Nature Genetics: doi: /ng Supplementary Figure 1. Alternative splicing events in the 5K panel.
|
|
|
- Holly Farmer
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1 Alternative splicing events in the 5K panel. The majority of cryptic splicing occurred by creation of an AG or GT (Type I). While some other mutations increased the usage of a nearby weaker splice-site (type II). Very few mutations were found to abolish alternative splice-site usage (type III).
2 Supplementary Figure 2 MaPSy performance. (a d) Agreement between allelic splicing ratios (log 2 ) of three cell culture replicates of MaPSy in vivo (a c) and two experimental replicates of MaPSy in vitro (d). (e) Stacked histogram of mutant (red) and wild-type (blue) relative splicing efficiency in MaPSy in vivo (top) and in vitro (bottom). (f) Full gel of output (spliced species) from MaPSy in vivo.
3 Supplementary Figure 3 MaPSy validation in patient samples and ENCODE data. (a f) MaPSy s identified ESMs in mutations causing inosine triphosphatase deficiency (a), galactosemia (b), haemorrhagic telangiectasia (c), Menkes syndrome (d) and Barth syndrome (e,f) were shown to exhibit splicing aberrations (exon skipping and/or intron retention) in RNAs derived from patient tissue samples. (g) Splicing efficiency in MaPSy corresponds to splicing in ENCODE data.
4 Supplementary Figure 4 Mode of inheritance in the 5K panel. (a) Percent ESM in the 5K panel stratified by modes of inheritance in haploinsufficient genes (prediction score = 1), haplosufficient genes (prediction score < 0.7) and moderately haploinsufficient genes (1 > prediction score 0.7) 8. Error bars, 95% confidence intervals. (b) Number of mutations in the different modes of inheritance in the 5K panel.
5 Supplementary Figure 5 Genes intolerant to protein-truncating variants (PTVs) in the ExAC population are predisposed to disease-associated splicing mutations. (a) Mean fraction of ESMs in PTV-intolerant (pli 0.9), semitolerant (0.1 < pli < 0.9) and tolerant (pli 0.1) genes in dominant and recessive traits. Error bars, s.e.m. (b) PTV-intolerant genes also have more introns than other genes, similar to disease genes that lose function via splicing mutations.
6 Supplementary Figure 6 Features of splicing. (a) The mean of relative splicing efficiency of wild-type species in vivo (n = 2,086) is plotted against increasing mean of feature measures in sliding window (size = 200, step = 1). Shaded regions represent 95% confidence intervals. Intron length is plotted on a log 10 scale. The mean of PhastCons score for all bases of the exon was used to measure conservation. Genomic features that have previously been associated with splicing are shown to display similar trends in MaPSy. P values were obtained from linear regression analyses. (b) The 5K panel is divided into five bins of increasing feature measures, and percent ESM in each bin is plotted. Error bars, 95% confidence intervals. Low differential GC content between exon and intron, less ESE, more ESS and less agreement with splicesite consensus sequence, which are all associated with weaker splicing are shown to sensitize exons to ESM. The Kruskal Wallis test was used to obtain P values.
7 Supplementary Figure 7 The role of PTBP1 and SRSF1 in ESM phenotypes. (a) The splicing phenotype of a mutation in exon 20 of COL1A2 that creates a PTBP1-binding motif was partially rescued when PTBP1 was knocked down. (b) A mutation that weaken a SRSF1-binding motif in exon 8 of MLH1 caused a modest but not significant increase of skipping events in the absence of SRSF1, whereas the wild-type exon that contains a SRSF1-binding site had a significant increase in skipping events when SRSF1 was knocked down.
8 Supplementary Figure 8 Overlap of intronic and exonic splicing regulatory motifs. (a) The density for each RBP motif was calculated in all wild-type species (n = 2,048). (b) Clustering of intronic data reveals similar trends in vivo and in vitro. (c) Intronic splicing activators and exonic splicing repressors show a high degree of overlap. (d) Intronic splicing repressor motifs and exonic splicing activator motifs display a high degree of overlap. (e) Table of exonic splicing repressors and exonic splicing activators that exhibit the same function in vivo and in vitro.
9 Supplementary Figure 9 In vitro functional SELEX. (a) Series of functional SELEX with MaPSy. (b) Mutant/wild type ratio in the B/C fraction in comparison to spliced species (left) and in the A fraction in comparison to spliced species (right). Enrichment in B/C complex is positively correlated with splicing, while enrichment in A complex is negatively correlated with splicing. (c) Clustering the effects of exonic mutation disruptions on different stages of spliceosomal assembly revealed mechanistic signatures of ESM. Only clusters with 8 members are shown.
10 Supplementary Figure 10 Mutant feature analyses in different clusters revealed distinct ESM mechanistic signatures. Horizontal dotted lines indicate the mean value of the features in the 5K panel. Box plots of feature values that are significantly different than background (permuted cluster assignment) are colored red. The medians are indicated as horizontal bold lines, and the means as black hollow dots.
11 Supplementary Figure 11 ESM visualization browser. A web browser was developed to visualize raw counts and information on individual mutations from original publications. Mutations can be queried by HGMD ID, gene or author.
12 A b Supplementary Figure 12 Common sequences of the 5K panel reporters. (a) In vivo reporter sequence: CMV enhancer and promoter sequence (blue), adenovirus sequence (green; exon in uppercase and intron in lowercase), 200-mer library (red), ACTN1 intron 15 (lowercase, purple) and exon 16 (uppercase, purple), bgh poly(a) (cyan). (b) In vitro reporter sequence: adenovirus sequence (green; including T7 promoter sequence in bold), 200-mer oligo library (red) and additional intronic sequence (purple).
13 Supplementary Table 2. Summary of MaPSy validation in patient samples HGMD or SNP id gene hg19 position nucleotide change AA change MaPSy result validation result Patient tissue/ RNA source class CM ATM chr11: :+ c.6095g>a p.r2032k loss of splicing in vivo (0 fold) and in vitro (0 fold) exon 44 skipping lymphoblastoid cell line TP loss of splicing in vivo (0 fold) and in vitro (0 fold), but 46 CM ATM chr11: :+ c.5932g>t p.e1978* exon 42 skipping lymphoblastoid cell line FN neither were significant due to low counts CM ATP7A chrx: :+ c.1933c>t p.r645* loss of splicing in vivo (0.4 fold) and in vitro (0.4 fold) exon 8 skipping fibroblasts TP Coriell CM BRCA1 chr17: :- c.211a>g p.r71g loss of splicing in vivo (0 fold) and in vitro (0.5 fold) exon 5 skipping T-lymphocytes TP CM BRCA1 chr17: :- c.5123c>a p.a1708e loss of splicing in vivo (0.3 fold) and in vitro (0.3 fold) exon 18 skipping T-lymphocytes TP CM BRCA1 chr17: :- c.5080g>t p.e1694* loss of splicing in vivo (0.03 fold) and in vitro (0.1 fold) exon 18 skipping T-lymphocytes TP Reference / source CM BRCA1 chr17: :- c.131g>a p.c44y no change no change lymphoblastoid cell line TN Amanda Toland CM BRCA2 chr13: :+ c.673a>g p.t225a no change no change T-lymphocytes TN CM BRCA2 chr13: :+ c.7007g>a p.r2336h loss of splicing in vivo (0.03 fold) and in vitro (0.03 fold) exon 13 skipping T-lymphocytes TP CM COL1A1 chr9: :- c.608g>t p.g203v no change no change fibroblast TN Joan Marini CM COL1A1 chr9: :- c.634g>a p.g212r Loss of splicing in vivo (0.1 fold) and in vitro (0.2 fold) no change fibroblast FP Joan Marini CM ENG chr9: :- c.1687g>t p.e563* loss of splicing in vivo (0.38 fold) and in vitro (0.2 fold) loss of exon 13 T-lymphocytes TP Authors CM FBN1 chr15: :- c.6339t>g p.y2113* loss of splicing in vivo (0.04 fold) and in vitro (0.2 fold) exon 51 skipping fibroblasts TP CM FBN1 chr15: :- c.4766g>t p.c1589f loss of splicing in vivo (0.3 fold) and in vitro (0.6 fold) no change fibroblasts FP Coriell CM GALT chr9: :+ c.563a>g p.q188r loss of splicing in vivo (0.01 fold) and in vitro (0.4 fold) missplicing lymphoblastoid cell line TP Coriell CM HPRT1 chrx: :+ c.590a>t p.e197v loss of splicing in vivo (0 fold) and in vitro (0.3 fold) exon 8 skipping lymphoblastoid cell line TP CM HPRT1 chrx: :+ c.602a>t p.d201v loss of splicing in vivo (0.2 fold) and in vitro (0.3 fold) exon 8 skipping T-lymphocytes TP CM HPRT1 chrx: :+ c.569g>a p.g190e no change no change lymphoblastoid cell line TN CM HPRT1 chrx: :+ c.601g>a p.d201n no change no change lymphoblastoid cell line TN CM HPRT1 chrx: :+ c.601g>t p.d201y loss of splicing in vivo (0.18 fold) but not in vitro no change CM ITPA chr20: :+ c.94c>a p.p32t loss of splicing in vivo (0.1 fold) and in vitro (0.3 fold) exon 3 skipping lymphoblastoid cell line and T lymphocytes lymphoblastoid cell line and T-lymphocytes CM MLH1 chr3: :+ c.677g>a p.r226q loss of splicing in vivo (0 fold) and in vitro (0.01 fold) loss of exon 8 whole blood TP CM MLH1 chr3: :+ c.350c>t p.t117m no change no change lymphoblastoid cell line TN CM MLH1 chr3: :+ c.338t>a p.v113d loss of splicing in vivo (0.6 fold) but no change in vitro no change lymphoblastoid cell line TN CM MLH1 chr3: :+ c.637g>a p.v213m loss of splicing in vivo (0.6 fold) and in vitro (0.65 fold) no change lymphoblastoid cell line FP CM PSEN1 chr14: :+ c.548g>t p.g1883v loss of splicing in vivo (0.3 fold) and in vitro (0.1 fold) brain-specific loss of exons 6 and 7 frontal cortex (post mortem) TP CM PSEN1 chr14: :+ c.871a>c p.t291p no change exon 9 skipping Whole blood FN CM PSEN1 chr14: :+ c.509c>t ps170f no change in vivo but loss of splicing in vitro (0.5 fold) no change frontal cortex and cerebellum (post mortem) TN rs RBM23 chr14: :- c.408g>a p.r136 loss of splicing in vivo (0.3 fold) and in vitro (0.5 fold) exon 6 skipping lymphoblastoid cell line TP CM SLC12A3 chr16: :+ c.433c>t p.r145c loss of splicing in vivo (0.4 fold) and in vitro (0.6 fold) but neither were significant CM TAZ chrx: :+ c.497t>a p.l166* loss of splicing in vivo (0.3 fold) and in vitro (0.5 fold) TN exon 7 skipping whole blood FN exon 6 and 7 skipping TP Coriell Alison Goate lymphoblastoid cell line TP Coriell CM TAZ chrx: :+ c.367c>t p.r123* loss of splicing in vivo (0.6 fold) and in vitro (0.4 fold) intron retention lymphoblastoid cell line TP Coriell 57 58
14 Supplementary Table 4. ENCODE Accession numbers used for the purpose of validating MaPSy results ENCODE Accession Number SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR SRR545700
15 References 46. Teraoka, S.N. et al. Splicing defects in the ataxia- telangiectasia gene, ATM: underlying mutations and consequences. American Journal of Human Genetics 64, (1999). 47. Sanz, D.J. et al. A high proportion of DNA variants of BRCA1 and BRCA2 is associated with aberrant splicing in breast/ovarian cancer patients. Clin Cancer Res 16, (2010). 48. Mazoyer, S. et al. A BRCA1 nonsense mutation causes exon skipping. Am J Hum Genet 62, (1998). 49. Sharp, A., Pichert, G., Lucassen, A. & Eccles, D. RNA analysis reveals splicing mutations and loss of expression defects in MLH1 and BRCA1. Hum Mutat 24, 272 (2004). 50. Dietz, H.C. et al. The skipping of constitutive exons in vivo induced by nonsense mutations. Science 259, (1993). 51. Tu, M., Tong, W., Perkins, R. & Valentine, C.R. Predicted changes in pre- mrna secondary structure vary in their association with exon skipping for mutations in exons 2, 4, and 8 of the Hprt gene and exon 51 of the fibrillin gene. Mutat Res 432, (2000). 52. Arenas, M., Duley, J., Sumi, S., Sanderson, J. & Marinaki, A. The ITPA c.94c>a and g.ivs2+21a>c sequence variants contribute to missplicing of the ITPA gene. Biochim Biophys Acta 1772, (2007). 53. Pagenstecher, C. et al. Aberrant splicing in MLH1 and MSH2 due to exonic and intronic variants. Human Genetics 119, 9-22 (2006). 54. Auclair, J. et al. Systematic mrna analysis for the effect of MLH1 and MSH2 missense and silent mutations on aberrant splicing. Hum Mutat 27, (2006). 55. Dermaut, B. et al. A novel presenilin 1 mutation associated with Pick's disease but not beta- amyloid plaques. Ann Neurol 55, (2004). 56. Dumanchin, C. et al. Biological effects of four PSEN1 gene mutations causing Alzheimer disease with spastic paraparesis and cotton wool plaques. Hum Mutat 27, 1063 (2006). 57. Hull, J. et al. Identification of common genetic variation that modulates alternative splicing. PLoS Genet 3, e99 (2007). 58. Riveira- Munoz, E. et al. Transcriptional and functional analyses of SLC12A3 mutations: new clues for the pathogenesis of Gitelman syndrome. J Am Soc Nephrol 18, (2007).
Supplementary Table 2. Identified causative mutations and/or mutation candidates.
 Supplementary Table 2. Identified causative mutations and/or mutation candidates. Nonsense mutations base change aa change Average depth Result of next generation in 432 patient Hereditary form of the
Supplementary Table 2. Identified causative mutations and/or mutation candidates. Nonsense mutations base change aa change Average depth Result of next generation in 432 patient Hereditary form of the
Whole Exome Sequenced Characteristics
 Supplementary Tables Supplementary Table 1: Patient characteristics of 45 whole exome sequenced HNSCC tumors Whole Exome Sequenced Characteristics Tumors (n=45) Age, years Median (range) 61.0 (19-90) Sex,
Supplementary Tables Supplementary Table 1: Patient characteristics of 45 whole exome sequenced HNSCC tumors Whole Exome Sequenced Characteristics Tumors (n=45) Age, years Median (range) 61.0 (19-90) Sex,
SUPPLEMENTARY INFORMATION. Intron retention is a widespread mechanism of tumor suppressor inactivation.
 SUPPLEMENTARY INFORMATION Intron retention is a widespread mechanism of tumor suppressor inactivation. Hyunchul Jung 1,2,3, Donghoon Lee 1,4, Jongkeun Lee 1,5, Donghyun Park 2,6, Yeon Jeong Kim 2,6, Woong-Yang
SUPPLEMENTARY INFORMATION Intron retention is a widespread mechanism of tumor suppressor inactivation. Hyunchul Jung 1,2,3, Donghoon Lee 1,4, Jongkeun Lee 1,5, Donghyun Park 2,6, Yeon Jeong Kim 2,6, Woong-Yang
Supplementary Table 1. PIK3CA mutation in colorectal cancer
 Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Beta Thalassemia Sami Khuri Department of Computer Science San José State University Spring 2015
 Bioinformatics in Medical Product Development SMPD 287 Three Beta Thalassemia Sami Khuri Department of Computer Science San José State University Hemoglobin Outline Anatomy of a gene Hemoglobinopathies
Bioinformatics in Medical Product Development SMPD 287 Three Beta Thalassemia Sami Khuri Department of Computer Science San José State University Hemoglobin Outline Anatomy of a gene Hemoglobinopathies
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Supplementary Figure 1
 Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first
 Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Beta Thalassemia Case Study Introduction to Bioinformatics
 Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Supplemental Information For: The genetics of splicing in neuroblastoma
 Supplemental Information For: The genetics of splicing in neuroblastoma Justin Chen, Christopher S. Hackett, Shile Zhang, Young K. Song, Robert J.A. Bell, Annette M. Molinaro, David A. Quigley, Allan Balmain,
Supplemental Information For: The genetics of splicing in neuroblastoma Justin Chen, Christopher S. Hackett, Shile Zhang, Young K. Song, Robert J.A. Bell, Annette M. Molinaro, David A. Quigley, Allan Balmain,
Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.
 Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.32 PCOS locus after conditioning for the lead SNP rs10993397;
Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.32 PCOS locus after conditioning for the lead SNP rs10993397;
Introduction. Introduction
 Introduction We are leveraging genome sequencing data from The Cancer Genome Atlas (TCGA) to more accurately define mutated and stable genes and dysregulated metabolic pathways in solid tumors. These efforts
Introduction We are leveraging genome sequencing data from The Cancer Genome Atlas (TCGA) to more accurately define mutated and stable genes and dysregulated metabolic pathways in solid tumors. These efforts
Nature Genetics: doi: /ng Supplementary Figure 1. TCGA data set on HNSCCs reanalyzed in this study.
 Supplementary Figure 1 TCGA data set on HNSCCs reanalyzed in this study. Summary of the TCGA dataset on HNSCCs re-analyzed in this study and the respective numbers of samples available within each. Supplementary
Supplementary Figure 1 TCGA data set on HNSCCs reanalyzed in this study. Summary of the TCGA dataset on HNSCCs re-analyzed in this study and the respective numbers of samples available within each. Supplementary
REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER
 REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER RNA Splicing Lecture 3, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice
REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER RNA Splicing Lecture 3, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
An Increased Specificity Score Matrix for the Prediction of. SF2/ASF-Specific Exonic Splicing Enhancers
 HMG Advance Access published July 6, 2006 1 An Increased Specificity Score Matrix for the Prediction of SF2/ASF-Specific Exonic Splicing Enhancers Philip J. Smith 1, Chaolin Zhang 1, Jinhua Wang 2, Shern
HMG Advance Access published July 6, 2006 1 An Increased Specificity Score Matrix for the Prediction of SF2/ASF-Specific Exonic Splicing Enhancers Philip J. Smith 1, Chaolin Zhang 1, Jinhua Wang 2, Shern
July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label)
 July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label) p AHLJ090 AKT1_33765 AKT1 33765 p.e17k c.49g>a C T 2 AHBKFRM BRAF_473
July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label) p AHLJ090 AKT1_33765 AKT1 33765 p.e17k c.49g>a C T 2 AHBKFRM BRAF_473
Analysis of individual differences in radiosensitivity using genome editing
 3 rd International Symposium on the System of Radiological Protection ICRP2015 Analysis of individual differences in radiosensitivity using genome editing Shinya Matsuura Department of Genetics and Cell
3 rd International Symposium on the System of Radiological Protection ICRP2015 Analysis of individual differences in radiosensitivity using genome editing Shinya Matsuura Department of Genetics and Cell
Supplementary Figure 1. Quantile-quantile (Q-Q) plot of the log 10 p-value association results from logistic regression models for prostate cancer
 Supplementary Figure 1. Quantile-quantile (Q-Q) plot of the log 10 p-value association results from logistic regression models for prostate cancer risk in stage 1 (red) and after removing any SNPs within
Supplementary Figure 1. Quantile-quantile (Q-Q) plot of the log 10 p-value association results from logistic regression models for prostate cancer risk in stage 1 (red) and after removing any SNPs within
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1
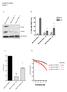 Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR 30 20 10 * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0
Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR 30 20 10 * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0
ESEfinder: a Web resource to identify exonic splicing enhancers
 ESEfinder: a Web resource to identify exonic splicing enhancers Luca Cartegni, Jinhua Wang, Zhengwei Zhu, Michael Q. Zhang, Adrian R. Krainer Cold Spring Harbor Laboratory, Cold Spring Harbor, New York,
ESEfinder: a Web resource to identify exonic splicing enhancers Luca Cartegni, Jinhua Wang, Zhengwei Zhu, Michael Q. Zhang, Adrian R. Krainer Cold Spring Harbor Laboratory, Cold Spring Harbor, New York,
Chapter II. Functional selection of intronic splicing elements provides insight into their regulatory mechanism
 23 Chapter II. Functional selection of intronic splicing elements provides insight into their regulatory mechanism Abstract Despite the critical role of alternative splicing in generating proteomic diversity
23 Chapter II. Functional selection of intronic splicing elements provides insight into their regulatory mechanism Abstract Despite the critical role of alternative splicing in generating proteomic diversity
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
Medium-Chain Acyl-CoA Dehydrogenase (MCAD) splicing mutations identified in newborns with an abnormal MS/MS profile
 EURASNET Workshop on RNA splicing and genetic diagnosis, London, UK Medium-Chain Acyl-CoA Dehydrogenase (MCAD) splicing mutations identified in newborns with an abnormal MS/MS profile Brage Storstein Andresen
EURASNET Workshop on RNA splicing and genetic diagnosis, London, UK Medium-Chain Acyl-CoA Dehydrogenase (MCAD) splicing mutations identified in newborns with an abnormal MS/MS profile Brage Storstein Andresen
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Nature Genetics: doi: /ng Supplementary Figure 1. TNFAIP3-associated haplotypes in family 1.
 Supplementary Figure 1 TNFAIP3-associated haplotypes in family 1. The p.leu227* mutation (shown as a star) arose de novo in the first affected member of the family (P1). Red haplotypes carry the TNFAIP3
Supplementary Figure 1 TNFAIP3-associated haplotypes in family 1. The p.leu227* mutation (shown as a star) arose de novo in the first affected member of the family (P1). Red haplotypes carry the TNFAIP3
Reporting TP53 gene analysis results in CLL
 Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Introduction to Cancer Biology
 Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep
 Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep Tom Walsh, PhD Division of Medical Genetics University of Washington Next generation sequencing Sanger sequencing gold
Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep Tom Walsh, PhD Division of Medical Genetics University of Washington Next generation sequencing Sanger sequencing gold
New Insights on Genetic Aspects of Thoracic Aortic Disease
 New Insights on Genetic Aspects of Thoracic Aortic Disease John A. Elefteriades, MD William W.L. Glenn Professor of Surgery Director, Aortic Institute at Yale-New Haven Yale University School of Medicine
New Insights on Genetic Aspects of Thoracic Aortic Disease John A. Elefteriades, MD William W.L. Glenn Professor of Surgery Director, Aortic Institute at Yale-New Haven Yale University School of Medicine
Nature Getetics: doi: /ng.3471
 Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
MSI positive MSI negative
 Pritchard et al. 2014 Supplementary Figure 1 MSI positive MSI negative Hypermutated Median: 673 Average: 659.2 Non-Hypermutated Median: 37.5 Average: 43.6 Supplementary Figure 1: Somatic Mutation Burden
Pritchard et al. 2014 Supplementary Figure 1 MSI positive MSI negative Hypermutated Median: 673 Average: 659.2 Non-Hypermutated Median: 37.5 Average: 43.6 Supplementary Figure 1: Somatic Mutation Burden
Clinical utility of the functional mrna evaluation of rare genetic variants in diagnostic practice
 Clinical utility of the functional mrna evaluation of rare genetic variants in diagnostic practice Celia Duff-Farrier: STP MSc project 2017 Funding provided by the Showering Fund mrna functional evaluation
Clinical utility of the functional mrna evaluation of rare genetic variants in diagnostic practice Celia Duff-Farrier: STP MSc project 2017 Funding provided by the Showering Fund mrna functional evaluation
Supplementary Table 3. 3 UTR primer sequences. Primer sequences used to amplify and clone the 3 UTR of each indicated gene are listed.
 Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) cancer DNA
 www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) DNA Supplementary Materials
www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) DNA Supplementary Materials
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project
 Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Genetic Testing and Analysis. (858) MRN: Specimen: Saliva Received: 07/26/2016 GENETIC ANALYSIS REPORT
 GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes.
 Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Using large-scale human genetic variation to inform variant prioritization in neuropsychiatric disorders
 Using large-scale human genetic variation to inform variant prioritization in neuropsychiatric disorders Kaitlin E. Samocha Hurles lab, Wellcome Trust Sanger Institute ACGS Summer Scientific Meeting 27
Using large-scale human genetic variation to inform variant prioritization in neuropsychiatric disorders Kaitlin E. Samocha Hurles lab, Wellcome Trust Sanger Institute ACGS Summer Scientific Meeting 27
Supplementary information to:
 Supplementary information to: Digital Sorting of Pure Cell Populations Enables Unambiguous Genetic Analysis of Heterogeneous Formalin-Fixed Paraffin Embedded Tumors by Next Generation Sequencing Authors
Supplementary information to: Digital Sorting of Pure Cell Populations Enables Unambiguous Genetic Analysis of Heterogeneous Formalin-Fixed Paraffin Embedded Tumors by Next Generation Sequencing Authors
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Supplementary Figure 1
 Supplementary Figure 1 5 microns C7 B6 unclassified H19 C7 signal H19 guide signal H19 B6 signal C7 SNP spots H19 RNA spots B6 SNP spots colocalization H19 RNA classification Supplementary Figure 1. Allele-specific
Supplementary Figure 1 5 microns C7 B6 unclassified H19 C7 signal H19 guide signal H19 B6 signal C7 SNP spots H19 RNA spots B6 SNP spots colocalization H19 RNA classification Supplementary Figure 1. Allele-specific
Nature Structural & Molecular Biology: doi: /nsmb.2419
 Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Supplemental Figure 1. Genes showing ectopic H3K9 dimethylation in this study are DNA hypermethylated in Lister et al. study.
 mc mc mc mc SUP mc mc Supplemental Figure. Genes showing ectopic HK9 dimethylation in this study are DNA hypermethylated in Lister et al. study. Representative views of genes that gain HK9m marks in their
mc mc mc mc SUP mc mc Supplemental Figure. Genes showing ectopic HK9 dimethylation in this study are DNA hypermethylated in Lister et al. study. Representative views of genes that gain HK9m marks in their
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Analysis of hair bundle morphology in Ush1c c.216g>a mice at P18 by SEM.
 Supplementary Figure 1 Analysis of hair bundle morphology in Ush1c c.216g>a mice at P18 by SEM. (a-c) Heterozygous c.216ga mice displayed normal hair bundle morphology at P18. (d-i) Disorganized hair bundles
Supplementary Figure 1 Analysis of hair bundle morphology in Ush1c c.216g>a mice at P18 by SEM. (a-c) Heterozygous c.216ga mice displayed normal hair bundle morphology at P18. (d-i) Disorganized hair bundles
Concurrent Practical Session ACMG Classification
 Variant Effect Prediction Training Course 6-8 November 2017 Prague, Czech Republic Concurrent Practical Session ACMG Classification Andreas Laner / Anna Benet-Pagès 1 Content 1. Background... 3 2. Aim
Variant Effect Prediction Training Course 6-8 November 2017 Prague, Czech Republic Concurrent Practical Session ACMG Classification Andreas Laner / Anna Benet-Pagès 1 Content 1. Background... 3 2. Aim
Genome 371, Autumn 2018 Quiz Section 9: Genetics of Cancer Worksheet
 Genome 371, Autumn 2018 Quiz Section 9: Genetics of Cancer Worksheet All cancer is due to genetic mutations. However, in cancer that clusters in families (familial cancer) at least one of these mutations
Genome 371, Autumn 2018 Quiz Section 9: Genetics of Cancer Worksheet All cancer is due to genetic mutations. However, in cancer that clusters in families (familial cancer) at least one of these mutations
indicated in shaded lowercase letters (hg19, Chr2: 217,955, ,957,266).
 Legend for Supplementary Figures Figure S1: Sequence of 2q35 encnv. The DNA sequence of the 1,375bp 2q35 encnv is indicated in shaded lowercase letters (hg19, Chr2: 217,955,892-217,957,266). Figure S2:
Legend for Supplementary Figures Figure S1: Sequence of 2q35 encnv. The DNA sequence of the 1,375bp 2q35 encnv is indicated in shaded lowercase letters (hg19, Chr2: 217,955,892-217,957,266). Figure S2:
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
Supplementary Online Content
 Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Finding subtle mutations with the Shannon human mrna splicing pipeline
 Finding subtle mutations with the Shannon human mrna splicing pipeline Presentation at the CLC bio Medical Genomics Workshop American Society of Human Genetics Annual Meeting November 9, 2012 Peter K Rogan
Finding subtle mutations with the Shannon human mrna splicing pipeline Presentation at the CLC bio Medical Genomics Workshop American Society of Human Genetics Annual Meeting November 9, 2012 Peter K Rogan
NGS for Cancer Predisposition
 NGS for Cancer Predisposition Colin Pritchard MD, PhD University of Washington Dept. of Lab Medicine AMP Companion Society Meeting USCAP Boston March 22, 2015 Disclosures I am an employee of the University
NGS for Cancer Predisposition Colin Pritchard MD, PhD University of Washington Dept. of Lab Medicine AMP Companion Society Meeting USCAP Boston March 22, 2015 Disclosures I am an employee of the University
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1. Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Nature Immunology: doi: /ni.
 Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
SUPPLEMENTARY FIGURES: Supplementary Figure 1
 SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
T he availability of fast sequencing protocols is
 737 REVIEW Splicing in action: assessing disease causing sequence changes D Baralle, M Baralle... Variations in new splicing regulatory elements are difficult to identify exclusively by sequence inspection
737 REVIEW Splicing in action: assessing disease causing sequence changes D Baralle, M Baralle... Variations in new splicing regulatory elements are difficult to identify exclusively by sequence inspection
fl/+ KRas;Atg5 fl/+ KRas;Atg5 fl/fl KRas;Atg5 fl/fl KRas;Atg5 Supplementary Figure 1. Gene set enrichment analyses. (a) (b)
 KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
Functional insights from genetic channelopathies Stephanie Schorge
 Functional Insights From Genetic Channelopathies Dr. 1 Royal Society University Research Fellow Department of Clinical and Experimental Epilepsy Aims of channelopathies lecture Describe channelopathies
Functional Insights From Genetic Channelopathies Dr. 1 Royal Society University Research Fellow Department of Clinical and Experimental Epilepsy Aims of channelopathies lecture Describe channelopathies
variant led to a premature stop codon p.k316* which resulted in nonsense-mediated mrna decay. Although the exact function of the C19L1 is still
 157 Neurological disorders primarily affect and impair the functioning of the brain and/or neurological system. Structural, electrical or metabolic abnormalities in the brain or neurological system can
157 Neurological disorders primarily affect and impair the functioning of the brain and/or neurological system. Structural, electrical or metabolic abnormalities in the brain or neurological system can
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
JULY 21, Genetics 101: SCN1A. Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology
 JULY 21, 2018 Genetics 101: SCN1A Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology Disclosures: I have no financial interests or relationships to disclose. Objectives 1. Review genetic
JULY 21, 2018 Genetics 101: SCN1A Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology Disclosures: I have no financial interests or relationships to disclose. Objectives 1. Review genetic
Mutational and phenotypical spectrum of phenylalanine hydroxylase deficiency in Denmark
 Clin Genet 2016: 90: 247 251 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12692 Mutational and
Clin Genet 2016: 90: 247 251 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12692 Mutational and
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins.
 Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
Investigating rare diseases with Agilent NGS solutions
 Investigating rare diseases with Agilent NGS solutions Chitra Kotwaliwale, Ph.D. 1 Rare diseases affect 350 million people worldwide 7,000 rare diseases 80% are genetic 60 million affected in the US, Europe
Investigating rare diseases with Agilent NGS solutions Chitra Kotwaliwale, Ph.D. 1 Rare diseases affect 350 million people worldwide 7,000 rare diseases 80% are genetic 60 million affected in the US, Europe
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Names for ABO (ISBT 001) Blood Group Alleles
 Names for ABO (ISBT 001) blood group alleles v1.1 1102 Names for ABO (ISBT 001) Blood Group Alleles General description: The ABO system was discovered as in 1900 and is considered the first and clinically
Names for ABO (ISBT 001) blood group alleles v1.1 1102 Names for ABO (ISBT 001) Blood Group Alleles General description: The ABO system was discovered as in 1900 and is considered the first and clinically
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research
 Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
RECAP (1)! In eukaryotes, large primary transcripts are processed to smaller, mature mrnas.! What was first evidence for this precursorproduct
 RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
CANCER GENETICS PROVIDER SURVEY
 Dear Participant, Previously you agreed to participate in an evaluation of an education program we developed for primary care providers on the topic of cancer genetics. This is an IRB-approved, CDCfunded
Dear Participant, Previously you agreed to participate in an evaluation of an education program we developed for primary care providers on the topic of cancer genetics. This is an IRB-approved, CDCfunded
REGULATED AND NONCANONICAL SPLICING
 REGULATED AND NONCANONICAL SPLICING RNA Processing Lecture 3, Biological Regulatory Mechanisms, Hiten Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice site consensus sequences do have
REGULATED AND NONCANONICAL SPLICING RNA Processing Lecture 3, Biological Regulatory Mechanisms, Hiten Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice site consensus sequences do have
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Whole exome sequencing as a first line test: Is there even a role for metabolic biochemists in the future?
 Whole exome sequencing as a first line test: Is there even a role for metabolic biochemists in the future? Tony Marinakii Purine Research Laboratory Biochemical Sciences Whole exome sequencing No doubt
Whole exome sequencing as a first line test: Is there even a role for metabolic biochemists in the future? Tony Marinakii Purine Research Laboratory Biochemical Sciences Whole exome sequencing No doubt
Belgian study into von Willebrand Disease (B-Will): First results
 Belgian study into von Willebrand Disease (B-Will): First results Inge Vangenechten VWD Research Unit, Antwerp University Hospital Supported by CSL Behring Chair in von Willebrand Disease, Antwerp University
Belgian study into von Willebrand Disease (B-Will): First results Inge Vangenechten VWD Research Unit, Antwerp University Hospital Supported by CSL Behring Chair in von Willebrand Disease, Antwerp University
Identification and characterization of multiple splice variants of Cdc2-like kinase 4 (Clk4)
 Identification and characterization of multiple splice variants of Cdc2-like kinase 4 (Clk4) Vahagn Stepanyan Department of Biological Sciences, Fordham University Abstract: Alternative splicing is an
Identification and characterization of multiple splice variants of Cdc2-like kinase 4 (Clk4) Vahagn Stepanyan Department of Biological Sciences, Fordham University Abstract: Alternative splicing is an
Supplementary Methods
 Supplementary Methods Short Read Preprocessing Reads are preprocessed differently according to how they will be used: detection of the variant in the tumor, discovery of an artifact in the normal or for
Supplementary Methods Short Read Preprocessing Reads are preprocessed differently according to how they will be used: detection of the variant in the tumor, discovery of an artifact in the normal or for
p.r623c p.p976l p.d2847fs p.t2671 p.d2847fs p.r2922w p.r2370h p.c1201y p.a868v p.s952* RING_C BP PHD Cbp HAT_KAT11
 ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
Asingle inherited mutant gene may be enough to
 396 Cancer Inheritance STEVEN A. FRANK Asingle inherited mutant gene may be enough to cause a very high cancer risk. Single-mutation cases have provided much insight into the genetic basis of carcinogenesis,
396 Cancer Inheritance STEVEN A. FRANK Asingle inherited mutant gene may be enough to cause a very high cancer risk. Single-mutation cases have provided much insight into the genetic basis of carcinogenesis,
Supplementary Tables. Supplementary Figures
 Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Advances in genetic diagnosis of neurological disorders
 Acta Neurol Scand 2014: 129 (Suppl. 198): 20 25 DOI: 10.1111/ane.12232 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd ACTA NEUROLOGICA SCANDINAVICA Review Article Advances in genetic diagnosis
Acta Neurol Scand 2014: 129 (Suppl. 198): 20 25 DOI: 10.1111/ane.12232 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd ACTA NEUROLOGICA SCANDINAVICA Review Article Advances in genetic diagnosis
Molecular Pathology of Ovarian Carcinoma with Morphological Correlation
 Molecular athology of Ovarian Carcinoma with Morphological Correlation Kathleen R. Cho, M.D. Comprehensive Cancer Center and Departments of athology and Internal Medicine University of Michigan Medical
Molecular athology of Ovarian Carcinoma with Morphological Correlation Kathleen R. Cho, M.D. Comprehensive Cancer Center and Departments of athology and Internal Medicine University of Michigan Medical
Nature Genetics: doi: /ng Supplementary Figure 1. PCA for ancestry in SNV data.
 Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Table S2. Baseline characteristics of the Rotterdam Study for interaction analysis
 Supplementary data Table S1. List of the primers for the cloning of precursor mir-4707 Table S2. Baseline characteristics of the Rotterdam Study for interaction analysis Table S3. List of the primers for
Supplementary data Table S1. List of the primers for the cloning of precursor mir-4707 Table S2. Baseline characteristics of the Rotterdam Study for interaction analysis Table S3. List of the primers for
Where Splicing Joins Chromatin And Transcription. 9/11/2012 Dario Balestra
 Where Splicing Joins Chromatin And Transcription 9/11/2012 Dario Balestra Splicing process overview Splicing process overview Sequence context RNA secondary structure Tissue-specific Proteins Development
Where Splicing Joins Chromatin And Transcription 9/11/2012 Dario Balestra Splicing process overview Splicing process overview Sequence context RNA secondary structure Tissue-specific Proteins Development
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Spectrum and Classification of ATP7B Variants in a Large Cohort of Chinese Patients with Wilson s Disease Guides Genetic Diagnosis
 Ivyspring International Publisher 638 Theranostics Research Paper 2016; 6(5): 638-649. doi: 10.7150/thno.14596 Spectrum and Classification of ATP7B Variants in a Large Cohort of Chinese Patients with Wilson
Ivyspring International Publisher 638 Theranostics Research Paper 2016; 6(5): 638-649. doi: 10.7150/thno.14596 Spectrum and Classification of ATP7B Variants in a Large Cohort of Chinese Patients with Wilson
Familial exudative vitreoretinopathy (FEVR) is an inheritable
 Genetics Spectrum of Variants in 389 Chinese Probands With Familial Exudative Vitreoretinopathy Jia-Kai Li, 1 Yian Li, 1 Xiang Zhang, 1 Chun-Li Chen, 2 Yu-Qing Rao, 1 Ping Fei, 1 Qi Zhang, 1 Peiquan Zhao,
Genetics Spectrum of Variants in 389 Chinese Probands With Familial Exudative Vitreoretinopathy Jia-Kai Li, 1 Yian Li, 1 Xiang Zhang, 1 Chun-Li Chen, 2 Yu-Qing Rao, 1 Ping Fei, 1 Qi Zhang, 1 Peiquan Zhao,
Clinical Genetics. Functional validation in a diagnostic context. Robert Hofstra. Leading the way in genetic issues
 Clinical Genetics Leading the way in genetic issues Functional validation in a diagnostic context Robert Hofstra Clinical Genetics Leading the way in genetic issues Future application of exome sequencing
Clinical Genetics Leading the way in genetic issues Functional validation in a diagnostic context Robert Hofstra Clinical Genetics Leading the way in genetic issues Future application of exome sequencing
SUPPLEMENTARY INFORMATION. Rare independent mutations in renal salt handling genes contribute to blood pressure variation
 SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
RNA splicing meets genetic testing: detection and interpretation of splicing defects in
 1 MEETING REPORT RNA splicing meets genetic testing: detection and interpretation of splicing defects in genetic diseases Mario Tosi 1, Stefan Stamm 2, Diana Baralle 3 1 Inserm U614, Faculty of Medicine,
1 MEETING REPORT RNA splicing meets genetic testing: detection and interpretation of splicing defects in genetic diseases Mario Tosi 1, Stefan Stamm 2, Diana Baralle 3 1 Inserm U614, Faculty of Medicine,
Potential Consequences on the RNA Level and using prediction tools. Andreas Laner MGZ München
 Potential Consequences on the RNA Level and using prediction tools Andreas Laner MGZ München laner@mgz-muenchen.de Potential Consequences on the RNA Level and using prediction tools A. Variants altering
Potential Consequences on the RNA Level and using prediction tools Andreas Laner MGZ München laner@mgz-muenchen.de Potential Consequences on the RNA Level and using prediction tools A. Variants altering
1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics
 1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics to gain mechanistic insight! 4. Return to step 2, as
1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics to gain mechanistic insight! 4. Return to step 2, as
MODULE 4: SPLICING. Removal of introns from messenger RNA by splicing
 Last update: 05/10/2017 MODULE 4: SPLICING Lesson Plan: Title MEG LAAKSO Removal of introns from messenger RNA by splicing Objectives Identify splice donor and acceptor sites that are best supported by
Last update: 05/10/2017 MODULE 4: SPLICING Lesson Plan: Title MEG LAAKSO Removal of introns from messenger RNA by splicing Objectives Identify splice donor and acceptor sites that are best supported by
Identifying Mutations Responsible for Rare Disorders Using New Technologies
 Identifying Mutations Responsible for Rare Disorders Using New Technologies Jacek Majewski, Department of Human Genetics, McGill University, Montreal, QC Canada Mendelian Diseases Clear mode of inheritance
Identifying Mutations Responsible for Rare Disorders Using New Technologies Jacek Majewski, Department of Human Genetics, McGill University, Montreal, QC Canada Mendelian Diseases Clear mode of inheritance
Mapping by recurrence and modelling the mutation rate
 Mapping by recurrence and modelling the mutation rate Shamil Sunyaev Broad Institute of M.I.T. and Harvard Current knowledge is from Comparative genomics Experimental systems: yeast reporter assays Potential
Mapping by recurrence and modelling the mutation rate Shamil Sunyaev Broad Institute of M.I.T. and Harvard Current knowledge is from Comparative genomics Experimental systems: yeast reporter assays Potential
Supplementary Document
 Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
