Analysis of neural stem cell self-renewal and differentiation by transgenic RNAi in Drosophila
|
|
|
- Kristian Gardner
- 5 years ago
- Views:
Transcription
1 Institutional Repository of the University of Basel University Library Schoenbeinstrasse CH-4056 Basel, Switzerland Year: 2013 Analysis of neural stem cell self-renewal and differentiation by transgenic RNAi in Drosophila Jiang, Yanrui and Reichert, Heinrich Posted at edoc, University of Basel Official URL: Originally published as: Jiang, Yanrui and Reichert, Heinrich. (2013) Analysis of neural stem cell self-renewal and differentiation by transgenic RNAi in Drosophila. Archives of biochemistry and biophysics, Vol. 534, H S
2 Analysis of neural stem cell self-renewal and differentiation by transgenic RNAi in Drosophila Yanrui Jiang*, Heinrich Reichert Biozentrum, University of Basel, Klingelbergstrasse 50/70, CH-4056 Basel, Switzerland *Corresponding author Address: Biozentrum, University of Basel, Klingelbergstrasse 50/70, CH-4056 Basel, Switzerland Tel: Fax: addresses: (Y. Jiang), (H. Reichert).
3 Abstract The fruit fly, Drosophila melanogaster, has proved to be a useful model organism for studying the biology of neural stem cells. Notably, significant progress has been made in identifying the molecular mechanisms that regulate the asymmetric cell divisions of the neural stem cell-like neuroblasts during brain development. Recently, the emerging technology of genome-wide transgenic RNA interference (RNAi), which makes it possible to analyze complicated developmental processes in a targeted, tissue-specific way, has been used for the analysis of gene function in Drosophila neuroblasts. Here, we review the key molecular mechanisms that regulate the asymmetric cell divisions of neuroblasts during brain development in Drosophila. We then summarize recent genome-wide transgenic RNAi screens in Drosophila and report on the identification of new regulators and gene networks that are required in balancing neuroblast selfrenewal and differentiation. Keywords: neural stem cell, neuroblast, self-renewal, differentiation, RNAi, Drosophila
4 Highlights Neuroblasts are neural stem cells in the Drosophila brain. Cell fate determinants control self-renewal and differentiation of neuroblasts. Genome-wide transgenic RNAi identify new genes regulating neuroblasts proliferation. Drosophila neuroblasts are a new model to study cancer stem cells. Abbreviations used: apkc, atypical PKC; Brat, Brain tumor; GMC, ganglion mother cell; INP, intermediate neural progenitor; Insc, Inscuteable; lgl, lethal (2) giant larvae; Loco, Locomotion defects; Mud, Mushroom body defect; Pins, Partner of inscuteable; Pon, Partner of Numb; Pros, Prospero; RISC, RNA-induced silencing complex; RNAi, RNA interference; TRiP, Transgenic RNAi Project; VDRC, Vienna Drosophila RNAi Center.
5 Introduction The central nervous system in both vertebrates and invertebrates comprises complex networks of neuronal circuits. The neural cells that make up these networks are derived from neural stem cells, which have the ability to self-renew and to give rise to differentiated neuronal and glial progeny through asymmetric and proliferative cell divisions. Balancing self-renewal and differentiation of neural stem cells is essential during development, because dysregulation of genes controlling asymmetric cell division can cause severe developmental defects, and furthermore, lead to the formation of brain tumors [1-3]. In the past two decades, the fruit fly, Drosophila melanogaster, has proved to be a powerful model organism for studying the biology of neural stem cells [4-7]. Drosophila neural stem cells are called neuroblasts for historical reasons. Many important genetic elements and signaling pathways that regulate the asymmetric cell divisions of Drosophila neuroblasts have been characterized and analyzed in great detail [5,8,9]. However, our understanding of the molecular and cellular mechanisms controlling self-renewal and differentiation of Drosophila neuroblasts is still incomplete. Recently, significant progress in analyzing these mechanisms has been attained through the use of genome-wide RNA interference (RNAi) for the identification and functional analysis of genes involved in neuroblast proliferation in the developing brain. This has been possible due to the generation and availability of several independent genome-wide transgenic RNAi libraries, which are based on the binary Gal4/UAS expression system that allow analysis of complicated developmental processes in a targeted, tissue-specific way [10-13]. In this review, we focus on the biology of Drosophila neuroblasts and summarize the key molecular mechanisms that regulate the asymmetric cell divisions of these neuroblasts during
6 brain development. We then introduce recent genome-wide RNAi screens in Drosophila. Finally, we review the application of genome-wide transgenic RNAi in the analysis of selfrenewal and differentiation of Drosophila neuroblasts and report on the identification of new regulators and gene networks that are required in balancing neuroblast self-renewal and differentiation. Neural stem cells in Drosophila Neuroblasts, the neural stem cells in Drosophila, have the characteristic feature of the neural stem cells in vertebrates, namely to divide asymmetrically to self-renew while generating a wide range of more differentiated progeny [3-7]. In Drosophila, neuroblasts give rise to differentiated neurons and glia cells in the central brain, optic lobes, and ventral nerve cord [14]. In this review, we will focus primarily on the mechanisms that underlie the specification and homeostasis of the neuroblasts in the central brain; similar mechanisms have been shown to operate in the neuroblasts of the optic lobes and ventral nerve cord, which have been recently reviewed elsewhere [3,5,6]. In Drosophila, neurogenesis starts during early embryonic development [15]. In the embryonic neuroectoderm, groups of cells acquire the potential to become neuroblasts by expressing proneural genes, such as the genes of the achaete-scute complex and daughterless (Fig. 1A) [15,16]. From each of these equivalence groups a single cell is selected to become a neuroblast through a process called lateral inhibition, which is mediated by the Notch signaling (Fig. 1A) [16]. The specified neuroblasts enlarge in size, delaminate basally from the neuroectoderm and then go on to divide asymmetrically along their apical-basal axis in a stem cell-like manner (Fig. 1A) [6,7].
7 In the developing brain, neuroblasts undergo two phases of proliferation. The first takes place during embryogenesis, and the neuroblasts only go through a limited number of cell divisions to generate the primary neurons of the larval brain [14,17,18]. After a quiescent phase at the end of embryogenesis, most of the brain neuroblasts reinitiate proliferation in a second phase of neurogenesis during larval development to produce numerous secondary neurons in the adult brain [18-21]. These secondary neurons make synaptic interconnections and form adult-specific neuropile structures during pupal development [18,22]. The clonal unit generated by a neuroblast, consisting of primary neurons as well as secondary neurons, is referred to as the neuroblast s lineage [18,22,23]. Approximately 100 neuroblasts have been identified in each hemisphere of the Drosophila brain, and these can be divided into two types (Fig. 1B) [7,8,24-27]. The type I neuroblasts, which constitute the majority of the brain neuroblasts, divide in a simple way. During each division, type I neuroblasts give rise to a larger daughter cell, the renewed neuroblast, and a smaller daughter cell called a ganglion mother cell (GMC). Each GMC divides only once to generate two postmitotic cells that further differentiate into neurons or glia cells (Fig. 2A) [7,8,24]. Recently, a second type of neuroblast, termed a type II neuroblast, has been identified in the Drosophila brain [25-27]. Unlike type I neuroblasts, there are only eight type II neuroblasts in each brain hemisphere. Six type II neuroblasts localize at the dorsomedial edge of the posterior brain lobe, and the other two are formed at a more lateral position (Fig. 1B) [25-27]. In contrast to type I neuroblasts, the type II neuroblasts divide in a more complex way to self-renew and to give rise to an intermediate neural progenitor (INP) cell, which undergoes several rounds of cell division, each of which results in self-renewal of the INP and in the generation of a GMC that further produces two progeny (Fig. 2B) [25-27]. Because of the amplification of proliferation through the divisions of INP cells, a significant increase in neural cell number occurs in type II
8 neuroblast lineages. Thus, whereas most type I neuroblast lineages consist of approximately 100 to 150 neurons, type II neuroblasts typically give rise to lineages that are 3 to 5 times larger [25]. In addition, to counter-balance the amplification of proliferation, excess neurons in type II lineages are eliminated through programmed cell death, a process that is essential for the correct neuropile innervation in the adult brain [28]. Asymmetric cell divisions of Drosophila neuroblasts Stem cell divisions in Drosophila, as in other animals, can be controlled by cell-intrinsic as well as cell-extrinsic mechanisms, to generate two daughter cells with different cell fates [29]. While the extrinsic regulation of stem cell division has been well-studied in the fly ovaries [30], the asymmetric cell divisions of the neuroblasts appear to be controlled primarily by cell-intrinsic regulatory mechanisms [8,31]. This intrinsic regulation of asymmetric neuroblast divisions is achieved by the asymmetric localization of a number of cell fate determinants within the neuroblast during mitosis and subsequently by the segregation of these proteins into only one of the two daughter cells [8,31]. During mitosis, two protein complexes are formed on opposite sides of the neuroblast. On the apical side, the proteins Bazooka, Par-6, atypical PKC (apkc), Inscuteable (Insc), Partner of inscuteable (Pins), Gαi, Locomotion defects (Loco), and Mushroom body defect (Mud) form an apical complex; whereas on the opposite basal side, the cell fate determinants Brain tumor (Brat), Prospero (Pros), and Numb, as well as two adaptor proteins Miranda and Partner of Numb (Pon) are localized in a basal complex [8,31]. In the apical complex, Bazooka, Par-6 and apkc constitute the Par complex, which already localizes to the apical cortex during interphase and establishes the apical-basal polarity in the neuroblast [32-36]. The Pins, Gαi, Loco, and Mud proteins are linked to the Par complex through
9 Insc and are required for spindle orientation [37-45]. Although all three proteins of the Par complex are preferentially segregated into the new neuroblast after cell division, and mutations in any of them affect the localization of the other apical proteins and the spindle orientation, the Par complex does not seem to control cell fate directly [8]. Instead, these proteins are thought to exert their function by inducing and restricting cell fate determinant proteins to localize to the basal side of the dividing neuroblast [8]. In Drosophila, three cell fate determinants, Brat, Pros, and Numb, have been characterized and studied in some detail [5-8]. All three proteins asymmetrically localize to the basal cortex during neuroblast division and subsequently are inherited only by the smaller GMC. In the GMC, these proteins are thought to play two essential roles: first, to inhibit cell proliferation by suppressing the expression of neuroblast-specific genes and promoting cell cycle exit; second, to trigger cell differentiation by activating a neural cell differentiative program [5-8]. Numb is a membraneassociated protein and it suppresses the Notch signaling pathway in the GMC [46,47]. Pros is a homeodomain containing transcription factor that is asymmetrically segregated into the GMC where it enters the nucleus and activates or represses more than 700 target genes that may be required for neuronal differentiation [48-51]. Brat is a RNA-binding protein that negatively regulates cell growth and ribosomal RNA synthesis [52-55]. Mutations in any of the three determinant genes result in the overproliferation of the neuroblasts and the formation of brain tumors [53-57]. The observation of overgrowth and tumorigenesis in the mutant brains have recently made Drosophila neuroblasts an emerging model to study cancer stem cells (see below) [8,9,31]. Although numerous studies have been made using classic genetic and molecular methods to investigate the functions of the three cell fate determinants in Drosophila neuroblasts, the complete gene networks controlled by these proteins in the regulation of neuroblast self-renewal
10 and differentiation are still poorly understood [5-8]. To obtain insight into these gene networks, large-scale genome-wide transgenic RNAi technology has recently been used in order to identify novel genes that are potential regulators in the self-renewal and proliferation of Drosophila neuroblasts [12,13]. Genome-wide RNAi screens in Drosophila RNA interference (RNAi) is a cellular process that can sufficiently suppress (knock down) the expression of genes at the post-transcription level. RNAi is mediated by the RNA-induced silencing complex (RISC) and short double-stranded RNA molecules that recognize their target genes in a sequence-specific manner [58,59]. These RNA molecules are either encoded endogenously (micrornas, sirnas) or introduced exogenously [60,61]. With the availability of a large number of genome sequences for different organisms, it is possible to construct RNAi libraries that target and silence every individual gene, thus facilitating large-scale genome-wide analysis of gene functions [10]. In the past decade, a variety of RNAi libraries have been generated and screens have been made using different cell cultures or model organisms including C. elegans, Planaria, and Drosophila. Early RNAi-based screens in Drosophila were mostly carried out in cultured cells, and a wide range of diverse biological processes have been studied in cell culture models [10]. To date, cellbased RNAi screens have been used to identify new candidate genes in genetic networks such as those that regulate growth and viability [62], cell cycle [63], cell death [64], cell signaling [65], and host-pathogen interactions [66]. Although this type of screen has provided new findings for many basic cellular processes, it is not suitable for the analysis of more complex multicellular developmental processes that occur within the intact organism.
11 In order to analyze complicated cellular and developmental processes in a tissue-specific manner in the intact animal, a second type of genome-wide screen has been developed which uses transgenic flies carrying RNAi transgenes that can be combined with the established Gal4/UAS binary expression system (Fig. 3) [67]. Currently, three independent sources of genome-wide UAS-RNAi transgenic lines are available [Vienna Drosophila RNAi Center (VDRC), Transgenic RNAi Project (TRiP) at the Harvard Medical School, and NGI-FLY RNAi Resources in Japan], such that altogether about 90% of the annotated genes in the Drosophila genome can now be targeted by these lines [67]. With the availability of these lines, genome-wide transgenic RNAi screens have been successfully used to study Notch signaling [68,69], muscle morphogenesis and functions [70], host-pathogen interactions [71], and self-renewal of neuroblasts [12,13]. Taken together, these studies show that even very complicated biological mechanisms can be systematically analyzed at the genome-wide level by transgenic RNAi. Transgenic RNAi analysis of neuroblast self-renewal and proliferation in Drosophila The suitability of large-scale genome-wide transgenic RNAi screens for the identification of candidate genes involved in the regulation of self-renewal and proliferation of Drosophila neuroblasts has been demonstrated in two recent reports [12,13]. In the first report, Knoblich and colleagues carried out a genome-wide RNAi screen to indentify genes that are involved in cytokinesis, cell growth, and differentiation in brain neuroblasts and their lineages (Fig. 4) [12]. Knock-down of candidate genes was targeted to the neuroblasts by crossing each individual UAS- RNAi line to the driver line insc-gal4 that is expressed in both type I and type II neuroblasts (Fig. 4A) [54]. In total, 17,362 RNAi lines were screened (corresponding to 89% of the Drosophila genome). 24.1% of the lines caused lethality and were further analyzed for brain phenotypes
12 (Fig. 4B). This screen identified about 600 candidate genes (832 lines) that cause an abnormal brain phenotype. The genes were then divided into subgroups according to a set of phenotypic categories based on the similarities of the abnormalities in the number, size and shape of the neuroblasts or their daughter cells (GMC, INP, or the entire lineage). For example, among these 600 candidates, a subgroup of 29 genes caused an overproliferation phenotype, suggesting that they are required for restricting neuroblast self-renewal. In addition, based on the annotated gene structure and the known or predicted molecular functions of the candidates, the authors also constructed gene networks using clustering algorithms. In this manner, they identified all putative transcription factors and chromatin regulators that form a transcriptional network for neuroblast self-renewal and also defined genes that interact with known regulators of asymmetric cell divisions and thus form an asymmetric cell division regulatory network. In a second study, Doe and colleagues combined both transcriptional profiling analysis and transgenic RNAi knock-down experiments, again with the aim of characterizing genes that regulate neuroblast self-renewal in Drosophila [13]. Using microarray-based transcriptome analysis, the authors first studied six different Drosophila mutants, including brat and lethal (2) giant larvae (lgl), which are known to generate ectopic neuroblasts in the third instar larval brains. By comparing the transcriptional profiling of these six mutants with wild-type, they identified around 1000 genes with elevated expression in the mutant brains, suggesting that they are likely to be expressed in neuroblasts and might be involved in neuroblast homeostasis. To investigate the function of these genes, transgenic RNAi knock-down experiments were carried out in which UAS-RNAi lines were crossed with another neuroblast driver line, wor-gal4 [72]. This functional analysis yielded a list of 84 potential regulators for neuroblast self-renewal in Drosophila.
13 The majority of genes identified in both of these screens are thought to promote neuroblast selfrenewal, as targeted knocking down of these genes generally showed an underproliferation phenotype (538 of 620 genes in [12], and 72 of 84 genes in [13]). Furthermore, for the genes with an underproliferation phenotype, 46 genes were identified by both groups. Interestingly, both screens identified a rather small number of genes for which RNAi knock-down resulted in an increase in the number of neuroblasts (18 in [12], and 12 in [13]). However, among these, only two genes (miranda and Ssrp) were found by both groups. The reason for this low degree of overlap between the two studies is not clear; differential expression patterns of the Gal4 driver lines, different genetic backgrounds of the fly strains used, as well as differences in the design of the two screens might be responsible, at least in part. The altered expression of specific genes was also different among the mutants with ectopic neuroblast phenotypes (with elevated expression of a given gene in one mutant, but decreased expression of the same gene in another), underscoring the fact that proliferation control in neuroblasts is a highly complex process. In summary, these two recent transgenic RNAi screens have identified a large number of novel genes that appear to play important roles in controlling the balance between self-renewal and differentiation of Drosophila neuroblasts. Although the precise function of each of these genes needs to be validated using classical genetic methods, these studies indeed provide a valuable source of candidate genes for further analysis. Moreover, since a large subset of the candidate genes identified in these screens have conserved orthologs in mammals, further analysis of these genes in murine models is likely to provide new insight into mammalian neural stem cell research as well. Drosophila neuroblasts as a model to study brain tumors
14 The investigation of genes involved in the balance between neuroblast self-renewal and differentiation is important for understanding the mechanisms of neural stem cell action in normal brain development. Moreover, there is an additional motivation for this type of research. In recent years, increasing evidence indicates that a small fraction of stem cells, or stem cell-like progenitor cells, might be the origin of certain cancers (cancer stem cells) including cancers of the brain [73]. However, the molecular mechanisms underlying normal and tumorigenic development in these stem cells are largely unknown. Therefore, understanding how the balance between self-renewal and proliferation is disrupted in tumor stem cells at the cellular and molecular level is likely to be essential for future therapeutic implications both in tumor biology and in stem cell-based regenerative medicine [8,9]. A first Drosophila model for cancer study was established more than three decades ago, when it was shown that mutations in genes such as lethal (1) disc large, lgl, lethal (2) giant disc, and lethal (3) giant larvae can lead to malignant overproliferation in Drosophila brains or discs [74]. More recently, comparable overgrowth phenotypes were also observed in brat, pros, and numb mutant brains, indicating that these cell fate determinants also act as tumor suppressor genes in Drosophila neuroblasts [53,55-57]. Indeed, the tumor suppressor function of genes involved in asymmetric cell division has been unequivocally demonstrated using a transplantation assay in which brain tissue from mutants of brat, pros, or numb transplanted into wild-type adult flies formed large malignant and metastatic tumors that eventually killed the hosts [75]. In view of these findings, it seems apparent that many other genes involved in neuroblast selfrenewal and differentiation may also act as tumor suppressors or oncogenes [9,75]. Thus an analysis of the potential role of the candidate genes involved in tumorigenesis from the recent and ongoing transgenic RNAi screens is likely to increase our mechanistic insight into the formation of brain tumors that derive from mutant neuroblasts. Indeed, given that transgenic RNAi can be
15 targeted to specific cells via Gal4 drivers and be activated or deactivated at specific times via Gal80 inhibitors, this emerging genetic technology seems to be ideally suited for investigating the molecular mechanisms underlying tumor formation and progression as well as tumor abrogation in the Drosophila neural stem cell model. Acknowledgements This work is supported by the Swiss National Science Foundation (SNSF) and SNSF-NFP63 Stem cells and regenerative medicine.
16 References [1] J.J. Breunig, T.F. Haydar, P. Rakic, Neuron 70 (2011) [2] F.T. Merkle, A. Alvarez-Buylla, Curr. Opin. Cell Biol. 18 (2006) [3] A.H. Brand, F.J. Livesay, Neuron 70 (2011) [4] R. Urbach, G.M. Technau, Bioessays 26 (2004) [5] H. Reichert, Results Probl. Cell Differ. 53 (2011) [6] B. Egger, J.M. Chell, A.H. Brand, Philos. Trans. R. Soc. B 363 (2008) [7] C.Q. Doe, Development 135 (2008) [8] J.A. Knoblich, Cell 132 (2008) [9] K.C. Chang, C. Wang, H. Wang, Bioessays 34 (2012) [10] S. Mohr, C. Bakal, N. Perrimon, Annu. Rev. Biochem. 79 (2010) [11] N. Perrimon, J.Q. Ni, L. Perkins, Cold Spring Harb. Perspect Biol. 2 (2010) a [12] R.A. Neumüller, C. Richter, A. Fischer, M. Novatchkova, K.G. Neumüller, J.A. Knoblich, Cell Stem Cell 8 (2011) [13] T.D. Carney, M.R. Miller, K.J. Robinson, O.A. Bayraktar, J.A. Osterhout, C.Q. Doe, Dev. Biol. 361 (2012) [14] J.A. Campos-Ortega, V. Hartenstein, The embryonic development of Drosophila melanogaster, second ed., Springer, Heidelberg, 1997.
17 [15] J.A. Campos-Ortega, Early neurogenesis in Drosophila melanogaster, in: M. Bate, A.M. Arias (Eds), The development of Drosophila melanogaster, Cold Spring Harbor Laboratory Press, New York, 1993, pp [16] C.S. Goodman, C.Q. Doe, Embryonic development of the Drosophila central nervous system, in: M. Bate, A.M. Arias (Eds), The development of Drosophila melanogaster, Cold Spring Harbor Laboratory Press, New York, 1993, pp [17] V. Hartenstein, E. Rudloff, J.A. Campos-Ortega, Rouxs Arch. Dev. Biol. 196 (1987) [18] V. Hartenstein, S. Spindler, W. Pereanu, S. Fung, The development of the Drosophila larval brain, in: G. Technau (Ed), Brain development in Drosophila melanogaster. Landes Bioscience, Austin, 2008, pp [19] J.W. Truman, M. Bate, Dev. Biol. 125 (1988) [20] A. Prokop, G.M. Technau, Development 111 (1991) [21] K. Ito, Y. Hotta, Dev. Biol. 149 (1992) [22] S.R. Spindler, V. Hartenstein, Dev. Genes Evol. 220 (2010) [23] K. Ito, T. Awasaki, Clonal unit architecture of the adult fly brain, in: G. Technau (Ed), Brain development in Drosophila melanogaster. Landes Bioscience, Austin, 2008, pp [24] M. Weng, C.Y. Lee, Curr. Opin. Neurobiol. 21 (2011) [25] B. Bello, N. Izergina, E. Caussinus, H. Reichert, Neural Dev. (2008) 3:5. [26] S.K. Bowman, V. Rolland, J. Betschinger, K.A. Kinesey, G. Emery, J.A. Knoblich, Dev. Cell 14 (2008)
18 [27] J.Q. Boone, C.Q. Doe, Dev. Neurobiol. 68 (2008) [28] Y. Jiang, H. Reichert, Neural Dev. (2012) 7:3. [29] H.R. Horvitz, I. Herskowitz, Cell 68 (1992) [30] L. Li, T. Xie, Annu. Rev. Cell Dev. Biol. 21 (2005) [31] R.A. Neumüller, J.A. Knoblich, Genes Dev 23 (2009) [32] A. Wodarz, A. Ramrath, U. Kuchinke, E. Knust, Nature 402 (1999) [33] M. Schober, M. Schaefer, J.A. Knoblich, Nature 402 (1999) [34] M. Petronczki, J.A. Knoblich, Nat Cell Biol. 3 (2001) [35] A. Wodarz, A. Ramrath, A. Grimm, E. Knust, J. Cell Biol. 150 (2000) [36] M.M. Rolls, R. Albertson, H.P. Shih, C.Y. Lee, C.Q. Doe, J. Cell Biol. 163 (2003) [37] M, Schaefer, M. Petronczki, D. Dorner, M. Forte, J.A. Knoblich, Cell 107 (2001) [38] F. Yu, Y. Cai, R. Kaushik, X. Yang, W. Chia, J. Cell Biol. 162 (2003) [39] Y. Izumi, N. Ohta, A. Itoh-Furuya, N. Fuse, F. Matsuzaki, J. Cell Biol. 164 (2004) [40] M. Schaefer, A. Shevchenko, A. Shevchenko, J.A. Knoblich, Curr. Biol. 10 (2000) [41] F. Yu, X. Morin, Y. Cai, X. Yang, W. Chia, Cell 100 (2000) [42] F. Yu, H. Wang, H. Qian, R. Kaushik, M. Bownes, X. Yang, W. Chia, Genes Dev. 19 (2005) [43] S.K. Bowman, R.A. Neumuller, M. Novatchkova, Q. Du, J.A. Knoblich, Dev. Cell 10 (2006)
19 [44] Y. Izumi, N. Ohta, K. Hisata, T. Raabe, F. Matsuzaki, Nat. Cell Biol. 8 (2006) [45] K.H. Siller, C. Cabernard, C.Q. Doe, Nat. Cell Biol. 8 (2006) [46] M.S. Rhyu, L.Y. Jan, Y.N. Jan Cell 76 (1994) [47] E.P. Spana, C. Kopczynski, C.S. Goodman, C.Q. Doe, Development 121 (1995) [48] J. Hirata, H. Nakagoshi, Y. Nabeshima, F. Matsuzaki, Nature 377 (1995) [49] J.A. Knoblich, L.Y. Jan, Y.N. Jan, Nature 377 (1995) [50] E.P. Spana, C.Q. Doe, Development 121 (1995) [51] S.P. Choksi, T.D. Southall, T. Bossing, K. Edoff, E. de Wit, B.E. Fischer, B. van Steensel, G. Micklem, A.H. Brand, Dev. Cell 11 (2006) [52] D.J. Frank, B.A. Edgar, M.B. Roth, Development 129 (2002) [53] B. Bello, H. Reichert, F. Hirth, Development 133 (2006) [54] J. Betschinger, K. Mechtler, J.A. Knoblich, Cell 124 (2006) [55] C.Y. Lee, B.D. Wilkinson, S.E. Siegrist, R.P. Wharton, C.Q. Doe, Dev. Cell 10 (2006) [56] C.Y. Lee, R.O. Andersen, C. Cabernard, L. Manning, K.D. Tran, M.J. Lanskey, A. Bashirullah, C.Q. Doe, Genes Dev. 20 (2006) [57] H. Wang, G.W. Somers, A. Bashirullah, U. Heberlein, F. Yu, W. Chia, Genes Dev. 20 (2006) [58] A. Fire, S. Xu, M.K. Montgomery, S.A. Kostas, S.E. Driver, C.C. Mello. Nature 391(1998) [59] P.D. Zamore, T. Tuschl, P.A. Sharp, D.P. Bartel. Cell 101 (2000)
20 [60] D.P. Bartel, Cell 136 (2009) [61] R.W. Carthew, E.J. Sontheimer, Cell 136 (2009) [62] M. Boutros, A.A. Kiger, S. Armknecht, K. Kerr, M. Hild, B. Koch, S.A. Haas, Heidelberg Fly Array Consortium, R. Paro, N. Perrimon, Science 303 (2004) [63] M. Bjorklund, M. Taipale, M. Varjosalo, J. Saharinen, J. Lahdenpera, J. Taipale, Nature 439 (2006) [64] C.H. Yi, D.K. Sogah, M. Boyce, A. Degterev, D.E. Christofferson, J. Yuan, J. Cell Biol. 179 (2007) [65] R. DasGupta, A. Kaykas, R.T. Moon, N. Perrimon, Science 308 (2005) [66] H. Agaisse, L.S. Burrack, J.A. Philips, E.J. Rubin, N. Perrimon, D.E. Higgins, Science 309 (2005) [67] G. Dietzl, D. Chen, F. Schnorrer, K.C. Su, Y. Barinova, M. Fellner, B. Gasser, K. Kinsey, S. Oppel, S. Scheiblauer, A. Couto, V. Marra, K. Keleman, B.J. Dickson, Nature 448 (2007) [68] J.L. Mummery-Widmer, M. Yamazaki, T. Stoeger, M. Novatchkova, S. Bhalerao, D. Chen, G. Dietzl, B.J. Dickson, J.A. Knoblich, Nature 458 (2009) [69] A. Saj, Z. Arziman, D. Stempfle, W. van Belle, U. Sauder, T. Horn, M. Dürrenberger, R. Paro, M. Boutros, G. Merdes, Dev. Cell 18 (2010) [70] F. Schnorrer, C. Schönbauer, C.C. Langer, G. Dietzl, M. Novatchkova, K. Schernhuber, M. Fellner, A. Azaryan, M. Radolf, A. Stark, K. Keleman, B.J. Dickson, Nature 464 (2010)
21 [71] S.J. Cronin, N.T. Nehme, S. Limmer, S. Liegeois, J.A. Pospisilik, D. Schramek, A. Leibbrandt, M. Simoes Rde, S. Gruber, U. Puc, I. Ebersberger, T. Zoranovic, G.G. Neely, A. von Haeseler, D. Ferrandon, J.M. Penninger, Science 325 (2009) [72] R. Albertson, C. Chabu, A. Sheehan, C.Q. Doe, J. Cell Sci. 117 (2004) [73] T. Reya, S.J. Morrison, M.F. Clarke, I.L. Weissman, Nature 414 (2001) [74] E.Gateff, Science 200 (1978) [75] E. Caussinus, C. Gonzalez, Nat. Genet. 37 (2005)
22 Figure Legends Fig. 1. (A) Specification of the neuroblasts during development. In the neuroectoderm, a group of cells acquire the potential to become a neuroblast by expressing proneural genes (green). Later, a single cell is selected to become a neuroblast through a Notch signaling mediated process called lateral inhibition (arrows). The specified neuroblast enlarges in size, delaminates basally from the neuroectoderm and then divides asymmetrically along the apical-basal axis. (B) Schematic representation of the Drosophila larval central nervous system, which consists of the central brain (CB), the optic lobes (OL), and the ventral nerve cord (VNC). The CB contains two types of neuroblasts (NBs). While the majorities are type I NBs (red), eight type II NBs are present in each brain hemisphere (blue). Fig. 2. (A) Mode of cell division of type I neuroblasts. The neuroblast (NB, pink) divides to self-renew and to give rise to a smaller ganglion mother cell (GMC, orange), which only divides once to generate two postmitotic neurons or glia cells (red). (B) Mode of cell division of type II neuroblasts. The neuroblast (NB, blue) divides to self-renew and to give rise to an intermediate neural progenitor cell (INP, light blue), which undergoes several rounds of cell division, each of which results in self-renewal of the INP and in the generation of a GMC (purple) that further produces two progeny (dark blue). Fig. 3. Schematic representation of transgenic RNAi in Drosophila. The driver line expresses the yeast protein Gal4, which is under the spatial and temporal control of a specific enhancer. The responder line contains an RNAi construct that is controlled by the UAS. After crossing the
23 two lines, Gal4 binds to UAS and leads to the transcription of the RNAi inverted repeat. The RNA hairpins are further processed through the RNAi pathway to generate functional RNAi molecules, which results in the tissue-specific target gene knock-down. Fig. 4. (A) Simplified workflow of the genome-wide transgenic RNAi screen in Drosophila neuroblasts [12]. The driver line co-expresses Dicer-2 to enhance the transgenic RNAi effect. Schematics show an overproliferative phenotype (left) and a normal brain (right). (B) Summary of the primary screen in [12]. In total, 17,362 lines were screened and 24.1% (4,182 lines) resulted in lethality. In these lethal lines, 832 showed abnormal brain phenotype.
Review Article Neural Stem Cells in Drosophila: Molecular Genetic Mechanisms Underlying Normal Neural Proliferation and Abnormal Brain Tumor Formation
 Stem Cells International Volume 2012, Article ID 486169, 10 pages doi:10.1155/2012/486169 Review Article Neural Stem Cells in Drosophila: Molecular Genetic Mechanisms Underlying Normal Neural Proliferation
Stem Cells International Volume 2012, Article ID 486169, 10 pages doi:10.1155/2012/486169 Review Article Neural Stem Cells in Drosophila: Molecular Genetic Mechanisms Underlying Normal Neural Proliferation
Control of neural stem cell self-renewal and differentiation in Drosophila
 Institutional Repository of the University of Basel University Library Schoenbeinstrasse 18-20 CH-4056 Basel, Switzerland http://edoc.unibas.ch/ Year: 2014 Control of neural stem cell self-renewal and
Institutional Repository of the University of Basel University Library Schoenbeinstrasse 18-20 CH-4056 Basel, Switzerland http://edoc.unibas.ch/ Year: 2014 Control of neural stem cell self-renewal and
The Tumor Suppressors Brat and Numb Regulate Transit-Amplifying Neuroblast Lineages in Drosophila
 Article The Tumor Suppressors Brat and Numb Regulate Transit-Amplifying Neuroblast Lineages in Drosophila Sarah K. Bowman, 1 Vivien Rolland, 1 Joerg Betschinger, 1,3 Kaolin A. Kinsey, 2 Gregory Emery,
Article The Tumor Suppressors Brat and Numb Regulate Transit-Amplifying Neuroblast Lineages in Drosophila Sarah K. Bowman, 1 Vivien Rolland, 1 Joerg Betschinger, 1,3 Kaolin A. Kinsey, 2 Gregory Emery,
Postembryonic Development of Amplifying Neuroblast Lineages in the Drosophila Brain: Proliferation, Differentiation and Projection Patterns.
 Postembryonic Development of Amplifying Neuroblast Lineages in the Drosophila Brain: Proliferation, Differentiation and Projection Patterns. Inauguraldissertation zur Erlangung der Würde eines Doktors
Postembryonic Development of Amplifying Neuroblast Lineages in the Drosophila Brain: Proliferation, Differentiation and Projection Patterns. Inauguraldissertation zur Erlangung der Würde eines Doktors
Brat Is a Miranda Cargo Protein that Promotes Neuronal Differentiation and Inhibits Neuroblast Self-Renewal
 Developmental Cell 10, 441 449, April, 2006 ª2006 Elsevier Inc. DOI 10.1016/j.devcel.2006.01.017 Brat Is a Miranda Cargo Protein that Promotes Neuronal Differentiation and Inhibits Neuroblast Self-Renewal
Developmental Cell 10, 441 449, April, 2006 ª2006 Elsevier Inc. DOI 10.1016/j.devcel.2006.01.017 Brat Is a Miranda Cargo Protein that Promotes Neuronal Differentiation and Inhibits Neuroblast Self-Renewal
Viktorin, G. and Riebli, N. and Popkova, A. and Giangrande, A. and Reichert, H.
 Institutional Repository of the University of Basel University Library Schoenbeinstrasse 18-20 CH-4056 Basel, Switzerland http://edoc.unibas.ch/ Year: 2011 Multipotent neural stem cells generate glial
Institutional Repository of the University of Basel University Library Schoenbeinstrasse 18-20 CH-4056 Basel, Switzerland http://edoc.unibas.ch/ Year: 2011 Multipotent neural stem cells generate glial
Cell Birth and Death. Chapter Three
 Cell Birth and Death Chapter Three Neurogenesis All neurons and glial cells begin in the neural tube Differentiated into neurons rather than ectoderm based on factors we have already discussed If these
Cell Birth and Death Chapter Three Neurogenesis All neurons and glial cells begin in the neural tube Differentiated into neurons rather than ectoderm based on factors we have already discussed If these
Regulation of Self-renewal and Differentiation in the Drosophila Nervous System
 Regulation of Self-renewal and Differentiation in the Drosophila Nervous System T.D. SOUTHALL, B. EGGER, K.S. GOLD, AND A.H. BRAND The Gurdon Institute and Department of hysiology, Development, and Neuroscience,
Regulation of Self-renewal and Differentiation in the Drosophila Nervous System T.D. SOUTHALL, B. EGGER, K.S. GOLD, AND A.H. BRAND The Gurdon Institute and Department of hysiology, Development, and Neuroscience,
Cell Type Nervous System I. Developmental Readout. Foundations. Stem cells. Organ formation. Human issues.
 7.72 10.11.06 Cell Type Nervous System I Human issues Organ formation Stem cells Developmental Readout Axes Cell type Axon guidance 3D structure Analysis Model + + organisms Foundations Principles 1 What
7.72 10.11.06 Cell Type Nervous System I Human issues Organ formation Stem cells Developmental Readout Axes Cell type Axon guidance 3D structure Analysis Model + + organisms Foundations Principles 1 What
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
A Genetic Program for Embryonic Development
 Concept 18.4: A program of differential gene expression leads to the different cell types in a multicellular organism During embryonic development, a fertilized egg gives rise to many different cell types
Concept 18.4: A program of differential gene expression leads to the different cell types in a multicellular organism During embryonic development, a fertilized egg gives rise to many different cell types
Control of proliferation activation in quiescent neuroblasts of the Drosophila central nervous system
 Development 121, 1173-1182 (1995) Printed in Great Britain The Company of Biologists Limited 1995 1173 Control of proliferation activation in quiescent neuroblasts of the Drosophila central nervous system
Development 121, 1173-1182 (1995) Printed in Great Britain The Company of Biologists Limited 1995 1173 Control of proliferation activation in quiescent neuroblasts of the Drosophila central nervous system
glial cells missing and gcm2 Cell-autonomously Regulate Both Glial and Neuronal
 glial cells missing and gcm2 Cell-autonomously Regulate Both Glial and Neuronal Development in the Visual System of Drosophila Carole Chotard, Wendy Leung and Iris Salecker Supplemental Data Supplemental
glial cells missing and gcm2 Cell-autonomously Regulate Both Glial and Neuronal Development in the Visual System of Drosophila Carole Chotard, Wendy Leung and Iris Salecker Supplemental Data Supplemental
LABORATÓRIUMI GYAKORLAT SILLABUSZ SYLLABUS OF A PRACTICAL DEMOSTRATION. financed by the program
 TÁMOP-4.1.1.C-13/1/KONV-2014-0001 projekt Az élettudományi-klinikai felsőoktatás gyakorlatorientált és hallgatóbarát korszerűsítése a vidéki képzőhelyek nemzetközi versenyképességének erősítésére program
TÁMOP-4.1.1.C-13/1/KONV-2014-0001 projekt Az élettudományi-klinikai felsőoktatás gyakorlatorientált és hallgatóbarát korszerűsítése a vidéki képzőhelyek nemzetközi versenyképességének erősítésére program
he micrornas of Caenorhabditis elegans (Lim et al. Genes & Development 2003)
 MicroRNAs: Genomics, Biogenesis, Mechanism, and Function (D. Bartel Cell 2004) he micrornas of Caenorhabditis elegans (Lim et al. Genes & Development 2003) Vertebrate MicroRNA Genes (Lim et al. Science
MicroRNAs: Genomics, Biogenesis, Mechanism, and Function (D. Bartel Cell 2004) he micrornas of Caenorhabditis elegans (Lim et al. Genes & Development 2003) Vertebrate MicroRNA Genes (Lim et al. Science
Phenomena first observed in petunia
 Vectors for RNAi Phenomena first observed in petunia Attempted to overexpress chalone synthase (anthrocyanin pigment gene) in petunia. (trying to darken flower color) Caused the loss of pigment. Bill Douherty
Vectors for RNAi Phenomena first observed in petunia Attempted to overexpress chalone synthase (anthrocyanin pigment gene) in petunia. (trying to darken flower color) Caused the loss of pigment. Bill Douherty
Polarity and Segmentation. Chapter Two
 Polarity and Segmentation Chapter Two Polarization Entire body plan is polarized One end is different than the other Head vs. Tail Anterior vs. Posterior Front vs. Back Ventral vs. Dorsal Majority of neural
Polarity and Segmentation Chapter Two Polarization Entire body plan is polarized One end is different than the other Head vs. Tail Anterior vs. Posterior Front vs. Back Ventral vs. Dorsal Majority of neural
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Neuroepithelial Cells and Neural Differentiation
 Neuroepithelial Cells and Neural Differentiation Neurulation The cells of the neural tube are NEUROEPITHELIAL CELLS Neural crest cells migrate out of neural tube Neuroepithelial cells are embryonic stem
Neuroepithelial Cells and Neural Differentiation Neurulation The cells of the neural tube are NEUROEPITHELIAL CELLS Neural crest cells migrate out of neural tube Neuroepithelial cells are embryonic stem
NIH Public Access Author Manuscript Dev Biol. Author manuscript; available in PMC 2007 April 30.
 NIH Public Access Author Manuscript Published in final edited form as: Dev Biol. 2007 January 1; 301(1): 287 297. Drosophila brain tumor metastases express both neuronal and glial cell type markers Michelle
NIH Public Access Author Manuscript Published in final edited form as: Dev Biol. 2007 January 1; 301(1): 287 297. Drosophila brain tumor metastases express both neuronal and glial cell type markers Michelle
Cancer Stem Cells & Glioblastoma
 Cancer Stem Cells & Glioblastoma JP Hugnot «Brain plasticity, Neural stem cells and Glial tumors» INSERM U1051-UM2 Institut des Neurosciences de Montpellier Montpellier 1-Stem cells and Brain Stem Cells
Cancer Stem Cells & Glioblastoma JP Hugnot «Brain plasticity, Neural stem cells and Glial tumors» INSERM U1051-UM2 Institut des Neurosciences de Montpellier Montpellier 1-Stem cells and Brain Stem Cells
Bidirectional Notch signaling regulates Drosophila intestinal stem cell multipotency
 RESEARCH RESEARCH ARTICLE SUMMARY STEM CELL REGULATION Bidirectional Notch signaling regulates Drosophila intestinal stem cell multipotency Zheng Guo and Benjamin Ohlstein* INTRODUCTION: In the Drosophila
RESEARCH RESEARCH ARTICLE SUMMARY STEM CELL REGULATION Bidirectional Notch signaling regulates Drosophila intestinal stem cell multipotency Zheng Guo and Benjamin Ohlstein* INTRODUCTION: In the Drosophila
Bi 8 Lecture 17. interference. Ellen Rothenberg 1 March 2016
 Bi 8 Lecture 17 REGulation by RNA interference Ellen Rothenberg 1 March 2016 Protein is not the only regulatory molecule affecting gene expression: RNA itself can be negative regulator RNA does not need
Bi 8 Lecture 17 REGulation by RNA interference Ellen Rothenberg 1 March 2016 Protein is not the only regulatory molecule affecting gene expression: RNA itself can be negative regulator RNA does not need
Lineage diversity in the Drosophila nervous system Yohanns Bellaïche* and François Schweisguth
 418 Lineage diversity in the Drosophila nervous system Yohanns Bellaïche* and François Schweisguth The detailed descriptions of cellular lineages in the Drosophila nervous system have provided the foundations
418 Lineage diversity in the Drosophila nervous system Yohanns Bellaïche* and François Schweisguth The detailed descriptions of cellular lineages in the Drosophila nervous system have provided the foundations
scientificreport Synergism between altered cortical polarity and the PI3K/TOR pathway in the suppression of tumour growth scientific report
 scientificreport Synergism between altered cortical polarity and the PI3K/TOR pathway in the suppression of tumour growth Fabrizio Rossi 1 &CayetanoGonzalez 1,2+ 1 Cell Division Group, Institute for Research
scientificreport Synergism between altered cortical polarity and the PI3K/TOR pathway in the suppression of tumour growth Fabrizio Rossi 1 &CayetanoGonzalez 1,2+ 1 Cell Division Group, Institute for Research
Transgenic Expression of the Helicobacter pylori Virulence Factor CagA Promotes Apoptosis or Tumorigenesis through JNK Activation in Drosophila
 Transgenic Expression of the Helicobacter pylori Virulence Factor CagA Promotes Apoptosis or Tumorigenesis through JNK Activation in Drosophila Anica M. Wandler, Karen Guillemin* Institute of Molecular
Transgenic Expression of the Helicobacter pylori Virulence Factor CagA Promotes Apoptosis or Tumorigenesis through JNK Activation in Drosophila Anica M. Wandler, Karen Guillemin* Institute of Molecular
The RNA binding protein Arrest (Bruno) regulates alternative splicing to enable myofibril maturation in Drosophila flight muscle
 Manuscript EMBOR-2014-39791 The RNA binding protein Arrest (Bruno) regulates alternative splicing to enable myofibril maturation in Drosophila flight muscle Maria L. Spletter, Christiane Barz, Assa Yeroslaviz,
Manuscript EMBOR-2014-39791 The RNA binding protein Arrest (Bruno) regulates alternative splicing to enable myofibril maturation in Drosophila flight muscle Maria L. Spletter, Christiane Barz, Assa Yeroslaviz,
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Supplemental Data SUPPLEMENTAL EXPERIMENTAL PROCEDURES
 Temporal transcription factors and their targets schedule the end of neural proliferation in Drosophila. Cédric Maurange, Louise Cheng and Alex P. Gould Supplemental Data SUPPLEMENTAL EXPERIMENTAL PROCEDURES
Temporal transcription factors and their targets schedule the end of neural proliferation in Drosophila. Cédric Maurange, Louise Cheng and Alex P. Gould Supplemental Data SUPPLEMENTAL EXPERIMENTAL PROCEDURES
Extrinsic cues orient the cell division axis in Drosophila embryonic neuroblasts
 RESEARCH ARTICLE 529 Development 133, 529-536 doi:10.1242/dev.02211 Extrinsic cues orient the cell division axis in Drosophila embryonic neuroblasts Sarah E. Siegrist and Chris Q. Doe* Cell polarity must
RESEARCH ARTICLE 529 Development 133, 529-536 doi:10.1242/dev.02211 Extrinsic cues orient the cell division axis in Drosophila embryonic neuroblasts Sarah E. Siegrist and Chris Q. Doe* Cell polarity must
SUPPLEMENTARY INFORMATION
 Supplementary Figure 1. Ras V12 expression in the entire eye-antennal disc does not cause invasive tumours. a, Eye-antennal discs expressing Ras V12 in all cells (marked with GFP, green) overgrow moderately
Supplementary Figure 1. Ras V12 expression in the entire eye-antennal disc does not cause invasive tumours. a, Eye-antennal discs expressing Ras V12 in all cells (marked with GFP, green) overgrow moderately
Developmental Biology
 Developmental iology 356 (2011) 553 565 ontents lists available at ScienceDirect Developmental iology journal homepage: www.elsevier.com/developmentalbiology Multipotent neural stem cells generate glial
Developmental iology 356 (2011) 553 565 ontents lists available at ScienceDirect Developmental iology journal homepage: www.elsevier.com/developmentalbiology Multipotent neural stem cells generate glial
Plasticity of Cerebral Cortex in Development
 Plasticity of Cerebral Cortex in Development Jessica R. Newton and Mriganka Sur Department of Brain & Cognitive Sciences Picower Center for Learning & Memory Massachusetts Institute of Technology Cambridge,
Plasticity of Cerebral Cortex in Development Jessica R. Newton and Mriganka Sur Department of Brain & Cognitive Sciences Picower Center for Learning & Memory Massachusetts Institute of Technology Cambridge,
effects on organ development. a-f, Eye and wing discs with clones of ε j2b10 show no
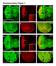 Supplementary Figure 1. Loss of function clones of 14-3-3 or 14-3-3 show no significant effects on organ development. a-f, Eye and wing discs with clones of 14-3-3ε j2b10 show no obvious defects in Elav
Supplementary Figure 1. Loss of function clones of 14-3-3 or 14-3-3 show no significant effects on organ development. a-f, Eye and wing discs with clones of 14-3-3ε j2b10 show no obvious defects in Elav
Role of Notch signaling in establishing the hemilineages of secondary neurons in Drosophila melanogaster
 RESEARCH ARTICLE 53 Development 137, 53-61 (2010) doi:10.1242/dev.041749 Role of Notch signaling in establishing the hemilineages of secondary neurons in Drosophila melanogaster James W. Truman*,, Wanda
RESEARCH ARTICLE 53 Development 137, 53-61 (2010) doi:10.1242/dev.041749 Role of Notch signaling in establishing the hemilineages of secondary neurons in Drosophila melanogaster James W. Truman*,, Wanda
CONTRACTING ORGANIZATION: Cold Spring Harbor Laboratory Cold Spring Harbor, NY 11724
 AD Award Number: W81XWH-10-1-0450 TITLE: The role of NF1 in memory retrieval PRINCIPAL INVESTIGATOR: Yi Zhong, Ph.D. CONTRACTING ORGANIZATION: Cold Spring Harbor Laboratory Cold Spring Harbor, NY 11724
AD Award Number: W81XWH-10-1-0450 TITLE: The role of NF1 in memory retrieval PRINCIPAL INVESTIGATOR: Yi Zhong, Ph.D. CONTRACTING ORGANIZATION: Cold Spring Harbor Laboratory Cold Spring Harbor, NY 11724
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins.
 Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
A. Incorrect! All the cells have the same set of genes. (D)Because different types of cells have different types of transcriptional factors.
 Genetics - Problem Drill 21: Cytogenetics and Chromosomal Mutation No. 1 of 10 1. Why do some cells express one set of genes while other cells express a different set of genes during development? (A) Because
Genetics - Problem Drill 21: Cytogenetics and Chromosomal Mutation No. 1 of 10 1. Why do some cells express one set of genes while other cells express a different set of genes during development? (A) Because
The Biology and Genetics of Cells and Organisms The Biology of Cancer
 The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 Expression of escargot (esg) and genetic approach for achieving IPC-specific knockdown. (a) esg MH766 -Gal4 UAS-cd8GFP (green) and esg-lacz B7-2-22 (red) show similar expression
Supplementary Figure 1 Expression of escargot (esg) and genetic approach for achieving IPC-specific knockdown. (a) esg MH766 -Gal4 UAS-cd8GFP (green) and esg-lacz B7-2-22 (red) show similar expression
Quantification of early stage lesions for loss of p53 should be shown in the main figures.
 Reviewer #1 (Remarks to the Author): Expert in prostate cancer The manuscript "Clonal dynamics following p53 loss of heterozygosity in Kras-driven cancers" uses a number of novel genetically engineered
Reviewer #1 (Remarks to the Author): Expert in prostate cancer The manuscript "Clonal dynamics following p53 loss of heterozygosity in Kras-driven cancers" uses a number of novel genetically engineered
Notch Maintains Drosophila Type II Neuroblasts by Suppressing the. Expression of the Fez Transcription Factor Earmuff
 First posted online on 5 May 2016 as 10.1242/dev.136184 Access the most recent version at http://dev.biologists.org/lookup/doi/10.1242/dev.136184 Notch Maintains Drosophila Type II Neuroblasts by Suppressing
First posted online on 5 May 2016 as 10.1242/dev.136184 Access the most recent version at http://dev.biologists.org/lookup/doi/10.1242/dev.136184 Notch Maintains Drosophila Type II Neuroblasts by Suppressing
RNA interference (RNAi)
 RN interference (RNi) Natasha aplen ene Silencing Section Office of Science and Technology Partnerships Office of the Director enter for ancer Research National ancer Institute ncaplen@mail.nih.gov Plants
RN interference (RNi) Natasha aplen ene Silencing Section Office of Science and Technology Partnerships Office of the Director enter for ancer Research National ancer Institute ncaplen@mail.nih.gov Plants
Chapter 5 A Dose Dependent Screen for Modifiers of Kek5
 Chapter 5 A Dose Dependent Screen for Modifiers of Kek5 "#$ ABSTRACT Modifier screens in Drosophila have proven to be a powerful tool for uncovering gene interaction and elucidating molecular pathways.
Chapter 5 A Dose Dependent Screen for Modifiers of Kek5 "#$ ABSTRACT Modifier screens in Drosophila have proven to be a powerful tool for uncovering gene interaction and elucidating molecular pathways.
Supplementary Figures
 J. Cell Sci. 128: doi:10.1242/jcs.173807: Supplementary Material Supplementary Figures Fig. S1 Fig. S1. Description and/or validation of reagents used. All panels show Drosophila tissues oriented with
J. Cell Sci. 128: doi:10.1242/jcs.173807: Supplementary Material Supplementary Figures Fig. S1 Fig. S1. Description and/or validation of reagents used. All panels show Drosophila tissues oriented with
Early events of neural development
 Early events of neural development Goals: 1) to discuss the origins of cells in the nervous system 2) to discuss how neural stem cells generate diverse cell types in the nervous system The next four lectures
Early events of neural development Goals: 1) to discuss the origins of cells in the nervous system 2) to discuss how neural stem cells generate diverse cell types in the nervous system The next four lectures
CONTRACTING ORGANIZATION: Cold Spring Harbor Laboratory Cold Spring Harbor, NY 11724
 AD Award Number: W81XWH-05-1-0256 TITLE: Dicer in Mammary Tumor Stem Cell Maintenance PRINCIPAL INVESTIGATOR: Elizabeth P. Murchison CONTRACTING ORGANIZATION: Cold Spring Harbor Laboratory Cold Spring
AD Award Number: W81XWH-05-1-0256 TITLE: Dicer in Mammary Tumor Stem Cell Maintenance PRINCIPAL INVESTIGATOR: Elizabeth P. Murchison CONTRACTING ORGANIZATION: Cold Spring Harbor Laboratory Cold Spring
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
Cell polarity in development and cancer
 Rr E Ve I E Wv i e w Cell polarity in development and cancer Andreas Wodarz and Inke Näthke The development of cancer is a multistep process in which the DNA of a single cell accumulates mutations in genes
Rr E Ve I E Wv i e w Cell polarity in development and cancer Andreas Wodarz and Inke Näthke The development of cancer is a multistep process in which the DNA of a single cell accumulates mutations in genes
Histones modifications and variants
 Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Hox genes. Discovered in Drosophila in 1923 by Bridges and Morgan Antennapaedia complex ANT-C Bithorax complex BX-C
 Hox genes Establish body plan during development Specify head to tail axis of animal embryos Head Hox genes, abdomen hox genes. Mutations can cause one body part to transform to another 39 transcription
Hox genes Establish body plan during development Specify head to tail axis of animal embryos Head Hox genes, abdomen hox genes. Mutations can cause one body part to transform to another 39 transcription
Control of neuronal cell fate and number by integration of distinct daughter cell proliferation modes with temporal progression
 678 AND STEM CELLS Development 139, 678-689 (2012) doi:10.1242/dev.074500 2012. Published by The Company of Biologists Ltd Control of neuronal cell fate and number by integration of distinct daughter cell
678 AND STEM CELLS Development 139, 678-689 (2012) doi:10.1242/dev.074500 2012. Published by The Company of Biologists Ltd Control of neuronal cell fate and number by integration of distinct daughter cell
Control of neuronal cell fate and number by integration of distinct daughter cell proliferation modes with temporal progression
 Control of neuronal cell fate and number by integration of distinct daughter cell proliferation modes with temporal progression Carina Ulvklo, Ryan MacDonald, Caroline Bivik, Magnus Baumgardt, Daniel Karlsson
Control of neuronal cell fate and number by integration of distinct daughter cell proliferation modes with temporal progression Carina Ulvklo, Ryan MacDonald, Caroline Bivik, Magnus Baumgardt, Daniel Karlsson
The genetics of heterochromatin. in metazoa. mutations by means of X-ray irradiation" "for the discovery of the production of
 The genetics of heterochromatin in metazoa 1 Hermann Joseph Muller 1946 Nobel Prize in Medicine: "for the discovery of the production of mutations by means of X-ray irradiation" 3 4 The true meaning of
The genetics of heterochromatin in metazoa 1 Hermann Joseph Muller 1946 Nobel Prize in Medicine: "for the discovery of the production of mutations by means of X-ray irradiation" 3 4 The true meaning of
Segmentation. Serial Homology. Hox genes
 Segmentation Hox genes Serial Homology William Bateson 1861-1926 Homeosis: a variation in which something has been changed into the likeness of something else Calvin Bridges at Columbia bithorax 1923 1
Segmentation Hox genes Serial Homology William Bateson 1861-1926 Homeosis: a variation in which something has been changed into the likeness of something else Calvin Bridges at Columbia bithorax 1923 1
MicroRNA in Cancer Karen Dybkær 2013
 MicroRNA in Cancer Karen Dybkær RNA Ribonucleic acid Types -Coding: messenger RNA (mrna) coding for proteins -Non-coding regulating protein formation Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
MicroRNA in Cancer Karen Dybkær RNA Ribonucleic acid Types -Coding: messenger RNA (mrna) coding for proteins -Non-coding regulating protein formation Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Mitosis and the Cell Cycle
 Mitosis and the Cell Cycle Chapter 12 The Cell Cycle: Cell Growth & Cell Division Where it all began You started as a cell smaller than a period at the end of a sentence Getting from there to here Cell
Mitosis and the Cell Cycle Chapter 12 The Cell Cycle: Cell Growth & Cell Division Where it all began You started as a cell smaller than a period at the end of a sentence Getting from there to here Cell
Computational Approaches to Transcriptome Signatures in the Human Brain
 Computational Approaches to Transcriptome Signatures in the Human Brain Agilent Technologies eseminar February 18, 2016 Mike Hawrylycz, Ph.D. ALLEN Adult Human Atlas Online tools An Anatomic Transcriptional
Computational Approaches to Transcriptome Signatures in the Human Brain Agilent Technologies eseminar February 18, 2016 Mike Hawrylycz, Ph.D. ALLEN Adult Human Atlas Online tools An Anatomic Transcriptional
Haematopoietic stem cells
 Haematopoietic stem cells Neil P. Rodrigues, DPhil NIH Centre for Biomedical Research Excellence in Stem Cell Biology Boston University School of Medicine neil.rodrigues@imm.ox.ac.uk Haematopoiesis: An
Haematopoietic stem cells Neil P. Rodrigues, DPhil NIH Centre for Biomedical Research Excellence in Stem Cell Biology Boston University School of Medicine neil.rodrigues@imm.ox.ac.uk Haematopoiesis: An
Why am I working in a Drosophila lab? Dr. Stefan Pulver, Brandeis University
 Why am I working in a Drosophila lab? Dr. Stefan Pulver, Brandeis University !Short life span The advantages of Drosophila as a model organism in neurocience!genetic manipulations are relatively easy!forward
Why am I working in a Drosophila lab? Dr. Stefan Pulver, Brandeis University !Short life span The advantages of Drosophila as a model organism in neurocience!genetic manipulations are relatively easy!forward
CONTRACTING ORGANIZATION: Mount Sinai School of Medicine New York, New York
 AD AWARD NUMBER: W81XWH-05-1-0475 TITLE: Restoration of Epithelial Polarity in Metastatic Tumors PRINCIPAL INVESTIGATOR: Sergei Sokol, Ph.D. CONTRACTING ORGANIZATION: Mount Sinai School of Medicine New
AD AWARD NUMBER: W81XWH-05-1-0475 TITLE: Restoration of Epithelial Polarity in Metastatic Tumors PRINCIPAL INVESTIGATOR: Sergei Sokol, Ph.D. CONTRACTING ORGANIZATION: Mount Sinai School of Medicine New
The ventral nervous system defective gene controls proneural gene expression at two distinct steps during neuroblast formation in Drosophila
 Development 120, 1517-1524 (1994) Printed in Great Britain The Company of Biologists Limited 1994 1517 The ventral nervous system defective gene controls proneural gene expression at two distinct steps
Development 120, 1517-1524 (1994) Printed in Great Britain The Company of Biologists Limited 1994 1517 The ventral nervous system defective gene controls proneural gene expression at two distinct steps
Citation for published version (APA): Martina-Mamber, C. E. (2014). GFAP as an understudy in adult neurogenesis. 's-hertogenbosch: Boxpress.
 UvA-DARE (Digital Academic Repository) GFAP as an understudy in adult neurogenesis Mamber, C.E. Link to publication Citation for published version (APA): Martina-Mamber, C. E. (2014). GFAP as an understudy
UvA-DARE (Digital Academic Repository) GFAP as an understudy in adult neurogenesis Mamber, C.E. Link to publication Citation for published version (APA): Martina-Mamber, C. E. (2014). GFAP as an understudy
Chapter 12 The Cell Cycle
 Chapter 12 The Cell Cycle Objectives Describe how cell reproduction contributes to repair and growth. Compare and contrast prokaryotic and eukaryotic cell division. Compare and contrast asexual and sexual
Chapter 12 The Cell Cycle Objectives Describe how cell reproduction contributes to repair and growth. Compare and contrast prokaryotic and eukaryotic cell division. Compare and contrast asexual and sexual
Introduction to Cancer Biology
 Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Chapter 18 Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences
Chapter 18 Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences
AWARD NUMBER: W81XWH TITLE: Sleep Homeostasis and Synaptic Plasticity. PRINCIPAL INVESTIGATOR: Ravi Allada
 AWARD NUMBER: W81XWH-16-1-0169 TITLE: Sleep Homeostasis and Synaptic Plasticity PRINCIPAL INVESTIGATOR: Ravi Allada CONTRACTING ORGANIZATION: Northwestern University Evanston, IL 60201 REPORT DATE: June
AWARD NUMBER: W81XWH-16-1-0169 TITLE: Sleep Homeostasis and Synaptic Plasticity PRINCIPAL INVESTIGATOR: Ravi Allada CONTRACTING ORGANIZATION: Northwestern University Evanston, IL 60201 REPORT DATE: June
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER
 REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER RNA Splicing Lecture 3, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice
REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER RNA Splicing Lecture 3, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice
Tramtrack controls glial number and identity in the Drosophila embryonic CNS
 Development 128, 4093-4101 (2001) Printed in Great Britain The Company of Biologists Limited 2001 DEV7891 4093 Tramtrack controls glial number and identity in the Drosophila embryonic CNS Paul Badenhorst
Development 128, 4093-4101 (2001) Printed in Great Britain The Company of Biologists Limited 2001 DEV7891 4093 Tramtrack controls glial number and identity in the Drosophila embryonic CNS Paul Badenhorst
Department of Cell and Developmental Biology, Vanderbilt University, Nashville, TN
 UNC-6/Netrin mediates dendritic self-avoidance Cody J. Smith 1, Joseph D. Watson 1,4,5, Miri K. VanHoven 2, Daniel A. Colón-Ramos 3 and David M. Miller III 1,4,6 1 Department of Cell and Developmental
UNC-6/Netrin mediates dendritic self-avoidance Cody J. Smith 1, Joseph D. Watson 1,4,5, Miri K. VanHoven 2, Daniel A. Colón-Ramos 3 and David M. Miller III 1,4,6 1 Department of Cell and Developmental
CONTRACTING ORGANIZATION: Cold Spring Harbor Laboratory Cold Spring Harbor, NY 11724
 AD Award Number: W81XWH-06-1-0249 TITLE: Detection of genes modifying sensitivity to proteasome inhibitors using a shrna Library in Breast Cancer PRINCIPAL INVESTIGATOR: Gregory J. Hannon, Ph.D. CONTRACTING
AD Award Number: W81XWH-06-1-0249 TITLE: Detection of genes modifying sensitivity to proteasome inhibitors using a shrna Library in Breast Cancer PRINCIPAL INVESTIGATOR: Gregory J. Hannon, Ph.D. CONTRACTING
Dr. Tian Xu INTERVIEW. Thomas Ni and Rachel Rosenstein
 YALE JOURNAL OF BIOLOGY AND MEDICINE 79 (2006), pp.193-197. Copyright 2006. INTERVIEW Dr. Tian Xu Thomas Ni and Rachel Rosenstein Yale University School of Medicine Tian Xu is professor and vice chairman
YALE JOURNAL OF BIOLOGY AND MEDICINE 79 (2006), pp.193-197. Copyright 2006. INTERVIEW Dr. Tian Xu Thomas Ni and Rachel Rosenstein Yale University School of Medicine Tian Xu is professor and vice chairman
MicroRNA and Male Infertility: A Potential for Diagnosis
 Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
G-TRACE: rapid Gal4-based cell lineage analysis in Drosophila
 nature methods G-TRACE: rapid Gal4-based cell lineage analysis in Drosophila Cory J Evans, John M Olson, Kathy T Ngo, Eunha Kim, Noemi E Lee, Edward Kuoy, Alexander N Patananan, Daniel Sitz, PhuongThao
nature methods G-TRACE: rapid Gal4-based cell lineage analysis in Drosophila Cory J Evans, John M Olson, Kathy T Ngo, Eunha Kim, Noemi E Lee, Edward Kuoy, Alexander N Patananan, Daniel Sitz, PhuongThao
Chapter 7 Conclusions
 VII-1 Chapter 7 Conclusions VII-2 The development of cell-based therapies ranging from well-established practices such as bone marrow transplant to next-generation strategies such as adoptive T-cell therapy
VII-1 Chapter 7 Conclusions VII-2 The development of cell-based therapies ranging from well-established practices such as bone marrow transplant to next-generation strategies such as adoptive T-cell therapy
Fragile X Syndrome. Genetics, Epigenetics & the Role of Unprogrammed Events in the expression of a Phenotype
 Fragile X Syndrome Genetics, Epigenetics & the Role of Unprogrammed Events in the expression of a Phenotype A loss of function of the FMR-1 gene results in severe learning problems, intellectual disability
Fragile X Syndrome Genetics, Epigenetics & the Role of Unprogrammed Events in the expression of a Phenotype A loss of function of the FMR-1 gene results in severe learning problems, intellectual disability
REGULATED AND NONCANONICAL SPLICING
 REGULATED AND NONCANONICAL SPLICING RNA Processing Lecture 3, Biological Regulatory Mechanisms, Hiten Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice site consensus sequences do have
REGULATED AND NONCANONICAL SPLICING RNA Processing Lecture 3, Biological Regulatory Mechanisms, Hiten Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice site consensus sequences do have
50mM D-Glucose. Percentage PI. L-Glucose
 Monica Dus, Minrong Ai, Greg S.B Suh. Taste-independent nutrient selection is mediated by a brain-specific Na+/solute cotransporter in Drosophila. Control + Phlorizin mm D-Glucose 1 2mM 1 L-Glucose Gr5a;Gr64a
Monica Dus, Minrong Ai, Greg S.B Suh. Taste-independent nutrient selection is mediated by a brain-specific Na+/solute cotransporter in Drosophila. Control + Phlorizin mm D-Glucose 1 2mM 1 L-Glucose Gr5a;Gr64a
Genes implicated in stem-cell identity and temporal-program are directly targeted by Notch in neuroblast tumours
 First posted online on 10 December 2015 as 10.1242/dev.126326 Access the most recent version at http://dev.biologists.org/lookup/doi/10.1242/dev.126326 Genes implicated in stem-cell identity and temporal-program
First posted online on 10 December 2015 as 10.1242/dev.126326 Access the most recent version at http://dev.biologists.org/lookup/doi/10.1242/dev.126326 Genes implicated in stem-cell identity and temporal-program
Cytoskelet Prednáška 6 Mikrotubuly a mitóza
 Cytoskelet Prednáška 6 Mikrotubuly a mitóza Polarity of tubulin polymerization Nuclei Tubulin > C C Preferential addition of tubulin ar (+) ends Tubulin < C C Preferential loss of tubulin ar (+) ends Kinesin
Cytoskelet Prednáška 6 Mikrotubuly a mitóza Polarity of tubulin polymerization Nuclei Tubulin > C C Preferential addition of tubulin ar (+) ends Tubulin < C C Preferential loss of tubulin ar (+) ends Kinesin
REVIEWS. Stem cell fate in cancer growth, progression and therapy resistance
 REVIEWS Stem cell fate in cancer growth, progression and therapy resistance Nikki K. Lytle 1,2,3, Alison G. Barber 1,2,3 and Tannishtha Reya 1,2,3 * Abstract Although we have come a long way in our understanding
REVIEWS Stem cell fate in cancer growth, progression and therapy resistance Nikki K. Lytle 1,2,3, Alison G. Barber 1,2,3 and Tannishtha Reya 1,2,3 * Abstract Although we have come a long way in our understanding
Figure S1. Analysis of endo-sirna targets in different microarray datasets. The
 Supplemental Figures: Figure S1. Analysis of endo-sirna targets in different microarray datasets. The percentage of each array dataset that were predicted endo-sirna targets according to the Ambros dataset
Supplemental Figures: Figure S1. Analysis of endo-sirna targets in different microarray datasets. The percentage of each array dataset that were predicted endo-sirna targets according to the Ambros dataset
Functional studies of Drosophila zinc transporters reveal the mechanism for dietary zinc absorption and regulation
 Qin et al. BMC Biology 2013, 11:101 RESEARCH ARTICLE Open Access Functional studies of Drosophila zinc transporters reveal the mechanism for dietary zinc absorption and regulation Qiuhong Qin, Xiaoxi Wang
Qin et al. BMC Biology 2013, 11:101 RESEARCH ARTICLE Open Access Functional studies of Drosophila zinc transporters reveal the mechanism for dietary zinc absorption and regulation Qiuhong Qin, Xiaoxi Wang
Review. Ageing 2: Cancer! Review: Mutations. Mutations 2/14/11. The Raw Material for Evolution. The Double Edged Sword
 Ageing 2: Cancer! Review: The force of natural selection declines with ageing due to increase in extrinsic mortality (= weakening of natural selection) and reduction in reproduction with age (selection
Ageing 2: Cancer! Review: The force of natural selection declines with ageing due to increase in extrinsic mortality (= weakening of natural selection) and reduction in reproduction with age (selection
Biochemistry of Cancer and Tumor Markers
 Biochemistry of Cancer and Tumor Markers The term cancer applies to a group of diseases in which cells grow abnormally and form a malignant tumor. It is a long term multistage genetic process. The first
Biochemistry of Cancer and Tumor Markers The term cancer applies to a group of diseases in which cells grow abnormally and form a malignant tumor. It is a long term multistage genetic process. The first
Early Embryonic Development
 Early Embryonic Development Maternal effect gene products set the stage by controlling the expression of the first embryonic genes. 1. Transcription factors 2. Receptors 3. Regulatory proteins Maternal
Early Embryonic Development Maternal effect gene products set the stage by controlling the expression of the first embryonic genes. 1. Transcription factors 2. Receptors 3. Regulatory proteins Maternal
Cross species analysis of genomics data. Computational Prediction of mirnas and their targets
 02-716 Cross species analysis of genomics data Computational Prediction of mirnas and their targets Outline Introduction Brief history mirna Biogenesis Why Computational Methods? Computational Methods
02-716 Cross species analysis of genomics data Computational Prediction of mirnas and their targets Outline Introduction Brief history mirna Biogenesis Why Computational Methods? Computational Methods
Ch 8 Practice Questions
 Ch 8 Practice Questions Multiple Choice Identify the choice that best completes the statement or answers the question. 1. What fraction of offspring of the cross Aa Aa is homozygous for the dominant allele?
Ch 8 Practice Questions Multiple Choice Identify the choice that best completes the statement or answers the question. 1. What fraction of offspring of the cross Aa Aa is homozygous for the dominant allele?
Lecture 14 - The cell cycle and cell death
 02.17.10 Lecture 14 - The cell cycle and cell death The cell cycle: cells duplicate their contents and divide The cell cycle may be divided into 4 phases The cell cycle triggers essential processes (DNA
02.17.10 Lecture 14 - The cell cycle and cell death The cell cycle: cells duplicate their contents and divide The cell cycle may be divided into 4 phases The cell cycle triggers essential processes (DNA
Stem Cell Epigenetics
 Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
Programmed Cell Death (apoptosis)
 Programmed Cell Death (apoptosis) Stereotypic death process includes: membrane blebbing nuclear fragmentation chromatin condensation and DNA framentation loss of mitochondrial integrity and release of
Programmed Cell Death (apoptosis) Stereotypic death process includes: membrane blebbing nuclear fragmentation chromatin condensation and DNA framentation loss of mitochondrial integrity and release of
Keywords: Daughter Cells Asexual Reproduction Sexual Reproduction Chromosomes Chromatin Homologous Chromosomes Diploid
 Name: CP Biology Unit 6: Cell Growth and Development Students will be able to: 6.1 Understand and explain the different aspects of the eukaryotic cell cycle. Explain how cell size is related to cell division
Name: CP Biology Unit 6: Cell Growth and Development Students will be able to: 6.1 Understand and explain the different aspects of the eukaryotic cell cycle. Explain how cell size is related to cell division
PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland
 AWARD NUMBER: W81XWH-15-1-0644 TITLE: Targeting Cell Polarity Machinery to Exhaust Breast Cancer Stem Cells PRINCIPAL INVESTIGATOR: Chun-Ju Chang CONTRACTING ORGANIZATION: Purdue University West Lafayette,
AWARD NUMBER: W81XWH-15-1-0644 TITLE: Targeting Cell Polarity Machinery to Exhaust Breast Cancer Stem Cells PRINCIPAL INVESTIGATOR: Chun-Ju Chang CONTRACTING ORGANIZATION: Purdue University West Lafayette,
Identification of novel Ras-cooperating oncogenes in Drosophila melanogaster: A. RhoGEF/Rho-family/JNK pathway is a central driver of tumorigenesis
 Genetics: Published Articles Ahead of Print, published on March 2, 2011 as 10.1534/genetics.111.127910 Identification of novel Ras-cooperating oncogenes in Drosophila melanogaster: A RhoGEF/Rho-family/JNK
Genetics: Published Articles Ahead of Print, published on March 2, 2011 as 10.1534/genetics.111.127910 Identification of novel Ras-cooperating oncogenes in Drosophila melanogaster: A RhoGEF/Rho-family/JNK
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
September 20, Submitted electronically to: Cc: To Whom It May Concern:
 History Study (NOT-HL-12-147), p. 1 September 20, 2012 Re: Request for Information (RFI): Building a National Resource to Study Myelodysplastic Syndromes (MDS) The MDS Cohort Natural History Study (NOT-HL-12-147).
History Study (NOT-HL-12-147), p. 1 September 20, 2012 Re: Request for Information (RFI): Building a National Resource to Study Myelodysplastic Syndromes (MDS) The MDS Cohort Natural History Study (NOT-HL-12-147).
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2419 Figure S1 NiGFP localization in Dl mutant dividing SOPs. a-c) time-lapse analysis of NiGFP (green) in Dl mutant SOPs (H2B-RFP, red; clones were identified by the loss of nlsgfp) showing
DOI: 10.1038/ncb2419 Figure S1 NiGFP localization in Dl mutant dividing SOPs. a-c) time-lapse analysis of NiGFP (green) in Dl mutant SOPs (H2B-RFP, red; clones were identified by the loss of nlsgfp) showing
Nature Neuroscience: doi: /nn Supplementary Figure 1. MADM labeling of thalamic clones.
 Supplementary Figure 1 MADM labeling of thalamic clones. (a) Confocal images of an E12 Nestin-CreERT2;Ai9-tdTomato brain treated with TM at E10 and stained for BLBP (green), a radial glial progenitor-specific
Supplementary Figure 1 MADM labeling of thalamic clones. (a) Confocal images of an E12 Nestin-CreERT2;Ai9-tdTomato brain treated with TM at E10 and stained for BLBP (green), a radial glial progenitor-specific
Huntington s Disease & MARY ET BOYLE, PH.D. DEPARTMENT OF COGNITIVE SCIENCE
 Huntington s Disease & Early Nervous System Development MARY ET BOYLE, PH.D. DEPARTMENT OF COGNITIVE SCIENCE UCSD The cups fell to the floor with a crash. Was this the alarm signal? Or was it forgetting
Huntington s Disease & Early Nervous System Development MARY ET BOYLE, PH.D. DEPARTMENT OF COGNITIVE SCIENCE UCSD The cups fell to the floor with a crash. Was this the alarm signal? Or was it forgetting
