Supplementary Notes. 2. Nine cell surface markers, CD33, CD20, CD3, CD4, CD7, CD123, CD14, IgM, and HLA-DR,
|
|
|
- Cody Hancock
- 5 years ago
- Views:
Transcription
1 Supplementary Notes Supplementary Note 1: Mass-tag cellular multiplexing workflow First, the cellular state is preserved by formaldehyde cross-linking. Second, the cells are permeabilized with methanol, making intracellular thiol groups and epitopes accessible. Third, the cells are washed with phosphate buffered saline (PBS), and then labeled with a unique combination of lanthanide isotope masses to barcode each sample. Fourth, the cells are washed to remove unreacted mdota-lanthanide(iii) reagent and then pooled into a single tube for antibody staining and mass cytometry measurements. Supplementary Note 2: Time course analysis of PBMC signaling SPADE To identify PBMC populations and their signaling response, Spanning-tree Progression Analysis of Density-normalized Events (SPADE) 1, 2 was applied to the generated time course dataset. SPADE is a computational tool that hierarchically clusters high-dimensional single-cell data, and connects these clusters of cells by a minimum spanning tree for two-dimensional visualization 1, 2. Nine cell surface markers, CD33, CD20, CD3, CD4, CD7, CD123, CD14, IgM, and HLA-DR, were used to generate cell clusters from the time course dataset; minimum spanning trees colored according to the expression level of each marker are shown in Fig. 2b. In combination, the cell surface marker expression levels of these trees were used to define 14 immune cell populations within the PBMC cellular hierarchy (Fig. 2c). For example, the middle branch displayed high CD20 expression (Fig. 2b), specifying B cell identity, but only a subset of this branch shows IgM expression, allowing separation of naïve (IgM + ) and memory (IgM - ) B cell subsets (Fig. 2b-c). In addition to cell surface markers, the SPADE-generated minimum spanning tree can also be used to display the levels of the 14 measured signaling molecules over the entire 4-hour stimulation time course, revealing subpopulation-specific signaling states and the signaling network dynamics of each cell type and subpopulation (Fig. 2d-e; Supplementary Results 1). Supplementary Note 3: Time course analysis of PBMC signaling analysis of feedback regulation The quantitative measurement of signaling network dynamics can be exploited to identify timedependent phenomena such as feedback regulation. PMA/ionomycin stimulates protein kinase C (PKC) and other calcium-related signaling pathways, which in T cells activate several downstream arms of the T cell receptor (TCR) signaling network (Supplementary Fig. 6), including phosphorylation of ERK, p38, NF B, ZAP70, and S6. Among this group, S6 responded most rapidly to PMA/ionomycin stimulation with a 3-fold increase in phosphorylation after one minute. Levels of phosphorylated ERK, p38 and ZAP70 rose after 5 minutes, and 1
2 NF B showed induction after 15 minutes (Supplementary Fig. 7). Conversely, phosphorylation of AKT, PLC 2 and SLP76, all membrane-proximal proteins upon activation, decreased after 1 min (AKT) or 5 min (PLC 2, SLP76), indicating rapid negative feedback regulation of the TCR signaling pathway coinciding with the initial network induction via PKC (Supplementary Fig. 7). A Ca 2+ signaling dependent feedback loop inhibiting PLC 2 via sequestration by the nonphosphatase Sprouty was described recently in T cells 3 and might explain the observed changes by making the phosphorylation epitope inaccessible for the antibody. PLC 2 also interacts with the PKC-activated phosphatase SHP2 4, which alternatively might dephosphorylate and inactivate PLC 2 5, 6. A similar mechanism may also apply to SLP76, which is targeted by the PKC-dependent SHP1 phosphatase 7. AKT phosphorylation is likely down-regulated after PKC stimulation by an S6K-mediated feedback loop similar to that operating on IRS-1 8, 9. Supplementary Note 4 Jak2 Inhibitor III selectivity IL3 or GM-CSF-induced, JAK2-dependent STAT5 phosphorylation in monocytes was not inhibited by JAK2 inhibitor III (Supplementary Fig. 16a; Supplementary Fig. 16b, columns 4 and 7/rows 4 and 8, blue boxes), indicating a lack of JAK2 inhibition. The inhibitor s impact on IFN- stimulated STAT1 phosphorylation in B cells and monocytes (Supplementary Fig. 16a; Supplementary Fig. 16b, columns 13 and 14/row 6, red boxes / columns 4 and 7/row 6, red boxes) revealed little effect, indicating a lack of efficacy against both JAK1 and JAK2. Furthermore only a slight inhibition of phosphorylation on STAT5 in IL-2 stimulated T cells was observed (Supplementary Fig. 16a; Supplementary Fig. 16b, columns 10 and 11/row 7, green box), excluding JAK3 and JAK1 as the prime inhibitor targets. However, when TYK2 is recruited to the receptor by IFN- or G-CSF, inhibition of phosphorylation on STAT5 and STAT3 in T cells and CD14 + HLA-DR mid, and CD14 - HLA-DR mid monocytes (Supplementary Fig. 16a; Supplementary Fig. 16b, column 10 and 11/row 5, grey box / columns 4 and 7/rows 3 and 5, yellow boxes) was observed. Supplementary Note 5 Jak3 Inhibitor VI selectivity Here no effect on JAK2-dependent STAT5 phosphorylation levels in monocytes activated by GM-CSF or IL-3 (Supplementary Fig. 17a; Supplementary Fig. 17b, columns 4 and 7/row 4, yellow boxes / columns 4 and 7/row 8, red boxes) or IL-3, thereby excluding JAK2 as the primary target. In contrast, in the presence of JAK3 inhibitor VI, IL2-activated STAT5 phosphorylation was inhibited in T cells, NK cells CD14 - surface neg. cells (Supplementary Fig. 17a; Supplementary Fig. 17b, columns 10 and 11/row 7, blue boxes / column 12/row 7, blue box / columns 1 and 2/row 7, blue box), suggesting inhibition of JAK1, JAK3, or both. To test whether the inhibitor was indeed specific for JAK3 and to exclude a contribution to signaling by 2
3 JAK1, IFN- stimulated cells were further analyzed. Here, JAK3 inhibitor VI inhibited phosphorylation of STAT1 in B cells (Supplementary Fig. 17a; Supplementary Fig. 17b, columns 13 and 14/row 6, green box) and of STAT1 and STAT5 in monocytes (Supplementary Fig. 17a; Supplementary Fig. 17b, columns 4 and 7/row 6, green boxes). These phosphorylations were induced by JAK1 and JAK2, but because JAK2 was excluded as a target the observed inhibitory effects must be due to JAK1 inhibition. JAK1/TYK2 phosphorylation of STAT1, STAT3, and STAT5 after IFN- stimulation was also inhibited in various cell types (Supplementary Fig. 17a; Supplementary Fig. 17b, row 5, grey box), further evidencing a lack of pure JAK3 specificity. Comparison with the inhibitors in vitro profile shows that this compound is 4-fold more potent against JAK3 than TYK2 and ~12-fold more compared to JAK1 and JAK2. While the inhibition of TYK2 agrees with our analysis, the low inhibition of JAK1 does not agree with our findings. Supplementary Fig. 17b shows that phosphorylation of other signaling molecules was also inhibited (e.g., SYK, PLC 2; Supplementary Fig. 17b, rows 1 and 3). These data indicate that the JAK3 inhibitor VI also inhibits JAK1/TYK2 and kinases outside the JAK-STAT signaling pathway, in agreement with the broad in vitro inhibition profile observed by Anastassiadis et all 10. Supplementary Note 6 Potent inhibition by Go-6983 (CAS # ) 11, a broad spectrum PKC inhibitor 10, of phosphorylation on S6 in most cell types, including B cells, after PKC activation with PMA/ionomycin was visible (Fig. 5b, column 7/row 11, grey box; Supplementary Fig. 18; Supplementary Fig. 19, column 7, green box; Supplementary Results 6). The mtor inhibitor rapamycin and the PI3K inhibitor GDC-0941 (CAS # ) 12, 13 lacked inhibition on the same phosphorylation site, if B cells were stimulated with PMA/ionomycin (Fig. 5b, red boxes; Supplementary Fig. 19, column 17/rows 13 and 14, blue box; Supplementary Fig. 19, column 6/rows 13 and 14, red box; Supplementary Fig. 20), which is in agreement with a model, in which PKC contributes to phosphorylation on S6 via the RAF-ERK-p90RSK signaling branch, independently of the PI3K-AKT-mTOR-p70S6K signaling pathway (Supplementary Fig. 6). After BCR/FcR-XL, rapamycin (Fig. 5b, column 17/row 2, pink box) and GDC-0941 (Fig. 5b, column 6/row 2, blue box) both inhibit S6 phosphorylation in B cells (Supplementary Fig. 20), achieving a 50% and 80% inhibition, respectively, again in agreement with the current model in which rapamycin only inhibits S6 phosphorylation via mtor-p70s6k, but GDC-0941 inhibits the PKC- RAF-ERK-p90RSK pathway in addition to PI3K-AKT-mTOR-p70S6K, therefore achieving greater percent inhibition of S6 ps235/6 phosphorylation (Supplementary Fig. 6). On the contrary, Go-6983 nearly completely abolished phosphorylation of S6 protein after BCR/FcR-XL in B cells (Fig. 5b, column 7/row 2, green box; Supplementary Fig. 18), although the current 3
4 model only assumes a partial contribution of PKC in parallel to the PI3K-AKT-mTOR-p70S6K pathway (Supplementary Fig. 6), indicating additional targets besides PKC. Indeed, recent in vitro data suggest 10, that Go-6983 also potently inhibits p70s6k and p90rsk, both kinases known to phosphorylate ps235/6 on S6, giving a potential explanation for this observation. Supplementary Note 7 - Cell type selectivity Imatinib 14, 15, an inhibitor of the constitutively active tyrosine kinase BCR-ABL used to treat patients with CML 14, 15, affected in CD14 + HLA-DR mid monocytes compared to other cell types consistently phosphorylation of SHP2, SYK, and PLC 2 under most conditions (Supplementary Fig. 23, column 7, green box). This might be due to inhibition of Lyn kinase by imatinib which was shown in vitro 10, 13, and suggesting why imatinib inhibits normal monocyte development and function 16, 17 and causes cytopenia in non-tumor blood cells at higher concentrations 18. The c-jun N-terminal kinase (JNK) inhibitor SP had a distinct bias toward inhibition of phosphorylation of ZAP70, a critical kinase for T cell receptor signaling 20, in CD4 + T cells compared to CD8 + T cells under most conditions studied (Supplementary Fig. 24, column 11, green box). SP impairs CD4 + T cell function, so far assumed via the inhibition of JNK 19. However, these results indicate inhibition of additional kinase targets involved in TCR activation contributing to this observation. For Streptoningrin, in CD14 - HLA-DR mid and CD14 + HLA-DR mid monocytes, reproducible inhibition was observed of SYK, PLC and BLNK phosphorylation in the CD14 + HLA-DR mid cells (Supplementary Fig. 25a; Supplementary Fig. 25d, columns 4 and 7, green boxes) whereas in CD14 - HLA-DR mid monocytes inhibition of these phosphorylation sites was only detected after PMA/ionomycin stimulation (Supplementary Fig. 25b; Supplementary Fig. 25d, columns 4 and 7/row 11, red boxes). Supplementary Note 8 limitation of the method Several points have to be kept in mind when using MCB to analyze an experimental system: mdota reagent levels have to be titrated for each sample type, to ensure maximum separation of positively and negatively labeled cell populations for gating and deconvolution (Supplementary Fig. 3). Factors influencing the mdota cell labeling levels include cell number, availability of free thiol groups and cell size. The latter can be influenced by long-term change in growth condition, long-term drug treatment (e.g. using growth signaling inhibitors such as GDC- 0941, rapamycin or AKTi-1/2), induction of apoptosis or osmotic shock. Potential solutions to this caveat include cell type gating before deconvolution (as done in this study) and normalization of barcode channels with an orthogonal measure of cell size. 4
5 Furthermore, in our experiments only phosphorylation but not protein abundance levels were quantified. While this usually is not a limitation in short term stimulation experiments as constant protein levels can be assumed, it is possible that in the long-term experiments (such as the four hour time course experiment), changes in phosphorylation abundance might be due to changes in protein levels. In addition, many proteins function as signaling relays (e.g. p53, AKT...) with a multitude of phosphorylation sites which act in a combinatorial manner resulting in an enormous number of protein signaling states, which are not completely described if only a single phosphorylation site is quantified per protein as in our experiments. Finally, the approach taken here does not reveal differences between multi-target drug effects and indirect adaptive changes of the signaling network occurring over time. Here the generation of high-resolution time course data beginning with the drug treatment could differentiate the effects that are directly caused by a drug from those due to the adaptation of the cellular signaling network. Finally, as any method relying on antibodies, MCB is fully dependent on their quality and availability. Many signaling molecules of great biological interest exist at low copy number in the cell, and can be difficult to quantify accurately at the single cell level, especially with antibodies that display poor binding characteristics. For this study, multiple antibody clones were tested, and in some cases antibodies with high background binding or low pathway-induced shift in binding were chosen because they were the best available choice to measure a pathway of high biological interest, such as AKT activation. The antibody staining panel used in this study already provides a broad overview of cell signaling state, and this analysis of cell signaling by MCB will only improve as more measurement channels become available through improved antibody-labeling chemistries, and as more antibodies become commercially available, particularly as more antibodies are developed specifically for intracellular flow cytometry. Despite these limitations, MCB allows a comprehensive study of signaling networks and cellular responses in highly complex cell mixtures with unprecedented throughput. 5
6 Supplementary Tables Cell type Donor 1 Donor 2 Donor 3 Donor 4 Donor 5 Donor 6 Donor 7 Donor 8 CD4 + T cells 24% 30% 19% 21% 28% 34% 15% 26% CD8 + T cells 22% 21% 10% 11% 13% 13% 16% 20% CD14 + HLADR + Monocytes 4% 7% 12% 8% 6% 5% 5% 6% CD14 + HLADR - Monocytes 1% 2% 4% 4% 1% 1% 3% 3% CD14 - HLADR + Monocytes 8% 7% 6% 4% 8% 6% 3% 4% CD14 - HLADR - Monocytes 8% 9% 4% 5% 6% 7% 4% 4% Dendritic cells 1% 2% 2% 3% 3% 3% 2% 2% IgM + B cells 7% 4% 10% 7% 5% 7% 5% 4% IgM - B cells 1% 1% 1% 1% 1% 1% 2% 1% NK cells 11% 8% 13% 15% 10% 8% 21% 11% Surf - CD14 + cells 1% 1% 8% 9% 2% 2% 9% 6% Surf - CD14 - cells 12% 10% 10% 14% 18% 15% 16% 13% Supplementary Table 1. Cell type percentages from 8 different donors as determined by MCB. 6
7 Stimulus JAKs STATs phosphorylated IFN- JAK1 TYK2 STAT1 STAT3 STAT5 IFN- JAK1 JAK2 STAT1 STAT5 G-CSF JAK1 JAK2 TYK2 STAT1 STAT3 STAT5 GM-CSF JAK2 STAT5 IL-2 JAK1 JAK3 STAT1 STAT3 STAT5 IL-3 JAK2 STAT5 IL-12 JAK2 TYK2 STAT4 Supplementary Table 2: JAK-STAT relationships. Each row shows the JAK combination resulting from a given stimulus, and the STAT proteins phosphorylated 21, 22. 7
8 Inhibitor JAK1* JAK2 JAK3 TYK2* JAK (pan) inhibitor I % 1.8% 0.44% 1.5% JAK2 inhibitor III $ >10,000 >10,000 >10,000 >10,000 JAK3 inhibitor VI % 58% 4.8% 20.4% Ruxolitinib Ruxolitinib Tofacitinib Tofacitinib Tofacitinib 23 N/A Tofacitinib % 5.1% 1.9% 12% Lestauritinib SP VX VX N/A >10000 VX Supplementary Table 3: IC 50 values (nm or percent inhibition) of JAK inhibitors analyzed in this study as determined by in vitro kinase assays or by 10. (*) only JH1domain-catalytic domain results shown. $ Two different batches of JAK2 inhibitor III were tested. Inhibitors highlighted in grey/without reference were tested for this manuscript 10. 8
9 Supplementary Figures Tb159 Supplementary Figure 1. Time course analysis of cellular labeling using mdotalanthanide(iii) loaded with terbium 159. Unstimulated Stimulated Ir193 Yb174 Supplementary Figure 2. Slp76 induction by orthovanadate. Level of phosphorylation on Tyr- 696 on Slp76 before (left) and after (right) orthovanadate stimulation of K562 cells is shown. 9
10 Supplementary Figure 3. All seven channels used for barcoding are shown for a typical PBMC inhibitor experiment (AKTi-1/2 (CAS # )) analysis. The clear separation between positively (populations to the right) and unlabeled (populations to the left) cells allows a clean deconvolution of the barcoded cell populations. The cell events indicated outside the contour lines are below 2% of all cell events. Nevertheless, for the PBMC experiments separation between mdota-lanthanide(iii) positive and negative populations was reduced relative to that of K562 cells, likely due to cell size variability in the PBMC sample
11 pstat1 pstat1 pstat1 pstat1 pstat3 pstat3 pstat3 pstat5 pstat5 pstat5 Supplementary Figure 4. Time-resolved response of PBMCs to IFN- treatment. X-axis plots time in min, y-axis plots signal intensity of indicated marker. Blue dot and blue bars plot median and standard deviation, respectively. 11
12 Supplementary Figure 5. Histogram overlay of STAT5 induction over time after IFN- stimulation in T-cells. Only part of the CD4 - T cell population responds to the stimulus. 12
13 Supplementary Figure 6. Immune cell signaling network nodes and inhibitors studied. Signaling molecules on which a phosphorylation site was quantified in this study are indicated with a red circle. All inhibitors used in this study and their in vitro target or targets are shown in a cell type independent immune signaling network diagram. Many inhibitors (indicated by *) have additional known targets. General inhibitors without defined targets or targets outside the shown network are listed at the bottom. 13
14 perk p-p38 pzap70 pnf B ps6 pakt pslp76 pplc 2 Supplementary Figure 7. PMA/ionomycin-stimulated TCR signaling components. X-axis plots time in min, y-axis plots signal intensity of indicated marker. Blue dot and blue bars plot median and standard deviation, respectively. 14
15 p-p38 perk pnf B ps6 Supplementary Figure 8. Time-resolved induction of phosphorylation of different signaling proteins in monocytes after LPS stimulation. X-axis plots time in min, y-axis indicates signal intensity of indicated marker. Blue dot and blue bars plot median and standard deviation, respectively. 15
16 pstat3 pstat3 pstat3 pstat3 pstat5 pstat5 pstat5 pstat5 Supplementary Figure 9a. Time-resolved induction of phosphorylation on different signaling proteins and PBMC subtypes after LPS stimulation (left) and corresponding control (right). X- axis plots time in min, y-axis plots signal intensity of indicated marker. Blue dot and blue bars plot median and standard deviation, respectively. 16
17 pitk pitk pitk pitk pstat1 pstat1 Supplementary Figure 9b. Time-resolved induction of phosphorylation on different signaling proteins and PBMC subtypes after LPS stimulation (left) and corresponding control (right). X- axis plots time in min, y-axis plots signal intensity of indicated marker. Blue dot and blue bars plot median and standard deviation, respectively. 17
18 Supplementary Figure 10. PBMC signaling response comparison of 8 different donors in CD8 + T cells after IFN- stimulation. For each donor the control (left) and stimulation (right) condition are shown. 18
19 Supplementary Figure 11. PBMC signaling response comparison of 8 different donors in CD14 + HLA-DR + Monocytes after IFN- stimulation. For each donor the control (left) and stimulation (right) condition are shown. 19
20 IFN- / pstat1 IFN- / pstat1 IL-2 / pstat5 IL-3 / pstat5 G-CSF / pstat3 GM-CSF / pstat5 Supplementary Figure 12a. Effect of tofacenib on the indicated cell type, stimulus, and phosphorylation site combinations. X-axis plots dose in M, y-axis plots the signal intensity of indicated marker. Blue dot and blue bars plot median and standard deviation, respectively. 20
21 Supplementary Figure 12b. Effect of tofacinib inhibitor on indicated cell type, stimulus, and phosphorylation site combinations. 21
22 Supplementary Figure 13. Effect of ruxolitinib inhibitor on indicated cell type, stimulus, and phosphorylation site combinations. White circles indicate inhibition, but the cell number in the reference control was too low for determination of unstimulated control. 22
23 Supplementary Figure 14. Effect of lestauritinib inhibitor on indicated cell type, stimulus, and phosphorylation site combinations. 23
24 Supplementary Figure 15. Effect of Jak(pan) inhibitor on indicated cell type, stimulus and phosphorylation site combinations. 24
25 GM-CSF / pstat5 GM-CSF / pstat5 IL-3 / pstat5 IL-3 / pstat5 IFN- / pstat1 IFN- / pstat1 IL-2 / pstat5 IL-2 / pstat5 Supplementary Figure 16a. Continues next page. X-axis plots dose in M, y-axis plots signal intensity of indicated marker. Blue dot and blue bars plot median and standard deviation, respectively. 25
26 IFN- / pstat5 IFN- / pstat5 IFN- / pstat3 IFN- / pstat3 Supplementary Figure 16a. Effect of Jak2 inhibitor III on the indicated cell type, stimulus, and phosphorylation site combinations. X-axis plots dose in M, y-axis plots the signal intensity of indicated marker. Blue dot and blue bars plot median and standard deviation, respectively. 26
27 Supplementary Figure 16b. Effect of Jak2 inhibitor on indicated cell type, stimulus, and phosphorylation site combinations. 27
28 GM-CSF / pstat5 IL3 / pstat5 IL2 / pstat5 IL2 / pstat5 IL2 / pstat5 IFN- / pstat1 IFN- / pstat1 IFN- / pstat5 IFN- / pstat1 IFN- / pstat3 IFN- / pstat5 Supplementary Figure 17a. Analysis of Jak3 inhibitor VI specificity. X-axis plots dose in M, y- axis plots the signal intensity of indicated marker. Blue dot and blue bars plot median and standard deviation, respectively. 28
29 Supplementary Figure 17b. Effect of Jak3 inhibitor VI on indicated cell type, stimulus and phosphorylation site combinations. 29
30 PMA/Iono./pS6 PMA/Iono./pS6 BCR/FcR-XL/pS6 BCR/FcR-XL/pS6 Supplementary Figure 18. Go-6983 impact on ps6 phosphorylation after PMA/ionomycin or BCR-XL treatment. X-axis plots dose in M, y-axis indicates signal intensity of indicated marker. Blue dot and blue bars plot median and standard deviation, respectively. 30
31 Supplementary Figure 19. Signaling response of all PBMC subtypes to PMA/ionomycin stimulation for all analyzed inhibitors. 31
32 Rapamycin BCR/FcR-XL/pS6 PMA/Iono./pS6 GDC-0941 BCR/FcR-XL/pS6 PMA/Iono./pS6 Supplementary Figure 20. Impact of rapamycin and GDC-0941 on S6 phosphorylation after BCR-XL or PMA/ionomycin stimulation. X-axis plots dose in M, y-axis plots signal intensity of indicated marker. Blue dot and blue bars plot median and standard deviation, respectively. 32
33 Supplementary Figure 21. Overview of inhibitor impact on CD14 - HLA-DR mid monocytes under indicated stimulation conditions. Supplementary Figure 22. Overview of inhibitor impact on CD14 + HLA-DR mid monocytes under indicated stimulation conditions. 33
34 Supplementary Figure 23. Effect of imatinib on indicated cell type, stimulus, and phosphorylation site combinations. 34
35 Supplementary Figure 24. Effect of SP on indicated cell type, stimulus, and phosphorylation site combinations. 35
36 G-CSF / psyk G-CSF / psyk G-CSF / pplc 2 G-CSF / pplc 2 GM-CSF / pblnk GM-CSF / pblnk Supplementary Figure 25a. Overview of streptonigrin impact on indicated phosphorylation sites in CD14 + and CD14 - HLA-DR mid monocytes in indicated conditions. X-axis plots dose in M, y-axis plots the signal intensity of indicated marker. Blue dot and blue bars plot median and standard deviation, respectively.. 36
37 PMA-Iono. / psyk PMA-Iono. / psyk PMA-Iono. / pplc 2 PMA-Iono. / pplc 2 PMA-Iono. / pblnk PMA-Iono. / pblnk Supplementary Figure 25b. Overview of streptonigrin impact on indicated phosphorylation sites in CD14 + and CD14 - HLA-DR mid monocytes under indicated conditions. X-axis plots dose in M, y-axis plots the signal intensity of indicated marker. Blue dot and blue bars plot median and standard deviation, respectively. 37
38 G-CSF / ps6 G-CSF / ps6 IL-2 / pplc 2 IL-2 / pplc 2 G-CSF / pblnk G-CSF / pblnk IL-2 / pstat5 IL-2 / pstat5 Supplementary Figure 25c. Overview of streptonigrin impact on indicated phosphorylation sites in T and B cells under various conditions. X-axis plots dose in M, y-axis plots the signal intensity of indicated marker. Blue dot and blue bars plot median and standard deviation, respectively. 38
39 Supplementary Figure 25d. Effect of streptoningrin on indicated cell type, stimulus, and phosphorylation site combinations. 39
40 CD14 - HLA-DR high Monocytes CD14 + HLA-DR high Monocytes CD14 - HLA-DR mid Monocytes CD14 + HLA-DR mid Monocytes CD14 + HLA-DR low Monocytes CD14 - HLA-DR low Monocytes CD14 + Surface neg. Dendritic cells CD4 + T cells CD8 + T cells CD14 - Surface neg. NK cells IgM + B cells IgM - B cells Supplementary Figure 26. Clustergram of first 30 principle components of cell-type PCA analysis. 40
41 IFN- / pstat1 IFN- / pstat1 IFN- / pstat1 IFN- / pstat3 IFN- / pstat3 IFN- / pstat3 IFN- / pstat5 IFN- / pstat5 IFN- / pstat5 Supplementary Figure 27. Overview of SP effects on STAT signaling in indicated cell types after IFN- stimulation. X-axis plots dose in M, y-axis plots the signal intensity of indicated marker. Blue dot and blue bars plot median and standard deviation, respectively. 41
42 IFN- / pstat3 IFN- / pstat3 IFN- / pstat3 IFN- / pstat5 IFN- / pstat5 IFN- / pstat5 Supplementary Figure 28a. Overview of VX680 inhibition of STAT3 and STAT5 signaling in indicated cell types after IFN- stimulation. X-axis plots dose in M, y-axis plots the signal intensity of indicated marker. Blue dot and blue bars plot median and standard deviation, respectively. 42
43 Supplementary Figure 28b. Effect of VX680 on indicated cell type, stimulus, and phosphorylation site combinations. 43
44 Supplementary Figure 29. PBMC response to ruxolitinib (CAS # ) inhibition in 4 different donors. Donor 1 showed a broader inhibition profile compared to donor
45 Supplementary References 1. Bendall, S.C. et al. Single-cell mass cytometry of differential immune and drug responses across a human hematopoietic continuum. Science 332, (2011). 2. Qiu, P. et al. Extracting a cellular hierarchy from high-dimensional cytometry data with SPADE. Nat Biotechnol 29, (2011). 3. Akbulut, S. et al. Sprouty proteins inhibit receptor-mediated activation of phosphatidylinositol-specific phospholipase C. Mol Biol Cell 21, (2010). 4. Boudot, C. et al. Phosphatidylinositol 3-kinase regulates glycosylphosphatidylinositol hydrolysis through PLC-gamma(2) activation in erythropoietin-stimulated cells. Cell Signal 14, (2002). 5. Strack, V. et al. The Protein-tyrosine-phosphatase SHP2 is phosphorylated on serine residues 576 and 591 by protein kinase C isoforms alpha, beta 1, beta 2, and eta. Biochemistry 41, (2002). 6. Mussig, K. et al. Shp2 is required for protein kinase C-dependent phosphorylation of serine 307 in insulin receptor substrate-1. J Biol Chem 280, (2005). 7. Binstadt, B.A. et al. SLP-76 is a direct substrate of SHP-1 recruited to killer cell inhibitory receptors. J Biol Chem 273, (1998). 8. Shah, O.J., Wang, Z. & Hunter, T. Inappropriate activation of the TSC/Rheb/mTOR/S6K cassette induces IRS1/2 depletion, insulin resistance, and cell survival deficiencies. Curr Biol 14, (2004). 9. Harrington, L.S. et al. The TSC1-2 tumor suppressor controls insulin-pi3k signaling via regulation of IRS proteins. J Cell Biol 166, (2004). 10. Anastassiadis, T., Deacon, S.W., Devarajan, K., Ma, H. & Peterson, J.R. Comprehensive assay of kinase catalytic activity reveals features of kinase inhibitor selectivity. Nat Biotechnol 29, (2011). 11. Gschwendt, M. et al. Inhibition of protein kinase C mu by various inhibitors. Differentiation from protein kinase c isoenzymes. FEBS Lett 392, (1996). 12. Folkes, A.J. et al. The identification of 2-(1H-indazol-4-yl)-6-(4-methanesulfonylpiperazin-1-ylmethyl)-4-morpholin -4-yl-thieno[3,2-d]pyrimidine (GDC-0941) as a potent, selective, orally bioavailable inhibitor of class I PI3 kinase for the treatment of cancer. J Med Chem 51, (2008). 13. Davis, M.I. et al. Comprehensive analysis of kinase inhibitor selectivity. Nat Biotechnol 29, (2011). 14. Buchdunger, E. et al. Inhibition of the Abl protein-tyrosine kinase in vitro and in vivo by a 2-phenylaminopyrimidine derivative. Cancer Res 56, (1996). 15. Druker, B.J. et al. Effects of a selective inhibitor of the Abl tyrosine kinase on the growth of Bcr-Abl positive cells. Nat Med 2, (1996). 16. Dewar, A.L., Doherty, K.V., Hughes, T.P. & Lyons, A.B. Imatinib inhibits the functional capacity of cultured human monocytes. Immunol Cell Biol 83, (2005). 17. Dewar, A.L., Domaschenz, R.M., Doherty, K.V., Hughes, T.P. & Lyons, A.B. Imatinib inhibits the in vitro development of the monocyte/macrophage lineage from normal human bone marrow progenitors. Leukemia 17, (2003). 18. Sawyers, C.L. et al. Imatinib induces hematologic and cytogenetic responses in patients with chronic myelogenous leukemia in myeloid blast crisis: results of a phase II study. Blood 99, (2002). 19. Bennett, B.L. et al. SP600125, an anthrapyrazolone inhibitor of Jun N-terminal kinase. Proceedings of the National Academy of Sciences of the United States of America 98, (2001). 20. Chan, A.C., Iwashima, M., Turck, C.W. & Weiss, A. ZAP-70: a 70 kd protein-tyrosine kinase that associates with the TCR zeta chain. Cell 71, (1992). 21. Shuai, K. & Liu, B. Regulation of JAK-STAT signalling in the immune system. Nat Rev Immunol 3, (2003). 45
46 22. Rawlings, J.S., Rosler, K.M. & Harrison, D.A. The JAK/STAT signaling pathway. J Cell Sci 117, (2004). 23. Karaman, M.W. et al. A quantitative analysis of kinase inhibitor selectivity. Nat Biotechnol 26, (2008). 24. Krutzik, P.O. & Nolan, G.P. Fluorescent cell barcoding in flow cytometry allows highthroughput drug screening and signaling profiling. Nat Methods 3, (2006). 46
47 Supplementary Methods Kinase inhibitors The following list shows all inhibitors analyzed in this study. The highest concentration used in the dose-titrations and the companies from which they were purchased are shown. Inhibitor Concentration ( M) CAS number Bought from AKTi-1/ Chemdea BTK inhibitor III Calbiochem/EMD/Merck crassin (NCI DTP; index.html) dasatinib LC Laboratories GDC Chemdea Go Tocris H LC Laboratories IKK inhibitor X Calbiochem/EMD/Merck imatinib LC Laboratories JAK inhibitor I Calbiochem/EMD/Merck JAK2 inhibitor III Calbiochem/EMD/Merck JAK3 inhibitor VI Calbiochem/EMD/Merck LCK inhibitor Calbiochem/EMD/Merck lestauritinib LC Laboratories PP Calbiochem/EMD/Merck rapamycin LC Laboratories ruxolitinib (INCB018424) LC Laboratories SB LC Laboratories sorafenib LC Laboratories SP LC Laboratories staurosporine LC Laboratories streptoningrin Sigma sunitinib LC Laboratories SYK inhibitor IV Calbiochem/EMD/Merck tofacitinib (CP ) LC Laboratories U LC Laboratories VX LC Laboratories Kinase inhibitor stocks were prepared in DMSO. Supplementary Methods Table 1. Inhibitors used in this study. 1
48 96 well plate Short name Column 1 Sodium orthovanadate 2 IL-2 3 IL-3 4 IL-12 5 Reference 6 G-CSF 7 GM-CSF 8 BCR/FcR-XL 9 IFN- 10 IFN- 11 LPS 12 PMA/Ionomycin Supplementary Methods Table 2. Overview of the stimulations used for this study and their locations on the 96-well plate. Reference indicates no stimulus. 2
49 Isotope Antigen Clone Final concentration Supplier g/ml Cd110, 111, 112, 114 CD3 UCHT1 Invitrogen In115 CD45 HI30 2 Biolegend La139 BC Pr141 BC Nd142 NF B (ps529) K BD Nd144 p38 (pt180/py182) 36/ p38 2 BD Nd145 CD4 RPA-T4 3 Biolegend Nd146 BC Sm147 CD20 (intracellular) H1 3 BD Nd148 CD33 WM53 3 Biolegend Nd150 Stat5 (py694) 47 2 BD Eu151 CD123 9f5 3 BD Sm152 Akt (pt308) J BD Eu153 Stat1 (py701) 4a 3 BD Sm154 SHP2 (py580) polyclonal 2 CST Gd156 Zap70/Syk (py319/py352) 17a 1 BD Gd158 Stat3 (py705) 4 2 BD Tb159 BC Gd160 CD14 M5E2 3 Biolegend Dy164 Slp76 J141- (py128)/blnk (py 72) 1 BD Ho165 Er166 Er167 Er168 Tm169 BC pbtk (py551/py511)/pitk (py511) pplc (py759) Erk1/2 (pt202/ py204) BC 24a/BTK 2 BD K BD 20A 2 BD Er170 plat (py226) J BD Yb171 IgM G BD Yb172 S6 (ps235/ps236) N BD Yb174 HLA-DR L243 3 Biolegend Lu175 BC Yb176 CD7 M-T701 5 BD Supplementary Methods Table 3. Antibodies used in this study (BD is Becton Dickinson). 3
50 Supplementary Methods Figure 1. Strategy to determine fold-change and R2 cut-offs for each analyzed dose response curve. 4
Supplementary Figure 1. Gating strategy for leukemic blasts and normal developing B
 Supplementary Information Supplementary Figure 1. Gating strategy for leukemic blasts and normal developing B cells. (a) Gating strategy for lineage-negative blasts (B + Lin - cells), the starting population
Supplementary Information Supplementary Figure 1. Gating strategy for leukemic blasts and normal developing B cells. (a) Gating strategy for lineage-negative blasts (B + Lin - cells), the starting population
Can we classify cancer using cell signaling?
 Can we classify cancer using cell signaling? Central hypotheses (big ideas) Alterations to signaling genes would cause leukemic cells to react in an inappropriate or sensitized manner to environmental
Can we classify cancer using cell signaling? Central hypotheses (big ideas) Alterations to signaling genes would cause leukemic cells to react in an inappropriate or sensitized manner to environmental
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE
 Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
CyTOF workflow: differential discovery in high-throughput high-dimensional cytometry datasets!
 Workshop!! CyTOF workflow: differential discovery in high-throughput high-dimensional cytometry datasets! Malgorzata Nowicka! University of Zurich! BioC 2017: Where Software and Biology Connect! Boston,
Workshop!! CyTOF workflow: differential discovery in high-throughput high-dimensional cytometry datasets! Malgorzata Nowicka! University of Zurich! BioC 2017: Where Software and Biology Connect! Boston,
Categorical analysis of human T cell heterogeneity with One-SENSE
 1 2 3 4 Supplementary Information Categorical analysis of human T cell heterogeneity with One-SENSE 5 6 Running title: T cell Analysis by One-SENSE 7 8 9 1 11 12 13 14 15 16 17 18 19 2 21 22 23 24 25 26
1 2 3 4 Supplementary Information Categorical analysis of human T cell heterogeneity with One-SENSE 5 6 Running title: T cell Analysis by One-SENSE 7 8 9 1 11 12 13 14 15 16 17 18 19 2 21 22 23 24 25 26
Nature Immunology: doi: /ni Supplementary Figure 1. Examples of staining for each antibody used for the mass cytometry analysis.
 Supplementary Figure 1 Examples of staining for each antibody used for the mass cytometry analysis. To illustrate the functionality of each antibody probe, representative plots illustrating the expected
Supplementary Figure 1 Examples of staining for each antibody used for the mass cytometry analysis. To illustrate the functionality of each antibody probe, representative plots illustrating the expected
Signaling Through Immune System Receptors (Ch. 7)
 Signaling Through Immune System Receptors (Ch. 7) 1. General principles of signal transduction and propagation. 2. Antigen receptor signaling and lymphocyte activation. 3. Other receptors and signaling
Signaling Through Immune System Receptors (Ch. 7) 1. General principles of signal transduction and propagation. 2. Antigen receptor signaling and lymphocyte activation. 3. Other receptors and signaling
Recommended Protocols for Phospho Protein Detection in Human Cells
 Recommended Protocols for Phospho Protein Detection in Human Cells Depending on subcellular localization of the phospho protein of interest, as well as epitope susceptibility to cell fixing and permeabilizing
Recommended Protocols for Phospho Protein Detection in Human Cells Depending on subcellular localization of the phospho protein of interest, as well as epitope susceptibility to cell fixing and permeabilizing
Application Information Bulletin: Cell Signaling
 Application Information Bulletin: Cell Signaling Detection of Both Surface Proteins and Intracellular Phospho-Determinants in Whole Blood Using the PerFix-p Reagent System William L. Godfrey 1, Li Yang
Application Information Bulletin: Cell Signaling Detection of Both Surface Proteins and Intracellular Phospho-Determinants in Whole Blood Using the PerFix-p Reagent System William L. Godfrey 1, Li Yang
Supplementary Figure 1. Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
 Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/3/114/ra23/dc1 Supplementary Materials for Regulation of Zap70 Expression During Thymocyte Development Enables Temporal Separation of CD4 and CD8 Repertoire Selection
www.sciencesignaling.org/cgi/content/full/3/114/ra23/dc1 Supplementary Materials for Regulation of Zap70 Expression During Thymocyte Development Enables Temporal Separation of CD4 and CD8 Repertoire Selection
Human Immunodeficiency Virus Type-1 Myeloid Derived Suppressor Cells Inhibit Cytomegalovirus Inflammation through Interleukin-27 and B7-H4
 Human Immunodeficiency Virus Type-1 Myeloid Derived Suppressor Cells Inhibit Cytomegalovirus Inflammation through Interleukin-27 and B7-H4 Ankita Garg, Rodney Trout and Stephen A. Spector,,* Department
Human Immunodeficiency Virus Type-1 Myeloid Derived Suppressor Cells Inhibit Cytomegalovirus Inflammation through Interleukin-27 and B7-H4 Ankita Garg, Rodney Trout and Stephen A. Spector,,* Department
BCR-ABL uncouples canonical JAK2-STAT5 signaling in chronic myeloid
 Supplementary Results BCR-ABL uncouples canonical JAK2-STAT5 signaling in chronic myeloid leukemia Oliver Hantschel*, Wolfgang Warsch*, Eva Eckelhart*, Ines Kaupe, Florian Grebien, Kay-Uwe Wagner, Giulio
Supplementary Results BCR-ABL uncouples canonical JAK2-STAT5 signaling in chronic myeloid leukemia Oliver Hantschel*, Wolfgang Warsch*, Eva Eckelhart*, Ines Kaupe, Florian Grebien, Kay-Uwe Wagner, Giulio
a b G75 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server.
 a b G75 2 2 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server. a. Overlay of top 10 models generated by I-TASSER illustrates the potential effect of 7 amino acid insertion
a b G75 2 2 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server. a. Overlay of top 10 models generated by I-TASSER illustrates the potential effect of 7 amino acid insertion
Next-Generation Immunohistochemistry: Multiplex tissue imaging with mass cytometry
 Nat Met, April 2014 Nat Med, April 2014 Next-Generation Immunohistochemistry: Multiplex tissue imaging with mass cytometry Journal Club Timo Böge Overview Introduction Conventional Immunohistochemistry
Nat Met, April 2014 Nat Med, April 2014 Next-Generation Immunohistochemistry: Multiplex tissue imaging with mass cytometry Journal Club Timo Böge Overview Introduction Conventional Immunohistochemistry
Muse Assays for Cell Analysis
 Muse Assays for Cell Analysis Multiple Assay Outputs for Cell Analysis Cell Health Cell Signalling Immunology Muse Count & Viability Kit Muse Cell Cycle Kit Muse Annexin V & Dead Cell Kit Muse Caspase
Muse Assays for Cell Analysis Multiple Assay Outputs for Cell Analysis Cell Health Cell Signalling Immunology Muse Count & Viability Kit Muse Cell Cycle Kit Muse Annexin V & Dead Cell Kit Muse Caspase
AbSeq on the BD Rhapsody system: Exploration of single-cell gene regulation by simultaneous digital mrna and protein quantification
 BD AbSeq on the BD Rhapsody system: Exploration of single-cell gene regulation by simultaneous digital mrna and protein quantification Overview of BD AbSeq antibody-oligonucleotide conjugates. High-throughput
BD AbSeq on the BD Rhapsody system: Exploration of single-cell gene regulation by simultaneous digital mrna and protein quantification Overview of BD AbSeq antibody-oligonucleotide conjugates. High-throughput
Improved Stability of the LANCE Ultra Signal in Kinase Assays
 Improved Stability of the LANCE Ultra Signal in Kinase Assays LANCE Ultra is a high throughput screening (HTS) technology platform optimized for homogeneous time-resolved fluorescence resonance energy
Improved Stability of the LANCE Ultra Signal in Kinase Assays LANCE Ultra is a high throughput screening (HTS) technology platform optimized for homogeneous time-resolved fluorescence resonance energy
CML CML CML. tyrosine kinase inhibitor CML. 22 t(9;22)(q34;q11) chronic myeloid leukemia CML ABL. BCR-ABL c- imatinib mesylate CML CML BCR-ABL
 1 Key Wordschronic myeloid leukemiaimatinib mesylate tyrosine kinase inhibitor chronic myeloid leukemia CML imatinib mesylate CML CML CML CML Ph 10 1 30 50 3 5 CML α IFNα Ph Ph cytogenetic response CRmajor
1 Key Wordschronic myeloid leukemiaimatinib mesylate tyrosine kinase inhibitor chronic myeloid leukemia CML imatinib mesylate CML CML CML CML Ph 10 1 30 50 3 5 CML α IFNα Ph Ph cytogenetic response CRmajor
Mass Cytometry Applications for Immunology Research. Pub Note PN 13 01_150505
 Mass Cytometry Applications for Immunology Research Pub Note PN 13 01_150505 Contents Continuum of CD8 + T cell phenotypes revealed by deep profiling with mass cytometry 3 The transcriptional landscape
Mass Cytometry Applications for Immunology Research Pub Note PN 13 01_150505 Contents Continuum of CD8 + T cell phenotypes revealed by deep profiling with mass cytometry 3 The transcriptional landscape
Supplementary Figure 1
 Supplementary Figure 1 Identification of IFN-γ-producing CD8 + and CD4 + T cells with naive phenotype by alternative gating and sample-processing strategies. a. Contour 5% probability plots show definition
Supplementary Figure 1 Identification of IFN-γ-producing CD8 + and CD4 + T cells with naive phenotype by alternative gating and sample-processing strategies. a. Contour 5% probability plots show definition
Bead Based Assays for Cytokine Detection
 Bead Based Assays for Cytokine Detection September 27, 2014 6 th EFIS-EJI South East European Immunology School SEEIS 2014 Timisoara, Romania The Cells of the Immune System The Immune Reaction (Th2) (Th1)
Bead Based Assays for Cytokine Detection September 27, 2014 6 th EFIS-EJI South East European Immunology School SEEIS 2014 Timisoara, Romania The Cells of the Immune System The Immune Reaction (Th2) (Th1)
Supplementary Figure 1. BMS enhances human T cell activation in vitro in a
 Supplementary Figure 1. BMS98662 enhances human T cell activation in vitro in a concentration-dependent manner. Jurkat T cells were activated with anti-cd3 and anti-cd28 antibody in the presence of titrated
Supplementary Figure 1. BMS98662 enhances human T cell activation in vitro in a concentration-dependent manner. Jurkat T cells were activated with anti-cd3 and anti-cd28 antibody in the presence of titrated
W/T Itgam -/- F4/80 CD115. F4/80 hi CD115 + F4/80 + CD115 +
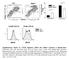 F4/8 % in the peritoneal lavage 6 4 2 p=.15 n.s p=.76 CD115 F4/8 hi CD115 + F4/8 + CD115 + F4/8 hi CD115 + F4/8 + CD115 + MHCII MHCII Supplementary Figure S1. CD11b deficiency affects the cellular responses
F4/8 % in the peritoneal lavage 6 4 2 p=.15 n.s p=.76 CD115 F4/8 hi CD115 + F4/8 + CD115 + F4/8 hi CD115 + F4/8 + CD115 + MHCII MHCII Supplementary Figure S1. CD11b deficiency affects the cellular responses
Dox. R26-rtTA Tyr-CreERT2. any ink/arf, no rtta (n=8) ink/arf +/+ (n=5) Day 0 Day 11 Day 18 Day 28
 A 4OHT Dox hraf iip tumors inras ddh 2 O -RT Ink/Arf / Pten l/ l R26-lsl-rtTA Tyr-reERT2 TetO-hRAF V6E Ink/Arf / Pten / R26-rtTA Tyr-reERT2 TetO-hRAF V6E Ink/Arf / Pten / R26-rtTA Tyr-reERT2 TetO-hRAF
A 4OHT Dox hraf iip tumors inras ddh 2 O -RT Ink/Arf / Pten l/ l R26-lsl-rtTA Tyr-reERT2 TetO-hRAF V6E Ink/Arf / Pten / R26-rtTA Tyr-reERT2 TetO-hRAF V6E Ink/Arf / Pten / R26-rtTA Tyr-reERT2 TetO-hRAF
Supplementary Table; Supplementary Figures and legends S1-S21; Supplementary Materials and Methods
 Silva et al. PTEN posttranslational inactivation and hyperactivation of the PI3K/Akt pathway sustain primary T cell leukemia viability Supplementary Table; Supplementary Figures and legends S1-S21; Supplementary
Silva et al. PTEN posttranslational inactivation and hyperactivation of the PI3K/Akt pathway sustain primary T cell leukemia viability Supplementary Table; Supplementary Figures and legends S1-S21; Supplementary
A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy se
 A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy serves as a defence mechanism that prevents or retards
A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy serves as a defence mechanism that prevents or retards
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/23854 holds various files of this Leiden University dissertation. Author: Marel, Sander van der Title: Gene and cell therapy based treatment strategies
Cover Page The handle http://hdl.handle.net/1887/23854 holds various files of this Leiden University dissertation. Author: Marel, Sander van der Title: Gene and cell therapy based treatment strategies
Supplementary Information. Table of contents
 Supplementary Information Table of contents Fig. S1. Inhibition of specific upstream kinases affects the activity of the analyzed readouts Fig. S2. Down-regulation of INCENP gene induces the formation
Supplementary Information Table of contents Fig. S1. Inhibition of specific upstream kinases affects the activity of the analyzed readouts Fig. S2. Down-regulation of INCENP gene induces the formation
Protein kinases are enzymes that add a phosphate group to proteins according to the. ATP + protein OH > Protein OPO 3 + ADP
 Protein kinase Protein kinases are enzymes that add a phosphate group to proteins according to the following equation: 2 ATP + protein OH > Protein OPO 3 + ADP ATP represents adenosine trisphosphate, ADP
Protein kinase Protein kinases are enzymes that add a phosphate group to proteins according to the following equation: 2 ATP + protein OH > Protein OPO 3 + ADP ATP represents adenosine trisphosphate, ADP
Detailed step-by-step operating procedures for NK cell and CTL degranulation assays
 Supplemental methods Detailed step-by-step operating procedures for NK cell and CTL degranulation assays Materials PBMC isolated from patients, relatives and healthy donors as control K562 cells (ATCC,
Supplemental methods Detailed step-by-step operating procedures for NK cell and CTL degranulation assays Materials PBMC isolated from patients, relatives and healthy donors as control K562 cells (ATCC,
blood mononuclear cells were stained with AAD, annexin, CD3, CD4, CD25, LAP and specific
 Supplementary figure legends Figure 1: Selective expression of regulatory markers on CD4 + LAP + T cells. Human peripheral blood mononuclear cells were stained with AAD, annexin, CD3, CD4, CD25, LAP and
Supplementary figure legends Figure 1: Selective expression of regulatory markers on CD4 + LAP + T cells. Human peripheral blood mononuclear cells were stained with AAD, annexin, CD3, CD4, CD25, LAP and
Nature Biotechnology: doi: /nbt.3828
 Supplementary Figure 1 Development of a FRET-based MCS. (a) Linker and MA2 modification are indicated by single letter amino acid code. indicates deletion of amino acids and N or C indicate the terminus
Supplementary Figure 1 Development of a FRET-based MCS. (a) Linker and MA2 modification are indicated by single letter amino acid code. indicates deletion of amino acids and N or C indicate the terminus
Supplementary Figure 1. Enhanced detection of CTLA-4 on the surface of HIV-specific
 SUPPLEMENTARY FIGURE LEGEND Supplementary Figure 1. Enhanced detection of CTLA-4 on the surface of HIV-specific CD4 + T cells correlates with intracellular CTLA-4 levels. (a) Comparative CTLA-4 levels
SUPPLEMENTARY FIGURE LEGEND Supplementary Figure 1. Enhanced detection of CTLA-4 on the surface of HIV-specific CD4 + T cells correlates with intracellular CTLA-4 levels. (a) Comparative CTLA-4 levels
Propagation of the Signal
 OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
Tbk1-TKO! DN cells (%)! 15! 10!
 a! T Cells! TKO! B Cells! TKO! b! CD4! 8.9 85.2 3.4 2.88 CD8! Tbk1-TKO! 1.1 84.8 2.51 2.54 c! DN cells (%)! 4 3 2 1 DP cells (%)! 9 8 7 6 CD4 + SP cells (%)! 5 4 3 2 1 5 TKO! TKO! TKO! TKO! 15 1 5 CD8
a! T Cells! TKO! B Cells! TKO! b! CD4! 8.9 85.2 3.4 2.88 CD8! Tbk1-TKO! 1.1 84.8 2.51 2.54 c! DN cells (%)! 4 3 2 1 DP cells (%)! 9 8 7 6 CD4 + SP cells (%)! 5 4 3 2 1 5 TKO! TKO! TKO! TKO! 15 1 5 CD8
Optimization of a LanthaScreen Kinase assay for CSF1R (FMS)
 Optimization of a LanthaScreen Kinase assay for CSF1R (FMS) Overview This protocol describes how to develop a LanthaScreen kinase assay designed to detect and characterize kinase inhibitors. The development
Optimization of a LanthaScreen Kinase assay for CSF1R (FMS) Overview This protocol describes how to develop a LanthaScreen kinase assay designed to detect and characterize kinase inhibitors. The development
Jakinibs 101: Theory, Practice and Prospects. References 10/27/2013
 Jakinibs 101: Theory, Practice and Prospects As a rheumatologist, what do you need to know? Why should you care? Understand what Jaks are Which cytokines care, which don t Mechanisms underlying Jakinib
Jakinibs 101: Theory, Practice and Prospects As a rheumatologist, what do you need to know? Why should you care? Understand what Jaks are Which cytokines care, which don t Mechanisms underlying Jakinib
Receptor mediated Signal Transduction
 Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Optimizing Intracellular Flow Cytometry
 Optimizing Intracellular Flow Cytometry Detection of Cytokines, Transcription Factors, and Phosphoprotein by Flow Cytometry Presented by Erika O Donnell, PhD, BD Biosciences 23-14876-00 Outline Basic principles
Optimizing Intracellular Flow Cytometry Detection of Cytokines, Transcription Factors, and Phosphoprotein by Flow Cytometry Presented by Erika O Donnell, PhD, BD Biosciences 23-14876-00 Outline Basic principles
Optimization of a LanthaScreen Kinase assay for ZAP70
 Optimization of a LanthaScreen Kinase assay for ZAP70 Overview This protocol describes how to develop a LanthaScreen kinase assay designed to detect and characterize kinase inhibitors. The development
Optimization of a LanthaScreen Kinase assay for ZAP70 Overview This protocol describes how to develop a LanthaScreen kinase assay designed to detect and characterize kinase inhibitors. The development
Cell Signaling part 2
 15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
Chapter 9: Biochemical Mechanisms for Information Storage at the Cellular Level. From Mechanisms of Memory, second edition By J. David Sweatt, Ph.D.
 Chapter 9: Biochemical Mechanisms for Information Storage at the Cellular Level From Mechanisms of Memory, second edition By J. David Sweatt, Ph.D. Chapter 9: Dendritic Spine Figure 1 Summary: Three Primary
Chapter 9: Biochemical Mechanisms for Information Storage at the Cellular Level From Mechanisms of Memory, second edition By J. David Sweatt, Ph.D. Chapter 9: Dendritic Spine Figure 1 Summary: Three Primary
Insulin/IGF1-IRS-PI-3 kinase-mtor signaling pathway
 Insulin/IGF1-IRS-PI-3 kinase-mtor signaling pathway Nutrients RAPAMYCIN (FKBP12) mtor/rictor complex phosphorylates Akt at the S473 activating site in a Rheb-independent fashion TSC1 or TSC2-null MEFs
Insulin/IGF1-IRS-PI-3 kinase-mtor signaling pathway Nutrients RAPAMYCIN (FKBP12) mtor/rictor complex phosphorylates Akt at the S473 activating site in a Rheb-independent fashion TSC1 or TSC2-null MEFs
MATERIALS AND METHODS. Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All
 MATERIALS AND METHODS Antibodies (Abs), flow cytometry analysis and cell lines Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All other antibodies used
MATERIALS AND METHODS Antibodies (Abs), flow cytometry analysis and cell lines Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All other antibodies used
Data Sheet. NFAT Reporter (Luc) Jurkat Cell line Catalog #: 60621
 Data Sheet NFAT Reporter (Luc) Jurkat Cell line Catalog #: 60621 Background The nuclear factor of activator T cells (NFAT) family of transcription factors plays an important role in immune response. T
Data Sheet NFAT Reporter (Luc) Jurkat Cell line Catalog #: 60621 Background The nuclear factor of activator T cells (NFAT) family of transcription factors plays an important role in immune response. T
Layered-IHC (L-IHC): A novel and robust approach to multiplexed immunohistochemistry So many markers and so little tissue
 Page 1 The need for multiplex detection of tissue biomarkers. There is a constant and growing demand for increased biomarker analysis in human tissue specimens. Analysis of tissue biomarkers is key to
Page 1 The need for multiplex detection of tissue biomarkers. There is a constant and growing demand for increased biomarker analysis in human tissue specimens. Analysis of tissue biomarkers is key to
SUPPLEMENTARY INFORMATION
 Complete but curtailed T-cell response to very-low-affinity antigen Dietmar Zehn, Sarah Y. Lee & Michael J. Bevan Supp. Fig. 1: TCR chain usage among endogenous K b /Ova reactive T cells. C57BL/6 mice
Complete but curtailed T-cell response to very-low-affinity antigen Dietmar Zehn, Sarah Y. Lee & Michael J. Bevan Supp. Fig. 1: TCR chain usage among endogenous K b /Ova reactive T cells. C57BL/6 mice
BD Flow Cytometry Reagents Multicolor Panels Designed for Optimal Resolution with the BD LSRFortessa X-20 Cell Analyzer
 Multicolor Panels Designed for Optimal Resolution with the BD LSRFortessa X-2 Cell Analyzer Proper multicolor panel design takes into account fluorochrome brightness, antigen density, co-expression, and
Multicolor Panels Designed for Optimal Resolution with the BD LSRFortessa X-2 Cell Analyzer Proper multicolor panel design takes into account fluorochrome brightness, antigen density, co-expression, and
T Cell Effector Mechanisms I: B cell Help & DTH
 T Cell Effector Mechanisms I: B cell Help & DTH Ned Braunstein, MD The Major T Cell Subsets p56 lck + T cells γ δ ε ζ ζ p56 lck CD8+ T cells γ δ ε ζ ζ Cα Cβ Vα Vβ CD3 CD8 Cα Cβ Vα Vβ CD3 MHC II peptide
T Cell Effector Mechanisms I: B cell Help & DTH Ned Braunstein, MD The Major T Cell Subsets p56 lck + T cells γ δ ε ζ ζ p56 lck CD8+ T cells γ δ ε ζ ζ Cα Cβ Vα Vβ CD3 CD8 Cα Cβ Vα Vβ CD3 MHC II peptide
Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in
 Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in Supplementary Fig. 2 Substitution Sequence Position variant Sequence original APNCYGNIPL original APNCYGNIPL
Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in Supplementary Fig. 2 Substitution Sequence Position variant Sequence original APNCYGNIPL original APNCYGNIPL
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity
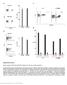 Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
Lecture 7: Signaling Through Lymphocyte Receptors
 Lecture 7: Signaling Through Lymphocyte Receptors Questions to Consider After recognition of its cognate MHC:peptide, how does the T cell receptor activate immune response genes? What are the structural
Lecture 7: Signaling Through Lymphocyte Receptors Questions to Consider After recognition of its cognate MHC:peptide, how does the T cell receptor activate immune response genes? What are the structural
The g c Family of Cytokines Prof. Warren J. Leonard M.D.
 The Family of Cytokines Chief, Laboratory of Molecular Immunology Director, Immunology Center National Heart, Lung, and Blood Institute National Institutes of Health Department of Health and Human Services
The Family of Cytokines Chief, Laboratory of Molecular Immunology Director, Immunology Center National Heart, Lung, and Blood Institute National Institutes of Health Department of Health and Human Services
T cell maturation. T-cell Maturation. What allows T cell maturation?
 T-cell Maturation What allows T cell maturation? Direct contact with thymic epithelial cells Influence of thymic hormones Growth factors (cytokines, CSF) T cell maturation T cell progenitor DN DP SP 2ry
T-cell Maturation What allows T cell maturation? Direct contact with thymic epithelial cells Influence of thymic hormones Growth factors (cytokines, CSF) T cell maturation T cell progenitor DN DP SP 2ry
Supporting Information Table of Contents
 Supporting Information Table of Contents Supporting Information Figure 1 Page 2 Supporting Information Figure 2 Page 4 Supporting Information Figure 3 Page 5 Supporting Information Figure 4 Page 6 Supporting
Supporting Information Table of Contents Supporting Information Figure 1 Page 2 Supporting Information Figure 2 Page 4 Supporting Information Figure 3 Page 5 Supporting Information Figure 4 Page 6 Supporting
Chapter 11. B cell generation, Activation, and Differentiation. Pro-B cells. - B cells mature in the bone marrow.
 Chapter B cell generation, Activation, and Differentiation - B cells mature in the bone marrow. - B cells proceed through a number of distinct maturational stages: ) Pro-B cell ) Pre-B cell ) Immature
Chapter B cell generation, Activation, and Differentiation - B cells mature in the bone marrow. - B cells proceed through a number of distinct maturational stages: ) Pro-B cell ) Pre-B cell ) Immature
G-Protein Signaling. Introduction to intracellular signaling. Dr. SARRAY Sameh, Ph.D
 G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
Nature Medicine: doi: /nm.2109
 HIV 1 Infects Multipotent Progenitor Cells Causing Cell Death and Establishing Latent Cellular Reservoirs Christoph C. Carter, Adewunmi Onafuwa Nuga, Lucy A. M c Namara, James Riddell IV, Dale Bixby, Michael
HIV 1 Infects Multipotent Progenitor Cells Causing Cell Death and Establishing Latent Cellular Reservoirs Christoph C. Carter, Adewunmi Onafuwa Nuga, Lucy A. M c Namara, James Riddell IV, Dale Bixby, Michael
staining and flow cytometry
 Detection of influenza virus-specific T cell responses by intracellular by cytokine intracellular staining cytokine staining and flow cytometry Detection of influenza virus-specific T cell responses and
Detection of influenza virus-specific T cell responses by intracellular by cytokine intracellular staining cytokine staining and flow cytometry Detection of influenza virus-specific T cell responses and
Optimization of a LanthaScreen Kinase assay for NTRK1 (TRKA)
 Optimization of a LanthaScreen Kinase assay for NTRK1 (TRKA) Overview This protocol describes how to develop a LanthaScreen kinase assay designed to detect and characterize kinase inhibitors. The development
Optimization of a LanthaScreen Kinase assay for NTRK1 (TRKA) Overview This protocol describes how to develop a LanthaScreen kinase assay designed to detect and characterize kinase inhibitors. The development
CD14 + S100A9 + Monocytic Myeloid-Derived Suppressor Cells and Their Clinical Relevance in Non-Small Cell Lung Cancer
 CD14 + S1A9 + Monocytic Myeloid-Derived Suppressor Cells and Their Clinical Relevance in Non-Small Cell Lung Cancer Po-Hao, Feng M.D., Kang-Yun, Lee, M.D. Ph.D., Ya-Ling Chang, Yao-Fei Chan, Lu- Wei, Kuo,Ting-Yu
CD14 + S1A9 + Monocytic Myeloid-Derived Suppressor Cells and Their Clinical Relevance in Non-Small Cell Lung Cancer Po-Hao, Feng M.D., Kang-Yun, Lee, M.D. Ph.D., Ya-Ling Chang, Yao-Fei Chan, Lu- Wei, Kuo,Ting-Yu
KEY CONCEPT QUESTIONS IN SIGNAL TRANSDUCTION
 Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to
 Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
Signal Transduction Pathway Smorgasbord
 Molecular Cell Biology Lecture. Oct 28, 2014 Signal Transduction Pathway Smorgasbord Ron Bose, MD PhD Biochemistry and Molecular Cell Biology Programs Washington University School of Medicine Outline 1.
Molecular Cell Biology Lecture. Oct 28, 2014 Signal Transduction Pathway Smorgasbord Ron Bose, MD PhD Biochemistry and Molecular Cell Biology Programs Washington University School of Medicine Outline 1.
MCB*4010 Midterm Exam / Winter 2008
 MCB*4010 Midterm Exam / Winter 2008 Name: ID: Instructions: Answer all 4 questions. The number of marks for each question indicates how many points you need to provide. Write your answers in point form,
MCB*4010 Midterm Exam / Winter 2008 Name: ID: Instructions: Answer all 4 questions. The number of marks for each question indicates how many points you need to provide. Write your answers in point form,
Supplementary Figure 1. IL-12 serum levels and frequency of subsets in FL patients. (A) IL-12
 1 Supplementary Data Figure legends Supplementary Figure 1. IL-12 serum levels and frequency of subsets in FL patients. (A) IL-12 serum levels measured by multiplex ELISA (Luminex) in FL patients before
1 Supplementary Data Figure legends Supplementary Figure 1. IL-12 serum levels and frequency of subsets in FL patients. (A) IL-12 serum levels measured by multiplex ELISA (Luminex) in FL patients before
Reviewers' comments: Reviewer #1 Expert in EAE and IL-17a (Remarks to the Author):
 Reviewers' comments: Reviewer #1 Expert in EAE and IL-17a (Remarks to the Author): This study shows that the inducible camp early repressor (ICER) is involved in development of Th17 cells that are pathogenic
Reviewers' comments: Reviewer #1 Expert in EAE and IL-17a (Remarks to the Author): This study shows that the inducible camp early repressor (ICER) is involved in development of Th17 cells that are pathogenic
Regulation of cell function by intracellular signaling
 Regulation of cell function by intracellular signaling Objectives: Regulation principle Allosteric and covalent mechanisms, Popular second messengers, Protein kinases, Kinase cascade and interaction. regulation
Regulation of cell function by intracellular signaling Objectives: Regulation principle Allosteric and covalent mechanisms, Popular second messengers, Protein kinases, Kinase cascade and interaction. regulation
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10134 Supplementary Figure 1. Anti-inflammatory activity of sfc. a, Autoantibody immune complexes crosslink activating Fc receptors, promoting activation of macrophages, and WWW.NATURE.COM/NATURE
doi:10.1038/nature10134 Supplementary Figure 1. Anti-inflammatory activity of sfc. a, Autoantibody immune complexes crosslink activating Fc receptors, promoting activation of macrophages, and WWW.NATURE.COM/NATURE
Supplementary Fig. 1 p38 MAPK negatively regulates DC differentiation. (a) Western blot analysis of p38 isoform expression in BM cells, immature DCs
 Supplementary Fig. 1 p38 MAPK negatively regulates DC differentiation. (a) Western blot analysis of p38 isoform expression in BM cells, immature DCs (idcs) and mature DCs (mdcs). A myeloma cell line expressing
Supplementary Fig. 1 p38 MAPK negatively regulates DC differentiation. (a) Western blot analysis of p38 isoform expression in BM cells, immature DCs (idcs) and mature DCs (mdcs). A myeloma cell line expressing
Lecture: CHAPTER 13 Signal Transduction Pathways
 Lecture: 10 17 2016 CHAPTER 13 Signal Transduction Pathways Chapter 13 Outline Signal transduction cascades have many components in common: 1. Release of a primary message as a response to a physiological
Lecture: 10 17 2016 CHAPTER 13 Signal Transduction Pathways Chapter 13 Outline Signal transduction cascades have many components in common: 1. Release of a primary message as a response to a physiological
CAN THE CD43 MOLECULE BE CONSIDERED AS A THERAPEUTIC TAGET? Yvonne Rosenstein Instituto de Biotecnologia, UNAM
 CAN THE CD43 MOLECULE BE CONSIDERED AS A THERAPEUTIC TAGET? Yvonne Rosenstein Instituto de Biotecnologia, UNAM CD43: the molecule and its signaling capacities favors a Th1 differentiation pattern and contributes
CAN THE CD43 MOLECULE BE CONSIDERED AS A THERAPEUTIC TAGET? Yvonne Rosenstein Instituto de Biotecnologia, UNAM CD43: the molecule and its signaling capacities favors a Th1 differentiation pattern and contributes
B-cell expansion and lymphomagenesis induced by chronic CD40-signaling is
 -cell expansion and lymphomagenesis induced by chronic CD4-signaling is strictly dependent on CD19 Caroline Hojer 1*, Samantha Frankenberger 1*, Lothar J. Strobl 1, Samantha Feicht 1, Kristina Djermanovic
-cell expansion and lymphomagenesis induced by chronic CD4-signaling is strictly dependent on CD19 Caroline Hojer 1*, Samantha Frankenberger 1*, Lothar J. Strobl 1, Samantha Feicht 1, Kristina Djermanovic
Synthesis and Biological Evaluation of Protein Kinase D Inhibitors
 Synthesis and Biological Evaluation of Protein Kinase D Inhibitors Celeste Alverez Topic Seminar October 26, 2013 Celeste Alverez @ Wipf Group 10/26/2013 1 Protein Kinase D (PKD) A novel family of serine/threonine
Synthesis and Biological Evaluation of Protein Kinase D Inhibitors Celeste Alverez Topic Seminar October 26, 2013 Celeste Alverez @ Wipf Group 10/26/2013 1 Protein Kinase D (PKD) A novel family of serine/threonine
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/381/ra59/dc1 Supplementary Materials for Analysis of single-cell cytokine secretion reveals a role for paracrine signaling in coordinating macrophage responses
www.sciencesignaling.org/cgi/content/full/8/381/ra59/dc1 Supplementary Materials for Analysis of single-cell cytokine secretion reveals a role for paracrine signaling in coordinating macrophage responses
Intracellular MHC class II molecules promote TLR-triggered innate. immune responses by maintaining Btk activation
 Intracellular MHC class II molecules promote TLR-triggered innate immune responses by maintaining Btk activation Xingguang Liu, Zhenzhen Zhan, Dong Li, Li Xu, Feng Ma, Peng Zhang, Hangping Yao and Xuetao
Intracellular MHC class II molecules promote TLR-triggered innate immune responses by maintaining Btk activation Xingguang Liu, Zhenzhen Zhan, Dong Li, Li Xu, Feng Ma, Peng Zhang, Hangping Yao and Xuetao
Spatially resolved multiparametric single cell analysis. Technical Journal Club 19th September 2017 Christina Müller (Group Speck)
 Spatially resolved multiparametric single cell analysis Technical Journal Club 19th September 2017 Christina Müller (Group Speck) Why spatially resolved multiparametric single cell analysis? Multiparametric
Spatially resolved multiparametric single cell analysis Technical Journal Club 19th September 2017 Christina Müller (Group Speck) Why spatially resolved multiparametric single cell analysis? Multiparametric
HD1 (FLU) HD2 (EBV) HD2 (FLU)
 ramer staining + anti-pe beads ramer staining a HD1 (FLU) HD2 (EBV) HD2 (FLU).73.11.56.46.24 1.12 b CD127 + c CD127 + d CD127 - e CD127 - PD1 - PD1 + PD1 + PD1-1 1 1 1 %CD127 + PD1-8 6 4 2 + anti-pe %CD127
ramer staining + anti-pe beads ramer staining a HD1 (FLU) HD2 (EBV) HD2 (FLU).73.11.56.46.24 1.12 b CD127 + c CD127 + d CD127 - e CD127 - PD1 - PD1 + PD1 + PD1-1 1 1 1 %CD127 + PD1-8 6 4 2 + anti-pe %CD127
Supplementary Figure 1) GABAergic enhancement by leptin hyperpolarizes POMC neurons A) Representative recording samples showing the membrane
 Supplementary Figure 1) GABAergic enhancement by leptin hyperpolarizes POMC neurons A) Representative recording samples showing the membrane potential recorded from POMC neurons following treatment with
Supplementary Figure 1) GABAergic enhancement by leptin hyperpolarizes POMC neurons A) Representative recording samples showing the membrane potential recorded from POMC neurons following treatment with
Titrations in Cytobank
 The Premier Platform for Single Cell Analysis (1) Titrations in Cytobank To analyze data from a titration in Cytobank, the first step is to upload your own FCS files or clone an experiment you already
The Premier Platform for Single Cell Analysis (1) Titrations in Cytobank To analyze data from a titration in Cytobank, the first step is to upload your own FCS files or clone an experiment you already
Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were
 Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were isolated from wild type (PKC-θ- WT) or PKC-θ null (PKC-θ-KO)
Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were isolated from wild type (PKC-θ- WT) or PKC-θ null (PKC-θ-KO)
Screening Protocol and Assay Conditions
 Page 1 of 20 OVERVIEW AND ASSAY THEORY 2 SELECTSCREEN ASSAY CONDITIONS 4 SELECTSCREEN ASSAY CONTROLS 6 SELECTSCREEN DATA ANALYSIS 7 PATHWAY PROFILING CELL LINES AVAILABLE FOR SCREENING 8 CELL LINE-SPECIFIC
Page 1 of 20 OVERVIEW AND ASSAY THEORY 2 SELECTSCREEN ASSAY CONDITIONS 4 SELECTSCREEN ASSAY CONTROLS 6 SELECTSCREEN DATA ANALYSIS 7 PATHWAY PROFILING CELL LINES AVAILABLE FOR SCREENING 8 CELL LINE-SPECIFIC
Nature Immunology: doi: /ni.3631
 Supplementary Figure 1 SPT analyses of Zap70 at the T cell plasma membrane. (a) Total internal reflection fluorescent (TIRF) excitation at 64-68 degrees limits single molecule detection to 100-150 nm above
Supplementary Figure 1 SPT analyses of Zap70 at the T cell plasma membrane. (a) Total internal reflection fluorescent (TIRF) excitation at 64-68 degrees limits single molecule detection to 100-150 nm above
T Cell Activation. Patricia Fitzgerald-Bocarsly March 18, 2009
 T Cell Activation Patricia Fitzgerald-Bocarsly March 18, 2009 Phases of Adaptive Immune Responses Phases of T cell responses IL-2 acts as an autocrine growth factor Fig. 11-11 Clonal Expansion of T cells
T Cell Activation Patricia Fitzgerald-Bocarsly March 18, 2009 Phases of Adaptive Immune Responses Phases of T cell responses IL-2 acts as an autocrine growth factor Fig. 11-11 Clonal Expansion of T cells
Nature Medicine: doi: /nm.3967
 Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
nature methods Organelle-specific, rapid induction of molecular activities and membrane tethering
 nature methods Organelle-specific, rapid induction of molecular activities and membrane tethering Toru Komatsu, Igor Kukelyansky, J Michael McCaffery, Tasuku Ueno, Lidenys C Varela & Takanari Inoue Supplementary
nature methods Organelle-specific, rapid induction of molecular activities and membrane tethering Toru Komatsu, Igor Kukelyansky, J Michael McCaffery, Tasuku Ueno, Lidenys C Varela & Takanari Inoue Supplementary
Mass Cytometry Publication List
 Mass Cytometry Publication List 2014 Publications 1 Becher, B., et al. High-dimensional analysis of the murine myeloid cell system. Nat Immunol 15 (12): 1181-1189, 2014. Behbehani, G.K., et al. Transient
Mass Cytometry Publication List 2014 Publications 1 Becher, B., et al. High-dimensional analysis of the murine myeloid cell system. Nat Immunol 15 (12): 1181-1189, 2014. Behbehani, G.K., et al. Transient
Therapeutic PD L1 and LAG 3 blockade rapidly clears established blood stage Plasmodium infection
 Supplementary Information Therapeutic PD L1 and LAG 3 blockade rapidly clears established blood stage Plasmodium infection Noah S. Butler, Jacqueline Moebius, Lecia L. Pewe, Boubacar Traore, Ogobara K.
Supplementary Information Therapeutic PD L1 and LAG 3 blockade rapidly clears established blood stage Plasmodium infection Noah S. Butler, Jacqueline Moebius, Lecia L. Pewe, Boubacar Traore, Ogobara K.
Defective STAT1 activation associated with impaired IFN-g production in NK and T lymphocytes from metastatic melanoma patients treated with IL-2
 Defective STAT1 activation associated with impaired IFN-g production in NK and T lymphocytes from metastatic melanoma patients treated with IL-2 SUPPLEMENTARY FIGURES AND TABLES Supplementary Figure S1:
Defective STAT1 activation associated with impaired IFN-g production in NK and T lymphocytes from metastatic melanoma patients treated with IL-2 SUPPLEMENTARY FIGURES AND TABLES Supplementary Figure S1:
Signaling changes in the stem cell factor AKT-S6 pathway in diagnostic AML samples are associated with disease relapse
 LETTER TO THE EDITOR Citation: (2011) 1, e3; doi:10.1038/bcj.2010.2 & 2011 Macmillan Publishers Limited All rights reserved 2044-5385/11 www.nature.com/bcj Signaling changes in the stem cell factor AKT-S6
LETTER TO THE EDITOR Citation: (2011) 1, e3; doi:10.1038/bcj.2010.2 & 2011 Macmillan Publishers Limited All rights reserved 2044-5385/11 www.nature.com/bcj Signaling changes in the stem cell factor AKT-S6
Supplementary Information. A vital role for IL-2 trans-presentation in DC-mediated T cell activation in humans as revealed by daclizumab therapy
 Supplementary Information A vital role for IL-2 trans-presentation in DC-mediated T cell activation in humans as revealed by daclizumab therapy Simone C. Wuest 1, Jehad Edwan 1, Jayne F. Martin 1, Sungpil
Supplementary Information A vital role for IL-2 trans-presentation in DC-mediated T cell activation in humans as revealed by daclizumab therapy Simone C. Wuest 1, Jehad Edwan 1, Jayne F. Martin 1, Sungpil
Phospho-AKT Sampler Kit
 Phospho-AKT Sampler Kit E 0 5 1 0 0 3 Kits Includes Cat. Quantity Application Reactivity Source Akt (Ab-473) Antibody E021054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit Akt (Phospho-Ser473) Antibody
Phospho-AKT Sampler Kit E 0 5 1 0 0 3 Kits Includes Cat. Quantity Application Reactivity Source Akt (Ab-473) Antibody E021054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit Akt (Phospho-Ser473) Antibody
CD25-PE (BD Biosciences) and labeled with anti-pe-microbeads (Miltenyi Biotec) for depletion of CD25 +
 Supplements Supplemental Materials and Methods Depletion of CD25 + T-cells from PBMC. Fresh or HD precultured PBMC were stained with the conjugate CD25-PE (BD Biosciences) and labeled with anti-pe-microbeads
Supplements Supplemental Materials and Methods Depletion of CD25 + T-cells from PBMC. Fresh or HD precultured PBMC were stained with the conjugate CD25-PE (BD Biosciences) and labeled with anti-pe-microbeads
Lecture 1, Fall 2014 The structure of the eukaryotic cell, as it relates to cells chemical and biological functions.
 Lecture 1, Fall 2014 The structure of the eukaryotic cell, as it relates to cells chemical and biological functions. There are several important themes that transcends just the chemistry and bring the
Lecture 1, Fall 2014 The structure of the eukaryotic cell, as it relates to cells chemical and biological functions. There are several important themes that transcends just the chemistry and bring the
a Beckman Coulter Life Sciences: White Paper
 a Beckman Coulter Life Sciences: White Paper An 8-color DuraClone IM panel for detection of Human blood dendritic cells by flow cytometry Nathalie Dupas 1, Snehita Sattiraju 2, Neha Girish 2, Murthy Pendyala
a Beckman Coulter Life Sciences: White Paper An 8-color DuraClone IM panel for detection of Human blood dendritic cells by flow cytometry Nathalie Dupas 1, Snehita Sattiraju 2, Neha Girish 2, Murthy Pendyala
Fluorochrome Panel 1 Panel 2 Panel 3 Panel 4 Panel 5 CTLA-4 CTLA-4 CD15 CD3 FITC. Bio) PD-1 (MIH4, BD) ICOS (C398.4A, Biolegend) PD-L1 (MIH1, BD)
 Additional file : Table S. Antibodies used for panel stain to identify peripheral immune cell subsets. Panel : PD- signaling; Panel : CD + T cells, CD + T cells, B cells; Panel : Tregs; Panel :, -T, cdc,
Additional file : Table S. Antibodies used for panel stain to identify peripheral immune cell subsets. Panel : PD- signaling; Panel : CD + T cells, CD + T cells, B cells; Panel : Tregs; Panel :, -T, cdc,
ANALYTISCHE STRATEGIE Tissue Imaging. Bernd Bodenmiller Institute of Molecular Life Sciences University of Zurich
 ANALYTISCHE STRATEGIE Tissue Imaging Bernd Bodenmiller Institute of Molecular Life Sciences University of Zurich Quantitative Breast single cancer cell analysis Switzerland Brain Breast Lung Colon-rectum
ANALYTISCHE STRATEGIE Tissue Imaging Bernd Bodenmiller Institute of Molecular Life Sciences University of Zurich Quantitative Breast single cancer cell analysis Switzerland Brain Breast Lung Colon-rectum
Easy50 PI3K Inhibitor Array*
 Easy50 PI3K Inhibitor Array* Catalog # IAP001 Instruction Manual For Research Use Only *Patent pending Introduction This Easy50 PI3K Inhibitor Array is a panel of serial dilutions of inhibitors that are
Easy50 PI3K Inhibitor Array* Catalog # IAP001 Instruction Manual For Research Use Only *Patent pending Introduction This Easy50 PI3K Inhibitor Array is a panel of serial dilutions of inhibitors that are
Akt and mtor pathways differentially regulate the development of natural and inducible. T H 17 cells
 Akt and mtor pathways differentially regulate the development of natural and inducible T H 17 cells Jiyeon S Kim, Tammarah Sklarz, Lauren Banks, Mercy Gohil, Adam T Waickman, Nicolas Skuli, Bryan L Krock,
Akt and mtor pathways differentially regulate the development of natural and inducible T H 17 cells Jiyeon S Kim, Tammarah Sklarz, Lauren Banks, Mercy Gohil, Adam T Waickman, Nicolas Skuli, Bryan L Krock,
