Supplementary information. Dual targeting of ANGPT1 and TGFBR2 genes by mir-204 controls
|
|
|
- Charles Patterson
- 5 years ago
- Views:
Transcription
1 Supplementary information Dual targeting of ANGPT1 and TGFBR2 genes by mir-204 controls angiogenesis in breast cancer Ali Flores-Pérez, Laurence A. Marchat, Sergio Rodríguez-Cuevas, Verónica Bautista-Piña, Alfredo Hidalgo-Miranda, Elena Aréchaga Ocampo, Mónica Sierra Martínez, Carlos Palma-Flores, Miguel A. Fonseca-Sánchez, Horacio Astudillo-de la Vega, Erika Ruiz-García, Juan Antonio González-Barrios, Carlos Pérez- Plasencia, María L. Streber, César López-Camarillo
2 mirna expression (TCGA RNA_seq level 3) Supplementary figure 1. Validation of expression of 33 out 54 mirnas detected by TLDAs (discovery cohort) using TCGA data sets (validation cohort) in 776 tumor and normal adjacent matched tissues. In bold are denoted 4 mirnas with discordant expression between the discovery and validation cohort N=776 matched tumor/normal N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T N T
3 Supplementary figure 2. Restoration of mir-204 expression in MDA-MB -231 breast cancer cell line MDA-MB-231 /premir-204 MDA-MB-231 scramble
4 Fold Change Supplementary figure 3. Validation of DNA microarrays analysis using RT-qPCR for ten selected genes Microarray data RT-qPCR PRKA1 TCERG1L CHRNA4 4 EGR ESR1 CCND IPO5 EGFR2-10 RPS6 TIMP2
5 Supplementary table 1. MicroRNAs regulated in the complete set of breast tumors (9/9). Fold change (log2rq) values are indicated for individual micrornas in breast cancer patients (T). Up-regulated Number of samples found Breast cancer patient (T) T 15 T 30 T 75 T 76 T 78 T 85 T 88 T 105 T 191 hsa-let-7g* 9/ hsa-mir-148b* 9/ hsa-mir-454* 9/ hsa-mir-592 9/ Downregulated Number of samples found Breast cancer patient (T) T 15 T 30 T 75 T 76 T 78 T 85 T 88 T 105 T 191 hsa-let-7c 9/ hsa-mir-100 9/ hsa-mir-10b* 9/ hsa-mir-10b 9/ hsa-mir-132 9/ hsa-mir-139-3p 9/ hsa-mir-1 9/ hsa-mir-195 9/ hsa-mir-204 9/ hsa-mir-218 9/ hsa-mir-26b* 9/ hsa-mir-376a 9/ hsa-mir-376c 9/ hsa-mir-424* 9/ hsa-mir-432 9/ hsa-mir-433 9/ hsa-mir-489 9/ hsa-mir-656 9/ hsa-mir-944 9/
6 Cellular pathway KEGG Total genes -ln (p-value) MAPK signaling pathway Focal adhesion Wnt signaling pathway Axon guidance Ubiquitin mediated proteolysis Colorectal cancer Prostate cancer Adherens junction TGF-beta signaling pathway ErbB signaling pathway Supplementary table 2. Cellular pathways potentially affected by micrornas deregulated in breast tumors
7 Gene Sequence (5' - 3') Product length (bp) Nucleotide position GAPDH IPO5 RPS6 TIMP2 EGFR2 ESR1 CCND1 PRKAA1 EGR3 ESR1 TCERG1L CHRNA4 ANGPT1 TGFBR2 CREB5 FW- CCCCACCACACTGAATCTCC RV- GTACATGACAAGGTGCGGCT FW- AGGAGAAATGCACGAGGCAA RV- TTAAGGCCCTTCACGCAGAG FW- TGTTACTCCACGTGTCCTGC RV- AAGTCTGCGTCTCTTCGCAA FW- AGCTTTGCTTTATCCGGGCT RV- ATGCTTAGCTGGCGTCACAT FW- GGTGGCATTTAGGGGTGACT RV- CAAGGGAACAGGAAATATGTCG FW- ACTCAACAGCGTGTCTCCG RV- CACATTTTCCCTGGTTCCTGTAG FW- GCTGTAGTGGGGTTCTAGGC RV- GGCACGCTACGCTACTGTAA FW- AGTGTCGCTTTTTGAATAGTTTG RV- AGCAGTCTAGGCCAACAAGC FW- GGTGACCATGAGCAGTTTGC RV- TAGGTCACGGTCTTGTTGCC FW- CGCTGCGTCGCCTCTAA RV- TCCAGCTCGTTCCCTTGGAT FW- TACAGGTTTCCAAGGGTGGC RV- TGCAAACTCCGGGCATACAA FW- AATGTCACCTCCATCCGCATC RV- GTCAGCATTGTTGTAGAGGG FW- CGTGAATCTGGAGCCGTTTG RV- AGTAGTTTGAAGCACAGCAAGC FW- TGTCAGAGGATACTGTGGCTTG RV- TGAGTACAGCTGAAGTGTTCCAT TCTTCAAAGACTGGCTTTCATTTT AAAGCACCAAAGCGTTTCCC UTR UTR UTR Supplementary table 4. Primers used to validate DNA microarrays data.
8 Supplementary table 5. Cell proliferation and migration genes modulated by mir-204 Expression Biological Process Number of genes Gene Symbol Downregulated Proliferation 38 E2F1, EXO1, TXNIP, CLSPN, DBF4B, PDS5A, PDPN, TIPIN, SKP2, CDK4, TACC1, TACC2, CDT1, PSMA2, TUBB, EREG, NPM1, TGFBR2*, SMAD4* CHTF18, MAPRE2*, FANCA, DSCC1, CDK4, ID2, NPM1, FOXC1, ASNS, ID3, GPR3, CREB5*,EXTL3, AKT1S1, ARHGAP5*, SPTBN1, NEFL, CDH4,RAB22A* Migration/Invasion 24 CAP2, MARK1, LLGL1, HOOK3, TACC2, EPB41L2, CORO2B, TUBB, ANK3, NPM1, CAP1, NEFL, LCP1, TUBA8, KIF21A, TACC2,FMOD, EID2, ID1, TGFBR2*, SMAD4*, FOXC1*, FRAS1*, LRP8* Upregulated Proliferation 32 btg1, GJA1, BDKRB2, TRIB1, GPNMB, ptch1, Wisp2, Dbp, BTG4, Hyal1, Nppc, mmp7, il6r, irs2, Spn, KCTD11, ESR2, egln3, TNFRSF14, APC, nupr1, ADRA1A, RARRES1, Hyal1, LAMA5, Ang, Kiss1r, Ifitm1, Hyal1, Scgb3a1,LAMA5,GPNMB Migration/Invasion 29 TMEM8B, RET, fn1, GPNMB, ITGA1, RHOB, ITGA7, CNTN2, Cntn4, Wisp2, ROR2, Itgax, PCDHB9, PCDHB10, PCDHB11, CD72, PCDHB5, Wisp2, Nedd9, sspn, CDH1, fn1, APC, RHOB, Selplg, LAMA5, GPNMB, LAMA5, CCR1 *Genes with mir-204 binding sites
9 Primer name Sequence 5-3 Nucleotide position in target gene ANGPT1.1-S GATCCGGATTTCAGAAGCCTAATTCCTTCAAGAGAGGAATTAGGCT TCTGAAATCCTTTTTT GGAAA ANGPT1.1-AS AGCTTTTCCAAAAAAGGATTTCAGAAGCCTAATTCCTCTCTTGAAG GAATTAGGCTTCTGAAATCCG ANGPT1.2-S GATCCGCCATAATGAACTGTAGTACATTCAAGAGA TGTACTACAGTTCATTATGGCTTTTTTGGAAA ANGPT1.2-AS AGCTTTTCCAAAAAAGCCATAATGAACTGTAGTACATCTCTTGAATG TACTACAGTTCATTATGGCG TGF R2 1.1-S GATCC GGAGGAGAAGATTCCTGAAGA TTCAAGAGA TCTTCAGGAATCTTCTCCTCC TTTT TT GGAAA TGF R2 1.1-AS AGCTTTTCCAAAAAAGGAGGAGAAGATTCCTGAAGATCTCTTGAAT CTTCAGGAATCTTCTCCTCCG TGF R2 1.2-S GATCCGGTGTTAAGATTTGAAGTTGGTTCAAGAGACCAACTTCAAA TCTTAACACCTTTTTTGGAAA TGF R2 1.2-AS AGCTTTTCCAAAAAAGGTGTTAAGATTTGAAGTTGGTCTCTTGAAC CAACTTCAAATCTTAACACCG Supplementary table 6. Primers used to silencing ANGPT1 and TGF R2 gene expression
Flores-Pérez A et al Suppression of cell migration is promoted by mir-944 through targeting of SIAH1 and PTP4A1 in breast cancer cells
 Author s response to reviews Title: Suppression of cell migration is promoted by mir-944 through targeting of SIAH1 and PTP4A1 in breast cancer cells Authors: Cesar Lopez-Camarillo (genomicas@yahoo.com.mx)
Author s response to reviews Title: Suppression of cell migration is promoted by mir-944 through targeting of SIAH1 and PTP4A1 in breast cancer cells Authors: Cesar Lopez-Camarillo (genomicas@yahoo.com.mx)
Late regulation of immune genes and micrornas in circulating leukocytes in a pig model of
 1 Supplementary material for: 2 3 4 5 6 Late regulation of immune genes and micrornas in circulating leukocytes in a pig model of influenza A (H1N2) infection Louise Brogaard, Peter M. H. Heegaard, Lars
1 Supplementary material for: 2 3 4 5 6 Late regulation of immune genes and micrornas in circulating leukocytes in a pig model of influenza A (H1N2) infection Louise Brogaard, Peter M. H. Heegaard, Lars
microrna-200b and microrna-200c promote colorectal cancer cell proliferation via
 Supplementary Materials microrna-200b and microrna-200c promote colorectal cancer cell proliferation via targeting the reversion-inducing cysteine-rich protein with Kazal motifs Supplementary Table 1.
Supplementary Materials microrna-200b and microrna-200c promote colorectal cancer cell proliferation via targeting the reversion-inducing cysteine-rich protein with Kazal motifs Supplementary Table 1.
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
The Role of micrornas in Pain. Phd Work of: Rehman Qureshi Advisor: Ahmet Sacan
 The Role of micrornas in Pain Phd Work of: Rehman Qureshi Advisor: Ahmet Sacan 1 Motivation for molecular analysis of Pain Chronic pain one of the most common causes for seeking medical treatment Approximate
The Role of micrornas in Pain Phd Work of: Rehman Qureshi Advisor: Ahmet Sacan 1 Motivation for molecular analysis of Pain Chronic pain one of the most common causes for seeking medical treatment Approximate
Supplemental Table 1. MicroRNAs regulated by ZEB1.
 Supplemental Table 1. MicroRNAs regulated by ZEB1. down-regulated mirnas function cancer roles [representative target] mir-34a TS prostate cancer (1) inhibits clonogenic expansion, tumor regeneration,
Supplemental Table 1. MicroRNAs regulated by ZEB1. down-regulated mirnas function cancer roles [representative target] mir-34a TS prostate cancer (1) inhibits clonogenic expansion, tumor regeneration,
Supplementary Figure 1: Func8onal Network Analysis of Kinases Significantly Modulated by MERS CoV Infec8on and Conserved Across All Time Points
 A. B. 8 4 Supplementary Figure : Func8onal Network Analysis of Kinases Significantly Modulated by MERS CoV Infec8on and Conserved Across All Time Points Examined. A) Venn diagram analysis of kinases significantly
A. B. 8 4 Supplementary Figure : Func8onal Network Analysis of Kinases Significantly Modulated by MERS CoV Infec8on and Conserved Across All Time Points Examined. A) Venn diagram analysis of kinases significantly
Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier
 Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier curves for progression-free survival (PFS) and overall survival (OS) in a cohort of patients (N=52) with stage III primary ovarian
Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier curves for progression-free survival (PFS) and overall survival (OS) in a cohort of patients (N=52) with stage III primary ovarian
Supplementary Material
 Supplementary Material Identification of mir-187 and mir-182 as biomarkers for early diagnosis and prognosis in prostate cancer patients treated with radical prostatectomy Irene Casanova-Salas 1, José
Supplementary Material Identification of mir-187 and mir-182 as biomarkers for early diagnosis and prognosis in prostate cancer patients treated with radical prostatectomy Irene Casanova-Salas 1, José
microrna PCR System (Exiqon), following the manufacturer s instructions. In brief, 10ng of
 SUPPLEMENTAL MATERIALS AND METHODS Quantitative RT-PCR Quantitative RT-PCR analysis was performed using the Universal mircury LNA TM microrna PCR System (Exiqon), following the manufacturer s instructions.
SUPPLEMENTAL MATERIALS AND METHODS Quantitative RT-PCR Quantitative RT-PCR analysis was performed using the Universal mircury LNA TM microrna PCR System (Exiqon), following the manufacturer s instructions.
MicroRNAs driving invasion and metastasis in ovarian cancer: Opportunities for translational medicine (Review)
 INTERNATIONAL JOURNAL OF ONCOLOGY 50: 1461-1476, 2017 MicroRNAs driving invasion and metastasis in ovarian cancer: Opportunities for translational medicine (Review) Carlos Palma Flores 1*, Raúl García-Vázquez
INTERNATIONAL JOURNAL OF ONCOLOGY 50: 1461-1476, 2017 MicroRNAs driving invasion and metastasis in ovarian cancer: Opportunities for translational medicine (Review) Carlos Palma Flores 1*, Raúl García-Vázquez
Supplementary Files. Novel blood-based microrna biomarker panel for early diagnosis of chronic pancreatitis
 Supplementary Files Novel blood-based microrna biomarker panel for early diagnosis of chronic pancreatitis Lei Xin 1 *, M.D., Jun Gao 1 *, Ph.D., Dan Wang 1 *, M.D., Jin-Huan Lin 1, M.D., Zhuan Liao 1,
Supplementary Files Novel blood-based microrna biomarker panel for early diagnosis of chronic pancreatitis Lei Xin 1 *, M.D., Jun Gao 1 *, Ph.D., Dan Wang 1 *, M.D., Jin-Huan Lin 1, M.D., Zhuan Liao 1,
TEB. Id4 p63 DAPI Merge. Id4 CK8 DAPI Merge
 a Duct TEB b Id4 p63 DAPI Merge Id4 CK8 DAPI Merge c d e Supplementary Figure 1. Identification of Id4-positive MECs and characterization of the Comma-D model. (a) IHC analysis of ID4 expression in the
a Duct TEB b Id4 p63 DAPI Merge Id4 CK8 DAPI Merge c d e Supplementary Figure 1. Identification of Id4-positive MECs and characterization of the Comma-D model. (a) IHC analysis of ID4 expression in the
MicroRNA expression profiling and functional analysis in prostate cancer. Marco Folini s.c. Ricerca Traslazionale DOSL
 MicroRNA expression profiling and functional analysis in prostate cancer Marco Folini s.c. Ricerca Traslazionale DOSL What are micrornas? For almost three decades, the alteration of protein-coding genes
MicroRNA expression profiling and functional analysis in prostate cancer Marco Folini s.c. Ricerca Traslazionale DOSL What are micrornas? For almost three decades, the alteration of protein-coding genes
Genomic Analyses across Six Cancer Types Identify Basal-like Breast Cancer as a Unique Molecular Entity
 Genomic Analyses across Six Cancer Types Identify Basal-like Breast Cancer as a Unique Molecular Entity Aleix Prat, Barbara Adamo, Cheng Fan, Vicente Peg, Maria Vidal, Patricia Galván, Ana Vivancos, Paolo
Genomic Analyses across Six Cancer Types Identify Basal-like Breast Cancer as a Unique Molecular Entity Aleix Prat, Barbara Adamo, Cheng Fan, Vicente Peg, Maria Vidal, Patricia Galván, Ana Vivancos, Paolo
SUPPLEMENTAY FIGURES AND TABLES
 SUPPLEMENTAY FIGURES AND TABLES Supplementary Figure S1: Validation of mir expression by quantitative real-time PCR and time course studies on mir- 29a-3p and mir-210-3p. A. The graphs illustrate the expression
SUPPLEMENTAY FIGURES AND TABLES Supplementary Figure S1: Validation of mir expression by quantitative real-time PCR and time course studies on mir- 29a-3p and mir-210-3p. A. The graphs illustrate the expression
U118MG. Supplementary Figure 1 U373MG U118MG 3.5 A CCF-SSTG
 A172 CCF-SSTG1 15 - - - 1 1 1 2 2 3 4 6 7 8 10101112131617192022-1 1 1 2 2 3 4 6 7 8 10 10 11 12 13 16 17 19 20 22 T98G U373MG - - - 1 1 1 2 2 3 4 6 7 8 10 10 11 12 13 16 17 19 20 22-1 1 1 2 2 3 4 6 7
A172 CCF-SSTG1 15 - - - 1 1 1 2 2 3 4 6 7 8 10101112131617192022-1 1 1 2 2 3 4 6 7 8 10 10 11 12 13 16 17 19 20 22 T98G U373MG - - - 1 1 1 2 2 3 4 6 7 8 10 10 11 12 13 16 17 19 20 22-1 1 1 2 2 3 4 6 7
SUPPLEMENTARY INFORMATION. Supp. Fig. 1. Autoimmunity. Tolerance APC APC. T cell. T cell. doi: /nature06253 ICOS ICOS TCR CD28 TCR CD28
 Supp. Fig. 1 a APC b APC ICOS ICOS TCR CD28 mir P TCR CD28 P T cell Tolerance Roquin WT SG Icos mrna T cell Autoimmunity Roquin M199R SG Icos mrna www.nature.com/nature 1 Supp. Fig. 2 CD4 + CD44 low CD4
Supp. Fig. 1 a APC b APC ICOS ICOS TCR CD28 mir P TCR CD28 P T cell Tolerance Roquin WT SG Icos mrna T cell Autoimmunity Roquin M199R SG Icos mrna www.nature.com/nature 1 Supp. Fig. 2 CD4 + CD44 low CD4
Supplementary Figure 1. The mir-182 binding site of SMAD7 3 UTR and the. mutated sequence.
 Supplementary Figure 1. The mir-182 binding site of SMAD7 3 UTR and the mutated sequence. 1 Supplementary Figure 2. Expression of mir-182 and SMAD7 in various cell lines. (A) Basal levels of mir-182 expression
Supplementary Figure 1. The mir-182 binding site of SMAD7 3 UTR and the mutated sequence. 1 Supplementary Figure 2. Expression of mir-182 and SMAD7 in various cell lines. (A) Basal levels of mir-182 expression
Supplemental Data. TGF-β-mediated mir-181a expression promotes breast cancer metastasis by targeting Bim.
 Supplemental Data TGF-β-mediated mir-181a expression promotes breast cancer metastasis by targeting Bim. Molly A. Taylor 1, Khalid Sossey-Alaoui 2, Cheryl L. Thompson 3, David Danielpour 4, and William
Supplemental Data TGF-β-mediated mir-181a expression promotes breast cancer metastasis by targeting Bim. Molly A. Taylor 1, Khalid Sossey-Alaoui 2, Cheryl L. Thompson 3, David Danielpour 4, and William
Profiles of gene expression & diagnosis/prognosis of cancer. MCs in Advanced Genetics Ainoa Planas Riverola
 Profiles of gene expression & diagnosis/prognosis of cancer MCs in Advanced Genetics Ainoa Planas Riverola Gene expression profiles Gene expression profiling Used in molecular biology, it measures the
Profiles of gene expression & diagnosis/prognosis of cancer MCs in Advanced Genetics Ainoa Planas Riverola Gene expression profiles Gene expression profiling Used in molecular biology, it measures the
GENE EXPRESSION AND microrna PROFILING IN LYMPHOMAS. AN INTEGRATIVE GENOMICS ANALYSIS
 GENE EXPRESSION AND microrna PROFILING IN LYMPHOMAS. AN INTEGRATIVE GENOMICS ANALYSIS Dra. Margarita Sánchez-Beato Lymphoma Group Molecular Pathology Programme CNIO Agilent Microarray and Sequencing Roadshow,
GENE EXPRESSION AND microrna PROFILING IN LYMPHOMAS. AN INTEGRATIVE GENOMICS ANALYSIS Dra. Margarita Sánchez-Beato Lymphoma Group Molecular Pathology Programme CNIO Agilent Microarray and Sequencing Roadshow,
The role of microrna in metastatic processes of non-small cell lung carcinoma
 The role of microrna in metastatic processes of non-small cell lung carcinoma Zuzana Pastorkova a, Jozef Skarda a, Jozef Andel b Background. MicroRNAs are small non-coding one-stranded RNA molecules that
The role of microrna in metastatic processes of non-small cell lung carcinoma Zuzana Pastorkova a, Jozef Skarda a, Jozef Andel b Background. MicroRNAs are small non-coding one-stranded RNA molecules that
Title: Single Cell Dual Adherent-Suspension Co-Culture Micro-Environment for Studying Tumor- Stromal Interactions
 Electronic Supplementary Material (ESI) for Lab on a Chip. This journal is The Royal Society of Chemistry 2016 Title: Single Cell Dual Adherent-Suspension Co-Culture Micro-Environment for Studying Tumor-
Electronic Supplementary Material (ESI) for Lab on a Chip. This journal is The Royal Society of Chemistry 2016 Title: Single Cell Dual Adherent-Suspension Co-Culture Micro-Environment for Studying Tumor-
Rho GTPase activating protein 8 /// PRR5- ARHGAP8 fusion
 Probe Set ID RefSeq Transcript ID Gene Title Gene Symbol 205980_s_at NM_001017526 /// NM_181334 /// NM_181335 Rho GTPase activating protein 8 /// PRR5- ARHGAP8 fusion ARHGAP8 /// LOC553158 FC ALK shrna
Probe Set ID RefSeq Transcript ID Gene Title Gene Symbol 205980_s_at NM_001017526 /// NM_181334 /// NM_181335 Rho GTPase activating protein 8 /// PRR5- ARHGAP8 fusion ARHGAP8 /// LOC553158 FC ALK shrna
About OMICS International Conferences
 About OMICS Group OMICS Group is an amalgamation of Open Access publications and worldwide international science conferences and events. Established in the year 2007 with the sole aim of making the information
About OMICS Group OMICS Group is an amalgamation of Open Access publications and worldwide international science conferences and events. Established in the year 2007 with the sole aim of making the information
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
Supplementary Table 1. Genes analysed for expression by angiogenesis gene-array.
 Supplementary Table 1. Genes analysed for expression by angiogenesis gene-array. Gene symbol Gene name TaqMan Assay ID UniGene ID 18S rrna 18S ribosomal RNA Hs99999901_s1 Actb actin, beta Mm00607939_s1
Supplementary Table 1. Genes analysed for expression by angiogenesis gene-array. Gene symbol Gene name TaqMan Assay ID UniGene ID 18S rrna 18S ribosomal RNA Hs99999901_s1 Actb actin, beta Mm00607939_s1
Supplementary Table S1. List of PTPRK-RSPO3 gene fusions in TCGA's colon cancer cohort. Chr. # of Gene 2. Chr. # of Gene 1
 Supplementary Tale S1. List of PTPRK-RSPO3 gene fusions in TCGA's colon cancer cohort TCGA Case ID Gene-1 Gene-2 Chr. # of Gene 1 Chr. # of Gene 2 Genomic coordiante of Gene 1 at fusion junction Genomic
Supplementary Tale S1. List of PTPRK-RSPO3 gene fusions in TCGA's colon cancer cohort TCGA Case ID Gene-1 Gene-2 Chr. # of Gene 1 Chr. # of Gene 2 Genomic coordiante of Gene 1 at fusion junction Genomic
Supplementary Figure 1. HeliScope CAGE revealed androgen-regulated signaling and differentially regulated promoters in hormone-refractory prostate
 Supplementary Figure 1. HeliScope CAGE revealed androgen-regulated signaling and differentially regulated promoters in hormone-refractory prostate cancer cells. (a) Cell proliferation of BicR cells in
Supplementary Figure 1. HeliScope CAGE revealed androgen-regulated signaling and differentially regulated promoters in hormone-refractory prostate cancer cells. (a) Cell proliferation of BicR cells in
Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3
 Supplemental Figure Legends. Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3 ErbB3 gene copy number gain. Supplemental Figure S1. ERBB3 mrna levels are elevated in
Supplemental Figure Legends. Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3 ErbB3 gene copy number gain. Supplemental Figure S1. ERBB3 mrna levels are elevated in
Supplementary Figure 1:
 Supplementary Figure 1: (A) Whole aortic cross-sections stained with Hematoxylin and Eosin (H&E), 7 days after porcine-pancreatic-elastase (PPE)-induced AAA compared to untreated, healthy control aortas
Supplementary Figure 1: (A) Whole aortic cross-sections stained with Hematoxylin and Eosin (H&E), 7 days after porcine-pancreatic-elastase (PPE)-induced AAA compared to untreated, healthy control aortas
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3021 Supplementary figure 1 Characterisation of TIMPless fibroblasts. a) Relative gene expression of TIMPs1-4 by real time quantitative PCR (RT-qPCR) in WT or ΔTimp fibroblasts (mean ±
DOI: 10.1038/ncb3021 Supplementary figure 1 Characterisation of TIMPless fibroblasts. a) Relative gene expression of TIMPs1-4 by real time quantitative PCR (RT-qPCR) in WT or ΔTimp fibroblasts (mean ±
Supplementary Table 3. 3 UTR primer sequences. Primer sequences used to amplify and clone the 3 UTR of each indicated gene are listed.
 Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
COMPUTATIONAL OPTIMISATION OF TARGETED DNA SEQUENCING FOR CANCER DETECTION
 COMPUTATIONAL OPTIMISATION OF TARGETED DNA SEQUENCING FOR CANCER DETECTION Pierre Martinez, Nicholas McGranahan, Nicolai Juul Birkbak, Marco Gerlinger, Charles Swanton* SUPPLEMENTARY INFORMATION SUPPLEMENTARY
COMPUTATIONAL OPTIMISATION OF TARGETED DNA SEQUENCING FOR CANCER DETECTION Pierre Martinez, Nicholas McGranahan, Nicolai Juul Birkbak, Marco Gerlinger, Charles Swanton* SUPPLEMENTARY INFORMATION SUPPLEMENTARY
ras Multikinase Inhibitor Multikinase Inhibitor 0.1
 a ras ** * ** * ** ** ** ** un in et m lu Se ib SL G 32 W 7 So 50 ra 74 fe ni b LY W 294 or 0 tm 02 R an ap n a in Ev my er cin ol im BE us Z2 3 En P 5 za I13 st 0 au rin D as SP ati 60 nib C 012 is 5
a ras ** * ** * ** ** ** ** un in et m lu Se ib SL G 32 W 7 So 50 ra 74 fe ni b LY W 294 or 0 tm 02 R an ap n a in Ev my er cin ol im BE us Z2 3 En P 5 za I13 st 0 au rin D as SP ati 60 nib C 012 is 5
An Integrative Pharmacogenomic Approach Identifies Two-drug. Combination Therapies for Personalized Cancer Medicine
 An Integrative Pharmacogemic Approach Identifies Two-drug Combination Therapies for Personalized Cancer Medicine Yin Liu 1,2,3, Teng Fei 4,5, Xiaoqi Zheng 6, Myles Brown 4,5, Peng Zhang 7*, X. Shirley
An Integrative Pharmacogemic Approach Identifies Two-drug Combination Therapies for Personalized Cancer Medicine Yin Liu 1,2,3, Teng Fei 4,5, Xiaoqi Zheng 6, Myles Brown 4,5, Peng Zhang 7*, X. Shirley
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Amelio et al., http://www.jcb.org/cgi/content/full/jcb.201203134/dc1 Figure S1. mir-24 regulates proliferation and by itself induces
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Amelio et al., http://www.jcb.org/cgi/content/full/jcb.201203134/dc1 Figure S1. mir-24 regulates proliferation and by itself induces
BWA alignment to reference transcriptome and genome. Convert transcriptome mappings back to genome space
 Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Case Study - Informatics
 bd@jubilantbiosys.com Case Study - Informatics www.jubilantbiosys.com Validating as a Prognostic marker and Therapeutic Target for Multiple Cancers Introduction Human genome projects and high throughput
bd@jubilantbiosys.com Case Study - Informatics www.jubilantbiosys.com Validating as a Prognostic marker and Therapeutic Target for Multiple Cancers Introduction Human genome projects and high throughput
Supplementary Information. Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase
 Journal: Nature Medicine Supplementary Information Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase 1,2 Peng Wang PhD, 1,2 Juan-Carlos Alvarez-Perez
Journal: Nature Medicine Supplementary Information Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase 1,2 Peng Wang PhD, 1,2 Juan-Carlos Alvarez-Perez
Supporting Information
 Supporting Information Retinal expression of small non-coding RNAs in a murine model of proliferative retinopathy Chi-Hsiu Liu 1, Zhongxiao Wang 1, Ye Sun 1, John Paul SanGiovanni 2, Jing Chen 1, * 1 Department
Supporting Information Retinal expression of small non-coding RNAs in a murine model of proliferative retinopathy Chi-Hsiu Liu 1, Zhongxiao Wang 1, Ye Sun 1, John Paul SanGiovanni 2, Jing Chen 1, * 1 Department
Original Article Identification of hsa-mir-21 as a target gene of tumorigenesis in neuroblastoma
 Int J Clin Exp Med 2016;9(2):2718-2727 www.ijcem.com /ISSN:1940-5901/IJCEM0016181 Original Article Identification of hsa-mir-21 as a target gene of tumorigenesis in neuroblastoma Zuopeng Wang *, Wei Yao
Int J Clin Exp Med 2016;9(2):2718-2727 www.ijcem.com /ISSN:1940-5901/IJCEM0016181 Original Article Identification of hsa-mir-21 as a target gene of tumorigenesis in neuroblastoma Zuopeng Wang *, Wei Yao
Plasma-Seq conducted with blood from male individuals without cancer.
 Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland
 AD Award Number: W81XWH-10-1-1029 TITLE: PRINCIPAL INVESTIGATOR: Mu-Shui Dai, M.D., Ph.D. CONTRACTING ORGANIZATION: Oregon Health Science niversity, Portland, Oregon 97239 REPORT DATE: October 2013 TYPE
AD Award Number: W81XWH-10-1-1029 TITLE: PRINCIPAL INVESTIGATOR: Mu-Shui Dai, M.D., Ph.D. CONTRACTING ORGANIZATION: Oregon Health Science niversity, Portland, Oregon 97239 REPORT DATE: October 2013 TYPE
EGFR: fundamenteel en klinisch
 EGFR: fundamenteel en klinisch Guido Lammering MAASTRO Clinic Maastricht, NL What is EGFR?? The EGFR some facts 1186 amino acids 170 kda Expressed by all cells of epithelial origin Increased activation
EGFR: fundamenteel en klinisch Guido Lammering MAASTRO Clinic Maastricht, NL What is EGFR?? The EGFR some facts 1186 amino acids 170 kda Expressed by all cells of epithelial origin Increased activation
SALSA MLPA Probemix P014-B1 Chromosome 8 Lot B and B
 SALSA MLPA Probemix P014-B1 Chromosome 8 Lot B1-0916 and B1-0713. Copy number changes of the human chromosome 8 are common in many types of tumours. In most cases, losses of 8p sequences and gains of 8q
SALSA MLPA Probemix P014-B1 Chromosome 8 Lot B1-0916 and B1-0713. Copy number changes of the human chromosome 8 are common in many types of tumours. In most cases, losses of 8p sequences and gains of 8q
Table 1 Demographic data of PD-patients. Data are represented as counts or median and full range.
 Table 1 Demographic data of PD-patients. Data are represented as counts or median and full range. Parameter Number Patients 5 Diabetes mellitus 0 Previous NTX 0 Steroids 0 Patients with residual renal
Table 1 Demographic data of PD-patients. Data are represented as counts or median and full range. Parameter Number Patients 5 Diabetes mellitus 0 Previous NTX 0 Steroids 0 Patients with residual renal
Supplementary Table 1: List of the 242 hypoxia/reoxygenation marker genes collected from literature and databases
 Supplementary Tables: Supplementary Table 1: List of the 242 hypoxia/reoxygenation marker genes collected from literature and databases Supplementary Table 2: Contingency tables used in the fisher exact
Supplementary Tables: Supplementary Table 1: List of the 242 hypoxia/reoxygenation marker genes collected from literature and databases Supplementary Table 2: Contingency tables used in the fisher exact
mir-509-5p and mir-1243 increase the sensitivity to gemcitabine by inhibiting
 mir-509-5p and mir-1243 increase the sensitivity to gemcitabine by inhibiting epithelial-mesenchymal transition in pancreatic cancer Hidekazu Hiramoto, M.D. 1,3, Tomoki Muramatsu, Ph.D. 1, Daisuke Ichikawa,
mir-509-5p and mir-1243 increase the sensitivity to gemcitabine by inhibiting epithelial-mesenchymal transition in pancreatic cancer Hidekazu Hiramoto, M.D. 1,3, Tomoki Muramatsu, Ph.D. 1, Daisuke Ichikawa,
Supplementary Material. Part I: Sample Information. Part II: Pathway Information
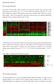 Supplementary Material Part I: Sample Information Three NPC cell lines, CNE1, CNE2, and HK1 were treated with CYC202. Gene expression of 380 selected genes were collected at 0, 2, 4, 6, 12 and 24 hours
Supplementary Material Part I: Sample Information Three NPC cell lines, CNE1, CNE2, and HK1 were treated with CYC202. Gene expression of 380 selected genes were collected at 0, 2, 4, 6, 12 and 24 hours
Supplementary figures legends
 Supplementary Information to Nijmegen Breakage Syndrome fibroblasts and ipscs: cellular models for uncovering diseaseassociated signaling pathways and establishing a screening platform for anti-oxidants
Supplementary Information to Nijmegen Breakage Syndrome fibroblasts and ipscs: cellular models for uncovering diseaseassociated signaling pathways and establishing a screening platform for anti-oxidants
CONTRACTING ORGANIZATION: Baylor College of Medicine Houston, TX 77030
 AD Award Number: W81XWH-05-1-0428 TITLE: MicroRNA and Breast Cancer Progression PRINCIPAL INVESTIGATOR: Konstantin Galaktionov, Ph.D. CONTRACTING ORGANIZATION: Baylor College of Medicine Houston, TX 77030
AD Award Number: W81XWH-05-1-0428 TITLE: MicroRNA and Breast Cancer Progression PRINCIPAL INVESTIGATOR: Konstantin Galaktionov, Ph.D. CONTRACTING ORGANIZATION: Baylor College of Medicine Houston, TX 77030
Supplemental Information. High-Throughput Microfluidic Labyrinth for the. Label-free Isolation of Circulating Tumor Cells
 Cell Systems, Volume 5 Supplemental Information High-Throughput Microfluidic Labyrinth for the Label-free Isolation of Circulating Tumor Cells Eric Lin, Lianette Rivera-Báez, Shamileh Fouladdel, Hyeun
Cell Systems, Volume 5 Supplemental Information High-Throughput Microfluidic Labyrinth for the Label-free Isolation of Circulating Tumor Cells Eric Lin, Lianette Rivera-Báez, Shamileh Fouladdel, Hyeun
Cardiovascular. Diseases. a simple and standardized kit qpcr analysis of circulating platelet-derived micrornas
 microrna Biomarkers of Platelet Function Neurodegenerative Diseases Cardiovascular Diseases Musculoskeletal Diseases a simple and standardized kit qpcr analysis of circulating platelet-derived micrornas
microrna Biomarkers of Platelet Function Neurodegenerative Diseases Cardiovascular Diseases Musculoskeletal Diseases a simple and standardized kit qpcr analysis of circulating platelet-derived micrornas
Supplementary Material. Table S1. Summary of mapping results
 Supplementary Material Table S1. Summary of mapping results Sample Total reads Mapped (%) HNE0-1 99687516 80786083 (81.0%) HNE0-2 94318720 77047792 (81.7%) HNE0-3 104033900 84348102 (81.1%) HNE15-1 94426598
Supplementary Material Table S1. Summary of mapping results Sample Total reads Mapped (%) HNE0-1 99687516 80786083 (81.0%) HNE0-2 94318720 77047792 (81.7%) HNE0-3 104033900 84348102 (81.1%) HNE15-1 94426598
Supplementary Figure 1. Characterization of ALDH-positive cell population in MCF-7 cells. (a) Expression level of stem cell markers in MCF-7 cells or
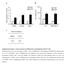 Supplementary Figure 1. Characterization of ALDH-positive cell population in MCF-7 cells. (a) Expression level of stem cell markers in MCF-7 cells or ALDH-positive cell population by qpcr. Data represent
Supplementary Figure 1. Characterization of ALDH-positive cell population in MCF-7 cells. (a) Expression level of stem cell markers in MCF-7 cells or ALDH-positive cell population by qpcr. Data represent
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Review Article Regulation of Breast Cancer and Bone Metastasis by MicroRNAs
 Disease Markers Volume 35 (2013), Issue 5, Pages 369 387 http://dx.doi.org/10.1155/2013/451248 Review Article Regulation of Breast Cancer and Bone Metastasis by MicroRNAs S. Vimalraj, P. J. Miranda, B.
Disease Markers Volume 35 (2013), Issue 5, Pages 369 387 http://dx.doi.org/10.1155/2013/451248 Review Article Regulation of Breast Cancer and Bone Metastasis by MicroRNAs S. Vimalraj, P. J. Miranda, B.
mir-125a-5p expression is associated with the age of breast cancer patients
 mir-125a-5p expression is associated with the age of breast cancer patients H. He 1 *, F. Xu 2 *, W. Huang 1, S.Y. Luo 1, Y.T. Lin 1, G.H. Zhang 1, Q. Du 1 and R.H. Duan 1 1 The State Key Laboratory of
mir-125a-5p expression is associated with the age of breast cancer patients H. He 1 *, F. Xu 2 *, W. Huang 1, S.Y. Luo 1, Y.T. Lin 1, G.H. Zhang 1, Q. Du 1 and R.H. Duan 1 1 The State Key Laboratory of
Supplementary figures
 Supplementary figures Supplementary figure 1: Pathway enrichment for each comparison. The first column shows enrichment with differentially expressed genes between the diet groups at t = 0, the second
Supplementary figures Supplementary figure 1: Pathway enrichment for each comparison. The first column shows enrichment with differentially expressed genes between the diet groups at t = 0, the second
Supplementary information
 Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Additional file 2 List of pathway from PID
 Additional file 2 List of pathway from PID Pathway ID Pathway name # components # enriched GO terms a4b1_paxdep_pathway Paxillin-dependent events mediated by a4b1 20 179 a4b1_paxindep_pathway Paxillin-independent
Additional file 2 List of pathway from PID Pathway ID Pathway name # components # enriched GO terms a4b1_paxdep_pathway Paxillin-dependent events mediated by a4b1 20 179 a4b1_paxindep_pathway Paxillin-independent
Accel-Amplicon Panels
 Accel-Amplicon Panels Amplicon sequencing has emerged as a reliable, cost-effective method for ultra-deep targeted sequencing. This highly adaptable approach is especially applicable for in-depth interrogation
Accel-Amplicon Panels Amplicon sequencing has emerged as a reliable, cost-effective method for ultra-deep targeted sequencing. This highly adaptable approach is especially applicable for in-depth interrogation
We are IntechOpen, the world s leading publisher of Open Access books Built by scientists, for scientists. International authors and editors
 We are IntechOpen, the world s leading publisher of Open Access books Built by scientists, for scientists 4,000 116,000 120M Open access books available International authors and editors Downloads Our
We are IntechOpen, the world s leading publisher of Open Access books Built by scientists, for scientists 4,000 116,000 120M Open access books available International authors and editors Downloads Our
Targeted Agent and Profiling Utilization Registry (TAPUR ) Study. February 2018
 Targeted Agent and Profiling Utilization Registry (TAPUR ) Study February 2018 Precision Medicine Therapies designed to target the molecular alteration that aids cancer development 30 TARGET gene alterations
Targeted Agent and Profiling Utilization Registry (TAPUR ) Study February 2018 Precision Medicine Therapies designed to target the molecular alteration that aids cancer development 30 TARGET gene alterations
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Jennifer Hauenstein Oncology Cytogenetics Emory University Hospital Atlanta, GA
 Comparison of Genomic Coverage using Affymetrix OncoScan Array and Illumina TruSight Tumor 170 NGS Panel for Detection of Copy Number Abnormalities in Clinical GBM Specimens Jennifer Hauenstein Oncology
Comparison of Genomic Coverage using Affymetrix OncoScan Array and Illumina TruSight Tumor 170 NGS Panel for Detection of Copy Number Abnormalities in Clinical GBM Specimens Jennifer Hauenstein Oncology
Dysregulation of Blimp1 transcriptional repressor unleashes p130cas/erbb2 breast cancer invasion (SREP T) Supplementary File
 Dysregulation of Blimp1 transcriptional repressor unleashes p130cas/erbb2 breast cancer invasion (SREP-16-42496-T) Marianna Sciortino 1, Maria del Pilar Camacho Leal 1, Francesca Orso 1, Elena Grassi 1,
Dysregulation of Blimp1 transcriptional repressor unleashes p130cas/erbb2 breast cancer invasion (SREP-16-42496-T) Marianna Sciortino 1, Maria del Pilar Camacho Leal 1, Francesca Orso 1, Elena Grassi 1,
Multistep Model of Cervical Cancer: Participation of mirnas and Coding Genes
 Int. J. Mol. Sci. 2014, 15, 15700-15733; doi:10.3390/ijms150915700 Review OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Multistep Model of Cervical Cancer:
Int. J. Mol. Sci. 2014, 15, 15700-15733; doi:10.3390/ijms150915700 Review OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Multistep Model of Cervical Cancer:
SUPPLEMENTARY DATA. Assessment of cell survival (MTT) Assessment of early apoptosis and cell death
 Overexpression of a functional calcium-sensing receptor dramatically increases osteolytic potential of MDA-MB-231 cells in a mouse model of bone metastasis through epiregulinmediated osteoprotegerin downregulation
Overexpression of a functional calcium-sensing receptor dramatically increases osteolytic potential of MDA-MB-231 cells in a mouse model of bone metastasis through epiregulinmediated osteoprotegerin downregulation
MicroRNA expression profiling studies on bronchopulmonary dysplasia: a systematic review and meta-analysis
 MicroRNA expression profiling studies on bronchopulmonary dysplasia: a systematic review and meta-analysis Y. Yang, J. Qiu, Q. Kan, X.-G. Zhou and X.-Y. Zhou Department of Neonates, Nanjing Children s
MicroRNA expression profiling studies on bronchopulmonary dysplasia: a systematic review and meta-analysis Y. Yang, J. Qiu, Q. Kan, X.-G. Zhou and X.-Y. Zhou Department of Neonates, Nanjing Children s
Biobehavioral Pathways in Epithelial Ovarian Cancer
 Biobehavioral Pathways in Epithelial Ovarian Cancer Susan K. Lutgendorf, Ph.D. Departments of Psychology, Obstetrics and Gynecology, and Urology and Holden Comprehensive Cancer Center University of Iowa
Biobehavioral Pathways in Epithelial Ovarian Cancer Susan K. Lutgendorf, Ph.D. Departments of Psychology, Obstetrics and Gynecology, and Urology and Holden Comprehensive Cancer Center University of Iowa
Figure S1: Heat map based on the relative expression of genes and mirnas in human
 Figure S1: Heat map based on the relative expression of genes and mirnas in human prostate tissue. Columns represent genes and mirnas; rows represent prostate tissue samples (red squares: Tu samples, green
Figure S1: Heat map based on the relative expression of genes and mirnas in human prostate tissue. Columns represent genes and mirnas; rows represent prostate tissue samples (red squares: Tu samples, green
Supplementary Figure 1. HOPX is hypermethylated in NPC. (a) Methylation levels of HOPX in Normal (n = 24) and NPC (n = 24) tissues from the
 Supplementary Figure 1. HOPX is hypermethylated in NPC. (a) Methylation levels of HOPX in Normal (n = 24) and NPC (n = 24) tissues from the genome-wide methylation microarray data. Mean ± s.d.; Student
Supplementary Figure 1. HOPX is hypermethylated in NPC. (a) Methylation levels of HOPX in Normal (n = 24) and NPC (n = 24) tissues from the genome-wide methylation microarray data. Mean ± s.d.; Student
B16F1 B16F10 Supplemental Figure S1
 B16F1 B16F1 Supplemental Figure S1 Representative microangiography images of B16F1 and B16F1 tumors grown in the cranial windows. FITC-dextran (2 million MW) was injected systemically to visualize blood
B16F1 B16F1 Supplemental Figure S1 Representative microangiography images of B16F1 and B16F1 tumors grown in the cranial windows. FITC-dextran (2 million MW) was injected systemically to visualize blood
The mutations that drive cancer. Paul Edwards. Department of Pathology and Cancer Research UK Cambridge Institute, University of Cambridge
 The mutations that drive cancer Paul Edwards Department of Pathology and Cancer Research UK Cambridge Institute, University of Cambridge Previously on Cancer... hereditary predisposition Normal Cell Slightly
The mutations that drive cancer Paul Edwards Department of Pathology and Cancer Research UK Cambridge Institute, University of Cambridge Previously on Cancer... hereditary predisposition Normal Cell Slightly
IMPLEMENTING NEXT GENERATION SEQUENCING IN A PATHOLOGY LABORATORY
 Madrid, Spain IMPLEMENTING NEXT GENERATION SEQUENCING IN A PATHOLOGY LABORATORY Dr. JL Rodríguez Peralto NGS Ion Torrent Oncomine Focus Assay - Implementation experience for EGFR mutation detection
Madrid, Spain IMPLEMENTING NEXT GENERATION SEQUENCING IN A PATHOLOGY LABORATORY Dr. JL Rodríguez Peralto NGS Ion Torrent Oncomine Focus Assay - Implementation experience for EGFR mutation detection
Supplementary Information
 Supplementary Information Index of Supplementary Materials Supplementary Figure. Distribution of methylation levels, characteristics of cis-meqtls and association plots for meqtls in methylation-related
Supplementary Information Index of Supplementary Materials Supplementary Figure. Distribution of methylation levels, characteristics of cis-meqtls and association plots for meqtls in methylation-related
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
P. Mathijs Voorhoeve, Carlos le Sage, Mariette Schrier, Ad J.M. Gillis, Hans Stoop,
 Supplemental Data A Genetic Screen Implicates mirna-372 and mirna-373 As Oncogenes in Testicular Germ Cell Tumors P. Mathijs Voorhoeve, Carlos le Sage, Mariette Schrier, Ad J.M. Gillis, Hans Stoop, Remco
Supplemental Data A Genetic Screen Implicates mirna-372 and mirna-373 As Oncogenes in Testicular Germ Cell Tumors P. Mathijs Voorhoeve, Carlos le Sage, Mariette Schrier, Ad J.M. Gillis, Hans Stoop, Remco
Gene-microRNA network module analysis for ovarian cancer
 Gene-microRNA network module analysis for ovarian cancer Shuqin Zhang School of Mathematical Sciences Fudan University Oct. 4, 2016 Outline Introduction Materials and Methods Results Conclusions Introduction
Gene-microRNA network module analysis for ovarian cancer Shuqin Zhang School of Mathematical Sciences Fudan University Oct. 4, 2016 Outline Introduction Materials and Methods Results Conclusions Introduction
The mir-199a/brm/egr1 axis is a determinant of anchorage-independent growth in epithelial tumor cell lines
 Supplementary information Supplementary Figure -9 Supplementary Table -4 The mir-99a/brm/egr axis is a determinant of anchorage-independent growth in epithelial tumor cell lines Kazuyoshi Kobayashi, Kouhei
Supplementary information Supplementary Figure -9 Supplementary Table -4 The mir-99a/brm/egr axis is a determinant of anchorage-independent growth in epithelial tumor cell lines Kazuyoshi Kobayashi, Kouhei
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature15260 Supplementary Data 1: Gene expression in individual basal/stem, luminal, and luminal progenitor cells. Box plots show expression levels for each gene from the 49-gene differentiation
doi:10.1038/nature15260 Supplementary Data 1: Gene expression in individual basal/stem, luminal, and luminal progenitor cells. Box plots show expression levels for each gene from the 49-gene differentiation
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
Inferring condition-specific mirna activity from matched mirna and mrna expression data
 Inferring condition-specific mirna activity from matched mirna and mrna expression data Junpeng Zhang 1, Thuc Duy Le 2, Lin Liu 2, Bing Liu 3, Jianfeng He 4, Gregory J Goodall 5 and Jiuyong Li 2,* 1 Faculty
Inferring condition-specific mirna activity from matched mirna and mrna expression data Junpeng Zhang 1, Thuc Duy Le 2, Lin Liu 2, Bing Liu 3, Jianfeng He 4, Gregory J Goodall 5 and Jiuyong Li 2,* 1 Faculty
Impact of NM23-M1 "knock-out" on metastatic potential of hepatocellular carcinoma: a transgenic approach
 Impact of NM23-M1 "knock-out" on metastatic potential of hepatocellular carcinoma: a transgenic approach NM23-M1 "knock- out" mice (without NDPK A) Experimental models of hepatocarcinogenesis Chemical
Impact of NM23-M1 "knock-out" on metastatic potential of hepatocellular carcinoma: a transgenic approach NM23-M1 "knock- out" mice (without NDPK A) Experimental models of hepatocarcinogenesis Chemical
Fusion Analysis of Solid Tumors Reveals Novel Rearrangements in Breast Carcinomas
 Fusion Analysis of Solid Tumors Reveals Novel Rearrangements in Breast Carcinomas Igor Astsaturov Philip Ellis Jeff Swensen Zoran Gatalica David Arguello Sandeep Reddy Wafik El-Deiry Disclaimers Dr. Igor
Fusion Analysis of Solid Tumors Reveals Novel Rearrangements in Breast Carcinomas Igor Astsaturov Philip Ellis Jeff Swensen Zoran Gatalica David Arguello Sandeep Reddy Wafik El-Deiry Disclaimers Dr. Igor
Cellecta Overview. Started Operations in 2007 Headquarters: Mountain View, CA
 Cellecta Overview Started Operations in 2007 Headquarters: Mountain View, CA Focus: Development of flexible, scalable, and broadly parallel genetic screening assays to expedite the discovery and characterization
Cellecta Overview Started Operations in 2007 Headquarters: Mountain View, CA Focus: Development of flexible, scalable, and broadly parallel genetic screening assays to expedite the discovery and characterization
Micro-RNA web tools. Introduction. UBio Training Courses. mirnas, target prediction, biology. Gonzalo
 Micro-RNA web tools UBio Training Courses Gonzalo Gómez//ggomez@cnio.es Introduction mirnas, target prediction, biology Experimental data Network Filtering Pathway interpretation mirs-pathways network
Micro-RNA web tools UBio Training Courses Gonzalo Gómez//ggomez@cnio.es Introduction mirnas, target prediction, biology Experimental data Network Filtering Pathway interpretation mirs-pathways network
Supplementary Figure 1
 Supplementary Figure 1 a Location of GUUUCA motif relative to RNA position 4e+05 3e+05 Reads 2e+05 1e+05 0e+00 35 40 45 50 55 60 65 Distance upstream b 1: Egg 2: L3 3: Adult 4: HES 4e+05 3e+05 2e+05 start
Supplementary Figure 1 a Location of GUUUCA motif relative to RNA position 4e+05 3e+05 Reads 2e+05 1e+05 0e+00 35 40 45 50 55 60 65 Distance upstream b 1: Egg 2: L3 3: Adult 4: HES 4e+05 3e+05 2e+05 start
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Transduction of lentivirus to human primary CD4+ T cells
 Transduction of lentivirus to human primary CD4 + T cells Human primary CD4 T cells were stimulated with anti-cd3/cd28 antibodies (10 µl/2 5 10^6 cells of Dynabeads CD3/CD28 T cell expander, Invitrogen)
Transduction of lentivirus to human primary CD4 + T cells Human primary CD4 T cells were stimulated with anti-cd3/cd28 antibodies (10 µl/2 5 10^6 cells of Dynabeads CD3/CD28 T cell expander, Invitrogen)
Figure S1. Analysis of endo-sirna targets in different microarray datasets. The
 Supplemental Figures: Figure S1. Analysis of endo-sirna targets in different microarray datasets. The percentage of each array dataset that were predicted endo-sirna targets according to the Ambros dataset
Supplemental Figures: Figure S1. Analysis of endo-sirna targets in different microarray datasets. The percentage of each array dataset that were predicted endo-sirna targets according to the Ambros dataset
Association of microrna-7 and its binding partner CDR1-AS with the prognosis and prediction of 1 st -line tamoxifen therapy in breast cancer
 www.nature.com/scientificreports Received: 1 March 2018 Accepted: 12 June 2018 Published: xx xx xxxx OPEN Association of microrna-7 and its binding partner CDR1-AS with the prognosis and prediction of
www.nature.com/scientificreports Received: 1 March 2018 Accepted: 12 June 2018 Published: xx xx xxxx OPEN Association of microrna-7 and its binding partner CDR1-AS with the prognosis and prediction of
MicroRNA and Male Infertility: A Potential for Diagnosis
 Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
SUPPLEMENTARY FIGURE LEGENDS
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1. Hippocampal sections from new-born Pten+/+ and PtenFV/FV pups were stained with haematoxylin and eosin (H&E) and were imaged at (a) low and (b) high
SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1. Hippocampal sections from new-born Pten+/+ and PtenFV/FV pups were stained with haematoxylin and eosin (H&E) and were imaged at (a) low and (b) high
Cancer Problems in Indonesia
 mirna and Cancer : mirna as a Key Regulator in Cancer Sofia Mubarika 2 nd Symposium Biomolecular Update in Cancer PERABOI Padang 18 Mei 2013 Cancer Problems in Indonesia 1. Chemoresistency / recurrency
mirna and Cancer : mirna as a Key Regulator in Cancer Sofia Mubarika 2 nd Symposium Biomolecular Update in Cancer PERABOI Padang 18 Mei 2013 Cancer Problems in Indonesia 1. Chemoresistency / recurrency
Supplement Results, Figures and Tables
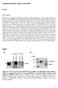 Supplement Results, Figures and Tables Results PET imaging The SUV was significantly higher in tumours of the parental HSC-3 cell line than tumours of the EV and the MMP-8 overexpressing cell lines (Figure
Supplement Results, Figures and Tables Results PET imaging The SUV was significantly higher in tumours of the parental HSC-3 cell line than tumours of the EV and the MMP-8 overexpressing cell lines (Figure
