Characterization of Micro RNA Signature in Peripheral Blood of Schizophrenia Patients using µparaflo TM mirna Microarray Assay
|
|
|
- Dylan Steven Simmons
- 6 years ago
- Views:
Transcription
1 International Journal of Current Microbiology and Applied Sciences ISSN: Volume 5 Number 7 (2016) pp Journal homepage: Original Research Article Characterization of Micro RNA Signature in Peripheral Blood of Schizophrenia Patients using µparaflo TM mirna Microarray Assay Tihomir I. Vachev 1,3*, Nikolay T. Popov 2, Danail Marchev 1, Hristo Ivanov 4 and Vili K. Stoyanova 3,4 1 Department of Plant Phisyology and Molecular Biology, University of Plovdiv Paisii Hilendarski, 24 Tzar Assen Str., Plovdiv, Bulgaria 2 Psychiatric ward for active treatment of men, State Phsychiatry Hospital Pazardzhik, 28 Bolnichna Str Pazardzhik, Bulgaria 3 University Hospital St. George Plovdiv, 66 Peshtersko Shuse Str., Plovdiv 4000, Bulgaria 4 Department of Pediatrics and Medical Genetics, Medical University, Plovdiv Vasil Aprilov 15- A Str., Plovdiv 4002, Bulgaria *Corresponding author A B S T R A C T K e y w o r d s micro RNA, Peripheral Blood of Schizophrenia, mirna Microarray Assay. Article Info Accepted: 15 June 2016 Available Online: 10 July 2016 Development of biomarkers for psychiatric disease still remains challenging. Recently, numerous alterations in the expression of micrornas (mirnas) in peripheral blood, serum and post-mortem brain tissue have been linked to schizophrenia. Additionally, it is known that mirnas can inactivate a target mrna sequence through binding to its 3 UTR region. Thus, mirnas might not only serve as prognostic biomarkers for diagnosis or prognosis of pathological conditions like schizophrenia, but could also actively modulate gene function. Therefore, the aim of this study was to test whether specific mirna display differential expression in patients with schizophrenia vs. healthy controls by using mirna microarray assay. Venous blood samples of 33 affected patients and 25 healthy controls were collected and the quantification of micrornas was established using mirna microarray assay. Moreover, we searched for potential validated target genes for each of the differentially expressed mirnas that might play a role in the regulation of schizophrenia susceptibility genes. Introduction As one of the most common psychopathological disorders, schizophrenia is considered to be a disease with an extensive social impact. Although symptoms and diagnostic criteria of this mental condition are well defined, the etiology of schizophrenia still remains incompletely elucidated. Both environmental and genetic factors are reported to play important role in schizophrenia development. Heritability of schizophrenia has already been described and its genetic background has been clearly established. Hersen and Beidal report higher co-occurrence of the disease among monozygotic in comparison with dizygotic twins (Herson et al., 2011). Additionally 503
2 several genome-wide association studies have revealed the involvement of many different loci in schizophrenia pathology (Mirnics et al., 2000). Abnormal gene expression connected with the disease has also been described by many groups (Perkins et al., 2007). MicroRNAs (mirnas) can be considered as master regulators of essential cellular processes (Bartel, 2009; Sun et al., 2010) and could therefore be affected by disease processes. Profiling of mirna is being undertaken in the search for potential biomarkers in a variety of diseases. Increasing evidence shows that mirnas take part in many biological processes, including cell proliferation, differentiation, migration, and apoptosis (Bartel, 2009). Thus, changes in the levels of expression of either mirna could have long-term consequences, making it plausible for them to play a role in the etiology of schizophrenia. Materials and Methods Participants The study design and the Inform Consent Form (ICF) were approved by the Ethics Committee of Plovdiv Medical University. The Institutional Review Board of the university approved the use of the samples for this study. Thirty schizophrenic patients from State Psychiatric Hospital - Pazardjik, Bulgaria were recruited аfter obtaining a written informed consent. All the patients were interviewed by certified Mini- International Neuropsychiatric Interview by a certified psychiatrist rater to evaluate diagnosis of schizophrenia on Diagnostic and Statistical Manual of Mental Disorders forth text revised edition (DSM IV TR) criteria and to exclude any mental disorder in the controls. The control group included 25 healthy subjects matched by gender and age. Main inclusion criteria was that the participants did not receive any medication before blood sampling for at least 2 weeks. People with other chronic medical conditions and current somatic/neurologic diseases, alcohol or drug abuse/dependancy were excluded. Blood collection and RNA extraction An aliquot of whole peripheral blood (2.5 ml) for each subject (schizophrenia and healthy controls) was collected directly into a PAXgene blood RNA tubes (PreAnalytiX) and stored at room temperature for minimum of 4 hours and than freezed at - 20 C, prior to total RNA extraction. Total RNA was isolated using PAXgene blood mirna kit (PreAnalytiX), according to the manufacturer s protocol. Assessment of absorbance ratio of 260/280 nm revealed that all RNA samples are with sufficient quality ( ) for further analysis, according to the service requirements. Pooled samples were created by adding an equivalent amount of total RNA from each individual sample to final concentration of 5 µg RNA samples. Pooled RNA samples were precipitated according to the service requirements, each pooled RNA sample was mixed with 1/10th volume of 3M NaOAc, ph 5.2 and 3 volume 100% ethanol, tо the final volume of 400 µl. Aliquots of pooled RNAs were frozen at -80 C and shiped on dry ice. RNA integrity of pooled samples was assessed by agarose gel electrophoresis and checked by Agilent 2100 Bioanalyzer. MicroRNA Expression Profiling (LC Science) The µparaflo TM mirna microarray assay was performed using a service provider (LC Sciences, Houston, TX) with a proprietary microfluidic array based on the Sanger mirbase v18.0 database ( 504
3 Rfam/mirna), designed to detect 1898 unique mature human mirna sequences. MiRNA microarray analysis was carried out to determine differential expression in blood mirnas between pooled samples of healthy individuals and pooled RNA samples derived from patients with schizophrenia. Fluorescence images were collected using a GenePix 4000B laser scanner (Molecular Device, Sunnyvale, CA) and digitized using Array-Pro image analysis software (Media Cybernetics, Bethesda, MD). Target gene analysis of differentially expressed micrornas in schizophrenia patients In order to study the potential modulation in gene expression that may be associated with specific mirnas changes identified in our study, mirna validated target studies were carried out using the publically available database, mirwalk available at ( The validated targets module hosted experimentally verified mirna interaction information associated with target genes were used. Results and Discussion Analysis of mirna microarray assay (LC Sciences) In order to study the potential significance of certain mirnas in schizophrenia, we analyzed global mirna expression profiles in pooled peripheral blood samples from 30 schizophrenia patients and 25 healthy controls using mirna microarray for mature human mirnas. To assess the reproducibility of the microarray technique, two replicates from the same preparation were used for each of the two microarray hybridizations conducted in this research. In order to statistically estimate significance of every observed change in microrna expression, a two sample independent t test was performed. Comparison between the micro-rna expression levels in these pooled samples identified 19 mirnas with significant change in expression (p < 0.05), the majority (13) of which showed increased expression in schizophrenia patients. Among them, 4 showed fold change greater than 2.5 compared with the controls. 11 micro-rnas appeared to be significantly downregulated (p < 0.05) in schizophrenia patients in comparison with healthy controls. These notable alterations in regulation of the respective micrornas suggest that they may play important role in the pathogenesis of schizophrenia. Moreover the heat map, which was generated to visualize the results of hierarchical clustering, clearly shows the distinction between schizophrenia and control samples. (Table 1). The cluster graph represents all micrornas that displayed statistically significant change with p value less than The clustering analysis revealed micrornas (such as hsa-mir p and hsa-mir-421) with very similar patterns of expression, despite the fact that their genes occupy very distant genome loci. High-throughput mirna expression analysis of 1898 unique annotated mature human mirna sequences (from mirbase 18.0) revealed a set of 19 differentially expressed mirnas. 13 of which was upregulated and 6 down regulated, respectevly (p<0,05) Significantly downregulated molecules includes, mir-183-5p, (log 2-0,65, p=0,026), mir p, (log 2-0,85, p=0,029), mir-125b-5p, (log 2-0,58, p= 0,033), mir-192-5p, (log 2-0,47, p=0,033), mir-1976, (log 2-0,42, p= 0,033), mir p, (log 2-0,54, p=0,044). A trend towards increased expression (upregulation) showed mir-4311, (log 2-0,96, p=0,014), mir p, (log 2-0,38, p=0,022), mir-421, (log 2-0,30, p=0,022), mir-4498, (log 2-0,98, p=0,027), mir- 505
4 4691-3p, (log 2-1,50, p=0,029), mir-3185, (log 2-1,01, p=0,035), mir p, (log 2-0,70, p=0,037), mir p, (log 2-2,54 p=0,039), mir-4479, (log 2-0,32, p=0,041), mir-30b-3p, (log 2-1,26, p=0,041), mir p, (log 2-1,45, p=0,041), mir-492, (log 2-1,97, p=0,042) and mir-30e-3p, (log 2-1,13, p=0,047). Target Analysis of Schizophrenia- Associated micrornas Validated target genes of differentially expressed micrornas identified by mirna microarray analysis are shown in Table 1. Abnormal mirna expression has widely been reported in numerous human diseases. A series of recent studies have shown that mirnas in whole blood serum and plasma are considered to be a biomarkers for several diseases (Scholer et al., 2010). Several recent reports have confirmed the principle that the peripheral blood is a potentially useful source of diagnostic biomarkers for neuropsychiatric diseases and disorders (Gardiner et al., 2012; Lai et al., 2011; Mellios et al., 2012; Ivanov et al. 2015). Some of them have indicated the usefulness of using whole blood for mirna expression studies and have demonstrated altered expression of blood and brain mirnas involved in brain plasticity and maturation associated to schizophrenia (Gardiner et al., 2012; Lai et al., 2011; Mellios et al., 2012). All these studies suggest that whole blood cells could reflect, or in part share, the molecular phenotype of neural cells and thus could be used in studies of psychiatric disorders. This study demonstrates the use of mirna signatures as an important advance in schizophrenia research. Elucidating mirna dysregulation in schizophrenia will promote understanding of the contribution of mirnas in neurocognitive, behavioral and other common disorders that demonstrate differential mirna expression. Recently, global analysis of the expression profile of micro RNA from peripheral blood mononuclear cells (PBMC) of 112 patients with schizophrenia and 76 non-psychiatric controls conducted by Gardiner and colleagues revealed a significant dysregulation of micro RNA expression in schizophrenia. In their study, the authors found 83 micro RNA molecules that showed statistically significant decreased expression in the group of schizophrenia patients. In addition, some of the differentially expressed micro RNA molecules (~ 20%) are transcribed from a single imprinting locus, maternally expressed in the DLK1- DIO3 region on chromosome 14q32 (Gardiner, 2011). All this suggests the possible involvement of major locus-specific genetic or epigenetic control mechanisms associated with this disorder. The micro RNA molecules, identified by us with a similar change in expression and mapped in single loci would be intriguing subject for clarification of the relationship between their expression and localization into the genome (locus-specific expression). In order to assess in in detail the biological consequences of the observed changes Gardiner et al. also analyze the predicted target genes of the differentially expressed micro RNA molecules. They found that the target genes are involved in biological pathways directly related to nerve function by axons guidance, regulation of actin cytoskeleton, long-term potentiation, longterm depression, adhesion (focal adhesions) and neurotropins (Gardiner, 2011). 506
5 Table.1 List of validated target genes of dysregylated mirnas MiRNAs Dysregulation Validated target genes mir-183-5p Downregulated AAMP, ABCF1, ARHGDIA, ASNS, AUH, CCND1, CAPNS1, CCNB1, SLC31A1, CR1, CS, CSNK2A1, DAP, DMWD, DOCK1, DTYMK, EGR1, FAT2, FEN1, FOXO1, FNTB, GAS1, GATA6, KAT2A, GLO1, GLUL, GSPT1, GSR, HARS, HCFC1, HINT1, HLA-A, HNRNPL, HOXA5, HSPA1B, HSP90AA1, FOXN2, IDH2, IGF1R, FOXK2, INSIG1, ITGA5, ITGB1, ITPKB, KIF2A, KIF5C, MCM4, MSH2, TRIM37, MYO10, HNRNPM, NEO1, NOTCH2, P4HB, PDE8A, PLOD2, POLH, PPP3R1, PSEN1, PSMD13, PTPN9, PTPN11, RAB5B, RALGDS, RCN2, REV3L, RPLP0, SRSF2, SRSF2, SFSWAP, SLC2A3, SLC5A3, SNRPD3, ST13, CCT3, TRO, UBP1, UCHL3, UQCRC1, EZR, BRPF1, ARHGEF5, PTP4A2, TTF2, ITGA8, API5, DGAT1, TIMELESS, BTRC, BTAF1, USP8, RPL23, BAG4, AKAP12, TRAF4, KIAA0430, PHF14, KNTC1, ARHGAP32, EIF4A3, SERTAD2, PLEKHM1, PDCD6, PREB, TRIM28, RNF41, CTDSPL, CEPT1, GNB2L1, LYPLA1, MYBBP1A, HYOU1, TXNIP, SRSF10, USP19, STIP1, C14orf1, MTF2, RBM34, MYCBP2, PPRC1, ERC1, TNRC6B, FAM175B, ZC3H4, USP22, LARP1, OPN3, TMEM245, NIPBL, UBXN7, GNL3, PDCD4, HIPK2, SENP1, RDH11, LARS, NUB1, RC3H2. OTUD4, KLHL24, TRIT1, COMMD8, RIF1, LAPTM4B, SMEK1, TSR1, SERTAD4, C10orf2, C15orf39, RTN4, THOC2, CNOT6, MTUS1, NUFIP2, ARHGAP21, MKL1, ZFAT, ABHD17C, ELAC2, FKBPL, TUT1, BCL11B, WNK1, UBE2Z, STK33, TMEM108, GCC1, ZFHX4, PEAK1, RPAP2, FAM188A, TUBB1, GUCD1, STK40, ARFGAP2, TNRC18, FAM104A, MASTL, ABCC10, FAM110B, LENG8, NAA30, ZC3H18, CYB5D2, IZUMO2, TPRG1L, C5orf24, KDELC2, C18orf25, KLHL23, SH3D19, NHSL2, WDR53, SHISA2, ZBTB34 mir-192-5p Downregulated ACADSB, ACVR2A, ACVR2B, ACVR2B, ADRBK2, AGL, ALCAM, ALCAM, ALDH9A1, ANG, APC, XIAP, TRIM23, ASPH, ATF1, ATP1B1, BARD1, BARD1, BCL2, BGLAP, BICD1, BLM, BMPR2, DST, BRCA1, BRCA2 BTC, BUB1B, BYSL, CA8, CAMK4, CASP7, CCNE1, ENTPD3, CDC20, CDC25A, CDKN1B, CDKN2A, CDKN2D, CDKN3, CENPA, CENPE, CENPF, CFL2, RCBTB2, CHM, CHML, CLIC1, CLK1, CREBL2, CRK, CTH, CYP24A1, DBT, DDOST, DDX3X, DDX3X, COCH, DHCR24, SLC26A2, HBEGF, DYRK1A, E2F5, ECT2, PHC2, EEF1A2, EFNB2, EGR1, EGR1, EIF2S1, EML1, EPS8, ERCC3, ERCC3, ERCC4, ERCC4, EXTL2, ACSL1, FBN1, FEN1, GPC4, FGF2, FOS, CENPI, GATM, GCH1, GNRH1, GPR19, GPR37, GRIA1, MSH6, HADH, HAS3, HMGCS1, HMMR, FOXA1, HOXA10, HOXA13, HOXB9, HRH1, HSPA2, ICAM3, ID1, ID2, ID4, IDI1, IFRD1, IGF1R, IL1R1, IL1RAP, IL6ST, IL7, IL15, IMPA1, INSIG1, IRAK1, ITGAV, ITGB1, KCNS3, KIF5B, KIF5C, KPNA5, LIMS1, LIPA, FADS1, LMAN1, LMNB1, LOXL2, MAD2L1, MAN2A2, MCAM, MCM3, MCM3, MCM6, DNAJB9, MDM4, MEIS2, MAP3K1, MFI2, MKI67, MSN, MSN, MT1F, MT1X, MXI1, MYB, PPP1R12A, NAB1, NBN, NCBP1, NEK1, NFIB, NFYA, NKX3-1, NSF, NUCB2, ODC1, OPA1, ORC1, OVOL1, PAFAH1B1, PARN, PAWR, PCDH7, PDE2A, PET112, CDK14, PGM3, PIM1, PIM1, PLAGL2, PLAU, PLOD1, PLS3, UBL3, PODXL, POU2F1, PPARG, PPAT, PPM1A, PPP1CA, PPP1CB, PRIM1, PRKAA2, PRKAR1A, MAPK9, DNAJC3, PRNP, PRPS2, LGMN, PTPRE, PTS, RAD1, PURA, RAB1A, RAB2A, RAB2A, RAB3B, RAB27A, RAD21, RAD51, RAD51, RAP1GAP, RB1, RB1, RBL2, RECQL, RET, RFC4, RNASE4, RNASEL, RNF6, RPS6KB1, RRM1, ATXN7, SCD, CCL3L1, TRAPPC2, SGCB, STIL, 507
6 SLC1A4, SLC7A2, SLC9A5, SMARCA2, SMARCB1, SNRPD1, SOAT1, SP4, SPARC, SPTBN1, SRD5A1, SS18, STK3, MED22, TBL1X, TCEB3, HNF1B, TCF7, TDG, THBD, THOP1, KLF10, TMPO, TOP1, TPM4, HSP90B1, TRPC1, TTK, TTPA, UBE2D1, UBE2D2, UBE2V2, UGT8, UNG, UPP1, UQCRC2, USP1, UVRAG, TRPV1, WHSC1, WNT3, XK, XPA, XPO1, ZBTB14, ZNF45, ZNF85, ZNF165, ZNF184, ZNF200, ZNF215, ZNF226, ZNF230, LRP8, PXDN, MANF, DEK, ALDH5A1, EPM2A, BRD3, STAM, CUL5, NCOA3, CDC7, FZD1, FZD4, FZD7, PIP4K2B, EEA1, STX7, STX7, CUL3, TTF2, RANBP3, PKP4, ENC1, CDC14A, DEGS1, TMEFF1, B3GALNT1, B3GALNT1, ABCC3, TNFSF13, PABPC4, CD164, CD164, SAP30, HESX1, IQGAP1, KAT2B, FUBP1, AP1S2, HERC2, MPZL1, SPAG9, USP14, NMI, SLC16A6, SYNGR1, SLC16A7, LRRFIP1, LRRFIP2, DLG5, STK17B, CD83, MMP20, NREP, TRIP13, ZNHIT3, B4GALT6, ARHGAP29, FAM189A2, TJP2, ABCG2, EIF2AK3, AKAP7, KIF23, ATP6V1G1, KIF20B, ARHGEF10, USP34, DLGAP5, MTSS1, ARHGAP11A, GAB2, EPM2AIP1, PPP1R26, PSMD6, SEC24D, FAM20B, KBTBD11, KIF14, AMMECR1, HS3ST3B1, ZBTB33, SH2B3, HCN4, TROAP, SMC4, ACTR2, KIF20A, CTR1B, ARL4C, ZNF263, AKAP9, LHFP, ZNF443, MFSD10, TRAIP, ZNF267, CCNO, TFG, ABCA8, CORO2B, ANAPC10, ST3GAL6, TUBGCP3, C1D, C1D, ZBTB18, SEC23B, SYNCRIP, SEMA4D, SEMA4D, CIB2, EAF1, LAMTOR5, SLC19A2, ARFGEF1, SMC2, ATP5L, KHDRBS3, CTCF, TNFSF13B, POLQ, TCFL5, PLK4, PHTF1, ZMYND11, ZBTB6, ZNF273, NMU, AP3M2, RAB40B, CKAP4, TMED10, PAPD7, CPSF6, TOPBP1, DNAJB4, PRSS23, PRDM4, IBADH, CIT, FGFR1OP, PTP4A3, ERLIN2, AKAP11, RASSF8, SEC63, USP18, ZNF652, ZFP30, RHOBTB3, COBLL1, RAB11FIP2, ICK, ENPP4, INPP5F,ANKRD6,CLSTN1,RUFY3, WDR47, NCBP2, ELL2, SCMH1, LIMCH1, FNBP1, ENDOD1, EMC1, SNX13, PASK, OTUD3, TMEM194A, FKBP15, SATB2, RPGRIP1L, NCAPH, DICER1, SIRT1, TARDBP, ABCB10, NPTXR, LEPROTL1, MACF1, GLTSCR1L, ZNF346, ARL2BP, RABGAP1, POLA2, SLC7A11, TMEM2, SH3BP4, PRKD3, CADM1, PITPNB, BCL2L13, PANX1, PANX1, DSTYK, RAD54B, DNAJB5, PARM1, ANAPC15, SAMHD1, TPGS2, LRIG1, TRNL1, APPL1, ARHGEF26, TOR1AIP1, C10orf137, PHF19, FBXW2, PITPNC1, FBXO25, FBXO5, FBXO4, FBXO3, PCOLCE2, NUFIP1, NPHP3, AGO2, SLC39A1, DIMT1, PDLIM3, KCNMB4, TOR1B, PPP2R3B, SLCO3A1, FLVCR1, NOB1, MRPL13, ATAD2, RACGAP1, GRHL1, TFCP2L1, SENP1, SNX10, PSMC3IP, GPSM2, UBIAD1, PIK3R4, CUZD1, IRX4, COPS7A, RNF141, CDON, MTERFD1, METTL9, NAA20, IRAK4, RNFT1, CKLF, PLCE1, NIN, ZDHHC2, CCDC174, RSRC1, THEM6, CRLF3, GULP1, PTPLAD1, DTL, LARS, TAF9B, CCZ1, TMEM66, NLK, RAB23, CMPK1, RAB8B, SIX4, SIX4, COQ3, BRWD1, SETD4, C21orf91, ANLN, WDR44, MRPL50, MTMR12, NDFIP2, FAM63B, RSBN1, EPDR1, NDE1, ERCC6L, VPS13C, BIVM, BCOR,UHRF1BP1, WHSC1L1, CMTM6, PARP16, PIGX, CLN6, GID8, CDCA4, UACA, CXorf57, TRIM68, LARP1B, RFWD3, CEP55, FAIM, RIC8B, BBS7, FANCI, NEIL3, NUDT15, PHF10, UBE2W, MIS18BP1, AGPAT5, MNS1, SLC39A9, CHDH, HJURP, MCM10, CSGALNACT2, CHST12, HES6, IL17RB, ZNF823, ENOSF1, BMP2K, RNPC3, FERMT1, G2E3, TBC1D22B, DEPDC1, CHD7, SLC48A1, KDM4D, RADIL, ARHGEF40, FAM178A, IQCC, N4BP2, C1orf112, TMEM30A, ZNF701, TDP1, SAYSD1, MBNL3, DCP1A, EAF2, WWC3, PPP2R2D, RCC2, NKRF, SLC25A40, SERTAD4, CTPS2, EMC7, PAK6, ARNTL2, NT5M, STOX2, KIF15, CHPT1, CTNNBIP1, CCDC47, CASC5, ENTPD7, PHTF2, ATP10D, ZNF248, GPR126, NHSL1, CLK4, KIAA1143, MKL2, KIDINS220, MIB1, KLHL42, NCEH1, STAMBPL1, KLHL8, HCN3, FANCM, DENND1A, IGDCC4, ZFYVE28, TRIB3, GATAD1, PTBP2, 508
7 CREBZF, INIP, ELOVL5, AASDHPPT, SPCS3, BCORL1, CLSPN, PERP, PERP, CENPK, MOAP1, SLC39A8, NOD2, TOR3A, PAPD5, NFKBIZ, ZMAT3, MPP5, NUCKS1, CCDC14, FNDC3B,ELOVL1, PLEKHG2, REEP1, SOWAHC, WNK1, ZBTB10, OTUB2, MIS12, DDX50, DDX54, KCTD15, PRRG4, C2orf49, RNF26, ZXDC, HAUS3, RNF219, OGFRL1, SCRN3, CCDC121, RPAP3, HSPBAP1, PLEKHF2, GALNT12, E2F8, CLIP4, FBXO31, VASH2, ZMYM1, TMEM180, BORA, RPAP2, ATAD5, MAP6D1, CNTNAP3, L2HGDH, ERMP1, PHF17, WDR76, DSN1, RMI1, WWC2, CCDC15, PIF1, CCDC176, ASRGL1, DBF4B, NAA50, C2orf44, KLHL15, TRAF3IP3, DDHD1, DUSP16, SGPP1, APOLD1, FAM83D, CAB39L, MAP1LC3B, C6orf62, LOC81691, VANGL1, SPRY4, ZNF611, KIF18A, NUF2, DNAL1, UCK1, ATG10, ROPN1L, GSG2, BRIP1, B3GNT5, KATNAL1, RAB6C, CEP78, TMEM164, MEX3B, LRRC8C, ANKRD32, TCHP, PRADC1, PHF6, TMEM246, PCGF5, HOOK3, KIAA1841, RPL43, FAXC, HIATL1, FAM126A, PSRC1, LMNB2, C1orf198, ATAD1, TMTC4, FAM136A, SLC35B4, FAM222A, ALG10, SERAC1, PRPF38A, CGNL1, UBASH3B, CEP19, ARHGAP19, USP45, KIAA1671, EAF1, ZIC5, LRRCC1, KIAA1731, FHDC1, ZC3H12C, ANKRD13A, LMLN, EFCAB11, C5orf30, TICRR, LYRM7, BTF3L4, C12orf65, SLC25A51, MTDH, TIFA, TMEM169, TEX30, PIGM, PIGM, TBC1D31, CADPS2, FAM71E1, GLCCI1, TMEM106A, TMEM200A, OSBPL6, OSBPL8, OSBPL10, C1QTNF3, ATG4A, KCTD12, DIS3L, LRRC58, LYSMD3, GINM1, RBP7, LEAP2, SFR1, ZNF641, NEDD1, NIPA1, C16orf46, NOXO1, GABPB2, CCSAP, CMPK2, LYPD6, AHSA2, FBXO36, DNAJC19, DCBLD2, C4orf32, EMB, PRRC1, AFAP1L1, HINT3, FAM199X, SPIN4, ZSWIM3, KDELC2, PRICKLE1, ALG10B, MIPOL1, CEP128, NRG4, EME1, SAMD13, SLC35F3, LSM14B, CKAP2L, GAREML, SEPT10, ZNF385B, FAM132B, KLHL23, SGOL2, SLC16A14, NIPAL1, BTNL9, SAMD3, ESCO2, CLVS1, C9orf163 FAM120AOS, KIAA1958, USP54, SLC35G1, TMTC2, TMTC3, GPR180, ZFP1, ZNF519, C18orf54, ZNF320, ZNF114, IFNLR1, SASS6, FAM171B, SGMS2, FUT11, ANO6, ACSF3, C1orf86, TMEM17, DHFRL1, RDM1, CCDC111, C8orf47, ZNF449, USP12, C10orf35, CCNY, RTKN2, HYLS1, TMEM136, SKA1, OTUD1, CCDC7, SPATA13, KDM1B, PXDC1, CCZ1B, RSBN1L, STXBP4, SLC25A30, CERS6, MSRB3, EHBP1L1, MCM9, C8orf46, ZDHHC23, PCSK9, NPNT, CNEP1R1, ASPM, GAS2L3, LC46A3, GPR137C, C17orf58, ZADH2, ZNF780A, RIMKLA, CCDC150, FAM126B, EOGT, CYP4V2, LIN9, TRIM59, MMAB, CHSY3, GOLGA6A, TMC3, CCDC18, KIF24, GEN1, TICAM2, FAM101B, FAM111B, LHFPL4, C5orf34, CA13, NHLRC3, C1orf53, VGLL3, ZBTB34, ZNF704, FAM110C, C16orf52 mir30e-3p Upregulated LNPEP, TRIM37, NFKBIA, SERPINE2, PPP2R2B, RBBP7, RPS3, SGSH, YWHAE, CLSTN1, TBC1D9B, UBXN1, DGCR8, FAM47E mir30b-3p Upregulated RORC, MYBBP1A, RNPS1, CXXC4 mir421 Upregulated SLC25A6, RHOB, RND3, ATM, ATP6V0C, ZFP36L2, CASP2, CEBPB, COPA, DDX3X, DFFB, DUSP3, EEF1A1, EIF4A1, EIF4G2, FAT2, NR6A1, GCSH, GRK6, NR3C1, HADHA, HTT, HSP90AA1, INPP4A, ITPK1, LBR, LIMS1, CD46, SEPT2, NPM1, PHKA1, POLR2D, PPP2CB, PRCC, MAPK8, MAPK8, PSME1, RANBP2, RBBP7, TRIM27, RPL18A, RPS10, RPS18, 509
8 SRSF7 SLIT1, SRP9, SRPR, ZXDA, RASAL1, AP3B1, USO1, VAPA, TTC37, CEP57, KIAA0100, DDX39A, DDX39A, TUBA1B, SLC9A6, CAP1, LEPREL4, MTHFD2, PPARGC1A, MSL3, CDC37, DDX20, ATXN2L, MLXIP, CNOT1, PHLPP2, NCDN, RHOBTB2, ZC3H7B, DNAJC16, SMCHD1, PPP1R13B, SPECC1L, LARS2, CBX7, ACOT9, ZMYND8, FKBP8, NIPBL, RAB26, NSL1, FAM32A, AGO1, AFF4, EIF3K, LMCD1, RPL26L1, DNAJC27, AMZ2, RNF138, PQLC2, GID8, ATG2B, RIF1, MIS18BP1, LIN7C, COPRS, ZNF280C, NDC1, DOK4, TSR1, CMAS, DUS3L, CDC42SE2, RALGAPB, SENP7, ISY1, FAM160B1, CREBZF, NCAPG, PCNXL4, MRPL38, GID4, AHNAK, TMEM109, CTC1, FBXO11, RAB11FIP1, CD276, PRR7, GFOD2, FAM172A, PARP10, RAB2B, ZNF30, MCFD2, RASL10B, NXPE3, SFXN1, PAXBP1, FAM122A, FAM168B, GAREML, CREBRF, OTUD1, SREK1IP1, RGMB, ZNF678, RBMXL1, HIST2H4B mir125b-5p Downregulated ABL1, ACLY, ADM, PARP1, AKT1, XIAP, ARF3, ATP5B, BACH1, BAK1, BAK1, BCL2, BCL3, PRDM1, BMPR1B, CBFB, CBFB, CCNE1, CDKN2A, CEBPA, CEBPG, CNGB1, COX7C, CREBBP, MAPK14, CYP24A1, E2F2, E2F3, E2F3, EEF1A1, EIF4EBP1, EIF5, EMD, EPHA7, ERBB2, ERBB2, ERBB2, ERBB3, ETF1, ETS1, EXTL3, FAT1, FBN1, FGFR2, FUS, GABRB3, GLI1, MKNK2, GRIN2A, GRINA, HDGF, HK2, HMGN1, HMGA1, HNRNPK, HOXA13, HSPA1B, HSPD1, IGF2, IRF4, KCNS3, KPNB1, KRT7, LIF, LSS, LTA4H, MMP13, ABCC1, MTF1, MUC1, NME2, NPM1, NRD1, NTRK3, PAM, PCBP2, PIGF, PKM, PMAIP1, MED1, PPAT, PRKAG1, PSMB1, RAF1, RANBP2, RNH1, RPL3, RPL29, RPL35A, RPLP0, RPS3A, RPS6KA1, RPS7, RPS12, RPS28, SORT1, S100A8, ATXN1, SDHB, SEL1L, SHMT1, SLC7A1, SLC19A1, SLC25A1, SMARCD2, SMO, SMO, SNRPB, SSR3, STAT3, SUPT6H, TDG, PRDX2, TEF, TNFAIP3, TP53, TP53, TP53, UNG, VCL, VDAC1, VDR, ZNF212, KMT2D, FXR1, HMGA2, TAF15, FZD4, IRS4, KHSRP, STC2, DGAT1, B4GALT3, PABPC4, SGPL1, MTMR3, SLC7A6, USP8, MTMR4, SLC16A4, AURKB, MFHAS1, SLC9A3R2, RASAL2, PTGES, ZNF592, NUP93, EDEM1, STARD8, RNF144A, MATR3, DHX38, IP6K1, KIAA0141, CEP170, UBAP2L, THRAP3, HUWE1, RASGRP1, SUGP2, ABCC4, RGS19, BCKDK, SMYD5, NEBL, RNASEH2A, SMC2, ARID3B, MAP3K2, NES, HBS1L, ZMYND11, SEC24A, ILVBL, ADRM1, AKAP2, LYPLA2, CASC3, CASC3, R3HDM2, BTBD3, SPEN, EMC1, TNRC6B, PLXND1, NUP205, GANAB, TBC1D1, SIK2, PDS5A, EXOC7, FAM208A, CRTC1, FRAT2, RYBP, TARDBP, SRRM2, NNT, CARHSP1, HSPBP1, TMEM2, BCL2L13, THUMPD3, ZDHHC5, ULK3, TIMM10, PABPC1, NPTN, BBC3, NDOR1, RRP7A, NKIRAS2, STRN3, PSAT1, MYEF2, MRTO4, PDZD11, CD320, KLF13, CHMP3, LUC7L3, ZFYVE1, PTOV1, CHPF2, MTRF1L, MTMR12, BCOR, NSUN2, C1orf109, C17orf80, RFWD3, UBA6, THUMPD1, NDC1, WDR11, ZNF395, PCDHGB2, MEPCE, C15orf39, SMIM8, PPM1H, CGN, CTDSP1, FAM60A, UNKL, RMND5A, WNK1, DDX50, DYNC2H1, SMC6, LIN28A, EHMT1, PIP4K2C, ESRP2, MYO19, ABTB1, FBXO38, CSRNP2, SPRTN, C2orf88, ACSS1, PLEKHA8, LTV1, FBXL20, TNKS1BP1, LBX2, THAP3, BMF, DHX57, FMNL3, TP53INP1, SLC35A4, LACTB, GPRIN1, CCDC124, FAM199X, SAMD10, RIMS4, E2F7, CMTM4, LSM14B, FAM91A1, ZNF483, QSOX2, RHOV, TMEM9, PLA2G4F, CDRT4, C19orf54, HKR1, MMAB, KIF24, GALNT18, LIN28B, HIST2H2BF, NRARP, TIMM23 510
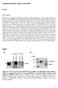
9 Fig.1 Differentially expressed micrornas identified by mirna microarray analysis in whole blood samples from SZ patients versus control subjects. In a similar study, Lai et al. (Lai et al. 2011) identified specific expression profile of seven micro RNA molecules in an initial cohort of 30 patients with schizophrenia and 30 controls. 6 of the micro RNA molecules were with increased expression (micro RNA-34a, micro RNA-449a, micro RNA- 564, micro RNA-548d, 572 micro-rna and micro RNA 652) and one - micro RNA 432 with decreased expression. These results were confirmed in an extended subsequently cohort of 60 patients with schizophrenia and 30 controls. Based on our results it can be assumed that changes in the expression levels of micro RNA molecules contribute to complex global changes in gene expression, that underlie the pathophysiology of schizophrenia. In this study we observed that micro RNA expression profile from peripheral blood is dysregulated in the patients diagnosed with the disorder. At the molecular level, our results can be explained by one or more of the following options: a) reduction of transcriptional and epigenetic repression of primary micro RNA transcripts b) changes in levels of Argonaute and homologous proteins, c) changes in micro RNA Processing (e.g., from DICER1, etc.), or d) an increased half-life of the micro RNA molecules. However, it remains to be determined if the 511
10 mirna dysregulation described here is also present in brain tissue and therefore playing a key role in the neuropsychiatric phenotype. Finally, the crucial challenge will be to identify and validate the target genes that are affected by those micrornas that are found dysregulated in schizophrenia and to characterize the involved pathways in a much more comprehensive manner in order to improve our understanding of how alterations in microrna mediated genetic networks can contribute to the pathophysiology of the disorder. MiR-let-7d suppresses neural stem cell proliferation by reducing TLX expression in neural stem cells, which may be a novel strategy for identifying potential interventions in relevant neurological diseases (Zhao et al., 2013). References Bartel, D.P MicroRNAs: target recognition and regulatory functions. Cell, 136: Gardiner, E., Beveridge, N.J., Wu, J.Q., Carr, V., Scott, R.J., Tooney, P.A., Cairns, M.J Imprinted DLK1-DIO3 region of 14q32 defines a schizophrenia-associated mirna signature in peripheral blood mononuclear cells. Mol. Psychiatry, 17: Herson, M Etiological considerations Adult psychopathology and diagnosis John Wiley & Sons ISBN Ivanov, H., Stoyanova, V., Popov, N., Vachev, T. Biodiscovery Blood-Based Gene Expression in children with Autism spectrum disorder; 17: 2; DOI: /BioDiscovery Lai, C.Y., Yu, S.L., Hsieh, M.H., Chen, C.H., Chen, H.Y., et al MicroRNA expression aberration as potential peripheral blood biomarkers for schizophrenia. PLoS One, 6: e Mellios, N., Sur, M The Emerging Role of micrornas in Schizophrenia and Autism Spectrum Disorders. Front Psychiatry, 3: 39. Mirnics Károly, Frank, A., Middleton, Adriana Marquez, David, A., Lewis. Molecular Characterization of Schizophrenia Viewed by Microarray Analysis of Gene Expression in Prefrontal Cortex Pat Levitt DOI: Perkins, D.O., Jeffries, C.D.,.Jarskog, L.F., et al microrna expression in the prefrontal cortex of individuals with schizophrenia and schizoaffective disorder. Genome Biol., 8(2):R27. doi: /gb r27 Scholer, N., Langer, C., Dohner, H., Buske, C., Kuchenbauer, F Serum micrornas as a novel class of biomarkers: a comprehensive review of the literature. Exp. Hematol., 38(12): doi: /j.exphem PMID: Zhao, C., Sun, G., Ye, P,. Li, S., Shi, Y MicroRNA let-7d regulates the TLX/microRNA-9 cascade to control neural cell fate and neurogenesis. Sci. Rep., 3: doi: /srep01329 PMID: How to cite this article: Tihomir I. Vachev, Nikolay T. Popov, Danail Marchev, Hristo Ivanov and Vili K. Stoyanova Characterization of Micro RNA Signature in Peripheral Blood of Schizophrenia Patients using µparaflo TM mirna Microarray Assay. Int.J.Curr.Microbiol.App.Sci. 5(7): doi: 512
devseek (Sequence(Analysis(Panel(for(Neurodevelopmental(Disorders
 ACSL4 Mental/retardation,/XBlinked/63 XLBR ADSL Adenylosuccinase/Deficiency AR AFF2 Mental/retardation,/XBlinked,/FRAXE/type XLBR ALG6 Congenital/disorder/of/glycosylation/type/Ic AR ANK3 Mental/retardation,/autosomal/recessive/37
ACSL4 Mental/retardation,/XBlinked/63 XLBR ADSL Adenylosuccinase/Deficiency AR AFF2 Mental/retardation,/XBlinked,/FRAXE/type XLBR ALG6 Congenital/disorder/of/glycosylation/type/Ic AR ANK3 Mental/retardation,/autosomal/recessive/37
Is genomic grading killing histological grading?
 Is genomic grading killing histological grading? Christos Sotiriou MD PhD Fonds National de Recherche Scientifique (FNRS) Université Libre de Bruxelles (ULB) Institut Jules Bordet Histological Grade and
Is genomic grading killing histological grading? Christos Sotiriou MD PhD Fonds National de Recherche Scientifique (FNRS) Université Libre de Bruxelles (ULB) Institut Jules Bordet Histological Grade and
Integrative Radiation Biology
 Dr. Kristian Unger Integrative Biology Group Research Unit of Radiation Cytogenetics Department of Radiation Sciences Helmholtz-Zentrum München 1 Key Questions Molecular mechanisms of radiation-induced
Dr. Kristian Unger Integrative Biology Group Research Unit of Radiation Cytogenetics Department of Radiation Sciences Helmholtz-Zentrum München 1 Key Questions Molecular mechanisms of radiation-induced
Supplementary Table 1. Information on the 174 single nucleotide variants identified by whole-exome sequencing
 Supplementary Table 1. Information on the 174 single nucleotide s identified by whole-exome sequencing Chr Position Type AF exomes Database 1 10163148 A/G UBE4B missense 0.000326 1534 2 98 1 4.44 Damaging
Supplementary Table 1. Information on the 174 single nucleotide s identified by whole-exome sequencing Chr Position Type AF exomes Database 1 10163148 A/G UBE4B missense 0.000326 1534 2 98 1 4.44 Damaging
Supplementary information: Forstner, Hofmann et al.
 Supplementary information: Forstner, Hofmann et al., Genome-wide analysis implicates micrornas and their target genes in the development of bipolar disorder Table of contents Supplementary Figure 1: Regional
Supplementary information: Forstner, Hofmann et al., Genome-wide analysis implicates micrornas and their target genes in the development of bipolar disorder Table of contents Supplementary Figure 1: Regional
Quantitative Real Time PCR (RT-qPCR) Differentiation of human preadipocytes to adipocytes
 Quantitative Real Time PCR (RT-qPCR) First strand cdna was synthesized and RT-qPCR was performed using RT2 first strand kits (Cat#330401) and RT2 qpcr Master Mix (SABioscience, Frederick, MD). Experiments
Quantitative Real Time PCR (RT-qPCR) First strand cdna was synthesized and RT-qPCR was performed using RT2 first strand kits (Cat#330401) and RT2 qpcr Master Mix (SABioscience, Frederick, MD). Experiments
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature12912 Figure S1. Distribution of mutation rates and spectra across 4,742 tumor-normal (TN) pairs from 21 tumor types, as in Figure 1 from Lawrence et al. Nature
SUPPLEMENTARY INFORMATION doi:1.138/nature12912 Figure S1. Distribution of mutation rates and spectra across 4,742 tumor-normal (TN) pairs from 21 tumor types, as in Figure 1 from Lawrence et al. Nature
Supplementary Material. Table S1. Summary of mapping results
 Supplementary Material Table S1. Summary of mapping results Sample Total reads Mapped (%) HNE0-1 99687516 80786083 (81.0%) HNE0-2 94318720 77047792 (81.7%) HNE0-3 104033900 84348102 (81.1%) HNE15-1 94426598
Supplementary Material Table S1. Summary of mapping results Sample Total reads Mapped (%) HNE0-1 99687516 80786083 (81.0%) HNE0-2 94318720 77047792 (81.7%) HNE0-3 104033900 84348102 (81.1%) HNE15-1 94426598
WERNER_FIBRO_DN LEI_MYB_REGULATED_GENES BRCA1_OVEREXP_DN SUBCLASSES_DIFF P21_P53_MIDDLE_DN
 Supplemental Table S1. Full List of Up-Regulated Conserved Biological Modules Between Zebrafish Liver Tumor with Human Liver, Lung, Gastric, and Prostate Tumors LvCMs (ZF liver cancer vs. human liver cancers
Supplemental Table S1. Full List of Up-Regulated Conserved Biological Modules Between Zebrafish Liver Tumor with Human Liver, Lung, Gastric, and Prostate Tumors LvCMs (ZF liver cancer vs. human liver cancers
Supplementary Table 1. Candidate loci that were associated with diabetes- and/or obesity-related traits.
 Supplementary Table 1. Candidate loci that were associated with diabetes- and/or obesity-related traits. All With Fst in the With long With SNP(s) at the 5' top 5% bracket haplotypes regulatory region
Supplementary Table 1. Candidate loci that were associated with diabetes- and/or obesity-related traits. All With Fst in the With long With SNP(s) at the 5' top 5% bracket haplotypes regulatory region
Jocelyn Chapman, MD Division of Gynecologic Oncology. Julie S. Mak, MS, MSc Genetic Counselor Hereditary Cancer Clinic
 Jocelyn Chapman, MD Division of Gynecologic Oncology Julie S. Mak, MS, MSc Genetic Counselor Hereditary Cancer Clinic Genetics underlies all cancers Somatic or tumor genetics Germline or inherited genetics
Jocelyn Chapman, MD Division of Gynecologic Oncology Julie S. Mak, MS, MSc Genetic Counselor Hereditary Cancer Clinic Genetics underlies all cancers Somatic or tumor genetics Germline or inherited genetics
WT-TP53 DLBCL WT-TP53 GCB-DLBCL
 SUPPLEMENTAL TABLES AND FIGURES Supplemental Table S1. Summary of gene expression profiling analysis results listing counts of significant differentially expressed transcripts between defined two groups
SUPPLEMENTAL TABLES AND FIGURES Supplemental Table S1. Summary of gene expression profiling analysis results listing counts of significant differentially expressed transcripts between defined two groups
Moyamoya disease susceptibility gene RNF213 links inflammatory and angiogenic
 Supplementary Information for Moyamoya disease susceptibility gene RNF213 links inflammatory and angiogenic signals in endothelial cells Kazuhiro Ohkubo 1,, Yasunari Sakai 1,*,, Hirosuke Inoue 1, Satoshi
Supplementary Information for Moyamoya disease susceptibility gene RNF213 links inflammatory and angiogenic signals in endothelial cells Kazuhiro Ohkubo 1,, Yasunari Sakai 1,*,, Hirosuke Inoue 1, Satoshi
Supplementary information. Dual targeting of ANGPT1 and TGFBR2 genes by mir-204 controls
 Supplementary information Dual targeting of ANGPT1 and TGFBR2 genes by mir-204 controls angiogenesis in breast cancer Ali Flores-Pérez, Laurence A. Marchat, Sergio Rodríguez-Cuevas, Verónica Bautista-Piña,
Supplementary information Dual targeting of ANGPT1 and TGFBR2 genes by mir-204 controls angiogenesis in breast cancer Ali Flores-Pérez, Laurence A. Marchat, Sergio Rodríguez-Cuevas, Verónica Bautista-Piña,
SALSA MLPA Probemix P014-B1 Chromosome 8 Lot B and B
 SALSA MLPA Probemix P014-B1 Chromosome 8 Lot B1-0916 and B1-0713. Copy number changes of the human chromosome 8 are common in many types of tumours. In most cases, losses of 8p sequences and gains of 8q
SALSA MLPA Probemix P014-B1 Chromosome 8 Lot B1-0916 and B1-0713. Copy number changes of the human chromosome 8 are common in many types of tumours. In most cases, losses of 8p sequences and gains of 8q
Table S1. CDE Sequences and Globin Reporter mrna Half-Lives in NIH3T3 Cells, Related to Figures 1 and 2 Construct Sequence t1/2 ± SD (hrs)
 Table S1. CDE Sequences and Globin Reporter mrna Half-Lives in NIH3T3 Cells, Related to Figures 1 and 2 Construct Sequence t 1/2 ± SD (hrs) n TNF -CDE 37 -V2 AGATCCTTCAGACAGACATGTTTTCTGTGAAAACGGAGCTGAGCTAGATCT
Table S1. CDE Sequences and Globin Reporter mrna Half-Lives in NIH3T3 Cells, Related to Figures 1 and 2 Construct Sequence t 1/2 ± SD (hrs) n TNF -CDE 37 -V2 AGATCCTTCAGACAGACATGTTTTCTGTGAAAACGGAGCTGAGCTAGATCT
Lions and Tigers and Bears.Oh My!
 Lions and Tigers and Bears.Oh My! TAKING THE FEAR OUT OF GENETIC TESTING Lisa Butterfield, MS, CGC Genetic Counselor Clinical Assistant Professor University of Kansas Health System Why is Genetic Testing
Lions and Tigers and Bears.Oh My! TAKING THE FEAR OUT OF GENETIC TESTING Lisa Butterfield, MS, CGC Genetic Counselor Clinical Assistant Professor University of Kansas Health System Why is Genetic Testing
Leukemia reconstitution in vivo is driven by cells in early cell cycle and low metabolic state
 Published Ahead of Print on March 8, 2018, as doi:10.3324/haematol.2017.167502. Copyright 2018 Ferrata Storti Foundation. Leukemia reconstitution in vivo is driven by cells in early cell cycle and low
Published Ahead of Print on March 8, 2018, as doi:10.3324/haematol.2017.167502. Copyright 2018 Ferrata Storti Foundation. Leukemia reconstitution in vivo is driven by cells in early cell cycle and low
Targeted Agent and Profiling Utilization Registry (TAPUR ) Study. February 2018
 Targeted Agent and Profiling Utilization Registry (TAPUR ) Study February 2018 Precision Medicine Therapies designed to target the molecular alteration that aids cancer development 30 TARGET gene alterations
Targeted Agent and Profiling Utilization Registry (TAPUR ) Study February 2018 Precision Medicine Therapies designed to target the molecular alteration that aids cancer development 30 TARGET gene alterations
Targeted Genes and Methodology Details for Epilepsy/Seizure Genetic Panels
 Targeted s and Methodology Details for Epilepsy/Seizure tic Panels Reference transcripts based on build GRCh37 (hg19) interrogated by Epilepsy/Seizure tic Panels Epilepsy Expanded Panel Epilepsy Expanded
Targeted s and Methodology Details for Epilepsy/Seizure tic Panels Reference transcripts based on build GRCh37 (hg19) interrogated by Epilepsy/Seizure tic Panels Epilepsy Expanded Panel Epilepsy Expanded
Sample Metrics. Allele Frequency (%) Read Depth Ploidy. Gene CDS Effect Protein Effect. LN Metastasis Tumor Purity Computational Pathology 80% 60%
 Supplemental Table 1: Estimated tumor purity, allele frequency, and independent read depth for all gene mutations classified as either potentially pathogenic or VUS in the metatastic and primary tumor
Supplemental Table 1: Estimated tumor purity, allele frequency, and independent read depth for all gene mutations classified as either potentially pathogenic or VUS in the metatastic and primary tumor
Supplemental Table 1. List of genes and mature shrna sequences identified from the shrna screen in BRCA2-mutant PEO1 cells.
 Guillemette_Supplemental Table 1 Gene Mature Sequence Gene Mature Sequence ATTCATAGGATGTCAGCAG LOC157193 TATGCGTAAGTAATAAATTGCT TTAGTTCTGTCTTGGAGG LOC152225 CGCATATATGTTCTGACTT ALDH2 CACTTCAGTGTATGCCT
Guillemette_Supplemental Table 1 Gene Mature Sequence Gene Mature Sequence ATTCATAGGATGTCAGCAG LOC157193 TATGCGTAAGTAATAAATTGCT TTAGTTCTGTCTTGGAGG LOC152225 CGCATATATGTTCTGACTT ALDH2 CACTTCAGTGTATGCCT
Supplementary Materials
 Supplementary Materials Using Multi objective Optimization to Identify Dynamical Network Biomarkers as Early warning Signals of Complex Diseases Fatemeh Vafaee 1, 2 1 Charles Perkins Centre, University
Supplementary Materials Using Multi objective Optimization to Identify Dynamical Network Biomarkers as Early warning Signals of Complex Diseases Fatemeh Vafaee 1, 2 1 Charles Perkins Centre, University
Supplementary Table 1. mirna modulated in D HF and ND HF patients. Table shows in a linear scale the same values of Fig. 2.
 Supplementary Table 1. mirna modulated in D HF and ND HF patients. Table shows in a linear scale the same values of Fig. 2. mir D HF VS CTR ND HF VS CTR D HF VS ND HF FOLD CHANGE p FOLD CHANGE p FOLD CHANGE
Supplementary Table 1. mirna modulated in D HF and ND HF patients. Table shows in a linear scale the same values of Fig. 2. mir D HF VS CTR ND HF VS CTR D HF VS ND HF FOLD CHANGE p FOLD CHANGE p FOLD CHANGE
Supplementary Information
 Supplementary Information Index of Supplementary Materials Supplementary Figure. Distribution of methylation levels, characteristics of cis-meqtls and association plots for meqtls in methylation-related
Supplementary Information Index of Supplementary Materials Supplementary Figure. Distribution of methylation levels, characteristics of cis-meqtls and association plots for meqtls in methylation-related
Table S2. Functional annotation of the downregulated, differentially expressed genes of Rasless MEFs.
 Table S2. Functional annotation of the downregulated, differentially expressed genes of Rasless MEFs. The GeneCodis (Gene Annotation Co-occurrence Discovery) functional annotation tool (http://genecodis.dacya.ucm.es)
Table S2. Functional annotation of the downregulated, differentially expressed genes of Rasless MEFs. The GeneCodis (Gene Annotation Co-occurrence Discovery) functional annotation tool (http://genecodis.dacya.ucm.es)
Provide your cancer patients personalized treatment options with ClariFind
 ONCOLOGY ClariFind employs next-generation sequencing (NGS) for comprehensive genomic profiling of a patient s tumor sample to identify potential clinically actionable targets and associated therapies.
ONCOLOGY ClariFind employs next-generation sequencing (NGS) for comprehensive genomic profiling of a patient s tumor sample to identify potential clinically actionable targets and associated therapies.
Comparison of prognostic signatures for ER positive breast cancer in TransATAC:
 Comparison of prognostic signatures for ER positive breast cancer in TransATAC: EndoPredict, a high performance test in node negative and node positive disease Ivana Sestak, PhD Centre for Cancer Prevention
Comparison of prognostic signatures for ER positive breast cancer in TransATAC: EndoPredict, a high performance test in node negative and node positive disease Ivana Sestak, PhD Centre for Cancer Prevention
Table SІ. List of genes upregulated by IL-4/IL-13-treatment of NHEK
 Supplemental legends Table SІ. List of genes upregulated by IL-4/IL-13-treatment of NHEK Table SП. List of genes in cluster 1 identified using two-dimensional hierarchical clustering analysis Table SШ.
Supplemental legends Table SІ. List of genes upregulated by IL-4/IL-13-treatment of NHEK Table SП. List of genes in cluster 1 identified using two-dimensional hierarchical clustering analysis Table SШ.
The Center for PERSONALIZED DIAGNOSTICS
 The Center for PERSONALIZED DIAGNOSTICS Precision Diagnostics for Personalized Medicine A joint initiative between The Department of Pathology and Laboratory Medicine & The Abramson Cancer Center The (CPD)
The Center for PERSONALIZED DIAGNOSTICS Precision Diagnostics for Personalized Medicine A joint initiative between The Department of Pathology and Laboratory Medicine & The Abramson Cancer Center The (CPD)
Supplement Results, Figures and Tables
 Supplement Results, Figures and Tables Results PET imaging The SUV was significantly higher in tumours of the parental HSC-3 cell line than tumours of the EV and the MMP-8 overexpressing cell lines (Figure
Supplement Results, Figures and Tables Results PET imaging The SUV was significantly higher in tumours of the parental HSC-3 cell line than tumours of the EV and the MMP-8 overexpressing cell lines (Figure
Table S1 A. Primers for validation of GABP binding at select gene loci using ChIP
 Table S1 A. Primers for validation of GABP binding at select gene loci using ChIP Gene symbols Hprt1 Bcl2 Bcl2l1 Mcl1 Zfx Etv6 Smad4 Foxo3 Gabpa Gabpb1 Atm Terf2 Pten Dnmt1 Ep300 Myst4 Smarca2 Smarca4
Table S1 A. Primers for validation of GABP binding at select gene loci using ChIP Gene symbols Hprt1 Bcl2 Bcl2l1 Mcl1 Zfx Etv6 Smad4 Foxo3 Gabpa Gabpb1 Atm Terf2 Pten Dnmt1 Ep300 Myst4 Smarca2 Smarca4
Plasma-Seq conducted with blood from male individuals without cancer.
 Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
A genome-wide association study identifies vitiligo
 A genome-wide association study identifies vitiligo susceptibility loci at MHC and 6q27 Supplementary Materials Index Supplementary Figure 1 The principal components analysis (PCA) of 2,546 GWAS samples
A genome-wide association study identifies vitiligo susceptibility loci at MHC and 6q27 Supplementary Materials Index Supplementary Figure 1 The principal components analysis (PCA) of 2,546 GWAS samples
Nature Genetics: doi: /ng Supplementary Figure 1. Depths and coverages in whole-exome and targeted deep sequencing data.
 Supplementary Figure 1 Depths and coverages in whole-exome and targeted deep sequencing data. Depth (top) and coverage (bottom) of whole-exome sequencing for 38 independent JPN cases (mean depth = 130)
Supplementary Figure 1 Depths and coverages in whole-exome and targeted deep sequencing data. Depth (top) and coverage (bottom) of whole-exome sequencing for 38 independent JPN cases (mean depth = 130)
Precision Oncology: Experience at UW
 Precision Oncology: Experience at UW Colin Pritchard MD, PhD University of Washington, Department of Lab Medicine WSMOS Meeting November 1, 2013 Conflict of Interest Disclosures I declare the following,
Precision Oncology: Experience at UW Colin Pritchard MD, PhD University of Washington, Department of Lab Medicine WSMOS Meeting November 1, 2013 Conflict of Interest Disclosures I declare the following,
Supplementary Information to
 Supplementary Information to Exome sequencing to identify de novo mutations in sporadic ALS trios Alessandra Chesi 1,11, Brett T. Staahl 2, Ana Jovicic 1, Julien Couthouis 1, Maria Fasolino 3, Alya R.
Supplementary Information to Exome sequencing to identify de novo mutations in sporadic ALS trios Alessandra Chesi 1,11, Brett T. Staahl 2, Ana Jovicic 1, Julien Couthouis 1, Maria Fasolino 3, Alya R.
# 1 India s Best Selling Cancer Genomics Test Precision treatment for every Cancer Patient Patient Guide
 # 1 India s Best Selling Cancer Genomics Test Precision treatment for every Cancer Patient Patient Guide PositiveSelect is India s widely used Cancer Genomics test with every 8 out of 10 patients opting
# 1 India s Best Selling Cancer Genomics Test Precision treatment for every Cancer Patient Patient Guide PositiveSelect is India s widely used Cancer Genomics test with every 8 out of 10 patients opting
SI Table. SI Table 3. SI Table 4. SI Table 5A. SI Table 5B. SI Table 5C. SI Table 6A. SI Table 6B. SI Table 6C
 SI Table SI Table 1 SI Table 2 SI Table 3 SI Table 4 SI Table 5A SI Table 5B SI Table 5C SI Table 6A SI Table 6B SI Table 6C Title List of primers used to clone HCV genes and gene fragments HCV genotype
SI Table SI Table 1 SI Table 2 SI Table 3 SI Table 4 SI Table 5A SI Table 5B SI Table 5C SI Table 6A SI Table 6B SI Table 6C Title List of primers used to clone HCV genes and gene fragments HCV genotype
Table S2. List of the genes upregulated in both CitK-/- and Magoh +/- developing neural tissue. Implicated in cell cycle and mitosis control
 Supplementary tables Table S2 List of the genes upregulated in both CitK-/- and Magoh +/- developing neural tissue. Gene Name Gene ID Implicated as p53 target gene Implicated in cell cycle and mitosis
Supplementary tables Table S2 List of the genes upregulated in both CitK-/- and Magoh +/- developing neural tissue. Gene Name Gene ID Implicated as p53 target gene Implicated in cell cycle and mitosis
SUPPLEMENTAL MATERIAL. Characterization and discovery of novel mirnas and mornas in JAK2V617F. mutated SET2 cells
 SUPPLEMENTAL MATERIAL Characterization and discovery of novel mirnas and mornas in JAK2V617F mutated SET2 cells Stefania Bortoluzzi 1*, Andrea Bisognin 1*, et al. 1 Supplemental methods Short RNA library
SUPPLEMENTAL MATERIAL Characterization and discovery of novel mirnas and mornas in JAK2V617F mutated SET2 cells Stefania Bortoluzzi 1*, Andrea Bisognin 1*, et al. 1 Supplemental methods Short RNA library
Jennifer Hauenstein Oncology Cytogenetics Emory University Hospital Atlanta, GA
 Comparison of Genomic Coverage using Affymetrix OncoScan Array and Illumina TruSight Tumor 170 NGS Panel for Detection of Copy Number Abnormalities in Clinical GBM Specimens Jennifer Hauenstein Oncology
Comparison of Genomic Coverage using Affymetrix OncoScan Array and Illumina TruSight Tumor 170 NGS Panel for Detection of Copy Number Abnormalities in Clinical GBM Specimens Jennifer Hauenstein Oncology
Supplementary Table 1: List of the 242 hypoxia/reoxygenation marker genes collected from literature and databases
 Supplementary Tables: Supplementary Table 1: List of the 242 hypoxia/reoxygenation marker genes collected from literature and databases Supplementary Table 2: Contingency tables used in the fisher exact
Supplementary Tables: Supplementary Table 1: List of the 242 hypoxia/reoxygenation marker genes collected from literature and databases Supplementary Table 2: Contingency tables used in the fisher exact
Supplementary Figure 1. Microarray data mining and validation
 Supplementary Figure 1. Microarray data mining and validation Microarray (>47,000 probes) Raw data Background subtraction NanoString (124 genes) Set cut-off and threshold values Raw data Normalized to
Supplementary Figure 1. Microarray data mining and validation Microarray (>47,000 probes) Raw data Background subtraction NanoString (124 genes) Set cut-off and threshold values Raw data Normalized to
Assessing the Economics of Genomic Medicine. Kenneth Offit, MD MPH Chief, Clinical Genetics Service Memorial Sloan-Kettering Cancer Center
 Assessing the Economics of Genomic Medicine Kenneth Offit, MD MPH Chief, Clinical Genetics Service Memorial Sloan-Kettering Cancer Center DISCLOSURES No conflicts Patents unenforced Consultancies unpaid
Assessing the Economics of Genomic Medicine Kenneth Offit, MD MPH Chief, Clinical Genetics Service Memorial Sloan-Kettering Cancer Center DISCLOSURES No conflicts Patents unenforced Consultancies unpaid
Supplemental Tables and Figures
 Supplemental Tables and Figures Post-zygotic de novo changes in glutamate and dopamine pathways may explain discordance of monozygotic twins for schizophrenia Castellani, CA., Melka, MG., Gui, JL., Gallo,
Supplemental Tables and Figures Post-zygotic de novo changes in glutamate and dopamine pathways may explain discordance of monozygotic twins for schizophrenia Castellani, CA., Melka, MG., Gui, JL., Gallo,
Copyright Warning & Restrictions
 Copyright Warning & Restrictions The copyright law of the United States (Title 17, United States Code) governs the making of photocopies or other reproductions of copyrighted material. Under certain conditions
Copyright Warning & Restrictions The copyright law of the United States (Title 17, United States Code) governs the making of photocopies or other reproductions of copyrighted material. Under certain conditions
Palindromic amplification of the ERBB2 oncogene in human primary breast tumors. A common pattern of copy number transition of chromosome 17 in HER2-
 Supplemental Figures Table of Contents Palindromic amplification of the ERBB2 oncogene in human primary breast tumors Michael Marotta 1,5, Taku Onodera 3,5, Jeffrey Johnson 4, Thomas Budd 2, Takaaki Watanabe
Supplemental Figures Table of Contents Palindromic amplification of the ERBB2 oncogene in human primary breast tumors Michael Marotta 1,5, Taku Onodera 3,5, Jeffrey Johnson 4, Thomas Budd 2, Takaaki Watanabe
Supplementary Figures. Supplementary Figure 1. Dinucleotide variant proportions. These are described and quantitated. for each lesion type.
 Supplementary Figures Supplementary Figure 1. Dinucleotide variant proportions. These are described and quantitated for each lesion type. a b Supplementary Figure 2. Non-negative matrix factorization-derived
Supplementary Figures Supplementary Figure 1. Dinucleotide variant proportions. These are described and quantitated for each lesion type. a b Supplementary Figure 2. Non-negative matrix factorization-derived
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature11986 relative IL-6 expression Viable intracellular Bp per well 18 16 1 1 5 5 3 1 6 +DG 8 8 8 Control DG Time (h) hours B. pertussis IL-6 (pg/ml) 15 Control DG
SUPPLEMENTARY INFORMATION doi:1.138/nature11986 relative IL-6 expression Viable intracellular Bp per well 18 16 1 1 5 5 3 1 6 +DG 8 8 8 Control DG Time (h) hours B. pertussis IL-6 (pg/ml) 15 Control DG
RNA sequencing of cancer reveals novel splicing alterations
 RNA sequencing of cancer reveals novel splicing alterations Jeyanthy Eswaran, Anelia Horvath, Sucheta Godbole, Sirigiri Divijendra Reddy, Prakriti Mudvari, Kazufumi Ohshiro, Dinesh Cyanam, Sujit Nair,
RNA sequencing of cancer reveals novel splicing alterations Jeyanthy Eswaran, Anelia Horvath, Sucheta Godbole, Sirigiri Divijendra Reddy, Prakriti Mudvari, Kazufumi Ohshiro, Dinesh Cyanam, Sujit Nair,
HO-1 inhibits preadipocyte proliferation and differentiation at the onset of obesity via ROS dependent activation of Akt2
 SUPPLEMENTAL INFORMATION referring to: HO-1 inhibits preadipocyte proliferation and differentiation at the onset of obesity via ROS dependent activation of Akt2 Gabriel Wagner 1, Josefine Lindroos-Christensen
SUPPLEMENTAL INFORMATION referring to: HO-1 inhibits preadipocyte proliferation and differentiation at the onset of obesity via ROS dependent activation of Akt2 Gabriel Wagner 1, Josefine Lindroos-Christensen
Changing the Culture of Cancer Care II. Eric Holland Fred Hutchinson Cancer Research Center University of Washington Seattle
 Changing the Culture of Cancer Care II Eric Holland Fred Hutchinson Cancer Research Center University of Washington Seattle Transforming Cancer Therapy Eric Holland Fred Hutchinson Cancer Research Center
Changing the Culture of Cancer Care II Eric Holland Fred Hutchinson Cancer Research Center University of Washington Seattle Transforming Cancer Therapy Eric Holland Fred Hutchinson Cancer Research Center
Regulation of eif4 and p70s6k signaling. Chondroitin and dermatan biosynthesis Interferon signaling
 Supplemental Information Table S1: Differentially regulated pathways in cortical versus cerebellar human astrocytes following treatment with PBS or IFN-β Pathway Regulation of eif4 and p70s6k signaling
Supplemental Information Table S1: Differentially regulated pathways in cortical versus cerebellar human astrocytes following treatment with PBS or IFN-β Pathway Regulation of eif4 and p70s6k signaling
HTG EdgeSeq Immuno-Oncology Assay Gene List
 A2M ABCB1 ABCB11 ABCC2 ABCG2 ABL1 ABL2 ACTB ADA ADAM17 ADGRE5 ADORA2A AICDA AKT3 ALCAM ALO5 ANA1 APAF1 APP ATF1 ATF2 ATG12 ATG16L1 ATG5 ATG7 ATM ATP5F1 AL BATF BA BCL10 BCL2 BCL2L1 BCL6 BID BIRC5 BLNK
A2M ABCB1 ABCB11 ABCC2 ABCG2 ABL1 ABL2 ACTB ADA ADAM17 ADGRE5 ADORA2A AICDA AKT3 ALCAM ALO5 ANA1 APAF1 APP ATF1 ATF2 ATG12 ATG16L1 ATG5 ATG7 ATM ATP5F1 AL BATF BA BCL10 BCL2 BCL2L1 BCL6 BID BIRC5 BLNK
p5+mir30: AATGATACGGCGACCACCGACTAAAGTAGCCCCTTGAATTC
 SUPPORTING INFORMATION METHODS Massively parallel sequencing and shrna screen data analysis pipeline Genomic DNA extraction and purification from surviving MCF7 cells in the genome wide screen was carried
SUPPORTING INFORMATION METHODS Massively parallel sequencing and shrna screen data analysis pipeline Genomic DNA extraction and purification from surviving MCF7 cells in the genome wide screen was carried
Supplementary Table 2b. List of spliced oncogenes and tumor suppressors identified in AML patients compared to normal donors
 1 Supplementary Table 2b. List of spliced oncogenes and tumor suppressors identified in AML patients compared to normal donors Gene Symbol Gene Name / Tumor Suppressors AKT1 v-akt murine thymoma viral
1 Supplementary Table 2b. List of spliced oncogenes and tumor suppressors identified in AML patients compared to normal donors Gene Symbol Gene Name / Tumor Suppressors AKT1 v-akt murine thymoma viral
Illumina s Cancer Research Portfolio and Dedicated Workflows
 Illumina s Cancer Research Portfolio and Dedicated Workflows Michael Sohn Clinical Sales Specialist Spain&Italy 2017 2017 Illumina, Inc. All rights reserved. Illumina, 24sure, BaseSpace, BeadArray, BlueFish,
Illumina s Cancer Research Portfolio and Dedicated Workflows Michael Sohn Clinical Sales Specialist Spain&Italy 2017 2017 Illumina, Inc. All rights reserved. Illumina, 24sure, BaseSpace, BeadArray, BlueFish,
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
Supplementary Information. PRC2 overexpression and PRC2-target gene repression relating to poorer prognosis in small
 Sato T, et al. Supplementary Information PRC2 overexpression and PRC2-target gene repression relating to poorer prognosis in small cell lung cancer Teruyuki Sato 1,2, Atsushi Kaneda 1,3,4 *, Shingo Tsuji
Sato T, et al. Supplementary Information PRC2 overexpression and PRC2-target gene repression relating to poorer prognosis in small cell lung cancer Teruyuki Sato 1,2, Atsushi Kaneda 1,3,4 *, Shingo Tsuji
SCIENCE CHINA Life Sciences
 SCIENCE CHINA Life Sciences RESEARCH PAPER April 2013 Vol.56 No.4: 1 7 doi: 10.1007/s11427-013-4460-x Table S1 Human paired-end RNA-Seq data sets from the Sequence Read Archive (SRA) database Experiment
SCIENCE CHINA Life Sciences RESEARCH PAPER April 2013 Vol.56 No.4: 1 7 doi: 10.1007/s11427-013-4460-x Table S1 Human paired-end RNA-Seq data sets from the Sequence Read Archive (SRA) database Experiment
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature14122 Supplementary Table 1: EZH2 Co-Expression Gene Signature Top 116 genes co-expressed with EZH2 across 9 Oncomine studies Gene Description Symbol ALS2CR4
SUPPLEMENTARY INFORMATION doi:10.1038/nature14122 Supplementary Table 1: EZH2 Co-Expression Gene Signature Top 116 genes co-expressed with EZH2 across 9 Oncomine studies Gene Description Symbol ALS2CR4
Supplemental Table 1: Enriched GO categories per cluster
 Supplemental Table 1: Enriched GO categories per cluster Cluster Enriched with #genes Raw p-value Corrected p-value Frequency in set (%) Cluster 1 immune response - GO:0006955 32 2.36E-14 0.001 13.5 Cluster
Supplemental Table 1: Enriched GO categories per cluster Cluster Enriched with #genes Raw p-value Corrected p-value Frequency in set (%) Cluster 1 immune response - GO:0006955 32 2.36E-14 0.001 13.5 Cluster
Analyzing the NCI-60 Cancer Cell Lines using Data Obtained from Genome-Wide ChIP-X Experiments. Jayanth (Jay) Krishnan. Mahopac High School
 1 Analyzing the NCI-60 Cancer Cell Lines using Data Obtained from Genome-Wide ChIP-X Experiments Jayanth (Jay) Krishnan Mahopac High School January 25, 2010 2 Analyzing the NCI-60 Cancer Cell Lines using
1 Analyzing the NCI-60 Cancer Cell Lines using Data Obtained from Genome-Wide ChIP-X Experiments Jayanth (Jay) Krishnan Mahopac High School January 25, 2010 2 Analyzing the NCI-60 Cancer Cell Lines using
Exons from 137 genes were targeted for hybrid selection and massively parallel sequencing.
 SUPPLEMENTARY TABLES Supplementary Table 1. Targeted Genes ABL1 ABL2 AKT1 AKT2 AKT3 ALK APC ATM AURKA BCL2 BRAF BRCA1 BRCA2 CCND1 CCNE1 CDC73 CDH1 CDK4 CDK6 CDK8 CDKN1A CDKN2A CEBPA CHEK1 CHEK2 CREBBP
SUPPLEMENTARY TABLES Supplementary Table 1. Targeted Genes ABL1 ABL2 AKT1 AKT2 AKT3 ALK APC ATM AURKA BCL2 BRAF BRCA1 BRCA2 CCND1 CCNE1 CDC73 CDH1 CDK4 CDK6 CDK8 CDKN1A CDKN2A CEBPA CHEK1 CHEK2 CREBBP
Subject ID: Date Test Started: Specimen Type: Peripheral Blood Date of Report: Date Specimen Obtained:
 TEST PERFORMED Whole exome sequencing (WES) with Focused Diagnostic Panel(s): Neuropathy, Leukodystrophy, Myopathy See below for a complete list of genes analyzed. RESULTS 1 sequence variant with potential
TEST PERFORMED Whole exome sequencing (WES) with Focused Diagnostic Panel(s): Neuropathy, Leukodystrophy, Myopathy See below for a complete list of genes analyzed. RESULTS 1 sequence variant with potential
Supplemental Figures/Tables. Methylation forward GTCGGGGCGTATTTAGTTC. Methylation reverse AACGACGTAAACGAAAATATCG
 Supplemental Figures/Tables Supplementary Tables Table S1. Primer sequences MSP Primers SPARC2 Unmethylation forward TTTTTTAGATTGTTTGGAGAGTG Unmethylation reverse AACTAACAACATAAACAAAAATATC Methylation
Supplemental Figures/Tables Supplementary Tables Table S1. Primer sequences MSP Primers SPARC2 Unmethylation forward TTTTTTAGATTGTTTGGAGAGTG Unmethylation reverse AACTAACAACATAAACAAAAATATC Methylation
Primer sequence CGGAAAGCGGCGGGGTGTTGGGGGAGGCGAGGAACAACCGAGGGGAAGC TTTCCTGATACCCATCCTACACAGGATGACGACGATAAGTA
 Supplemental Table 1. Sequences of the primers used for generation of H1/H6 virus primer mirna H1/6 kan F mirna H1/6 kan R mirna H1/6 zeo F mirna H1/6 zeo R Primer sequence CGGAAAGCGGCGGGGTGTTGGGGGAGGCGAGGAACAACCGAGGGGAAGC
Supplemental Table 1. Sequences of the primers used for generation of H1/H6 virus primer mirna H1/6 kan F mirna H1/6 kan R mirna H1/6 zeo F mirna H1/6 zeo R Primer sequence CGGAAAGCGGCGGGGTGTTGGGGGAGGCGAGGAACAACCGAGGGGAAGC
U118MG. Supplementary Figure 1 U373MG U118MG 3.5 A CCF-SSTG
 A172 CCF-SSTG1 15 - - - 1 1 1 2 2 3 4 6 7 8 10101112131617192022-1 1 1 2 2 3 4 6 7 8 10 10 11 12 13 16 17 19 20 22 T98G U373MG - - - 1 1 1 2 2 3 4 6 7 8 10 10 11 12 13 16 17 19 20 22-1 1 1 2 2 3 4 6 7
A172 CCF-SSTG1 15 - - - 1 1 1 2 2 3 4 6 7 8 10101112131617192022-1 1 1 2 2 3 4 6 7 8 10 10 11 12 13 16 17 19 20 22 T98G U373MG - - - 1 1 1 2 2 3 4 6 7 8 10 10 11 12 13 16 17 19 20 22-1 1 1 2 2 3 4 6 7
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/9/e1600220/dc1 Supplementary Materials for Portraying breast cancers with long noncoding RNAs Olivier Van Grembergen, Martin Bizet, Eric J. de Bony, Emilie Calonne,
advances.sciencemag.org/cgi/content/full/2/9/e1600220/dc1 Supplementary Materials for Portraying breast cancers with long noncoding RNAs Olivier Van Grembergen, Martin Bizet, Eric J. de Bony, Emilie Calonne,
I have no conflicts of interest
 Deploying lung cancer molecular pathology guidelines in real life Neal Lindeman, MD Director, Molecular Pathology Brigham & Women s Hospital USCAP Annual Meeting AMP Companion Meeting San Antonio, T Sunday,
Deploying lung cancer molecular pathology guidelines in real life Neal Lindeman, MD Director, Molecular Pathology Brigham & Women s Hospital USCAP Annual Meeting AMP Companion Meeting San Antonio, T Sunday,
Supplementary Table S1. Histopathologic findings for multi-region analysis.
 SUPPLEMENTARY INFORMATION SUPPLEMENTARY TABLES Supplementary Table S1. Histopathologic findings for multi-region analysis. Patient Histology R1: clear cell RCC, Fuhrman 2-3, acinar pattern 1 R2: clear
SUPPLEMENTARY INFORMATION SUPPLEMENTARY TABLES Supplementary Table S1. Histopathologic findings for multi-region analysis. Patient Histology R1: clear cell RCC, Fuhrman 2-3, acinar pattern 1 R2: clear
A brain tumor and NGS/multiplexing
 A brain tumor and NGS/multiplexing Santiago Ramón y Cajal Jefe de Servicio. Hospital Vall d Hebron Catedrático de Anatomía Patológica U.A.B. Académico de Número de la Real Academia Nacional de Medicina
A brain tumor and NGS/multiplexing Santiago Ramón y Cajal Jefe de Servicio. Hospital Vall d Hebron Catedrático de Anatomía Patológica U.A.B. Académico de Número de la Real Academia Nacional de Medicina
Expression Profiling of KLF4
 Expression Profiling of KLF4 AJCR0000006 Supplemental Data Figure S1. Snapshot of enriched gene sets identified by GSEA in Klf4-null MEFs. Figure S2. Snapshot of enriched gene sets identified by GSEA in
Expression Profiling of KLF4 AJCR0000006 Supplemental Data Figure S1. Snapshot of enriched gene sets identified by GSEA in Klf4-null MEFs. Figure S2. Snapshot of enriched gene sets identified by GSEA in
A. Complete list of 177 genes overexpressed in replicative senescence
 Table S1. A. Complete list of 177 genes overexpressed in replicative senescence Value Gene Description UniGene RefSeq 2.440 WNT16 wingless-type MMTV integration site family, member 16 (WNT16), transcript
Table S1. A. Complete list of 177 genes overexpressed in replicative senescence Value Gene Description UniGene RefSeq 2.440 WNT16 wingless-type MMTV integration site family, member 16 (WNT16), transcript
Patricia Aoun MD, MPH Professor and Vice-Chair for Clinical Affairs Medical Director, Clinical Laboratories Department of Pathology City of Hope
 Patricia Aoun MD, MPH Professor and Vice-Chair for Clinical Affairs Medical Director, Clinical Laboratories Department of Pathology City of Hope National Medical Center Disclosures I have no disclosures
Patricia Aoun MD, MPH Professor and Vice-Chair for Clinical Affairs Medical Director, Clinical Laboratories Department of Pathology City of Hope National Medical Center Disclosures I have no disclosures
103.5 Membrane metalloendopeptidase TGA GGG GTC ACG ATT TTA GG ATG ATG GTG AGG AGC AGG AC
 Supplementary Table SA Primers used for real-time RT-PCR gene regulation and their confirmation Gene Name Accession no Size Region Forward Primer Reverse Primer Efficiency% AKT v-akt oncogene homolog NM_00563
Supplementary Table SA Primers used for real-time RT-PCR gene regulation and their confirmation Gene Name Accession no Size Region Forward Primer Reverse Primer Efficiency% AKT v-akt oncogene homolog NM_00563
Cell cycle arrest through indirect transcriptional repression by p53: I have a DREAM
 Review (2018) 25, 114 132 Official journal of the Cell Death Differentiation Association www.nature.com/cdd Cell cycle arrest through indirect transcriptional repression by p53: I have a DREAM OPEN Kurt
Review (2018) 25, 114 132 Official journal of the Cell Death Differentiation Association www.nature.com/cdd Cell cycle arrest through indirect transcriptional repression by p53: I have a DREAM OPEN Kurt
Table S1. Demographics of patients and tumor characteristics.
 Supplemental Tables for: A Clinical Actionability of Comprehensive Genomic Profiling for Management of Rare or Refractory Cancers Kim Hirshfield et al. Table S1. Demographics of patients and tumor characteristics.
Supplemental Tables for: A Clinical Actionability of Comprehensive Genomic Profiling for Management of Rare or Refractory Cancers Kim Hirshfield et al. Table S1. Demographics of patients and tumor characteristics.
What Do We Know About Individual Variability and Its
 What Do We Know About Individual Variability and Its Contribution to Disease? Nathaniel Rothman, MD, MPH, MHS Occupational and Environmental Epidemiology Branch, Division of Cancer Epidemiology and Genetics,
What Do We Know About Individual Variability and Its Contribution to Disease? Nathaniel Rothman, MD, MPH, MHS Occupational and Environmental Epidemiology Branch, Division of Cancer Epidemiology and Genetics,
ChromoFic, Chromosome SNP Microarray 750K, High Resolution
 SPECIMEN Peripheral blood ChromoFic, Chromosome SNP Microarray 750K, High Resolution CLINICAL INDICATION Failure to thrive, dysmorphism, triangular face, long philtrum, congenital heart disease, increased
SPECIMEN Peripheral blood ChromoFic, Chromosome SNP Microarray 750K, High Resolution CLINICAL INDICATION Failure to thrive, dysmorphism, triangular face, long philtrum, congenital heart disease, increased
Thyroid Tumor Management in the Era of Precision Oncology. Bryan Haugen, MD Former Fellow
 Thyroid Tumor Management in the Era of Precision Oncology Bryan Haugen, MD Former Fellow Outline/Timeline Disclosures Led Afirma GEC study Writing group Thyroseqv3 study Rosetta Genomics research support
Thyroid Tumor Management in the Era of Precision Oncology Bryan Haugen, MD Former Fellow Outline/Timeline Disclosures Led Afirma GEC study Writing group Thyroseqv3 study Rosetta Genomics research support
Genomic and transcriptomic landscapes of acute promyelocy3c leukemia Kankan Wang
 Genomic and transcriptomic landscapes of acute promyelocy3c leukemia Kankan Wang State Key Laboratory of Medical Genomics Rui-Jin Hospital Shanghai Jiaotong University School of Medicine Understanding
Genomic and transcriptomic landscapes of acute promyelocy3c leukemia Kankan Wang State Key Laboratory of Medical Genomics Rui-Jin Hospital Shanghai Jiaotong University School of Medicine Understanding
Oncology Update: The 10 Most Talked About Breast Cancer Topics of The Top 10 10/24/2013. The Awareness Debate
 Oncology Update: The 10 Most Talked About Breast Cancer Topics of 2013 Michaela Tsai, MD Martha Bacon Stimpson Chair of Breast Oncology Virginia Piper Cancer Institute Minnesota Oncology The Top 10 1.
Oncology Update: The 10 Most Talked About Breast Cancer Topics of 2013 Michaela Tsai, MD Martha Bacon Stimpson Chair of Breast Oncology Virginia Piper Cancer Institute Minnesota Oncology The Top 10 1.
Micro-RNAs in cancer: novel origins and sequence variation. Feng Yu
 Micro-RNAs in cancer: novel origins and sequence variation Feng Yu M.Sc., B.Sc. This thesis is submitted in fulfilment of the requirements for the Doctor of Philosophy School of Medicine The University
Micro-RNAs in cancer: novel origins and sequence variation Feng Yu M.Sc., B.Sc. This thesis is submitted in fulfilment of the requirements for the Doctor of Philosophy School of Medicine The University
Proteome Analysis of Human Perilymph using an Intraoperative Sampling Method
 Supporting Information Article Title Proteome Analysis of Human Perilymph using an Intraoperative Sampling Method Authors Heike A. Schmitt 1,3, Andreas Pich 2, Anke Schröder 2, Verena Scheper 1,3, Giorgio
Supporting Information Article Title Proteome Analysis of Human Perilymph using an Intraoperative Sampling Method Authors Heike A. Schmitt 1,3, Andreas Pich 2, Anke Schröder 2, Verena Scheper 1,3, Giorgio
Whole Exome Sequencing Test for Cancer - EXaCT-1
 Patient ID: PMTEST type: tumor type test Primary site: primary site test Report date: Apr. 11, 2016 Whole Exome Sequencing Test for Cancer - EXaCT-1 CLINICAL INFORMATION Patient ID: Requesting physician:
Patient ID: PMTEST type: tumor type test Primary site: primary site test Report date: Apr. 11, 2016 Whole Exome Sequencing Test for Cancer - EXaCT-1 CLINICAL INFORMATION Patient ID: Requesting physician:
Supplementary Materials for
 Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 217 Supplementary Materials for Chemical genomic analysis of GPR35 signaling Heidi (Haibei)
Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 217 Supplementary Materials for Chemical genomic analysis of GPR35 signaling Heidi (Haibei)
Pushing%PET%Imaging%to%the% Cellular%Level:%Development%of%a% Radioluminescence%Microscope%
 Stanford University School of Medicine Department of Radiation Oncology Division of Radiation Physics Pushing%PET%Imaging%to%the% Cellular%Level:%Development%of%a% Radioluminescence%Microscope%! Guillem!Pratx,!PhD!
Stanford University School of Medicine Department of Radiation Oncology Division of Radiation Physics Pushing%PET%Imaging%to%the% Cellular%Level:%Development%of%a% Radioluminescence%Microscope%! Guillem!Pratx,!PhD!
Editorial Type 2 Diabetes and More Gene Panel: A Predictive Genomics Approach for a Polygenic Disease
 Cronicon OPEN ACCESS EC DIABETES AND METABOLIC RESEARCH Editorial Type 2 Diabetes and More Gene Panel: A Predictive Genomics Approach for a Polygenic Disease Amr TM Saeb* University Diabetes Center, College
Cronicon OPEN ACCESS EC DIABETES AND METABOLIC RESEARCH Editorial Type 2 Diabetes and More Gene Panel: A Predictive Genomics Approach for a Polygenic Disease Amr TM Saeb* University Diabetes Center, College
Nature Immunology: doi: /ni.3745
 Supplementary Figure 1 Experimental settings. (a) Experimental setup and bioinformatic data analysis of transcriptomic changes induced in neonatal and adult monocytes by stimulation with 10 ng/ml LPS for
Supplementary Figure 1 Experimental settings. (a) Experimental setup and bioinformatic data analysis of transcriptomic changes induced in neonatal and adult monocytes by stimulation with 10 ng/ml LPS for
Supplemental Table 1. 3 types of signals that are involved in T cell activation
 Supplemental Table 1. 3 types of signals that are involved in T cell activation Syn-T cells Allo-T cells CD3/28- T cells First signal MHC-TCR-CD3 pathway Not engaged MHC-alloantigen- TCR-CD3 Engaged CD3-Engaged
Supplemental Table 1. 3 types of signals that are involved in T cell activation Syn-T cells Allo-T cells CD3/28- T cells First signal MHC-TCR-CD3 pathway Not engaged MHC-alloantigen- TCR-CD3 Engaged CD3-Engaged
15. Supplementary Figure 9. Predicted gene module expression changes at 24hpi during HIV
 Supplementary Information Table of content 1. Supplementary Table 1. Summary of RNAseq data and mapping statistics 2. Supplementary Table 2. Biological functions enriched in 12 hpi DE genes, derived from
Supplementary Information Table of content 1. Supplementary Table 1. Summary of RNAseq data and mapping statistics 2. Supplementary Table 2. Biological functions enriched in 12 hpi DE genes, derived from
APPLICATIONS OF NEXT GENERATION SEQUENCING IN SOLID TUMORS - PATHOLOGIST PROSPECTIVE
 AMP COMPANION MEETING SYMPOSIUM AT USCAP 2015 NEXT-GENERATION OF PATHOLOGY: ROLE OF PATHOLOGIST IN NGS-BASED PERSONALIZED MEDICINE APPLICATIONS OF NEXT GENERATION SEQUENCING IN SOLID TUMORS - PATHOLOGIST
AMP COMPANION MEETING SYMPOSIUM AT USCAP 2015 NEXT-GENERATION OF PATHOLOGY: ROLE OF PATHOLOGIST IN NGS-BASED PERSONALIZED MEDICINE APPLICATIONS OF NEXT GENERATION SEQUENCING IN SOLID TUMORS - PATHOLOGIST
Inferring the Mammalian Cardiogenic Gene Regulatory Network. Jason Bazil 10/18/13 Cardiac Physiome Workshop 2013
 Inferring the Mammalian Cardiogenic Gene Regulatory Network Jason Bazil 1/18/13 Cardiac Physiome Workshop 213 Reverse Engineering Gene Regulatory Networks Two main questions need addressed: 1) What s connected
Inferring the Mammalian Cardiogenic Gene Regulatory Network Jason Bazil 1/18/13 Cardiac Physiome Workshop 213 Reverse Engineering Gene Regulatory Networks Two main questions need addressed: 1) What s connected
Supporting Information
 Supporting Information Lin et al. 10.1073/pnas.1408759111 Fig. S1. Safety and potential toxicity evaluation of M1 in immunocompetent mice. Immunocompetent BALB/c mice received two doses of i.v. infusion
Supporting Information Lin et al. 10.1073/pnas.1408759111 Fig. S1. Safety and potential toxicity evaluation of M1 in immunocompetent mice. Immunocompetent BALB/c mice received two doses of i.v. infusion
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Frampton et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing Supplementary Figure 1A Base substitution detection
SUPPLEMENTARY INFORMATION Frampton et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing Supplementary Figure 1A Base substitution detection
The International Association for the Study of Lung Cancer (IASLC) Lung Cancer Staging Project, Data Elements
 Page 1 Contents 1.1. Registration... 2 1.2. Patient Characteristics... 3 1.3. Laboratory Values at Diagnosis... 5 1.4. Lung Cancers with Multiple Lesions... 8 1.5. Primary Tumour Description... 10 1.6.
Page 1 Contents 1.1. Registration... 2 1.2. Patient Characteristics... 3 1.3. Laboratory Values at Diagnosis... 5 1.4. Lung Cancers with Multiple Lesions... 8 1.5. Primary Tumour Description... 10 1.6.
Table 1 Demographic data of PD-patients. Data are represented as counts or median and full range.
 Table 1 Demographic data of PD-patients. Data are represented as counts or median and full range. Parameter Number Patients 5 Diabetes mellitus 0 Previous NTX 0 Steroids 0 Patients with residual renal
Table 1 Demographic data of PD-patients. Data are represented as counts or median and full range. Parameter Number Patients 5 Diabetes mellitus 0 Previous NTX 0 Steroids 0 Patients with residual renal
Epigenetic Regulation by Chromatin Activation Mark H3K4me3 in Primate Progenitor
 Epigenetic Regulation by Chromatin Activation Mark H3K4me3 in Primate Progenitor Cells within Adult Neurogenic Niche Richard S. Sandstrom 1, Michael R. Foret 2, Douglas A. Grow 2,3, Eric Haugen 1, Christopher
Epigenetic Regulation by Chromatin Activation Mark H3K4me3 in Primate Progenitor Cells within Adult Neurogenic Niche Richard S. Sandstrom 1, Michael R. Foret 2, Douglas A. Grow 2,3, Eric Haugen 1, Christopher
