CyTOF workflow: differential discovery in high-throughput high-dimensional cytometry datasets!
|
|
|
- Mervin Black
- 5 years ago
- Views:
Transcription
1 Workshop!! CyTOF workflow: differential discovery in high-throughput high-dimensional cytometry datasets! Malgorzata Nowicka! University of Zurich! BioC 2017: Where Software and Biology Connect! Boston, 28 July 2017!
2 Have you analyzed CyTOF or flow cytometry data before?!
3 My approach to this workshop! This is just a shorter version of the original workflow! The text is basically the same but maybe try to not focus on it so much, as you can carefully read it later! Ideally, I would do the life coding but because of the time constrains, I will:! explain all the code in the vignette (html file)! copy-paste to R, to follow with you and have the opportunity to modify some things if needed!
4 Introduction! CyTOF (mass cytometry) experiment! Measuring ~40 parameters (metals)! Hundreds of cells per second Figure adapted from Bendall et al. Science 2011!
5 Introduction! High-dimensional cytometry because it is higher than before (from ~12 to ~40)! We refer to this differential approach as classic! Common strategy:! identify cell populations of interest by manual gating or automated clustering! determine which of the cell subpopulations or protein markers are associated with a phenotype of interest using statistical tests! Other approaches:! Citrus (Bruggner et al. 2014)! CellCnn (Arvaniti and Claassen 2016)! Hybrid approaches (thanks to the modularity)! This workflow can serve as a template!
6 Overview! 1. Data description! 2. Data pre-processing (CATALYST pckg)! 3. Data import flowcore pckg! 4. Data transformation - arcsinh with cofactor 5! 5. Spot checks! MDS plots (limma pckg)! 6. Cell clustering with FlowSOM and ClusterConsensusPlus pckgs! overclustering + manual merging! 7. Visualization of the clusters with! t-sne - Rtsne pckg! heatmaps - pheatmap pckg! ggplot pckg! 8. Differential analysis with linear mixed models (lme4 and multcomp pckgs)! diff. cell population abundance! diff. marker expression!
7 Data description! Subset of data from the Bodenmiller et al study; used mass cytometry to measure PBMC's! from 8 different patients! after 11 different stimulation conditions as well as an unstimulated "Reference" state! Here, we use the samples that correspond to the BCR-crosslinking and Reference! For each sample, 10 cell surface markers and 14 signaling markers were measured!
8 Data pre-processing! The pre-processing steps in the Bodenmiller data, as we can download it, included removal of debris and de-barcoding! In general, pre-processing steps may involve:! normalization using bead standards! de-barcoding! compensation! CATALYST pckg!
9 Data import! All the data is available at the Robinson Lab server cytofworkflow/! Downloading using the download.file()! Metadata! FCS files! Panel file! Files with cluster merging!
10 Data import: expr! S1 c1 c2 c3 c4 M1 M2 M3 S2 c5 c6 c7 S3 c8 c9 c10 c11 c12
11 Data transformation! Mass cytometry Linear Arcsinh cofactor 5 Skewed distributions! Hard to distinguish between positive and negative populations! Magic transformation - arcsinh (hyperbolic inverse sine) with cofactor 5! linear for the lower expression,! logarithmic for the higher expression,! works for the negative and zero values (they do not have to be excluded from the analysis)! Figure adapted from Bendall et al. Science 2011!
12 Diagnostic plots quick look on! Plot with per-sample marker expression distributions, colored by condition! Identify problematic samples or markers! Cell counts in samples! MDS plot (or PCA plot)! Standard usage in the RNA-seq analysis! Generated with the plotmds() function from the limma pckg! As we want to plot samples we need to summarize the information about cells to the sample level!
13 MDS plot: expr_median_sample_tbl! M1 M2 M3 c1 S1 c2 c3 * * * c4 M1 M2 M3 c5 S1 * * * S2 c6 c7 * * * S2 * * * c8 c9 S3 * * * S3 c10 * * * c11 c12 * - median!
14 MDS plot!
15 Cell population identification! Manual gating! Comparison of clustering algorithms by Lukas Weber et al., Cytometry Part A, 2016! FlowSOM (+ ClusterConsensusPlus) one of the best performing methods and super fast! 3 steps:! 1. Building of the self-organizing map (SOM) with the BuildSOM function - cells are assigned according to their similarities to 100 grid points (or, so-called codes) of the SOM! 2. Building of a minimal spanning tree, which is mainly used for graphical representation of the clusters! 3. Metaclustering of the SOM codes performed directly with the ConsensusClusterPlus function!
16 Over-clustering! Some level of over-clustering is necessary, in order to detect somewhat rare populations! In addition, merging can always follow an overclustering step, but splitting of existing clusters is generally not feasible!
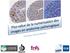
17 Over-clustering! Over-clustering into 20 groups! Median marker expression of data normalized to 0-1! (3.85%) 5 (1.89%) 20 (0.34%) 15 (7.8%) 12 (24.25%) 18 (1.85%) 3 (14.19%) 4 (1.97%) 9 (16.62%) 2 (2.26%) 17 (0.64%) 19 (1.51%) 8 (8.82%) 14 (2.2%) 10 (2.08%) 6 (3.58%) 7 (4.37%) 11 (0.64%) 13 (0.42%) 16 (0.72%) CD14 CD45 CD33 CD123 IgM CD20 HLA_DR CD3 CD4 CD7 cluster Figures adapted from Nowicka et al. F1000Research 2017!
18 Merging by an expert! cluster cluster_merging Median marker expression of data normalized to 0-1! CD CD CD HLA_DR CD IgM CD CD CD CD14 1 (3.85%) 5 (1.89%) 20 (0.34%) 15 (7.8%) 12 (24.25%) 18 (1.85%) 3 (14.19%) 4 (1.97%) 9 (16.62%) 2 (2.26%) 17 (0.64%) 19 (1.51%) 8 (8.82%) 14 (2.2%) 10 (2.08%) 6 (3.58%) 7 (4.37%) 11 (0.64%) 13 (0.42%) 16 (0.72%) cluster_merging B cells IgM+ B cells IgM CD4 T cells CD8 T cells DC NK cells monocytes surface Figures adapted from Nowicka et al. F1000Research 2017!
19 Statistical testing differential abundance!
20 Statistical testing differential abundance! Ref BCRXL cluster B cells IgM+ B cells IgM CD4 T cells CD8 T cells DC NK cells monocytes surface 0 Ref1 Ref2 Ref3 Ref4 Ref5 Ref6 Ref7 Ref8 BCRXL1 BCRXL2 BCRXL3 BCRXL4 BCRXL5 BCRXL6 BCRXL7 BCRXL8 proportion sample_id
21 proportion Statistical testing differential abundance! Ref1 Ref2 Ref3 Ref Ref4 Ref5 proportion Ref6 Ref7 Ref B cells IgM+ BCRXL B cells IgM CD4 T cells CD8 T cells BCRXL1 BCRXL2 BCRXL3 B cells IgM+ B cells IgM CD4 T cells CD8 T cells BCRXL4 BCRXL5 BCRXL6 BCRXL7 BCRXL cluster 2.0 B cells IgM B cells IgM CD4 T cells CD8 T cells DC DC NK cells NK cells monocytes surface condition monocytes Patient surface Patient condition DC NK cells monocytes surface sample_id 2.0 Ref BCRXL The generalized linear mixed model (GLMM) for each cell 1.5 population! proportion Y number 1.5 of CD4 cells! π proportion of CD4 cells! 1.0 m total number of cells! 0.5 i = 1,,8 patient ID! j = 1, 2 condition! observation-level random effects!! random intercept for each patient! proportion! condition20 Y Bin(m, ) Ref BCRXL logit( 1.5 ij )= x ij + ij + i condition ij N(0, i N(0, 2 ) 2 ) patient_id Patient1 Patient2 Patient3 Patient4 Patient5 Patient6 p c
22 Statistical testing differential expression of signaling markers!
23 Statistical testing differential expression of signaling markers!
24 ! proportio DC NK cells monocytes surface 25 Statistical testing differential expression 3.0 of signaling Patient8 markers! B cells IgM+ 0 B cells IgM CD4 T cells CD8 T cells CD4 monocytes T cells condition patient_id CD8 surface T cells Ref Patient BCRXL Patient Patient condition Patient Patient Patient6 DC NK cells monocytes surface 1 Patient Patient DC 2.5 NK cells condition Ref antigen BCRXL The linear mixed model 2 (LMM) for each cell population! condition Y median expression 1 of 1 proportion marker in CD4 cells! median_expression Median expression! pnfkb pp38 pstat5 pakt pstat1 pshp2 pzap70 pstat3 pslp76 pbtk pplcg2 perk Y N(µ, plat ps6 2 ) 1 Patient6 Patient7 pnfkb pp38 pstat5 pakt pstat1 pshp2 pzap70 pstat3 pslp76 pbtk i = 1,,8 patient ID! j = 1, 2 condition! random intercept for each patient! Y ij = x ij + ij + i monocytes ij N(0, i N(0, 2 ) 2 ) surface
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE
 Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
AbSeq on the BD Rhapsody system: Exploration of single-cell gene regulation by simultaneous digital mrna and protein quantification
 BD AbSeq on the BD Rhapsody system: Exploration of single-cell gene regulation by simultaneous digital mrna and protein quantification Overview of BD AbSeq antibody-oligonucleotide conjugates. High-throughput
BD AbSeq on the BD Rhapsody system: Exploration of single-cell gene regulation by simultaneous digital mrna and protein quantification Overview of BD AbSeq antibody-oligonucleotide conjugates. High-throughput
Title: High dimensional single cell analysis predicts response to anti-pd-1 immunotherapy. SUPPLEMENTARY INFORMATION - Content list
 Title: High dimensional single cell analysis predicts response to anti-pd- immunotherapy Authors: Carsten Krieg*, Malgorzata Nowicka,, Silvia Guglietta, Sabrina Schindler 5, Felix J. Hartmann, Lukas M.
Title: High dimensional single cell analysis predicts response to anti-pd- immunotherapy Authors: Carsten Krieg*, Malgorzata Nowicka,, Silvia Guglietta, Sabrina Schindler 5, Felix J. Hartmann, Lukas M.
Supplementary Notes. 2. Nine cell surface markers, CD33, CD20, CD3, CD4, CD7, CD123, CD14, IgM, and HLA-DR,
 Supplementary Notes Supplementary Note 1: Mass-tag cellular multiplexing workflow First, the cellular state is preserved by formaldehyde cross-linking. Second, the cells are permeabilized with methanol,
Supplementary Notes Supplementary Note 1: Mass-tag cellular multiplexing workflow First, the cellular state is preserved by formaldehyde cross-linking. Second, the cells are permeabilized with methanol,
New technologies for studying human immunity. Lisa Wagar Postdoctoral fellow, Mark Davis lab Stanford University School of Medicine
 New technologies for studying human immunity Lisa Wagar Postdoctoral fellow, Mark Davis lab Stanford University School of Medicine New strategies: Human immunology is ideal for a systems approach We have
New technologies for studying human immunity Lisa Wagar Postdoctoral fellow, Mark Davis lab Stanford University School of Medicine New strategies: Human immunology is ideal for a systems approach We have
Bangor University Laboratory Exercise 1, June 2008
 Laboratory Exercise, June 2008 Classroom Exercise A forest land owner measures the outside bark diameters at.30 m above ground (called diameter at breast height or dbh) and total tree height from ground
Laboratory Exercise, June 2008 Classroom Exercise A forest land owner measures the outside bark diameters at.30 m above ground (called diameter at breast height or dbh) and total tree height from ground
Next-Generation Immunohistochemistry: Multiplex tissue imaging with mass cytometry
 Nat Met, April 2014 Nat Med, April 2014 Next-Generation Immunohistochemistry: Multiplex tissue imaging with mass cytometry Journal Club Timo Böge Overview Introduction Conventional Immunohistochemistry
Nat Met, April 2014 Nat Med, April 2014 Next-Generation Immunohistochemistry: Multiplex tissue imaging with mass cytometry Journal Club Timo Böge Overview Introduction Conventional Immunohistochemistry
ANALYTISCHE STRATEGIE Tissue Imaging. Bernd Bodenmiller Institute of Molecular Life Sciences University of Zurich
 ANALYTISCHE STRATEGIE Tissue Imaging Bernd Bodenmiller Institute of Molecular Life Sciences University of Zurich Quantitative Breast single cancer cell analysis Switzerland Brain Breast Lung Colon-rectum
ANALYTISCHE STRATEGIE Tissue Imaging Bernd Bodenmiller Institute of Molecular Life Sciences University of Zurich Quantitative Breast single cancer cell analysis Switzerland Brain Breast Lung Colon-rectum
CyTOF analyses in rheumatoid arthritis. Deepak Rao, MD PhD Rheumatology, Immunology, Allergy, BWH
 CyTOF analyses in rheumatoid arthritis Deepak Rao, MD PhD Rheumatology, Immunology, Allergy, BWH Agenda: CyTOF analyses Analysis of synovial tissue Panel for analysis of synovial cells Comparison of CyTOF
CyTOF analyses in rheumatoid arthritis Deepak Rao, MD PhD Rheumatology, Immunology, Allergy, BWH Agenda: CyTOF analyses Analysis of synovial tissue Panel for analysis of synovial cells Comparison of CyTOF
Gene expression analysis. Roadmap. Microarray technology: how it work Applications: what can we do with it Preprocessing: Classification Clustering
 Gene expression analysis Roadmap Microarray technology: how it work Applications: what can we do with it Preprocessing: Image processing Data normalization Classification Clustering Biclustering 1 Gene
Gene expression analysis Roadmap Microarray technology: how it work Applications: what can we do with it Preprocessing: Image processing Data normalization Classification Clustering Biclustering 1 Gene
Chapter 1. Introduction
 Chapter 1 Introduction 1.1 Motivation and Goals The increasing availability and decreasing cost of high-throughput (HT) technologies coupled with the availability of computational tools and data form a
Chapter 1 Introduction 1.1 Motivation and Goals The increasing availability and decreasing cost of high-throughput (HT) technologies coupled with the availability of computational tools and data form a
Assigning B cell Maturity in Pediatric Leukemia Gabi Fragiadakis 1, Jamie Irvine 2 1 Microbiology and Immunology, 2 Computer Science
 Assigning B cell Maturity in Pediatric Leukemia Gabi Fragiadakis 1, Jamie Irvine 2 1 Microbiology and Immunology, 2 Computer Science Abstract One method for analyzing pediatric B cell leukemia is to categorize
Assigning B cell Maturity in Pediatric Leukemia Gabi Fragiadakis 1, Jamie Irvine 2 1 Microbiology and Immunology, 2 Computer Science Abstract One method for analyzing pediatric B cell leukemia is to categorize
Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD
 Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD Department of Biomedical Informatics Department of Computer Science and Engineering The Ohio State University Review
Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD Department of Biomedical Informatics Department of Computer Science and Engineering The Ohio State University Review
Mass Cytometry Applications for Immunology Research. Pub Note PN 13 01_150505
 Mass Cytometry Applications for Immunology Research Pub Note PN 13 01_150505 Contents Continuum of CD8 + T cell phenotypes revealed by deep profiling with mass cytometry 3 The transcriptional landscape
Mass Cytometry Applications for Immunology Research Pub Note PN 13 01_150505 Contents Continuum of CD8 + T cell phenotypes revealed by deep profiling with mass cytometry 3 The transcriptional landscape
The power of single cells: Building a tumor immune atlas Dana Pe er Department of Biological Science Department of Systems Biology Columbia
 The power of single cells: Building a tumor immune atlas Dana Pe er Department of Biological Science Department of Systems Biology Columbia University The Precision Medicine Initiative Most personalized
The power of single cells: Building a tumor immune atlas Dana Pe er Department of Biological Science Department of Systems Biology Columbia University The Precision Medicine Initiative Most personalized
Titrations in Cytobank
 The Premier Platform for Single Cell Analysis (1) Titrations in Cytobank To analyze data from a titration in Cytobank, the first step is to upload your own FCS files or clone an experiment you already
The Premier Platform for Single Cell Analysis (1) Titrations in Cytobank To analyze data from a titration in Cytobank, the first step is to upload your own FCS files or clone an experiment you already
Nature Immunology: doi: /ni Supplementary Figure 1. Examples of staining for each antibody used for the mass cytometry analysis.
 Supplementary Figure 1 Examples of staining for each antibody used for the mass cytometry analysis. To illustrate the functionality of each antibody probe, representative plots illustrating the expected
Supplementary Figure 1 Examples of staining for each antibody used for the mass cytometry analysis. To illustrate the functionality of each antibody probe, representative plots illustrating the expected
List of Figures. List of Tables. Preface to the Second Edition. Preface to the First Edition
 List of Figures List of Tables Preface to the Second Edition Preface to the First Edition xv xxv xxix xxxi 1 What Is R? 1 1.1 Introduction to R................................ 1 1.2 Downloading and Installing
List of Figures List of Tables Preface to the Second Edition Preface to the First Edition xv xxv xxix xxxi 1 What Is R? 1 1.1 Introduction to R................................ 1 1.2 Downloading and Installing
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INORMATION Activation of the reward system boosts innate and adaptive immunity Tamar L. Ben-Shaanan 1,2, Hilla Azulay-Debby 1,2, Tania Dubovik 1, Elina Starosvetsky 1, Ben Korin 1,2, Maya
SUPPLEMENTARY INORMATION Activation of the reward system boosts innate and adaptive immunity Tamar L. Ben-Shaanan 1,2, Hilla Azulay-Debby 1,2, Tania Dubovik 1, Elina Starosvetsky 1, Ben Korin 1,2, Maya
MHC MULTIMER PROFICIENCY PANEL 2015
 MHC MULTIMER PROFICIENCY PANEL 2015 July 2015 CONTACT Charlotte Halgreen ProficiencyPanel@immudex.com FOR MORE INFORMATION www.proficiencypanel.com MHC MULTIMER PROFICIENCY PANEL 2015 This report summarizes
MHC MULTIMER PROFICIENCY PANEL 2015 July 2015 CONTACT Charlotte Halgreen ProficiencyPanel@immudex.com FOR MORE INFORMATION www.proficiencypanel.com MHC MULTIMER PROFICIENCY PANEL 2015 This report summarizes
January 25, 2017 Scientific Research Process Name of Journal: ESPS Manuscript NO: Manuscript Type: Title: Authors: Correspondence to
 January 25, 2017 Scientific Research Process Name of Journal: World Journal of Gastroenterology ESPS Manuscript NO: 31928 Manuscript Type: ORIGINAL ARTICLE Title: Thiopurine use associated with reduced
January 25, 2017 Scientific Research Process Name of Journal: World Journal of Gastroenterology ESPS Manuscript NO: 31928 Manuscript Type: ORIGINAL ARTICLE Title: Thiopurine use associated with reduced
Digitizing the Proteomes From Big Tissue Biobanks
 Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
DURACLONE RE THERE ARE EVENTS YOU CANNOT AFFORD TO MISS
 DURACLONE RE THERE ARE EVENTS YOU CANNOT AFFORD TO MISS Your clinical research trial companion For Reseach Use Only - Not for use in Diagnostic procedures THERE ARE EVENTS YOU CANNOT AFFORD TO MISS Rare
DURACLONE RE THERE ARE EVENTS YOU CANNOT AFFORD TO MISS Your clinical research trial companion For Reseach Use Only - Not for use in Diagnostic procedures THERE ARE EVENTS YOU CANNOT AFFORD TO MISS Rare
Graphical display of population data
 Graphical display of population data E. Niclas Jonsson Division of Pharmacokinetics and Drug Therapy Department of Pharmaceutical Biosciences Uppsala University Nature of population data Hierarchical Variable
Graphical display of population data E. Niclas Jonsson Division of Pharmacokinetics and Drug Therapy Department of Pharmaceutical Biosciences Uppsala University Nature of population data Hierarchical Variable
Visual interpretation in pathology
 13 Visual interpretation in pathology Tissue architecture (alteration) evaluation e.g., for grading prostate cancer Immunohistochemistry (IHC) staining scoring e.g., HER2 in breast cancer (companion diagnostic
13 Visual interpretation in pathology Tissue architecture (alteration) evaluation e.g., for grading prostate cancer Immunohistochemistry (IHC) staining scoring e.g., HER2 in breast cancer (companion diagnostic
Spatially resolved multiparametric single cell analysis. Technical Journal Club 19th September 2017 Christina Müller (Group Speck)
 Spatially resolved multiparametric single cell analysis Technical Journal Club 19th September 2017 Christina Müller (Group Speck) Why spatially resolved multiparametric single cell analysis? Multiparametric
Spatially resolved multiparametric single cell analysis Technical Journal Club 19th September 2017 Christina Müller (Group Speck) Why spatially resolved multiparametric single cell analysis? Multiparametric
Data Analysis Using Regression and Multilevel/Hierarchical Models
 Data Analysis Using Regression and Multilevel/Hierarchical Models ANDREW GELMAN Columbia University JENNIFER HILL Columbia University CAMBRIDGE UNIVERSITY PRESS Contents List of examples V a 9 e xv " Preface
Data Analysis Using Regression and Multilevel/Hierarchical Models ANDREW GELMAN Columbia University JENNIFER HILL Columbia University CAMBRIDGE UNIVERSITY PRESS Contents List of examples V a 9 e xv " Preface
Accessing and Using ENCODE Data Dr. Peggy J. Farnham
 1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
MHC MULTIMER PROFICIENCY PANEL 2017
 MHC MULTIMER PROFICIENCY PANEL 2017 August 2017 CONTACT Charlotte Halgreen ProficiencyPanel@immudex.com FOR MORE INFORMATION www.proficiencypanel.com MHC MULTIMER PROFICIENCY PANEL 2017 This report summarizes
MHC MULTIMER PROFICIENCY PANEL 2017 August 2017 CONTACT Charlotte Halgreen ProficiencyPanel@immudex.com FOR MORE INFORMATION www.proficiencypanel.com MHC MULTIMER PROFICIENCY PANEL 2017 This report summarizes
Metabolomic and Proteomics Solutions for Integrated Biology. Christine Miller Omics Market Manager ASMS 2015
 Metabolomic and Proteomics Solutions for Integrated Biology Christine Miller Omics Market Manager ASMS 2015 Integrating Biological Analysis Using Pathways Protein A R HO R Protein B Protein X Identifies
Metabolomic and Proteomics Solutions for Integrated Biology Christine Miller Omics Market Manager ASMS 2015 Integrating Biological Analysis Using Pathways Protein A R HO R Protein B Protein X Identifies
Bayesian Inference for Single-cell ClUstering and ImpuTing (BISCUIT) Elham Azizi
 Bayesian Inference for Single-cell ClUstering and ImpuTing (BISCUIT) Elham Azizi BioC 2017: Where Software and Biology Connect Profiling Tumor-Immune Ecosystem in Breast Cancer Immunotherapy treatments
Bayesian Inference for Single-cell ClUstering and ImpuTing (BISCUIT) Elham Azizi BioC 2017: Where Software and Biology Connect Profiling Tumor-Immune Ecosystem in Breast Cancer Immunotherapy treatments
DURACLONE IF BE CERTAIN ABOUT THE RESPONSE. l res. a il n c n. For Research Use Only - Not for use in Diagnostic procedures
 DURACLONE IF earch tria l res lc a om ic il n c n nio pa Yo ur BE CERTAIN ABOUT THE RESPONSE For Research Use Only - Not for use in Diagnostic procedures BE CERTAIN ABOUT THE RESPONSE The sensitive and
DURACLONE IF earch tria l res lc a om ic il n c n nio pa Yo ur BE CERTAIN ABOUT THE RESPONSE For Research Use Only - Not for use in Diagnostic procedures BE CERTAIN ABOUT THE RESPONSE The sensitive and
User Guide. Association analysis. Input
 User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
Human Immunodeficiency Virus Type-1 Myeloid Derived Suppressor Cells Inhibit Cytomegalovirus Inflammation through Interleukin-27 and B7-H4
 Human Immunodeficiency Virus Type-1 Myeloid Derived Suppressor Cells Inhibit Cytomegalovirus Inflammation through Interleukin-27 and B7-H4 Ankita Garg, Rodney Trout and Stephen A. Spector,,* Department
Human Immunodeficiency Virus Type-1 Myeloid Derived Suppressor Cells Inhibit Cytomegalovirus Inflammation through Interleukin-27 and B7-H4 Ankita Garg, Rodney Trout and Stephen A. Spector,,* Department
BD CBA on the BD Accuri C6: Bringing Multiplexed Cytokine Detection to the Benchtop
 BD CBA on the BD Accuri C6: Bringing Multiplexed Cytokine Detection to the Benchtop Maria Dinkelmann, PhD Senior Marketing Applications Specialist BD Biosciences, Ann Arbor, MI 23-14380-00 Cellular Communication
BD CBA on the BD Accuri C6: Bringing Multiplexed Cytokine Detection to the Benchtop Maria Dinkelmann, PhD Senior Marketing Applications Specialist BD Biosciences, Ann Arbor, MI 23-14380-00 Cellular Communication
MethylMix An R package for identifying DNA methylation driven genes
 MethylMix An R package for identifying DNA methylation driven genes Olivier Gevaert May 3, 2016 Stanford Center for Biomedical Informatics Department of Medicine 1265 Welch Road Stanford CA, 94305-5479
MethylMix An R package for identifying DNA methylation driven genes Olivier Gevaert May 3, 2016 Stanford Center for Biomedical Informatics Department of Medicine 1265 Welch Road Stanford CA, 94305-5479
Class discovery in Gene Expression Data: Characterizing Splits by Support Vector Machines
 Class discovery in Gene Expression Data: Characterizing Splits by Support Vector Machines Florian Markowetz and Anja von Heydebreck Max-Planck-Institute for Molecular Genetics Computational Molecular Biology
Class discovery in Gene Expression Data: Characterizing Splits by Support Vector Machines Florian Markowetz and Anja von Heydebreck Max-Planck-Institute for Molecular Genetics Computational Molecular Biology
Supplementary Materials
 Supplementary Materials Supplementary figure 1. Taxonomic representation summarized at genus level. Fecal microbiota from a separate set of Jackson and Harlan mice prior to irradiation. A taxon was included
Supplementary Materials Supplementary figure 1. Taxonomic representation summarized at genus level. Fecal microbiota from a separate set of Jackson and Harlan mice prior to irradiation. A taxon was included
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
T. R. Golub, D. K. Slonim & Others 1999
 T. R. Golub, D. K. Slonim & Others 1999 Big Picture in 1999 The Need for Cancer Classification Cancer classification very important for advances in cancer treatment. Cancers of Identical grade can have
T. R. Golub, D. K. Slonim & Others 1999 Big Picture in 1999 The Need for Cancer Classification Cancer classification very important for advances in cancer treatment. Cancers of Identical grade can have
a Beckman Coulter Life Sciences: White Paper
 a Beckman Coulter Life Sciences: White Paper An 8-color DuraClone IM panel for detection of Human blood dendritic cells by flow cytometry Nathalie Dupas 1, Snehita Sattiraju 2, Neha Girish 2, Murthy Pendyala
a Beckman Coulter Life Sciences: White Paper An 8-color DuraClone IM panel for detection of Human blood dendritic cells by flow cytometry Nathalie Dupas 1, Snehita Sattiraju 2, Neha Girish 2, Murthy Pendyala
Clinical question. Screening tube. Diagnostic panel MRD. Clinical question
 OW CYTOMETRY UPDATES IN LYMPHOPROLIFERATIVE DISORDERS CANCER RESEARCH CENTER IBSAL UNIVERSITY & UNIVERSITY HOSPITAL, SALAMANCA (SPAIN) DISCLOSURES The EuroFlow Scientific Consortium Iamco-chairof receives
OW CYTOMETRY UPDATES IN LYMPHOPROLIFERATIVE DISORDERS CANCER RESEARCH CENTER IBSAL UNIVERSITY & UNIVERSITY HOSPITAL, SALAMANCA (SPAIN) DISCLOSURES The EuroFlow Scientific Consortium Iamco-chairof receives
WDHS Curriculum Map Probability and Statistics. What is Statistics and how does it relate to you?
 WDHS Curriculum Map Probability and Statistics Time Interval/ Unit 1: Introduction to Statistics 1.1-1.3 2 weeks S-IC-1: Understand statistics as a process for making inferences about population parameters
WDHS Curriculum Map Probability and Statistics Time Interval/ Unit 1: Introduction to Statistics 1.1-1.3 2 weeks S-IC-1: Understand statistics as a process for making inferences about population parameters
Mass Cytometry Publication List
 Mass Cytometry Publication List 2014 Publications 1 Becher, B., et al. High-dimensional analysis of the murine myeloid cell system. Nat Immunol 15 (12): 1181-1189, 2014. Behbehani, G.K., et al. Transient
Mass Cytometry Publication List 2014 Publications 1 Becher, B., et al. High-dimensional analysis of the murine myeloid cell system. Nat Immunol 15 (12): 1181-1189, 2014. Behbehani, G.K., et al. Transient
Supplemental Figure S1
 Supplemental Figure S1 Children from Nandi (low pathogen burden area) and Kisumu (high pathogen burden area) have significantly different pathogen-specific serological profiles. Serum antibody titers for
Supplemental Figure S1 Children from Nandi (low pathogen burden area) and Kisumu (high pathogen burden area) have significantly different pathogen-specific serological profiles. Serum antibody titers for
Statistics: A Brief Overview Part I. Katherine Shaver, M.S. Biostatistician Carilion Clinic
 Statistics: A Brief Overview Part I Katherine Shaver, M.S. Biostatistician Carilion Clinic Statistics: A Brief Overview Course Objectives Upon completion of the course, you will be able to: Distinguish
Statistics: A Brief Overview Part I Katherine Shaver, M.S. Biostatistician Carilion Clinic Statistics: A Brief Overview Course Objectives Upon completion of the course, you will be able to: Distinguish
Overview of role of immunologic markers in HIV diagnosis
 Overview of role of immunologic markers in HIV diagnosis Savita Pahwa, M.D. Departments of Microbiology & Immunology and Pediatrics University of Miami, Miller School of Medicine, Miami, Florida Background:
Overview of role of immunologic markers in HIV diagnosis Savita Pahwa, M.D. Departments of Microbiology & Immunology and Pediatrics University of Miami, Miller School of Medicine, Miami, Florida Background:
Supplemental Information. Aryl Hydrocarbon Receptor Controls. Monocyte Differentiation. into Dendritic Cells versus Macrophages
 Immunity, Volume 47 Supplemental Information Aryl Hydrocarbon Receptor Controls Monocyte Differentiation into Dendritic Cells versus Macrophages Christel Goudot, Alice Coillard, Alexandra-Chloé Villani,
Immunity, Volume 47 Supplemental Information Aryl Hydrocarbon Receptor Controls Monocyte Differentiation into Dendritic Cells versus Macrophages Christel Goudot, Alice Coillard, Alexandra-Chloé Villani,
AP Statistics. Semester One Review Part 1 Chapters 1-5
 AP Statistics Semester One Review Part 1 Chapters 1-5 AP Statistics Topics Describing Data Producing Data Probability Statistical Inference Describing Data Ch 1: Describing Data: Graphically and Numerically
AP Statistics Semester One Review Part 1 Chapters 1-5 AP Statistics Topics Describing Data Producing Data Probability Statistical Inference Describing Data Ch 1: Describing Data: Graphically and Numerically
Next-Gen Analytics in Digital Pathology
 Next-Gen Analytics in Digital Pathology Cliff Hoyt, CTO Cambridge Research & Instrumentation April 29, 2010 Seeing life in a new light 1 Digital Pathology Today Acquisition, storage, dissemination, remote
Next-Gen Analytics in Digital Pathology Cliff Hoyt, CTO Cambridge Research & Instrumentation April 29, 2010 Seeing life in a new light 1 Digital Pathology Today Acquisition, storage, dissemination, remote
Supplementary Figure 1. Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
 Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Introduction to LOH and Allele Specific Copy Number User Forum
 Introduction to LOH and Allele Specific Copy Number User Forum Jonathan Gerstenhaber Introduction to LOH and ASCN User Forum Contents 1. Loss of heterozygosity Analysis procedure Types of baselines 2.
Introduction to LOH and Allele Specific Copy Number User Forum Jonathan Gerstenhaber Introduction to LOH and ASCN User Forum Contents 1. Loss of heterozygosity Analysis procedure Types of baselines 2.
Introduction to Machine Learning. Katherine Heller Deep Learning Summer School 2018
 Introduction to Machine Learning Katherine Heller Deep Learning Summer School 2018 Outline Kinds of machine learning Linear regression Regularization Bayesian methods Logistic Regression Why we do this
Introduction to Machine Learning Katherine Heller Deep Learning Summer School 2018 Outline Kinds of machine learning Linear regression Regularization Bayesian methods Logistic Regression Why we do this
CNV PCA Search Tutorial
 CNV PCA Search Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Data Preparation 2 A. Join Log Ratio Data with Phenotype Information.............................. 2 B. Activate only
CNV PCA Search Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Data Preparation 2 A. Join Log Ratio Data with Phenotype Information.............................. 2 B. Activate only
AC40. The true clinical. hybrid
 AC40 The true clinical hybrid The true clinical hybrid audiometer Stand alone & PCbased audiometry in one box A true hybrid audiometer The AC40 full two channel audiometer includes all the advanced features
AC40 The true clinical hybrid The true clinical hybrid audiometer Stand alone & PCbased audiometry in one box A true hybrid audiometer The AC40 full two channel audiometer includes all the advanced features
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity
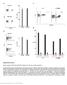 Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
AVENIO ctdna Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB
 Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB Analysis Kits Next-generation performance in liquid biopsies 2 Accelerating clinical research From liquid biopsy to next-generation
Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB Analysis Kits Next-generation performance in liquid biopsies 2 Accelerating clinical research From liquid biopsy to next-generation
The Simulacrum. What is it, how is it created, how does it work? Michael Eden on behalf of Sally Vernon & Cong Chen NAACCR 21 st June 2017
 The Simulacrum What is it, how is it created, how does it work? Michael Eden on behalf of Sally Vernon & Cong Chen NAACCR 21 st June 2017 sally.vernon@phe.gov.uk & cong.chen@phe.gov.uk 1 Overview What
The Simulacrum What is it, how is it created, how does it work? Michael Eden on behalf of Sally Vernon & Cong Chen NAACCR 21 st June 2017 sally.vernon@phe.gov.uk & cong.chen@phe.gov.uk 1 Overview What
Chapter 9. Tests, Procedures, and Diagnosis Codes The McGraw-Hill Companies, Inc. All rights reserved.
 Chapter 9 Tests, Procedures, and Diagnosis Codes Chapter 9 Content: Overview Ordering A Test SpringLabsTM & Reference Lab Results Managing and Charting Tests Creating A New Test Documenting and Activating
Chapter 9 Tests, Procedures, and Diagnosis Codes Chapter 9 Content: Overview Ordering A Test SpringLabsTM & Reference Lab Results Managing and Charting Tests Creating A New Test Documenting and Activating
Equinox 2.0 PC-based audiometer. Not just an audiometer - A full counseling and reporting solution
 Equinox 2.0 PC-based audiometer Not just an audiometer - A full counseling and reporting solution The PC-based Advantage The Equinox 2.0 is a state of the art two channel clinical audiometer. Counseling
Equinox 2.0 PC-based audiometer Not just an audiometer - A full counseling and reporting solution The PC-based Advantage The Equinox 2.0 is a state of the art two channel clinical audiometer. Counseling
Agilent GeneSpring/MPP Metadata Analysis Framework
 Agilent GeneSpring/MPP Metadata Analysis Framework Technical Overview Authors Srikanthi R., Pritha Aggarwal, Durairaj R., Maria Kammerer, and Pramila Tata Strand Life Sciences Bangalore, India Michael
Agilent GeneSpring/MPP Metadata Analysis Framework Technical Overview Authors Srikanthi R., Pritha Aggarwal, Durairaj R., Maria Kammerer, and Pramila Tata Strand Life Sciences Bangalore, India Michael
Nature Immunology: doi: /ni Supplementary Figure 1. RNA-Seq analysis of CD8 + TILs and N-TILs.
 Supplementary Figure 1 RNA-Seq analysis of CD8 + TILs and N-TILs. (a) Schematic representation of the tumor and cell types used for the study. HNSCC, head and neck squamous cell cancer; NSCLC, non-small
Supplementary Figure 1 RNA-Seq analysis of CD8 + TILs and N-TILs. (a) Schematic representation of the tumor and cell types used for the study. HNSCC, head and neck squamous cell cancer; NSCLC, non-small
Systematic Analysis of Cell-to-Cell Expression Variation of T Lymphocytes in a Human Cohort Identifies Aging and Genetic Associations
 Resource Systematic Analysis of Cell-to-Cell Expression Variation of T Lymphocytes in a Human Cohort Identifies Aging and Genetic Associations Graphical Abstract Authors Yong Lu, Angelique Biancotto, Foo
Resource Systematic Analysis of Cell-to-Cell Expression Variation of T Lymphocytes in a Human Cohort Identifies Aging and Genetic Associations Graphical Abstract Authors Yong Lu, Angelique Biancotto, Foo
Bayesian Nonparametric Methods for Precision Medicine
 Bayesian Nonparametric Methods for Precision Medicine Brian Reich, NC State Collaborators: Qian Guan (NCSU), Eric Laber (NCSU) and Dipankar Bandyopadhyay (VCU) University of Illinois at Urbana-Champaign
Bayesian Nonparametric Methods for Precision Medicine Brian Reich, NC State Collaborators: Qian Guan (NCSU), Eric Laber (NCSU) and Dipankar Bandyopadhyay (VCU) University of Illinois at Urbana-Champaign
ENZYME ACTIVITY. Readings: Review pp , and in your text (POHS, 5 th ed.).
 ENZYME ACTIVITY Readings: Review pp. 51-58, and 128-139 in your text (POHS, 5 th ed.). Introduction Enzymes are biological catalysts; that is, enzymes are able to mediate the conversion of substrate into
ENZYME ACTIVITY Readings: Review pp. 51-58, and 128-139 in your text (POHS, 5 th ed.). Introduction Enzymes are biological catalysts; that is, enzymes are able to mediate the conversion of substrate into
Diurnal Pattern of Reaction Time: Statistical analysis
 Diurnal Pattern of Reaction Time: Statistical analysis Prepared by: Alison L. Gibbs, PhD, PStat Prepared for: Dr. Principal Investigator of Reaction Time Project January 11, 2015 Summary: This report gives
Diurnal Pattern of Reaction Time: Statistical analysis Prepared by: Alison L. Gibbs, PhD, PStat Prepared for: Dr. Principal Investigator of Reaction Time Project January 11, 2015 Summary: This report gives
EXPression ANalyzer and DisplayER
 EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
VUmc Basispresentatie
 Clinical diagnostic cytometry Gerrit J Schuurhuis Dept of Hematology VU University Medical Center Amsterdam, Netherlands Use of immunophenotyping at diagnosis to trace residual disease after therapy 1.
Clinical diagnostic cytometry Gerrit J Schuurhuis Dept of Hematology VU University Medical Center Amsterdam, Netherlands Use of immunophenotyping at diagnosis to trace residual disease after therapy 1.
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded
 Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
RNA-seq: filtering, quality control and visualisation. COMBINE RNA-seq Workshop
 RNA-seq: filtering, quality control and visualisation COMBINE RNA-seq Workshop QC and visualisation (part 1) Slide taken from COMBINE RNAseq workshop on 23/09/2016 RNA-seq of Mouse mammary gland Basal
RNA-seq: filtering, quality control and visualisation COMBINE RNA-seq Workshop QC and visualisation (part 1) Slide taken from COMBINE RNAseq workshop on 23/09/2016 RNA-seq of Mouse mammary gland Basal
A step-by-step approach to build and analyze a multicolor panel
 Analyze A step-by-step approach to build and analyze a multicolor panel For Research Use Only. Not for use in diagnostic or therapeutic procedures. Alexa Fluor is a registered trademark of Life Technologies
Analyze A step-by-step approach to build and analyze a multicolor panel For Research Use Only. Not for use in diagnostic or therapeutic procedures. Alexa Fluor is a registered trademark of Life Technologies
RNA-seq Introduction
 RNA-seq Introduction DNA is the same in all cells but which RNAs that is present is different in all cells There is a wide variety of different functional RNAs Which RNAs (and sometimes then translated
RNA-seq Introduction DNA is the same in all cells but which RNAs that is present is different in all cells There is a wide variety of different functional RNAs Which RNAs (and sometimes then translated
STATISTICS & PROBABILITY
 STATISTICS & PROBABILITY LAWRENCE HIGH SCHOOL STATISTICS & PROBABILITY CURRICULUM MAP 2015-2016 Quarter 1 Unit 1 Collecting Data and Drawing Conclusions Unit 2 Summarizing Data Quarter 2 Unit 3 Randomness
STATISTICS & PROBABILITY LAWRENCE HIGH SCHOOL STATISTICS & PROBABILITY CURRICULUM MAP 2015-2016 Quarter 1 Unit 1 Collecting Data and Drawing Conclusions Unit 2 Summarizing Data Quarter 2 Unit 3 Randomness
The Role of Face Parts in Gender Recognition
 The Role of Face Parts in Gender Recognition Yasmina Andreu Ramón A. Mollineda Pattern Analysis and Learning Section Computer Vision Group University Jaume I of Castellón (Spain) Y. Andreu, R.A. Mollineda
The Role of Face Parts in Gender Recognition Yasmina Andreu Ramón A. Mollineda Pattern Analysis and Learning Section Computer Vision Group University Jaume I of Castellón (Spain) Y. Andreu, R.A. Mollineda
GPS Observation of Continent-size (or super-size ) Traveling TEC Pulsations at the Start of Geomagnetic Storms
 GPS Observation of Continent-size (or super-size ) Traveling TEC Pulsations at the Start of Geomagnetic Storms Rezy Pradipta Boston College Institute for Scientific Research Tuesday, 18 November 2014 An
GPS Observation of Continent-size (or super-size ) Traveling TEC Pulsations at the Start of Geomagnetic Storms Rezy Pradipta Boston College Institute for Scientific Research Tuesday, 18 November 2014 An
TITLE: Immunology, Systems Biology, and Immunotherapy of Breast Cancer. Stanford, CA 94305
 AD AWARD NUMBER: W81XWH-06-1-0417 TITLE: Immunology, Systems Biology, and Immunotherapy of Breast Cancer PRINCIPAL INVESTIGATOR: Peter P. Lee, M.D. CONTRACTING ORGANIZATION: Stanford University Stanford,
AD AWARD NUMBER: W81XWH-06-1-0417 TITLE: Immunology, Systems Biology, and Immunotherapy of Breast Cancer PRINCIPAL INVESTIGATOR: Peter P. Lee, M.D. CONTRACTING ORGANIZATION: Stanford University Stanford,
Development of the Web-Based Child Asthma Risk Assessment Tool
 Development of the Web-Based Child Asthma Risk Assessment Tool Herman Mitchell, Ph.D. Rho, Inc Chapel Hill, NC hmitchell@rhoworld.com (919) 408-8000 x223 Overview: One of the clearest results of the Phase
Development of the Web-Based Child Asthma Risk Assessment Tool Herman Mitchell, Ph.D. Rho, Inc Chapel Hill, NC hmitchell@rhoworld.com (919) 408-8000 x223 Overview: One of the clearest results of the Phase
SUPPLEMENTARY APPENDIX
 SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
International Journal of Computer Science Trends and Technology (IJCST) Volume 5 Issue 1, Jan Feb 2017
 RESEARCH ARTICLE Classification of Cancer Dataset in Data Mining Algorithms Using R Tool P.Dhivyapriya [1], Dr.S.Sivakumar [2] Research Scholar [1], Assistant professor [2] Department of Computer Science
RESEARCH ARTICLE Classification of Cancer Dataset in Data Mining Algorithms Using R Tool P.Dhivyapriya [1], Dr.S.Sivakumar [2] Research Scholar [1], Assistant professor [2] Department of Computer Science
Figure S2. Distribution of acgh probes on all ten chromosomes of the RIL M0022
 96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
DeconRNASeq: A Statistical Framework for Deconvolution of Heterogeneous Tissue Samples Based on mrna-seq data
 DeconRNASeq: A Statistical Framework for Deconvolution of Heterogeneous Tissue Samples Based on mrna-seq data Ting Gong, Joseph D. Szustakowski April 30, 2018 1 Introduction Heterogeneous tissues are frequently
DeconRNASeq: A Statistical Framework for Deconvolution of Heterogeneous Tissue Samples Based on mrna-seq data Ting Gong, Joseph D. Szustakowski April 30, 2018 1 Introduction Heterogeneous tissues are frequently
Applied Analysis of Variance and Experimental Design. Lukas Meier, Seminar für Statistik
 Applied Analysis of Variance and Experimental Design Lukas Meier, Seminar für Statistik About Me Studied mathematics at ETH. Worked at the statistical consulting service and did a PhD in statistics (at
Applied Analysis of Variance and Experimental Design Lukas Meier, Seminar für Statistik About Me Studied mathematics at ETH. Worked at the statistical consulting service and did a PhD in statistics (at
Laboratory. Mendelian Genetics
 Laboratory 9 Mendelian Genetics Biology 171L FA17 Lab 9: Mendelian Genetics Student Learning Outcomes 1. Predict the phenotypic and genotypic ratios of a monohybrid cross. 2. Determine whether a gene is
Laboratory 9 Mendelian Genetics Biology 171L FA17 Lab 9: Mendelian Genetics Student Learning Outcomes 1. Predict the phenotypic and genotypic ratios of a monohybrid cross. 2. Determine whether a gene is
Cancer Informatics Lecture
 Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Data mining with Ensembl Biomart. Stéphanie Le Gras
 Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
DuraClone IM. Standardized phenotyping panels for studies of the human immune system
 DuraClone IM Standardized phenotyping panels for studies of the human immune system Standardize with the experts, adopt DuraClone IM Tubes. Clinical research studies require accurate, reproducible results
DuraClone IM Standardized phenotyping panels for studies of the human immune system Standardize with the experts, adopt DuraClone IM Tubes. Clinical research studies require accurate, reproducible results
LAB ASSIGNMENT 4 INFERENCES FOR NUMERICAL DATA. Comparison of Cancer Survival*
 LAB ASSIGNMENT 4 1 INFERENCES FOR NUMERICAL DATA In this lab assignment, you will analyze the data from a study to compare survival times of patients of both genders with different primary cancers. First,
LAB ASSIGNMENT 4 1 INFERENCES FOR NUMERICAL DATA In this lab assignment, you will analyze the data from a study to compare survival times of patients of both genders with different primary cancers. First,
Flow Cytometry Critical Assessment of Population Identification Methods
 Flow Cytometry Critical Assessment of Population Identification Methods Richard H. Scheuermann, Ph.D. Director of Informatics J. Craig Venter Institute Single Cell Phenotypes Different cell types play
Flow Cytometry Critical Assessment of Population Identification Methods Richard H. Scheuermann, Ph.D. Director of Informatics J. Craig Venter Institute Single Cell Phenotypes Different cell types play
Application of Local Control Strategy in analyses of the effects of Radon on Lung Cancer Mortality for 2,881 US Counties
 Application of Local Control Strategy in analyses of the effects of Radon on Lung Cancer Mortality for 2,881 US Counties Bob Obenchain, Risk Benefit Statistics, August 2015 Our motivation for using a Cut-Point
Application of Local Control Strategy in analyses of the effects of Radon on Lung Cancer Mortality for 2,881 US Counties Bob Obenchain, Risk Benefit Statistics, August 2015 Our motivation for using a Cut-Point
Simple, rapid, and reliable RNA sequencing
 Simple, rapid, and reliable RNA sequencing RNA sequencing applications RNA sequencing provides fundamental insights into how genomes are organized and regulated, giving us valuable information about the
Simple, rapid, and reliable RNA sequencing RNA sequencing applications RNA sequencing provides fundamental insights into how genomes are organized and regulated, giving us valuable information about the
ChIP-seq hands-on. Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs
 ChIP-seq hands-on Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs Main goals Becoming familiar with essential tools and formats Visualizing and contextualizing raw data Understand
ChIP-seq hands-on Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs Main goals Becoming familiar with essential tools and formats Visualizing and contextualizing raw data Understand
USER GUIDE: NEW CIR APP. Technician User Guide
 USER GUIDE: NEW CIR APP. Technician User Guide 0 Table of Contents 1 A New CIR User Interface Why?... 3 2 How to get started?... 3 3 Navigating the new CIR app. user interface... 6 3.1 Introduction...
USER GUIDE: NEW CIR APP. Technician User Guide 0 Table of Contents 1 A New CIR User Interface Why?... 3 2 How to get started?... 3 3 Navigating the new CIR app. user interface... 6 3.1 Introduction...
UN Handbook Ch. 7 'Managing sources of non-sampling error': recommendations on response rates
 JOINT EU/OECD WORKSHOP ON RECENT DEVELOPMENTS IN BUSINESS AND CONSUMER SURVEYS Methodological session II: Task Force & UN Handbook on conduct of surveys response rates, weighting and accuracy UN Handbook
JOINT EU/OECD WORKSHOP ON RECENT DEVELOPMENTS IN BUSINESS AND CONSUMER SURVEYS Methodological session II: Task Force & UN Handbook on conduct of surveys response rates, weighting and accuracy UN Handbook
Technical Specifications
 Technical Specifications In order to provide summary information across a set of exercises, all tests must employ some form of scoring models. The most familiar of these scoring models is the one typically
Technical Specifications In order to provide summary information across a set of exercises, all tests must employ some form of scoring models. The most familiar of these scoring models is the one typically
Chapter 1: Managing workbooks
 Chapter 1: Managing workbooks Module A: Managing worksheets You use the Insert tab on the ribbon to insert new worksheets. True or False? Which of the following are options for moving or copying a worksheet?
Chapter 1: Managing workbooks Module A: Managing worksheets You use the Insert tab on the ribbon to insert new worksheets. True or False? Which of the following are options for moving or copying a worksheet?
Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit
 APPLICATION NOTE Ion PGM System Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit Key findings The Ion PGM System, in concert with the Ion ReproSeq PGS View Kit and Ion Reporter
APPLICATION NOTE Ion PGM System Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit Key findings The Ion PGM System, in concert with the Ion ReproSeq PGS View Kit and Ion Reporter
Tutorial: RNA-Seq Analysis Part II: Non-Specific Matches and Expression Measures
 : RNA-Seq Analysis Part II: Non-Specific Matches and Expression Measures March 15, 2013 CLC bio Finlandsgade 10-12 8200 Aarhus N Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com support@clcbio.com
: RNA-Seq Analysis Part II: Non-Specific Matches and Expression Measures March 15, 2013 CLC bio Finlandsgade 10-12 8200 Aarhus N Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com support@clcbio.com
EBCC Data Analysis Tool (EBCC DAT) Introduction
 Instructor: Paul Wolfgang Faculty sponsor: Yuan Shi, Ph.D. Andrey Mavrichev CIS 4339 Project in Computer Science May 7, 2009 Research work was completed in collaboration with Michael Tobia, Kevin L. Brown,
Instructor: Paul Wolfgang Faculty sponsor: Yuan Shi, Ph.D. Andrey Mavrichev CIS 4339 Project in Computer Science May 7, 2009 Research work was completed in collaboration with Michael Tobia, Kevin L. Brown,
Multiple Treatments on the Same Experimental Unit. Lukas Meier (most material based on lecture notes and slides from H.R. Roth)
 Multiple Treatments on the Same Experimental Unit Lukas Meier (most material based on lecture notes and slides from H.R. Roth) Introduction We learned that blocking is a very helpful technique to reduce
Multiple Treatments on the Same Experimental Unit Lukas Meier (most material based on lecture notes and slides from H.R. Roth) Introduction We learned that blocking is a very helpful technique to reduce
Using Multiplex Assays and Assay Development to Enhance your Research Program James A. Lederer, PhD
 Using Multiplex Assays and Assay Development to Enhance your Research Program James A. Lederer, PhD Department of Surgery Brigham and Women s Hospital (BWH), BWH Biomedical Research Institute (BWH-BRI)
Using Multiplex Assays and Assay Development to Enhance your Research Program James A. Lederer, PhD Department of Surgery Brigham and Women s Hospital (BWH), BWH Biomedical Research Institute (BWH-BRI)
