Controlling Macromolecular Topology with Genetically Encoded. SpyTag-SpyCatcher Chemistry
|
|
|
- Randolf Hoover
- 5 years ago
- Views:
Transcription
1 Electronic Supporting Information for Controlling Macromolecular Topology with Genetically Encoded SpyTag-SpyCatcher Chemistry Wen-Bin Zhang, Fei Sun, David A. Tirrell,* and Frances H. Arnold* Division of Chemistry and Chemical Engineering, California Institute of Technology, Pasadena, California 91125, United States These two authors contributed equally. S-1
2 Table of Contents Page 1. Figure S1. (a) Cartoon illustration of the original ELP construct. It contains four restriction sites: SalI (GTCGAC), SacI (GAGCTC), SpeI (ACTAGT), and XhoI (CTCGAG) as shown. (b) The gene inserts can be made in a way that is either flanked by SalI and XhoI for insertion into the N-terminus or C-terminus or by SacI and SpeI for insertion into the middle of the chain. The restriction sites SalI and XhoI share the same sticky ends. Therefore, the insert can be used to ligate the vector that is digested by SalI or XhoI. The inserts can be SpyTag or SpyCatcher.... S4 2. Figure S2. Full amino acid sequence of AB20D (351 a.a. MW = 34002). The reactive aspartic acid (20D) is shown in the box at the first line. It is changed to nonreactive alanine (20A) in AB20A. The reactive lysine (255K) is shown in the box at the 7 th line. The segments highlighted in red, yellow, and purple are the SpyTag, TEV protease recognition sequence, and SpyCatcher, respectively.... S5 3. Figure S3. MALDI-TOF mass spectra of AB20D expressed at 37 o C with induction: (a) monomer; (b) dimer; (c) trimer; and (d) tetramer.... S6 4. Figure S4. MALDI-TOF mass spectrum of AB20A... S7 5. Figure S5. MALDI-TOF mass spectrum of relinearized AB20D... S8 6. Figure S6. Full amino acid sequence of EAEB (332 a.a. MW = 32172). The reactive aspartic acid (102D) and lysine (236K) are shown in the box at the 3 th and 6 th lines, respectively. The segments highlighted in red, yellow, and purple are the SpyTag, TEV protease recognition sequence, and SpyCatcher, respectively... S9 7. Figure S7. MALDI-TOF mass spectra of EAEB expressed at 37 o C with 1 mm IPTG induction: (a) monomer; (b) dimer; and (c) trimer.... S10 8. Figure S8. Full amino acid sequence of (a) EA and (b) EB. The reactive aspartic acid (203D) and lysine (240K) are shown in the box. The segments highlighted in red and purple are SpyTag and SpyCatcher, respectively... S11 9. Figure S9. MALDI-TOF mass spectra of EA (a) and EB (b).... S Figure S10. Full amino acid sequence of (a) EAE and (b) EBE. The reactive aspartic acid (102D) and lysine (139K) are shown in the box. The segments highlighted in red and purple are SpyTag and SpyCatcher, respectively... S Figure S11. MALDI-TOF mass spectra of EAE (a) and EBE (b).... S14 S-2
3 12. Figure S12. Full amino acid sequence of (a) AA and (b) BB. The reactive aspartic acid (20D and 218D) and lysine (57K and 377K) are shown in the box. The segments highlighted in red and purple are SpyTag and SpyCatcher, respectively.... S Figure S13. MALDI-TOF mass spectra of AA (a) and BB (b)... S Figure S14. MALDI-TOF mass spectrum of the block protein (EA+EB).... S Figure S15. MALDI-TOF mass spectra of the 3-arm star proteins EA+EBE (a) and EAE+EB (b) and the 4-arm star protein EAE+EBE (c)... S Figure S16. MALDI-TOF mass spectra of the H-shaped proteins AA+2EBE (a) and BB+2EAE (b) and the corresponding intermediate 3-arm proteins AA+EBE (c) and BB+EBE (d).... S Table S1. Summary of bacterial strains and plasmids used in this study... S20 S-3
4 Figure S1. (a) Cartoon illustration of the original ELP construct. It contains four restriction sites: SalI (GTCGAC), SacI (GAGCTC), SpeI (ACTAGT), and XhoI (CTCGAG) as shown. (b) The gene inserts can be made in a way that is either flanked by SalI and XhoI for insertion into the N-terminus or C-terminus or by SacI and SpeI for insertion into the middle of the chain. The restriction sites SalI and XhoI share the same sticky ends. Therefore, the insert can be used to ligate the vector that is digested by SalI or XhoI. The inserts can be SpyTag or SpyCatcher. S-4
5 1 MKGSSHHHHHHVDAHIVMV D AYKPTKLDGHGVGVPGVGVP 41 GVGVPGEGVPGVGVPGVGVPGVGVPGVGVPGEGVPGVGVP 81 GVGVPGVGVPGVGVPGEGVPGVGVPGVGELYAVTGRGDSP 121 ASSAPIATSVPGVGVPGVGVPGEGVPGVGVPGVGVPGVGV 161 PGVGVPGEGVPGVGVPGVGVPGVGVPGVGVPGEGVPGVGV 201 PGVGVPGGLLDIPTTENLYFQGAMVDTLSGLSSEQGQSGD 241 MTIEEDSATHIKFS K RDEDGKELAGATMELRDSSGKTIST 281 WISDGQVKDFYLYPGKYTFVETAAPDGYEVATAITFTVNE 321 QGQVTVNGKATKGDAHIDGPQGIWGQLEWKK Figure S2. Full amino acid sequence of AB20D (351 a.a. MW = 34002). The reactive aspartic acid (20D) is shown in the box at the first line. It is changed to nonreactive alanine (20A) in AB20A. The reactive lysine (255K) is shown in the box at the 7 th line. The segments highlighted in red, yellow, and purple are the SpyTag, TEV protease recognition sequence, and SpyCatcher, respectively. S-5
6 Figure S3. MALDI-TOF mass spectra of AB20D expressed at 37 o C with induction: (a) monomer; (b) dimer; (c) trimer; and (d) tetramer. S-6
7 Figure S4. MALDI-TOF mass spectrum of AB20A. S-7
8 Figure S5. MALDI-TOF mass spectrum of relinearized AB20D. S-8
9 1 MKGSSHHHHHHVDGHGVGVPGVGVPGVGVPGEGVPGVGVP 41 GVGVPGVGVPGVGVPGEGVPGVGVPGVGVPGVGVPGVGVP 81 GEGVPGVGVPGVGELAHIVMV D AYKPTKTSVPGVGVPGVG 121 VPGEGVPGVGVPGVGVPGVGVPGVGVPGEGVPGVGVPGVG 161 VPGVGVPGVGVPGEGVPGVGVPGVGVPGGLLDIPTTENLY 201 FQGAMVDTLSGLSSEQGQSGDMTIEEDSATHIKFS K RDED 241 GKELAGATMELRDSSGKTISTWISDGQVKDFYLYPGKYTF 281 VETAAPDGYEVATAITFTVNEQGQVTVNGKATKGDAHIDG 321 PQGIWGQLEWKK Figure S6. Full amino acid sequence of EAEB (332 a.a. MW = 32172). The reactive aspartic acid (102D) and lysine (236K) are shown in the box at the 3 th and 6 th lines, respectively. The segments highlighted in red, yellow, and purple are the SpyTag, TEV protease recognition sequence, and SpyCatcher, respectively. S-9
10 Figure S7. MALDI-TOF mass spectra of EAEB expressed at 37 o C with 1 mm IPTG induction: (a) monomer; (b) dimer; and (c) trimer. S-10
11 (a) EA: 214 a.a. MW = MKGSSHHHHHHVDGHGVGVPGVGVPGVGVPGEGVPGVGVP 41 GVGVPGVGVPGVGVPGEGVPGVGVPGVGVPGVGVPGVGVP 81 GEGVPGVGVPGVGELYAVTGRGDSPASSAPIATSVPGVGV 121 PGVGVPGEGVPGVGVPGVGVPGVGVPGVGVPGEGVPGVGV 161 PGVGVPGVGVPGVGVPGEGVPGVGVPGVGVPGGLLDAHIV 201 MV D AYKPTKLEWKK (b) EB: 336 a.a. MW = MKGSSHHHHHHVDGHGVGVPGVGVPGVGVPGEGVPGVGVP 41 GVGVPGVGVPGVGVPGEGVPGVGVPGVGVPGVGVPGVGVP 81 GEGVPGVGVPGVGELYAVTGRGDSPASSAPIATSVPGVGV 121 PGVGVPGEGVPGVGVPGVGVPGVGVPGVGVPGEGVPGVGV 161 PGVGVPGVGVPGVGVPGEGVPGVGVPGVGVPGGLLDIPTT 201 ENLYFQGAMVDTLSGLSSEQGQSGDMTIEEDSATHIKFS K 241 RDEDGKELAGATMELRDSSGKTISTWISDGQVKDFYLYPG 281 KYTFVETAAPDGYEVATAITFTVNEQGQVTVNGKATKGDA 321 HIDGPQGIWGQLEWKK Figure S8. Full amino acid sequence of (a) EA and (b) EB. The reactive aspartic acid (203D) and lysine (240K) are shown in the box. The segments highlighted in red and purple are SpyTag and SpyCatcher, respectively. S-11
12 Figure S9. MALDI-TOF mass spectra of EA (a) and EB (b). S-12
13 (a) EAE: 205 a.a. MW = MKGSSHHHHHHVDGHGVGVPGVGVPGVGVPGEGVPGVGVP 41 GVGVPGVGVPGVGVPGEGVPGVGVPGVGVPGVGVPGVGVP 81 GEGVPGVGVPGVGELAHIVMV D AYKPTKTSVPGVGVPGVG 121 VPGEGVPGVGVPGVGVPGVGVPGVGVPGEGVPGVGVPGVG 161 VPGVGVPGVGVPGEGVPGVGVPGVGVPGGLLDGPQGIWGQ 201 LEKKM (b) EBE: 327 a.a. MW = MKGSSHHHHHHVDGHGVGVPGVGVPGVGVPGEGVPGVGVP 41 GVGVPGVGVPGVGVPGEGVPGVGVPGVGVPGVGVPGVGVP 81 GEGVPGVGVPGVGELIPTTENLYFQGAMVDTLSGLSSEQG 121 QSGDMTIEEDSATHIKFS K RDEDGKELAGATMELRDSSGK 161 TISTWISDGQVKDFYLYPGKYTFVETAAPDGYEVATAITF 201 TVNEQGQVTVNGKATKGDAHIDGPQGIWGQTSVPGVGVPG 241 VGVPGEGVPGVGVPGVGVPGVGVPGVGVPGEGVPGVGVPG 281 VGVPGVGVPGVGVPGEGVPGVGVPGVGVPGGLLDGPQGIW 321 GQLEKKM Figure S10. Full amino acid sequence of (a) EAE and (b) EBE. The reactive aspartic acid (102D) and lysine (139K) are shown in the box. The segments highlighted in red and purple are SpyTag and SpyCatcher, respectively. S-13
14 Figure S11. MALDI-TOF mass spectra of EAE (a) and EBE (b). S-14
15 (a) AA: 229 a.a. MW = MKGSSHHHHHHVDAHIVMV D AYKPTKLDGHGVGVPGVGVP 41 GVGVPGEGVPGVGVPGVGVPGVGVPGVGVPGEGVPGVGVP 81 GVGVPGVGVPGVGVPGEGVPGVGVPGVGELYAVTGRGDSP 121 ASSAPIATSVPGVGVPGVGVPGEGVPGVGVPGVGVPGVGV 161 PGVGVPGEGVPGVGVPGVGVPGVGVPGVGVPGEGVPGVGV 201 PGVGVPGGLLDAHIVMV D AYKPTKLEWKK* (b) BB: 473 a.a. MW = MKGSSHHHHHHVDIPTTENLYFQGAMVDTLSGLSSEQGQS 41 GDMTIEEDSATHIKFS K RDEDGKELAGATMELRDSSGKTI 81 STWISDGQVKDFYLYPGKYTFVETAAPDGYEVATAITFTV 121 NEQGQVTVNGKATKGDAHIDGPQGIWGQLDGHGVGVPGVG 161 VPGVGVPGEGVPGVGVPGVGVPGVGVPGVGVPGEGVPGVG 201 VPGVGVPGVGVPGVGVPGEGVPGVGVPGVGELYAVTGRGD 241 SPASSAPIATSVPGVGVPGVGVPGEGVPGVGVPGVGVPGV 281 GVPGVGVPGEGVPGVGVPGVGVPGVGVPGVGVPGEGVPGV 321 GVPGVGVPGGLLDIPTTENLYFQGAMVDTLSGLSSEQGQS 361 GDMTIEEDSATHIKFS K RDEDGKELAGATMELRDSSGKTI 401 STWISDGQVKDFYLYPGKYTFVETAAPDGYEVATAITFTV 441 NEQGQVTVNGKATKGDAHIDGPQGIWGQLEWKK* Figure S12. Full amino acid sequence of (a) AA and (b) BB. The reactive aspartic acid (20D and 218D) and lysine (57K and 377K) are shown in the box. The segments highlighted in red and purple are SpyTag and SpyCatcher, respectively. S-15
16 Figure S13. MALDI-TOF mass spectra of AA (a) and BB (b). S-16
17 Figure S14. MALDI-TOF mass spectrum of the block protein (EA+EB). S-17
18 Figure S15. MALDI-TOF mass spectra of the 3-arm star proteins EA+EBE (a) and EAE+EB (b) and the 4-arm star protein EAE+EBE (c). S-18
19 Figure S16. MALDI-TOF mass spectra of the H-shaped proteins AA+2EBE (a) and BB+2EAE (b) and the corresponding intermediate 3-arm proteins AA+EBE (c) and BB+EBE (d). S-19
20 Table S1. Summary of bacterial strains and plasmids used in this study Strains Relevant Characteristics Sources E. coli DH5α Stratagene BL21 star(de3) Invitrogen Plasmids pqe-80l T5 promoter-operator, N-terminal Qiagen His tag, Amp r pqe-elp The pqe-80l plasmid containing The starting vector for all the the gene encoding elastin constructions of recombinant proteins in this study pqe-ab20d The plasmid for the expression of This study circular protein pqe-ab20a The plasmid mutant for the This study expression of the control linear protein pqe-eaeb The plasmid for the expression of This study tadpole protein pqe-ea The plasmid for the expression of This study Elastin-SpyTag pqe-eb The plasmid for the expression of This study Elastin-SpyCatcher pqe-eae The plasmid for the expression of This study Elastin-SpyTag-Elastin pqe-ebe The plasmid for the expression of This study Elastin-SpyCatcher-Elastin pqe-aa The plasmid for the expression of This study SpyTag-Elastin-SpyTag pqe-bb The plasmid for the expression of SpyCatcher-Elastin-SpyCatcher This study S-20
Overview of the Expressway Cell-Free Expression Systems. Expressway Mini Cell-Free Expression System
 Overview of the Expressway Cell-Free Expression Systems The Expressway Cell-Free Expression Systems use an efficient coupled transcription and translation reaction to produce up to milligram quantities
Overview of the Expressway Cell-Free Expression Systems The Expressway Cell-Free Expression Systems use an efficient coupled transcription and translation reaction to produce up to milligram quantities
Supplemental Data Functional characterization of core components of the Bacillus subtilis c- di-
 Supplemental Data Functional characterization of core components of the Bacillus subtilis c- di- GMP signaling pathway Xiaohui Gao 1,3, Sampriti Mukherjee 2, Paige M. Matthews 1, Loubna A. Hammad 1, Daniel
Supplemental Data Functional characterization of core components of the Bacillus subtilis c- di- GMP signaling pathway Xiaohui Gao 1,3, Sampriti Mukherjee 2, Paige M. Matthews 1, Loubna A. Hammad 1, Daniel
Lecture 6. Burr BIO 4353/6345 HIV/AIDS. Tetramer staining of T cells (CTL s) Andrew McMichael seminar: Background
 Lecture 6 Burr BIO 4353/6345 HIV/AIDS Andrew McMichael seminar: Background Tetramer staining of T cells (CTL s) 1. Vβ 19: There are 52 T cell receptor (TCR) Vβ gene segments in germ line DNA (See following
Lecture 6 Burr BIO 4353/6345 HIV/AIDS Andrew McMichael seminar: Background Tetramer staining of T cells (CTL s) 1. Vβ 19: There are 52 T cell receptor (TCR) Vβ gene segments in germ line DNA (See following
Mass Spectrometry. Mass spectrometer MALDI-TOF ESI/MS/MS. Basic components. Ionization source Mass analyzer Detector
 Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Catalysis & specificity: Proteins at work
 Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin
 SUPPORTING INFORMATION Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin Anna R. Arnold, Andy Zhou, and Jacqueline K. Barton Division of Chemistry and Chemical Engineering, California
SUPPORTING INFORMATION Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin Anna R. Arnold, Andy Zhou, and Jacqueline K. Barton Division of Chemistry and Chemical Engineering, California
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB
 Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Enzyme Catalytic Mechanisms. Dr. Kevin Ahern
 Enzyme Catalytic Mechanisms Dr. Kevin Ahern Cleave Peptide Bonds Specificity of Cutting Common Active Site Composition/Structure Mechanistically Well Studied Chymotrypsin Chymotrypsin Catalysis H2O Chymotrypsin
Enzyme Catalytic Mechanisms Dr. Kevin Ahern Cleave Peptide Bonds Specificity of Cutting Common Active Site Composition/Structure Mechanistically Well Studied Chymotrypsin Chymotrypsin Catalysis H2O Chymotrypsin
Gene Vaccine Dr. Sina Soleimani
 Gene Vaccine Dr. Sina Soleimani Human Viral Vaccines Quality Control Laboratory (HVVQC) Titles 1. A short Introduction of Vaccine History 2. First Lineage of Vaccines 3. Second Lineage of Vaccines 3. New
Gene Vaccine Dr. Sina Soleimani Human Viral Vaccines Quality Control Laboratory (HVVQC) Titles 1. A short Introduction of Vaccine History 2. First Lineage of Vaccines 3. Second Lineage of Vaccines 3. New
Molecular Graphics Perspective of Protein Structure and Function
 Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Identification of Mutation(s) in. Associated with Neutralization Resistance. Miah Blomquist
 Identification of Mutation(s) in the HIV 1 gp41 Subunit Associated with Neutralization Resistance Miah Blomquist What is HIV 1? HIV-1 is an epidemic that affects over 34 million people worldwide. HIV-1
Identification of Mutation(s) in the HIV 1 gp41 Subunit Associated with Neutralization Resistance Miah Blomquist What is HIV 1? HIV-1 is an epidemic that affects over 34 million people worldwide. HIV-1
Section 1 Proteins and Proteomics
 Section 1 Proteins and Proteomics Learning Objectives At the end of this assignment, you should be able to: 1. Draw the chemical structure of an amino acid and small peptide. 2. Describe the difference
Section 1 Proteins and Proteomics Learning Objectives At the end of this assignment, you should be able to: 1. Draw the chemical structure of an amino acid and small peptide. 2. Describe the difference
SUPPLEMENTARY INFORMATION. An orthogonal ribosome-trnas pair by the engineering of
 SUPPLEMENTARY INFORMATION An orthogonal ribosome-trnas pair by the engineering of peptidyl transferase center Naohiro Terasaka 1 *, Gosuke Hayashi 1 *, Takayuki Katoh 1, and Hiroaki Suga 1,2 1 Department
SUPPLEMENTARY INFORMATION An orthogonal ribosome-trnas pair by the engineering of peptidyl transferase center Naohiro Terasaka 1 *, Gosuke Hayashi 1 *, Takayuki Katoh 1, and Hiroaki Suga 1,2 1 Department
Supporting Information Parsimonious Charge Deconvolution for Native Mass Spectrometry
 Supporting Information Parsimonious Charge Deconvolution for Native Mass Spectrometry Marshall Bern* 1, Tomislav Caval 2, Yong J. Kil 1, Wilfred Tang 1, Christopher Becker 1, Eric Carlson 1, Doron Kletter
Supporting Information Parsimonious Charge Deconvolution for Native Mass Spectrometry Marshall Bern* 1, Tomislav Caval 2, Yong J. Kil 1, Wilfred Tang 1, Christopher Becker 1, Eric Carlson 1, Doron Kletter
Supplementary Information. Amino group carrier protein-mediated secondary metabolite biosynthesis in Streptomyces
 Supplementary Information Amino group carrier protein-mediated secondary metabolite biosynthesis in Streptomyces Fumihito Hasebe 1, Kenichi Matsuda 1, Taro Shiraishi 1, Yushi Futamura 2, Takeshi Nakano
Supplementary Information Amino group carrier protein-mediated secondary metabolite biosynthesis in Streptomyces Fumihito Hasebe 1, Kenichi Matsuda 1, Taro Shiraishi 1, Yushi Futamura 2, Takeshi Nakano
Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance
 Bruker Daltonics Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance The new ultraflextreme exceeds all current expectations of MALDI-TOF/TOF technology: A proprietary khz smartbeam-ii TM MALDI
Bruker Daltonics Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance The new ultraflextreme exceeds all current expectations of MALDI-TOF/TOF technology: A proprietary khz smartbeam-ii TM MALDI
Post-PKS tailoring steps of the spiramycin. macrolactone ring in Streptomyces ambofaciens
 Post-PKS tailoring steps of the spiramycin macrolactone ring in Streptomyces ambofaciens Hoang-Chuong NGUYEN, Emmanuelle DARBN, Robert THAI, Jean-Luc PERNDET, Sylvie LAUTRU SUPPLEMENTARY DATA 1 Figure
Post-PKS tailoring steps of the spiramycin macrolactone ring in Streptomyces ambofaciens Hoang-Chuong NGUYEN, Emmanuelle DARBN, Robert THAI, Jean-Luc PERNDET, Sylvie LAUTRU SUPPLEMENTARY DATA 1 Figure
LAB 4 Macromolecules
 LAB 4 Macromolecules Overview In addition to water and minerals, living things contain a variety of organic molecules. Most of the organic molecules in living organisms are of 4 basic types: carbohydrate,
LAB 4 Macromolecules Overview In addition to water and minerals, living things contain a variety of organic molecules. Most of the organic molecules in living organisms are of 4 basic types: carbohydrate,
Luminescent platforms for monitoring changes in the solubility of amylin and huntingtin in living cells
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2016 Contents Supporting Information Luminescent platforms for monitoring changes in the
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2016 Contents Supporting Information Luminescent platforms for monitoring changes in the
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL
 Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
Biological Mass spectrometry in Protein Chemistry
 Biological Mass spectrometry in Protein Chemistry Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi MASS SPECTROMETRY is an analytical technique that identifies the chemical composition of
Biological Mass spectrometry in Protein Chemistry Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi MASS SPECTROMETRY is an analytical technique that identifies the chemical composition of
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2988 Supplementary Figure 1 Kif7 L130P encodes a stable protein that does not localize to cilia tips. (a) Immunoblot with KIF7 antibody in cell lysates of wild-type, Kif7 L130P and Kif7
DOI: 10.1038/ncb2988 Supplementary Figure 1 Kif7 L130P encodes a stable protein that does not localize to cilia tips. (a) Immunoblot with KIF7 antibody in cell lysates of wild-type, Kif7 L130P and Kif7
Allergen of Japanese Cedar Pollen, Cryj2, in Escherichia coli
 Chaperone Coexpression Plasmids: Differential and Synergistic Roles of DnaK-DnaJ-GrpE and GroELGroES in Assisting Folding of an Allergen of Japanese Cedar Pollen, Cryj2, in Escherichia coli K. Nishihara
Chaperone Coexpression Plasmids: Differential and Synergistic Roles of DnaK-DnaJ-GrpE and GroELGroES in Assisting Folding of an Allergen of Japanese Cedar Pollen, Cryj2, in Escherichia coli K. Nishihara
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction
 Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
An Exploration to Determine if Fab Molecules are Efficacious in Neutralizing Influenza H1 and H3 Subtypes. Nick Poulton June September 2012
 An Exploration to Determine if Fab Molecules are Efficacious in Neutralizing Influenza H1 and H3 Subtypes Nick Poulton June 2012- September 2012 Epidemiology of Influenza Infection Causes between 250,000
An Exploration to Determine if Fab Molecules are Efficacious in Neutralizing Influenza H1 and H3 Subtypes Nick Poulton June 2012- September 2012 Epidemiology of Influenza Infection Causes between 250,000
PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland
 AD Award Number: DAMD17-02-1-0427 TITLE: Energetics and structure prediction of the network of homo- and hetero- Oligomers Formed by the Transmembrane Domains of the ErbBReceptor Family of Proteins PRINCIPAL
AD Award Number: DAMD17-02-1-0427 TITLE: Energetics and structure prediction of the network of homo- and hetero- Oligomers Formed by the Transmembrane Domains of the ErbBReceptor Family of Proteins PRINCIPAL
Cellular Neurobiology BIPN140. 1st Midterm Exam October 18 th, Tuesday Material covered: Lectures 1-6 & Reading
 Cellular Neurobiology BIPN140 1st Midterm Exam October 18 th, Tuesday Material covered: Lectures 1-6 & Reading Review session October 17 th 3500 Pacitic Hall, 6-8 pm (access code is 127895) Come with questions!
Cellular Neurobiology BIPN140 1st Midterm Exam October 18 th, Tuesday Material covered: Lectures 1-6 & Reading Review session October 17 th 3500 Pacitic Hall, 6-8 pm (access code is 127895) Come with questions!
Double charge of 33kD peak A1 A2 B1 B2 M2+ M/z. ABRF Proteomics Research Group - Qualitative Proteomics Study Identifier Number 14146
 Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
SUPPLEMENTARY INFORMATION
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2919 Direct observation of the influence of cardiolipin and antibiotics on lipid II binding to MurJ Jani
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2919 Direct observation of the influence of cardiolipin and antibiotics on lipid II binding to MurJ Jani
1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled
 Protein Targeting Objectives 1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled As a protein is being synthesized, decisions
Protein Targeting Objectives 1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled As a protein is being synthesized, decisions
Lecture 3. Tandem MS & Protein Sequencing
 Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
MALDI-TOF analysis of whole blood: its usefulness and potential in the assessment of HbA1c levels
 MALDI-TOF analysis of whole blood: its usefulness and potential in the assessment of HbA1c levels Jane Y. Yang, David A. Herold Department of Pathology, University of California San Diego, 9500 Gilman
MALDI-TOF analysis of whole blood: its usefulness and potential in the assessment of HbA1c levels Jane Y. Yang, David A. Herold Department of Pathology, University of California San Diego, 9500 Gilman
Peptide sequencing using chemically assisted fragmentation (CAF) and Ettan MALDI-ToF Pro mass spectrometry
 application note Ett MALDI-ToF Pro Peptide sequencing using chemically assisted fragmentation (CAF) d Ett MALDI-ToF Pro mass spectrometry Key words: MALDI-ToF, PSD, peptide sequencing, chemically assisted
application note Ett MALDI-ToF Pro Peptide sequencing using chemically assisted fragmentation (CAF) d Ett MALDI-ToF Pro mass spectrometry Key words: MALDI-ToF, PSD, peptide sequencing, chemically assisted
Organic Chemistry - Problem Drill 23: Amino Acids, Peptides, and Proteins
 rganic Chemistry - Problem Drill 23: Amino Acids, Peptides, and Proteins No. 1 of 10 1. Which amino acid does not contain a chiral center? (A) Serine (B) Proline (C) Alanine (D) Phenylalanine (E) Glycine
rganic Chemistry - Problem Drill 23: Amino Acids, Peptides, and Proteins No. 1 of 10 1. Which amino acid does not contain a chiral center? (A) Serine (B) Proline (C) Alanine (D) Phenylalanine (E) Glycine
Supporting Information. Lysine Propionylation to Boost Proteome Sequence. Coverage and Enable a Silent SILAC Strategy for
 Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Crystallization-grade After D After V3 cocktail. Time (s) Time (s) Time (s) Time (s) Time (s) Time (s)
 Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Proteins? Protein function. Protein folding. Protein folding diseases. Protein interactions. Macromolecular assemblies. The end product of Genes
 Proteins? Protein function Protein folding Protein folding diseases Protein interactions Macromolecular assemblies The end product of Genes Protein Unfolding DOD Acid Catalysis DOD HDOD + N H N D C N C
Proteins? Protein function Protein folding Protein folding diseases Protein interactions Macromolecular assemblies The end product of Genes Protein Unfolding DOD Acid Catalysis DOD HDOD + N H N D C N C
Study of different types of ubiquitination
 Study of different types of ubiquitination Rudi Beyaert (rudi.beyaert@irc.vib-ugent.be) VIB UGent Center for Inflammation Research Ghent, Belgium VIB Training Novel Proteomics Tools: Identifying PTMs October
Study of different types of ubiquitination Rudi Beyaert (rudi.beyaert@irc.vib-ugent.be) VIB UGent Center for Inflammation Research Ghent, Belgium VIB Training Novel Proteomics Tools: Identifying PTMs October
Protein Modeling Event
 Protein Modeling Event School Name: School Number: Team Member 1: Team Member 2: : Pre-Build Score: On-Site Build Score: Test Score: Tie Breaker: Total: Final Rank: Part I: Pre-Build (40% of total score)
Protein Modeling Event School Name: School Number: Team Member 1: Team Member 2: : Pre-Build Score: On-Site Build Score: Test Score: Tie Breaker: Total: Final Rank: Part I: Pre-Build (40% of total score)
Chemistry 135, First Exam. September 23, Chem 135, Exam 1 SID:
 Chemistry 135, First Exam September 23, 2015 This exam will be worth 15% of your overall grade. Please read all instructions/questions carefully and provide answers in the space provided. There should
Chemistry 135, First Exam September 23, 2015 This exam will be worth 15% of your overall grade. Please read all instructions/questions carefully and provide answers in the space provided. There should
Amino Acids and Proteins (2) Professor Dr. Raid M. H. Al-Salih
 Amino Acids and Proteins (2) Professor Dr. Raid M. H. Al-Salih 1 Some important biologically active peptides 2 Proteins The word protein is derived from Greek word, proteios which means primary. As the
Amino Acids and Proteins (2) Professor Dr. Raid M. H. Al-Salih 1 Some important biologically active peptides 2 Proteins The word protein is derived from Greek word, proteios which means primary. As the
SUPPLEMENTARY INFORMATION. Single-molecule site-specific detection of protein phosphorylation with a nanopore
 SUPPLEMENTRY INFORMTION Single-molecule site-specific detection of protein phosphorylation with a nanopore Christian B. Rosen 1,2*, David Rodriguez-Larrea 1* and Hagan Bayley 1 Department of Chemistry,
SUPPLEMENTRY INFORMTION Single-molecule site-specific detection of protein phosphorylation with a nanopore Christian B. Rosen 1,2*, David Rodriguez-Larrea 1* and Hagan Bayley 1 Department of Chemistry,
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11700 Figure 1: RIP3 as a potential Sirt2 interacting protein. Transfected Flag-tagged Sirt2 was immunoprecipitated from cells and eluted from the Sepharose beads using Flag peptide.
doi:10.1038/nature11700 Figure 1: RIP3 as a potential Sirt2 interacting protein. Transfected Flag-tagged Sirt2 was immunoprecipitated from cells and eluted from the Sepharose beads using Flag peptide.
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplemental Material for. Figure S1. Identification of TetR responsive promoters in F. novicida and E. coli.
 Supplemental Material for Synthetic promoters functional in Francisella novicida and Escherichia coli Ralph L. McWhinnie and Francis E. Nano Department of Biochemistry and Microbiology, University of Victoria,
Supplemental Material for Synthetic promoters functional in Francisella novicida and Escherichia coli Ralph L. McWhinnie and Francis E. Nano Department of Biochemistry and Microbiology, University of Victoria,
Enhanced pinocembrin production in Escherichia coli by regulating cinnamic acid metabolism
 1 Enhanced pinocembrin production in Escherichia coli by regulating cinnamic acid metabolism 2 Weijia Cao 1,2, Weichao Ma 1,2,3, Xin wang 1,2, Bowen Zhang 1,2, Xun Cao 1,2, Kequan Chen 1,2*, Yan Li 1,2
1 Enhanced pinocembrin production in Escherichia coli by regulating cinnamic acid metabolism 2 Weijia Cao 1,2, Weichao Ma 1,2,3, Xin wang 1,2, Bowen Zhang 1,2, Xun Cao 1,2, Kequan Chen 1,2*, Yan Li 1,2
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation
 Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Near-infrared fluorescent proteins
 nature methods Near-infrared fluorescent proteins Dmitry Shcherbo, Irina I Shemiakina, Anastasiya V Ryabova, Kathryn E Luker, Bradley T Schmidt, Ekaterina A Souslova, Tatiana V Gorodnicheva, Lydia Strukova,
nature methods Near-infrared fluorescent proteins Dmitry Shcherbo, Irina I Shemiakina, Anastasiya V Ryabova, Kathryn E Luker, Bradley T Schmidt, Ekaterina A Souslova, Tatiana V Gorodnicheva, Lydia Strukova,
Structural biology of viruses
 Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Sensing Mechanism of Globin-coupled Oxygen Sensor AfGcHK
 Sensing Mechanism of Globin-coupled Oxygen Sensor AfGcHK 1 Two-component signal transduction system (1) Signal redox, temperature, osmolarity etc. sensor (2) Autophospholylation ATP HK His - P ADP (HK)
Sensing Mechanism of Globin-coupled Oxygen Sensor AfGcHK 1 Two-component signal transduction system (1) Signal redox, temperature, osmolarity etc. sensor (2) Autophospholylation ATP HK His - P ADP (HK)
Figure S1 Time-dependent down-modulation of HER3 by EZN No Treatment. EZN-3920, 2 μm. Time, h
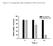 Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
Unveiling transient protein-protein interactions that modulate inhibition of alpha-synuclein aggregation
 Supplementary information Unveiling transient protein-protein interactions that modulate inhibition of alpha-synuclein aggregation by beta-synuclein, a pre-synaptic protein that co-localizes with alpha-synuclein.
Supplementary information Unveiling transient protein-protein interactions that modulate inhibition of alpha-synuclein aggregation by beta-synuclein, a pre-synaptic protein that co-localizes with alpha-synuclein.
ESI and MALDI Mass Spectrometry. S. Sankararaman Department of Chemistry Indian Institute of Technology Madras Chennai
 ESI and MALDI Mass Spectrometry S. Sankararaman Department of Chemistry Indian Institute of Technology Madras Chennai 600036 sanka@iitm.ac.in MODULE 22 ESI-Q MASS SPECTROMETRY - examples Mol. Wt = 1140
ESI and MALDI Mass Spectrometry S. Sankararaman Department of Chemistry Indian Institute of Technology Madras Chennai 600036 sanka@iitm.ac.in MODULE 22 ESI-Q MASS SPECTROMETRY - examples Mol. Wt = 1140
Ch 07. Microbial Metabolism
 Ch 07 Microbial Metabolism SLOs Differentiate between metabolism, catabolism, and anabolism. Fully describe the structure and function of enzymes. Differentiate between constitutive and regulated enzymes.
Ch 07 Microbial Metabolism SLOs Differentiate between metabolism, catabolism, and anabolism. Fully describe the structure and function of enzymes. Differentiate between constitutive and regulated enzymes.
Chymotrypsin Lecture. Aims: to understand (1) the catalytic strategies used by enzymes and (2) the mechanism of chymotrypsin
 Chymotrypsin Lecture Aims: to understand (1) the catalytic strategies used by enzymes and (2) the mechanism of chymotrypsin What s so great about enzymes? They accomplish large rate accelerations (10 10-10
Chymotrypsin Lecture Aims: to understand (1) the catalytic strategies used by enzymes and (2) the mechanism of chymotrypsin What s so great about enzymes? They accomplish large rate accelerations (10 10-10
Transient Ribosomal Attenuation Coordinates Protein Synthesis and Co-translational Folding
 SUPPLEMENTARY INFORMATION: Transient Ribosomal Attenuation Coordinates Protein Synthesis and Co-translational Folding Gong Zhang 1,2, Magdalena Hubalewska 1 & Zoya Ignatova 1,2 1 Department of Cellular
SUPPLEMENTARY INFORMATION: Transient Ribosomal Attenuation Coordinates Protein Synthesis and Co-translational Folding Gong Zhang 1,2, Magdalena Hubalewska 1 & Zoya Ignatova 1,2 1 Department of Cellular
Study of Amino Acids in DDGS
 Study of Amino Acids in DDGS Y. Zhang, J. V. Simpson and B. A. Wrenn National Corn-to-Ethanol Research Center Edwardsville, IL 62025 Hans Stein University of Illinois Urbana Champaign Gerald C. Shurson
Study of Amino Acids in DDGS Y. Zhang, J. V. Simpson and B. A. Wrenn National Corn-to-Ethanol Research Center Edwardsville, IL 62025 Hans Stein University of Illinois Urbana Champaign Gerald C. Shurson
AA s are the building blocks of proteins
 Chamras Chemistry 106 Lecture otes Chapter 24: Amino Acids, Peptides, and Proteins General Formula: () n (') α-amino Acids: (n = 1) Example: Amino Acids and Proteins: Glycine Alanine Valine AA s are the
Chamras Chemistry 106 Lecture otes Chapter 24: Amino Acids, Peptides, and Proteins General Formula: () n (') α-amino Acids: (n = 1) Example: Amino Acids and Proteins: Glycine Alanine Valine AA s are the
CHEM-643 Biochemistry Mid-term Examination 8:00 10:00, Friday, 2 November 2007
 EM-643 Biochemistry Mid-term Examination 8:00 10:00, Friday, 2 November 2007 Name Dr.. White - Instructor There are 10 pages to this examination including this page. Write your name on each new page. Read
EM-643 Biochemistry Mid-term Examination 8:00 10:00, Friday, 2 November 2007 Name Dr.. White - Instructor There are 10 pages to this examination including this page. Write your name on each new page. Read
2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry
 Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Engineering of Ministry of Education, Institute of Molecular Science, Shanxi University, Taiyuan , China.
 Indicator approach to develop a chemosensor for the colorimetric sensing of thiol-containing in water and its application for the thiol detection in plasma Fang-Jun Huo, a Yu-Tao Yang, b Jing Su, b Yuan-Qiang
Indicator approach to develop a chemosensor for the colorimetric sensing of thiol-containing in water and its application for the thiol detection in plasma Fang-Jun Huo, a Yu-Tao Yang, b Jing Su, b Yuan-Qiang
Practice Problems 3. a. What is the name of the bond formed between two amino acids? Are these bonds free to rotate?
 Life Sciences 1a Practice Problems 3 1. Draw the oligopeptide for Ala-Phe-Gly-Thr-Asp. You do not need to indicate the stereochemistry of the sidechains. Denote with arrows the bonds formed between the
Life Sciences 1a Practice Problems 3 1. Draw the oligopeptide for Ala-Phe-Gly-Thr-Asp. You do not need to indicate the stereochemistry of the sidechains. Denote with arrows the bonds formed between the
Components of a Mass Spectrometer
 Components of a Mass Spectrometer Sample Introduction Inlet GC LC Direct Insertion (Syringe/Probe) Ionization Ion Separation Ion Detection Ion Source EI,CI,,, MALDI Mass Analyzer Under vacuum TOF, Quadrupole,
Components of a Mass Spectrometer Sample Introduction Inlet GC LC Direct Insertion (Syringe/Probe) Ionization Ion Separation Ion Detection Ion Source EI,CI,,, MALDI Mass Analyzer Under vacuum TOF, Quadrupole,
AND JEREMY LUBAN 1,2 *
 JOURNAL OF VIROLOGY, Oct. 1999, p. 8527 8540 Vol. 73, No. 10 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Human Immunodeficiency Virus Type 1 Gag Polyprotein
JOURNAL OF VIROLOGY, Oct. 1999, p. 8527 8540 Vol. 73, No. 10 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Human Immunodeficiency Virus Type 1 Gag Polyprotein
Insulin mrna to Protein Kit
 Insulin mrna to Protein Kit A 3DMD Paper BioInformatics and Mini-Toober Folding Activity Student Handout www.3dmoleculardesigns.com Insulin mrna to Protein Kit Contents Becoming Familiar with the Data...
Insulin mrna to Protein Kit A 3DMD Paper BioInformatics and Mini-Toober Folding Activity Student Handout www.3dmoleculardesigns.com Insulin mrna to Protein Kit Contents Becoming Familiar with the Data...
Genetic information flows from mrna to protein through the process of translation
 Genetic information flows from mrn to protein through the process of translation TYPES OF RN (RIBONUCLEIC CID) RN s job - protein synthesis (assembly of amino acids into proteins) Three main types: 1.
Genetic information flows from mrn to protein through the process of translation TYPES OF RN (RIBONUCLEIC CID) RN s job - protein synthesis (assembly of amino acids into proteins) Three main types: 1.
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Supplementary Information
 Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Biomolecular Mass Spectrometry
 Lipids ot different than other organic small molecules Carbohydrates Polymers of monosaccharides linked via glycosidic bonds (acetals/ ketals) many different combinationsvery interesting no time ucleic
Lipids ot different than other organic small molecules Carbohydrates Polymers of monosaccharides linked via glycosidic bonds (acetals/ ketals) many different combinationsvery interesting no time ucleic
Supporting Information
 Supporting Information Keller et al. 10.1073/pnas.1222607110 MB1 Ori puc18 Ori AscI (8713) pdad downstream flank pdad PpdaD SphI (6970) Msed1993 ApaI (10721) Apr ApaLI (9115) NcoI (8) PmeI (8796) PvuI
Supporting Information Keller et al. 10.1073/pnas.1222607110 MB1 Ori puc18 Ori AscI (8713) pdad downstream flank pdad PpdaD SphI (6970) Msed1993 ApaI (10721) Apr ApaLI (9115) NcoI (8) PmeI (8796) PvuI
Viral vaccines. Lec. 3 أ.د.فائزة عبد هللا مخلص
 Lec. 3 أ.د.فائزة عبد هللا مخلص Viral vaccines 0bjectives 1-Define active immunity. 2-Describe the methods used for the preparation of attenuated live & killed virus vaccines. 3- Comparison of Characteristics
Lec. 3 أ.د.فائزة عبد هللا مخلص Viral vaccines 0bjectives 1-Define active immunity. 2-Describe the methods used for the preparation of attenuated live & killed virus vaccines. 3- Comparison of Characteristics
Protein Identification and Phosphorylation Site Determination by de novo sequencing using PepFrag TM MALDI-Sequencing kit
 Application Note Tel: +82-54-223-2463 Fax : +82-54-223-2460 http://www.genomine.com venture ldg 306 Pohang techno park Pohang, kyungbuk, Korea(ROK) Protein Identification and Phosphorylation Site Determination
Application Note Tel: +82-54-223-2463 Fax : +82-54-223-2460 http://www.genomine.com venture ldg 306 Pohang techno park Pohang, kyungbuk, Korea(ROK) Protein Identification and Phosphorylation Site Determination
PROTEIN. By: Shamsul Azahari Zainal Badari Department of Resource Management and Consumer Studies Faculty of Human Ecology UPM
 PROTEIN By: Shamsul Azahari Zainal Badari Department of Resource Management and Consumer Studies Faculty of Human Ecology UPM OBJECTIVES OF THE LECTURE By the end of this lecture, student can: Define
PROTEIN By: Shamsul Azahari Zainal Badari Department of Resource Management and Consumer Studies Faculty of Human Ecology UPM OBJECTIVES OF THE LECTURE By the end of this lecture, student can: Define
PREDICTING THE EXPRESSION AND SOLUBILITY OF MEMBRANE PROTEINS
 PREDICTING THE EXPRESSION AND SOLUBILITY OF MEMBRANE PROTEINS Mark E. Dumont, Michael A. White, Kathy Clark, Elizabeth J. Grayhack, and Eric. M. Phizicky Unknowns in Membrane Protein Expression, and Solubilization
PREDICTING THE EXPRESSION AND SOLUBILITY OF MEMBRANE PROTEINS Mark E. Dumont, Michael A. White, Kathy Clark, Elizabeth J. Grayhack, and Eric. M. Phizicky Unknowns in Membrane Protein Expression, and Solubilization
Objective: You will be able to explain how the subcomponents of
 Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
H 2 C H 2 N C CH O N C CH 3 CH 2 H O. aspartame
 1 The addition of sucrose, table sugar, to food and drink has been linked to the increased risk of obesity and insulin resistance. Aspartame is used as an alternative to sugar. The structure of aspartame
1 The addition of sucrose, table sugar, to food and drink has been linked to the increased risk of obesity and insulin resistance. Aspartame is used as an alternative to sugar. The structure of aspartame
Cloning and Expression of Influenza H1N1 NS1 Protein in Escherichia Coli BL21
 Iran J Biotech. 2014 March; 12(1): e12625. Published online 2014 March 19. DOI: 10.5812/ijb.12625 Research Article Cloning and Expression of Influenza H1N1 NS1 Protein in Escherichia Coli BL21 Marzieh
Iran J Biotech. 2014 March; 12(1): e12625. Published online 2014 March 19. DOI: 10.5812/ijb.12625 Research Article Cloning and Expression of Influenza H1N1 NS1 Protein in Escherichia Coli BL21 Marzieh
7.012 Quiz 3 Answers
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
Biotechnology-Based Vaccines. Dr. Aws Alshamsan Department of Pharmaceutics Office: AA87 Tel:
 Biotechnology-Based Vaccines Dr. Aws Alshamsan Department of Pharmaceutics Office: AA87 Tel: 4677363 aalshamsan@ksu.edu.sa Objectives of this lecture By the end of this lecture you will be able to: 1.
Biotechnology-Based Vaccines Dr. Aws Alshamsan Department of Pharmaceutics Office: AA87 Tel: 4677363 aalshamsan@ksu.edu.sa Objectives of this lecture By the end of this lecture you will be able to: 1.
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
BCH Graduate Survey of Biochemistry
 BCH 5045 Graduate Survey of Biochemistry Instructor: Charles Guy Producer: Ron Thomas Director: Glen Graham Lecture 7 Slide sets available at: http://hort.ifas.ufl.edu/teach/guyweb/bch5045/index.html David
BCH 5045 Graduate Survey of Biochemistry Instructor: Charles Guy Producer: Ron Thomas Director: Glen Graham Lecture 7 Slide sets available at: http://hort.ifas.ufl.edu/teach/guyweb/bch5045/index.html David
Stable isotope labeled Media products
 www.isotope.com RESEARCH PRODUCTS Stable isotope labeled Media products Bacterial Cell Growth Insect Cell Growth Mammalian Cell Growth Yeast Cell Growth Minimal Media for Bacterial Cell Growth Spectra
www.isotope.com RESEARCH PRODUCTS Stable isotope labeled Media products Bacterial Cell Growth Insect Cell Growth Mammalian Cell Growth Yeast Cell Growth Minimal Media for Bacterial Cell Growth Spectra
Supporting Information. Chemoenzymatic Synthesis of Galectin Binding. Glycopolymers
 Supporting Information Chemoenzymatic Synthesis of Galectin Binding Glycopolymers Jessica H. Ennist, Henry R. Termuehlen, Samuel P. Bernhard, Mackenzie S. Fricke, and Mary J. Cloninger* Department of Chemistry
Supporting Information Chemoenzymatic Synthesis of Galectin Binding Glycopolymers Jessica H. Ennist, Henry R. Termuehlen, Samuel P. Bernhard, Mackenzie S. Fricke, and Mary J. Cloninger* Department of Chemistry
WHAT IS GENETIC ENGINEERING?
 WHAT IS GENETIC ENGINEERING? CONSIDER: Preview the title, subtitles, and illustrations found on pages A-5 through A-12 and then list the topics that you think will be covered in this reading. TREATING
WHAT IS GENETIC ENGINEERING? CONSIDER: Preview the title, subtitles, and illustrations found on pages A-5 through A-12 and then list the topics that you think will be covered in this reading. TREATING
Multiple Anti-Interferon Actions of the Influenza A Virus NS1 Protein
 JOURNAL OF VIROLOGY, July 2007, p. 7011 7021 Vol. 81, No. 13 0022-538X/07/$08.00 0 doi:10.1128/jvi.02581-06 Copyright 2007, American Society for Microbiology. All Rights Reserved. Multiple Anti-Interferon
JOURNAL OF VIROLOGY, July 2007, p. 7011 7021 Vol. 81, No. 13 0022-538X/07/$08.00 0 doi:10.1128/jvi.02581-06 Copyright 2007, American Society for Microbiology. All Rights Reserved. Multiple Anti-Interferon
Protein regulation Protein motion
 Lecture 13 Protein regulation Protein motion Antoine van Oijen BCMP201 Spring 2008 04/02 Section IV 04/09 Hands-on methods session / PS 4 due 1 Today s lecture 1) Mechanisms of protein regulation 2) Molecular
Lecture 13 Protein regulation Protein motion Antoine van Oijen BCMP201 Spring 2008 04/02 Section IV 04/09 Hands-on methods session / PS 4 due 1 Today s lecture 1) Mechanisms of protein regulation 2) Molecular
Characterization of Influenza Hemagglutinin Mutants for the Elucidation of Key Residues Effect on Activation
 University of Tennessee, Knoxville Trace: Tennessee Research and Creative Exchange University of Tennessee Honors Thesis Projects University of Tennessee Honors Program 5-2013 Characterization of Influenza
University of Tennessee, Knoxville Trace: Tennessee Research and Creative Exchange University of Tennessee Honors Thesis Projects University of Tennessee Honors Program 5-2013 Characterization of Influenza
APRIL 1, 2011 VOLUME 286 NUMBER 13 JOURNAL OF BIOLOGICAL CHEMISTRY 11415
 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 286, NO. 13, pp. 11415 11426, April 1, 2011 2011 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. Intensity of Deoxycytidine
THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 286, NO. 13, pp. 11415 11426, April 1, 2011 2011 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. Intensity of Deoxycytidine
Biology 2E- Zimmer Protein structure- amino acid kit
 Biology 2E- Zimmer Protein structure- amino acid kit Name: This activity will use a physical model to investigate protein shape and develop key concepts that govern how proteins fold into their final three-dimensional
Biology 2E- Zimmer Protein structure- amino acid kit Name: This activity will use a physical model to investigate protein shape and develop key concepts that govern how proteins fold into their final three-dimensional
The mechanism of patellamide macrocyclization revealed by study of the Prochloron sp PatG macrocyclase domain
 The mechanism of patellamide macrocyclization revealed by study of the Prochloron sp PatG macrocyclase domain Jesko Koehnke 1,2, Andrew Bent 1,2, Wael E. Houssen 2,3,4, David Zollman 1, Falk Morawitz 1,
The mechanism of patellamide macrocyclization revealed by study of the Prochloron sp PatG macrocyclase domain Jesko Koehnke 1,2, Andrew Bent 1,2, Wael E. Houssen 2,3,4, David Zollman 1, Falk Morawitz 1,
Trans-stereospecific polymerization of butadiene and random copolymerization with styrene using borohydrido rare earths / magnesium dialkyl catalysts
 Trans-stereospecific polymerization of butadiene and random copolymerization with styrene using borohydrido rare earths / magnesium dialkyl catalysts A. Ventura a, T. Chenal b,c,d,e, *, M. Bria b,f, F.
Trans-stereospecific polymerization of butadiene and random copolymerization with styrene using borohydrido rare earths / magnesium dialkyl catalysts A. Ventura a, T. Chenal b,c,d,e, *, M. Bria b,f, F.
Supporting Information. Enzymatic synthesis of electrospinnable poly(butylene succinate-co-dilinoleic. succinate) thermoplastic elastomer
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2017 Supporting Information Enzymatic synthesis of electrospinnable poly(butylene succinate-co-dilinoleic
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2017 Supporting Information Enzymatic synthesis of electrospinnable poly(butylene succinate-co-dilinoleic
Proteolytic processing at a novel cleavage site in the N-terminal region of the tomato ringspot nepovirus RNA-1-encoded polyprotein in vitro
 Journal of General Virology (2000), 81, 2771 2781. Printed in Great Britain... Proteolytic processing at a novel cleavage site in the N-terminal region of the tomato ringspot nepovirus RNA-1-encoded polyprotein
Journal of General Virology (2000), 81, 2771 2781. Printed in Great Britain... Proteolytic processing at a novel cleavage site in the N-terminal region of the tomato ringspot nepovirus RNA-1-encoded polyprotein
Cambridge International Examinations Cambridge Ordinary Level
 Cambridge International Examinations Cambridge Ordinary Level *9076754804* BIOLOGY 5090/21 Paper 2 Theory May/June 2016 1 hour 45 minutes Candidates answer on the Question Paper. No Additional Materials
Cambridge International Examinations Cambridge Ordinary Level *9076754804* BIOLOGY 5090/21 Paper 2 Theory May/June 2016 1 hour 45 minutes Candidates answer on the Question Paper. No Additional Materials
Comparison of mass spectrometers performances
 Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Chapter 8. Interaction between the phosphatidylinositol 3- kinase SH3 domain and a photocleavable cyclic peptide
 Interaction between the phosphatidylinositol 3- kinase SH3 domain and a photocleavable cyclic peptide 129 Abstract The interaction of the PI3K SH3 domain with a cyclic photocleavable peptide and the linear
Interaction between the phosphatidylinositol 3- kinase SH3 domain and a photocleavable cyclic peptide 129 Abstract The interaction of the PI3K SH3 domain with a cyclic photocleavable peptide and the linear
Re-configuring Yarrowia lipolytica lipogenesis platform towards free fatty acid and biofuel production
 Re-configuring Yarrowia lipolytica lipogenesis platform towards free fatty acid and biofuel production Yonghong Meng, Ali Abghari, Rishikesh Ghogare, Xiaochao Xiong, GuokunWang, and Shulin Chen Bioprocessing
Re-configuring Yarrowia lipolytica lipogenesis platform towards free fatty acid and biofuel production Yonghong Meng, Ali Abghari, Rishikesh Ghogare, Xiaochao Xiong, GuokunWang, and Shulin Chen Bioprocessing
The ubiquitin-proteasome system. in bone biology. University of Nottingham. ECTS PhD Training July 5 th Dr Rob Layfield
 The ubiquitin-proteasome system in bone biology Dr Rob Layfield University of Nottingham ECTS PhD Training July 5 th 2009 Images reproduced from: http://www.bostonbiochem.com/overview.php?prod=ubchains
The ubiquitin-proteasome system in bone biology Dr Rob Layfield University of Nottingham ECTS PhD Training July 5 th 2009 Images reproduced from: http://www.bostonbiochem.com/overview.php?prod=ubchains
