Supplemental Figure legends
|
|
|
- Samson Higgins
- 5 years ago
- Views:
Transcription
1 Supplemental Figure legends Supplemental Figure S1 Frequently mutated genes. Frequently mutated genes (mutated in at least four patients) with information about mutation frequency, RNA-expression and copy-number. Tumors from NPD patients are shown to the left and tumors from PD patients to the right. Supplemental Figure S2 Analysis of differences and changes in mutations types. Only patients having paired tumors were included in the analysis (14 NPD and 15 PD patients). (A-D), Diagrams showing the proportions of the six mutation types in the first and last tumors for the NPD patients (A), the PD patients (B), the smokers only (C) and the combined former and non-smokers (D). In cases where more than two tumors were available from one patient the first resected and last resected tumors were used. The bars represent lowest, highest and median proportions across patients. (E) Differences in intra patient mutation type proportions. Proportions are calculated as the mean of absolute change across NPD and PD patients. (F) Differences in intra patient mutation type proportions. Change is calculated as the mean of Tumor2-Tumor1 across NPD and PD patients. In cases where more than two tumors were available from one patient, the change in proportion was calculated between tumors having highest difference in stage. Supplemental Figure S3 Mutational signatures based on trinucleotide contexts The mutational context is represented for each sample and each patient. Each bar represents the relative ratio of a given mutation in a given trinucleotide context (dark color when including only cat 1 SNVs and light colors when including all SNVs called). Supplemental Figure S4 Contributions and comparison of most prevalent mutational signatures across samples Contributions of signatures 1A, 1B, 2, 6, 12, 13 and 2+13 across all samples. Samples from NPD patients are shown in green and samples of the PD patients are shown in blue. Signatures 2 and 13 are a hallmark of APOBEC enzyme activity and a plot of signature 2+13 is included to represent the combined APOBEC effect. 13
2 Supplemental Figure S5 Comparisons of contributions of most prevalent mutational signatures from initial tumor to most recent tumor in NPD and PD patients Signatures 1A, 1B, 2, 6, 12, 13, 2+13 plotted against Tumor 1(initial tumor) and Tumor 2 (most recent tumor) are shown as bar plots. For patients having more than two tumor samples, the tumor with the highest stage is selected as tumor 2. Samples from NPD patients are shown in light green (tumor 1) and dark green (tumor 2). Samples from PD patients are shown in light red (tumor 1) and dark red (tumor 2). Supplemental Figure S6 Comparison of mutation rate and APOBEC score. (A) Correlation plot of APOBEC score (Cat 1-2) versus the mutation rate (cat1+2 SNV/Mb) for all samples. (B) Relationship between APOBEC score (Bars, primary Y-axis; Cat 1-2) and the mutation rate (Black lines, secondary y-axis) for tumors from NPD patients (green) and PD patients (red). Supplemental Figure S7 APOBEC enzymes expression Bar plots displaying gene expression of Apobec1, Apobec2, Apobec3a, Apobec3b, Apobec3b-as1, Apobec3c, Apobec3d, Apobec3f, Apobec3g, Apobec3h and Apobec4 for the initial Ta tumors from NPD patients (green bars) and PD patients (blue bars). Supplemental Figure S8 Comparison of deleterious mutations in different sub-groups. (A) Comparison of the initial Ta tumors from each group (omitting patients with primary invasive tumors) identified ten early drivers of progression (LRP1B, RYR3, PCDHGA12, TLN1, RP1L1, BBX, ZNF717, ADAMTS18, TENM1 and SSPO) only mutated in Ta tumors from the PD group. Of these LRP1B and RYR3, both related to cancer cell migration, were mutated in 26% (4/15) of the PD patients (P<0.05). The LRP1B gene is often deleted in bladder cancer and is thought to be a tumor suppressor gene (13). Recently, LRP1B was found to be a possible cancer driver gene in clear cell kidney carcinoma, present as a subclonal mutation (9). (B) Comparison of the initial Ta tumors from NPD patients versus the T1 or T2 tumors from PD patients (most recent tumor) identified 14 genes that were only mutated in the PD group. The most frequently mutated gene, TRRAP, was mutated in four (24%) of the PD samples and is involved in MYC signaling (14). TRRAP encodes a kinase that is part of several histone acetyltransferase (HAT) complexes e.g. the TIP60 complex (14), which is involved in bladder cancer (15). Loss of TRRAP activates the WNT pathway and is frequently seen in melanomas (15). (C) A comparison of paired Ta tumors from NPD patients revealed frequent mutations in PIK3CA, RBM10, KDM6A, RANBP2 and FGFR3 were early events present in the initial Ta, whereas mutations in MKI67, CCDC168, MIA3 and OPN1LW primarily occurred late in tumorigenesis of recurrent Ta tumors. (D) Finally, we compared metachronous tumors from PD patients and we found that genes like RYR3, RP1L1, KDM6A, FGFR3, PIK3CA and ZNF717 were frequently mutated in the initial tumor. Interestingly only 35% (7/20) of the mutations in these genes seemed to be retained in the progressed samples. However, visual inspection (SNVs only) resulted in detection of additional recurrent low frequency SNVs (marked by gray) in early (n=7) and late PD samples (n=8), increasing the retained mutations to 50% (10/20) in 14
3 these genes. In contrast, mutations in ADRA2B and TRRAP are predominantly present in the progressed samples. However, after visual inspection, 100% (3/3) of ADRA2B and 33% (1/3) of TRRAP mutations were observed in the initial tumor, at very low frequency (marked by an orange box). Supplemental Figure S9 Disease evolution inferred by phylogenetic tree analysis for patients with two tumors analyzed. The total number of mutations and the percentages of SNVs belonging to each of the branches are shown. Tumor suppressor genes, oncogenes and bladder intogen genes with Tier 0-1 SNVs are highlighted on the right side (ancestral: black font; sample-specific: green font for the first tumor and blue font for the second) together with their clonal status as defined in (9) (red: clonal, blue: subclonal and grey: not defined). Genes potentially actionable using FDA approved drugs are marked with an asterisk. The time-scale indicates the time between the tumor removals. Supplemental Figure S10 Mutational heat maps for patients with three or more tumor samples analyzed For each patient with three or more tumor sample analyzed, a heat map of presence (green) or absence (white) of mutations (left) and a heat map of allele frequencies (right) are shown. Supplemental Figure S11 Clonality status of all Cat 1 Tier 0-1 mutations found in NPD patients. For each sample, the cancer cell fraction (CCF) was estimated using information about the number of reads for the reference and alternate alleles, an estimation of the cellularity and the copy number status of the region harboring the mutation. The mean and confidence intervals from all mutations called in Cat1 Tier 0-1 are plotted. A mutation was called clonal and colored in red if the upper limit of the confidence interval was above If not, it was called sub-clonal and colored in blue. Supplemental Figure S12 Clonality status of all Cat 1 Tier 0-1 mutations found in PD patients. For each sample, the cancer cell fraction (CCF) was estimated using information about the number of reads for the reference and alternate alleles, an estimation of the cellularity and the copy number status of the region harboring the mutation. The mean and confidence intervals from all mutations called in Cat1 Tier 0-1 are plotted. A mutation was called clonal and colored in red if the upper limit of the confidence interval was above If not, it was called sub-clonal and colored in blue. Supplemental Figure S13 List of genes with their clonality status in all samples from NPD patients. For NPD patients with two tumor samples analyzed, we show all Cat 1 Tier 0-1 mutations (defined by the gene they are affecting) in the phylogenetic tree context (Fig. 3). The genes in the ancestral branches are defined by a black font, the genes in the first sample branch by a green font and the genes in the second branch by a blue font. The clonal status is defined in the first column for the first tumor and in the second column for the last tumor: red for clonal, blue for sub-clonal and grey for not defined. 15
4 Supplemental Figure S14 List of genes with their clonality status in all samples from PD patients. For PD patients with two tumor samples analyzed, we show all Cat 1 Tier 0-1 mutations (defined by the gene they are affecting) in the phylogenetic tree context (Fig. 3). The genes in the ancestral branches are defined by a black font, the genes in the first sample branch by a green font and the genes in the second branch by a blue font. The clonal status is defined in the first column for the first tumor and in the second column for the last tumor: red for clonal, blue for sub-clonal and grey for not defined. Supplemental Figure S15 Correlation between allele frequencies estimated from exome-seq versus ampliconseq. All SNVs used to infer the subclonal populations are plotted. Overall we observed a very good correlation (correlation = 0.91) between the frequencies estimated in the ampliconseq data (x-axis) and in the exome-seq data (y-axis). The overall density is indicated by the red shading. Supplemental Figure S16 Subclonal populations inferred by PyClone in patients with two samples analyzed. For 17 patients with two tumor samples analyzed, the cellular prevalence of a selection of SNVs was inferred by PyClone and plotted (black dots). The overall density is indicated by the red shading and the name of driver genes (oncogenes, tumor suppressor genes or intogen genes) is written next to their cluster. Supplemental Figure S17 Subclonal populations inferred by PyClone in patients with three samples analyzed. For four patients with three tumor samples analyzed, the cellular prevalence of a selection of SNVs was inferred by PyClone and plotted (black dots). The overall density is indicated by the red shading and the name of driver genes (oncogenes, tumor suppressor genes or intogen genes) is written next to their cluster. Supplemental Figure S18 Subclonal populations inferred by PyClone in the patient with five samples analyzed. For one patient with five tumor samples analyzed, the cellular prevalence of a selection of SNVs was inferred by PyClone and plotted (black dots). The overall density is indicated by the red shading and the name of driver genes (oncogenes, tumor suppressor genes or intogen genes) is written next to their cluster. Supplemental Figure S19 APOBEC enzymes expression in tumor samples from patients with three or more metachronous tumor samples. Bar plots displaying gene expression of APOBEC1, APOBEC2, APOBEC3A, APOBEC3B, APOBEC3B-AS1, APOBEC3C, APOBEC3D, APOBEC3F, APOBEC3G, APOBEC3H and APOBEC4 for all samples from the six patients with three or more tumor samples analyzed. 16
5 Supplemental Table legends Supplemental Table 1 Clinical and histopathological information together with information on exome and RNA sequencing (A) Overview of clinical data for the 75 tumor samples included in the paired-whole exome sequencing set. (B) Summary of samples and clinical data. (C) Summary statistics on Paired-WES data set. (D) Summary statistics of RNA sequencing data. Supplemental Table 2 Mutations called by MuTect The table displays all mutations called by MuTect for all the samples annotated by SnpEff and associated categories, Cat 1 and Cat 2. Supplemental Table 3 Allele specific expression (ASE) (A) Differences in ASE between cancerous and normal bladder cells as a function of gene class. (B) Significant genes with a different ASE between cancerous and normal bladder cells. (C) Differences in ASE between initial Ta tumors from NPD versus PD patients as a function of gene class. (D) Significant genes with a different ASE between initial Ta tumors from NPD versus PD patients. Supplemental Table 4 Heat maps of mutations and gene expression for patients with three or more tumor sample analyzed. For each of the six patients with more than two metachronous tumors analyzed, we show a heat map of presence (green) or absence (white), a heat map with allele frequencies information and the gene expression values for the corresponding mutations. Supplemental Table 5 Mutations included in the 1530 amplicons target panel. (A) The 1800 SNVs included in the 1530 amplicons target panel. For each sample, information about the category of the call is given if the mutation has been called in this sample. (B) Summary statistic for the 1800 SNVs included in the 1536 amplicons target panel for all analyzed patients samples. The table includes alternative and reference call for both the amplicon sequencing and exome sequencing. (C) Comparison of SNV calls from WES and deep amplicon sequencing for patient 11. The table displays heat map, allele frequency of alterations for patient 19 calculated from the WES data. The reference and alternate allele counts from the SNV positions that were included on the targeted deep sequencing panel for this patient are shown to the right. This demonstrates that we can fully trust a negative SNV in the WES data as the ultra-deep sequencing do not pick up low frequency SNV not detected in the WES data. (D) All primer/probe position and sequence for 1530 amplicons. Supplemental Table 6 Potential therapeutic targets. All altered genes that are potential targets identified by search in the Drug-Gene Interaction database (DGIdb), the Target database, and the IntOGen database are listed 17
6 for each patient. The genes altered, the observed alterations, and the samples affected are shown. Additionally, IntOGen genes, oncogenes, or tumor suppressor genes are highlighted in green, red, and blue font, respectively. All FDA approved drugs for the given target are listed (identified by the DrugBank database). 18
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Nature Getetics: doi: /ng.3471
 Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
 Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
ARTICLE RESEARCH. Macmillan Publishers Limited. All rights reserved
 Extended Data Figure 6 Annotation of drivers based on clinical characteristics and co-occurrence patterns. a, Putative drivers affecting greater than 10 patients were assessed for enrichment in IGHV mutated
Extended Data Figure 6 Annotation of drivers based on clinical characteristics and co-occurrence patterns. a, Putative drivers affecting greater than 10 patients were assessed for enrichment in IGHV mutated
Supplementary Materials for
 www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
Supplementary Tables. Supplementary Figures
 Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Research Strategy: 1. Background and Significance
 Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
Nature Genetics: doi: /ng Supplementary Figure 1. Mutational signatures in BCC compared to melanoma.
 Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Supplementary Figure 1. Estimation of tumour content
 Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/1/8/e1500296/dc1 Supplementary Materials for Transcriptional regulation of APOBEC3 antiviral immunity through the CBF- /RUNX axis This PDF file includes: Brett
advances.sciencemag.org/cgi/content/full/1/8/e1500296/dc1 Supplementary Materials for Transcriptional regulation of APOBEC3 antiviral immunity through the CBF- /RUNX axis This PDF file includes: Brett
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first
 Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs.
 Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs. (a) CNA analysis of expression microarray data obtained from 15 tumors in the SV40Tag
Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs. (a) CNA analysis of expression microarray data obtained from 15 tumors in the SV40Tag
Nature Structural & Molecular Biology: doi: /nsmb.2419
 Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
CDH1 truncating alterations were detected in all six plasmacytoid-variant bladder tumors analyzed by whole-exome sequencing.
 Supplementary Figure 1 CDH1 truncating alterations were detected in all six plasmacytoid-variant bladder tumors analyzed by whole-exome sequencing. Whole-exome sequencing of six plasmacytoid-variant bladder
Supplementary Figure 1 CDH1 truncating alterations were detected in all six plasmacytoid-variant bladder tumors analyzed by whole-exome sequencing. Whole-exome sequencing of six plasmacytoid-variant bladder
Figure S2. Distribution of acgh probes on all ten chromosomes of the RIL M0022
 96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Nature Genetics: doi: /ng Supplementary Figure 1. Clinical timeline for the discovery WES cases.
 Supplementary Figure 1 Clinical timeline for the discovery WES cases. This illustrates the timeline of the disease events during the clinical course of each patient s disease, further indicating the available
Supplementary Figure 1 Clinical timeline for the discovery WES cases. This illustrates the timeline of the disease events during the clinical course of each patient s disease, further indicating the available
Figure S4. 15 Mets Whole Exome. 5 Primary Tumors Cancer Panel and WES. Next Generation Sequencing
 Figure S4 Next Generation Sequencing 15 Mets Whole Exome 5 Primary Tumors Cancer Panel and WES Get coverage of all variant loci for all three Mets Variant Filtering Sequence Alignments Index and align
Figure S4 Next Generation Sequencing 15 Mets Whole Exome 5 Primary Tumors Cancer Panel and WES Get coverage of all variant loci for all three Mets Variant Filtering Sequence Alignments Index and align
Myeloma Genetics what do we know and where are we going?
 in partnership with Myeloma Genetics what do we know and where are we going? Dr Brian Walker Thames Valley Cancer Network 14 th September 2015 Making the discoveries that defeat cancer Myeloma Genome:
in partnership with Myeloma Genetics what do we know and where are we going? Dr Brian Walker Thames Valley Cancer Network 14 th September 2015 Making the discoveries that defeat cancer Myeloma Genome:
fraction (soluble sugars, starch and total NSC). Conditional and marginal R 2 values are given for
 1 2 Martínez-Vilalta, Sala, Asensio, Galiano, Hoch, Palacio, Piper and Lloret. Dynamics of non- structural carbohydrates in terrestrial plants: a global synthesis. Ecological Monographs. 3 4 APPENDIX S2.
1 2 Martínez-Vilalta, Sala, Asensio, Galiano, Hoch, Palacio, Piper and Lloret. Dynamics of non- structural carbohydrates in terrestrial plants: a global synthesis. Ecological Monographs. 3 4 APPENDIX S2.
Nature Medicine: doi: /nm.3967
 Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
NGS in Cancer Pathology After the Microscope: From Nucleic Acid to Interpretation
 NGS in Cancer Pathology After the Microscope: From Nucleic Acid to Interpretation Michael R. Rossi, PhD, FACMG Assistant Professor Division of Cancer Biology, Department of Radiation Oncology Department
NGS in Cancer Pathology After the Microscope: From Nucleic Acid to Interpretation Michael R. Rossi, PhD, FACMG Assistant Professor Division of Cancer Biology, Department of Radiation Oncology Department
Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes.
 Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Computer Science, Biology, and Biomedical Informatics (CoSBBI) Outline. Molecular Biology of Cancer AND. Goals/Expectations. David Boone 7/1/2015
 Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies
 OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies
 Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies Dr. Maricel G. Kann Assistant Professor Dept of Biological Sciences UMBC 2 The term protein domain
Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies Dr. Maricel G. Kann Assistant Professor Dept of Biological Sciences UMBC 2 The term protein domain
Nature Genetics: doi: /ng Supplementary Figure 1. Rates of different mutation types in CRC.
 Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
DNA-seq Bioinformatics Analysis: Copy Number Variation
 DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
Chronic HIV-1 Infection Frequently Fails to Protect against Superinfection
 Chronic HIV-1 Infection Frequently Fails to Protect against Superinfection Anne Piantadosi 1,2[, Bhavna Chohan 1,2[, Vrasha Chohan 3, R. Scott McClelland 3,4,5, Julie Overbaugh 1,2* 1 Division of Human
Chronic HIV-1 Infection Frequently Fails to Protect against Superinfection Anne Piantadosi 1,2[, Bhavna Chohan 1,2[, Vrasha Chohan 3, R. Scott McClelland 3,4,5, Julie Overbaugh 1,2* 1 Division of Human
Nature Genetics: doi: /ng Supplementary Figure 1. HOX fusions enhance self-renewal capacity.
 Supplementary Figure 1 HOX fusions enhance self-renewal capacity. Mouse bone marrow was transduced with a retrovirus carrying one of three HOX fusion genes or the empty mcherry reporter construct as described
Supplementary Figure 1 HOX fusions enhance self-renewal capacity. Mouse bone marrow was transduced with a retrovirus carrying one of three HOX fusion genes or the empty mcherry reporter construct as described
Nature Genetics: doi: /ng Supplementary Figure 1. Somatic coding mutations identified by WES/WGS for 83 ATL cases.
 Supplementary Figure 1 Somatic coding mutations identified by WES/WGS for 83 ATL cases. (a) The percentage of targeted bases covered by at least 2, 10, 20 and 30 sequencing reads (top) and average read
Supplementary Figure 1 Somatic coding mutations identified by WES/WGS for 83 ATL cases. (a) The percentage of targeted bases covered by at least 2, 10, 20 and 30 sequencing reads (top) and average read
Frequency(%) KRAS G12 KRAS G13 KRAS A146 KRAS Q61 KRAS K117N PIK3CA H1047 PIK3CA E545 PIK3CA E542K PIK3CA Q546. EGFR exon19 NFS-indel EGFR L858R
 Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Cancer Genomics. Nic Waddell. Winter School in Mathematical and Computational Biology. July th
 Cancer Genomics Nic Waddell Winter School in Mathematical and Computational Biology 6th July 2015 Time Line of Key Events in Cancer Genomics Michael R. Stratton Science 2011;331:1553-1558 The Cancer Genome
Cancer Genomics Nic Waddell Winter School in Mathematical and Computational Biology 6th July 2015 Time Line of Key Events in Cancer Genomics Michael R. Stratton Science 2011;331:1553-1558 The Cancer Genome
Plasma-Seq conducted with blood from male individuals without cancer.
 Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma.
 Supplementary Figure 1 Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma. (a) Breeding scheme. R26-LSL-SB11 homozygous mice were bred to Trp53 LSL-R270H/+
Supplementary Figure 1 Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma. (a) Breeding scheme. R26-LSL-SB11 homozygous mice were bred to Trp53 LSL-R270H/+
NGS in tissue and liquid biopsy
 NGS in tissue and liquid biopsy Ana Vivancos, PhD Referencias So, why NGS in the clinics? 2000 Sanger Sequencing (1977-) 2016 NGS (2006-) ABIPrism (Applied Biosystems) Up to 2304 per day (96 sequences
NGS in tissue and liquid biopsy Ana Vivancos, PhD Referencias So, why NGS in the clinics? 2000 Sanger Sequencing (1977-) 2016 NGS (2006-) ABIPrism (Applied Biosystems) Up to 2304 per day (96 sequences
Supplemental Information. Molecular, Pathological, Radiological, and Immune. Profiling of Non-brainstem Pediatric High-Grade
 Cancer Cell, Volume 33 Supplemental Information Molecular, Pathological, Radiological, and Immune Profiling of Non-brainstem Pediatric High-Grade Glioma from the HERBY Phase II Randomized Trial Alan Mackay,
Cancer Cell, Volume 33 Supplemental Information Molecular, Pathological, Radiological, and Immune Profiling of Non-brainstem Pediatric High-Grade Glioma from the HERBY Phase II Randomized Trial Alan Mackay,
SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer.
 Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Journal: Nature Methods
 Journal: Nature Methods Article Title: Network-based stratification of tumor mutations Corresponding Author: Trey Ideker Supplementary Item Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure
Journal: Nature Methods Article Title: Network-based stratification of tumor mutations Corresponding Author: Trey Ideker Supplementary Item Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure
CONTRACTING ORGANIZATION: The Chancellor, Masters and Scholars of the University of Cambridge, Clara East, The Old Schools, Cambridge CB2 1TN
 AWARD NUMBER: W81XWH-14-1-0110 TITLE: A Molecular Framework for Understanding DCIS PRINCIPAL INVESTIGATOR: Gregory Hannon CONTRACTING ORGANIZATION: The Chancellor, Masters and Scholars of the University
AWARD NUMBER: W81XWH-14-1-0110 TITLE: A Molecular Framework for Understanding DCIS PRINCIPAL INVESTIGATOR: Gregory Hannon CONTRACTING ORGANIZATION: The Chancellor, Masters and Scholars of the University
Nature Immunology: doi: /ni Supplementary Figure 1. RNA-Seq analysis of CD8 + TILs and N-TILs.
 Supplementary Figure 1 RNA-Seq analysis of CD8 + TILs and N-TILs. (a) Schematic representation of the tumor and cell types used for the study. HNSCC, head and neck squamous cell cancer; NSCLC, non-small
Supplementary Figure 1 RNA-Seq analysis of CD8 + TILs and N-TILs. (a) Schematic representation of the tumor and cell types used for the study. HNSCC, head and neck squamous cell cancer; NSCLC, non-small
Genomic tests to personalize therapy of metastatic breast cancers. Fabrice ANDRE Gustave Roussy Villejuif, France
 Genomic tests to personalize therapy of metastatic breast cancers Fabrice ANDRE Gustave Roussy Villejuif, France Future application of genomics: Understand the biology at the individual scale Patients
Genomic tests to personalize therapy of metastatic breast cancers Fabrice ANDRE Gustave Roussy Villejuif, France Future application of genomics: Understand the biology at the individual scale Patients
A general framework for analyzing tumor subclonality using SNP array and DNA sequencing data
 Li and Li Genome Biology 2014, 15:473 METHOD A general framework for analyzing tumor subclonality using SNP array and DNA sequencing data Bo Li 1 and Jun Z Li 2 Open Access Abstract Intra-tumor heterogeneity
Li and Li Genome Biology 2014, 15:473 METHOD A general framework for analyzing tumor subclonality using SNP array and DNA sequencing data Bo Li 1 and Jun Z Li 2 Open Access Abstract Intra-tumor heterogeneity
AD (Leave blank) TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients
 AD (Leave blank) Award Number: W81XWH-12-1-0444 TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients PRINCIPAL INVESTIGATOR: Mark A. Watson, MD PhD CONTRACTING ORGANIZATION:
AD (Leave blank) Award Number: W81XWH-12-1-0444 TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients PRINCIPAL INVESTIGATOR: Mark A. Watson, MD PhD CONTRACTING ORGANIZATION:
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits
 AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
OncoPhase: Quantification of somatic mutation cellular prevalence using phase information
 OncoPhase: Quantification of somatic mutation cellular prevalence using phase information Donatien Chedom-Fotso 1, 2, 3, Ahmed Ashour Ahmed 1, 2, and Christopher Yau 3, 4 1 Ovarian Cancer Cell Laboratory,
OncoPhase: Quantification of somatic mutation cellular prevalence using phase information Donatien Chedom-Fotso 1, 2, 3, Ahmed Ashour Ahmed 1, 2, and Christopher Yau 3, 4 1 Ovarian Cancer Cell Laboratory,
Supplementary Figure 1. Copy Number Alterations TP53 Mutation Type. C-class TP53 WT. TP53 mut. Nature Genetics: doi: /ng.
 Supplementary Figure a Copy Number Alterations in M-class b TP53 Mutation Type Recurrent Copy Number Alterations 8 6 4 2 TP53 WT TP53 mut TP53-mutated samples (%) 7 6 5 4 3 2 Missense Truncating M-class
Supplementary Figure a Copy Number Alterations in M-class b TP53 Mutation Type Recurrent Copy Number Alterations 8 6 4 2 TP53 WT TP53 mut TP53-mutated samples (%) 7 6 5 4 3 2 Missense Truncating M-class
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded
 Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Liposarcoma*Genome*Project*
 LiposarcomaGenomeProject July2015! Submittedby: JohnMullen,MD EdwinChoy,MD,PhD GregoryCote,MD,PhD G.PeturNielsen,MD BradBernstein,MD,PhD Liposarcoma Background Liposarcoma is the most common soft tissue
LiposarcomaGenomeProject July2015! Submittedby: JohnMullen,MD EdwinChoy,MD,PhD GregoryCote,MD,PhD G.PeturNielsen,MD BradBernstein,MD,PhD Liposarcoma Background Liposarcoma is the most common soft tissue
Reviewers' comments: Reviewer #1 (Remarks to the Author):
 Reviewers' comments: Reviewer #1 (Remarks to the Author): In this study the authors analysed 18 deep penetrating nevi for oncogenic genomic changes (single nucleotide variations, insertions/deletions,
Reviewers' comments: Reviewer #1 (Remarks to the Author): In this study the authors analysed 18 deep penetrating nevi for oncogenic genomic changes (single nucleotide variations, insertions/deletions,
Clustered mutations of oncogenes and tumor suppressors.
 Supplementary Figure 1 Clustered mutations of oncogenes and tumor suppressors. For each oncogene (red dots) and tumor suppressor (blue dots), the number of mutations found in an intramolecular cluster
Supplementary Figure 1 Clustered mutations of oncogenes and tumor suppressors. For each oncogene (red dots) and tumor suppressor (blue dots), the number of mutations found in an intramolecular cluster
Supplementary Materials for
 www.sciencemag.org/content/355/6332/eaai8478/suppl/dc1 Supplementary Materials for Decoupling genetics, lineages, and microenvironment in IDH-mutant gliomas by single-cell RNA-seq Andrew S. Venteicher,
www.sciencemag.org/content/355/6332/eaai8478/suppl/dc1 Supplementary Materials for Decoupling genetics, lineages, and microenvironment in IDH-mutant gliomas by single-cell RNA-seq Andrew S. Venteicher,
period. The distribution of PREs and TCs, stratified by synoptic category
 3. Climatology of PREs during 1988 2008 3.1 Overview A total of 56 PREs associated with 38 Atlantic basin TCs were identified for the 1988 2008 period. The distribution of PREs and TCs, stratified by synoptic
3. Climatology of PREs during 1988 2008 3.1 Overview A total of 56 PREs associated with 38 Atlantic basin TCs were identified for the 1988 2008 period. The distribution of PREs and TCs, stratified by synoptic
SiFit: inferring tumor trees from single-cell sequencing data under finite-sites models
 Zafar et al. Genome Biology (2017) 18:178 DOI 10.1186/s13059-017-1311-2 METHOD Open Access SiFit: inferring tumor trees from single-cell sequencing data under finite-sites models Hamim Zafar 1,2, Anthony
Zafar et al. Genome Biology (2017) 18:178 DOI 10.1186/s13059-017-1311-2 METHOD Open Access SiFit: inferring tumor trees from single-cell sequencing data under finite-sites models Hamim Zafar 1,2, Anthony
of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed.
 Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Cancer Informatics Lecture
 Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity
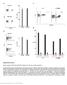 Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
Supplementary Methods
 Supplementary Methods Short Read Preprocessing Reads are preprocessed differently according to how they will be used: detection of the variant in the tumor, discovery of an artifact in the normal or for
Supplementary Methods Short Read Preprocessing Reads are preprocessed differently according to how they will be used: detection of the variant in the tumor, discovery of an artifact in the normal or for
Module 3: Pathway and Drug Development
 Module 3: Pathway and Drug Development Table of Contents 1.1 Getting Started... 6 1.2 Identifying a Dasatinib sensitive cancer signature... 7 1.2.1 Identifying and validating a Dasatinib Signature... 7
Module 3: Pathway and Drug Development Table of Contents 1.1 Getting Started... 6 1.2 Identifying a Dasatinib sensitive cancer signature... 7 1.2.1 Identifying and validating a Dasatinib Signature... 7
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma. Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute
 Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH
 EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
Module 2: Target Discovery and Validation
 Module 2: Target Discovery and Validation Table of Contents 1.1 Getting Started... 6 1.2 Nomination, validation, and associations of AGTR1 as an outlier in a subset of breast cancers... 7 1.2.1 AGTR1 discovery
Module 2: Target Discovery and Validation Table of Contents 1.1 Getting Started... 6 1.2 Nomination, validation, and associations of AGTR1 as an outlier in a subset of breast cancers... 7 1.2.1 AGTR1 discovery
COSMIC - Catalogue of Somatic Mutations in Cancer
 COSMIC - Catalogue of Somatic Mutations in Cancer http://cancer.sanger.ac.uk/cosmic https://academic.oup.com/nar/articl e-lookup/doi/10.1093/nar/gkw1121 Data In Large-scale systematic screens Detailed
COSMIC - Catalogue of Somatic Mutations in Cancer http://cancer.sanger.ac.uk/cosmic https://academic.oup.com/nar/articl e-lookup/doi/10.1093/nar/gkw1121 Data In Large-scale systematic screens Detailed
R2 Training Courses. Release The R2 support team
 R2 Training Courses Release 2.0.2 The R2 support team Nov 08, 2018 Students Course 1 Student Course: Investigating Intra-tumor Heterogeneity 3 1.1 Introduction.............................................
R2 Training Courses Release 2.0.2 The R2 support team Nov 08, 2018 Students Course 1 Student Course: Investigating Intra-tumor Heterogeneity 3 1.1 Introduction.............................................
The feasibility of circulating tumour DNA as an alternative to biopsy for mutational characterization in Stage III melanoma patients
 The feasibility of circulating tumour DNA as an alternative to biopsy for mutational characterization in Stage III melanoma patients ASSC Scientific Meeting 13 th October 2016 Prof Andrew Barbour UQ SOM
The feasibility of circulating tumour DNA as an alternative to biopsy for mutational characterization in Stage III melanoma patients ASSC Scientific Meeting 13 th October 2016 Prof Andrew Barbour UQ SOM
Supplementary Figure 1: Classification scheme for non-synonymous and nonsense germline MC1R variants. The common variants with previously established
 Supplementary Figure 1: Classification scheme for nonsynonymous and nonsense germline MC1R variants. The common variants with previously established classifications 1 3 are shown. The effect of novel missense
Supplementary Figure 1: Classification scheme for nonsynonymous and nonsense germline MC1R variants. The common variants with previously established classifications 1 3 are shown. The effect of novel missense
Although the authors have addressed some of my comments form the previous round of reviews, I still have major concerns:
 Editorial Note: this manuscript has been previously reviewed at another journal that is not operating a transparent peer review scheme. This document only contains reviewer comments and rebuttal letters
Editorial Note: this manuscript has been previously reviewed at another journal that is not operating a transparent peer review scheme. This document only contains reviewer comments and rebuttal letters
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
Characterisation of structural variation in breast. cancer genomes using paired-end sequencing on. the Illumina Genome Analyser
 Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Abstract. Optimization strategy of Copy Number Variant calling using Multiplicom solutions APPLICATION NOTE. Introduction
 Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
PBZ FT01_PBZ FT01_TZ FT01_NZ. interface zone (I) tumor zone (TZ) necrotic zone (NZ)
 Oncotarget, Supplementary Materials www.impactjournals.com/oncotarget/ SUPPLEMENTRY FLES ndividuals factor map (P) FT_ FT_ FT_ Dim (.%) Dim (.%) >% peripheral brain zone () around % interface zone () FT
Oncotarget, Supplementary Materials www.impactjournals.com/oncotarget/ SUPPLEMENTRY FLES ndividuals factor map (P) FT_ FT_ FT_ Dim (.%) Dim (.%) >% peripheral brain zone () around % interface zone () FT
Nature Genetics: doi: /ng Supplementary Figure 1. Workflow of CDR3 sequence assembly from RNA-seq data.
 Supplementary Figure 1 Workflow of CDR3 sequence assembly from RNA-seq data. Paired-end short-read RNA-seq data were mapped to human reference genome hg19, and unmapped reads in the TCR regions were extracted
Supplementary Figure 1 Workflow of CDR3 sequence assembly from RNA-seq data. Paired-end short-read RNA-seq data were mapped to human reference genome hg19, and unmapped reads in the TCR regions were extracted
Genomic Medicine: What every pathologist needs to know
 Genomic Medicine: What every pathologist needs to know Stephen P. Ethier, Ph.D. Professor, Department of Pathology and Laboratory Medicine, MUSC Director, MUSC Center for Genomic Medicine Genomics and
Genomic Medicine: What every pathologist needs to know Stephen P. Ethier, Ph.D. Professor, Department of Pathology and Laboratory Medicine, MUSC Director, MUSC Center for Genomic Medicine Genomics and
Biomarkers in Imunotherapy: RNA Signatures as predictive biomarker
 Biomarkers in Imunotherapy: RNA Signatures as predictive biomarker Joan Carles, MD PhD Director GU, CNS and Sarcoma Program Department of Medical Oncology Vall d'hebron University Hospital Outline Introduction
Biomarkers in Imunotherapy: RNA Signatures as predictive biomarker Joan Carles, MD PhD Director GU, CNS and Sarcoma Program Department of Medical Oncology Vall d'hebron University Hospital Outline Introduction
README file for GRASTv1.0.pl
 README file for GRASTv.0.pl Genome Reduction Analysing Software Tool (GRAST). Produced by Christina Toft and Mario A. Fares Date 03/04/06 Reference and more information: Toft, C and Fares, MA (2006). GRAST:
README file for GRASTv.0.pl Genome Reduction Analysing Software Tool (GRAST). Produced by Christina Toft and Mario A. Fares Date 03/04/06 Reference and more information: Toft, C and Fares, MA (2006). GRAST:
Supplementary Figure 1. Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Nature Immunology: doi: /ni.
 Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 LD (r 2 ) between the A3AB deletion and all markers in a 400-kb APOBEC3 region in 1000 Genomes Project populations. Populations: CEU, individuals of European ancestry from Utah,
Supplementary Figure 1 LD (r 2 ) between the A3AB deletion and all markers in a 400-kb APOBEC3 region in 1000 Genomes Project populations. Populations: CEU, individuals of European ancestry from Utah,
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Table S1. Relative abundance of AGO1/4 proteins in different organs. Table S2. Summary of smrna datasets from various samples.
 Supplementary files Table S1. Relative abundance of AGO1/4 proteins in different organs. Table S2. Summary of smrna datasets from various samples. Table S3. Specificity of AGO1- and AGO4-preferred 24-nt
Supplementary files Table S1. Relative abundance of AGO1/4 proteins in different organs. Table S2. Summary of smrna datasets from various samples. Table S3. Specificity of AGO1- and AGO4-preferred 24-nt
Table S1: Kinetic parameters of drug and substrate binding to wild type and HIV-1 protease variants. Data adapted from Ref. 6 in main text.
 Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
BWA alignment to reference transcriptome and genome. Convert transcriptome mappings back to genome space
 Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
SUPPLEMENTARY INFORMATION
 Somatic ERCC2 Mutations Are Associated with a Distinct Genomic Signature in Urothelial Tumors Jaegil Kim, Kent W Mouw, Paz Polak, Lior Z Braunstein, Atanas Kamburov, Grace Tiao, David J Kwiatkowski, Jonathan
Somatic ERCC2 Mutations Are Associated with a Distinct Genomic Signature in Urothelial Tumors Jaegil Kim, Kent W Mouw, Paz Polak, Lior Z Braunstein, Atanas Kamburov, Grace Tiao, David J Kwiatkowski, Jonathan
Qué hemos aprendido hasta hoy? What have we learned so far?
 Qué hemos aprendido hasta hoy? What have we learned so far? Luís Costa Hospital de Santa Maria & Instituto de Medicina Molecular Faculdade de Medicina de Lisboa Disclosures Research Grants: Amgen; Novartis;
Qué hemos aprendido hasta hoy? What have we learned so far? Luís Costa Hospital de Santa Maria & Instituto de Medicina Molecular Faculdade de Medicina de Lisboa Disclosures Research Grants: Amgen; Novartis;
Integration of Cancer Genome into GECCO- Genetics and Epidemiology of Colorectal Cancer Consortium
 Integration of Cancer Genome into GECCO- Genetics and Epidemiology of Colorectal Cancer Consortium Ulrike Peters Fred Hutchinson Cancer Research Center University of Washington U01-CA137088-05, PI: Peters
Integration of Cancer Genome into GECCO- Genetics and Epidemiology of Colorectal Cancer Consortium Ulrike Peters Fred Hutchinson Cancer Research Center University of Washington U01-CA137088-05, PI: Peters
Supplementary figure 1: LII/III GIN-cells show morphological characteristics of MC
 1 2 1 3 Supplementary figure 1: LII/III GIN-cells show morphological characteristics of MC 4 5 6 7 (a) Reconstructions of LII/III GIN-cells with somato-dendritic compartments in orange and axonal arborizations
1 2 1 3 Supplementary figure 1: LII/III GIN-cells show morphological characteristics of MC 4 5 6 7 (a) Reconstructions of LII/III GIN-cells with somato-dendritic compartments in orange and axonal arborizations
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Dr Yvonne Wallis Consultant Clinical Scientist West Midlands Regional Genetics Laboratory
 Dr Yvonne Wallis Consultant Clinical Scientist West Midlands Regional Genetics Laboratory Personalised Therapy/Precision Medicine Selection of a therapeutic drug based on the presence or absence of a specific
Dr Yvonne Wallis Consultant Clinical Scientist West Midlands Regional Genetics Laboratory Personalised Therapy/Precision Medicine Selection of a therapeutic drug based on the presence or absence of a specific
Department of Chemistry, Université de Montréal, C.P. 6128, Succursale centre-ville, Montréal, Québec, H3C 3J7, Canada.
 Phosphoproteome dynamics of Saccharomyces cerevisiae under heat shock and cold stress Evgeny Kanshin 1,5, Peter Kubiniok 1,2,5, Yogitha Thattikota 1,3, Damien D Amours 1,3 and Pierre Thibault 1,2,4 * 1
Phosphoproteome dynamics of Saccharomyces cerevisiae under heat shock and cold stress Evgeny Kanshin 1,5, Peter Kubiniok 1,2,5, Yogitha Thattikota 1,3, Damien D Amours 1,3 and Pierre Thibault 1,2,4 * 1
Tutorial: RNA-Seq Analysis Part II: Non-Specific Matches and Expression Measures
 : RNA-Seq Analysis Part II: Non-Specific Matches and Expression Measures March 15, 2013 CLC bio Finlandsgade 10-12 8200 Aarhus N Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com support@clcbio.com
: RNA-Seq Analysis Part II: Non-Specific Matches and Expression Measures March 15, 2013 CLC bio Finlandsgade 10-12 8200 Aarhus N Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com support@clcbio.com
Supplementary Figure 1. Schematic diagram of o2n-seq. Double-stranded DNA was sheared, end-repaired, and underwent A-tailing by standard protocols.
 Supplementary Figure 1. Schematic diagram of o2n-seq. Double-stranded DNA was sheared, end-repaired, and underwent A-tailing by standard protocols. A-tailed DNA was ligated to T-tailed dutp adapters, circularized
Supplementary Figure 1. Schematic diagram of o2n-seq. Double-stranded DNA was sheared, end-repaired, and underwent A-tailing by standard protocols. A-tailed DNA was ligated to T-tailed dutp adapters, circularized
Supplementary Figure 1
 Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Importance of minor TP53 mutated clones in the clinic
 Importance of minor TP53 mutated clones in the clinic Davide Rossi, M.D., Ph.D. Hematology IOSI - Oncology Institute of Southern Switzerland IOR - Institute of Oncology Reserach Bellinzona - Switzerland
Importance of minor TP53 mutated clones in the clinic Davide Rossi, M.D., Ph.D. Hematology IOSI - Oncology Institute of Southern Switzerland IOR - Institute of Oncology Reserach Bellinzona - Switzerland
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
COMPUTATIONAL OPTIMISATION OF TARGETED DNA SEQUENCING FOR CANCER DETECTION
 COMPUTATIONAL OPTIMISATION OF TARGETED DNA SEQUENCING FOR CANCER DETECTION Pierre Martinez, Nicholas McGranahan, Nicolai Juul Birkbak, Marco Gerlinger, Charles Swanton* SUPPLEMENTARY INFORMATION SUPPLEMENTARY
COMPUTATIONAL OPTIMISATION OF TARGETED DNA SEQUENCING FOR CANCER DETECTION Pierre Martinez, Nicholas McGranahan, Nicolai Juul Birkbak, Marco Gerlinger, Charles Swanton* SUPPLEMENTARY INFORMATION SUPPLEMENTARY
