EBV encoded mir-bart15-3p promotes cell apoptosis partially by targeting BRUCE
|
|
|
- Reynold Sutton
- 5 years ago
- Views:
Transcription
1 JVI Accepts, published online ahead of print on 15 May 2013 J. Virol. doi: /jvi Copyright 2013, American Society for Microbiology. All Rights Reserved. 1 EBV encoded mir-bart15-3p promotes cell apoptosis partially by targeting BRUCE 2 3 Hoyun Choi, 1 Hanna Lee, 1 Sae Rom Kim, 2 Yong Song Gho, 2 and Suk Kyeong Lee 1 * Research Institute of Immunobiology, Department of Medical Lifescience, College of Medicine, The Catholic University of Korea, Seoul, Republic of Korea 1 ; Department of Life Sciences, Pohang University of Science and Technology, Pohang, Republic of Korea 2 Running Title: EBV mir-bart15-3p targets BRUCE Keywords: Epstein-Barr Virus; MicroRNAs; Cell Proliferation; Inhibitor of Apoptosis Proteins; Apoptosis; 3' Untranslated Regions; Gastric Carcinoma; mir-bart15-3p; BRUCE; Exosome # Corresponding author: Suk Kyeong Lee, Ph.D. Research Institute of Immunobiology, Catholic University of Korea 505 Banpo-dong, Seocho-gu, Seoul, , Korea Tel: Fax: sukklee@catholic.ac.kr Word count: Abstract: 248; Text: 4389 ABSTRACT
2 25 Epstein-Barr Virus (EBV) generates a variety of viral micrornas (mirnas) by processing the 26 BHRF1 and BamHI A rightward (BART) transcripts. BART mirnas are expressed in all cells latently 27 infected with EBV, but the functions of most BART mirnas remain unknown. The results of a cell proliferation assay revealed that mir-bart15-3p inhibited cell proliferation. Fluorescence-activated cell sorting following staining with annexin V or propidium iodide showed that mir-bart15-3p promoted apoptosis. Furthermore, the inhibitor for mir-bart15-3p increased cell growth and reduced apoptosis in EBV-infected cells. Using bioinformatic analyses, we predicted that mir- BART15-3p may target the anti-apoptotic B-cell lymphoma 2 (BCL2), BCL2L2, DEAD (Asp-Glu-Ala- Asp) box polypeptide 42 (DDX42), and baculovirus inhibitor of apoptosis repeat-containing ubiquitinconjugating enzyme (BRUCE) mrnas. The luciferase reporter assay showed that only the 3 - untranslated region (UTR) of BRUCE was affected by mir-bart15-3p. Two putative seed-matched sites for mir-bart15-3p were evident on the BRUCE 3 -UTR. The results of a mutation study indicated that mir-bart15-3p hybridized only with the first seed-matched site on the BRUCE 3 -UTR. mir-bart15-3p down-regulated the BRUCE protein in EBV-negative cells, while the inhibitor for mir-bart15-3p up-regulated the BRUCE protein in EBV-infected cells without affecting the BRUCE mrna level. mir-bart15-3p was secreted from EBV-infected gastric carcinoma cells, and the level of mir-bart15-3p was 2- to 16-fold higher in exosomes than in the corresponding cells. Our data 42 suggest that mir-bart15-3p can induce apoptosis partially by inhibiting the translation of the 43 apoptosis inhibitor BRUCE. Further study is warranted to understand the role of mir-bart15-3p in 44 the EBV life cycle.
3 45 46
4 47 INTRODUCTION 48 MicroRNAs (mirnas) are small non-coding RNAs approximately nucleotides in length that 49 can modulate gene expression in multiple species. Primary mirna transcripts are processed consecutively by the enzymes Drosha and Dicer. Mature mirnas function as negative gene regulators through complementary sequence pairing to the 3' untranslated region (3'-UTR) of the target gene (1). Epstein Barr Virus (EBV) is a herpesvirus that infects more than 90% of the adult population and which has transforming activity (2). It establishes latent infection in most people and is closely associated with a variety of malignancies, including Burkitt s lymphoma (3), Hodgkin s disease, gastric carcinoma, nasopharyngeal carcinoma, and nasal natural killer/t-cell lymphoma (2). There are three types of latency in EBV infections depending on the expression patterns of the latent proteins (4). EBV-encoded RNAs (EBERs) and BamHI A rightward transcripts (BARTs) are expressed in all three latency types (4, 5). EBV expresses 25 different pre-micrornas (6-8). BamHI fragment H rightward open reading frame 1 (BHRF1) mirnas processed mainly from the long transcripts of the Epstein- Barr virus nuclear antigen (EBNA) are expressed in latency type III, while 22 pre-mirnas generated from the BART transcripts are detected in most EBV-associated tumors and cell lines (8-11). The functions of several EBV BART mirnas have been identified. mir-bart5-5p reduces the 64 expression of p53 up-regulated modulator of apoptosis (PUMA), a pro-apoptotic protein, resulting in 65 increased cell survival (12). mir-bart1-5p, 16-5p, and 17-5p decrease the expression of latent 66 membrane protein 1 (LMP1), which usually triggers cell growth and transformation, but inhibits cell
5 67 growth and potentiates apoptosis when over-expressed (13). mir-bart22-3p targets latent 68 membrane protein 2A (LMP2A) of EBV to contribute to immune evasion, but does not affect cell 69 proliferation and apoptosis (14). mir-bart2-5p down-regulates the EBV DNA polymerase BALF5 to produce persistent EBV latency (15), and the natural killer cell ligand MICB, which enables evasion of the immune response (16). The expression of Dicer, which is associated with mirna biogenesis, is decreased by mir-bart6-5p (17). BART cluster 1 and 2 mirnas inhibit the expression of proapoptotic Bim to reduce apoptosis. However, which specific BART mirna targets Bim is unclear (18). The functions of the majority of the BART mirnas remain unknown. As part of a larger effort to determine the function of each individual BART mirna, a total of 44 BART mirna mimics were prepared and transfected into AGS (gastric adenocarcinoma) cells. Unexpectedly, unlike the majority of the BART mirnas, a few BART mirnas increased cell apoptosis and inhibited cell proliferation. The functional mechanism of mir-bart15-3p, which showed the strongest apoptotic activity among the BART mirnas, was further investigated. MATERIALS AND METHODS Cell lines. AGS is an EBV-negative gastric cancer (GC) cell line, while AGS-EBV is an AGS cell line infected with a recombinant Akata virus. AGS and SNU-719 cells were cultured in RPMI-1640 (Gibco 84 BRL, Grand Island, NY, USA) supplemented with 10% fetal bovine serum (FBS) and antibiotics ( U/ml penicillin and 100 μg/ml streptomycin; Gibco BRL). AGS-EBV was maintained in the culture 86 medium containing 400 μg/ml G418 (Gibco BRL).
6 87 88 Transfection of mirna mimic and LNA-miRNA inhibitor. All the BART mirna mimics and the 89 scrambled control, which was used as a negative control, were purchased from Genolution Pharmaceuticals (Seoul, Korea). The locked nucleic acid (LNA)-miR-BART15-3p inhibitor and the negative control LNA-miRNA inhibitor were purchased from Exiqon (Vedbaek, Denmark). All transfection experiments were performed using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) according to the manufacturer s protocol. Protein or RNA was extracted 48 h after transfection. Cell proliferation assay. Cell proliferation was analyzed using a Cell Counting Kit-8 (CCK-8; Dojindo Molecular Technologies, Tokyo, Japan). AGS cells (1x10 3 cells/well) were seeded in a 96-well plate and transfected with each BART mirna (10 nm) or the scrambled control (10 nm). AGS-EBV cells (1x10 3 cells/well) were plated in a 96-well plate and transfected with LNA-miR-BART15-3p inhibitor (50 nm) or the negative control (50 nm). After 72 h incubation, 10 μl of CCK-8 solution was added to each well. The absorbance was measured after 2 h using a SoftMax apparatus (Molecular Devices, Sunnyvale, CA, USA) at a wavelength of 450 nm. Annexin V staining. Cells were washed with cold phosphate buffered saline (PBS) and resuspended 104 in 500 μl annexin V binding buffer (PE Annexin V apoptosis detection kit; BD Biosciences, San Diego, 105 CA, USA), containing phycoerythrin (PE)-labeled annexin V and 7-amino-actinomycin (7-AAD). 106 Annexin V was used to label cells undergoing apoptosis by detecting phosphatidylserine (PS) on the
7 107 outer plasma membrane, while 7-AAD was used to detect dead cells. After incubating for 10 min at 108 room temperature in an area shielded from light, the specimens were analyzed by fluorescence- 109 activated cell sorting (FACS) using a FACSCalibur apparatus (BD Biosciences), acquiring 10, events. Cells positive for annexin V and negative for 7-AAD were considered to be undergoing early apoptosis. Sub G1 population analysis using propidium iodide (PI) staining. Cells were harvested, washed with PBS, and fixed in 70% ethanol at -20 C overnight. The cells were washed twice with PBS, and then resuspended in PBS containing 10 μg/ml RNase A (Invitrogen) and 50 μg/ml PI (Sigma-Aldrich, St. Louis, MO, USA). The distribution of cells in each phase of the cell cycle was analyzed using a FACScalibur apparatus (BD Biosciences) as described previously (19). Plasmid constructs. The full length 3'-UTR of each putative mir-bart 15-3p target gene was amplified from the genomic DNA of AGS-EBV cells and cloned into the XhoI/NotI sites between the Renilla luciferase coding sequence and the poly(a) site of the psicheck-2 plasmid (Promega, Madison, WI, USA). The primers used for the amplification were baculovirus inhibitor of apoptosis repeat-containing ubiquitin-conjugating enzyme (BRUCE); 5'- 124 CCGCTCGAGTGCATTGATGTGGACTTCATAGA-3' and 5'- 125 ATAAGAATGCGGCCGCAAATGAGCCTGTATGGCAGGT-3', B-cell lymphoma 2 (BCL2); 5'- 126 CCGCTCGAGCCCTGGCCTGAAGAAGAGAC-3' and 5'-
8 127 ATAAGAATGCGGCCGCAGGGACGAGGAAACCTTCAA-3', BCL2L2; 5'- 128 CCGCTCGAGGTTCTCTGTCCCTCCTCCCA-3' and 5'- 129 ATAAGAATGCGGCCGCTGCAGCTCCTCTTGGCTAAA-3', DEAD (Asp-Glu-Ala-Asp) box polypeptide 42 (DDX42); 5'-CCGCTCGAGAGGGGATGTGCTAAAGCGT-3' and 5'- ATAAGAATGCGGCCGCCCCGGTAGTAAAACATTTACTAGA-3'. Mutations to the seed sequences of psic-bruce were introduced using an EZchange site-directed mutagenesis kit (Enzynomics, Daejeon, Korea). The primers used for the amplification were: BRUCEm1; 5'- TGGTATGTTCAACAAATTTGTGTATACAAAG-3' and 5'- AAGTGTCGTTCTCACAATTGAAAAATAAAAG-3', BRUCEm2; 5'- TTTGATAGATTTTATGTTTGGCCATATCTTCATG-3' and 5'- TAGTGTCAAAAGTTGCTGACTTTAAATAGTAGTTG-3'. Luciferase reporter assay. To test the effect of mir-bart15-3p on the expression of putative target genes, HEK293T cells were seeded in a 96-well plate (5x10 3 cells/well). After 24 h, the cells were cotransfected with the psicheck reporter vector containing the 3'-UTR fragment of the putative target gene, and mir-bart15-3p or the mutated mir-bart15-3p (mir-bart15-3pm). To test the effect of an inhibitor of mir-bart15-3p on the luciferase activity of psic-bruce in EBV-infected gastric 144 carcinoma cell lines, AGS-EBV and SNU-719 were seeded in a 96-well plate (5x10 3 cells/well). After h, the cells were co-transfected with the psic-bruce reporter vector and the LNA-miR-BART p inhibitor. Luciferase activities were measured 48 h post-transfection using the Dual-Glo
9 147 luciferase reporter assay system (Promega). Renilla luciferase activity was normalized using firefly 148 luciferase activity for each sample Quantitative reverse transcription-polymerase chain reaction (qrt-pcr). AGS and AGS-EBV cells were harvested and total RNA was extracted using the RNAzol B reagent (Tel-Test, Friendswood, TX, USA) according to the manufacturer's instructions. cdna was synthesized using 1 μg total RNA, oligo(dt) (Ahram Biosystems, Seoul, Korea), and M-MLV reverse transcriptase (Invitrogen). Real-time PCR for the indicated genes was carried out using a SYBR green qpcr kit (Takara, Tokyo, Japan) with a Mx3000P Real-Time PCR System (Stratagene, La Jolla, CA, USA). The sequences of the primers were: BRUCE; 5'-CTTGGTCTGAACACGAAAGACA-3' and 5'- TCCATCCGTACAAGGAAACTGT-3', glyceraldehyde phosphate dehydrogenase (GAPDH); 5'- CATGAGAAGTATGACAACAGCCT-3' and 5'-AGTCCTTCCACGATACCAAAGT-3'. The PCR conditions were 95 C for 10 min, followed by 40 cycles at 95 C for 30 s, 60 C for 30 s, and 72 C for 30 s. For the dissociation curve, reactions were incubated at 95 C for 10 s and ramped up from 60 C to 95 C with a heating rate of 0.1 C/s, and fluorescence was measured continuously. Relative gene expression was calculated according to the comparative Ct method using GAPDH as an internal standard Stem-loop real-time PCR for mirna analysis. Real-time reverse transcription quantitative PCR (qrt- 166 PCR) reagent kits were purchased from Applied Biosystems (Foster City, CA, USA). The kit includes
10 167 TaqMan assays for EBV-miR-BART15-3p and a control assay (RNU6B), the TaqMan microrna 168 reverse transcription kit, and UNG-free TaqMan Universal PCR Master Mix II. All TaqMan assays 169 were performed using a two-step procedure. First, single-stranded cdna synthesis from total RNA was performed using the reverse transcription kit. Briefly, for each 7 μl of reverse transcription (RT) master mixture, 0.15 μl of 100 mm dntps, 1 μl of MultiScribe reverse transcriptase, 1.5 μl of 10 RT buffer, 0.19 μl of RNase inhibitor, and 4.16 μl of nuclease-free water were combined. The 7 μl of RT master mixture was then combined in a fresh tube with 5 μl total RNA (1 ng) and 3 μl RT primer (specific to each TaqMan assay). The RT reactions were then performed in a DNA Engine thermal cycler (Bio-Rad Laboratories, Hercules, CA, USA) programmed to incubate the reactions at 16 C for 30 min, 42 C for 30 min, and 85 C for 5 min. In the second step, TaqMan qrt-pcr assays were carried out using a Mx3000P real-time PCR system (Stratagene). Each qrt-pcr reaction was performed in a 20 μl volume containing 10 μl of TaqMan Universal PCR Master Mix II, 1 μl of 20X TaqMan mirna assay mix, 8 μl of RNase free water, and 1 μl single-stranded cdna product. Three replicates were used for each sample. Relative gene expression was calculated according to the comparative Ct method using RNU6B as an internal standard. Small interfering RNA (sirna)-specific knock-down of BRUCE expression. sirna specific for BRUCE 184 (sibruce) was synthesized by Genolution Pharmaceuticals (Seoul, South Korea). A negative control 185 sirna lacking any known gene product was purchased from Genolution Pharmaceuticals. AGS cells 186 at 30 50% confluency were transfected with 10 nm of the sirna using Lipofectamine 2000
11 187 (Invitrogen) in 60 mm-diameter dishes. The cells were harvested 48 h after transfection to analyze 188 BRUCE expression Western blot. To detect the BRUCE protein, the cell lysate in RIPA buffer (50 μg) was mixed with NuPAGE LDS sample buffer (4 ) and heated at 70 C for 10 min. The samples were electrophoretically separated on precast 3 8% NuPAGE Novex Tris-Acetate gels (Invitrogen), and the separated proteins were transferred to a nitrocellulose membrane (Invitrogen). Rabbit polyclonal antibody against BRUCE (Abcam, Cambridge, UK) was used as the primary antibody at a dilution of 1:1000. After washing, the blots were incubated with horseradish peroxidase-conjugated anti-rabbit secondary antibody (Amersham Biosciences, Piscataway, NJ, USA) at a dilution of 1:1000 for 2 h at room temperature. Protein bands were visualized using an enhanced chemiluminescence detection system (Amersham Biosciences). β-actin antibody (Cell Signaling Technology, Danvers, MA, USA) was used to confirm comparable loading. Exosome isolation and characterization. Cells were cultured for 48 h with RPMI containing 10% FBS. To avoid contamination with the bovine serum exosome, FBS was pre-depleted of exosome by ultracentrifuging at 150,000 g for 16 h at 4. The cell culture medium was collected after 48 h and concentrated by ultracentrifugation using the QuixStand Benchtop System (Amersham) with a kda hollow fiber membrane (Amersham Biosciences). To remove the remaining cell debris, the 205 concentrated culture medium was sequentially centrifuged at 500 g for 5 min and then at 3,000 g for min at 4. The concentrated medium was then ultracentrifuged in a 70Ti rotor (Beckman
12 207 Instruments, Fullerton, CA, USA) at 100,000 g for 2 h at 4 and the pelleted exosome was 208 resuspended with 1 PBS. Exosomes were stored at -70 before RNA extraction or Western blot 209 analysis (20) To characterize the exosomes, proteins of whole cell lysates (50 μg) and exosomes (1 μg) were separated by SDS-PAGE and transferred to PVDF membranes (Thermo Fisher Scientific, Rockford, IL, USA). The blocked membrane was incubated with the indicated antibodies. The immunoreactive bands were visualized using enhanced chemiluminescence substrate (GE Healthcare, Buckinghamshire, UK). Transmission electron microscopy (TEM). The purified exosomes were applied to glow-discharged carbon-coated copper grids (EMS, Matfield, PA, USA). After allowing the exosomes to be absorbed onto the grid for 3 min, the grids were rinsed with droplets of de-ionized water and negative-stained with 2% uranylacetate (Ted Pella, Redding, CA, USA). Electron micrographs were recorded with a JEM 1011 microscope (Jeol, Tokyo, Japan) at an acceleration voltage of 100 kv (21). Statistical analyses. The data were analyzed using one-way repeated-measure analysis of variance (ANOVA) or Student s t test. Curve fit and analysis were performed using GraphPad Prism 224 (GraphPad Software, San Diego, CA, USA). P-values < 0.05 were considered statistically significant. 225 All results are expressed as the mean ± standard deviation (SD). 226
13 227 RESULTS 228 BART mirnas affected cell proliferation in AGS cells. In order to investigate the effects of BART 229 mirnas on cell growth, we purchased all BART mirna mimics (a total of 44 mimics). Figure 1A shows the sequence of the mir-bart15-3p mimic. AGS cells were transfected with each of the BART mirnas (10 nm) and cell proliferation was accessed 72 h after transfection using the CCK-8 kit (Figure 1B). The majority of BART mirnas enhanced cell proliferation while five BART mirnas (mir- BART15-3p, 5-5p, 16-5p, 17-3p, and 20-3p) reduced cell growth. Among the five mirnas, mir- BART15-3p suppressed cell proliferation most strongly and was further studied. mir-bart15-3p inhibits cell proliferation and induces apoptosis in AGS cells. In order to investigate the effects of mir-bart15-3p on cell growth, AGS cells were transfected with 10 nm mir- BART15-3p. Cell growth was analyzed using the cell proliferation assay at 6, 12, 24, 48, and 72 h after transfection. Cell proliferation of AGS cells transfected with mir-bart15-3p decreased significantly after 48~72 h compared to that of cells transfected with the scrambled control (Figure 2A). When AGS cells were transfected with increasing concentrations of mir-bart15-3p, the cell proliferation measured after 72 h was significantly lower than that in control cells at concentrations over 3 nm. Cell growth was almost completely inhibited by mir-bart15-3p at concentrations higher 244 than 10 nm (Figure 2B). 245 mir-bart15-3p was transfected into AGS cells in order to analyze apoptotic activity. After 72 h, 246 the ratio of the sub-g1 population in AGS cells transfected with mir-bart15-3p was ± 0.34%,
14 247 while that in AGS cells transfected with the scrambled control was 2.57 ± 0.05%. (Figure 2C). Similar 248 results were obtained in two more independent experiments; the mean ± SD values of all three 249 independent experiments are shown in Figure 2D. The effect of mir-bart15-3p on cell apoptosis was further analyzed using a PE-Annexin V apoptosis detection kit. AGS cells transfected with mir- BART15-3p showed an increased early apoptotic cell population compared to cells transfected with the scrambled control (Figure 2E). Similar results were obtained in two more independent experiments; the mean ± SD values of all three independent experiments are shown in Figure 2F. mir-bart15-3p inhibitor increases cell proliferation and inhibits apoptosis in EBV-infected cells. To check the effect of the LNA inhibitor on the mir-bart15-3p expression level, qrt-pcr was carried out in AGS-EBV cells transfected with LNA-miR-BART15-3p inhibitor. The inhibitor reduced the endogenous level of mir-bart15-3p to 9.1% of the level in control LNA-transfected cells (Figure 3A). As expected, mir-bart15-3p expression was not detected in the EBV-negative cell line, AGS (Figure 3A). AGS cells transfected with mir-bart15-3p mimic showed 7-fold higher levels of mir- BART15-3p than the endogenous level found in AGS-EBV cells. Cell proliferation measured 72 h after inhibitor transfection was higher than in control LNA transfected cells (Figure 3B). To analyze the effects of reduced mir-bart15-3p function on cell apoptosis, the inhibitor was transfected into AGS- 264 EBV cells. After 72 h, the proportion of sub-g1 cells was measured by FACS analysis following PI 265 staining. The proportion of the sub G1 population was lower in AGS-EBV cells transfected with the
15 266 inhibitor than in cells transfected with the control LNA (Figure 3C). Figure 3D shows the summarized 267 results of three independent PI staining/facs experiments mir-bart15-3p directly targets the BRUCE 3'-UTR. We used three publicly available programs to predict putative targets of mir-bart15-3p: TargetScan ( DIANA-microT ( and RNA hybrid ( Among the many predicted targets, BCL2, BCL2L2, DDX42, and BRUCE were chosen for further analysis based on their gene functions and the free energy provided by RNA hybrid program. BRUCE is a member of the inhibitor of apoptosis (IAP) family containing an IAP repeat region at its N-terminal domain (22). The 3'-UTR of the four putative target genes for mir- BART15-3p were individually cloned into the psicheck-2 plasmid (Figure 4A). The resulting 3'-UTR reporter vectors were then individually co-transfected with mir-bart15-3p into HEK293T cells. The luciferase activity of the reporter vector containing the 3'-UTR of BRUCE (psic-bruce) was inhibited by mir-bart15-3p, whereas the activities of the other vectors were not. However, the luciferase activity of the psic-bruce vector was not affected by mutated mir-bart15-3p (mir-bart15-3pm) or by the scrambled control (Figure 4B). There were two putative seed-matched regions for mir- BART15-3p in the BRUCE 3'-UTR. Site-directed mutagenesis was performed to mutate one (psic- 283 BRUCEm1, psic-brucem2) or both of these sites (psic-brucem1m2) (Figure 4C). These plasmids 284 were co-transfected into HEK293T cells with mir-bart15-3p, and a luciferase assay was carried out. 285 The luciferase activity of cells transfected with either psic-brucem1 or psic-brucem1m2 was not
16 286 altered by mir-bart15-3p, whereas that of psic-brucem2-transfected cells was reduced by mir- 287 BART15-3p (Figure 4D). As expected, the luciferase activity of the wild-type or mutated BRUCE UTR reporter vector was not affected by mir-bart15-3pm or the scrambled control To investigate the regulation of luciferase activity of psic-bruce by endogenously expressed mir-bart15-3p, EBV-infected cells were co-transfected with the reporters and LNA-miR-BART15-3p inhibitor (Figure 5). The luciferase activity of psic-bruce was inhibited when it was introduced into EBV-infected cells. In addition, co-transfection of LNA-miR-BART15-3p inhibitor with psic-bruce resulted in recovery of luciferase activity to the control level (Figure 5). Interestingly, real-time RT-PCR analysis revealed that BRUCE mrna expression was not reduced by mir-bart15-3p (Figure 6A). However, Western blot analysis showed that the expression of the BRUCE protein in AGS cells was impaired following mir-bart15-3p transfection (Figures 6B, 6C, and 6D). To confirm these results, we transfected AGS-EBV cells with the LNA-miR-BART15-3p inhibitor, and measured the expression levels of BRUCE mrna and protein after 48 h. The level of BRUCE mrna was not altered (Figure 7A), but an increase in the BRUCE protein level was observed in AGS-EBV cells (Figures 7B, 7C, and 7D) following inhibitor transfection. Similar Western blot results were obtained in SNU-719 cells (Figure 7E and 7F). 303 Knockdown of BRUCE induces cell death in AGS. We examined whether sirna against BRUCE 304 also causes apoptosis similar to mir-bart15-3p. AGS cells were transfected with 10 nm sibruce, 305 and the fraction of the sub-g1 population was analyzed 72 h later by PI staining. sibruce reduced
17 306 the mrna level and protein level of BRUCE as expected (Figures 8A and 8B). sibruce-transfected 307 AGS cells showed an increased sub-g1 population (Figure 8C). However, the fraction of the sub-g1 308 population was higher in AGS cells transfected with mir-bart15-3p than with sibruce mir-bart15-3p is present in exosomes from EBV-infected gastric cell lines. In order to check the possibility that mir-bart15-3p is secreted from the cells, exosomes were isolated from the culture media of the AGS and AGS-EBV cell lines by the ultracentrifugation method. TEM analysis of exosome preparations from both cells showed circular and cup-like vesicles with sizes ranging between 30~50 nm (Figure 9A). The expression of the exosome markers CD8 and CD81 was analyzed by Western blot. CD8 and CD81 were detected in the exosome preparations, while the cellular marker cytochrome C was detected only in the whole cell lysates (Figure 9B). To test whether mir-bart15-3p is secreted via exosomes, quantitative real-time RT-PCR for mir-bart15-3p was carried out using the same amount of RNAs purified from the exosomes and cell pellets. A two-fold higher level of mir-bart15-3p was detected in the exosomal RNA than in the cellular RNA of AGS- EBV. mir-bart1-3p, which was used as a exosomally secreted mirna control (23, 24), was enriched 12-fold in the exosomal RNA. In the case of SNU-719, mir-bart 15-3p was enriched 16- fold in the exosomal RNA; and mir-bart1-3p was enriched in exosomal RNA compared to cellular 323 RNA (Figure 9C). mir-bart5-5p was not significantly enriched in the exosome (Figure 9C), as 324 previously reported (23, 24). 325
18 326 DISCUSSION 327 In the process of screening the effect of BART mirnas on cell growth, we found that a few BART 328 mirnas inhibited cell growth, while majority of the BART mirnas enhanced cell proliferation. Among the BART-miRNAs that inhibited cell growth, mir-bart15-3p showed the most potent effect. In addition, an inhibitor of mir-bart15-3p promoted cell proliferation and inhibited cell death in EBVinfected cells. We also found that mir-bart15-3p mimic repressed the expression of the reporter fused to the 3 -UTR of BRUCE in HEK293T cells. The 3 -UTR of BRUCE has two potential seed matches for mir- BART15-3p. However, mir-bart15-3p seemed to directly target only the first predicted site (NM_016252: ) of BRUCE 3 -UTR. The mir-bart15-3p binding sites are well conserved among mammals (data not shown), even though EBV is a human herpesvirus. The luciferase activity of psic-bruce was reduced in AGS-EBV cells endogenously expressing mir- BART15-3p, while the reduction in psic-bruce activity was abrogated by the inhibitor for mir- BART15-3p. Thus, the endogenous expression of mir-bart15-3p in the EBV-infected gastric carcinoma cell lines AGS-EBV and SNU-719 seems to suppress BRUCE expression. Interestingly, we observed that mir-bart15-3p down-regulated the level of BRUCE protein without affecting the level of BRUCE mrna. This agrees with the previous finding that some mirnas 343 suppress the translation of mrna without affecting their stability (25). 344 In this study, custom predictions of TargetScan and DIANA-microT were used to predict targets for 345 mir-bart15-3p. Four anti-apoptotic genes (BCL2, BCL2L2, DDX42, and BRUCE) were chosen as
19 346 putative mir-bart15-3p targets for further analysis. However, these four genes were not predicted 347 targets of mir-bart15-3p based on the high-throughput sequencing (HITS) cross-linking 348 immunoprecipitation (CLIP) and photoactivatable-ribonucleoside-enhanced (PAR)-CLIP data of other researchers (26-28). The free energy of BRUCE was noticeably the lowest among the four chosen potential target genes. BRUCE, also known as APOLLON or BIRC6, is a member of the inhibitor of apoptosis (IAP) family containing an IAP repeat region at its N-terminal domain (22). BRUCE has a single BIR domain, which protects cells from apoptosis by inhibiting the activities of caspase and proapoptotic elements through pairing and ubiquitin-dependent degradation (29). Similar to previous observations that BRUCE-knockout cells can be sensitized to cell death and easily induced to undergo cell death (30), the effect of mir-bart15-3p on cell apoptosis through BRUCE was confirmed in the present study using BRUCE sirna in AGS cells. Considering the dramatic effect of BRUCE sirna on BRUCE expression, its effect on cell death was not very potent compared to mir-bart15-3p. This might be due to the fact that mirnas can have multiple targets. Other apoptosis-related target genes of mir- BART15-3p are now under investigation in our laboratory. The expression level of mir-bart15-3p detected in EBV-infected NPC cells using qrt-pcr and deep sequencing has been reported to be intermediate compared with those of other BART mirnas 363 (31, 32). In EBV-infected natural killer/t cell lymphomas and lymphoblastoid cell lines (LCLs), mir- 364 BART15-3p can be detected at low frequencies by deep sequencing (27, 33). In contrast, mir-bart p is not detected in EBV-infected diffuse large B-cell lymphomas, germinal center B cells, and
20 366 memory B cells (34, 35). In general, the expression level of mir-bart15-3p seems to be lower in 367 immune cells than in epithelial cells. 368 The effect of mir-bart15-3p on cell apoptosis was rather unexpected as EBV is a tumorigenic virus and some of its mirnas have been reported to increase cell proliferation and inhibit apoptosis (12, 13, 18, 26). It is unclear why mir-bart15-3p would cause cell apoptosis and inhibit cell proliferation. mir-bart15-3p may accelerate the virus lytic cycle and/or spreading of progeny viruses by inducing host cell apoptosis, as caspase cleavage of some viral proteins was shown to favor viral replication and spreading (36). For example, Aleutian mink disease virus promotes caspase activation during permissive infection and cleaves the NS1 protein to facilitate entry into the viral lytic cycle (37, 38). In addition, human papillomavirus (HPV) induces caspase 3- dependent apoptosis, and caspase 3 cleaves the HPV E1 protein enhancing viral amplification (36, 39). The effects observed following transfection with the mir-bart15-3p mimic should be evaluated with care, because the level of transfected mir-bart15-3p was 7-fold higher than its physiological level detected in AGS-EBV cells. However, based on the fact that inhibition of mir-bart15-3p enhanced cell survival and reduced the sub-g1 population in AGS-EBV cells (Fig 3), mir-bart15-3p seems to induce cell apoptosis at physiological conditions. EBV-infected cells would be able to withstand the detrimental effect of mir-bart15-3p as there are many BART mirnas supportive for 383 cell proliferation (Fig 1B). Thus, BART mirna expression following EBV infection would potentiate 384 cell survival rather than induce cell apoptosis in total.
21 385 Recently, several mirnas are known to be secreted selectively via microvesicles, especially 386 exosomes (40). Furthermore, secreted mirnas can be transported to adjacent cells and function in 387 repressing target mrnas (40). This suggests that mirnas could be used in communication between cells. EBV viral mirnas are also secreted and transported to adjacent cells via exosomes (23, 40, 41). In EBV-positive nasopharyngeal cancer, several viral mirnas including mir-bart4, 1, 7, 16, 9, 12, and 13 were enriched in the exosomes and transported to adjacent endothelial cells (23). EBVtransformed lymphoblastoid B cells also secreted several viral mirnas including mir-bhrf1-1, 1-2, mir-bart1-3p, 1-5p, and 2-5p (24). While we were preparing this manuscript, NACHT, LRR and PYD domains-containing protein 3 (NLRP3) was identified as a target of mir-bart15-3p. The authors reported that mir-bart15-3p can be secreted from EBV-infected B cells via exosomes, and a small but significant amount of EBV mir-bart15 was taken up by non-infected cells. Furthermore, the delivered mir-bart15-3p was able to inhibit the NLRP3 inflammasome in the PMA-differentiated macrophage cell line Thp-1 (41). In this study, we observed that mir-bart15-3p increases apoptosis partially by inhibiting translation of BRUCE mrna without affecting its stability. We also found that mir-bart15-3p exists in exosomes from an EBV-positive gastric cancer cell line, and that this mirna is enriched in the exosomes compared to the corresponding cell pellets. Because mir-bart15-3p inhibited BRUCE 402 expression, its transfer to adjacent immune cells would induce their apoptosis. Our data suggest the 403 possibility that exosomal mir-bart15-3p can provide favorable microenvironment for the growth of
22 404 EBV-associated tumors by transmitted to neighboring immune cells. Further investigation of the 405 communication of EBV-positive gastric cancer cells with adjacent immune cells is warranted ACKNOWLEDGEMENTS This research was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education, Science and Technology (2012R1A1A ) and by grants from the Gyeonggi Regional Research Center (GRRC) of the Catholic University of Korea [(GRRC Catholic 2012 B05), RNA-based development of biopharmaceutical lead molecules]. Downloaded from on January 1, 2019 by guest
23 413 REFERENCES Bartel DP MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116: Young LS, Rickinson AB Epstein-Barr virus: 40 years on. Nat Rev Cancer 4: Young LS, Murray PG Epstein-Barr virus and oncogenesis: from latent genes to tumours. Oncogene 22: Middeldorp JM, Brink AA, van den Brule AJ, Meijer CJ Pathogenic roles for Epstein- Barr virus (EBV) gene products in EBV-associated proliferative disorders. Crit Rev Oncol Hematol 45: van Beek J, Brink AA, Vervoort MB, van Zijp MJ, Meijer CJ, van den Brule AJ, Middeldorp JM In vivo transcription of the Epstein-Barr virus (EBV) BamHI-A region without associated in vivo BARF0 protein expression in multiple EBV-associated disorders. J Gen Virol 84: Pfeffer S, Zavolan M, Grasser FA, Chien M, Russo JJ, Ju J, John B, Enright AJ, Marks D, Sander C, Tuschl T Identification of virus-encoded micrornas. Science 304: Grundhoff A, Sullivan CS, Ganem D A combined computational and microarray-based approach identifies novel micrornas encoded by human gamma-herpesviruses. RNA 12: Cai X, Schafer A, Lu S, Bilello JP, Desrosiers RC, Edwards R, Raab-Traub N, Cullen BR Epstein-Barr virus micrornas are evolutionarily conserved and differentially expressed. PLoS Pathog 2:e Zhu JY, Pfuhl T, Motsch N, Barth S, Nicholls J, Grasser F, Meister G Identification of novel Epstein-Barr virus microrna genes from nasopharyngeal carcinomas. J Virol 83: Kim do N, Chae HS, Oh ST, Kang JH, Park CH, Park WS, Takada K, Lee JM, Lee WK, Lee SK Expression of viral micrornas in Epstein-Barr virus-associated gastric carcinoma. J Virol 81: Jun SM, Hong YS, Seo JS, Ko YH, Yang CW, Lee SK Viral microrna profile in Epstein-Barr virus-associated peripheral T cell lymphoma. Br J Haematol 142: Choy EY, Siu KL, Kok KH, Lung RW, Tsang CM, To KF, Kwong DL, Tsao SW, Jin DY An Epstein-Barr virus-encoded microrna targets PUMA to promote host cell survival. J Exp Med 205: Lo AK, To KF, Lo KW, Lung RW, Hui JW, Liao G, Hayward SD Modulation of LMP1 protein expression by EBV-encoded micrornas. Proc Natl Acad Sci U S A 104: Lung RW, Tong JH, Sung YM, Leung PS, Ng DC, Chau SL, Chan AW, Ng EK, Lo KW, To KF Modulation of LMP2A expression by a newly identified Epstein-Barr virus-encoded
24 microrna mir-bart22. Neoplasia 11: Barth S, Pfuhl T, Mamiani A, Ehses C, Roemer K, Kremmer E, Jaker C, Hock J, Meister G, Grasser FA Epstein-Barr virus-encoded microrna mir-bart2 down-regulates the viral DNA polymerase BALF5. Nucleic Acids Res 36: Nachmani D, Stern-Ginossar N, Sarid R, Mandelboim O Diverse herpesvirus micrornas target the stress-induced immune ligand MICB to escape recognition by natural killer cells. Cell Host Microbe 5: Iizasa H, Wulff BE, Alla NR, Maragkakis M, Megraw M, Hatzigeorgiou A, Iwakiri D, Takada K, Wiedmer A, Showe L, Lieberman P, Nishikura K Editing of Epstein-Barr virus-encoded BART6 micrornas controls their dicer targeting and consequently affects viral latency. J Biol Chem 285: Marquitz AR, Mathur A, Nam CS, Raab-Traub N The Epstein-Barr Virus BART micrornas target the pro-apoptotic protein Bim. Virology 412: Shin HJ, Kim do N, Lee SK Association between Epstein-Barr virus infection and chemoresistance to docetaxel in gastric carcinoma. Mol Cells 32: Kim CW, Lee HM, Lee TH, Kang C, Kleinman HK, Gho YS Extracellular membrane vesicles from tumor cells promote angiogenesis via sphingomyelin. Cancer Res 62: Choi DS, Park JO, Jang SC, Yoon YJ, Jung JW, Choi DY, Kim JW, Kang JS, Park J, Hwang D, Lee KH, Park SH, Kim YK, Desiderio DM, Kim KP, Gho YS Proteomic analysis of microvesicles derived from human colorectal cancer ascites. Proteomics 11: Chen Z, Naito M, Hori S, Mashima T, Yamori T, Tsuruo T A human IAP-family gene, apollon, expressed in human brain cancer cells. Biochem Biophys Res Commun 264: Pegtel DM, Cosmopoulos K, Thorley-Lawson DA, van Eijndhoven MA, Hopmans ES, Lindenberg JL, de Gruijl TD, Wurdinger T, Middeldorp JM Functional delivery of viral mirnas via exosomes. Proc Natl Acad Sci U S A 107: Meckes DG, Jr., Shair KH, Marquitz AR, Kung CP, Edwards RH, Raab-Traub N Human tumor virus utilizes exosomes for intercellular communication. Proc Natl Acad Sci U S A 107: Bartel DP MicroRNAs: target recognition and regulatory functions. Cell 136: Riley KJ, Rabinowitz GS, Yario TA, Luna JM, Darnell RB, Steitz JA EBV and human micrornas co-target oncogenic and apoptotic viral and human genes during latency. EMBO J 31: Skalsky RL, Corcoran DL, Gottwein E, Frank CL, Kang D, Hafner M, Nusbaum JD, Feederle R, Delecluse HJ, Luftig MA, Tuschl T, Ohler U, Cullen BR The viral and cellular microrna targetome in lymphoblastoid cell lines. PLoS Pathog 8:e Gottwein E, Corcoran DL, Mukherjee N, Skalsky RL, Hafner M, Nusbaum JD, Shamulailatpam P, Love CL, Dave SS, Tuschl T, Ohler U, Cullen BR Viral microrna targetome of KSHV-infected primary effusion lymphoma cell lines. Cell Host Microbe 10:515-
25 Verhagen AM, Coulson EJ, Vaux DL Inhibitor of apoptosis proteins and their relatives: IAPs and other BIRPs. Genome Biol 2:REVIEWS Hao Y, Sekine K, Kawabata A, Nakamura H, Ishioka T, Ohata H, Katayama R, Hashimoto C, Zhang X, Noda T, Tsuruo T, Naito M Apollon ubiquitinates SMAC and caspase-9, and has an essential cytoprotection function. Nat Cell Biol 6: Cosmopoulos K, Pegtel M, Hawkins J, Moffett H, Novina C, Middeldorp J, Thorley-Lawson DA Comprehensive profiling of Epstein-Barr virus micrornas in nasopharyngeal carcinoma. J Virol 83: Chen SJ, Chen GH, Chen YH, Liu CY, Chang KP, Chang YS, Chen HC Characterization of Epstein-Barr virus mirnaome in nasopharyngeal carcinoma by deep sequencing. PLoS One Motsch N, Alles J, Imig J, Zhu J, Barth S, Reineke T, Tinguely M, Cogliatti S, Dueck A, Meister G, Renner C, Grasser FA MicroRNA Profiling of Epstein-Barr Virus-Associated NK/T-Cell Lymphomas by Deep Sequencing. PLoS One 7:e Imig J, Motsch N, Zhu JY, Barth S, Okoniewski M, Reineke T, Tinguely M, Faggioni A, Trivedi P, Meister G, Renner C, Grasser FA microrna profiling in Epstein-Barr virusassociated B-cell lymphoma. Nucleic Acids Res 39: Qiu J, Cosmopoulos K, Pegtel M, Hopmans E, Murray P, Middeldorp J, Shapiro M, Thorley- Lawson DA A novel persistence associated EBV mirna expression profile is disrupted in neoplasia. PLoS Pathog 7:e Richard A, Tulasne D Caspase cleavage of viral proteins, another way for viruses to make the best of apoptosis. Cell Death Dis 3:e Best SM, Wolfinbarger JB, Bloom ME Caspase activation is required for permissive replication of Aleutian mink disease parvovirus in vitro. Virology 292: Best SM, Shelton JF, Pompey JM, Wolfinbarger JB, Bloom ME Caspase cleavage of the nonstructural protein NS1 mediates replication of Aleutian mink disease parvovirus. J Virol 77: Moody CA, Laimins LA Human papillomaviruses activate the ATM DNA damage pathway for viral genome amplification upon differentiation. PLoS Pathog 5:e Valadi H, Ekstrom K, Bossios A, Sjostrand M, Lee JJ, Lotvall JO Exosome-mediated transfer of mrnas and micrornas is a novel mechanism of genetic exchange between cells. Nat Cell Biol 9: Haneklaus M, Gerlic M, Kurowska-Stolarska M, Rainey AA, Pich D, McInnes IB, Hammerschmidt W, O'Neill LA, Masters SL Cutting edge: mir-223 and EBV mir- BART15 regulate the NLRP3 inflammasome and IL-1beta production. J Immunol 189:
26 528
27 529 Figure legends 530 FIG 1 Effects of each individual BART mirna on cell growth in AGS. Cells were transfected with each 531 individual BART mirna or the scrambled control. (A) The mir-bart15-3p mimic, which has a 2-u overhang at the 3' end of both strands, is shown. All the other BART mirna mimics were designed in a similar pattern. (B) The degree of cell proliferation was analyzed using the CCK-8 assay kit after 72 h (n = 9). Error bars indicate SD. * P < 0.05; P < FIG 2 Effect of mir-bart15-3p on cell proliferation and apoptosis. (A) The sequence of the LNA inhibitor for mir-bart15-3p is shown in the upper panel. Cells were seeded in a 96-well plate and then transfected with the mir-bart15-3p mimic or the scrambled control. After the indicated periods, 10 μl of CCK-8 solution was added to each well to check cell proliferation. (B) Cells were transfected with the indicated concentrations of the mir-bart15-3p mimic or the scrambled control. After 72 h, 10 μl of CCK-8 solution was added to each well. (C) Cells were transfected with the mir-bart15-3p mimic or the scrambled control and analyzed after 72 h. The proportion of the sub-g1 population was evaluated following propidium iodide staining. (D) Similar results to those in (C) were obtained in two more independent experiments, and the mean ± SD values of all three independent experiments are plotted. (E) Cells were transfected with the mir-bart15-3p mimic or the scrambled control and 546 analyzed after 72 h. The proportions of apoptotic cells were accessed by PE-Annexin V staining. (F) 547 Similar results to those in (E) were obtained in two more independent experiments, and the mean ±
28 548 SD values of all three independent experiments are plotted. Error bars indicate SD (n = 3). * P < 0.05; 549 P < FIG 3 Effect of the inhibitor for mir-bart15-3p on AGS-EBV cells. (A) The sequence of the LNA inhibitor for mir-bart15-3p is shown in upper panel. The endogenous expression of mir-bart15-3p in AGS-EBV cells was analyzed by the TaqMan mirna assay 72 h after transfection with the LNAmiR-BART15-3p inhibitor (Inhibitor) or the LNA control (Control). (B) To determine the effect of the mir-bart15-3p inhibitor on cell proliferation, AGS-EBV cells in a 96-well plate were transfected with the inhibitor or the control LNA. After 72 h, 10 μl of CCK-8 solution was added to each well. (C) AGS- EBV cells were transfected with the inhibitor or the control LNA. The proportion of the sub-g1 population was evaluated 72 h later by PI staining. (D) Similar results to those in (C) were obtained in two more independent experiments, and the mean ± SD values of all three independent experiments are plotted. P < FIG 4 Target search for mir-bart15-3p. (A) Seed-matched region of the putative target genes for mir-bart15-3p are shown. (B) To test whether the putative target genes are regulated by mir- BART15-3p, the 3'-UTR fragment of each gene was cloned into a luciferase reporter (psicheck-2) 565 vector. HEK293T cells were co-transfected with the mir-bart15-3p mimic and the appropriate UTR reporter plasmid. To confirm the sequence-specific function of mir-bart15-3p, mir-bart pm (the sequence is shown at the bottom of Figure A), which has substituted nucleotides at 4~6
29 568 sites of mir-bart15-3p, was also used. Luciferase activity was analyzed 48 h after transfection (n = 569 3). (C) An illustration showing the location of the possible seed-match sites for mir-bart15-3p on 570 the BRUCE 3'-UTR and the changed sites to produce mutated forms of psic-bruce. (D) HEK293T cells were co-transfected with mir-bart15-3p and the indicated psic-bruce vector containing the wild-type or mutated 3'-UTR of BRUCE. Luciferase activity was normalized using firefly luciferase activity. Error bars indicate SD (n = 3). P < FIG 5 Effect of Inhibitor for mir-bart15-3p on the luciferase activity of the psic-bruce reporter vector co-transfected into EBV-infected cells. AGS-EBV cells were co-transfected with the inhibitor or the control LNA, and the psic-bruce vector containing the wild-type or mutated 3'-UTR of BRUCE (upper panel). Similar experiments as described above were also performed using a naturally EBVinfected gastric carcinoma cell line, SNU-719 (lower panel). The observed luciferase activity was normalized to that of firefly luciferase. Error bars indicate SD (n = 3). * P < 0.05; P < FIG 6 Effect of mir-bart15-3p on the mrna and protein levels of BRUCE. AGS cells were transfected with the mir-bart15-3p mimic or the scrambled control. Cells were harvested after 48 h to extract RNA or protein. (A) Real-time RT-PCR for BRUCE mrna was carried out using the SYBR 585 green qpcr kit. Relative gene expression was calculated according to the comparative Ct method 586 using GAPDH as an internal control. (B) Anti-BRUCE (1:1000) and anti-β-actin (1:1000) antibodies 587 were used for Western blot analysis. β-actin antibody was used to confirm comparable loading. (C)
30 588 The effect of mir-bart15-3p on BRUCE expression was analyzed by Western blots using three 589 independently transfected AGS cell batches and similar results were obtained from them. (D) The 590 Western blot of BRUCE shown in (C) was quantified and normalized to β-actin and presented as a bar graph. * P < FIG 7 Effect of LNA-miR-BART15-3p inhibitor on BRUCE mrna and protein levels. AGS-EBV cells were transfected with the LNA-miR-BART15-3p inhibitor or the control LNA. The cells were harvested after 48 h to extract RNA or protein. (A) Real-time RT-PCR for BRUCE mrna was carried out using the SYBR green qpcr kit. Relative gene expression was calculated according to the comparative Ct method using GAPDH as an internal control. (B and D) Anti-BRUCE (1:1000) and anti-β-actin (1:1000) antibodies were used for Western blot analysis. The β-actin antibody was used to confirm comparable loading. The effect of LNA-miR-BART15-3p inhibitor on the expression of BRUCE in the EBV-infected gastric cancer cells was analyzed by Western blots using three independently transfected AGS-EBV (B) and SNU-719 (D) cell batches. Similar results were obtained from both cell lines. (C and E) The results of the Western blot of BRUCE expression shown in (B) and (D) were quantified and normalized to β-actin and presented as bar graphs. * P < FIG 8 Effect of sirna for BRUCE (sibruce) on BRUCE expression and cell apoptosis. The 606 expression levels of BRUCE mrna (A) and protein (B) in AGS cells 48 h after transfection with the 607 mir-bart15-3p mimic sibruce or the scrambled control were measured by qrt-pcr and Western
MicroRNAs (mirnas) are small noncoding RNAs approximately
 Epstein-Barr Virus-Encoded MicroRNA BART15-3p Promotes Cell Apoptosis Partially by Targeting BRUCE Hoyun Choi, a Hanna Lee, a Sae Rom Kim, b Yong Song Gho, b Suk Kyeong Lee a Research Institute of Immunobiology,
Epstein-Barr Virus-Encoded MicroRNA BART15-3p Promotes Cell Apoptosis Partially by Targeting BRUCE Hoyun Choi, a Hanna Lee, a Sae Rom Kim, b Yong Song Gho, b Suk Kyeong Lee a Research Institute of Immunobiology,
An Epstein-Barr virus-encoded microrna targets PUMA to promote host cell survival
 An Epstein-Barr virus-encoded microrna targets to promote host cell survival The Journal of Experimental Medicine 205(11): 2551-2560, 2008. 1 Elizabeth Yee-Wai Choy, Kam-Leung Siu, Kin-Hang Kok, Raymond
An Epstein-Barr virus-encoded microrna targets to promote host cell survival The Journal of Experimental Medicine 205(11): 2551-2560, 2008. 1 Elizabeth Yee-Wai Choy, Kam-Leung Siu, Kin-Hang Kok, Raymond
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
HEK293FT cells were transiently transfected with reporters, N3-ICD construct and
 Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
The Viral and Cellular MicroRNA Targetome in Lymphoblastoid Cell Lines
 The Viral and Cellular MicroRNA Targetome in Lymphoblastoid Cell Lines Rebecca L. Skalsky 1, David L. Corcoran 2, Eva Gottwein 3, Christopher L. Frank 1, Dong Kang 1, Markus Hafner 4, Jeffrey D. Nusbaum
The Viral and Cellular MicroRNA Targetome in Lymphoblastoid Cell Lines Rebecca L. Skalsky 1, David L. Corcoran 2, Eva Gottwein 3, Christopher L. Frank 1, Dong Kang 1, Markus Hafner 4, Jeffrey D. Nusbaum
Supplementary data Supplementary Figure 1 Supplementary Figure 2
 Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
High level of viral microrna-bart20-5p expression is associated with worse survival of patients with Epstein-Barr virusassociated
 /, 2017, Vol. 8, (No. 9), pp: 14988-14994 High level of viral microrna-bart20-5p expression is associated with worse survival of patients with Epstein-Barr virusassociated gastric cancer Byung Woog Kang
/, 2017, Vol. 8, (No. 9), pp: 14988-14994 High level of viral microrna-bart20-5p expression is associated with worse survival of patients with Epstein-Barr virusassociated gastric cancer Byung Woog Kang
Determination of the temporal pattern and importance of BALF1 expression in Epstein-Barr viral infection
 Determination of the temporal pattern and importance of BALF1 expression in Epstein-Barr viral infection Melissa Mihelidakis May 6, 2004 7.340 Research Proposal Introduction Apoptosis, or programmed cell
Determination of the temporal pattern and importance of BALF1 expression in Epstein-Barr viral infection Melissa Mihelidakis May 6, 2004 7.340 Research Proposal Introduction Apoptosis, or programmed cell
York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
 MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO
 Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Islet viability assay and Glucose Stimulated Insulin Secretion assay RT-PCR and Western Blot
 Islet viability assay and Glucose Stimulated Insulin Secretion assay Islet cell viability was determined by colorimetric (3-(4,5-dimethylthiazol-2-yl)-2,5- diphenyltetrazolium bromide assay using CellTiter
Islet viability assay and Glucose Stimulated Insulin Secretion assay Islet cell viability was determined by colorimetric (3-(4,5-dimethylthiazol-2-yl)-2,5- diphenyltetrazolium bromide assay using CellTiter
HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates
 HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates Department of Microbiology, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, USA
HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates Department of Microbiology, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, USA
Epstein-Barr Virus BART MicroRNAs Are Produced from a Large Intron prior to Splicing
 JOURNAL OF VIROLOGY, Sept. 2008, p. 9094 9106 Vol. 82, No. 18 0022-538X/08/$08.00 0 doi:10.1128/jvi.00785-08 Copyright 2008, American Society for Microbiology. All Rights Reserved. Epstein-Barr Virus BART
JOURNAL OF VIROLOGY, Sept. 2008, p. 9094 9106 Vol. 82, No. 18 0022-538X/08/$08.00 0 doi:10.1128/jvi.00785-08 Copyright 2008, American Society for Microbiology. All Rights Reserved. Epstein-Barr Virus BART
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Purification and Fluorescent Labeling of Exosomes Asuka Nanbo 1*, Eri Kawanishi 2, Ryuji Yoshida 2 and Hironori Yoshiyama 3
 Purification and Fluorescent Labeling of Exosomes Asuka Nanbo 1*, Eri Kawanishi 2, Ryuji Yoshida 2 and Hironori Yoshiyama 3 1 Graduate School of Medicine, Hokkaido University, Sapporo, Japan; 2 Graduate
Purification and Fluorescent Labeling of Exosomes Asuka Nanbo 1*, Eri Kawanishi 2, Ryuji Yoshida 2 and Hironori Yoshiyama 3 1 Graduate School of Medicine, Hokkaido University, Sapporo, Japan; 2 Graduate
Supplementary Materials and Methods
 Supplementary Materials and Methods Immunoblotting Immunoblot analysis was performed as described previously (1). Due to high-molecular weight of MUC4 (~ 950 kda) and MUC1 (~ 250 kda) proteins, electrophoresis
Supplementary Materials and Methods Immunoblotting Immunoblot analysis was performed as described previously (1). Due to high-molecular weight of MUC4 (~ 950 kda) and MUC1 (~ 250 kda) proteins, electrophoresis
HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation
 SUPPLEMENTARY INFORMATION Materials and Methods Human cell lines and culture conditions HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation in exon 20 of BRCA1
SUPPLEMENTARY INFORMATION Materials and Methods Human cell lines and culture conditions HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation in exon 20 of BRCA1
Supplementary information
 Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using
 Supplementary Information Materials and Methods RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using Trizol reagent (Invitrogen,Carlsbad, CA) according to the manufacturer's instructions.
Supplementary Information Materials and Methods RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using Trizol reagent (Invitrogen,Carlsbad, CA) according to the manufacturer's instructions.
Comprehensive Profiling of Epstein-Barr Virus MicroRNAs in Nasopharyngeal Carcinoma
 JOURNAL OF VIROLOGY, Mar. 2009, p. 2357 2367 Vol. 83, No. 5 0022-538X/09/$08.00 0 doi:10.1128/jvi.02104-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Comprehensive Profiling
JOURNAL OF VIROLOGY, Mar. 2009, p. 2357 2367 Vol. 83, No. 5 0022-538X/09/$08.00 0 doi:10.1128/jvi.02104-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Comprehensive Profiling
Supplementary Information
 Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Data Sheet TIGIT / NFAT Reporter - Jurkat Cell Line Catalog #60538
 Data Sheet TIGIT / NFAT Reporter - Jurkat Cell Line Catalog #60538 Background: TIGIT is a co-inhibitory receptor that is highly expressed in Natural Killer (NK) cells, activated CD4+, CD8+ and regulatory
Data Sheet TIGIT / NFAT Reporter - Jurkat Cell Line Catalog #60538 Background: TIGIT is a co-inhibitory receptor that is highly expressed in Natural Killer (NK) cells, activated CD4+, CD8+ and regulatory
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
EBV infection B cells and lymphomagenesis. Sridhar Chaganti
 EBV infection B cells and lymphomagenesis Sridhar Chaganti How EBV infects B-cells How viral genes influence the infected B cell Differences and similarities between in vitro and in vivo infection How
EBV infection B cells and lymphomagenesis Sridhar Chaganti How EBV infects B-cells How viral genes influence the infected B cell Differences and similarities between in vitro and in vivo infection How
Doctoral Degree Program in Marine Biotechnology, College of Marine Sciences, Doctoral Degree Program in Marine Biotechnology, Academia Sinica, Taipei,
 Cyclooxygenase 2 facilitates dengue virus replication and serves as a potential target for developing antiviral agents Chun-Kuang Lin 1,2, Chin-Kai Tseng 3,4, Yu-Hsuan Wu 3,4, Chih-Chuang Liaw 1,5, Chun-
Cyclooxygenase 2 facilitates dengue virus replication and serves as a potential target for developing antiviral agents Chun-Kuang Lin 1,2, Chin-Kai Tseng 3,4, Yu-Hsuan Wu 3,4, Chih-Chuang Liaw 1,5, Chun-
Oncolytic Adenovirus Complexes Coated with Lipids and Calcium Phosphate for Cancer Gene Therapy
 Oncolytic Adenovirus Complexes Coated with Lipids and Calcium Phosphate for Cancer Gene Therapy Jianhua Chen, Pei Gao, Sujing Yuan, Rongxin Li, Aimin Ni, Liang Chu, Li Ding, Ying Sun, Xin-Yuan Liu, Yourong
Oncolytic Adenovirus Complexes Coated with Lipids and Calcium Phosphate for Cancer Gene Therapy Jianhua Chen, Pei Gao, Sujing Yuan, Rongxin Li, Aimin Ni, Liang Chu, Li Ding, Ying Sun, Xin-Yuan Liu, Yourong
Epstein-Barr viral micrornas target caspase 3
 Harold et al. Virology Journal (2016) 13:145 DOI 10.1186/s12985-016-0602-7 SHORT REPORT Epstein-Barr viral micrornas target caspase 3 Cecelia Harold 1,2, Diana Cox 1,3 and Kasandra J. Riley 1 Open Access
Harold et al. Virology Journal (2016) 13:145 DOI 10.1186/s12985-016-0602-7 SHORT REPORT Epstein-Barr viral micrornas target caspase 3 Cecelia Harold 1,2, Diana Cox 1,3 and Kasandra J. Riley 1 Open Access
SUPPORTING MATREALS. Methods and Materials
 SUPPORTING MATREALS Methods and Materials Cell Culture MC3T3-E1 (subclone 4) cells were maintained in -MEM with 10% FBS, 1% Pen/Strep at 37ºC in a humidified incubator with 5% CO2. MC3T3 cell differentiation
SUPPORTING MATREALS Methods and Materials Cell Culture MC3T3-E1 (subclone 4) cells were maintained in -MEM with 10% FBS, 1% Pen/Strep at 37ºC in a humidified incubator with 5% CO2. MC3T3 cell differentiation
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/2/1/ra81/dc1 Supplementary Materials for Delivery of MicroRNA-126 by Apoptotic Bodies Induces CXCL12- Dependent Vascular Protection Alma Zernecke,* Kiril Bidzhekov,
www.sciencesignaling.org/cgi/content/full/2/1/ra81/dc1 Supplementary Materials for Delivery of MicroRNA-126 by Apoptotic Bodies Induces CXCL12- Dependent Vascular Protection Alma Zernecke,* Kiril Bidzhekov,
Original Article Differential expression profile analysis of mirnas with HER-2 overexpression and intervention in breast cancer cells
 Int J Clin Exp Pathol 2017;10(5):5039-5062 www.ijcep.com /ISSN:1936-2625/IJCEP0052419 Original Article Differential expression profile analysis of mirnas with HER-2 overexpression and intervention in breast
Int J Clin Exp Pathol 2017;10(5):5039-5062 www.ijcep.com /ISSN:1936-2625/IJCEP0052419 Original Article Differential expression profile analysis of mirnas with HER-2 overexpression and intervention in breast
Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2)
 Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
Phosphate buffered saline (PBS) for washing the cells TE buffer (nuclease-free) ph 7.5 for use with the PrimePCR Reverse Transcription Control Assay
 Catalog # Description 172-5080 SingleShot Cell Lysis Kit, 100 x 50 µl reactions 172-5081 SingleShot Cell Lysis Kit, 500 x 50 µl reactions For research purposes only. Introduction The SingleShot Cell Lysis
Catalog # Description 172-5080 SingleShot Cell Lysis Kit, 100 x 50 µl reactions 172-5081 SingleShot Cell Lysis Kit, 500 x 50 µl reactions For research purposes only. Introduction The SingleShot Cell Lysis
mirna in Plasma Exosome is Stable under Different Storage Conditions
 Molecules 2014, 19, 1568-1575; doi:10.3390/molecules19021568 Article OPEN ACCESS molecules ISSN 1420-3049 www.mdpi.com/journal/molecules mirna in Plasma Exosome is Stable under Different Storage Conditions
Molecules 2014, 19, 1568-1575; doi:10.3390/molecules19021568 Article OPEN ACCESS molecules ISSN 1420-3049 www.mdpi.com/journal/molecules mirna in Plasma Exosome is Stable under Different Storage Conditions
Department of Pharmaceutical Sciences, School of Pharmacy, Northeastern University, Boston, MA 02115, USA 2
 Pancreatic Cancer Cell Exosome-Mediated Macrophage Reprogramming and the Role of MicroRNAs 155 and 125b2 Transfection using Nanoparticle Delivery Systems Mei-Ju Su 1, Hibah Aldawsari 2, and Mansoor Amiji
Pancreatic Cancer Cell Exosome-Mediated Macrophage Reprogramming and the Role of MicroRNAs 155 and 125b2 Transfection using Nanoparticle Delivery Systems Mei-Ju Su 1, Hibah Aldawsari 2, and Mansoor Amiji
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
Supporting Information
 Supporting Information Identification of Novel ROS Inducers: Quinone Derivatives Tethered to Long Hydrocarbon Chains Yeonsun Hong,, Sandip Sengupta,, Wooyoung Hur, *, Taebo Sim,, * KU-KIST Graduate School
Supporting Information Identification of Novel ROS Inducers: Quinone Derivatives Tethered to Long Hydrocarbon Chains Yeonsun Hong,, Sandip Sengupta,, Wooyoung Hur, *, Taebo Sim,, * KU-KIST Graduate School
supplementary information
 DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
Data Sheet IL-2-Luciferase Reporter (Luc) - Jurkat Cell Line Catalog # 60481
 642 Cornerstone Court W, Ste B Tel: 1.858.829.382 Data Sheet IL-2-Luciferase Reporter (Luc) - Jurkat Cell Line Catalog # 6481 Description Human IL-2 reporter construct is stably integrated into the genome
642 Cornerstone Court W, Ste B Tel: 1.858.829.382 Data Sheet IL-2-Luciferase Reporter (Luc) - Jurkat Cell Line Catalog # 6481 Description Human IL-2 reporter construct is stably integrated into the genome
MicroRNA and Male Infertility: A Potential for Diagnosis
 Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Detection of Apoptosis in Primary Cells by Annexin V Binding Using the Agilent 2100 Bioanalyzer. Application Note
 Detection of Apoptosis in Primary Cells by Annexin V Binding Using the Agilent 2100 Bioanalyzer Application Note Samuel D. H. Chan Marc Valer and Tobias Preckel, Introduction The Agilent 2100 bioanalyzer
Detection of Apoptosis in Primary Cells by Annexin V Binding Using the Agilent 2100 Bioanalyzer Application Note Samuel D. H. Chan Marc Valer and Tobias Preckel, Introduction The Agilent 2100 bioanalyzer
mir 375 inhibits the proliferation of gastric cancer cells by repressing ERBB2 expression
 EXPERIMENTAL AND THERAPEUTIC MEDICINE 7: 1757-1761, 2014 mir 375 inhibits the proliferation of gastric cancer cells by repressing ERBB2 expression ZHI YONG SHEN *, ZI ZHEN ZHANG *, HUA LIU, EN HAO ZHAO
EXPERIMENTAL AND THERAPEUTIC MEDICINE 7: 1757-1761, 2014 mir 375 inhibits the proliferation of gastric cancer cells by repressing ERBB2 expression ZHI YONG SHEN *, ZI ZHEN ZHANG *, HUA LIU, EN HAO ZHAO
(a) Significant biological processes (upper panel) and disease biomarkers (lower panel)
 Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
VIRUSES. Biology Applications Control. David R. Harper. Garland Science Taylor & Francis Group NEW YORK AND LONDON
 VIRUSES Biology Applications Control David R. Harper GS Garland Science Taylor & Francis Group NEW YORK AND LONDON vii Chapter 1 Virus Structure and 2.2 VIRUS MORPHOLOGY 26 Infection 1 2.3 VIRAL CLASSIFICATION
VIRUSES Biology Applications Control David R. Harper GS Garland Science Taylor & Francis Group NEW YORK AND LONDON vii Chapter 1 Virus Structure and 2.2 VIRUS MORPHOLOGY 26 Infection 1 2.3 VIRAL CLASSIFICATION
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation
 Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
JVI Accepts, published online ahead of print on 19 May 2010 J. Virol. doi: /jvi
 JVI Accepts, published online ahead of print on 19 May 2010 J. Virol. doi:10.1128/jvi.00379-10 Copyright 2010, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JVI Accepts, published online ahead of print on 19 May 2010 J. Virol. doi:10.1128/jvi.00379-10 Copyright 2010, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
Berberine Sensitizes Human Ovarian Cancer Cells to Cisplatin Through mir-93/ PTEN/Akt Signaling Pathway
 Chen Accepted: et al.: February Berberine 24, Sensitizes 2015 Ovarian Cancer Cells to Cisplatin www.karger.com/cpb 956 1421-9778/15/0363-0956$39.50/0 Original Paper This is an Open Access article licensed
Chen Accepted: et al.: February Berberine 24, Sensitizes 2015 Ovarian Cancer Cells to Cisplatin www.karger.com/cpb 956 1421-9778/15/0363-0956$39.50/0 Original Paper This is an Open Access article licensed
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Analysis of small RNAs from Drosophila Schneider cells using the Small RNA assay on the Agilent 2100 bioanalyzer. Application Note
 Analysis of small RNAs from Drosophila Schneider cells using the Small RNA assay on the Agilent 2100 bioanalyzer Application Note Odile Sismeiro, Jean-Yves Coppée, Christophe Antoniewski, and Hélène Thomassin
Analysis of small RNAs from Drosophila Schneider cells using the Small RNA assay on the Agilent 2100 bioanalyzer Application Note Odile Sismeiro, Jean-Yves Coppée, Christophe Antoniewski, and Hélène Thomassin
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay
 Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Complete genomic sequence of Epstein-Barr virus in nasopharyngeal carcinoma cell line C666-1
 Title Complete genomic sequence of Epstein-Barr virus in nasopharyngeal carcinoma cell line C666-1 Author(s) Tso, KK; Yip, KY; Mak, CK; Cheung, ST Citation Infectious Agents and Cancer, 2013, v. 8 n. 1,
Title Complete genomic sequence of Epstein-Barr virus in nasopharyngeal carcinoma cell line C666-1 Author(s) Tso, KK; Yip, KY; Mak, CK; Cheung, ST Citation Infectious Agents and Cancer, 2013, v. 8 n. 1,
Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice
 Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
Materials and Methods , The two-hybrid principle.
 The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
Dynamic expression of viral and cellular micrornas in infectious mononucleosis caused by primary Epstein-Barr virus infection in children
 Gao et al. Virology Journal (2015) 12:208 DOI 10.1186/s12985-015-0441-y RESEARCH Dynamic expression of viral and cellular micrornas in infectious mononucleosis caused by primary Epstein-Barr virus infection
Gao et al. Virology Journal (2015) 12:208 DOI 10.1186/s12985-015-0441-y RESEARCH Dynamic expression of viral and cellular micrornas in infectious mononucleosis caused by primary Epstein-Barr virus infection
Herpesviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Herpesviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped icosahedral capsid (T=16), diameter 125 nm Diameter of enveloped virion 200 nm Capsid
Herpesviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped icosahedral capsid (T=16), diameter 125 nm Diameter of enveloped virion 200 nm Capsid
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in
 Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in Supplementary Fig. 2 Substitution Sequence Position variant Sequence original APNCYGNIPL original APNCYGNIPL
Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in Supplementary Fig. 2 Substitution Sequence Position variant Sequence original APNCYGNIPL original APNCYGNIPL
Intrinsic cellular defenses against virus infection
 Intrinsic cellular defenses against virus infection Detection of virus infection Host cell response to virus infection Interferons: structure and synthesis Induction of antiviral activity Viral defenses
Intrinsic cellular defenses against virus infection Detection of virus infection Host cell response to virus infection Interferons: structure and synthesis Induction of antiviral activity Viral defenses
Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases
 Brief Communication Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases Qinghai Zeng 1 *, Cuihong Jin 2 *, Wenhang Chen 2, Fang Xia 3, Qi Wang 3, Fan Fan 4,
Brief Communication Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases Qinghai Zeng 1 *, Cuihong Jin 2 *, Wenhang Chen 2, Fang Xia 3, Qi Wang 3, Fan Fan 4,
Online Data Supplement. Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2
 Online Data Supplement Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2 Yi Lin and Zhongjie Sun Department of physiology, college of
Online Data Supplement Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2 Yi Lin and Zhongjie Sun Department of physiology, college of
EXOSOMES & MICROVESICLES
 The Sample Preparation Experts EXOSOMES & MICROVESICLES The New Standard in Exosome Purification and RNA Isolation Best-in-Class, Pure & Simple Exosome Purification, Fractionation of Exosomal Free & Circulating
The Sample Preparation Experts EXOSOMES & MICROVESICLES The New Standard in Exosome Purification and RNA Isolation Best-in-Class, Pure & Simple Exosome Purification, Fractionation of Exosomal Free & Circulating
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
Instructions for Use. APO-AB Annexin V-Biotin Apoptosis Detection Kit 100 tests
 3URGXFW,QIRUPDWLRQ Sigma TACS Annexin V Apoptosis Detection Kits Instructions for Use APO-AB Annexin V-Biotin Apoptosis Detection Kit 100 tests For Research Use Only. Not for use in diagnostic procedures.
3URGXFW,QIRUPDWLRQ Sigma TACS Annexin V Apoptosis Detection Kits Instructions for Use APO-AB Annexin V-Biotin Apoptosis Detection Kit 100 tests For Research Use Only. Not for use in diagnostic procedures.
MTC-TT and TPC-1 cell lines were cultured in RPMI medium (Gibco, Breda, The Netherlands)
 Supplemental data Materials and Methods Cell culture MTC-TT and TPC-1 cell lines were cultured in RPMI medium (Gibco, Breda, The Netherlands) supplemented with 15% or 10% (for TPC-1) fetal bovine serum
Supplemental data Materials and Methods Cell culture MTC-TT and TPC-1 cell lines were cultured in RPMI medium (Gibco, Breda, The Netherlands) supplemented with 15% or 10% (for TPC-1) fetal bovine serum
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Research Communication
 IUBMB Life, 64(7): 628 635, July 2012 Research Communication MicroRNA-181b Targets camp Responsive Element Binding Protein 1 in Gastric Adenocarcinomas Lin Chen*, Qian Yang*, Wei-Qing Kong*, Tao Liu, Min
IUBMB Life, 64(7): 628 635, July 2012 Research Communication MicroRNA-181b Targets camp Responsive Element Binding Protein 1 in Gastric Adenocarcinomas Lin Chen*, Qian Yang*, Wei-Qing Kong*, Tao Liu, Min
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
University of Oklahoma Health Sciences Center, Oklahoma City, OK, USA; e Boone Pickens
 Nanosomes carrying doxorubicin exhibit potent anticancer activity against human lung cancer cells Akhil Srivastava a,h,+, Narsireddy Amreddy a,h,+, Anish Babu a,h, Janani Panneerselvam a,h, Meghna Mehta
Nanosomes carrying doxorubicin exhibit potent anticancer activity against human lung cancer cells Akhil Srivastava a,h,+, Narsireddy Amreddy a,h,+, Anish Babu a,h, Janani Panneerselvam a,h, Meghna Mehta
Figure S1 Time-dependent down-modulation of HER3 by EZN No Treatment. EZN-3920, 2 μm. Time, h
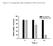 Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
Epstein-Barr Virus: Stimulation By 5 '-Iododeoxy uridine or 5 '-Brom odeoxy uridine in Human Lymphoblastoid Cells F ro m a Rhabdom yosarcom a*
 A n n a ls o f C l i n i c a l L a b o r a t o r y S c i e n c e, Vol. 3, No. 6 Copyright 1973, Institute for Clinical Science Epstein-Barr Virus: Stimulation By 5 '-Iododeoxy uridine or 5 '-Brom odeoxy
A n n a ls o f C l i n i c a l L a b o r a t o r y S c i e n c e, Vol. 3, No. 6 Copyright 1973, Institute for Clinical Science Epstein-Barr Virus: Stimulation By 5 '-Iododeoxy uridine or 5 '-Brom odeoxy
Blocking the expression of the hepatitis B virus S gene in hepatocellular carcinoma cell lines with an anti-gene locked nucleic acid in vitro
 Blocking the expression of the hepatitis B virus S gene in hepatocellular carcinoma cell lines with an anti-gene locked nucleic acid in vitro Y.-B. Deng, H.-J. Qin, Y.-H. Luo, Z.-R. Liang and J.-J. Zou
Blocking the expression of the hepatitis B virus S gene in hepatocellular carcinoma cell lines with an anti-gene locked nucleic acid in vitro Y.-B. Deng, H.-J. Qin, Y.-H. Luo, Z.-R. Liang and J.-J. Zou
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Electron micrograph of phosphotungstanic acid-stained exosomes derived from murine
 1 SUPPLEMENTARY INFORMATION SUPPLEMENTARY FIGURES Supplementary Figure 1. Physical properties of murine DC-derived exosomes. a, Electron micrograph of phosphotungstanic acid-stained exosomes derived from
1 SUPPLEMENTARY INFORMATION SUPPLEMENTARY FIGURES Supplementary Figure 1. Physical properties of murine DC-derived exosomes. a, Electron micrograph of phosphotungstanic acid-stained exosomes derived from
Data Sheet. Notch Pathway Reporter Kit Catalog # 60509
 Data Sheet Notch Pathway Reporter Kit Catalog # 60509 6042 Cornerstone Court W, Ste B Background The Notch signaling pathway controls cell fate decisions in vertebrate and invertebrate tissues. NOTCH signaling
Data Sheet Notch Pathway Reporter Kit Catalog # 60509 6042 Cornerstone Court W, Ste B Background The Notch signaling pathway controls cell fate decisions in vertebrate and invertebrate tissues. NOTCH signaling
Protection against doxorubicin-induced myocardial dysfunction in mice by cardiac-specific expression of carboxyl terminus of hsp70-interacting protein
 Protection against doxorubicin-induced myocardial dysfunction in mice by cardiac-specific expression of carboxyl terminus of hsp70-interacting protein Lei Wang 1, Tian-Peng Zhang 1, Yuan Zhang 2, Hai-Lian
Protection against doxorubicin-induced myocardial dysfunction in mice by cardiac-specific expression of carboxyl terminus of hsp70-interacting protein Lei Wang 1, Tian-Peng Zhang 1, Yuan Zhang 2, Hai-Lian
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism SUPPLEMENTARY FIGURES, LEGENDS AND METHODS
 A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
Supplemental information
 Carcinoemryonic antigen-related cell adhesion molecule 6 (CEACAM6) promotes EGF receptor signaling of oral squamous cell carcinoma metastasis via the complex N-glycosylation y Chiang et al. Supplemental
Carcinoemryonic antigen-related cell adhesion molecule 6 (CEACAM6) promotes EGF receptor signaling of oral squamous cell carcinoma metastasis via the complex N-glycosylation y Chiang et al. Supplemental
Supporting Information Table of content
 Supporting Information Table of content Supporting Information Fig. S1 Supporting Information Fig. S2 Supporting Information Fig. S3 Supporting Information Fig. S4 Supporting Information Fig. S5 Supporting
Supporting Information Table of content Supporting Information Fig. S1 Supporting Information Fig. S2 Supporting Information Fig. S3 Supporting Information Fig. S4 Supporting Information Fig. S5 Supporting
m 6 A mrna methylation regulates AKT activity to promote the proliferation and tumorigenicity of endometrial cancer
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
Utility of Circulating micrornas in Cardiovascular Disease
 Utility of Circulating micrornas in Cardiovascular Disease Pil-Ki Min, MD, PhD Cardiology Division, Gangnam Severance Hospital, Yonsei University College of Medicine Introduction Biology of micrornas Circulating
Utility of Circulating micrornas in Cardiovascular Disease Pil-Ki Min, MD, PhD Cardiology Division, Gangnam Severance Hospital, Yonsei University College of Medicine Introduction Biology of micrornas Circulating
Proteomic profiling of small-molecule inhibitors reveals dispensability of MTH1 for cancer cell survival
 Supplementary Information for Proteomic profiling of small-molecule inhibitors reveals dispensability of MTH1 for cancer cell survival Tatsuro Kawamura 1, Makoto Kawatani 1, Makoto Muroi, Yasumitsu Kondoh,
Supplementary Information for Proteomic profiling of small-molecule inhibitors reveals dispensability of MTH1 for cancer cell survival Tatsuro Kawamura 1, Makoto Kawatani 1, Makoto Muroi, Yasumitsu Kondoh,
Supplementary Material
 Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
Supplementary Data. (continued)
 Supplementary Data SUPPLEMENTARY FIG. S1. Gating strategy for Figure 1. Splenocytes were gated for dimension and for positivity to the different markers representing different cell populations. (A) B cells,
Supplementary Data SUPPLEMENTARY FIG. S1. Gating strategy for Figure 1. Splenocytes were gated for dimension and for positivity to the different markers representing different cell populations. (A) B cells,
mir-18a Inhibits CDC42 and Plays a Tumour Suppressor Role in Colorectal Cancer Cells
 mir-18a Inhibits CDC42 and Plays a Tumour Suppressor Role in Colorectal Cancer Cells Karen J. Humphreys, Ross A. McKinnon, Michael Z. Michael* Flinders Centre for Innovation in Cancer, School of Medicine,
mir-18a Inhibits CDC42 and Plays a Tumour Suppressor Role in Colorectal Cancer Cells Karen J. Humphreys, Ross A. McKinnon, Michael Z. Michael* Flinders Centre for Innovation in Cancer, School of Medicine,
Extracellular Vesicle RNA isolation Kits
 Extracellular Vesicle RNA isolation Kits Summary Section 4 Introduction 42 Exo-TotalRNA and TumorExo-TotalRNA isolation kits 43 Extracellular Vesicle RNA extraction kits Ordering information Products can
Extracellular Vesicle RNA isolation Kits Summary Section 4 Introduction 42 Exo-TotalRNA and TumorExo-TotalRNA isolation kits 43 Extracellular Vesicle RNA extraction kits Ordering information Products can
INTERNATIONAL JOURNAL OF ONCOLOGY 54: , 2019
 326 Paclitaxel resistant gastric cancer MGC 803 cells promote epithelial to mesenchymal transition and chemoresistance in paclitaxel sensitive cells via exosomal delivery of mir 155 5p MEI WANG 1,2, RONG
326 Paclitaxel resistant gastric cancer MGC 803 cells promote epithelial to mesenchymal transition and chemoresistance in paclitaxel sensitive cells via exosomal delivery of mir 155 5p MEI WANG 1,2, RONG
Supplementary webappendix
 Supplementary webappendix This webappendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Ueda T, Volinia S, Okumura H, et al.
Supplementary webappendix This webappendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Ueda T, Volinia S, Okumura H, et al.
High AU content: a signature of upregulated mirna in cardiac diseases
 https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
Annals of Oncology Advance Access published January 10, 2005
 Annals of Oncology Advance Access published January 10, 2005 Original article Annals of Oncology doi:10.1093/annonc/mdi077 Expression of survivin and bax/bcl-2 in peroxisome proliferator activated receptor-g
Annals of Oncology Advance Access published January 10, 2005 Original article Annals of Oncology doi:10.1093/annonc/mdi077 Expression of survivin and bax/bcl-2 in peroxisome proliferator activated receptor-g
Life Science Journal 2016;13(8)
 Structural and Functional Conservation of Epstein Barr Virus microrna Using Phylogenetic Study Ruchi Yadav* AMITY Institute of Biotechnology, AMITY University, Uttar Pradesh Lucknow 226028, UP, INDIA Abstract:
Structural and Functional Conservation of Epstein Barr Virus microrna Using Phylogenetic Study Ruchi Yadav* AMITY Institute of Biotechnology, AMITY University, Uttar Pradesh Lucknow 226028, UP, INDIA Abstract:
Graveley Lab shrna knockdown followed by RNA-seq Biosample Preparation and Characterization Document
 Graveley Lab shrna knockdown followed by RNA-seq Biosample Preparation and Characterization Document Wet Lab: Sara Olson and Lijun Zhan Computational Lab: Xintao Wei and Michael Duff PI: Brenton Graveley
Graveley Lab shrna knockdown followed by RNA-seq Biosample Preparation and Characterization Document Wet Lab: Sara Olson and Lijun Zhan Computational Lab: Xintao Wei and Michael Duff PI: Brenton Graveley
FOXO Reporter Kit PI3K/AKT Pathway Cat. #60643
 Data Sheet FOXO Reporter Kit PI3K/AKT Pathway Cat. #60643 Background The PI3K/AKT signaling pathway is essential for cell growth and survival. Disruption of this pathway or its regulation has been linked
Data Sheet FOXO Reporter Kit PI3K/AKT Pathway Cat. #60643 Background The PI3K/AKT signaling pathway is essential for cell growth and survival. Disruption of this pathway or its regulation has been linked
microrna targets, rhesus lymphocryptovirus, herpesvirus papio, BHRF1, LMP1
 JVI Accepts, published online ahead of print on 20 November 2013 J. Virol. doi:10.1128/jvi.02071-13 Copyright 2013, American Society for Microbiology. All Rights Reserved. 1 Title: 2 Evolutionary Conservation
JVI Accepts, published online ahead of print on 20 November 2013 J. Virol. doi:10.1128/jvi.02071-13 Copyright 2013, American Society for Microbiology. All Rights Reserved. 1 Title: 2 Evolutionary Conservation
Single Cell Quantitative Polymer Chain Reaction (sc-qpcr)
 Single Cell Quantitative Polymer Chain Reaction (sc-qpcr) Analyzing gene expression profiles from a bulk population of cells provides an average profile which may obscure important biological differences
Single Cell Quantitative Polymer Chain Reaction (sc-qpcr) Analyzing gene expression profiles from a bulk population of cells provides an average profile which may obscure important biological differences
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Chairoungdua et al., http://www.jcb.org/cgi/content/full/jcb.201002049/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Expression of CD9 and CD82 inhibits Wnt/ -catenin
Supplemental material Chairoungdua et al., http://www.jcb.org/cgi/content/full/jcb.201002049/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Expression of CD9 and CD82 inhibits Wnt/ -catenin
Notch Signaling Pathway Notch CSL Reporter HEK293 Cell line Catalog #: 60652
 Notch Signaling Pathway Notch CSL Reporter HEK293 Cell line Catalog #: 60652 Background The Notch signaling pathway controls cell fate decisions in vertebrate and invertebrate tissues. Notch signaling
Notch Signaling Pathway Notch CSL Reporter HEK293 Cell line Catalog #: 60652 Background The Notch signaling pathway controls cell fate decisions in vertebrate and invertebrate tissues. Notch signaling
ExoQuick Exosome Isolation and RNA Purification Kits
 ExoQuick Exosome Isolation and RNA Purification Kits Cat # EQ806A-1, EQ806TC-1, EQ808A-1 User Manual Storage: Please see individual components Version 2 8/14/2018 A limited-use label license covers this
ExoQuick Exosome Isolation and RNA Purification Kits Cat # EQ806A-1, EQ806TC-1, EQ808A-1 User Manual Storage: Please see individual components Version 2 8/14/2018 A limited-use label license covers this
Supplementary Figure 1. Confocal immunofluorescence showing mitochondrial translocation of Drp1. Cardiomyocytes treated with H 2 O 2 were prestained
 Supplementary Figure 1. Confocal immunofluorescence showing mitochondrial translocation of Drp1. Cardiomyocytes treated with H 2 O 2 were prestained with MitoTracker (red), then were immunostained with
Supplementary Figure 1. Confocal immunofluorescence showing mitochondrial translocation of Drp1. Cardiomyocytes treated with H 2 O 2 were prestained with MitoTracker (red), then were immunostained with
