- digestion (Xrn1/Rat1, exosome) nuclear RNA surveillance: polyadenylation by TRAMP (Trf4/5) followed by degradation by the exosome, Xrn1 or Rat1 post-transcriptional gene silencing mrna cleavage translation inhibition/mrna decay Processing and degradation are often carried out by the same machineries
Pol II - mrna synthesis
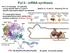 Pol II - mrna synthesis Pol II, S. cerevisiae (12 subunits) - core by specific Rpb1-3, 9 and 11 - Rpb5-6, 8, 10 and 12 - shared by Pol I-III - specific subcomplex Rpb4/7 not essential - TF: transcription
Pol II - mrna synthesis Pol II, S. cerevisiae (12 subunits) - core by specific Rpb1-3, 9 and 11 - Rpb5-6, 8, 10 and 12 - shared by Pol I-III - specific subcomplex Rpb4/7 not essential - TF: transcription
Novel RNAs along the Pathway of Gene Expression. (or, The Expanding Universe of Small RNAs)
 Novel RNAs along the Pathway of Gene Expression (or, The Expanding Universe of Small RNAs) Central Dogma DNA RNA Protein replication transcription translation Central Dogma DNA RNA Spliced RNA Protein
Novel RNAs along the Pathway of Gene Expression (or, The Expanding Universe of Small RNAs) Central Dogma DNA RNA Protein replication transcription translation Central Dogma DNA RNA Spliced RNA Protein
RNA-quality control by the exosome
 RNA-quality control by the exosome Jonathan Houseley, John LaCava and David Tollervey Abstract The exosome complex of 3 5 exonucleases is an important component of the RNA-processing machinery in eukaryotes.
RNA-quality control by the exosome Jonathan Houseley, John LaCava and David Tollervey Abstract The exosome complex of 3 5 exonucleases is an important component of the RNA-processing machinery in eukaryotes.
Chapter 10 - Post-transcriptional Gene Control
 Chapter 10 - Post-transcriptional Gene Control Chapter 10 - Post-transcriptional Gene Control 10.1 Processing of Eukaryotic Pre-mRNA 10.2 Regulation of Pre-mRNA Processing 10.3 Transport of mrna Across
Chapter 10 - Post-transcriptional Gene Control Chapter 10 - Post-transcriptional Gene Control 10.1 Processing of Eukaryotic Pre-mRNA 10.2 Regulation of Pre-mRNA Processing 10.3 Transport of mrna Across
Messenger RNA (mrna) is never alone
 Messenger RNA (mrna) is never alone mrna is coated and compacted by RNA-binding proteins (RBPs), forming large messenger ribonucleoprotein particles (mrnps). RBPs assemble on nascent and mature mrnas.
Messenger RNA (mrna) is never alone mrna is coated and compacted by RNA-binding proteins (RBPs), forming large messenger ribonucleoprotein particles (mrnps). RBPs assemble on nascent and mature mrnas.
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004
 Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
The meaning of nonsense
 Opinion The meaning of nonsense Lukas Stalder and Oliver Mühlemann Institute of Cell Biology, University of Berne, Baltzerstrabe 4, 3012 Berne, Switzerland To ensure the accuracy of gene expression, eukaryotes
Opinion The meaning of nonsense Lukas Stalder and Oliver Mühlemann Institute of Cell Biology, University of Berne, Baltzerstrabe 4, 3012 Berne, Switzerland To ensure the accuracy of gene expression, eukaryotes
Eukaryotic mrna is covalently processed in three ways prior to export from the nucleus:
 RNA Processing Eukaryotic mrna is covalently processed in three ways prior to export from the nucleus: Transcripts are capped at their 5 end with a methylated guanosine nucleotide. Introns are removed
RNA Processing Eukaryotic mrna is covalently processed in three ways prior to export from the nucleus: Transcripts are capped at their 5 end with a methylated guanosine nucleotide. Introns are removed
The Nonsense-Mediated Decay RNA Surveillance Pathway
 Annu. Rev. Biochem. 2007. 76:51 74 First published online as a Review in Advance on March 12, 2007 The Annual Review of Biochemistry is online at biochem.annualreviews.org This article s doi: 10.1146/annurev.biochem.76.050106.093909
Annu. Rev. Biochem. 2007. 76:51 74 First published online as a Review in Advance on March 12, 2007 The Annual Review of Biochemistry is online at biochem.annualreviews.org This article s doi: 10.1146/annurev.biochem.76.050106.093909
Assembly of mrnp Complexes During Stress and Nonsense-Mediated mrna Decay Quality Control in Saccharomyces cerevisiae
 Assembly of mrnp Complexes During Stress and Nonsense-Mediated mrna Decay Quality Control in Saccharomyces cerevisiae Item Type text; Electronic Dissertation Authors Swisher, Kylie Publisher The University
Assembly of mrnp Complexes During Stress and Nonsense-Mediated mrna Decay Quality Control in Saccharomyces cerevisiae Item Type text; Electronic Dissertation Authors Swisher, Kylie Publisher The University
Genetics. Instructor: Dr. Jihad Abdallah Transcription of DNA
 Genetics Instructor: Dr. Jihad Abdallah Transcription of DNA 1 3.4 A 2 Expression of Genetic information DNA Double stranded In the nucleus Transcription mrna Single stranded Translation In the cytoplasm
Genetics Instructor: Dr. Jihad Abdallah Transcription of DNA 1 3.4 A 2 Expression of Genetic information DNA Double stranded In the nucleus Transcription mrna Single stranded Translation In the cytoplasm
RNA decapping inside and outside of processing bodies Christy Fillman and Jens Lykke-Andersen
 RNA decapping inside and outside of processing bodies Christy Fillman and Jens Lykke-Andersen Decapping is a central step in eukaryotic mrna turnover. Recent studies have identified several factors involved
RNA decapping inside and outside of processing bodies Christy Fillman and Jens Lykke-Andersen Decapping is a central step in eukaryotic mrna turnover. Recent studies have identified several factors involved
Molecular Cell Biology - Problem Drill 10: Gene Expression in Eukaryotes
 Molecular Cell Biology - Problem Drill 10: Gene Expression in Eukaryotes Question No. 1 of 10 1. Which of the following statements about gene expression control in eukaryotes is correct? Question #1 (A)
Molecular Cell Biology - Problem Drill 10: Gene Expression in Eukaryotes Question No. 1 of 10 1. Which of the following statements about gene expression control in eukaryotes is correct? Question #1 (A)
Interrelations between translation and general mrna degradation in yeast Susanne Huch and Tracy Nissan
 Interrelations between translation and general mrna degradation in yeast Susanne Huch and Tracy Nissan Messenger RNA (mrna) degradation is an important element of gene expression that can be modulated
Interrelations between translation and general mrna degradation in yeast Susanne Huch and Tracy Nissan Messenger RNA (mrna) degradation is an important element of gene expression that can be modulated
Bi 8 Lecture 17. interference. Ellen Rothenberg 1 March 2016
 Bi 8 Lecture 17 REGulation by RNA interference Ellen Rothenberg 1 March 2016 Protein is not the only regulatory molecule affecting gene expression: RNA itself can be negative regulator RNA does not need
Bi 8 Lecture 17 REGulation by RNA interference Ellen Rothenberg 1 March 2016 Protein is not the only regulatory molecule affecting gene expression: RNA itself can be negative regulator RNA does not need
Processing of RNA II Biochemistry 302. February 13, 2006
 Processing of RNA II Biochemistry 302 February 13, 2006 Precursor mrna: introns and exons Intron: Transcribed RNA sequence removed from precursor RNA during the process of maturation (for class II genes:
Processing of RNA II Biochemistry 302 February 13, 2006 Precursor mrna: introns and exons Intron: Transcribed RNA sequence removed from precursor RNA during the process of maturation (for class II genes:
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression. Chromatin Array of nuc
 Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER
 REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER RNA Splicing Lecture 3, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice
REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER RNA Splicing Lecture 3, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice
Processing of RNA II Biochemistry 302. February 18, 2004 Bob Kelm
 Processing of RNA II Biochemistry 302 February 18, 2004 Bob Kelm What s an intron? Transcribed sequence removed during the process of mrna maturation Discovered by P. Sharp and R. Roberts in late 1970s
Processing of RNA II Biochemistry 302 February 18, 2004 Bob Kelm What s an intron? Transcribed sequence removed during the process of mrna maturation Discovered by P. Sharp and R. Roberts in late 1970s
RNA Processing in Eukaryotes *
 OpenStax-CNX module: m44532 1 RNA Processing in Eukaryotes * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you
OpenStax-CNX module: m44532 1 RNA Processing in Eukaryotes * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you
Processing of RNA II Biochemistry 302. February 14, 2005 Bob Kelm
 Processing of RNA II Biochemistry 302 February 14, 2005 Bob Kelm What s an intron? Transcribed sequence removed during the process of mrna maturation (non proteincoding sequence) Discovered by P. Sharp
Processing of RNA II Biochemistry 302 February 14, 2005 Bob Kelm What s an intron? Transcribed sequence removed during the process of mrna maturation (non proteincoding sequence) Discovered by P. Sharp
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins.
 Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
1. Investigate the structure of the trna Synthase in complex with a trna molecule. (pdb ID 1ASY).
 Problem Set 11 (Due Nov 25 th ) 1. Investigate the structure of the trna Synthase in complex with a trna molecule. (pdb ID 1ASY). a. Why don t trna molecules contain a 5 triphosphate like other RNA molecules
Problem Set 11 (Due Nov 25 th ) 1. Investigate the structure of the trna Synthase in complex with a trna molecule. (pdb ID 1ASY). a. Why don t trna molecules contain a 5 triphosphate like other RNA molecules
RECAP (1)! In eukaryotes, large primary transcripts are processed to smaller, mature mrnas.! What was first evidence for this precursorproduct
 RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
L I F E S C I E N C E S
 1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 NOVEMBER 2, 2006
1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 NOVEMBER 2, 2006
Molecular Biology (BIOL 4320) Exam #2 April 22, 2002
 Molecular Biology (BIOL 4320) Exam #2 April 22, 2002 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Biology (BIOL 4320) Exam #2 April 22, 2002 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
The Human Nuclear Poly(A)-Binding Protein Promotes RNA Hyperadenylation and Decay
 The Human Nuclear Poly(A)-Binding Protein Promotes RNA Hyperadenylation and Decay Stefan M. Bresson, Nicholas K. Conrad* Department of Microbiology, University of Texas Southwestern Medical Center, Dallas,
The Human Nuclear Poly(A)-Binding Protein Promotes RNA Hyperadenylation and Decay Stefan M. Bresson, Nicholas K. Conrad* Department of Microbiology, University of Texas Southwestern Medical Center, Dallas,
Mechanism of splicing
 Outline of Splicing Mechanism of splicing Em visualization of precursor-spliced mrna in an R loop Kinetics of in vitro splicing Analysis of the splice lariat Lariat Branch Site Splice site sequence requirements
Outline of Splicing Mechanism of splicing Em visualization of precursor-spliced mrna in an R loop Kinetics of in vitro splicing Analysis of the splice lariat Lariat Branch Site Splice site sequence requirements
Where Splicing Joins Chromatin And Transcription. 9/11/2012 Dario Balestra
 Where Splicing Joins Chromatin And Transcription 9/11/2012 Dario Balestra Splicing process overview Splicing process overview Sequence context RNA secondary structure Tissue-specific Proteins Development
Where Splicing Joins Chromatin And Transcription 9/11/2012 Dario Balestra Splicing process overview Splicing process overview Sequence context RNA secondary structure Tissue-specific Proteins Development
V16: involvement of micrornas in GRNs
 What are micrornas? V16: involvement of micrornas in GRNs How can one identify micrornas? What is the function of micrornas? Elisa Izaurralde, MPI Tübingen Huntzinger, Izaurralde, Nat. Rev. Genet. 12,
What are micrornas? V16: involvement of micrornas in GRNs How can one identify micrornas? What is the function of micrornas? Elisa Izaurralde, MPI Tübingen Huntzinger, Izaurralde, Nat. Rev. Genet. 12,
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
RECAP (1)! In eukaryotes, large primary transcripts are processed to smaller, mature mrnas.! What was first evidence for this precursorproduct
 RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
Pre-mRNA has introns The splicing complex recognizes semiconserved sequences
 Adding a 5 cap Lecture 4 mrna splicing and protein synthesis Another day in the life of a gene. Pre-mRNA has introns The splicing complex recognizes semiconserved sequences Introns are removed by a process
Adding a 5 cap Lecture 4 mrna splicing and protein synthesis Another day in the life of a gene. Pre-mRNA has introns The splicing complex recognizes semiconserved sequences Introns are removed by a process
REGULATED AND NONCANONICAL SPLICING
 REGULATED AND NONCANONICAL SPLICING RNA Processing Lecture 3, Biological Regulatory Mechanisms, Hiten Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice site consensus sequences do have
REGULATED AND NONCANONICAL SPLICING RNA Processing Lecture 3, Biological Regulatory Mechanisms, Hiten Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice site consensus sequences do have
Interaktionen von RNAs und Proteinen
 Sonja Prohaska Computational EvoDevo Universitaet Leipzig May 5, 2017 RNA-protein binding vs. DNA-protein binding Is binding of proteins to RNA the same as binding of proteins to DNA? Will the same rules
Sonja Prohaska Computational EvoDevo Universitaet Leipzig May 5, 2017 RNA-protein binding vs. DNA-protein binding Is binding of proteins to RNA the same as binding of proteins to DNA? Will the same rules
Turnover in the Alps: an mrna perspective. Workshop on Mechanisms and Regulation of mrna Turnover
 Turnover in the Alps: an mrna perspective Workshop on Mechanisms and Regulation of mrna Turnover Sarah F. Newbury 1, Oliver Mühlemann 2 & Georg Stoecklin 3+ 1 Institute of Cell and Molecular Biosciences,
Turnover in the Alps: an mrna perspective Workshop on Mechanisms and Regulation of mrna Turnover Sarah F. Newbury 1, Oliver Mühlemann 2 & Georg Stoecklin 3+ 1 Institute of Cell and Molecular Biosciences,
RECAP FROM MONDAY AND TUESDAY LECTURES
 RECAP FROM MONDAY AND TUESDAY LECTURES Nucleo-cytoplasmic transport factors: interact with the target macromolecule (signal recognition) interact with the NPC (nucleoporin Phe - Gly repeats) NPC cytoplasmic
RECAP FROM MONDAY AND TUESDAY LECTURES Nucleo-cytoplasmic transport factors: interact with the target macromolecule (signal recognition) interact with the NPC (nucleoporin Phe - Gly repeats) NPC cytoplasmic
Objectives: Prof.Dr. H.D.El-Yassin
 Protein Synthesis and drugs that inhibit protein synthesis Objectives: 1. To understand the steps involved in the translation process that leads to protein synthesis 2. To understand and know about all
Protein Synthesis and drugs that inhibit protein synthesis Objectives: 1. To understand the steps involved in the translation process that leads to protein synthesis 2. To understand and know about all
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
MCB Chapter 11. Topic E. Splicing mechanism Nuclear Transport Alternative control modes. Reading :
 MCB Chapter 11 Topic E Splicing mechanism Nuclear Transport Alternative control modes Reading : 419-449 Topic E Michal Linial 14 Jan 2004 1 Self-splicing group I introns were the first examples of catalytic
MCB Chapter 11 Topic E Splicing mechanism Nuclear Transport Alternative control modes Reading : 419-449 Topic E Michal Linial 14 Jan 2004 1 Self-splicing group I introns were the first examples of catalytic
MicroRNA in Cancer Karen Dybkær 2013
 MicroRNA in Cancer Karen Dybkær RNA Ribonucleic acid Types -Coding: messenger RNA (mrna) coding for proteins -Non-coding regulating protein formation Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
MicroRNA in Cancer Karen Dybkær RNA Ribonucleic acid Types -Coding: messenger RNA (mrna) coding for proteins -Non-coding regulating protein formation Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
TRANSCRIPTION CAPPING
 Messenger RNA Eucaryotic gene expression transcription START site exons upstream elements promoter elements TRANSCRIPTION CAPPING introns gene m 7 G pre-mrna m 7 G SPLICING POLYADENYLATION AAAAAAAAAn
Messenger RNA Eucaryotic gene expression transcription START site exons upstream elements promoter elements TRANSCRIPTION CAPPING introns gene m 7 G pre-mrna m 7 G SPLICING POLYADENYLATION AAAAAAAAAn
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Lecture 4 Regulation of Initiation: mrna-binding
 Institut für Biochemie und Molekulare Medizin Lecture 4 Regulation of Initiation: mrna-binding Michael Altmann FS 2010 Regulation of mrna binding to ribosomes mrna structure Cap accessibility RNA secondary
Institut für Biochemie und Molekulare Medizin Lecture 4 Regulation of Initiation: mrna-binding Michael Altmann FS 2010 Regulation of mrna binding to ribosomes mrna structure Cap accessibility RNA secondary
How introns influence and enhance eukaryotic gene expression q
 Review TRENDS in Biochemical Sciences Vol.28 No.4 April 2003 215 How introns influence and enhance eukaryotic gene expression q Hervé Le Hir 1,2, Ajit Nott 1 and Melissa J. Moore 1 1 Howard Hughes Medical
Review TRENDS in Biochemical Sciences Vol.28 No.4 April 2003 215 How introns influence and enhance eukaryotic gene expression q Hervé Le Hir 1,2, Ajit Nott 1 and Melissa J. Moore 1 1 Howard Hughes Medical
1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics
 1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics to gain mechanistic insight! 4. Return to step 2, as
1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics to gain mechanistic insight! 4. Return to step 2, as
Abstract. comment reviews reports deposited research refereed research interactions information
 http://genomebiology.com/2002/3/3/reviews/1006.1 Minireview Integration of splicing, transport and translation to achieve mrna quality control by the nonsense-mediated decay pathway Thomas Schell*, Andreas
http://genomebiology.com/2002/3/3/reviews/1006.1 Minireview Integration of splicing, transport and translation to achieve mrna quality control by the nonsense-mediated decay pathway Thomas Schell*, Andreas
Computational Biology I LSM5191
 Computational Biology I LSM5191 Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression II TRANSLATION RIBOSOMES: protein synthesizing machines Translation takes place on defined
Computational Biology I LSM5191 Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression II TRANSLATION RIBOSOMES: protein synthesizing machines Translation takes place on defined
Life Sciences 1A Midterm Exam 2. November 13, 2006
 Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Canonical Poly(A) Polymerase Activity Promotes the Decay of a Wide Variety of Mammalian Nuclear RNAs
 RESEARCH ARTICLE Canonical Poly(A) Polymerase Activity Promotes the Decay of a Wide Variety of Mammalian Nuclear RNAs Stefan M. Bresson, Olga V. Hunter, Allyson C. Hunter, Nicholas K. Conrad* Department
RESEARCH ARTICLE Canonical Poly(A) Polymerase Activity Promotes the Decay of a Wide Variety of Mammalian Nuclear RNAs Stefan M. Bresson, Olga V. Hunter, Allyson C. Hunter, Nicholas K. Conrad* Department
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Messenger RNA maturation. Dr.ssa Mariangela Morlando
 Messenger RNA maturation Dr.ssa Mariangela Morlando Features of RNA molecule Synthe(sed from a DNA template (transcrip(on) Single strand molecule Contains ribose in place of deoxyribose Contains Uracil
Messenger RNA maturation Dr.ssa Mariangela Morlando Features of RNA molecule Synthe(sed from a DNA template (transcrip(on) Single strand molecule Contains ribose in place of deoxyribose Contains Uracil
mrnp quality control goes regulatory
 *Manuscript Click here to download Manuscript: RN quality control_revised0.docx Mühlemann & Jensen, TiG review, 0 0 This is the peer reviewed version of the following article: Mühlemann O, Jensen TH (0).
*Manuscript Click here to download Manuscript: RN quality control_revised0.docx Mühlemann & Jensen, TiG review, 0 0 This is the peer reviewed version of the following article: Mühlemann O, Jensen TH (0).
CELLS. Cells. Basic unit of life (except virus)
 Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Transcriptome-wide Analysis of Exosome Targets
 Article Claudia Schneider, 1,2, * Grzegorz Kudla, 1,3 Wiebke Wlotzka, 1 Alex Tuck, 1 and David Tollervey 1, * 1 Wellcome Trust Centre for Cell Biology, The University of Edinburgh, Edinburgh UK 2 Present
Article Claudia Schneider, 1,2, * Grzegorz Kudla, 1,3 Wiebke Wlotzka, 1 Alex Tuck, 1 and David Tollervey 1, * 1 Wellcome Trust Centre for Cell Biology, The University of Edinburgh, Edinburgh UK 2 Present
scientific report ARE-mRNA degradation requires the decay pathway scientificreport
 scientificreport ARE-mRNA degradation requires the 5 0 3 0 decay pathway Georg Stoecklin +,ThomasMayo&PaulAnderson Division of Rheumatology, Immunology & Allergy, Brigham & Women s Hospital, Harvard Medical
scientificreport ARE-mRNA degradation requires the 5 0 3 0 decay pathway Georg Stoecklin +,ThomasMayo&PaulAnderson Division of Rheumatology, Immunology & Allergy, Brigham & Women s Hospital, Harvard Medical
The amount of a particular mrna present at a specific time in a
 MINIREVIEW Regulation of Natural mrnas by the Nonsense-Mediated mrna Decay Pathway Megan Peccarelli, Bessie W. Kebaara Department of Biology, Baylor University, Waco, Texas, USA The nonsense-mediated mrna
MINIREVIEW Regulation of Natural mrnas by the Nonsense-Mediated mrna Decay Pathway Megan Peccarelli, Bessie W. Kebaara Department of Biology, Baylor University, Waco, Texas, USA The nonsense-mediated mrna
A garbage can for ribosomes: how eukaryotes degrade their ribosomes
 Review A garbage can for ribosomes: how eukaryotes degrade their ribosomes Denis L.J. Lafontaine 1,2 1 Fonds de la Recherche Scientifique (FRS-F.N.R.S.), Institut de Biologie et de Médecine Moléculaire
Review A garbage can for ribosomes: how eukaryotes degrade their ribosomes Denis L.J. Lafontaine 1,2 1 Fonds de la Recherche Scientifique (FRS-F.N.R.S.), Institut de Biologie et de Médecine Moléculaire
1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled
 Protein Targeting Objectives 1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled As a protein is being synthesized, decisions
Protein Targeting Objectives 1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled As a protein is being synthesized, decisions
Bio 111 Study Guide Chapter 17 From Gene to Protein
 Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Alan Weiner BIOCHEM 530 Friday, MEB 248 October 23, 2015 RNA structure, the ribosome, structure-based drug design
 Alan Weiner BIOCHEM 530 Friday, MEB 248 October 23, 2015 RNA structure, the ribosome, structure-based drug design Crick's "Central Dogma"? DNA makes RNA makes protein Crick's "Central Dogma"? DNA makes
Alan Weiner BIOCHEM 530 Friday, MEB 248 October 23, 2015 RNA structure, the ribosome, structure-based drug design Crick's "Central Dogma"? DNA makes RNA makes protein Crick's "Central Dogma"? DNA makes
Translation. Host Cell Shutoff 1) Initiation of eukaryotic translation involves many initiation factors
 Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
Protein Synthesis
 Protein Synthesis 10.6-10.16 Objectives - To explain the central dogma - To understand the steps of transcription and translation in order to explain how our genes create proteins necessary for survival.
Protein Synthesis 10.6-10.16 Objectives - To explain the central dogma - To understand the steps of transcription and translation in order to explain how our genes create proteins necessary for survival.
BIO 5099: Molecular Biology for Computer Scientists (et al)
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
TRANSCRIPTION. DNA à mrna
 TRANSCRIPTION DNA à mrna Central Dogma Animation DNA: The Secret of Life (from PBS) http://www.youtube.com/watch? v=41_ne5ms2ls&list=pl2b2bd56e908da696&index=3 Transcription http://highered.mcgraw-hill.com/sites/0072507470/student_view0/
TRANSCRIPTION DNA à mrna Central Dogma Animation DNA: The Secret of Life (from PBS) http://www.youtube.com/watch? v=41_ne5ms2ls&list=pl2b2bd56e908da696&index=3 Transcription http://highered.mcgraw-hill.com/sites/0072507470/student_view0/
Cell Quality Control. Peter Takizawa Department of Cell Biology
 Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Viral Control of SR Protein Activity
 Comprehensive Summaries of Uppsala Dissertations from the Faculty of Medicine 1078 Viral Control of SR Protein Activity BY CAMILLA ESTMER NILSSON ACTA UNIVERSITATIS UPSALIENSIS UPPSALA 2001 Dissertation
Comprehensive Summaries of Uppsala Dissertations from the Faculty of Medicine 1078 Viral Control of SR Protein Activity BY CAMILLA ESTMER NILSSON ACTA UNIVERSITATIS UPSALIENSIS UPPSALA 2001 Dissertation
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Post-transcriptional regulation of an intronic microrna
 Post-transcriptional regulation of an intronic microrna Carl Novina Dana-Farber Cancer Institute Harvard Medical School Broad Institute of Harvard and MIT Qiagen Webinar 05-17-11 Outline 1. The biology
Post-transcriptional regulation of an intronic microrna Carl Novina Dana-Farber Cancer Institute Harvard Medical School Broad Institute of Harvard and MIT Qiagen Webinar 05-17-11 Outline 1. The biology
Transcription and RNA processing
 Transcription and RNA processing Lecture 7 Biology 3310/4310 Virology Spring 2018 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at the
Transcription and RNA processing Lecture 7 Biology 3310/4310 Virology Spring 2018 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at the
Investigating the regulation of mirna biogenesis and Argonaute2 by RNA binding proteins
 Investigating the regulation of mirna biogenesis and Argonaute2 by RNA binding proteins Patrick Peter Connerty Supervisor: Gyorgy Hutvagner Thesis Submitted for the Degree of Doctor of Philosophy (Science)
Investigating the regulation of mirna biogenesis and Argonaute2 by RNA binding proteins Patrick Peter Connerty Supervisor: Gyorgy Hutvagner Thesis Submitted for the Degree of Doctor of Philosophy (Science)
TRANSLATION. Translation is a process where proteins are made by the ribosomes on the mrna strand.
 TRANSLATION Dr. Mahesha H B, Yuvaraja s College, University of Mysore, Mysuru. Translation is a process where proteins are made by the ribosomes on the mrna strand. Or The process in the ribosomes of a
TRANSLATION Dr. Mahesha H B, Yuvaraja s College, University of Mysore, Mysuru. Translation is a process where proteins are made by the ribosomes on the mrna strand. Or The process in the ribosomes of a
This thesis is based on the following papers, referred to in the text by their roman numerals:
 List of papers This thesis is based on the following papers, referred to in the text by their roman numerals: I II Margaret Rush, Xiaomin Zhao, Stefan Schwartz. (2005) A splicing enhancer in the E4 coding
List of papers This thesis is based on the following papers, referred to in the text by their roman numerals: I II Margaret Rush, Xiaomin Zhao, Stefan Schwartz. (2005) A splicing enhancer in the E4 coding
MicroRNAs, RNA Modifications, RNA Editing. Bora E. Baysal MD, PhD Oncology for Scientists Lecture Tue, Oct 17, 2017, 3:30 PM - 5:00 PM
 MicroRNAs, RNA Modifications, RNA Editing Bora E. Baysal MD, PhD Oncology for Scientists Lecture Tue, Oct 17, 2017, 3:30 PM - 5:00 PM Expanding world of RNAs mrna, messenger RNA (~20,000) trna, transfer
MicroRNAs, RNA Modifications, RNA Editing Bora E. Baysal MD, PhD Oncology for Scientists Lecture Tue, Oct 17, 2017, 3:30 PM - 5:00 PM Expanding world of RNAs mrna, messenger RNA (~20,000) trna, transfer
MicroRNA and Male Infertility: A Potential for Diagnosis
 Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Eukaryotic Gene Regulation
 Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being A Eukaryote. Eukaryotic Cells. Basic eukaryotes have:
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
RNA (Ribonucleic acid)
 RNA (Ribonucleic acid) Structure: Similar to that of DNA except: 1- it is single stranded polunucleotide chain. 2- Sugar is ribose 3- Uracil is instead of thymine There are 3 types of RNA: 1- Ribosomal
RNA (Ribonucleic acid) Structure: Similar to that of DNA except: 1- it is single stranded polunucleotide chain. 2- Sugar is ribose 3- Uracil is instead of thymine There are 3 types of RNA: 1- Ribosomal
Human Genome: Mapping, Sequencing Techniques, Diseases
 Human Genome: Mapping, Sequencing Techniques, Diseases Lecture 4 BINF 7580 Fall 2005 1 Let us review what we talked about at the previous lecture. Please,... 2 The central dogma states that the transfer
Human Genome: Mapping, Sequencing Techniques, Diseases Lecture 4 BINF 7580 Fall 2005 1 Let us review what we talked about at the previous lecture. Please,... 2 The central dogma states that the transfer
MECHANISMS OF ALTERNATIVE PRE-MESSENGER RNA SPLICING
 Annu. Rev. Biochem. 2003. 72:291 336 doi: 10.1146/annurev.biochem.72.121801.161720 Copyright 2003 by Annual Reviews. All rights reserved First published online as a Review in Advance on February 27, 2003
Annu. Rev. Biochem. 2003. 72:291 336 doi: 10.1146/annurev.biochem.72.121801.161720 Copyright 2003 by Annual Reviews. All rights reserved First published online as a Review in Advance on February 27, 2003
Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN
 Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN Institute for Brain Disorders and Neural Regeneration F.M. Kirby Program in Neural
Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN Institute for Brain Disorders and Neural Regeneration F.M. Kirby Program in Neural
Cellular MicroRNA and P Bodies Modulate Host-HIV-1 Interactions. 指導教授 : 張麗冠博士 演講者 : 黃柄翰 Date: 2009/10/19
 Cellular MicroRNA and P Bodies Modulate Host-HIV-1 Interactions 指導教授 : 張麗冠博士 演講者 : 黃柄翰 Date: 2009/10/19 1 MicroRNA biogenesis 1. Pri mirna: primary mirna 2. Drosha: RNaseIII 3. DCR 1: Dicer 1 RNaseIII
Cellular MicroRNA and P Bodies Modulate Host-HIV-1 Interactions 指導教授 : 張麗冠博士 演講者 : 黃柄翰 Date: 2009/10/19 1 MicroRNA biogenesis 1. Pri mirna: primary mirna 2. Drosha: RNaseIII 3. DCR 1: Dicer 1 RNaseIII
Explain that each trna molecule is recognised by a trna-activating enzyme that binds a specific amino acid to the trna, using ATP for energy
 7.4 - Translation 7.4.1 - Explain that each trna molecule is recognised by a trna-activating enzyme that binds a specific amino acid to the trna, using ATP for energy Each amino acid has a specific trna-activating
7.4 - Translation 7.4.1 - Explain that each trna molecule is recognised by a trna-activating enzyme that binds a specific amino acid to the trna, using ATP for energy Each amino acid has a specific trna-activating
Transcription and RNA processing
 Transcription and RNA processing Lecture 7 Biology W3310/4310 Virology Spring 2016 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at
Transcription and RNA processing Lecture 7 Biology W3310/4310 Virology Spring 2016 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at
MicroRNAs Silence Gene Expression by Repressing Protein Expression and/or by Promoting mrna Decay
 MicroRNAs Silence Gene Expression by Repressing Protein Expression and/or by Promoting mrna Decay I. BEHM-ANSMANT, J. REHWINKEL, AND E. IZAURRALDE MPI for Developmental Biology, D-72076 Tuebingen, Germany
MicroRNAs Silence Gene Expression by Repressing Protein Expression and/or by Promoting mrna Decay I. BEHM-ANSMANT, J. REHWINKEL, AND E. IZAURRALDE MPI for Developmental Biology, D-72076 Tuebingen, Germany
Pre-mRNA Processing Reaches Back to Transcription and Ahead to Translation
 Leading Edge Review Pre-mRNA Processing Reaches Back to Transcription and Ahead to Translation Melissa J. Moore 1,2, * and Nick J. Proudfoot 3, * 1 Howard Hughes Medical Institute 2 Department of Biochemistry
Leading Edge Review Pre-mRNA Processing Reaches Back to Transcription and Ahead to Translation Melissa J. Moore 1,2, * and Nick J. Proudfoot 3, * 1 Howard Hughes Medical Institute 2 Department of Biochemistry
Organization of genetic material in eukaryotes
 Organization of genetic material in eukaryotes biologiemoleculara.usmf.md pass.: bmgu e.usmf.md 1 DNA in eukaryotes Location: In nucleus In mitochondria biologiemoleculara.usmf.md e.usmf.md pass.: bmgu
Organization of genetic material in eukaryotes biologiemoleculara.usmf.md pass.: bmgu e.usmf.md 1 DNA in eukaryotes Location: In nucleus In mitochondria biologiemoleculara.usmf.md e.usmf.md pass.: bmgu
Chapter 9. Cellular Signaling
 Chapter 9 Cellular Signaling Cellular Messaging Page 215 Cells can signal to each other and interpret the signals they receive from other cells and the environment Signals are most often chemicals The
Chapter 9 Cellular Signaling Cellular Messaging Page 215 Cells can signal to each other and interpret the signals they receive from other cells and the environment Signals are most often chemicals The
Getting to the Root of mirna-mediated Gene Silencing
 Leading Edge Essay Getting to the Root of mirna-mediated Gene Silencing Ana Eulalio, 1 Eric Huntzinger, 1 and Elisa Izaurralde 1, * 1 Max-Planck-Institute for Developmental Biology, Spemannstrasse 35,
Leading Edge Essay Getting to the Root of mirna-mediated Gene Silencing Ana Eulalio, 1 Eric Huntzinger, 1 and Elisa Izaurralde 1, * 1 Max-Planck-Institute for Developmental Biology, Spemannstrasse 35,
Introduction retroposon
 17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
The influence of micrornas and poly(a) tail length on endogenous mrna protein complexes
 Rissland et al. Genome Biology (7 8: DOI.86/s359-7-33-z RESEARCH Open Access The influence of micrornas and poly(a tail length on endogenous mrna protein complexes Olivia S. Rissland,,3,4,5,6*, Alexander
Rissland et al. Genome Biology (7 8: DOI.86/s359-7-33-z RESEARCH Open Access The influence of micrornas and poly(a tail length on endogenous mrna protein complexes Olivia S. Rissland,,3,4,5,6*, Alexander
Retroviral RNA Processing and stability
 Retroviral RN Processing and stability m 7 gag pol env src Karen Beemon Johns Hopkins niversity m 7 env src m 7 src Retroviruses hijack host cell gene expression machinery to generate progeny virions Simple
Retroviral RN Processing and stability m 7 gag pol env src Karen Beemon Johns Hopkins niversity m 7 env src m 7 src Retroviruses hijack host cell gene expression machinery to generate progeny virions Simple
Transcript-Specific Cytoplasmic Degradation of YRA1 Pre-mRNA Mediated by the Yeast EDC3 Protein: A Dissertation
 University of Massachusetts Medical School escholarship@umms GSBS Dissertations and Theses Graduate School of Biomedical Sciences 12-17-2007 Transcript-Specific Cytoplasmic Degradation of YRA1 Pre-mRNA
University of Massachusetts Medical School escholarship@umms GSBS Dissertations and Theses Graduate School of Biomedical Sciences 12-17-2007 Transcript-Specific Cytoplasmic Degradation of YRA1 Pre-mRNA
Islamic University Faculty of Medicine
 Islamic University Faculty of Medicine 2012 2013 2 RNA is a modular structure built from a combination of secondary and tertiary structural motifs. RNA chains fold into unique 3 D structures, which act
Islamic University Faculty of Medicine 2012 2013 2 RNA is a modular structure built from a combination of secondary and tertiary structural motifs. RNA chains fold into unique 3 D structures, which act
THE EFFECT OF HSA-MIR-105 ON PROSTATE CANCER GROWTH
 THE EFFECT OF HSA-MIR-105 ON PROSTATE CANCER GROWTH by David Rice Honeywell Bachelor of Science, Queen s University, 2010 THESIS SUBMITTED IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF MASTER
THE EFFECT OF HSA-MIR-105 ON PROSTATE CANCER GROWTH by David Rice Honeywell Bachelor of Science, Queen s University, 2010 THESIS SUBMITTED IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF MASTER
Utility of Circulating micrornas in Cardiovascular Disease
 Utility of Circulating micrornas in Cardiovascular Disease Pil-Ki Min, MD, PhD Cardiology Division, Gangnam Severance Hospital, Yonsei University College of Medicine Introduction Biology of micrornas Circulating
Utility of Circulating micrornas in Cardiovascular Disease Pil-Ki Min, MD, PhD Cardiology Division, Gangnam Severance Hospital, Yonsei University College of Medicine Introduction Biology of micrornas Circulating
DNA codes for RNA, which guides protein synthesis.
 Section 3: DNA codes for RNA, which guides protein synthesis. K What I Know W What I Want to Find Out L What I Learned Vocabulary Review synthesis New RNA messenger RNA ribosomal RNA transfer RNA transcription
Section 3: DNA codes for RNA, which guides protein synthesis. K What I Know W What I Want to Find Out L What I Learned Vocabulary Review synthesis New RNA messenger RNA ribosomal RNA transfer RNA transcription
Signal Transduction Cascades
 Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
