An esterase from anaerobic Clostridium hathewayi can hydrolyze aliphatic-aromatic polyesters
|
|
|
- Wilfred Hensley
- 5 years ago
- Views:
Transcription
1 Supporting Information An esterase from anaerobic Clostridium hathewayi can hydrolyze aliphatic-aromatic polyesters Veronika Perz a, Altijana Hromic b,c, Armin Baumschlager b, Georg Steinkellner b, Tea Pavkov-Keller b, Karl Gruber b,c, Klaus Bleymaier b, Sabine Zitzenbacher b, Armin Zankel d, Claudia Mayrhofer d, Carsten Sinkel e, Ulf Kueper e, Katharina Schlegel e, Doris Ribitsch a,f,*, Georg M. Guebitz a,f a Austrian Centre of Industrial Biotechnology ACIB, Konrad Lorenz Strasse 20, 3430 Tulln, Austria b Austrian Centre of Industrial Biotechnology ACIB, Petersgasse 14, 8010, Graz, Austria c Institute of Molecular Biosciences, University of Graz, Humboldtstrasse 50/III, 8010 Graz, Austria d Institute of Electron Microscopy and Nanoanalysis, Graz University of Technology, Steyrergasse 17/III, 8010 Graz, Austria e BASF SE, Carl-Bosch-Straße 38, Ludwigshafen, Germany f Institute of Environmental Biotechnology, University of Natural Resources and Life Sciences, Vienna, Konrad Lorenz Strasse 20, 3430 Tulln, Austria *Corresponding author: Doris Ribitsch, doris.ribitsch@acib.at, phone: Content Crystallographic parameters of Chath_Est1 from Clostridium hathewayi (Table S1).. S2 Alignment of homologous sequences starting with amino acid 351 (Figure S1)..S3 SEM image of opbat pellets (Figure S2) S4 RP-HPLC profile of the opbat batch after 28 days of incubation (Figure S3)...S5 RP-HPLC profiles of C. hathewayi cell supernatant samples (Figure S4)....S6 Expression profile of the heterologous gene Chath_Est1 in E. coli BL21(DE3) (Figure S5)...S7 Released molecules after incubation of BaETaEBa with 6 µm Chath_Est1 at different phs (Figure S6)...S8 Hydrolysis of PBAT, BTaBTaB and BTaB by 0.6 µm Chath_Est1 at ph 7.0 and 37 C (Figure S7)... S9 α/β hydrolase fold represented on chain A (Figure S8) S10 Alignment of Chath_Est1 (green) with Geobacillus stearothermophilus carboxylesterase Est55 (Figure S9)...S11 Cut through the dimer structure to the active site pocket of Chath_Est1 (Figure S10) S12 S1
2 Table S1. Crystallographic parameters of Chath_Est1 from Clostridium hathewayi X-ray source PETRA III, DESY Hamburg wavelength (Å) Temperature 100 K Space group P2 1 Cell dimensions a, b, c (Å) 62.05, , β ( ) 90.0 Resolution (Å) high resolution shell ( ) Total no. reflections Unique no. reflections Multiplicity 2.9 (2.7) Completeness (%) 84.1 (74.3) R merge (%) 8.8 (59.2) <I/σI> 7.8 (2.1) R work / R free 18.86/23.74 No. atoms Protein Water 1592 Mean B factor 24.9 R.m.s. deviations Bond lengths (Å) Bond angles ( ) Ramachandran outliers (%) 0.2 Most favored residues (%) 95.6 PDB 5A2G S2
3 EFC WP_ WP_ WP_ GHGKELKTVF AEAYPGKSPV DLLTLDTIFR GPTKEFVRSL AAAGGSVYSY GHGKELKAVF AEAYPGKSPV DLLTLDTIFR GPTKEFVRSL AAAGGSVYSY GRGKELKAVF AEAYPGKNPA DLLTLDTIFR GPTREFVRAL AAAGGSVYSY GRGKELKAVF AEAYPGKNPA DLLTLDTIFR GPTREFVRAL AAAGGSVYSY EFC LFALEFPYQN QKTA WP_ LFALEFPYQN QKTAWHCSDI PFIFHNTELV PVANIPEVSD RLEEQMFGAV WP_ LFALEFPYQN QKTAWHCSDI PFVFHNTELV PVANIPGVSD RLEEQIFGAV WP_ LFALEFPYQN QKTAWHCSDI PFVFHNTELV PAANIPEVSD RLQEQIFGAV EFC WP_ MAFARSGKPE YGGLPQWPAS RENDEATMIF DRVCEVRHNH DNRLLKLHAE WP_ MAFARTGKPE YEGLPQWPAS REDDEATMIF DRVCEVRHNH DDRLLKLHAE WP_ MAFARTGKPE YEGLPQWPAS REDDEATMIF DRVCEVRHNH DDRLLKLHAE EFC WP_ LSPKFDLAAV MAEMGDEIQH WP_ ASPKFDLAAM MAKMGDEIQH WP_ ASPKFDLAAM MTKMGDEIQH Figure S1. Alignment of homologous sequences starting with amino acid 351. EFC partial para-nitrobenzyl esterase from Clostridium hathewayi DSM-13479; WP_ para-nitrobenzyl esterase from Clostridium hathewayi CAG:224; WP_ carboxylesterase from Hungatella hathewayi; WP_ hypothetical protein from Hungatella hathewayi. S3
4 Figure S2. SEM images of opbat pellets. A: untreated opbat sample. B: opbat sample after batch-wise incubation in abiotic medium. C: opbat sample with biofilm after batch-wise incubation in biogas sludge. The incubation was performed under the conditions described in the experimental section. S4
5 Figure S3. RP-HPLC profile of the opbat batch after 28 days of incubation. Liberated Ta: terephthalic acid was quantified. S5
6 Figure S4. RP-HPLC profiles of C. hathewayi cell supernatant samples PYX + Glucose (above) and PYX + PBAT + opbat (below). Liberated hydrolysis products Ta: terephthalic acid; BTa: mono(4-hydroxybutyl) terephthalate and BTaB: bis(4-hydroxybutyl) terephthalate were quantified. S6
7 Lysate Pellet kda STD,,,,2h,,,,,4h,,,,20h,,,,2h,,,,4h,,,,20h, Figure S5. Expression profile of the heterologous gene Chath_Est1 in E. coli BL21(DE3) at 20 C and 160 rpm for 20 h after induction with 0.05 M IPTG. Lysates and pellets were withdrawn and analyzed after 2, 4 h, and 20 h after induction with IPTG. S7
8 Total released molecules [µm] ETaE ETa Ta Ba BaE ph4 ph5 ph6 ph7 ph8 ph9 ph10 Figure S6. Released molecules after incubation of BaETaEBa with 6 µm Chath_Est1 for 23 h and 37 C at different phs. The release molecules are shown in µm, ETaE: bis(2-hydroxyethyl) terephthalate, ETa: mono(2- hydroxyethyl) terephthalate, Ta: terephthalic acid, BaE: 2-hydroxyethyl benzoate and Ba: benzoic acid were quantified by means of HPLC analysis. Each bar represents the average of three independent samples; error bars indicate the standard deviation. S8
9 PBAT released molecules [µm] BTaBTaB released molecules [µm] BTaB released molecules [µm] Ta BTa 48#h# 72#h# 48#h# 72#h# 48 h 72 h Figure S7. Hydrolysis of PBAT, BTaBTaB and BTaB by 0.6 μm Chath_Est1 at ph 7.0 and 37 C. Released products (in µm) were quantified by means of RP-HPLC analysis. Ta: terephthalic acid; BTa: mono(4-hydroxybutyl) terephthalate; BTaB: bis(4-hydroxybutyl) terephthalate. Each bar represents the average of three independent samples; error bars indicate the standard deviation. S9
10 Figure S8. α/β hydrolase fold represented on chain A: alpha helices (cyan), beta sheets (magenta) and loops (salmon). S10
11 Figure S9. Alignment of Chath_Est1 (green) with Geobacillus stearothermophilus carboxylesterase Est55 (cyan, left) and pnb esterase (blue, right) S11
12 Figure S10. Cut through the dimer structure to the active site pocket of Chath_Est1. S12
Table S1. X-ray data collection and refinement statistics
 Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Enzymatic hydrolysis of polyester thin films at the nanoscale: effects of polyester structure and enzyme active-site accessibility
 Supporting information Enzymatic hydrolysis of polyester thin films at the nanoscale: effects of polyester structure and enzyme active-site accessibility Michael Thomas Zumstein, Daniela Rechsteiner, Nicolas
Supporting information Enzymatic hydrolysis of polyester thin films at the nanoscale: effects of polyester structure and enzyme active-site accessibility Michael Thomas Zumstein, Daniela Rechsteiner, Nicolas
FCC2 5CY7 FCC1 5CY8. Actinonin 5CVQ
 Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Supplementary Information Janssen et al.
 Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplemental Information. An Atlas of b-glucuronidases. in the Human Intestinal Microbiome
 Structure, Volume 2 Supplemental Information An Atlas of b-glucuronidases in the Human Intestinal Microbiome Rebecca M. Pollet, Emma H. D'Agostino, William G. Walton, Yongmei Xu, Michael S. Little, Kristen
Structure, Volume 2 Supplemental Information An Atlas of b-glucuronidases in the Human Intestinal Microbiome Rebecca M. Pollet, Emma H. D'Agostino, William G. Walton, Yongmei Xu, Michael S. Little, Kristen
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
Supporting Information
 Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
SUPPLEMENTARY INFORMATION FOR. (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation. by COX-2
 SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/9/e1600292/dc1 Supplementary Materials for Native phasing of x-ray free-electron laser data for a G protein coupled receptor Alexander Batyuk, Lorenzo Galli,
advances.sciencemag.org/cgi/content/full/2/9/e1600292/dc1 Supplementary Materials for Native phasing of x-ray free-electron laser data for a G protein coupled receptor Alexander Batyuk, Lorenzo Galli,
Figure S1. HP1α localizes to centromeres in mitosis and interacts with INCENP. (A&B) HeLa
 SUPPLEMENTARY FIGURES Figure S1. HP1α localizes to centromeres in mitosis and interacts with INCENP. (A&B) HeLa tet-on cells that stably express HP1α-CFP, HP1β-CFP, or HP1γ-CFP were monitored with livecell
SUPPLEMENTARY FIGURES Figure S1. HP1α localizes to centromeres in mitosis and interacts with INCENP. (A&B) HeLa tet-on cells that stably express HP1α-CFP, HP1β-CFP, or HP1γ-CFP were monitored with livecell
Comparison of Biochemical Properties of the Original. and Newly Identified Oleate Hydratases from
 Comparison of Biochemical Properties of the Original and Newly Identified Oleate Hydratases from Stenotrophomonas maltophilia Woo-Ri Kang, a Min-Ju Seo, a Kyung-Chul Shin, a Jin-Byung Park, b Deok-Kun
Comparison of Biochemical Properties of the Original and Newly Identified Oleate Hydratases from Stenotrophomonas maltophilia Woo-Ri Kang, a Min-Ju Seo, a Kyung-Chul Shin, a Jin-Byung Park, b Deok-Kun
Supporting Information
 Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
Bioinformatics for molecular biology
 Bioinformatics for molecular biology Structural bioinformatics tools, predictors, and 3D modeling Structural Biology Review Dr Research Scientist Department of Microbiology, Oslo University Hospital -
Bioinformatics for molecular biology Structural bioinformatics tools, predictors, and 3D modeling Structural Biology Review Dr Research Scientist Department of Microbiology, Oslo University Hospital -
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
Adaptable Lipid Matrix Promotes Protein Protein Association in Membranes
 Supporting information Adaptable Lipid Matrix Promotes Protein Protein Association in Membranes Andrey S. Kuznetsov, Anton A. Polyansky,, Markus Fleck, Pavel E. Volynsky, and Roman G. Efremov *,, M. M.
Supporting information Adaptable Lipid Matrix Promotes Protein Protein Association in Membranes Andrey S. Kuznetsov, Anton A. Polyansky,, Markus Fleck, Pavel E. Volynsky, and Roman G. Efremov *,, M. M.
PAPER No. : 16, Bioorganic and biophysical chemistry MODULE No. : 22, Mechanism of enzyme catalyst reaction (I) Chymotrypsin
 Subject Paper No and Title 16 Bio-organic and Biophysical Module No and Title 22 Mechanism of Enzyme Catalyzed reactions I Module Tag CHE_P16_M22 Chymotrypsin TABLE OF CONTENTS 1. Learning outcomes 2.
Subject Paper No and Title 16 Bio-organic and Biophysical Module No and Title 22 Mechanism of Enzyme Catalyzed reactions I Module Tag CHE_P16_M22 Chymotrypsin TABLE OF CONTENTS 1. Learning outcomes 2.
SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES
 SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors:
SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors:
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
The Binding Mode of by Electron Crystallography
 The Binding Mode of Epothilone A on α,β-tubulin by Electron Crystallography James H. Nettles, Huilin Li, Ben Cornett, Joseph M. Krahn, James P. Snyder, Kenneth H. Downing Science, Volume 305, August 6,
The Binding Mode of Epothilone A on α,β-tubulin by Electron Crystallography James H. Nettles, Huilin Li, Ben Cornett, Joseph M. Krahn, James P. Snyder, Kenneth H. Downing Science, Volume 305, August 6,
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Previous Class. Today. Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism. Protein Kinase Catalytic Properties
 Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
SUPPLEMENTAL MATERIAL. UNC119 is required for G protein trafficking in sensory neurons
 1 SUPPLEMENTAL MATERIAL UNC119 is required for G protein trafficking in sensory neurons Houbin Zhang, Ryan N. Constantine, Sergey Vorobiev, Yang Chen, Jayaraman Seetharaman, Yuanpeng Janet Huang, Rong
1 SUPPLEMENTAL MATERIAL UNC119 is required for G protein trafficking in sensory neurons Houbin Zhang, Ryan N. Constantine, Sergey Vorobiev, Yang Chen, Jayaraman Seetharaman, Yuanpeng Janet Huang, Rong
The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism
 Supplementary Information The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism Address correspondence to Yong Li (yongli@xmu.edu.cn, Tel: 86-592-218151) GW464 CDCA Supplementary
Supplementary Information The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism Address correspondence to Yong Li (yongli@xmu.edu.cn, Tel: 86-592-218151) GW464 CDCA Supplementary
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Secondary Structure. by hydrogen bonds
 Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
KDM2A. Reactions. containing. Reactions
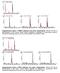 Supplementary Figure 1 KDM2A catalyses only lysine demethylation.. MALDI-TOF MS of demethylation of the shown variant histone peptides as catalysed by recombinant KDM2A. Reactions containing enzyme are
Supplementary Figure 1 KDM2A catalyses only lysine demethylation.. MALDI-TOF MS of demethylation of the shown variant histone peptides as catalysed by recombinant KDM2A. Reactions containing enzyme are
Catalysis & specificity: Proteins at work
 Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
Secondary Structure North 72nd Street, Wauwatosa, WI Phone: (414) Fax: (414) dmoleculardesigns.com
 Secondary Structure In the previous protein folding activity, you created a generic or hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously
Secondary Structure In the previous protein folding activity, you created a generic or hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Levels of Protein Structure:
 Levels of Protein Structure: PRIMARY STRUCTURE (1 ) - Defined, non-random sequence of amino acids along the peptide backbone o Described in two ways: Amino acid composition Amino acid sequence M-L-D-G-C-G
Levels of Protein Structure: PRIMARY STRUCTURE (1 ) - Defined, non-random sequence of amino acids along the peptide backbone o Described in two ways: Amino acid composition Amino acid sequence M-L-D-G-C-G
Lecture 10 More about proteins
 Lecture 10 More about proteins Today we're going to extend our discussion of protein structure. This may seem far-removed from gene cloning, but it is the path to understanding the genes that we are cloning.
Lecture 10 More about proteins Today we're going to extend our discussion of protein structure. This may seem far-removed from gene cloning, but it is the path to understanding the genes that we are cloning.
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.
 Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Chemistry 135, First Exam. September 23, Chem 135, Exam 1 SID:
 Chemistry 135, First Exam September 23, 2015 This exam will be worth 15% of your overall grade. Please read all instructions/questions carefully and provide answers in the space provided. There should
Chemistry 135, First Exam September 23, 2015 This exam will be worth 15% of your overall grade. Please read all instructions/questions carefully and provide answers in the space provided. There should
BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached
 BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached 1). 20 points total T or F (2 points each; if false, briefly state why
BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached 1). 20 points total T or F (2 points each; if false, briefly state why
Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex
 John Clizer & Greg Ralph Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex The turkey ovomucoid third domain (OMTKY3) is considered to be one of the most studied protein inhibitors. 1 Ovomucin
John Clizer & Greg Ralph Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex The turkey ovomucoid third domain (OMTKY3) is considered to be one of the most studied protein inhibitors. 1 Ovomucin
Porphyrins: Chemistry and Biology
 Porphyrins: Chemistry and Biology 20.109 Lecture 6 24 February, 2011 Goals Explore some essential roles of heme in biology Appreciate how ature has used the same cofactor to achieve diverse functions Gain
Porphyrins: Chemistry and Biology 20.109 Lecture 6 24 February, 2011 Goals Explore some essential roles of heme in biology Appreciate how ature has used the same cofactor to achieve diverse functions Gain
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
This student paper was written as an assignment in the graduate course
 77:222 Spring 2003 Free Radicals in Biology and Medicine Page 0 This student paper was written as an assignment in the graduate course Free Radicals in Biology and Medicine (77:222, Spring 2003) offered
77:222 Spring 2003 Free Radicals in Biology and Medicine Page 0 This student paper was written as an assignment in the graduate course Free Radicals in Biology and Medicine (77:222, Spring 2003) offered
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
SUPPLEMENTARY INFORMATION
 Supplementary Discussion The HGGG sequence motif forms an oxyanion hole in HLSs. In the case of GID1, the third Gly residue of the sequence motif is replaced with Ser (Ser123 in OsGID1), i.e., the sequence
Supplementary Discussion The HGGG sequence motif forms an oxyanion hole in HLSs. In the case of GID1, the third Gly residue of the sequence motif is replaced with Ser (Ser123 in OsGID1), i.e., the sequence
H C. C α. Proteins perform a vast array of biological function including: Side chain
 Topics The topics: basic concepts of molecular biology elements on Python overview of the field biological databases and database searching sequence alignments phylogenetic trees microarray data analysis
Topics The topics: basic concepts of molecular biology elements on Python overview of the field biological databases and database searching sequence alignments phylogenetic trees microarray data analysis
Supplementary Information
 Supplementary Information Structural Basis of Sialidase in Complex with Geranylated Flavonoids as Potent Natural Inhibitors Youngjin Lee, a,b Young Bae Ryu, d Hyung-Seop Youn, a,b Jung Keun Cho, e Young
Supplementary Information Structural Basis of Sialidase in Complex with Geranylated Flavonoids as Potent Natural Inhibitors Youngjin Lee, a,b Young Bae Ryu, d Hyung-Seop Youn, a,b Jung Keun Cho, e Young
Chapter 6. X-ray structure analysis of D30N tethered HIV-1 protease. dimer/saquinavir complex
 Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Chem Lecture 2 Protein Structure
 Chem 452 - Lecture 2 Protein Structure 110923 Proteins are the workhorses of a living cell and involve themselves in nearly all of the activities that take place in a cell. Their wide range of structures
Chem 452 - Lecture 2 Protein Structure 110923 Proteins are the workhorses of a living cell and involve themselves in nearly all of the activities that take place in a cell. Their wide range of structures
<Supplemental information>
 The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
Section 1 Lecture 1- Origins of Life Life probably started by Hydrothermal Vents.
 Section 1 Lecture 1- Origins of Life Life probably started by Hydrothermal Vents. Photosynthesis originated around 3GA, as cells figured out how to fix CO2 and release O2. Eukaryotes originates 1.5-2.5
Section 1 Lecture 1- Origins of Life Life probably started by Hydrothermal Vents. Photosynthesis originated around 3GA, as cells figured out how to fix CO2 and release O2. Eukaryotes originates 1.5-2.5
Protein Secondary Structure
 Protein Secondary Structure Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 2, pp. 37-45 Problems in textbook: chapter 2, pp. 63-64, #1,5,9 Directory of Jmol structures of proteins: http://www.biochem.arizona.edu/classes/bioc462/462a/jmol/routines/routines.html
Protein Secondary Structure Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 2, pp. 37-45 Problems in textbook: chapter 2, pp. 63-64, #1,5,9 Directory of Jmol structures of proteins: http://www.biochem.arizona.edu/classes/bioc462/462a/jmol/routines/routines.html
List of Figures. List of Tables
 Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Lesson 5 Proteins Levels of Protein Structure
 Lesson 5 Proteins Levels of Protein Structure Primary 1º Structure The primary structure is simply the sequence of amino acids in a protein. Chains of amino acids are written from the amino terminus (N-terminus)
Lesson 5 Proteins Levels of Protein Structure Primary 1º Structure The primary structure is simply the sequence of amino acids in a protein. Chains of amino acids are written from the amino terminus (N-terminus)
Structural insights into the potential of 4-fluoroproline to modulate biophysical properties of proteins
 SUPPLEMENTARY INFORMATION Structural insights into the potential of 4-fluoroproline to modulate biophysical properties of proteins Bastian Holzberger, Samra Obeid, Wolfram Welte, Kay Diederichs, and Andreas
SUPPLEMENTARY INFORMATION Structural insights into the potential of 4-fluoroproline to modulate biophysical properties of proteins Bastian Holzberger, Samra Obeid, Wolfram Welte, Kay Diederichs, and Andreas
172R 172K TAM-2/172R TAM-2/172K. AZT concentration [nm] AZT concentration [nm] MgCl 2 2.5K 2.5K 5K 2.5K 5K 2.5K K 5K 2.5K 5K 2.5K 50 2.
![172R 172K TAM-2/172R TAM-2/172K. AZT concentration [nm] AZT concentration [nm] MgCl 2 2.5K 2.5K 5K 2.5K 5K 2.5K K 5K 2.5K 5K 2.5K 50 2. 172R 172K TAM-2/172R TAM-2/172K. AZT concentration [nm] AZT concentration [nm] MgCl 2 2.5K 2.5K 5K 2.5K 5K 2.5K K 5K 2.5K 5K 2.5K 50 2.](/thumbs/82/85138523.jpg) 5 5 5 5 A MgCl 2 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] B 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] ATP + ATP - Supplemental Figure 1. Primer extension of HIV-1 RT polymorphisms
5 5 5 5 A MgCl 2 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] B 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] ATP + ATP - Supplemental Figure 1. Primer extension of HIV-1 RT polymorphisms
Ras and Cell Signaling Exercise
 Ras and Cell Signaling Exercise Learning Objectives In this exercise, you will use, a protein 3D- viewer, to explore: the structure of the Ras protein the active and inactive state of Ras and the amino
Ras and Cell Signaling Exercise Learning Objectives In this exercise, you will use, a protein 3D- viewer, to explore: the structure of the Ras protein the active and inactive state of Ras and the amino
APOLs with low ph dependence can kill all African trypanosomes
 SUPPLEMENTARY INFORMATION Letters DOI: 1.138/s41564-17-34-1 In the format provided by the authors and unedited. APOLs with low ph dependence can kill all African trypanosomes Frédéric Fontaine 1, Laurence
SUPPLEMENTARY INFORMATION Letters DOI: 1.138/s41564-17-34-1 In the format provided by the authors and unedited. APOLs with low ph dependence can kill all African trypanosomes Frédéric Fontaine 1, Laurence
Proteins consist of joined amino acids They are joined by a Also called an Amide Bond
 Lecture Two: Peptide Bond & Protein Structure [Chapter 2 Berg, Tymoczko & Stryer] (Figures in Red are for the 7th Edition) (Figures in Blue are for the 8th Edition) Proteins consist of joined amino acids
Lecture Two: Peptide Bond & Protein Structure [Chapter 2 Berg, Tymoczko & Stryer] (Figures in Red are for the 7th Edition) (Figures in Blue are for the 8th Edition) Proteins consist of joined amino acids
Table S1: Kinetic parameters of drug and substrate binding to wild type and HIV-1 protease variants. Data adapted from Ref. 6 in main text.
 Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular
 Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
F-actin VWF Vinculin. F-actin. Vinculin VWF
 a F-actin VWF Vinculin b F-actin VWF Vinculin Supplementary Fig. 1. WPBs in HUVECs are located along stress fibers and at focal adhesions. (a) Immunofluorescence images of f-actin (cyan), VWF (yellow),
a F-actin VWF Vinculin b F-actin VWF Vinculin Supplementary Fig. 1. WPBs in HUVECs are located along stress fibers and at focal adhesions. (a) Immunofluorescence images of f-actin (cyan), VWF (yellow),
Carbohydrates. Organic compounds which comprise of only C, H and O. C x (H 2 O) y
 Carbohydrates Organic compounds which comprise of only C, H and O C x (H 2 O) y Carbohydrates Monosaccharides Simple sugar Soluble in water Precursors in synthesis triose sugars of other (C3) molecules
Carbohydrates Organic compounds which comprise of only C, H and O C x (H 2 O) y Carbohydrates Monosaccharides Simple sugar Soluble in water Precursors in synthesis triose sugars of other (C3) molecules
Ch5: Macromolecules. Proteins
 Ch5: Macromolecules Proteins Essential Knowledge 4.A.1 The subcomponents of biological molecules and their sequence determine the properties of that molecule A. Structure and function of polymers are derived
Ch5: Macromolecules Proteins Essential Knowledge 4.A.1 The subcomponents of biological molecules and their sequence determine the properties of that molecule A. Structure and function of polymers are derived
This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth 2 points.
 MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
Manja Henze, Dorothee Merker and Lothar Elling. 1. Characteristics of the Recombinant β-glycosidase from Pyrococcus
 S1 of S17 Supplementary Materials: Microwave-Assisted Synthesis of Glycoconjugates by Transgalactosylation with Recombinant Thermostable β-glycosidase from Pyrococcus Manja Henze, Dorothee Merker and Lothar
S1 of S17 Supplementary Materials: Microwave-Assisted Synthesis of Glycoconjugates by Transgalactosylation with Recombinant Thermostable β-glycosidase from Pyrococcus Manja Henze, Dorothee Merker and Lothar
Lecture 15. Membrane Proteins I
 Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Supplementary Fig. 1 Atomic force microscopy topography images Two-dimensional atomic force microscopy images (with an area of 1 m 1 m) of Cu and
 Supplementary Fig. 1 Atomic force microscopy topography images Two-dimensional atomic force microscopy images (with an area of 1 m 1 m) of Cu and Cu(O = 5.0%) films deposited on 20-nm-thick ZnO films during
Supplementary Fig. 1 Atomic force microscopy topography images Two-dimensional atomic force microscopy images (with an area of 1 m 1 m) of Cu and Cu(O = 5.0%) films deposited on 20-nm-thick ZnO films during
Review II: The Molecules of Life
 Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
Guided Inquiry Skills Lab. Additional Lab 1 Making Models of Macromolecules. Problem. Introduction. Skills Focus. Materials.
 Additional Lab 1 Making Models of Macromolecules Guided Inquiry Skills Lab Problem How do monomers join together to form polymers? Introduction A small number of elements make up most of the mass of your
Additional Lab 1 Making Models of Macromolecules Guided Inquiry Skills Lab Problem How do monomers join together to form polymers? Introduction A small number of elements make up most of the mass of your
Supplementary Figure 1. Properties of various IZUMO1 monoclonal antibodies and behavior of SPACA6. (a) (b) (c) (d) (e) (f) (g) .
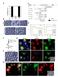 Supplementary Figure 1. Properties of various IZUMO1 monoclonal antibodies and behavior of SPACA6. (a) The inhibitory effects of new antibodies (Mab17 and Mab18). They were investigated in in vitro fertilization
Supplementary Figure 1. Properties of various IZUMO1 monoclonal antibodies and behavior of SPACA6. (a) The inhibitory effects of new antibodies (Mab17 and Mab18). They were investigated in in vitro fertilization
Proteins. These are polymers of 20 common amino acids linked in various sequences. Proteins differ in molecular mass, molecular structure and shape
 Proteins These are polymers of 20 common amino acids linked in various sequences. Proteins differ in molecular mass, molecular structure and shape Characteristics if protein Amino acids are linked by covalent
Proteins These are polymers of 20 common amino acids linked in various sequences. Proteins differ in molecular mass, molecular structure and shape Characteristics if protein Amino acids are linked by covalent
Supporting Information
 Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
BIO 311C Spring Lecture 15 Friday 26 Feb. 1
 BIO 311C Spring 2010 Lecture 15 Friday 26 Feb. 1 Illustration of a Polypeptide amino acids peptide bonds Review Polypeptide (chain) See textbook, Fig 5.21, p. 82 for a more clear illustration Folding and
BIO 311C Spring 2010 Lecture 15 Friday 26 Feb. 1 Illustration of a Polypeptide amino acids peptide bonds Review Polypeptide (chain) See textbook, Fig 5.21, p. 82 for a more clear illustration Folding and
Near-infrared fluorescent proteins
 nature methods Near-infrared fluorescent proteins Dmitry Shcherbo, Irina I Shemiakina, Anastasiya V Ryabova, Kathryn E Luker, Bradley T Schmidt, Ekaterina A Souslova, Tatiana V Gorodnicheva, Lydia Strukova,
nature methods Near-infrared fluorescent proteins Dmitry Shcherbo, Irina I Shemiakina, Anastasiya V Ryabova, Kathryn E Luker, Bradley T Schmidt, Ekaterina A Souslova, Tatiana V Gorodnicheva, Lydia Strukova,
Kinetics analysis of β-fructofuranosidase enzyme. 1-Effect of Time Incubation On The Rate Of An Enzymatic Reaction
 Kinetics analysis of β-fructofuranosidase enzyme 1-Effect of Time Incubation On The Rate Of An Enzymatic Reaction Enzyme kinetics It is the study of the chemical reactions that are catalyzed by enzymes.
Kinetics analysis of β-fructofuranosidase enzyme 1-Effect of Time Incubation On The Rate Of An Enzymatic Reaction Enzyme kinetics It is the study of the chemical reactions that are catalyzed by enzymes.
Data are contained in multiple tabs in Excel spreadsheets and in CSV files.
 Contents Overview Curves Methods Measuring enzymatic activity (figure 2) Enzyme characterisation (Figure S1, S2) Enzyme kinetics (Table 3) Effect of ph on activity (figure 3B) Effect of metals and inhibitors
Contents Overview Curves Methods Measuring enzymatic activity (figure 2) Enzyme characterisation (Figure S1, S2) Enzyme kinetics (Table 3) Effect of ph on activity (figure 3B) Effect of metals and inhibitors
FOR OPTIMAL GUT HEALTH KEMIN.COM/GUTHEALTH
 FOR OPTIMAL GUT HEALTH KEMIN.COM/GUTHEALTH ALETA A SOURCE OF 1,3-BETA GLUCANS Aleta is highly bioavailable, offering a concentration greater than 5% of 1,3-beta glucans. Aleta provides a consistent response
FOR OPTIMAL GUT HEALTH KEMIN.COM/GUTHEALTH ALETA A SOURCE OF 1,3-BETA GLUCANS Aleta is highly bioavailable, offering a concentration greater than 5% of 1,3-beta glucans. Aleta provides a consistent response
BCH Graduate Survey of Biochemistry
 BCH 5045 Graduate Survey of Biochemistry Instructor: Charles Guy Producer: Ron Thomas Director: Glen Graham Lecture 10 Slide sets available at: http://hort.ifas.ufl.edu/teach/guyweb/bch5045/index.html
BCH 5045 Graduate Survey of Biochemistry Instructor: Charles Guy Producer: Ron Thomas Director: Glen Graham Lecture 10 Slide sets available at: http://hort.ifas.ufl.edu/teach/guyweb/bch5045/index.html
Proteins are linear polymers built of monomer units called amino acids. Proteins contain a wide range of functional groups.
 Chapter 2: Protein Structure and Function Proteins arevery versatile with regards to functions for the cell Uses? Proteins are linear polymers built of monomer units called amino acids. One dimensional
Chapter 2: Protein Structure and Function Proteins arevery versatile with regards to functions for the cell Uses? Proteins are linear polymers built of monomer units called amino acids. One dimensional
Supplementary Figure S1. Supplementary Figure S1. Chemical structure of CH
 Supplementary Figure S1 Supplementary Figure S1. Chemical structure of CH7057288. Supplementary Figure S2 Kinase % competition TRKA 99 TRKC 95 TRKB 92 MEK5 87 AXL 84 CLK1 82 MERTK 82 GCN2 (Kin.Dom.2,S808G)
Supplementary Figure S1 Supplementary Figure S1. Chemical structure of CH7057288. Supplementary Figure S2 Kinase % competition TRKA 99 TRKC 95 TRKB 92 MEK5 87 AXL 84 CLK1 82 MERTK 82 GCN2 (Kin.Dom.2,S808G)
Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector
 kda 1 2 3 4 5 26 14 1 7 Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector 5 4 35 25 15 1 Supplementary Figure 1. SDS-PAGE of acterially expressed and purified proteins.
kda 1 2 3 4 5 26 14 1 7 Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector 5 4 35 25 15 1 Supplementary Figure 1. SDS-PAGE of acterially expressed and purified proteins.
FIRST MIDTERM EXAMINATION
 FIRST MIDTERM EXAMINATION 1. True or false: because enzymes are produced by living organisms and because they allow chemical reactions to occur that would not otherwise occur, enzymes represent an exception
FIRST MIDTERM EXAMINATION 1. True or false: because enzymes are produced by living organisms and because they allow chemical reactions to occur that would not otherwise occur, enzymes represent an exception
Course Content
 Biology Induction Course Content AS Biology A-Level Biology AS Practical Work Career options Degree options Research Based IS Task Due date: 1 st lesson back after the summer holidays 1. Compare and contrast
Biology Induction Course Content AS Biology A-Level Biology AS Practical Work Career options Degree options Research Based IS Task Due date: 1 st lesson back after the summer holidays 1. Compare and contrast
Nafith Abu Tarboush DDS, MSc, PhD
 Nafith Abu Tarboush DDS, MSc, PhD natarboush@ju.edu.jo www.facebook.com/natarboush Protein conformation Many conformations are possible for proteins due to flexibility of amino acids linked by peptide
Nafith Abu Tarboush DDS, MSc, PhD natarboush@ju.edu.jo www.facebook.com/natarboush Protein conformation Many conformations are possible for proteins due to flexibility of amino acids linked by peptide
Unit 3: Chemistry of Life Mr. Nagel Meade High School
 Unit 3: Chemistry of Life Mr. Nagel Meade High School IB Syllabus Statements 3.2.1 Distinguish between organic and inorganic compounds. 3.2.2 Identify amino acids, glucose, ribose and fatty acids from
Unit 3: Chemistry of Life Mr. Nagel Meade High School IB Syllabus Statements 3.2.1 Distinguish between organic and inorganic compounds. 3.2.2 Identify amino acids, glucose, ribose and fatty acids from
We are going to talk about two classifications of proteins: fibrous & globular.
 Slide # 13 (fibrous proteins) : We are going to talk about two classifications of proteins: fibrous & globular. *fibrous proteins: (dense fibers) *Their structures are mainly formed of the secondary structure
Slide # 13 (fibrous proteins) : We are going to talk about two classifications of proteins: fibrous & globular. *fibrous proteins: (dense fibers) *Their structures are mainly formed of the secondary structure
Hemoglobin & Sickle Cell Anemia Exercise
 Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Learning Objectives In this exercise, you will use StarBiochem, a protein 3D viewer, to explore the structure of the normal hemoglobin protein
Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Learning Objectives In this exercise, you will use StarBiochem, a protein 3D viewer, to explore the structure of the normal hemoglobin protein
The three important structural features of proteins:
 The three important structural features of proteins: a. Primary (1 o ) The amino acid sequence (coded by genes) b. Secondary (2 o ) The interaction of amino acids that are close together or far apart in
The three important structural features of proteins: a. Primary (1 o ) The amino acid sequence (coded by genes) b. Secondary (2 o ) The interaction of amino acids that are close together or far apart in
Introduction to Protein Structure Collection
 Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
So where were we? But what does the order mean? OK, so what's a protein? 4/1/11
 So where were we? We know that DNA is responsible for heredity Chromosomes are long pieces of DNA DNA turned out to be the transforming principle We know that DNA is shaped like a long double helix, with
So where were we? We know that DNA is responsible for heredity Chromosomes are long pieces of DNA DNA turned out to be the transforming principle We know that DNA is shaped like a long double helix, with
Reducing Intestinal Digestion and Absorption of Fat Using a Nature-Derived Biopolymer: Interference of Triglyceride Hydrolysis by Nanocellulose.
 Reducing Intestinal Digestion and Absorption of Fat Using a Nature-Derived Biopolymer: Interference of Triglyceride Hydrolysis by Nanocellulose. Glen M. DeLoid 1 *, Ikjot Singh Soha1 2, Laura R. Lorente
Reducing Intestinal Digestion and Absorption of Fat Using a Nature-Derived Biopolymer: Interference of Triglyceride Hydrolysis by Nanocellulose. Glen M. DeLoid 1 *, Ikjot Singh Soha1 2, Laura R. Lorente
The high efficiency of sub-2 µm UHPLC columns combined with the low pressure and high speed of monolith columns.
 The high efficiency of sub-2 µm UHPLC columns combined with the low pressure and high speed of monolith columns. Fast High peak capacity Low back pressure PROTEIN PEPTIDE UHPLC COLUMNS Put these new, high-performance
The high efficiency of sub-2 µm UHPLC columns combined with the low pressure and high speed of monolith columns. Fast High peak capacity Low back pressure PROTEIN PEPTIDE UHPLC COLUMNS Put these new, high-performance
The Structure and Function of Macromolecules
 The Structure and Function of Macromolecules Macromolecules are polymers Polymer long molecule consisting of many similar building blocks. Monomer the small building block molecules. Carbohydrates, proteins
The Structure and Function of Macromolecules Macromolecules are polymers Polymer long molecule consisting of many similar building blocks. Monomer the small building block molecules. Carbohydrates, proteins
Journal of Chemical and Pharmaceutical Research, 2015, 7(12): Research Article
 Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2015, 7(12):1082-1086 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Determination of three dimensional structure
Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2015, 7(12):1082-1086 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Determination of three dimensional structure
Polar bodies are either introduced or unmasked, which results in more polar metabolites Phase I reactions can lead either to activation or
 Polar bodies are either introduced or unmasked, which results in more polar metabolites Phase I reactions can lead either to activation or inactivation of the drug (i.e. therapeutic effects or toxicity)
Polar bodies are either introduced or unmasked, which results in more polar metabolites Phase I reactions can lead either to activation or inactivation of the drug (i.e. therapeutic effects or toxicity)
Org/Biochem Final Lec Form, Spring 2012 Page 1 of 6
 Page 1 of 6 Missing Complete Protein and Question #45 Key Terms: Fill in the blank in the following 25 statements with one of the key terms in the table. Each key term may only be used once. Print legibly.
Page 1 of 6 Missing Complete Protein and Question #45 Key Terms: Fill in the blank in the following 25 statements with one of the key terms in the table. Each key term may only be used once. Print legibly.
Statin inhibition of HMG-CoA reductase: a 3-dimensional view
 Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
